Abstract
Purpose of the Review
Cardiovascular disease remains the leading cause of death worldwide, resulting in cardiac dysfunction and, subsequently, heart failure (HF). Single-cell RNA sequencing (scRNA-seq) is a rapidly developing tool for studying the transcriptional heterogeneity in both healthy and diseased hearts. Wide applications of techniques like scRNA-seq could significantly contribute to uncovering the molecular mechanisms involved in the onset and progression to HF and contribute to the development of new, improved therapies. This review discusses several studies that successfully applied scRNA-seq to the mouse and human heart using various methods of tissue processing and downstream analysis.
Recent Findings
The application of scRNA-seq in the cardiovascular field is continuously expanding, providing new detailed insights into cardiac pathophysiology.
Summary
Increased understanding of cardiac pathophysiology on the single-cell level will contribute to the development of novel, more effective therapeutic strategies. Here, we summarise the possible application of scRNA-seq to the adult mammalian heart.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In the past few years, single-cell sequencing has become a very powerful method to study gene expression changes in different organs [1, 2]. Even though the application of this technique to many tissues appeared to be fairly straightforward, the use of this method in the heart seemed to be quite challenging [3]. This is probably because of the unique nature of the heart tissue. On the one hand, cardiomyocytes are large and can be easily damaged by the enzymatic digestion and sorting, and on the other hand, nonmyocytes are smaller and more resistant and thus need more harsh treatment. Also, while studying a diseased heart, the pathological remodelling will have an impact on the procedure. For instance, during ischemic injury of the heart, there is formed a rigid fibrotic scar that is very difficult to dissociate into the single-cell suspension, often resulting in a poor RNA quality.
Single-Cell RNA Sequencing, Why Do We Need It?
For decades, researchers all over the world have used traditional bulk sequencing methods to study gene expression changes in different tissues [4]. The obtained information was sufficient to study the total content of RNAs, including mRNA, rRNA, t-RNA, and noncoding RNA [5]. Undoubtfully, bulk RNA sequencing has greatly advanced our understanding of the molecular signalling involved in cardiac homeostasis and disease, but it has some limitations. The most significant advantage of single-cell sequencing over bulk sequencing is the ability to obtain the gene expression profile of individual cells. It can reveal complex and rare cell populations, uncover cell type–specific differences in gene expression, study heterogeneity within one cell type, and uncover new and potentially unexpected biological processes that could be overlooked with the traditional bulk sequencing [2].
The very first study using scRNA-seq was described in 2009 in Nature Methods by Tang and colleagues [6]. They used a labour-intense and time-consuming protocol to manually pick a single oocyte or blastomere under the microscope, followed by individual processing. Luckily in the following years, the scRNA-seq methods were much improved, increasing the stability and reproducibility and, at the same time, reducing costs and workload [1]. Microfluidics and robotics-based approached supported simultaneous isolation of thousands of cells, followed by rapid RNA extraction and downstream processing for sequencing [7,8,9]. Also, developments of more elaborate analysis methods led to a significant increase in studies using scRNA-seq. The first reports of using scRNA-seq in the heart came in 2016 from Sean Wu and Christine Seidman’s labs [10, 11]. They used scRNA-seq to profile the transcriptome of the mouse embryonic heart and identify lineage-specific gene programs that underlie early cardiac development. After that, in 2018, the first studies were published using scRNA-seq on adult healthy and diseased hearts [12,13,14]. Nowadays, multiple researchers use this technique in various cardiac disease models and even applied it to human tissue. The use of scRNA-seq in the cardiovascular field is continuously expanding, providing new detailed insights into cardiac pathophysiology. Here, we discuss the most commonly used application of scRNA-seq to the adult mammalian heart (Fig. 1).
