Abstract
Rice crops are vulnerable to low temperatures. During development, the reproductive stage is particularly sensitive to cold exposure, which causes abnormal pollen development and a high degree of male sterility. In this study, shotgun proteomic analysis was used to analyze rice anthers containing pollen grains from a cold-tolerant variety, Dianxi 4. Protein expression was compared between normal anthers and anthers exposed to cold temperatures at the young microspore stage. In total, 3835 non-redundant proteins were identified in the rice anther. Of these, 441 proteins were differentially expressed between normal and cold-treated anthers. Pollen allergens, ATP synthase, actin, profilin, and β-expansin proteins were highly abundant, reflecting anther development, pollen germination, and pollen tube elongation. Starch and sucrose metabolic proteins such as α-amylase precursor and 4-α-glucanotransferase exhibited reduced expression after cold exposure. Among the proteins that exhibited increased expression after cold exposure, C2 domain proteins, and GRPs were identified as candidate signaling factors for mediation of the cold tolerance response. Through high-throughput proteomic analysis we were able to reveal proteomic changes against cold stress and suggest two signaling factors as the candidate genes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plants are affected by various environmental stresses during their life-cycles, such as low temperature, high temperature, high soil salt concentration, and drought. Exposure of plants to low temperatures induces physical and biochemical changes that have particularly serious effects. At the cellular level, low temperature exposure leads to several changes, including alterations to protein and carbohydrate composition and abundance of unsaturated fatty acids that result in physiological symptoms such as wilting, chlorosis, and leaf expansion (Schutzki and Cregg 2007). Low temperature stress experienced during the reproductive phase causes structural and functional abnormalities in reproductive organs that lead to fertilization failure. Rice is vulnerable to low temperatures, particularly during the reproductive stage (Nishiyama 1995). Cytological and histological abnormalities observed in cold-treated rice were more severe in the anthers than in the pistil or other organs of the flower, and fertility was rescued by artificial pollination with healthy pollen collected from non-stressed florets (Satake and Hayase 1970). Exposure to low temperature at the onset of microspore release caused tapetum hypertrophy and disordered microspore development that consequently gave rise to a high degree of male sterility (Hayase et al. 1969; Nishiyama 1970). The young microspore stage of pollen development, including meiosis, tetrad development, and the first contraction phase, was found to be particularly vulnerable to cold stress (Satake and Hayase 1970).
The molecular changes experienced by developed rice anthers after exposure to cold stress were previously examined using cDNA microarray and proteomic analysis. Microarray analysis of cold-stressed rice anthers in the early microspore stage showed that 160 ESTs were differentially expressed, 38 of which were upregulated and 122 of which were downregulated compared to control anthers. The molecular functions of the differentially expressed genes were related to primary metabolism, proteolysis, signal transduction defense, and secondary metabolism. Approximately 36% of the differentially expressed genes were of unknown function (Yamaguchi et al. 2004). Proteomic analysis of rice anther maturation after cold exposure showed that 70 protein spots were differentially expressed, including several ascorbate peroxidases that probably detoxified activated oxygen species by scavenging hydrogen peroxide generated under cold conditions (Imin et al. 2004). This hypothesis also suggested that programmed cell death and protein degradation might be involved in responses to cold stress during rice anther development.
Abnormal pollen development after cold exposure leads to a high degree of male sterility in rice. However, the very small size of individual pollen grains and technical difficulties in isolating pollen from anthers precludes direct proteomic analysis of pollen. Here, rice anthers containing pollen grains were used for proteomic analysis. Quantitative shotgun proteomic analysis was used to detect changes of protein expression patterns at the trinucleate stage where cold temperature treated during young microspore stage in a cold-tolerant rice variety.
Materials and methods
Plant materials and low temperature treatment
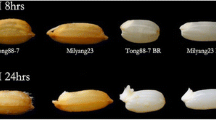
A Chinese low-temperature-tolerant variety of Oryza sativa, L., Dianxi 4, was used for proteomic analysis. To confirm the cold tolerance of Dianxi 4, spikelet fertility after low temperature treatment was compared between Dianxi 4 and Mihyang, a low-temperature-susceptible variety, as follows. Dianxi 4 and Mihyang seed were germinated and grown under natural conditions in 15 × 30 cm rows at the Konkuk University experimental field. Individual plants with auricle distances (the distance between the ligule of the flag leaf and the penultimate leaf) of approximately 10 mm were selected and transferred, along with field soil, to a 15 cm diameter pot. Six rice plants were grown in a growth chamber and treated at 12 °C, with 70% relative humidity, under a 12 h day/12 h night cycle for 4 days. Plants were then transferred to a greenhouse and cultivated until rice grains were fully matured. Control plants were collected and transferred to pots at the same stage as test plants, but remained under natural conditions in the field for 4 days prior to transfer to the greenhouse. Spikelet fertility was assessed using the fully matured spikelet of three replicate rice plants. For Dianxi 4 proteomic analysis, anthers from control and low-temperature-treated plants were harvested from the spikelets of the top three primary branches 1 day prior to anthesis.
