Abstract
The 9p21.3 locus was the first to yield to genome-wide association studies (GWAS) seeking common genetic variants predisposing to increased risk of coronary artery atherosclerotic disease (CAD). The 59 single nucleotide polymorphisms that show highest association with CAD are clustered in a region 100,000 to 150,000 base pairs 5′ to the cyclin-dependent kinase inhibitors CDKN2B (coding for p15ink4b) and CDKN2A (coding for p16ink4a and p14ARF). This region also covers the 3′ end of a long noncoding RNA transcribed antisense to CDKN2B (CDKN2BAS, aka ANRIL for antisense noncoding RNA at the ink4 locus) whose expression has been linked to chromatin remodeling at the locus. Despite intensive investigation over the past 7 years, the functional significance of the 9p21.3 locus remains elusive. Other variants at this locus have been associated with glaucoma, glioma, and type 2 diabetes mellitus, diseases that implicate tissue-resident macrophages. Here, we review the evidence that genetic variants at 9p21.3 disrupt tissue-specific enhancers and propose new insights to guide future studies.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Coronary artery disease (CAD) is the leading cause of death in the world, according to World Health Organization statistics [1]. Although fewer than 5 % of individuals show symptoms of CAD before the age of 50 [2], intravascular ultrasound revealed that 1 in 6 adolescents and 85 % of persons over the age of 50 have measurable coronary atherosclerosis [3]. Modifiable risk factors such as high blood cholesterol, diabetes, high blood pressure, smoking, and obesity contribute to the risk of CAD [4], but genetic risk is equally important [5]. The notion that common genetic variants could contribute to the risk of CAD was borne out by the discovery of the first common CAD risk variant to be discovered by the genome-wide association study (GWAS) approach, located on chromosome 9 at 9p21.3 [6–8].
The 9p21 risk alleles are carried by 75 % of the world population (excluding black Africans) and confer risk for coronary atherosclerosis independently of known risk factors [6–8]. To date, more than 40 loci have been reported to contribute to CAD risk [9], but the 9p21.3 locus remains the second most significant (after LPA, encoding lipoprotein (a)). The population-attributable risk of CAD conferred by homozygosity for the 9p21.3 risk allele is 21 % [10]. It is important to realize that this is on the same order of magnitude as the population-attributable risk of hypertension (28 % in men and 29 % in women) [11] or of elevated total cholesterol (27 % in men and 34 % in women) [11]. The quest to identify the mechanism whereby the 9p21.3 confers risk for CAD has been ongoing for the past 7 years, and progress has been limited. Here, we will review what we know about the 9p21.3 locus and how evidence has gradually emerged to point to possible mechanisms.
Atherosclerosis—a Common Inflammatory Disease of Arteries
CAD is an age progressive disease caused by atherosclerosis of the coronary arteries, an inflammatory condition in which the artery becomes occluded due to the accumulation of fatty material called plaque. Plaque is an aggregate of cholesterol and triglycerides taken up by tissue macrophages within the vessel wall to form foam cells [12]. Macrophage foam cells produce high levels of inflammatory cytokines and chemokines. That macrophages accumulate at atherosclerotic lesions is well documented [13], but the prevailing view that this is due to monocytes being recruited by local production of chemokines [14–20] has recently been challenged with evidence that recruited monocytes account for only about 15 % of the proliferating macrophages in atherosclerotic lesions [21]. Indeed, several recent studies indicate that tissue macrophages, while exhibiting distinct mature differentiated phenotypes, are endowed with the capacity for self-renewal similar to that of stem cells [22]. Genetic variants that influence macrophage self-renewal by affecting cell cycle control genes would be expected to affect the severity of atherosclerosis plaque formation.
The 9p21.3 Genetic Risk Locus Promotes Atherosclerosis Rather Than Myocardial Infarction
We reported that the 9p21.3 risk alleles associated with angiographic CAD [8], whereas the deCODE Genetics group reported that it associated with myocardial infarction (MI) [7], and the Wellcome Trust Case Control Consortium found that it associated with CAD broadly defined (but predominantly recruited from patients with MI) [6]. It is well recognized that MI occurs on a substrate of coronary atherosclerosis. Thus, it was unclear whether these risk alleles promote the deposition of atherosclerotic lesions (CAD per se) or the formation of vulnerable plaque that are prone to rupture triggering thrombosis and MI. To address this question, we conducted a cross-sectional survey of 1714 patients with angiographically documented CAD and found that the risk allele associated with severity of CAD as measured by the number of occluded coronaries rather than MI among cases with pre-existing CAD [23•]. Shortly thereafter, the same finding was also reported by Patel et al. [24] who examined 2334 angiographic CAD cases. A recent meta-analysis comprising 21 studies and 33,673 subjects with angiographic data and MI status also confirmed that the 9p21.3 risk locus increases the burden of coronary atherosclerosis but not the risk of MI in the presence of underlying CAD [25]. Other vascular phenotypic manifestations of the 9p21.3 risk locus include abdominal aortic aneurysm [26–29], intracranial aneurysm [30, 31], the ankle-brachial pressure index, an indication of peripheral vascular disease [32], late onset Alzheimer’s disease and vascular dementia [33], and ischemic stroke [34–37]. Thus, these findings are consistent with an effect of the 9p21.3 risk alleles to stimulate coronary atherosclerosis. Whether the effect is mediated through resident macrophages of the vessel wall or by modifying the cellular properties of arterial vascular smooth muscle cells (VSMCs) or a combination thereof remains unclear.