Schematic representation of a commonly used application of single-cell sequencing in the adult mammalian heart. a Heterogeneity within the same cell type, including disease-specific populations. b Lineage trajectory reconstitution. c Network analysis. d Cellular communication. e Identification of rare cells
Studying the Heterogeneity Within the Same Cell Type, Including Disease-Specific Populations
One of the first studies that applied scRNA-seq to healthy and diseased hearts observed heterogeneity within the same cardiac cell type [14]. The authors found that there is heterogeneity in the cardiomyocyte population and that only a subset of them expresses myozenin 2 (Myoz2). MYOZ2 is a sarcomeric protein that tether a-actinin to calcineurin, a well-known driver of cardiac hypertrophy, therefore preventing the pathological hypertrophic response of cardiomyocytes [15]. MYOZ2 is expressed in the subset of cardiomyocytes close to the epicardial region of the heart, suggesting region- and function-dependent differences in the expression already in the healthy tissue. Another report using scRNA-seq aimed to investigate the heterogeneity among adult cardiomyocytes under baseline and pressure overload-induced cardiac hypertrophy [16]. They specifically focused on determining the transcriptional profile of mono- and multi-nucleated cardiomyocytes. In this study, Yekelchyk and colleagues reported that adult rod-shaped cardiomyocytes are homogenous and that mono-, bi-, and multi-nucleated cells express a nearly identical set of genes, which was an unexpected observation. In contrast, induction of hypertrophy induced significant transcriptional changes in cardiomyocytes and generated substantial heterogeneity [16]. Cardiac fibroblasts are another population that displays extensive transcriptional heterogeneity in the heart. In the manuscript by Vidal et al., they used snRNA-seq to investigate the impact of ageing on cardiac cells [17]. They discovered that aged fibroblasts displayed the most pronounced differential gene expression pattern among all cell types, including dysregulation of inflammatory, extracellular matrix, angiogenesis, and osteogenic-related genes [17]. The study by Gladka et al. reported a disease-specific subpopulation of fibroblasts, and cytoskeleton-associated protein 4 (Ckap4) was identified as a novel marker specifically expressed in activated fibroblasts [14]. Single-cell transcriptome analysis was also used to generate the expression atlas of resident cardiac endothelial cells that promote neovascularization following ischemic injury [18]. Li et al. demonstrated that the structural integrity of adult cardiac endothelium post-myocardial infarction is mainly sustained through a clonal proliferation of resident endothelial cells [18]. By defining region-, age-, or disease-specific in different cell populations and corresponding gene expression signature, we might be able to identify novel molecular mechanisms relevant for cardiac disease and new therapies.
Studying Cellular Communication Between Cardiac Cell Types
Characterisation of the cardiac interaction network of cells form the heart is necessary for understanding cardiac homeostasis and disease. In 2018, Skelly and colleges reported that scRNA-seq is a great tool to study cellular communication in the heart. They performed scRNA-seq on a noncardiomyocyte fraction from a mouse heart, and based on a previously published dataset from humans [19], they generated a list of potential ligand-receptor pairs and applied it to their mouse scRNA-seq data. The extensive analysis revealed a dense intercellular communication network between all cardiac cell types, with fibroblasts being the most communicative cells [13]. Farbehi et al. also constructed a cell-to-cell communication network of ligand and receptor interactions based on the previously mentioned human ligand-receptor pair dataset [19]. They performed scRNA-seq on the noncardiomyocyte fraction of the healthy and injured heart. The obtained data showed dynamics in myeloid and fibroblast lineage post-myocardial infarction and provide insights for the analysis of cardiac homeostasis, inflammation, fibrosis, and regeneration. They also investigated a previously undescribed fibroblast lineage trajectory present in both sham and ischemic hearts [20]. Most cells express tens to hundreds of ligands and receptors to create a highly connected signalling network by multiple ligand-receptor pairs, yet the functional readout of this interaction on target gene expression in the recipient cells is not fully understood. Browaeys et al. [21] recently developed a tool called NicheNet for modelling intercellular communication on the single-cell level by linking ligands to target genes. By doing so, it is not only possible to model how gene expression of one cell is influenced by interaction with another cell, but also which signalling mediator may be involved [22].
