Abstract
Central nervous system (CNS) infections are potentially life threatening if not diagnosed and treated early. The initial clinical presentations of many CNS infections are non-specific, making a definitive etiologic diagnosis challenging. Nucleic acid in vitro amplification-based molecular methods are increasingly being applied for routine microbial detection. These methods are a vast improvement over conventional techniques with the advantage of rapid turnaround and higher sensitivity and specificity. Additionally, molecular methods performed on cerebrospinal fluid samples are considered the new gold standard for diagnosis of CNS infection caused by pathogens, which are otherwise difficult to detect. Commercial diagnostic platforms offer various monoplex and multiplex PCR assays for convenient testing of targets that cause similar clinical illness. Pan-omic molecular platforms possess potential for use in this area. Although molecular methods are predicted to be widely used in diagnosing and monitoring CNS infections, results generated by these methods need to be carefully interpreted in combination with clinical findings. This review summarizes the currently available armamentarium of molecular assays for diagnosis of central nervous system infections, their application, and future approaches.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The central nervous system (CNS) has unique anatomic and immunologic characteristics that play an important role in the pathogenesis and detection of infection. CNS is protected by the blood–brain barrier (BBB) but is still highly vulnerable to microbial invasion by extension from a contiguous focus; hematogenous dissemination; or less commonly, intraneural passage of organisms [1]. Various environmental or commensal bacteria, viruses, fungi, protozoa, or parasites can migrate into CNS causing a variety of infections [2••]. Clinical manifestation of CNS infection can consist of fever, headache, vomiting, photophobia, stiff neck, and focal neurological presentations.
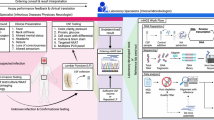
CNS infections are classified according to their anatomic localization (Fig. 1) [2••, 3, 4 5•]. Infection of the meninges, brain, and spinal cord results in meningitis, encephalitis, brain abscess, and myelitis, respectively. Infection may be limited to a single anatomic compartment or may involve multiple sites (e.g., meningoencephalitis and encephalomyelitis). Based on the duration, infection can be classified as acute, sub-acute, chronic, or recurrent. Meningitis is characterized by the onset of fever, headache, neck stiffness, and photophobia over a period of hours to days. Encephalitis is characterized by brain parenchymal inflammation, and the clinical hallmark is alteration in mental status, ranging from lethargy to coma [6]. Myelitis is characterized by the inflammation of the spinal cord with symptoms including fever, headache, and paraparesis or paralysis. Based on the duration, infection can be classified as acute, sub-acute, chronic, or recurrent.
A diagram of nervous system anatomy and the specific pathogens associated with each structure. Modified with permission from Swanson and McGavern [5•]. AVs alphaviruses, BVs bunyaviruses, CMV cytomegalovirus, HEVs human enteroviruses, HIV human immunodeficiency virus, HSV herpes simplex virus, JCV John Cunningham virus, JEV Japanese encephalitis virus, LCMV lymphocytic choriomeningitis virus, MeV measles virus, Mum mumps virus, Nip Nipah virus, PV poliovirus, RV rabies virus, SLEV St. Louis encephalitis virus, TBEV tick-borne encephalitis virus, WNV West Nile virus, Lm Listeria monocytogenes, Nm Neisseria meningitidis, Ec Escherichia coli, Cn Cryptococcus neoformans, Hc Histoplasma capsulatum, Bh Blastocystis hominis, Ci Coccidioides immitis, Td Toxoplasma gondii, GBS Guillain-Barré syndrome, HIB Haemophilus influenzae type b, MTB Mycobacterium tuberculosis, Sp Streptococcus pneumoniae, Tp Treponema pallidum
The most rapidly fatal infection of the CNS is acute bacterial meningitis, with an annual incidence of 3 to 5 cases per 100,000 persons in the USA and an associated mortality rate of 6 to 26 % [2••]. Annually, approximately 4000 cases of acute bacterial meningitis occur in the USA with 500 deaths. The leading causes of bacterial meningitis across all age groups include Streptococcus pneumoniae, group B Streptococcus, Neisseria meningitidis, Haemophilus influenzae, and Listeria monocytogenes.
CNS infections caused by viruses are more common and mostly mild and self limited. These can clinically manifest as meningitis and/or encephalitis [7]. The incidence of viral CNS infections can vary by geographic region and season. Non-polio enteroviruses account for majority of meningitis/encephalitis cases that typically peak between late spring to fall [8•]. The more serious CNS infections due to herpes simplex viruses (HSVs) are associated with sporadic encephalitis and meningitis with severe sequelae if not treated promptly [9].
Rapid and accurate detection and identification of microbial pathogens is essential in directing timely clinical intervention. Conventional methods used in clinical microbiology laboratories include direct microscopic examination, culture techniques, antigen, and antibody detection assays (Table 1). These methods although currently applied have several important limitations—using the example of enteroviruses, the most common cause of meningitis [10••, 11]. Direct microscopic examination of cerebrospinal fluid (CSF) has limited sensitivity and specificity. The sensitivity of culture for enteroviruses is between 65 and 75 % with a mean retrieval time of 3.7 to 8.2 days [12]. Furthermore, some serotypes of enteroviruses, especially Coxsackievirus A strains, are known to grow poorly or are non-cultivable [13]. Enteroviruses lack a common antigen among various serotypes making a universal antigen or antibody detection impossible. Similar issues occur around the diagnosis of CNS HSV infections by conventional methods—culture sensitivity from CSF is extremely poor. Presence of HSV IgG antibodies in CSF can be used in diagnosis; however, production is delayed until day 10 or 12 after infection and is therefore not ideal for early diagnosis [14].