The 9p21 Risk Locus Disrupts a Tissue-Specific Enhancer
The genetic risk locus at 9p21.3 consists of a cluster of 59 linked single nucleotide polymorphisms (SNPs) over a 53,202 bp region (Fig. 1). A growing consensus is that 1 or several of the 59 linked SNPs alter the DNA sequence and disrupt or create transcription factor binding sites that would alter the control of regional gene expression. The nearest protein coding genes are the cyclin-dependent kinase inhibitors CDKN2A (coding for p16ink4a and p14ARF) and CDKN2B (coding for p15ink4b) that lie about 100,000 base pairs upstream of the 9p21.3 locus. In addition, we find methylthioadenosine phosphorylase (MTAP) and much farther upstream nearly the entire type I interferon gene cluster.
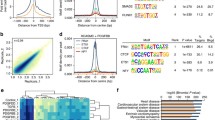
The 9p21.3 locus is a hotbed of haplotypes linked to several diseases: are these cell-specific enhancers? The coronary artery disease (CAD, in red) haplotype block spans 53,000 bp and includes 59 linked SNPs. Other GWAS loci include 1 for primary open angle glaucoma/normal pressure glaucoma (POAG/NPG) spanning 63,000 bp, another for glioma spanning 28,000 bp, and 1 for type 2 diabetes mellitus (T2D) spanning approximately 12,000 bp. It is noteworthy that the POAG/NPG and glioma haplotypes are partially dependent (eg, rs2157719 for glaucoma and rs4977756 for glioma have a D’ of 0.905, r2 = 0.718 and a LOD score of 31.79 according to HapMap). Similarly, the Glioma and CAD haplotypes are also partially dependent (rs4977756 of the glioma haplotype and rs1333049 of the CAD haplotype have a D’ of 0.824, an r2 = 0.422 and a LOD score of 16.53). In contrast, the T2D haplotype is independent of the other haplotypes (eg, rs1333049 and rs10811661 have a D’ of 0.018, r2 = 0 and LOD score of 0)
We identified several functional enhancer elements at the 9p21 region [38•]. Deletion of the same region in the mouse genome was associated with reduced expression of Cdkn2a and Cdkn2b mRNAs, demonstrating the presence of regulatory enhancers at this locus [39••]. A recent study published in Nature by Kelly Frazer’s group [40•] used the technique of chromatin conformation capture and identified short- and long-range interactions between sequences at the 9p21.3 locus and sequences in the vicinity of the genes encoding CDKN2A and CDKN2B and methylthioadenosine phosphorylase (MTAP) in the short range, and between IFNW1 and interferon-α21 (IFNA21), in the long range, approximately 1 million base pairs upstream on chromosome 9. This finding is remarkable because it suggests that the influence of the enhancer sequences at 9p21.3 act at considerably greater distances than previously thought. Although these authors did not examine the expression of the type I interferon genes, a more recent study failed to find association between the circulating levels of various type 1 interferons, including IFNA21, and the 9p21 risk allele genotype in healthy individuals [41•]. Whereas these findings cast doubt as to the significance of the long-range interactions between the 9p21 risk alleles and distant enhancers, the effect of the 9p21.3 locus may reflect a tissue-specific enhancer or a disease-associated effect that has not yet been replicated in cultured cells. A recent study performed expression profiling of 9p21.3 genotyped monocyte-derived macrophages in culture and remarkably, found differential expression of genes not at the 9p21.3 locus, but rather of the chemokines CCL8 and CCL2 and the lectins CLEC4E and CLEC5A [42•]. This study also did not find significant changes in type 1 interferon expression by 9p21.3 genotype.
Haplotype Analysis Reveals Multiple Phenotypes at the 9p21.3 Locus
Tissue-specific enhancers are typically located at some distance from gene promoters [43]. If the 9p21.3 CAD risk locus disrupts sequences of a tissue-specific enhancer that control tissue-resident macrophage proliferation in the arterial wall and worsen coronary atherosclerosis, might other regions of the 9p21.3 locus affect cellular functions in other tissues?
When we further examined the 9p21.3 locus for association of specific haplotypes (groups of co-inherited SNPs) to different phenotypes we found 2 haplotypes, the well-known locus tagged by rs1333049 that associated with atherosclerosis and another tagged by rs518394 closer to CDKN2B that associated with MI, but only among cases of clinically significant (>50 % stenosis) angiographic CAD who were carriers of the rs1333049 non-risk allele [44•] (Fig. 1). Another surprise was that the MI haplotype was inversely associated with the severity of coronary atherosclerosis, suggesting that it exerts its effect on a minimal burden of plaque. A SNP linked to this region was previously associated with elevated platelet reactivity [45•]. Intriguingly, recent evidence points to a cross-talk between platelets and macrophages to clear blood borne bacterial pathogens, so that this dual haplotype may relate to an important component of the innate immune response [46].
Macrophages are the progeny of the myeloid lineage that includes Kupffer cells of the liver, tissue-resident macrophages of arteries, adipose tissue and pancreas, and microglia of the central nervous system. Genome-wide association studies have identified several diseases affected by polymorphisms at the 9p21.3 locus, including primary open angle glaucoma (PAOG) [47] and normal pressure glaucoma (NPG) [48, 49], glioma [50, 51], and type 2 diabetes (T2D) [52, 53] (Fig. 1). It is intriguing that in every case tissue macrophages are implicated in the disease process. Microglia worsen the outcome of glaucoma [49, 54, 55], glioma is often difficult to distinguish from gliosis [56], a condition in which microglia proliferate in response to pathogens [57], and in type 2 diabetes, where inflammatory macrophages proliferate in the vicinity of islets [58].
Mice hemizygous for Mtap (with 1 copy of Mtap deleted) were found to have more significant atherosclerotic lesions when bred on a susceptible background [59]. Whether this observation is the result of deficient MTAP function or reflects altered chromatin remodeling due to loss of one allele remains to be determined.