Network and Trajectory Analysis
Analysis of network and trajectory is tightly connected to cell-to-cell communication, but it requires more extensive data analysis. Researchers use these analyses to define the pattern of a dynamic process encountered by single cells, based on their gene expression, and then organise cells depending on their progression through that process [23]. Weighted gene co-expression network analysis (WGCNA) is a commonly used data mining method for studying biological networks based on pairwise correlations between variables [24]. In the study of Nomura et al., they applied WGCNA to their single-cardiomyocyte transcriptomes and identified several co-expression gene modules [12]. This was followed by Monocle analysis which allows studying how cells choose between one of several possible end states [25, 26]. They found that after the reconstitution, the branched trajectory of cardiomyocytes from pressure overload mouse model was able to distinguish gene modules for cardiomyocyte hypertrophy and failure. The extensive analysis showed that during the early hypertrophic response in cardiomyocytes, there is an activation of mitochondrial genes closely correlated with ERK1/2 and NRF1/2. On the other hand, persistent overload leads to bifurcation into adaptive and failing stage via p57 signalling [12]. Another study used a trajectory analysis to investigate the molecular signature at different stages during the progression of pressure overload-induced cardiac hypertrophy. In the manuscript by Ren et al., it was reported that macrophage activation is a crucial factor for the development of cardiac hypertrophy, indicating cellular cross-talk between different cardiac cell types. The authors also showed that macrophage activation and subtype switching was successfully targeted by therapeutic intervention [27]. Wang et al. were able to construct cellular interactomes and regulatory signalling networks at multiple postnatal stages in the mouse heart [28]. In-depth analysis of the cellular network identified fibroblasts as a critical constituent in the microenvironment, promoting cardiomyocyte maturity. They also provide insights on how the manipulation of cardiomyocyte maturity impacts disease development and cardiac regeneration [28]. Another study by See et al. employed microfluidic flatform for single nuclear RNA-seq (snRNA-seq) to the frozen mouse and human failing and nonfailing adult hearts and revealed significant cellular heterogeneity [29]. They characterised these subgroups by WGCNA and discovered long intergenic noncoding RNAs (lincRNAs) as key nodal regulators. Knockdown of nodal lincRNAs affected the expression levels of genes related to dedifferentiation and proliferation within the same gene regulatory network. Additionally, they found that subpopulations of cardiomyocytes upregulate cell cycle–related genes, suggesting a unique endogenous regenerative potential [29]. There is no doubt that understanding how gene expression programs are controlled involves identifying regulatory interactions between transcription factors and target genes [30]. Numerous trajectory reconstitution algorithms are used to study the gradual transcriptomic response of cells to disease over time [12, 31]. Moreover, the application of WGCNA can characterise a network of co-expressed genes and together drive biological or pathological processes. Applying these analyses to the single-cell dataset not only can significantly advance the understanding of the processes driving cardiac pathologies but also shed light on new strategies for cell type- and stage-specific therapeutic interventions.
Identification of Rare Cells in the Heart
scRNA-seq allows us to characterise rare cell populations that are often overlooked in bulk sequencing approaches. Identifying rare cells is crucial to acquire a better understanding of normal or diseased tissue biology. To address this challenge, Grün et al. developed a RaceID algorithm suitable for rare cell type identification in a complex population of cells. They applied it for the first time to the intestine and identified regenerating islet-derived protein 4 (Reg4) as a novel marker for enteroendocrine cells, a rare population of hormone-producing intestinal cells [32].
Regenerative properties of the adult mammalian heart are insufficient to repair the heart post-ischemic injury. Many researchers seek to find evidence of cardiomyocyte renewal in the adult heart. Kretzschmar et al. attempted to look for resident cardiac stem cells (CSCs) that could contribute to cardiac regeneration [33]. Since the existence and significance of such cells in the heart are highly debated, they used scRNA-seq followed by the RaceID algorithm analysis and genetic lineage tracing to search for these rare cells. Unfortunately, they found no evidence for the existence of a quiescent CSC population or de novo generated cardiomyocytes that could contribute to cardiac repair [33]. Another study by Zhang and colleagues used multi-reporter mouse models and snRNA-seq to study cardiomyocyte dedifferentiation and proliferation. The obtained data suggest that increased endogenous cardiomyocyte renewal in post-ischemic hearts arises from the dedifferentiation and proliferation of pre-existing cardiomyocytes and not by cardiac differentiation of putative adult CPCs [34]. They pointed out that even prolonged cardiomyocyte dedifferentiation and subsequent proliferation can adversely affect cardiac function [35]. Thus, additional knowledge of specific regulators of both dedifferentiation and cell cycle reactivation will be required to be able to promote endogenous cardiomyocyte proliferation without diminishing heart function. More ongoing studies aim to find and characterise the rare population of proliferative cardiomyocytes that may explain why this process is so inefficient in the adult heart and perhaps how it can be manipulated for therapeutic benefit.