Rice anther protein extraction
Anthers were placed into 1.5-mL tubes with 2–3 small metal balls and ground with a vortexer to powder in liquid nitrogen. Proteins were extracted by incubating the resultant powder with extraction buffer as 8 M urea/5 mM DTT/1% lithium dodecyl sulfate/100 mM Tris pH 8.5 at room temperature for 30 min. The mixture was then centrifuged at 14,000 g for 15 min, after which the supernatant was retained and filtered through a 0.45 µm membrane filter (Millipore, Billerica, MA, USA). The extracted proteins were precipitated overnight with 20% (v/v) trichloroacetic acid (TCA) and then washed several times with cold acetone until pigments were removed. The extracted proteins were resuspended in resolubilization buffer as 8 M urea/Tris–HCl pH 8.5. Protein concentration was assayed using a 2D-Protein Quant Kit (GE Healthcare, Piscataway, NJ, USA) as described previously (Lee et al. 2007).
1D LDS-PAGE and in-gel digestion
Protein samples (50 µg) were prepared using NuPage® LDS Sample Buffer according to the manufacturer’s instructions (Invitrogen, Carlsbad, CA, USA) and loaded onto 4–12% Bis–Tris acrylamide gradient gels (Invitrogen, Carlsbad, CA, USA). After electrophoresis, gels were fixed in fixing solution (40% deionized water/50% methanol/10% acetic acid) for 10 min with shaking, after which the fixing solution was drained and the gel stained with colloidal Coomassie blue (NOVEX® Colloidal Blue Stain Kit: Invitrogen, Carlsbad, CA, USA). Stained protein lanes were sliced into seven pieces of equal size. Each piece was diced further (approximate size 1 mm2) and transferred to an Eppendorf tube. Destaining buffer [50% acetonitrile (ACN) in 50 mM ammonium bicarbonate (ABC) pH 7.8] was added to remove Coomassie blue and samples were dehydrated in a SpeedVac. Dehydrated samples were reduced in reduction buffer (10 mM DTT in 25 mM NH4HCO3) at 56 °C for 45 min, then alkylated for 30 min with alkylation buffer (55 mM iodoacetamide in 25 mM NH4HCO3) at room temperature in the dark. Reduced and alkylated samples were digested with trypsin as trypsin 12.5 ng/µL in 50 mM NH4HCO3 overnight at 36 °C. After digestion, tryptic peptides were recovered from the gel by incubation with harvest buffer as 5% formic acid in 50% acetonitrile for 20 min at room temperatures. Supernatants were dried using a SpeedVac, desalted using Pierce® C18 spin columns (Thermo Scientific, Rockford, IL, USA), and finally resuspended in 5% ACN/0.1% formic acid.
LC MS/MS analysis
A nanoflow HPLC instrument (Easy nLC, Thermo Fisher Scientific, San Jose, CA, USA) was coupled on-line to a Q-Exactive mass spectrometer (Thermo Fisher Scientific, Bremen, Germany). Columns for analysis (12 cm, 75 µm inner diameter) were packed in-house with Alltima C18-AQ 5 µm resin. Samples were separated using a binary buffer system (Buffer A: 0.1% formic acid; Buffer B: acetonitrile in 0.1% formic acid) with a linear gradient of 3–50% buffer B at a flow rate of 250 nL/min. The total run time for LC MS/MS was 110 min. MS data were acquired using the data-dependent top 8 method, dynamically choosing the most abundant precursor ions from the survey scan (300–2000 Da) for higher-energy collisional dissociation (HCD) fragmentation. The dynamic exclusion duration was 60 s, and the isolation window for precursors was set to 4 m/z. Survey scans were acquired at a resolution of 70,000 at m/z 200 and the resolution for HCD spectra was set to 17,500 at m/z 200.