Chromatin Remodeling at the 9p21.3 Locus
CTCF binding sequences are found between the CDKN2A gene and the antisense ANRIL transcript forming a bidirectional promoter [60–62]. Expression of ANRIL and CDKN2A are both dependent on CTCF [61]. CTCF is a multifunctional transcription regulator that can not only activate 1 gene, but at the same time prevent inappropriate activation of another gene on the same chromosomal region. Thus, CTCF also acts as an insulator [63]. CTCF binds to unmethylated DNA target sequences. CTCF contains 11 zinc fingers and through its ability to bind DNA and homodimerize, CTCF can bring together distant sequences to form higher order chromatin structure [64]. CTCF has been implicated in long range intra- and even inter-chromosomal interactions [64, 65].
CTCF binds and activates the CDKN2A locus in a manner that is sensitive to DNA methylation [61, 62]. DNA methylation is not restricted to imprinted genes but is widely present across the genome [66, 67]. Importantly, increased genomic DNA methylation has been described in atherosclerotic lesions [68] and in peripheral lymphocytes of patients with CAD relative to controls [69]. The 9p21 CAD risk locus enhancer sequences located within the 3′ region of the ANRIL gene strongly upregulate transcription of ANRIL, and alter the levels of its multiple alternatively spliced isoforms [38•, 40•]. As a consequence, components of the polycomb repressor complex (PRC1 and PRC2) are recruited to the ANRIL transcript and silence gene expression from the CDKN2A locus, in part by histone 3 lysine 27 trimethylation by polycomb complex protein EZH2 [70, 71]. EZH2 also recruits the DNA methyltransferase DNMT1 [72] and this would also increase DNA methylation and inactivation of the CDKN2A locus. Zhuan et al. found that methylation of CDKN2B and elevated expression of ANRIL were associated with coronary artery disease in a Chinese angiographic study, but this was not tightly correlated to the 9p21 risk genotype [73]. This result suggests that allele-specific methylation is unlikely to participate in gene regulation at the 9p21.3 locus.
When cells differentiate or age they typically stop proliferating, and this requires activation of the cyclin dependent kinase inhibitors like p16ink4a [74]. Deletion of the 9p21.3 syntenic region in the mouse reversed cellular senescence of primary fibroblasts and VSMCs [39••]. Primary cultures of human arterial VSMCs showed reduced expression of p16ink4a and p15ink4b and increased cellular proliferation [75•]. Thus, the 9p21.3 locus appears to cause loss of CDKN2A and CDKN2B suppression. Activation of the CDKN2A locus requires displacement and eviction of PRC1 and PRC2, and the chromatin remodeling protein BRG1 plays a critical role in this process [76]. BRG1 interacts with CTCF and participates in remodeling long range interactions at the major histocompatibility complex locus [77]. BRG1 is also known to interact with several transcription factors, including GATA1 [78], STAT2 [79], and TEAD1 [80], so that these transcription factors are good candidates to regulate chromatin structure at the 9p21.3 locus.
The Nature of the Altered Regulatory Sequences at the 9p21.3 Risk Locus
We reported that the risk allele of one of the SNPs (rs1333045) alters a TGFβ consensus sequence within an enhancer and reduced its function [38•]. Although we have not formally tested whether this is a true functional TGFβ site, this result suggests that sequences at or in the vicinity of this SNP are functional. A different hypothesis was proposed by the Frazer group. They suggested that another SNP (rs10757278) disrupts a putative binding site for the signal transducer of activated T cells 1 (STAT1), the transcription factor that mediates IFN-γ-inducible gene expression [40•]. However, our recent study showed that the upregulated expression of p15ink4b and p16ink4a in response to IFN-γ was not affected by the 9p21.3 risk allele, and in fact occurs largely by a posttranscriptional mechanism [81•]. Thus, a differential response to IFN-γ does not account for the 9p21 risk effect.
TGFβ Signaling Inhibits Macrophage Foam Cell Formation
TGFβ is a cytokine that inhibits macrophage foam cell formation [82–84]. Low levels of plasma TGFβ are associated with a poor outcome in CAD [85]. Expression of a dominant negative mutant of the TGFβ receptor increases foam cell formation and worsens atherosclerosis [86]. Conversely, over-expression of TGFβ in macrophages reduces foam cell formation and atherosclerosis [87]. Together, these findings establish TGFβ as a potent anti-atherogenic factor.
What underlies the anti-atherogenic effect of TGFβ? Evidence suggests this comes about several ways. TGFβ suppresses the uptake and accumulation of oxidized LDL-cholesterol in macrophages by lowering the expression of the scavenger receptor CD36 [82, 83]. TGFβ also increases the export of cholesterol from macrophages by upregulating the expression of the cholesterol transporter ABCA1 [82, 84]. Macrophage proliferation also contributes to atherosclerosis. Ablation of the cell cycle inhibitor p27Kip1 in hematopoietic progenitors accelerates macrophage proliferation and atherosclerosis [88]. Conversely, TGFβ blocks proliferation of human macrophages derived from myeloid cells stimulated by macrophage-colony stimulating factor (M-CSF) [89]. TGFβ has been shown to activate the expression of the cyclin-dependent kinase inhibitors CDKN2B (p15ink4b) and CDKN2A (p16ink4a) [90–92]. Of these TGFβ-responsive genes, only CDKN2A and CDKN2B are located in the vicinity and are under the direct control of the 9p21.3 genetic risk locus. Thus, genetic polymorphisms like rs1333045 that disrupt TGFβ-responsive elements at the 9p21.3 risk locus would be predicted to interfere with TGFβ-mediated suppression of macrophage or VSMC proliferation and worsen atherosclerosis.