Application for Human Cardiac Tissue
Naturally, to confirm the relevance of all discoveries related to cardiac diseases performed in mouse models, they must be validated in human samples. Unfortunately, accessibility to fresh human cardiac tissue is not always possible. Biopsies from patients suffering from cardiac diseases could be a valuable option but to obtain cardiac tissue from healthy individuals is nearly impossible. Despite these challenges, several studies managed to apply scRNA-seq to the adult healthy and diseased human heart. In the manuscript of Nomura et al., they performed sequencing on intact cardiomyocytes manually isolated from healthy individuals and patients with dilated cardiomyopathy. They were able to validate the conservation of pathological transcriptional signatures that they previously identified in mouse [12]. In the study by Wang et al., the authors performed a comparative analysis of atrial and ventricular cells from healthy, failing, and partially recovered adult human hearts. They revealed the presence of cellular heterogeneity in cardiomyocytes and compartment-specific properties of the noncardiomyocyte fraction. Additionally, the authors showed that based on the cellular interaction network, cardiomyocytes’ contractility and metabolism are the essential features affected by cardiac function deterioration [36]. Commonly used droplet-based scRNA-seq systems (e.g., Drop-seq) are not applicable for intact adult human cardiomyocytes because of their large size. Therefore, many studies utilise frozen nuclei instead of intact cells for droplet-based systems. These approaches have high throughput and are highly suitable for frozen cardiac tissues stored in biobanks, which are much more accessible than fresh tissue. Selewa et al. applied Drop-seq technology to frozen nuclei from human heart tissue (DropNuc-seq) and demonstrated that this method could be successfully applied to identify different cardiac cells from frozen tissue [37]. The recent study of Litvinukova et al. used scRNA-seq and snRNA-seq to provide comprehensive transcriptomic data on six distinct cardiac regions of the adult human heart. Similar to other reports, they observed the cellular heterogeneity of cardiomyocytes, pericytes, and fibroblasts, and additionally, they analysed cell-to-cell interactions [38]. Microfluidic was also used by Tucker et al. to perform snRNA-seq of healthy human donors to identify and study the cellular and transcriptional diversity of the nonfailing human heart [39]. Hence, extensive analysis of the sequencing data from healthy adult human hearts on the single-cell level provides a comprehensive dataset of human cardiac cells and increases our understanding of cardiac biology.
Proper Platform Selection for the Best Results
Dispersing and sorting cardiac tissue into single cell are crucial steps for successful scRNA-seq. Unfortunately, there are considerable limitations for all available methods to dissociate and sort cells, especially important when selecting large adult mouse or human cardiomyocytes. Manual cell picking was employed in some studies [12], but this method is labour-intense and might be biased toward cells that are picked. Plate-based platforms like flow cytometry [13, 14, 33, 38] or integrated fluidic circuit [29] are commonly used techniques for large and fragile cells like cardiomyocytes, yet the number of cells that can be processed at once is limited. On the other hand, microfluidic platforms like 10x Genomics [17, 18, 20, 34, 38] or Drop-seq [37] are commercially available droplet-based platforms that allow for rapid profiling of thousands of individual cells. These microfluidic platforms are not suitable for cells bigger than 30 μm in diameter. Therefore, they are commonly used for smaller noncardiomyocytes or nuclei isolated from cardiomyocytes rather than intact cardiomyocytes. ICELL8 [16, 27, 28, 36] systems also enable high-throughput processing of hundreds of single cells without the typical microfluidic cell size restrains or the imaging limitations of droplet-based systems. Thereby, a careful selection of the most suitable platform that fits our needs is a crucial step for successful single-cell transcriptomic analysis.