Analysis of proteomic data
Spectral fragment data from the Q-Exactive analysis were searched against the TIGR Rice Pseudomolecule protein database RELEASE v7.0 (http://rice.plantbiology.msu.edu/annotation_pseudo_current.shtml) using Proteome Discoverer (version 1.3) software (Thermo Fisher Scientific), which provided the pI value and molecular weight of each identified protein. Carbamidomethylation of cysteine was used as a fixed modification and oxidation of methionine was set as a variable modification for data searching. Both identifications were filtered using a 1% false discovery rate.
Comparative analysis of relative protein abundance
Output data from Proteome Discoverer (version 1.3) were exported to Microsoft Excel and normalized spectral counts (NSpC) were calculated (Paoletti et al. 2006; Zybailov et al. 2006; Griffin et al. 2010). The NSpC for each protein k is given by:
where the total number of MS/MS spectra matching peptides from protein k (SpC) is divided by the protein length (L), then divided by SpC/L for all N proteins in the experiment.
Bioinformatic analysis of proteomic data
Predicted protein molecular weights and pI values were calculated from the rice genome at EMBOSS Pepstats, EMBL-EBI (http://www.ebi.ac.uk/Tools/seqstats/emboss_pepstats/) and the EMBOSS Pepstats algorithm (http://emboss.bioinformatics.nl/cgi-bin/emboss/pepstats). Gene ontologies (GO) were retrieved from the TIGR Rice Pseudomolecule protein database Release V7.0. and GO enrichment analysis was performed in agriGO (Du et al. 2010). Differential protein expression was analyzed using Mapman (Thimm et al. 2004).
Results and discussion
Cold tolerance in Dianxi 4
The cold-susceptible rice variety Mihyang was used as a control to verify the cold tolerance of Dianxi 4 and the effectiveness of cold treatments. Spikelet fertility under control conditions was 88.39 ± 2.5% for Dianxi 4 and 94.2 ± 2.5% for Mihyang. After exposure to low temperature, spikelet fertility decreased to 52.57 ± 7.9% for Dianxi 4 and 1.5 ± 0.86% for Mihyang (Fig. 1), indicating that the low temperature treatment was effective. Although Dianxi 4 exhibited lower spikelet fertility after cold treatment than under normal conditions, by comparison with Mihyang, Dianxi 4 was relatively cold-tolerant. Therefore, comparative proteomic analysis between control and cold-treated Dianxi 4 anthers would effectively capture the protein expression response to cold temperature exposure at the early young microspore stage and the effects on anther maturation.
Identification of rice anther proteins
Mature anthers at the trinucleate stage were harvested from cold-treated and control Dianxi 4 rice plants. Proteomes were identified using high-throughput shotgun proteomic analysis at a 0.01 false discovery rate. Three biological replicates were analyzed per treatment. Approximately 2000 proteins were identified per sample, and 3853 non-redundant proteins were identified in total from the six samples (Supplementary file S1). The analytical incompleteness phenomenon, in which single analytical runs may identify only a fraction of the associated peptides in a highly complex mixture, may explain why all of the proteins were not identified in every sample (Wilkins et al. 2006).
The physiochemical properties of the 3853 proteins were compared with those of the whole predicted rice proteome at TIGR (http://rice.plantbiology.msu.edu/annotation_pseudo_current.shtml). Protein molecular weights spanned a wide range, from 5.28 kDa (LOC_Os04g02670.1: Expressed protein) to 486.00 kDa (LOC_Os09g07300.1: BIG, putative, expressed) (Fig. 2a). Although the proportions of identified proteins < 20 and > 120 kDa were lower than in the predicated rice proteome, the overall molecular weight distribution was similar. The molecular weight range and distribution showed that the shotgun proteomic analysis method used did not exhibit bias in identifying rice anther proteins. The pI values of the identified proteins also covered a wide range, from pH 3.93 (LOC_Os07g41694.1: acidic leucine-rich nuclear phosphoprotein 32-related protein 1, putative, expressed) to pH 12.48 (LOC_Os01g69020.1: retrotransposon protein, putative, unclassified, expressed) (Fig. 2b). The proportion of proteins with pI values of pH 5–8 was higher for the anther proteins than for the whole predicted rice proteome.