TEAD Transcription Factors Interact with SMAD3 and Cooperate in TGFβ Signaling
TGFβ signaling activates SMAD transcription factors to regulate gene expression. TGFβ-responsive genes can be activated directly by SMADs through SMAD-responsive elements, or by other transcription factors via their interaction with SMAD proteins [93]. For example, TEAD transcription factors interact with SMAD3 and mediate TGFβ-dependent gene activation [94–96]. Recent evidence shows that TEAD/SMAD interaction controls proliferation of self-renewing cells [97] and that selective knockdown of TEAD transcription factors affects cellular senescence though a p16inka-dependent pathway. Of particular interest, the activation of p14ARF by TGFβ2 in the eye is abrogated by deletion of the 9p21.3 risk locus syntenic region on mouse chromosome 4 [98•].
TEAD transcription factors are one of a small group of transcription factors that not only can regulate expression of genes in the proximity of TEAD-binding regulatory promoter/enhancer elements (in the short-range) [99], but can also activate genes at a distance by recruiting chromatin remodeling proteins [80, 100]. A recent study found that a chromatin remodeling protein is recruited to 2 TEAD binding sites within the 9p21.3 risk locus [100]. This result shows that TEAD factors are present at the 9p21.3 risk locus and may participate in its long-range regulation of gene expression.
Conclusions
The mechanism whereby the 9p21.3 locus confers increased susceptibility to coronary atherosclerosis remains elusive. SNPs likely disrupt specific regulatory sequences within tissue-specific enhancers. Identifying these functional SNPs and the cells in which they are functional is a challenge not just for atherosclerosis but for other diseases that also associate with this locus.
References
Papers of particular interest, published recently, have been highlighted as: • Of importance •• Of major importance
2011 [cited December 09, 2012]; Available at: http://www.who.int/mediacentre/factsheets/fs310/en/index.html. Accessed 7 Sept 2013.
Prevalence of coronary heart disease—United States, 2006–2010. Morb Mortal Wkly Rep. 2011;60:1377–81.
Tuzcu EM, Kapadia SR, Tutar E, Ziada KM, Hobbs RE, McCarthy PM, et al. High prevalence of coronary atherosclerosis in asymptomatic teenagers and young adults: evidence from intravascular ultrasound. Circulation. 2001;103:2705–10.
Yusuf S, Hawken S, Ounpuu S, Dans T, Avezum A, Lanas F, et al. Effect of potentially modifiable risk factors associated with myocardial infarction in 52 countries (the INTERHEART study): case–control study. Lancet. 2004;364:937–52.
Roberts R, Stewart AFR. Genes and coronary artery disease: where are we? J Am Coll Cardiol. 2012;60:1715–21.
Wellcome Trust Case Control Consortium. Genome-wide association study of 14,000 cases of seven common diseases and 3,000 shared controls. Nature. 2007;447:661–78.
Helgadottir A, Thorleifsson G, Manolescu A, Gretarsdottir S, Blondal T, Jonasdottir A, et al. A common variant on chromosome 9p21 affects the risk of myocardial infarction. Science. 2007;316:1491–3.
McPherson R, Pertsemlidis A, Kavaslar N, Stewart A, Roberts R, Cox DR, et al. A common allele on chromosome 9 associated with coronary heart disease. Science. 2007;316:1488–91.
Deloukas P, Kanoni S, Willenborg C, Farrall M, Assimes TL, Thompson JR, et al. Large-scale association analysis identifies new risk loci for coronary artery disease. Nat Genet. 2013;45:25–33.
Anderson JL, Horne BD, Kolek MJ, Muhlestein JB, Mower CP, Park JJ, et al. Genetic variation at the 9p21 locus predicts angiographic coronary artery disease prevalence but not extent and has clinical utility. Am Heart J. 2008;156:1155–62, e2.
Wilson PW, D’Agostino RB, Levy D, Belanger AM, Silbershatz H, Kannel WB. Prediction of coronary heart disease using risk factor categories. Circulation. 1998;97:1837–47.
Ouimet M, Marcel YL. Regulation of lipid droplet cholesterol efflux from macrophage foam cells. Arterioscler Thromb Vasc Biol. 2012;32:575–81.
Libby P, Ridker PM, Maseri A. Inflammation and atherosclerosis. Circulation. 2002;105:1135–43.
Kim CJ, Khoo JC, Gillotte-Taylor K, Li A, Palinski W, Glass CK, et al. Polymerase chain reaction-based method for quantifying recruitment of monocytes to mouse atherosclerotic lesions in vivo: enhancement by tumor necrosis factor-alpha and interleukin-1 beta. Arterioscler Thromb Vasc Biol. 2000;20:1976–82.
Lessner SM, Prado HL, Waller EK, Galis ZS. Atherosclerotic lesions grow through recruitment and proliferation of circulating monocytes in a murine model. Am J Pathol. 2002;160:2145–55.
Swirski FK, Pittet MJ, Kircher MF, Aikawa E, Jaffer FA, Libby P, et al. Monocyte accumulation in mouse atherogenesis is progressive and proportional to extent of disease. Proc Natl Acad Sci U S A. 2006;103:10340–5.
Swirski FK, Libby P, Aikawa E, Alcaide P, Luscinskas FW, Weissleder R, et al. Ly-6Chi monocytes dominate hypercholesterolemia-associated monocytosis and give rise to macrophages in atheromata. J Clin Invest. 2007;117:195–205.