To overcome the several limitations of conventional diagnostic techniques, molecular methods, dominantly PCR-based amplification, have gradually become mainstay tools in detection and identification of microbial pathogens in CSF (Table 1) [2••, 15]. When compared to conventional methods, molecular methods show greater detection rates; one study reported that 16S ribosomal ribonucleic acid (rRNA) PCR-based assays were able to accurately detect the causative organism in 65 % of banked CSF samples, compared to 35 % when using culture and microscopy [16]. In another report, improved diagnostic yield based on molecular methods was used to optimize antibiotic treatment of patients with infectious meningitis when conventional methods provided a negative result [17]. Currently, molecular methods performed on CSF samples are considered a “platinum” standard, in contrast of the culture gold standard, in diagnosis of CNS infections caused by viruses which are difficult to detect and identify [18–20].
Since we provided a review on the topic in 2002 [21•], the diagnostic landscape in CNS infections has changed dramatically. PCR-based molecular methods made their way into clinical microbiology laboratory, providing tools for rapid and accurate diagnosis. In parallel, several commercial molecular assays as summarized in Table 2 have been cleared by the Food and Drug Administration (FDA) for detection of target microbial pathogens in CSF.
Despite the advances in molecular techniques, several challenges remain. Using a combination of conventional and molecular diagnostic methods, Glaser et al. showed that in approximately 62 % of patients with clinical encephalitis, an etiologic organism could not be identified [22]. The focus is now shifting towards development of advanced techniques beyond nucleic acid-based detection. In this review, we give an update on the existing conventional and molecular platforms for the diagnosis of CNS infections. We also provide a preview on the potential clinical application of future technologies including pan-omic assays. The emphasis is given to optimal test selection based on the clinical scenario.
Conventional Microbiology Methods
Microscopic Examination
A positive CSF Gram stain is highly suggestive of bacterial meningitis [23, 24]. The reported sensitivity of the Gram stain for diagnosis of bacterial meningitis is 60∼80 % in patients who have not received antimicrobial treatment and 40∼60 % among those on antibacterial treatment [25]. In one study, Gram stain detected as many as 90 % S. pneumoniae and 50 % L. monocytogenes in CSF collected from patients with bacterial meningitis confirmed by PCR [26]. Two organisms which are often diagnosed by microscopy are Mycobacterium tuberculosis by acid-fast bacillus (AFB) staining and Cryptococcus neoformans by India ink or Gram stain. While these methods maintain satisfactory specificities, the sensitivities are relatively poor; thereby, a culture is usually performed in parallel.
Culture
Culture of brain tissue can provide definitive diagnosis of CNS infections; however, obtaining biopsies is highly invasive and often avoided unless deemed necessary by a clinician. CSF sampling is most commonly performed to determine etiology of suspected CNS infection [25, 27]. CSF viral, bacterial (including mycobacterial), and fungal cultures remain the mainstay in the diagnosis of infectious meningitis. However, the yield of CSF cultures in suspected cases is low [28]. Another disadvantage of CSF bacterial culture is that it requires up to 72 h for final identification. A recent study reported that CSF mycobacterial culture had a sensitivity of 22 % and a specificity of 100 % in diagnosis of tuberculosis meningitis [29]. For viruses, the application of monoclonal antibodies in shell vial culture has increased the speed and specificity. However, due to the long time and low sensitivity, CSF viral culture is often unable to provide the timely diagnosis required for optimum patient management [11, 30].
Rapid Antigen Detection
Among the antigen assays for CNS infections, Cryptococcal antigen is the most widely used. The test relies on detection of Cryptococcus capsular polysaccharide antigens in CSF by enzyme immunoassay [31]. In a single report that examined patients >35 years of age with CNS cryptocossis, an overall sensitivity and specificity of 93∼100 and 93∼98 %, respectively, were reported [32]. Cryptococcus is a neurotropic fungus; polysaccharide serum antigen titers in conjunction with host immune status are often used as a diagnostic aid to determine need for lumbar puncture to evaluate patient for CNS involvement. The baseline peak titer of polysaccharide antigen in serum or CSF has demonstrated important prognostic significance [33] with higher titer (peak titer >1:1024) associated with antifungal therapy failure [34].
Detection of galactomannan (GM) antigen and (1,3)-β-D-glucan (BDG) in CSF can aid in the diagnosis of CNS aspergillosis or other invasive fungal infection such as fusariosis [10••, 35]. Elevated BDG in serum as well as CSF is associated with fungal infections; measuring the levels of BDG might be a useful biomarker in the evaluation of fungal CNS disease [36]. It was recently reported that patients receiving effective antifungal therapy showed reduction in CSF BDG concentration (<31 pg/ml), and for this reason, BDG titers in CSF can be as a useful biomarker in monitoring response to treatment [37].
For acute bacterial meningitis, a rapid antigen assay is available to detect pneumococcal capsular antigen [38]. Recently, several reports revealed potential application of detection of M. tuberculosis-specific antigens in CSF as rapid diagnosis of tuberculosis meningitis [39, 40]. The level of M. tuberculosis early secreted antigenic target 6 (ESAT-6) was associated with clinical severity and may be used for the prognosis of tuberculosis meningitis [41, 42].
Serology
Definitive serological diagnosis of CNS infections is established by detecting IgM antibodies or demonstrating a at least a fourfold increase in neutralizing antibody titers between acute- and convalescent-phase CSF [43, 44]. In general, due to delay in antibody response after symptom onset, a negative antibody test cannot be used to rule out infections and retesting may be required. In addition, in certain selected populations such as immunocompromised individuals, the tests may not offer optimum sensitivity. In most circumstances, nucleic acid amplification tests have surpassed antibody-based detection as the test of choice [11]. For certain infections, these assays still have a valuable role. CSF IgM is the most widely used test for West Nile virus (WNV) infections; antibody may appear as early as 3 days and persist for up to 3 months. However, its accuracy is complicated by high cross-reactivity with other clinically relevant flaviviruses and related vaccines [45]. Antibodies against recombinant WNV E proteins have been proposed as a potential solution to make this important distinction in areas where cross-reacting viruses co-circulate or in individuals who have been immunized [45].