Conclusions
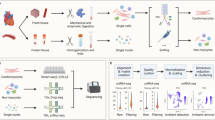
The exceptional resolution and valuable data obtained by scRNA-seq studies summarised here and by others (Fig. 2) continue to provide a new perspective into cardiac biology of healthy and diseased mammalian hearts. Increased understanding of molecular signalling specific for cardiac repair and regeneration [29, 33, 34], intercellular communication networks [12, 13, 20, 27,28,29], rare cell populations [33, 34], and new endogenous markers and mechanistic regulators of disease-associated processes [12, 14] will contribute to the development of novel, more effective therapeutic strategies. Perhaps in the future, combining scRNA-seq with spatial transcriptomics will further reveal spatial gene expression patterns in the single-cell resolution.
Change history
02 April 2021
A Correction to this paper has been published: https://doi.org/10.1007/s11897-021-00507-0
References
Wang Y, Navin NE. Advances and applications of single-cell sequencing technologies. Mol Cell. 2015;58(4):598–609. https://doi.org/10.1016/j.molcel.2015.05.005.
Hwang B, Lee JH, Bang D. Single-cell RNA sequencing technologies and bioinformatics pipelines. Exp Mol Med. 2018;50(8):96. https://doi.org/10.1038/s12276-018-0071-8.
Molenaar B, van Rooij E. Single-cell sequencing of the mammalian heart. Circ Res 2018;123(9):1033-1035. https://doi.org/10.1161/CIRCRESAHA.118.313531.
Stark R, Grzelak M, Hadfield J. RNA sequencing: the teenage years. Nat Rev Genet. 2019;20(11):631–56. https://doi.org/10.1038/s41576-019-0150-2.
Wang Z, Gerstein M, Snyder M. RNA-Seq: a revolutionary tool for transcriptomics. Nat Rev Genet. 2009;10(1):57–63. https://doi.org/10.1038/nrg2484.
Tang F, Barbacioru C, Wang Y, Nordman E, Lee C, Xu N, et al. mRNA-Seq whole-transcriptome analysis of a single cell. Nat Methods. 2009;6(5):377–82. https://doi.org/10.1038/nmeth.1315.
Klein AM, Mazutis L, Akartuna I, Tallapragada N, Veres A, Li V, et al. Droplet barcoding for single-cell transcriptomics applied to embryonic stem cells. Cell. 2015;161(5):1187–201. https://doi.org/10.1016/j.cell.2015.04.044.
Macosko EZ, Basu A, Satija R, Nemesh J, Shekhar K, Goldman M, et al. Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell. 2015;161(5):1202–14. https://doi.org/10.1016/j.cell.2015.05.002.
Muraro MJ, Dharmadhikari G, Grun D, Groen N, Dielen T, Jansen E, et al. A single-cell transcriptome atlas of the human pancreas. Cell Syst. 2016;3(4):385–94 e3. https://doi.org/10.1016/j.cels.2016.09.002.
DeLaughter DM, Bick AG, Wakimoto H, McKean D, Gorham JM, Kathiriya IS, et al. Single-cell resolution of temporal gene expression during heart development. Dev Cell. 2016;39(4):480–90. https://doi.org/10.1016/j.devcel.2016.10.001.
Li G, Xu A, Sim S, Priest JR, Tian X, Khan T, et al. Transcriptomic profiling maps anatomically patterned subpopulations among single embryonic cardiac cells. Dev Cell. 2016;39(4):491–507. https://doi.org/10.1016/j.devcel.2016.10.014.
Nomura S, Satoh M, Fujita T, Higo T, Sumida T, Ko T, et al. Cardiomyocyte gene programs encoding morphological and functional signatures in cardiac hypertrophy and failure. Nat Commun. 2018;9(1):4435. https://doi.org/10.1038/s41467-018-06639-7.
Skelly DA, Squiers GT, McLellan MA, Bolisetty MT, Robson P, Rosenthal NA, et al. Single-cell transcriptional profiling reveals cellular diversity and intercommunication in the mouse heart. Cell Rep. 2018;22(3):600–10. https://doi.org/10.1016/j.celrep.2017.12.072.