Relative abundance of identified proteins in each sample was estimated using a normalized spectral count method. Several pollen allergens were highly abundant in the matured anther, as well as ATP synthase, UTP-glucose-1-phosphate uridylyltransferase, glyceraldehyde-3-phosphate dehydrogenase, and cupin domain containing proteins (Supplementary table S2, Table 1). The presence of these proteins at high levels was indicative of preparation for pollen tube germination and elongation. Pollen allergens, which were the most abundant proteins, were previously shown to be involved in pollen germination and growth (Songnuan 2013). High levels of carbohydrate metabolism related proteins such as ATP synthase and glyceraldehyde-3-phosphate dehydrogenase are needed for energy supply during tube germination and elongation (Dai et al. 2007). Actin and profilin proteins, as well as expansin precursors, were also highly abundant in mature anthers. Actin is a major component of microfilaments, and is thus important for cell structure, expansion and development, and profilin is a small actin monomer-binding protein (Kovar et al. 2000; McCurdy et al. 2001). β-expansin is involved in loosening the cell wall, which promotes pollen development and pollen tube growth (Kerim et al. 2003). Cupin domain proteins are involved in seed germination and development (Lapik and Kaufman 2003) and have no known role in pollen development. Nevertheless, cupin proteins were highly abundant in mature anthers and might therefore play a hitherto unsuspected role in pollen germination.
Protein expression changes in mature anthers after cold exposure at the early young microspore stage
Relative expression analysis was used to assess the differences in protein expression between control anthers and those exposed to cold stress at the early young microspore stage. Although a total of 3853 proteins were identified from the six anther samples, not all proteins were identified in all samples. Only proteins that were detected in all three biological replicates of either control or treated anthers, with at least two spectral counts per replicate, were used for the comparative analysis: this resulted in 2359 proteins. Of these, 441 proteins exhibited at least twofold differential expression between cold-treated and control anthers, as assessed by t test (α = 0.05) (Supplementary table S3). Of these, 371 proteins were overexpressed and 70 proteins were underexpressed in cold-treated anthers compared to the control. GO category enrichment analysis of the 441 proteins revealed that 31 GO categories were enriched, comprising three biological process, two molecular function, and 26 cellular component GO terms (Table 2). The enriched biological process categories were establishment of localization, localization, and transport. For molecular function, catalytic activity and hydrolase activity were enriched. The 26 enriched cellular component terms encompassed the majority of the proteomic changes that occurred in response to low temperature exposure in the cold-tolerant Dianxi 4 variety. Although spikelet fertility in Dianxi 4 decreased to 52% after exposure to cold treatment at the early microspore stage, fertility remained significantly higher than in the cold-susceptible variety Milyang. Maintenance of cellular functions during pollen development after meiosis is fundamental for proper development. Several of these proteins were upregulated in Dianxi 4 anthers after cold treatment, which may have partially compensated for the effects of cold exposure. We therefore hypothesized that the differentially expressed proteins associated with GO cellular component categories were responsible for cold tolerance in Dianxi 4.
Analysis of differential protein expression was performed using Mapman analysis (Fig. 3). Expression of the majority of metabolic pathway proteins, particularly those involved in cell division and the cell cycle, increased after cold exposure. Other proteins upregulated in response to cold stress included proteins responsive to abiotic drought or salt stress (LOC_Os04g57890: oligosaccharyl transferase, putative, expressed; and LOC_Os07g05570: ERD4 protein, putative, expressed), miscellaneous proteins (LOC_Os02g13330: bet v I allergen family protein, putative, expressed; and LOC_Os06g02420: ATOZI1, putative, expressed), heat stress proteins (LOC_Os12g27070: heat shock protein DnaJ, putative, expressed; LOC_Os05g26902: chaperone protein DnaJ, putative, expressed; and LOC_Os06g11610: heat shock 22 kDa protein, mitochondrial precursor, putative, expressed), and cold stress proteins (LOC_Os07g39620: C2 domain containing protein, putative, expressed; and LOC_Os02g02870: glycine-rich protein 2, putative, expressed). Expression of proteins involved in redox reactions also increased after cold exposure.
Differential protein expression between cold stress and normal conditions in a cold-tolerant rice variety
Consideration of prior known functions for proteins upregulated after cold exposure allowed identification of candidate cold-tolerance genes (Table 3). Expression of glycine-rich protein 2 increased 6.86-fold after cold exposure. The glycine-rich protein (GRP) family has a range of functions in plants. All GRPs contain semi-repetitive glycine-rich motifs, and plant GRPs are classified based on the presence of conserved motifs and motif arrangements (Sachetto-Martins et al. 2000; Fusaro et al. 2001; Bocca et al. 2005). GRPs are structurally diverse, exhibit variable expression and modulation patterns, and are found in a range of subcellular locations, all of which contribute to their diverse functions in plants (Mangeon et al. 2010). GRPs are involved in flower development (Flores Fusaro and Sachetto-Martins 2007) and their expression increases under conditions of abiotic or biotic stress or after abscisic acid (ABA) treatment (Long et al. 2013; Sachetto-Martins et al. 2000; Mangeon et al. 2010). In Arabidopsis, overexpression of AtGRP2, a member of the cold-shock domain protein family, enhanced tolerance to freezing (Kim et al. 2005).