Tacke F, Alvarez D, Kaplan TJ, Jakubzick C, Spanbroek R, Llodra J, et al. Monocyte subsets differentially employ CCR2, CCR5, and CX3CR1 to accumulate within atherosclerotic plaques. J Clin Invest. 2007;117:185–94.
Combadiere C, Potteaux S, Rodero M, Simon T, Pezard A, Esposito B, et al. Combined inhibition of CCL2, CX3CR1, and CCR5 abrogates Ly6C(hi) and Ly6C(lo) monocytosis and almost abolishes atherosclerosis in hypercholesterolemic mice. Circulation. 2008;117:1649–57.
Saederup N, Chan L, Lira SA, Charo IF. Fractalkine deficiency markedly reduces macrophage accumulation and atherosclerotic lesion formation in CCR2−/− mice: evidence for independent chemokine functions in atherogenesis. Circulation. 2008;117:1642–8.
Robbins CS, Hilgendorf I, Weber GF, Theurl I, Iwamoto Y, Figueiredo JL, et al. Local proliferation dominates lesional macrophage accumulation in atherosclerosis. Nat Med. 2013;19:1166–72.
Sieweke MH, Allen JE. Beyond stem cells: self-renewal of differentiated macrophages. Science. 2013;342:1242974.
Dandona S, Stewart AFR, Chen L, Williams K, So D, O’Brien E, et al. Gene dosage of the common variant 9p21 predicts severity of coronary artery disease. J Am Coll Cardiol. 2010;56:479–86. This study was the first to document that the 9p21.3 risk locus associates with atherosclerosis rather than myocardial infarction.
Patel RS, Su S, Neeland IJ, Ahuja A, Veledar E, Zhao J, et al. The chromosome 9p21 risk locus is associated with angiographic severity and progression of coronary artery disease. Eur Heart J. 2010;31:3017–23.
Chan K, Patel RS, Newcombe P, Nelson CP, Qasim A, Epstein SE, et al. Association between the chromosome 9p21 locus and angiographic coronary artery disease burden: a collaborative meta-analysis. J Am Coll Cardiol. 2013;61:957–70.
Bjorck HM, Lanne T, Alehagen U, Persson K, Rundkvist L, Hamsten A, et al. Association of genetic variation on chromosome 9p21.3 and arterial stiffness. J Intern Med. 2009;265:373–81.
Bown MJ, Braund PS, Thompson J, London NJ, Samani NJ, Sayers RD. Association between the coronary artery disease risk locus on chromosome 9p21.3 and abdominal aortic aneurysm. Circ Cardiovasc Genet. 2008;1:39–42.
Helgadottir A, Thorleifsson G, Magnusson KP, Gretarsdottir S, Steinthorsdottir V, Manolescu A, et al. The same sequence variant on 9p21 associates with myocardial infarction, abdominal aortic aneurysm and intracranial aneurysm. Nat Genet. 2008;40:217–24.
Thompson AR, Golledge J, Cooper JA, Hafez H, Norman PE, Humphries SE. Sequence variant on 9p21 is associated with the presence of abdominal aortic aneurysm disease but does not have an impact on aneurysmal expansion. Eur J Hum Genet. 2009;17:391–4.
Yasuno K, Bilguvar K, Bijlenga P, Low SK, Krischek B, Auburger G, et al. Genome-wide association study of intracranial aneurysm identifies three new risk loci. Nat Genet. 2010;42:420–5.
Foroud T, Koller DL, Lai D, Sauerbeck L, Anderson C, Ko N, et al. Genome-wide association study of intracranial aneurysms confirms role of Anril and SOX17 in disease risk. Stroke. 2012;43:2846–52.
Murabito JM, White CC, Kavousi M, Sun YV, Feitosa MF, Nambi V, et al. Association between chromosome 9p21 variants and the ankle-brachial index identified by a meta-analysis of 21 genome-wide association studies. Circ Cardiovasc Genet. 2012;5:100–12.
Emanuele E, Lista S, Ghidoni R, Binetti G, Cereda C, Benussi L, et al. Chromosome 9p21.3 genotype is associated with vascular dementia and Alzheimer’s disease. Neurobiol Aging. 2011;32:1231–5.
Dichgans M, Malik R, Konig IR, Rosand J, Clarke R, Gretarsdottir S, et al. Shared genetic susceptibility to ischemic stroke and coronary artery disease: a genome-wide analysis of common variants. Stroke. 2014;45:24–36.
Ding H, Xu Y, Wang X, Wang Q, Zhang L, Tu Y, et al. 9p21 is a shared susceptibility locus strongly for coronary artery disease and weakly for ischemic stroke in Chinese Han population. Circ Cardiovasc Genet. 2009;2:338–46.
Gschwendtner A, Bevan S, Cole JW, Plourde A, Matarin M, Ross-Adams H, et al. Sequence variants on chromosome 9p21.3 confer risk for atherosclerotic stroke. Ann Neurol. 2009;65:531–9.
Matarin M, Brown WM, Singleton A, Hardy JA, Meschia JF. Whole genome analyses suggest ischemic stroke and heart disease share an association with polymorphisms on chromosome 9p21. Stroke. 2008;39:1586–9.
Jarinova O, Stewart AFR, Roberts R, Wells G, Lau P, Naing T, et al. Functional analysis of the chromosome 9p21.3 coronary artery disease risk locus. Arterioscler Thromb Vasc Biol. 2009;29:1671–7. This study was the first to identify functional enhancers at the 9p21.3 risk locus.
Visel A, Zhu Y, May D, Afzal V, Gong E, Attanasio C, et al. Targeted deletion of the 9p21 noncoding coronary artery disease risk interval in mice. Nature. 2010;464:409–12. This study demonstrated that the 9p21.3 syntenic sequences in mouse chromosome 4 contain enhancer sequences that regulate CDKN2A and CDKN2B expression.