Other highly important serological assays for CNS infections are tests used for screening and diagnosis of neurosyphilis. Neurosyphilis can be confirmed by a positive CSF venereal disease research laboratory (VDRL) test [46]. Detection of antibodies to varicella zoster virus (VZV) IgG and/or DNA in CSF remain as the most commonly used methods to establish a diagnosis of VZV-related CNS disease [47].
Molecular Methods in Diagnosis of CNS Infections
Because of higher sensitivity and specificity, nucleic acid in vitro amplification-based molecular techniques are now widely implemented across clinical laboratories in the USA. Molecular methods have dramatically improved the ability to diagnose CNS infections in a reasonable and effective time frame. Several PCR-derived techniques have collectively expanded the flexibility and rigor of currently available laboratory diagnostic methods [48, 49].
Reverse transcriptase (RT)-PCR was developed to amplify RNA targets; its application has played an important role in diagnosing RNA-virus infections and in some cases monitoring response to therapy. Timely access to enterovirus RT-PCR results have been shown to facilitate shorter hospital stays, reduce unnecessary antibiotic use, and lessens ancillary laboratory testing [50–52]. Broad-range rRNA PCR techniques, which use single pair of primers targeting conserved regions of genes, are successfully used for rapid detection and identification of bacterial pathogens and herpesviruses in the CSF [16, 53, 54]. Isothermal amplification-based techniques including loop-mediated isothermal amplification (LAMP) has been successfully developed to provide point-of-care diagnosis within minutes to hours [55]. Table 2 provides an overview and comparison of commercial molecular in vitro diagnostic devices (IVDs) that have been cleared by the US FDA for detection and identification of microbial pathogens in CSF. Details and updates concerning these devices are available from the FDA website (http://www.fda.gov/MedicalDevices/ProductsandMedicalProcedures/InVitroDiagnostics/ucm330711.htm#microbial). In the following section, we will discuss three different types of molecular testing with focuses on CNS infection diagnosis.
Monoplex Assays
A conventional molecular procedure includes the following three separate steps: sample extraction, target nucleic acid amplification, and amplicon detection. One of the first molecular assays used successfully for CNS infection diagnosis was for detection of HSV in CSF [56•]. PCR quickly became the test of choice when studies demonstrated that CSF PCR was equivalent to culture of brain tissue for diagnosis of HSV encephalitis and meningitis [15]. Since then, numerous PCR-based methods for herpesvirus and enterovirus have become available with superior sensitivity compared to viral culture [57, 58].
Real-time PCR with simultaneous nucleic acid amplification and amplicon detection further accelerated the transition to molecular testing in clinical laboratories. Unlike conventional PCR, the real-time system is a “closed” system and therefore overcomes the important concern of carryover contamination. At the time of manuscript preparation, three molecular assays to detect HSV [59] and enteroviruses [51, 60] in CSF have been approved by FDA as shown in Table 2 [61••].
Real-time PCR-based methods are the main format to detect Zika virus, which was first reported in Uganda in 1947 and is now a worldwide concern after the virus spread widely in Brazil and Central America [62]. Faye et al. developed a one-step RT-PCR assay to detect Zika virus in human serum with limit of detection of 7.7 pfu/reaction [63, 64]. In addition to plasma, Zika virus RNA can be detected in the urine and plasma within first 2 weeks after symptom onset [65]. In March 2016, the FDA approved a trioplex-PCR assay under emergency use authorization for the simultaneous detection of Zika, Chikungunya, and dengue viruses in serum, urine, CSF, and amniotic fluid. The RT-PCR assay uses dual-labeled hydrolysis probes with a LOD of 1.54 × 10 GCE/ml [4] of Zika virus in serum (http://www.fda.gov/downloads/MedicalDevices/Safety/EmergencySituations/UCM491592.pdf).
Introduction of real-time PCR-based diagnostic assays has had a substantial impact on early and effective diagnosis of certain bacterial infections [66, 67]. Isothermal amplification-based molecular assays have excellent performance characteristics and have the distinct advantage of not requiring any specialized equipment. These assays are therefore ideal for use as on or near point-of-care testing. Using this technology, LAMP-based methods have been used to detect Neisseria meningitis, S. pneumoniae, H. influenzae type b, M. tuberculosis, and Japanese encephalitis virus (JEV) in the CSF [68–71]. The Xpert MTB/RIF assay has revolutionized the landscape of global tuberculosis control by providing an integrated and automated system that enables rapid clinical decision making in a POC or near-care context [67]. Several studies have applied the Xpert MTB/RIF to evaluate detection of M. tuberculosis in CSF from cases of TB meningitis [72•, 73, 74]. In a meta-analysis of 13 studies, the pooled sensitivity of Xpert assay was 80.5 % (95 % CI 59.0–92.2) against culture and 62.8 % (95 % CI 47.7–75.8) against composite standard. Using a large volume of sample (at least 8–10 ml) is required for testing CSF, and centrifugation can lead to modest improvement in yield [75]. Despite the lack of standardization for sample processing, WHO has endorsed testing CSF with the automated Xpert MTB/RIF assay as a first-line test over conventional microscopy in patients with suspected TB meningitis.
Multiplex Assays
Relative simplicity and high-throughput detection make multiplex molecular assays an attractive option for screening and detection of a panel of microbial targets [76]. Several multiplex PCR assays have been developed to identify bacterial pathogens in CSF, targeting the most common causes of meningitis, S. pneumoniae, N. meningitis, H. influenzae [26, 48, 76, 77], L. monocytogenes [26, 77], Streptococcus agalactiae, Staphylococcus aureus, Escherichia coli [26], and Mycoplasma pneumoniae [26]. A multiplex PCR followed by Luminex suspension array can simultaneously detect eight bacterial and viral pathogens in CSF including N. meningitis, S. pueumoniae, E. coli, S. aureus, L. monocytogenes, S. agalactiae, HSV-1/2, and VZV [78].
