Abstract
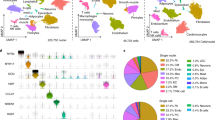
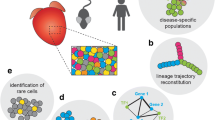
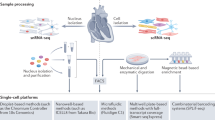
Heart failure is caused by a complicated pathogenic process and has a poor prognosis. Quality of life is often impaired due to repeated hospitalization. Integrative analysis of the morphological, physiological, and molecular profiles of cardiomyocytes, which are responsible mainly for heart contraction, may lead to a deeper understanding of the pathogenesis of heart failure. However, unlike other types of cells, cardiomyocytes are relatively large, vulnerable to stress, and difficult to use for single-cell analysis. With some ingenuity, we have established a single-cardiomyocyte analysis pipeline. Here, we describe the procedure for single-cell RNA sequencing of adult mouse cardiomyocytes from isolation to analysis.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Picelli S, Faridani OR, Björklund AK, Winberg G, Sagasser S, Sandberg R (2014) Full-length RNA-seq from single cells using smart-seq2. Nat Protoc 9:171–181
Andrews S (2010) FastQC: a quality control tool for high throughput sequence data. http://www.bioinformatics.babraham.ac.uk/projects/fastqc
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for illumina sequence data. Bioinformatics 30:2114–2120
Chen S, Zhou Y, Chen Y, Gu Y (2018) Fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34:i884–i890
Dobin A, Davis CA, Schlesinger F, Drenkow J, Zaleski C, Jha S, Batut P, Chaisson M, Gingeras TR (2013) STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29:15–21
Liao Y, Smyth GK, Shi W (2014) featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30:923–930
Stuart T, Butler A, Hoffman P, Hafemeister C, Papalexi E, Mauck WM III, Hao Y, Stoeckius M, Smibert P, Satija R (2019) Comprehensive integration of single-cell data. Cell 177:1888–1902
Qiu X, Mao Q, Tang Y, Wang L, Chawla R, Pliner HA, Trapnell C (2017) Reversed graph embedding resolves complex single-cell trajectories. Nat Methods 14:979–982. http://cole-trapnell-lab.github.io/monocle-release/docs/
Luecken MD, Theis FJ (2019) Current best practices in single-cell RNA-seq analysis: a tutorial. Mol Syst Biol 15:e8746
Acknowledgments
This work was supported by Grant-in-Aid for Young Scientists from the Japan Society for the Promotion of Science (to M.K.), Grant-in-Aid for Scientific Research (B) (to S.N.), Grant-in-Aid for Scientific Research (A) from the Japan Society for the Promotion of Science (to I.K.), and grants from the Japan Foundation for Applied Enzymology (to S.N.), the SENSHIN Medical Research Foundation (to S.N.), the KANAE Foundation for the Promotion of Medical Science (to S.N.), MSD Life Science Foundation (to S.N.), The Tokyo Biomedical Research Foundation (to S.N.), Astellas Foundation for Research on Metabolic Disorders (to S.N.), The Novartis Foundation (Japan) for the Promotion of Science (to S.N.), the Japanese Circulation Society (to S.N.), Takeda Science Foundation (to S.N.), and AMED (JP20gm0810013, JP20ek0109440, JP20ek0109487, JP20ek0109406, JP20km0405209, JP20bm0704026, JP20gm6210010, JP20ek0210141, JP20ek0210152, JP19bm0804010) (to S.N. and I.K.). The authors report that they have no relationships relevant to the contents of this chapter to disclose.
Author information
Authors and Affiliations
Corresponding authors
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Katoh, M., Nomura, S., Yamada, S., Aburatani, H., Komuro, I. (2021). Single-Cardiomyocyte RNA Sequencing to Dissect the Molecular Pathophysiology of the Heart. In: Yoshida, Y. (eds) Pluripotent Stem-Cell Derived Cardiomyocytes. Methods in Molecular Biology, vol 2320. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-1484-6_18
Download citation
DOI: https://doi.org/10.1007/978-1-0716-1484-6_18
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-1483-9
Online ISBN: 978-1-0716-1484-6
eBook Packages: Springer Protocols