Harismendy O, Notani D, Song X, Rahim NG, Tanasa B, Heintzman N, et al. 9p21 DNA variants associated with coronary artery disease impair interferon-gamma signalling response. Nature. 2011;470:264–8. This study proposed a mechanism whereby a SNP at the 9p21.3 locus confers risk for CAD. This study also demonstrated long-range effects of the enhancer sequences at the 9p21.3 locus, modifying chromatin structure in the vicinity of type 1 interferon genes, but see references 41, 42, and 81.
Erridge C, Gracey J, Braund PS, Samani NJ. The 9p21 locus does not affect risk of coronary artery disease through induction of type 1 interferons. J Am Coll Cardiol. 2013;62:1376–81. This study failed to find an effect of the 9p21.3 locus on type 1 interferon expression in plasma of healthy donors or in monocyte-derived macrophages.
Zollbrecht C, Grassl M, Fenk S, Hocherl R, Hubauer U, Reinhard W, et al. Expression pattern in human macrophages dependent on 9p21.3 coronary artery disease risk locus. Atherosclerosis. 2013;227:244–9. This study compared whole genome RNA expression profiles of monocyte-derived macrophages from 9p21.3-genotyped CAD patients and found no difference in type 1 interferon expression.
Pennacchio LA, Loots GG, Nobrega MA, Ovcharenko I. Predicting tissue-specific enhancers in the human genome. Genome Res. 2007;17:201–11.
Fan M, Dandona S, McPherson R, Allayee H, Hazen SL, Wells GA, et al. Two chromosome 9p21 haplotype blocks distinguish between coronary artery disease and myocardial infarction risk. Circ Cardiovasc Genet. 2013;6:372–80. This study identified an additional haplotype at the 9p21.3 locus that associates with myocardial infarction in carriers of the 9p21.3 non-risk allele, consistent with reference 45.
Musunuru K, Post WS, Herzog W, Shen H, O’Connell JR, McArdle PF, et al. Association of single nucleotide polymorphisms on chromosome 9p21.3 with platelet reactivity: a potential mechanism for increased vascular disease. Circ Cardiovasc Genet. 2010;3:445–53. This study was the first to identify additional phenotypes tied to platelet reactivity in the vicinity of the 9p21.3 CAD risk locus.
Wong CH, Jenne CN, Petri B, Chrobok NL, Kubes P. Nucleation of platelets with blood-borne pathogens on Kupffer cells precedes other innate immunity and contributes to bacterial clearance. Nat Immunol. 2013;14:785–92.
Osman W, Low SK, Takahashi A, Kubo M, Nakamura Y. A genome-wide association study in the Japanese population confirms 9p21 and 14q23 as susceptibility loci for primary open angle glaucoma. Hum Mol Genet. 2012;21:2836–42.
Takamoto M, Kaburaki T, Mabuchi A, Araie M, Amano S, Aihara M, et al. Common variants on chromosome 9p21 are associated with normal tension glaucoma. PLoS One. 2012;7:e40107.
Wiggs JL, Yaspan BL, Hauser MA, Kang JH, Allingham RR, Olson LM, et al. Common variants at 9p21 and 8q22 are associated with increased susceptibility to optic nerve degeneration in glaucoma. PLoS Genet. 2012;8:e1002654.
Rajaraman P, Melin BS, Wang Z, McKean-Cowdin R, Michaud DS, Wang SS, et al. Genome-wide association study of glioma and meta-analysis. Hum Genet. 2012;131:1877–88.
Shete S, Hosking FJ, Robertson LB, Dobbins SE, Sanson M, Malmer B, et al. Genome-wide association study identifies five susceptibility loci for glioma. Nat Genet. 2009;41:899–904.
Parra EJ, Below JE, Krithika S, Valladares A, Barta JL, Cox NJ, et al. Genome-wide association study of type 2 diabetes in a sample from Mexico City and a meta-analysis of a Mexican-American sample from Starr County, Texas. Diabetologia. 2011;54:2038–46.
Manning AK, Hivert MF, Scott RA, Grimsby JL, Bouatia-Naji N, Chen H, et al. A genome-wide approach accounting for body mass index identifies genetic variants influencing fasting glycemic traits and insulin resistance. Nat Genet. 2012;44:659–69.
Bosco A, Crish SD, Steele MR, Romero CO, Inman DM, Horner PJ, et al. Early reduction of microglia activation by irradiation in a model of chronic glaucoma. PLoS One. 2012;7:e43602.
Ebneter A, Casson RJ, Wood JP, Chidlow G. Microglial activation in the visual pathway in experimental glaucoma: spatiotemporal characterization and correlation with axonal injury. Invest Ophthalmol Vis Sci. 2010;51:6448–60.
Rivera-Zengotita M, Yachnis AT. Gliosis vs glioma? Don’t grade until you know. Adv Anat Pathol. 2012;19:239–49.
Ajami B, Bennett JL, Krieger C, Tetzlaff W, Rossi FM. Local self-renewal can sustain CNS microglia maintenance and function throughout adult life. Nat Neurosci. 2007;10:1538–43.
Ehses JA, Perren A, Eppler E, Ribaux P, Pospisilik JA, Maor-Cahn R, et al. Increased number of islet-associated macrophages in type 2 diabetes. Diabetes. 2007;56:2356–70.
Kim JB, Deluna A, Mungrue IN, Vu C, Pouldar D, Civelek M, et al. Effect of 9p21.3 coronary artery disease locus neighboring genes on atherosclerosis in mice. Circulation. 2012;126:1896–906.