Considering the breadth of pathogens implicated in CNS infection, application of comprehensive molecular panels with multiple bacterial and viral targets have improved the diagnostic efficiency. The BioFire FilmArray Meningitis/Encephalitis panel is currently the only FDA-cleared multiplex assay for the detection of six bacterial (E. coli K1, H. influenzae, L. monocytogenes, N. meningitidis, S. agalactiae, and S. pneumoniae), seven viral (cytomegalovirus, enterovirus, HSV-1, HSV-2, human herpesvirus 6 (HHV-6), human parechovirus, and VZV), and single fungal (C. neoformans/gattii) target in CSF (Table 2). The integrated FilmArray system has a turnaround time of about an hour, with only 2 min of hands-on time. At the time of the manuscript preparation, two studies have reported on the performance of this assay [79, 80]. Using 48 samples from Gram stain-negative CSF samples from suspected cases of meningitis, Wootton et al. showed that this system detected more viral pathogens especially EBV. Four cases of WNV and single case of Histoplasma were not detected by this assay. Among HIV-infected patients in Uganda, the test performance demonstrated superior sensitivity and specificity for detection of Cryptococcus [79, 80]. Although the FilmArray Meningitis/Encephalitis panel offers a promising platform for rapid diagnosis of CNS infections, further clinical studies are needed to determine its performance for various targets and among other high-risk populations.
Co-infections are frequently encountered among immunocompromised patients and present a difficult diagnostic challenge for clinicians. Multiplex design enables simultaneous detection and identification of multiple targets on the same sample. Rajasingham et al. [81] used a panel of monoplex and multiplex molecular assays to conduct a prospective cohort study in Uganda to comprehensively evaluate the etiology of meningitis among HIV-infected adults. Among the 314 HIV-infected patients with suspected meningitis, EBV co-infection was detected with Cryptococcus, M. tuberculosis, or other viral pathogens [81]. The clinical significance of EBV in CSF in these settings is not completely understood, although a single study associated high EBV viral load as a marker of poor outcome in individuals with bacterial meningitis and EBV co-infection/reactivation [82].
Pan-Omic Molecular Assays
Technological improvements in metagenomic deep sequencing have led to its potential application for clinical diagnosis of infections [83, 84 85••]. Several reports have demonstrated its utility into solving diagnostic dilemmas that challenge the limits of traditional laboratory testing [83, 86, 87 85••, ]. Due to sterile status and protection by BBB, CSF and brain biopsies are ideal to further explore the application of this technology for pathogen detection and discovery. As demonstrated in a highly challenging clinical situation, metagenomics was successfully used to establish a timely diagnosis of neuroleptospirosis in a 14-year-old boy with severe combined immunodeficiency, who suffered from recurrent bouts of fever, headache, and coma [85••]. Similarly, high-throughput RNA sequencing performed on brain biopsy from an 18-month-old boy with encephalopathy was able to identify a new astrovirus as the cause [83]. Despite the enormously attractive potential of metagenomics for infectious disease diagnosis, there are many technological and practical concerns that need to be addressed before this form of diagnostic testing can become mainstream and part of the clinical standard of care.
Other promising advances have occurred in transcriptomics, proteomics, and metabolomics. Host and microbial microRNA (miRNA) profiles have been used for various inflammatory and infectious diseases [88]. Two miRNAs, miR-155 and miRNA-29b, were reported as potential biomarkers for JEV infection as well as therapeutic targets for anti-JEV therapy [89, 90]. Host neural epidermal growth factor like 2 and apolipoprotein B in CSF were able to diagnose tuberculous meningitis with 83.3–89.3 % sensitivity and 75–92 % specificity [91, 92]. CSF metabolite profiling has been reported useful in classification, diagnosis, epidemiology, and treatment assessment of CNS infections in HIV patients [93–95]. CSF metabolic profile analysis implicated bioenergetic adaptation as a neural mechanism regulating shifts in cognitive states of HIV-infected patients [96].
Selective Testing Results to Interpret Correlation with Clinical Diseases
Identification of an etiologic agent in patients with CNS infections requires consideration of the most likely causative organisms, the available diagnostic tests for these agents, and the highest-yield clinical specimens for testing. Knowledge of the epidemiology and clinical presentation of specific agents is critical in selecting which diagnostic methods are appropriate for a given patient. In particular, animal or vector exposures, geographic location, recent travel history, season of the year, exposure of ill contacts, and occupational exposures should be considered.
When selecting appropriate pathogen-specific molecular diagnostic methods, the following factors should be considered. CSF is the optimal specimen for PCR testing for patients with suspected meningitis or meningoencephalitis. While indirect evidence can be gained by testing of other specimen types, attempts should be made to obtain CSF samples early before commencement of treatment that can compromise yield. Time of testing from symptom onset is essential to understand and rule out false negative results and recommend retesting within a certain time frame. For example, HSV PCR can commonly render false negative results if CSF sample is obtained very early or late in the process of HSE infection. Host health condition is known to influence the test performance characteristics. Immunocompromised patients are at risk for infection by a much wider array of opportunistic pathogens, for example, HHV-6, JC virus, Toxoplasma encephalitis in bone marrow transplant recipients, and patients with HIV. Often, infection can be more severe (e.g., WNV) and difficult to diagnose in this population. Table 3 provides the practical recommendations on application and pitfalls of molecular test for diagnosis of CNS infections.
Finally, a positive nucleic acid amplification testing result is complicated by the fact that some viruses survive latently in macrophages or neurologic tissues and are incidentally detected by sensitive molecular techniques without an actual pathogenic role and can potentially lead to overtreatment. Uses of adjunctive biomarkers that depict active replication are being explored to overcome this drawback.
Conclusion
Historically, identification of microbiologic agents in patients with CNS infections has been hindered by the low yield of CSF culture for viral and fastidious bacterial organisms, delays in CNS production of organism-specific antibodies, and difficulties in obtaining optimum samples for testing. Nucleic acid in vitro amplification-based molecular diagnosis methods have a wider and better application in clinical microbiology practice. The monoplex assay will likely be the main platform for urgent, random-access, low-throughput assays. Multiplex assays have the additional advantage of detecting multiple targets and mixed infections. As volume of CSF sample retrieved is often small, multiplex assays enable comprehensive diagnostic analysis with low amount of sample, obviating need for repeated lumbar punctures. The clinical relevance and cost-effectiveness of simultaneous multipathogen detection and identification strategies merit further investigation. Application of pan-omic techniques in difficult-to-diagnose CNS infections is the new exciting frontier; the technology is promising, but routine implementation is expected to be slow due to various challenges such as lack of applicable regulatory guidelines and adaptation in the clinical setting.