Gladka MM, Molenaar B, de Ruiter H, van der Elst S, Tsui H, Versteeg D, et al. Single-cell sequencing of the healthy and diseased heart reveals cytoskeleton-associated protein 4 as a new modulator of fibroblasts activation. Circulation. 2018;138(2):166–80. https://doi.org/10.1161/CIRCULATIONAHA.117.030742.
Panther F, Williams T, Ritter O. Inhibition of the calcineurin-NFAT signalling cascade in the treatment of heart failure. Recent Pat Cardiovasc Drug Discov. 2009;4(3):180–6. https://doi.org/10.2174/157489009789152276.
Yekelchyk M, Guenther S, Preussner J, Braun T. Mono- and multi-nucleated ventricular cardiomyocytes constitute a transcriptionally homogenous cell population. Basic Res Cardiol. 2019;114(5):36. https://doi.org/10.1007/s00395-019-0744-z.
Vidal R, Wagner JUG, Braeuning C, Fischer C, Patrick R, Tombor L, et al. Transcriptional heterogeneity of fibroblasts is a hallmark of the aging heart. JCI Insight. 2019;4(22). https://doi.org/10.1172/jci.insight.131092.
Li Z, Solomonidis EG, Meloni M, Taylor RS, Duffin R, Dobie R, et al. Single-cell transcriptome analyses reveal novel targets modulating cardiac neovascularization by resident endothelial cells following myocardial infarction. Eur Heart J. 2019;40(30):2507–20. https://doi.org/10.1093/eurheartj/ehz305.
Ramilowski JA, Goldberg T, Harshbarger J, Kloppmann E, Lizio M, Satagopam VP, et al. A draft network of ligand-receptor-mediated multicellular signalling in human. Nat Commun. 2015;6:7866. https://doi.org/10.1038/ncomms8866.
Farbehi N, Patrick R, Dorison A, Xaymardan M, Janbandhu V, Wystub-Lis K, et al. Single-cell expression profiling reveals dynamic flux of cardiac stromal, vascular and immune cells in health and injury. Elife. 2019;8. https://doi.org/10.7554/eLife.43882.
Browaeys R, Saelens W, Saeys Y. NicheNet: modeling intercellular communication by linking ligands to target genes. Nat Methods. 2020;17(2):159–62. https://doi.org/10.1038/s41592-019-0667-5.
Gladka MM. Cellular communication in a ‘virtual lab’: going beyond the classical ligand-receptor interaction. Cardiovasc Res. 2020;116(7):e67–e9. https://doi.org/10.1093/cvr/cvaa076.
Van den Berge K, Roux de Bezieux H, Street K, Saelens W, Cannoodt R, Saeys Y, et al. Trajectory-based differential expression analysis for single-cell sequencing data. Nat Commun. 2020;11(1):1201. https://doi.org/10.1038/s41467-020-14766-3.
Langfelder P, Horvath S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinformatics. 2008;9:559. https://doi.org/10.1186/1471-2105-9-559.
Qiu X, Mao Q, Tang Y, Wang L, Chawla R, Pliner HA, et al. Reversed graph embedding resolves complex single-cell trajectories. Nat Methods. 2017;14(10):979–82. https://doi.org/10.1038/nmeth.4402.
Qiu X, Hill A, Packer J, Lin D, Ma YA, Trapnell C. Single-cell mRNA quantification and differential analysis with census. Nat Methods. 2017;14(3):309–15. https://doi.org/10.1038/nmeth.4150.
Ren Z, Yu P, Li D, Li Z, Liao Y, Wang Y, et al. Single-cell reconstruction of progression trajectory reveals intervention principles in pathological cardiac hypertrophy. Circulation. 2020;141(21):1704–19. https://doi.org/10.1161/CIRCULATIONAHA.119.043053.
Wang Y, Yao F, Wang L, Li Z, Ren Z, Li D, et al. Single-cell analysis of murine fibroblasts identifies neonatal to adult switching that regulates cardiomyocyte maturation. Nat Commun. 2020;11(1):2585. https://doi.org/10.1038/s41467-020-16204-w.