Kim TH, Abdullaev ZK, Smith AD, Ching KA, Loukinov DI, Green RD, et al. Analysis of the vertebrate insulator protein CTCF-binding sites in the human genome. Cell. 2007;128:1231–45.
Rodriguez C, Borgel J, Court F, Cathala G, Forne T, Piette J. CTCF is a DNA methylation-sensitive positive regulator of the INK/ARF locus. Biochem Biophys Res Commun. 2010;392:129–34.
Witcher M, Emerson BM. Epigenetic silencing of the p16(INK4a) tumor suppressor is associated with loss of CTCF binding and a chromatin boundary. Mol Cell. 2009;34:271–84.
Wallace JA, Felsenfeld G. We gather together: insulators and genome organization. Curr Opin Genet Dev. 2007;17:400–7.
Gaszner M, Felsenfeld G. Insulators: exploiting transcriptional and epigenetic mechanisms. Nat Rev Genet. 2006;7:703–13.
Phillips JE, Corces VG. CTCF: master weaver of the genome. Cell. 2009;137:1194–211.
Schalkwyk LC, Meaburn EL, Smith R, Dempster EL, Jeffries AR, Davies MN, et al. Allelic skewing of DNA methylation is widespread across the genome. Am J Hum Genet. 2010;86:196–212.
Shoemaker R, Deng J, Wang W, Zhang K. Allele-specific methylation is prevalent and is contributed by CpG-SNPs in the human genome. Genome Res. 2010;20:883–9.
Kim J, Kim JY, Song KS, Lee YH, Seo JS, Jelinek J, et al. Epigenetic changes in estrogen receptor beta gene in atherosclerotic cardiovascular tissues and in-vitro vascular senescence. Biochim Biophys Acta. 2007;1772:72–80.
Sharma P, Kumar J, Garg G, Kumar A, Patowary A, Karthikeyan G, et al. Detection of altered global DNA methylation in coronary artery disease patients. DNA Cell Biol. 2008;27:357–65.
Kotake Y, Nakagawa T, Kitagawa K, Suzuki S, Liu N, Kitagawa M, et al. Long noncoding RNA ANRIL is required for the PRC2 recruitment to and silencing of p15(INK4B) tumor suppressor gene. Oncogene. 2011;30:1956–62.
Yap KL, Li S, Munoz-Cabello AM, Raguz S, Zeng L, Mujtaba S, et al. Molecular interplay of the noncoding RNA ANRIL and methylated histone H3 lysine 27 by polycomb CBX7 in transcriptional silencing of INK4a. Mol Cell. 2010;38:662–74.
Vire E, Brenner C, Deplus R, Blanchon L, Fraga M, Didelot C, et al. The Polycomb group protein EZH2 directly controls DNA methylation. Nature. 2006;439:871–4.
Zhuang J, Peng W, Li H, Wang W, Wei Y, Li W, et al. Methylation of p15INK4b and expression of ANRIL on chromosome 9p21 are associated with coronary artery disease. PLoS One. 2012;7:e47193.
Krishnamurthy J, Torrice C, Ramsey MR, Kovalev GI, Al-Regaiey K, Su L, et al. Ink4a/Arf expression is a biomarker of aging. J Clin Invest. 2004;114:1299–307.
Motterle A, Pu X, Wood H, Xiao Q, Gor S, Liang Ng F, et al. Functional analyses of coronary artery disease associated variation on chromosome 9p21 in vascular smooth muscle cells. Hum Mol Genet. 2012;21:4021–9. This study found that p15ink4b and p16ink4a expression is markedly reduced in arterial vascular smooth muscle cells and associated with high cell proliferation in carriers of the 9p21.3 risk allele.
Kia SK, Gorski MM, Giannakopoulos S, Verrijzer CP. SWI/SNF mediates polycomb eviction and epigenetic reprogramming of the INK4b-ARF-INK4a locus. Mol Cell Biol. 2008;28:3457–64.
Ottaviani D, Lever E, Mao S, Christova R, Ogunkolade BW, Jones TA, et al. CTCF binds to sites in the major histocompatibility complex that are rapidly reconfigured in response to interferon-gamma. Nucleic Acids Res. 2012;40:5262–70.
Hu G, Schones DE, Cui K, Ybarra R, Northrup D, Tang Q, et al. Regulation of nucleosome landscape and transcription factor targeting at tissue-specific enhancers by BRG1. Genome Res. 2011;21:1650–8.
Huang M, Qian F, Hu Y, Ang C, Li Z, Wen Z. Chromatin-remodelling factor BRG1 selectively activates a subset of interferon-alpha-inducible genes. Nat Cell Biol. 2002;4:774–81.
Cuddapah S, Cui K, Zhao K. Transcriptional enhancer factor 1 (TEF-1/TEAD1) mediates activation of IFITM3 gene by BRG1. FEBS Lett. 2008;582:391–7.
Almontashiri NA, Fan M, Cheng BL, Chen HH, Roberts R, Stewart AFR. Interferon-gamma activates expression of p15 and p16 regardless of 9p21.3 coronary artery disease risk genotype. J Am Coll Cardiol. 2013;61:143–7. This study showed that interferon gamma does elevate p15ink4b and p16ink4a expression, but that this occurs primarily through a post-transcriptional mechanism that is independent of the 9p21.3 risk genotype.
Argmann CA, Van Den Diepstraten CH, Sawyez CG, Edwards JY, Hegele RA, Wolfe BM, et al. Transforming growth factor-beta1 inhibits macrophage cholesteryl ester accumulation induced by native and oxidized VLDL remnants. Arterioscler Thromb Vasc Biol. 2001;21:2011–8.