References
Papers of particular interest, published recently, have been highlighted as: • Of importance •• Of major importance
Varatharaj A, Galea I. The blood–brain barrier in systemic inflammation. Brain Behav Immun 2016.
Bloch KC, Tang YW. Molecular approaches to the diagnosis of meningitis and encephalitis. In: Persing DH et al., editors. Molecular microbiology: diagnostic principles and practice. III edth ed. Washington: American Society for Microbiology Press; 2016. p. 287–305. This recently published book chapter described molecular methods to detect microbial pathogens and their pros and cons in diagnosis meningitis and encephalitis.
Bale Jr JF. Virus and immune-mediated encephalitis: epidemiology, diagnosis, treatment, and prevention. Pediatr Neurol. 2015;53(1):3–12.
Han SH, Choi HY, Kim JM, Park KR, Youn YC, Shin HW. Etiology of aseptic meningitis and clinical characteristics in immune-competent adults. J Med Virol. 2016;88(1):175–9.
Swanson 2nd P, McGavern DB. Viral diseases of the central nervous system. Curr Opin Virol. 2015;11:44–54. This review highlighted a list of neurotropic viruses that can cause meningitis, encephalitis, and myelitis.
Venkatesan A. Epidemiology and outcomes of acute encephalitis. Curr Opin Neurol. 2015;28(3):277–82.
Lipkin WI, Hornig M. Diagnostics and discovery in viral central nervous system infections. Brain Pathol. 2015;25(5):600–4.
Ramers C, Billman G, Hartin M, Ho S, Sawyer MH. Impact of a diagnostic cerebrospinal fluid enterovirus polymerase chain reaction test on patient management. JAMA. 2000;283(20):2680–5. This was the first article demonstrating that a positive molecular enterovirus result affects clinical decision making and promotes rapid discharge of patients.
Whitley RJ. Herpes simplex virus infections of the central nervous system. Continuum (Minneap Minn) 2015; 21(6 Neuroinfectious disease): 1704–13.
Caliendo AM, Gilbert DN, Ginocchio CC, et al. Better tests, better care: improved diagnostics for infectious diseases. Clin Infect Dis: Off Publ Infect Dis Soc Am. 2013;57 Suppl 3:S139–70. An important review on the current diagnostic landscape of CNS infections, including unmet needs and emerging technologies.
Storch GA. Diagnostic virology. Clin Infect Dis: Off Publ Infect Dis Soc Am. 2000;31(3):739–51.
Ginocchio CC, Zhang F, Malhotra A, et al. Development, technical performance, and clinical evaluation of a NucliSens basic kit application for detection of enterovirus RNA in cerebrospinal fluid. J Clin Microbiol. 2005;43(6):2616–23.
Tyler KL. Herpes simplex virus infections of the central nervous system: encephalitis and meningitis, including Mollaret’s. Herpes: J IHMF. 2004;11 Suppl 2:57A–64A.
Liermann K, Schafler A, Henke A, Sauerbrei A. Evaluation of commercial herpes simplex virus IgG and IgM enzyme immunoassays. J Virol Methods. 2014;199:29–34.
Tang YW. Laboratory diagnosis of CNS infections by molecular amplification techniques. Expert Opin Med Diagn. 2007;1(4):489–509.
Meyer T, Franke G, Polywka SK, et al. Improved detection of bacterial central nervous system infections by use of a broad-range PCR assay. J Clin Microbiol. 2014;52(5):1751–3.
Bahr NC, Marais S, Caws M, et al. GeneXpert MTB/RIF to diagnose tuberculous meningitis: perhaps the first test but not the last. Clin Infect Dis: Off Publ Infect Dis Soc Am. 2016;62(9):1133–5.
Jaramillo-Gutierrez G, Benschop KS, Claas EC, et al. September through October 2010 multi-centre study in the Netherlands examining laboratory ability to detect enterovirus 68, an emerging respiratory pathogen. J Virol Methods. 2013;190(1–2):53–62.
Launes C, Armero G, Anton A, et al. Molecular epidemiology of severe respiratory disease by human rhinoviruses and enteroviruses at a tertiary paediatric hospital in Barcelona, Spain. Clin Microbiol Infect: Off Publ Eur Soc Clin Microbiol Infect Dis. 2015;21(8):799.e5–7.
McAllister SC, Schleiss MR, Arbefeville S, et al. Epidemic 2014 enterovirus D68 cross-reacts with human rhinovirus on a respiratory molecular diagnostic platform. PLoS One. 2015;10(3):e0118529.
Lu HZ, Bloch KC, Tang YW. Molecular techniques in the diagnosis of central nervous system infections. Curr Infect Dis Rep. 2002;4(4):339–50. An original article that our review articles based and extended.
Glaser CA, Gilliam S, Schnurr D, et al. In search of encephalitis etiologies: diagnostic challenges in the California Encephalitis Project, 1998–2000. Clin Infect Dis: Off Publ Infect Dis Soc Am. 2003;36(6):731–42.
Cattamanchi A, Davis JL, Pai M, Huang L, Hopewell PC, Steingart KR. Does bleach processing increase the accuracy of sputum smear microscopy for diagnosing pulmonary tuberculosis? J Clin Microbiol. 2010;48(7):2433–9.
Musher DM, Schell RF. Letter: false-positive gram stains of cerebrospinal fluid. Ann Intern Med. 1973;79(4):603–4.
Dunbar SA, Eason RA, Musher DM, Clarridge 3rd JE. Microscopic examination and broth culture of cerebrospinal fluid in diagnosis of meningitis. J Clin Microbiol. 1998;36(6):1617–20.