See K, Tan WLW, Lim EH, Tiang Z, Lee LT, Li PYQ, et al. Single cardiomyocyte nuclear transcriptomes reveal a lincRNA-regulated de-differentiation and cell cycle stress-response in vivo. Nat Commun. 2017;8(1):225. https://doi.org/10.1038/s41467-017-00319-8.
Jackson CA, Castro DM, Saldi GA, Bonneau R, Gresham D. Gene regulatory network reconstruction using single-cell RNA sequencing of barcoded genotypes in diverse environments. Elife. 2020;9. https://doi.org/10.7554/eLife.51254.
Dick SA, Macklin JA, Nejat S, Momen A, Clemente-Casares X, Althagafi MG, et al. Self-renewing resident cardiac macrophages limit adverse remodeling following myocardial infarction. Nat Immunol. 2019;20(1):29–39. https://doi.org/10.1038/s41590-018-0272-2.
Grun D, Lyubimova A, Kester L, Wiebrands K, Basak O, Sasaki N, et al. Single-cell messenger RNA sequencing reveals rare intestinal cell types. Nature. 2015;525(7568):251–5. https://doi.org/10.1038/nature14966.
Kretzschmar K, Post Y, Bannier-Helaouet M, Mattiotti A, Drost J, Basak O, et al. Profiling proliferative cells and their progeny in damaged murine hearts. Proc Natl Acad Sci U S A. 2018;115(52):E12245–E54. https://doi.org/10.1073/pnas.1805829115.
Zhang Y, Gago-Lopez N, Li N, Zhang Z, Alver N, Liu Y, et al. Single-cell imaging and transcriptomic analyses of endogenous cardiomyocyte dedifferentiation and cycling. Cell Discov. 2019;5:30. https://doi.org/10.1038/s41421-019-0095-9.
Kubin T, Poling J, Kostin S, Gajawada P, Hein S, Rees W, et al. Oncostatin M is a major mediator of cardiomyocyte dedifferentiation and remodeling. Cell Stem Cell. 2011;9(5):420–32. https://doi.org/10.1016/j.stem.2011.08.013.
Wang L, Yu P, Zhou B, Song J, Li Z, Zhang M, et al. Single-cell reconstruction of the adult human heart during heart failure and recovery reveals the cellular landscape underlying cardiac function. Nat Cell Biol. 2020;22(1):108–19. https://doi.org/10.1038/s41556-019-0446-7.
Selewa A, Dohn R, Eckart H, Lozano S, Xie B, Gauchat E, et al. Systematic comparison of high-throughput single-cell and single-nucleus transcriptomes during cardiomyocyte differentiation. Sci Rep. 2020;10(1):1535. https://doi.org/10.1038/s41598-020-58327-6.
Litvinukova M, Talavera-Lopez C, Maatz H, Reichart D, Worth CL, Lindberg EL, et al. Cells of the adult human heart. Nature. 2020. https://doi.org/10.1038/s41586-020-2797-4.
Tucker NR, Chaffin M, Fleming SJ, Hall AW, Parsons VA, Bedi KC Jr, et al. Transcriptional and cellular diversity of the human heart. Circulation. 2020;142(5):466–82. https://doi.org/10.1161/CIRCULATIONAHA.119.045401.
Funding
MG was supported by Dr. Dekker fellowship from the Dutch Heart Foundation (NHS2016T009).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
No potential conflicts of interest relevant to this article were reported.
Human and Animal Rights and Informed Consent
All reported studies/experiments with human or animal subjects performed by the author have been previously published and complied with all applicable ethical standards (including the Helsinki declaration and its amendments, institutional/national research committee standards and international/national/institutional guidelines).
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article is part of the Topical Collection on Translational Research in Heart Failure
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Gladka, M.M. Single-Cell RNA Sequencing of the Adult Mammalian Heart—State-of-the-Art and Future Perspectives. Curr Heart Fail Rep 18, 64–70 (2021). https://doi.org/10.1007/s11897-021-00504-3
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11897-021-00504-3