Han J, Hajjar DP, Tauras JM, Feng J, Gotto Jr AM, Nicholson AC. Transforming growth factor-beta1 (TGF-beta1) and TGF-beta2 decrease expression of CD36, the type B scavenger receptor, through mitogen-activated protein kinase phosphorylation of peroxisome proliferator-activated receptor-gamma. J Biol Chem. 2000;275:1241–6.
Hu YW, Wang Q, Ma X, Li XX, Liu XH, Xiao J, et al. TGF-beta1 up-regulates expression of ABCA1, ABCG1 and SR-BI through liver X receptor alpha signaling pathway in THP-1 macrophage-derived foam cells. J Atheroscler Thromb. 2010;17:493–502.
Tashiro H, Shimokawa H, Sadamatu K, Yamamoto K. Prognostic significance of plasma concentrations of transforming growth factor-beta in patients with coronary artery disease. Coron Artery Dis. 2002;13:139–43.
Lievens D, Habets KL, Robertson AK, Laouar Y, Winkels H, Rademakers T, et al. Abrogated transforming growth factor beta receptor II (TGFbetaRII) signalling in dendritic cells promotes immune reactivity of T cells resulting in enhanced atherosclerosis. Eur Heart J. 2013;34:3717–27.
Reifenberg K, Cheng F, Orning C, Crain J, Kupper I, Wiese E, et al. Overexpression of TGF-b1 in macrophages reduces and stabilizes atherosclerotic plaques in ApoE-deficient mice. PLoS One. 2012;7:e40990.
Diez-Juan A, Perez P, Aracil M, Sancho D, Bernad A, Sanchez-Madrid F, et al. Selective inactivation of p27(Kip1) in hematopoietic progenitor cells increases neointimal macrophage proliferation and accelerates atherosclerosis. Blood. 2004;103:158–61.
Fan K, Ruan Q, Sensenbrenner L, Chen B. Transforming growth factor-beta 1 bifunctionally regulates murine macrophage proliferation. Blood. 1992;79:1679–85.
Hannon GJ, Beach D. p15INK4B is a potential effector of TGF-beta-induced cell cycle arrest. Nature. 1994;371:257–61.
Reynisdottir I, Polyak K, Iavarone A, Massague J. Kip/Cip and Ink4 Cdk inhibitors cooperate to induce cell cycle arrest in response to TGF-beta. Genes Dev. 1995;9:1831–45.
Vijayachandra K, Higgins W, Lee J, Glick A. Induction of p16ink4a and p19ARF by TGFbeta1 contributes to growth arrest and senescence response in mouse keratinocytes. Mol Carcinog. 2009;48:181–6.
Brown KA, Pietenpol JA, Moses HL. A tale of two proteins: differential roles and regulation of SMAD2 and SMAD3 in TGF-beta signaling. J Cell Biochem. 2007;101:9–33.
MacLellan WR, Lee TC, Schwartz RJ, Schneider MD. Transforming growth factor-b response elements of the skeletal a-actin gene. Combinatorial action of serum response factor, YY1, and the SV40 enhancer-binding protein, TEF-1. J Biol Chem. 1994;269:16754–60.
Leask A, Holmes A, Black CM, Abraham DJ. Connective tissue growth factor gene regulation. Requirements for its induction by transforming growth factor-beta 2 in fibroblasts. J Biol Chem. 2003;278:13008–15.
Fujii M, Toyoda T, Nakanishi H, Yatabe Y, Sato A, Matsudaira Y, et al. TGF-beta synergizes with defects in the Hippo pathway to stimulate human malignant mesothelioma growth. J Exp Med. 2012;209:479–94.
Beyer TA, Weiss A, Khomchuk Y, Huang K, Ogunjimi AA, Varelas X, et al. Switch enhancers interpret TGF-beta and Hippo signaling to control cell fate in human embryonic stem cells. Cell Rep. 2013;5:1611–24.
Zheng Y, Devitt C, Liu J, Mei J, Skapek SX. A distant, cis-acting enhancer drives induction of Arf by Tgfbeta in the developing eye. Dev Biol. 2013;380:49–57. This article shows the requirement for the 9p21.3 syntenic region in mouse chromosome 4 for TGFβ-dependent activation of p14ARF from the CDKN2A gene.
Stewart AFR, Larkin SB, Farrance IKG, Mar JH, Hall DE, Ordahl CP. Muscle-enriched TEF-1 isoforms bind M-CAT elements from muscle-specific promoters and differentially activate transcription. J Biol Chem. 1994;269:3147–50.
Gray LT, Fong KK, Pavelitz T, Weiner AM. Tethering of the conserved piggyBac transposase fusion protein CSB-PGBD3 to chromosomal AP-1 proteins regulates expression of nearby genes in humans. PLoS Genet. 2012;8:e1002972.
Compliance with Ethics Guidelines
Conflict of Interest
Hsiao-Huei Chen, Naif A. M. Almontashiri, Darlène Antoine, and Alexandre F. R. Stewart declare that they have no conflict of interest.
Human and Animal Rights and Informed Consent
This article does not contain any studies with human or animal subjects performed by any of the authors.
Author information
Authors and Affiliations
Corresponding author
Additional information
This article is part of the Topical Collection on Cardiovascular Genomics
Rights and permissions
About this article
Cite this article
Chen, HH., Almontashiri, N.A.M., Antoine, D. et al. Functional Genomics of the 9p21.3 Locus for Atherosclerosis: Clarity or Confusion?. Curr Cardiol Rep 16, 502 (2014). https://doi.org/10.1007/s11886-014-0502-7
Published:
DOI: https://doi.org/10.1007/s11886-014-0502-7