Ubukata K. Rapid identification of meningitis due to bacterial pathogens. Rinsho Shinkeigaku. 2013;53(11):1184–6.
Polage CR, Petti CA. Assessment of the utility of viral culture of cerebrospinal fluid. Clin Infect Dis: Off Publ Infect Dis Soc Am. 2006;43(12):1578–9.
Ku LC, Boggess KA, Cohen-Wolkowiez M. Bacterial meningitis in infants. Clin Perinatol. 2015;42(1):29–45. vii-viii.
Solomons RS, Visser DH, Friedrich SO, et al. Improved diagnosis of childhood tuberculous meningitis using more than one nucleic acid amplification test. Int J Tuberc Lung Dis: Off J Int Union Against Tuberc Lung Dis. 2015;19(1):74–80.
Kelley VA, Caliendo AM. Successful testing protocols in virology. Clin Chem. 2001;47(8):1559–62.
Antinori S, Radice A, Galimberti L, Magni C, Fasan M, Parravicini C. The role of cryptococcal antigen assay in diagnosis and monitoring of cryptococcal meningitis. J Clin Microbiol. 2005;43(11):5828–9.
Ji S, Ni L, Zhang J, Huang J, Zhou Z, Yu Y. Value of three capsular antigen detection methods in diagnosis and efficacy assessment in patients with cryptococcal meningoencephalitis. Zhonghua Yi Xue Za Zhi. 2015;95(46):3733–6.
Kaplan JE, Vallabhaneni S, Smith RM, Chideya-Chihota S, Chehab J, Park B. Cryptococcal antigen screening and early antifungal treatment to prevent cryptococcal meningitis: a review of the literature. J Acquir Immune Defic Syndr. 2015;68 Suppl 3:S331–9.
Jarvis JN, Percival A, Bauman S, et al. Evaluation of a novel point-of-care cryptococcal antigen test on serum, plasma, and urine from patients with HIV-associated cryptococcal meningitis. Clin Infect Dis: Off Publ Infect Dis Soc Am. 2011;53(10):1019–23.
Marchetti O, Lamoth F, Mikulska M, Viscoli C, Verweij P, Bretagne S. ECIL recommendations for the use of biological markers for the diagnosis of invasive fungal diseases in leukemic patients and hematopoietic SCT recipients. Bone Marrow Transplant. 2012;47(6):846–54.
Lyons JL, Thakur KT, Lee R, et al. Utility of measuring (1,3)-beta-d-glucan in cerebrospinal fluid for diagnosis of fungal central nervous system infection. J Clin Microbiol. 2015;53(1):319–22.
Salvatore CM, Chen TK, Toussi SS, et al. (1→3)-beta-d-Glucan in cerebrospinal fluid as a biomarker for Candida and Aspergillus infections of the central nervous system in pediatric patients. J Pediatric Infect Dis Soc 2015.
Marcos MA, Martinez E, Almela M, Mensa J, de Jimenez Anta MT. New rapid antigen test for diagnosis of pneumococcal meningitis. Lancet (London, England). 2001;357(9267):1499–500.
Haldar S, Sankhyan N, Sharma N, et al. Detection of Mycobacterium tuberculosis GlcB or HspX antigens or devR DNA impacts the rapid diagnosis of tuberculous meningitis in children. PLoS One. 2012;7(9):e44630.
Silva BD, Tannus-Silva DG, Rabahi MF, Kipnis A, Junqueira-Kipnis AP. The use of Mycobacterium tuberculosis HspX and GlcB proteins to identify latent tuberculosis in rheumatoid arthritis patients. Mem Inst Oswaldo Cruz. 2014;109(1):29–37.
Peng X, Jiang G, Liu W, Zhang Q, Qian W, Sun J. Characterization of differential pore-forming activities of ESAT-6 proteins from Mycobacterium tuberculosis and Mycobacterium smegmatis. FEBS Lett. 2016;590(4):509–19.
Song F, Sun X, Wang X, Nai Y, Liu Z. Early diagnosis of tuberculous meningitis by an indirect ELISA protocol based on the detection of the antigen ESAT-6 in cerebrospinal fluid. Ir J Med Sci. 2014;183(1):85–8.
Kuroda H. Update on herpes simplex encephalitis. Brain Nerve. 2015;67(7):931–9.
Steingart KR, Ramsay A, Dowdy DW, Pai M. Serological tests for the diagnosis of active tuberculosis: relevance for India. Indian J Med Res. 2012;135(5):695–702.
Chabierski S, Barzon L, Papa A, et al. Distinguishing West Nile virus infection using a recombinant envelope protein with mutations in the conserved fusion-loop. BMC Infect Dis. 2014;14:246.
Tuddenham S, Ghanem KG. Emerging trends and persistent challenges in the management of adult syphilis. BMC Infect Dis. 2015;15:351.
Gilden D, Cohrs RJ, Mahalingam R, Nagel MA. Varicella zoster virus vasculopathies: diverse clinical manifestations, laboratory features, pathogenesis, and treatment. Lancet Neurol. 2009;8(8):731–40.
Conca N, Santolaya ME, Farfan MJ, et al. Etiologic diagnosis in meningitis and encephalitis molecular biology techniques. Rev Chil Pediatr. 2016;87(1):24–30.
Ohkusu K. Molecular approaches for the diagnosis of central nervous system infections. Brain Nerve. 2015;67(7):799–811.
Jin D, Heo TH, Byeon JH, et al. Analysis of clinical information and reverse transcriptase-polymerase chain reaction for early diagnosis of enteroviral meningitis. Korean J Pediatr. 2015;58(11):446–50.
Nolte FS, Rogers BB, Tang YW, et al. Evaluation of a rapid and completely automated real-time reverse transcriptase PCR assay for diagnosis of enteroviral meningitis. J Clin Microbiol. 2011;49(2):528–33.
Shaker OG, Abdelhamid N. Detection of enteroviruses in pediatric patients with aseptic meningitis. Clin Neurol Neurosurg. 2015;129:67–71.
Mishra AK, Dufour H, Roche PH, Lonjon M, Raoult D, Fournier PE. Molecular revolution in the diagnosis of microbial brain abscesses. Eur J Clin Microbiol Infect Dis: Off Publ Eur Soc Clin Microbiol. 2014;33(12):2083–93.
Moayedi AR, Nejatizadeh A, Mohammadian M, Rahmati MB, Namardizadeh V. Accuracy of universal polymerase chain reaction (PCR) for detection of bacterial meningitis among suspected patients. Electron Physician. 2015;7(8):1609–12.
Notomi T, Mori Y, Tomita N, Kanda H. Loop-mediated isothermal amplification (LAMP): principle, features, and future prospects. J Microbiol. 2015;53(1):1–5.
Rowley AH, Whitley RJ, Lakeman FD, Wolinsky SM. Rapid detection of herpes-simplex-virus DNA in cerebrospinal fluid of patients with herpes simplex encephalitis. Lancet. 1990;335(8687):440–1. One of the early articles demonstrating that PCR assay provided rapid and accurate diagnosis of herpes simplex encephalitis.
Gitman MR, Ferguson D, Landry ML. Comparison of Simplexa HSV 1 & 2 PCR with culture, immunofluorescence, and laboratory-developed TaqMan PCR for detection of herpes simplex virus in swab specimens. J Clin Microbiol. 2013;51(11):3765–9.
Giulieri SG, Chapuis-Taillard C, Manuel O, et al. Rapid detection of enterovirus in cerebrospinal fluid by a fully-automated PCR assay is associated with improved management of aseptic meningitis in adult patients. J Clin Virol: Off Publ Pan Am Soc Clin Virol. 2015;62:58–62.
Kuypers J, Boughton G, Chung J, et al. Comparison of the Simplexa HSV1 & 2 Direct kit and laboratory-developed real-time PCR assays for herpes simplex virus detection. J Clin Virol: Off Publ Pan Am Soc Clin Virol. 2015;62:103–5.
Capaul SE, Gorgievski-Hrisoho M. Detection of enterovirus RNA in cerebrospinal fluid (CSF) using NucliSens EasyQ Enterovirus assay. J Clin Virol: Off Publ Pan Am Soc Clin Virol. 2005;32(3):236–40.
Emmadi R, Boonyaratanakornkit JB, Selvarangan R, et al. Molecular methods and platforms for infectious diseases testing a review of FDA-approved and cleared assays. J Mol Diagn. 2011;13(6):583–604. This wonderful review article covered infectious disease molecular diagnostic tests which were approved or cleared by FDA. Indications and limitations of each assay were described and contrasted.
Mlakar J, Korva M, Tul N, et al. Zika virus associated with microcephaly. N Engl J Med. 2016;374(10):951–8.
Faye O, Faye O, Diallo D, Diallo M, Weidmann M, Sall AA. Quantitative real-time PCR detection of Zika virus and evaluation with field-caught mosquitoes. Virol J. 2013;10:311.
Faye O, Faye O, Dupressoir A, Weidmann M, Ndiaye M, Alpha SA. One-step RT-PCR for detection of Zika virus. J Clin Virol: Off Publ Pan Am Soc Clin Virol. 2008;43(1):96–101.
Musso D, Roche C, Nhan TX, Robin E, Teissier A, Cao-Lormeau VM. Detection of Zika virus in saliva. J Clin Virol: Off Publ Pan Am Soc Clin Virol. 2015;68:53–5.
McHugh MP, Gray SJ, Kaczmarski EB, Guiver M. Reduced turnaround time and improved diagnosis of invasive serogroup B Neisseria meningitidis and Streptococcus pneumoniae infections using a lyophilized quadruplex quantitative PCR. J Med Microbiol. 2015;64(11):1321–8.
Dheda K, Ruhwald M, Theron G, Peter J, Yam WC. Point-of-care diagnosis of tuberculosis: past, present and future. Respirology (Carlton, Vic). 2013;18(2):217–32.
Deng J, Pei J, Gou H, Ye Z, Liu C, Chen J. Rapid and simple detection of Japanese encephalitis virus by reverse transcription loop-mediated isothermal amplification combined with a lateral flow dipstick. J Virol Methods. 2015;213:98–105.
Kim DW, Kilgore PE, Kim EJ, et al. The enhanced pneumococcal LAMP assay: a clinical tool for the diagnosis of meningitis due to Streptococcus pneumoniae. PLoS One. 2012;7(8):e42954.
Kim DW, Kilgore PE, Kim EJ, Kim SA, Anh DD, Seki M. Loop-mediated isothermal amplification assay for detection of Haemophilus influenzae type b in cerebrospinal fluid. J Clin Microbiol. 2011;49(10):3621–6.
Lee D, Kim EJ, Kilgore PE, et al. A novel loop-mediated isothermal amplification assay for serogroup identification of Neisseria meningitidis in cerebrospinal fluid. Front Microbiol. 2015;6:1548.
Denkinger CM, Schumacher SG, Boehme CC, Dendukuri N, Pai M, Steingart KR. Xpert MTB/RIF assay for the diagnosis of extrapulmonary tuberculosis: a systematic review and meta-analysis. Eur Respir J. 2014;44(2):435–46. This is a systematic review and meta-analysis to assess the accuracy of GeneXpert system for the laboratory diagnosis of extrapulmonary tuberculosis.
Nhu NT, Heemskerk D, Thu do DA, et al. Evaluation of GeneXpert MTB/RIF for diagnosis of tuberculous meningitis. J Clin Microbiol. 2014;52(1):226–33.
Patel VB, Theron G, Lenders L, et al. Diagnostic accuracy of quantitative PCR (Xpert MTB/RIF) for tuberculous meningitis in a high burden setting: a prospective study. PLoS Med. 2013;10(10):e1001536.
Bahr NC, Tugume L, Rajasingham R, et al. Improved diagnostic sensitivity for tuberculous meningitis with Xpert((R)) MTB/RIF of centrifuged CSF. Int J Tuberc Lung Dis: Off J Int Union Against Tuberc Lung Dis. 2015;19(10):1209–15.
Huy NT, le Hang TT, Boamah D, et al. Development of a single-tube loop-mediated isothermal amplification assay for detection of four pathogens of bacterial meningitis. FEMS Microbiol Lett. 2012;337(1):25–30.
Palmiere C, Vanhaebost J, Ventura F, Bonsignore A, Bonetti LR. Cerebrospinal fluid PCR analysis and biochemistry in bodies with severe decomposition. J Forensic Leg Med. 2015;30:21–4.
Boving MK, Pedersen LN, Moller JK. Eight-plex PCR and liquid-array detection of bacterial and viral pathogens in cerebrospinal fluid from patients with suspected meningitis. J Clin Microbiol. 2009;47(4):908–13.
Rhein J, Bahr NC, Hemmert AC, et al. Diagnostic performance of a multiplex PCR assay for meningitis in an HIV-infected population in Uganda. Diagn Microbiol Infect Dis. 2016;84(3):268–73.
Wootton SH, Aguilera E, Salazar L, Hemmert AC, Hasbun R. Enhancing pathogen identification in patients with meningitis and a negative Gram stain using the BioFire FilmArray((R)) Meningitis/Encephalitis panel. Ann Clin Microbiol Antimicrob. 2016;15(1):26.
Rajasingham R, Rhein J, Klammer K, et al. Epidemiology of meningitis in an HIV-infected Ugandan cohort. Am J Trop Med Hyg. 2015;92(2):274–9.
Kelly C, Sohal A, Michael BD, Riordan A, Solomon T, Kneen R. Suboptimal management of central nervous system infections in children: a multi-centre retrospective study. BMC Pediatr. 2012;12:145.
Brown JR, Morfopoulou S, Hubb J, et al. Astrovirus VA1/HMO-C: an increasingly recognized neurotropic pathogen in immunocompromised patients. Clin Infect Dis: Off Publ Infect Dis Soc Am. 2015;60(6):881–8.
Goldberg B, Sichtig H, Geyer C, Ledeboer N, Weinstock GM. Making the leap from research laboratory to clinic: challenges and opportunities for next-generation sequencing in infectious disease diagnostics. mBio. 2015;6(6):e01888–15.
Wilson MR, Naccache SN, Samayoa E, et al. Actionable diagnosis of neuroleptospirosis by next-generation sequencing. N Engl J Med. 2014;370(25):2408–17. This was the first clinical case that metagenomics was successfully used to establish a timely etiologic diagnosis, which directed a targeted antimicrobial agent administration resulting the patient’s status close to his premorbid condition.
Naccache SN, Peggs KS, Mattes FM, et al. Diagnosis of neuroinvasive astrovirus infection in an immunocompromised adult with encephalitis by unbiased next-generation sequencing. Clin Infect Dis: Off Publ Infect Dis Soc Am. 2015;60(6):919–23.
Wilson MR, Shanbhag NM, Reid MJ, et al. Diagnosing Balamuthia mandrillaris encephalitis with metagenomic deep sequencing. Ann Neurol. 2015;78(5):722–30.
Panackal AA, Williamson KC, van de Beek D, Boulware DR, Williamson PR. Fighting the monster: applying the host damage framework to human central nervous system infections. mBio. 2016;7(1):e01906–15.
Thounaojam MC, Kundu K, Kaushik DK, et al. MicroRNA 155 regulates Japanese encephalitis virus-induced inflammatory response by targeting Src homology 2-containing inositol phosphatase 1. J Virol. 2014;88(9):4798–810.
Zhu B, Ye J, Nie Y, et al. MicroRNA-15b modulates Japanese encephalitis virus-mediated inflammation via targeting RNF125. J Immunol (Baltimore, Md : 1950). 2015;195(5):2251–62.
Mu J, Yang Y, Chen J, et al. Elevated host lipid metabolism revealed by iTRAQ-based quantitative proteomic analysis of cerebrospinal fluid of tuberculous meningitis patients. Biochem Biophys Res Commun. 2015;466(4):689–95.
Yang Y, Mu J, Chen G, et al. iTRAQ-based quantitative proteomic analysis of cerebrospinal fluid reveals NELL2 as a potential diagnostic biomarker of tuberculous meningitis. Int J Mol Med. 2015;35(5):1323–32.
Cassol E, Misra V, Dutta A, Morgello S, Gabuzda D. Cerebrospinal fluid metabolomics reveals altered waste clearance and accelerated aging in HIV patients with neurocognitive impairment. AIDS (London, England). 2014;28(11):1579–91.
Haughey NJ, Zhu X, Bandaru VV. A biological perspective of CSF lipids as surrogate markers for cognitive status in HIV. J Neuroimmune Pharm: Off J Soc NeuroImmune Pharm. 2013;8(5):1136–46.
Shawahna R. Physical and metabolic integrity of the blood–brain barrier in HIV infection: a special focus on intercellular junctions, influx and efflux transporters and metabolizing enzymes. Curr Drug Metab. 2015;16(2):105–23.
Dickens AM, Anthony DC, Deutsch R, et al. Cerebrospinal fluid metabolomics implicate bioenergetic adaptation as a neural mechanism regulating shifts in cognitive states of HIV-infected patients. AIDS (London, England). 2015;29(5):559–69.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Human and Animal Rights and Informed Consent
This article does not contain any studies with human or animal subjects performed by the author.
Additional information
This article is part of the Topical Collection on Central Nervous System Infections
Rights and permissions
About this article
Cite this article
He, T., Kaplan, S., Kamboj, M. et al. Laboratory Diagnosis of Central Nervous System Infection. Curr Infect Dis Rep 18, 35 (2016). https://doi.org/10.1007/s11908-016-0545-6
Published:
DOI: https://doi.org/10.1007/s11908-016-0545-6