Expression of several heat shock proteins and chaperone proteins such as DnaJ, chaperone proteins, heat shock 22 kDa proteins and T-complex proteins increased after cold exposure. Similar increases in expression of these proteins were reported in rice anther after exposure to high temperatures (Kim et al. 2015). This suggested that high and low unfavorable temperatures elicited similar responses in rice anthers. Heat tolerance responses in plants are mediated by increased expression of heat shock proteins and chaperone proteins. These proteins also cooperate with other cellular components to decrease cellular damage in a range of conditions (Wang et al. 2004).
Expression of a C2-domain-containing protein, LOC_Os07g39620.1, which was previously shown to contain a Ca2+-dependent membrane-binding motif, increased 33-fold after cold exposure. Calcium ions play important roles in defense signaling in plant cells (Nalefski and Falke 1996), and C2-domain-containing proteins, which are targeted to membranes, are associated with tolerance to abiotic and biotic stress (Kim et al. 2003; Yokotani et al. 2009). Overexpression of a single C2 domain protein in Arabidopsis enhanced tolerance to abiotic stresses including high salinity, osmotic stress, dehydration, and oxidative stress, as well as biotic stress caused by Pseudomonas syringae.
Expression of putative peroxidase precursor proteins LOC_Os03g02920.1, LOC_Os11g10460.1, LOC_Os02g14180.1, and LOC_Os01g16450.1 increased 27.68-, 22.73-, 15.65-, and 2.56-fold, respectively, after cold exposure. These proteins enhance resistance to abiotic and biotic by catalyzing reactions with reactive oxygen species (ROS) such as H2O2. Salicylic acid and nitric oxide produced by these processes are active in plant defenses. Peroxidase proteins participate in signaling and release of ROS in response to various stresses (Mittler 2002). Together, these results indicate that several signaling systems are active under cold stress conditions (Dat et al. 2000; Zhu 2016). Peroxidases are also involved in stress-induced phenylpropanoid metabolism (Dixon and Paiva 1995). Phenylpropanoid metabolism increased under cold stress conditions and abundance of the resultant phenolic compounds increased (Leyva et al. 1995). Phenolic compounds were oxidized by peroxidase, and these enzyme activities contributed to enhanced stress resistance (Rivero et al. 2001).
Several proteins exhibited reduced expression levels after cold exposure. These included proteins associated with starch metabolism. Expression levels of two α-amylase precursors, LOC_Os01g51754.1 and LOC_Os01g51754.3, were 0.48- and 0.004-fold lower, respectively, in cold-exposed anthers than in control anthers. The role of α-amylase is to produce smaller oligosaccharides from complex carbohydrates such as starch and cellulose. Expression of another starch metabolism protein, 4-α-glucanotransferase (LOC_Os07g43390), decreased 0.49-fold after cold exposure. The role of 4-α-glucanotransferase in carbohydrate metabolism is to transfer a segment of 1,4-α-d-glucan to other carbohydrates such as glucose or 1,4-α-d-glucan (Takaha et al. 1993).
During rice anther development, large amounts of sugars are mobilized to the anthers to support early development (Castro and Clément 2007). Physiological damage caused by exposure to low temperatures can block sugar mobilization to the young microspore, culminating in abortion of pollen development and sterility (Oliver et al. 2007). ABA was previously shown to inhibit α-amylase synthesis and growth (Khan and Downing 1968), and cold stress resulted in an increase in ABA synthesis (Chinnusamy et al. 2007). Although protein expression changes were examined in a cold-tolerant rice variety, Dianxi 4, in this study, the detected proteomic changes nevertheless reflect the impacts of cold stress. The observed decrease in α-amylase was consistent with ABA responses to cold stress. (Oliver et al. 2007).
Conclusions
Rice variety Dianxi 4 exhibited cold tolerance at the early young microspore stage. High-throughput shotgun proteomic analysis identified 3835 non-redundant proteins in matured rice anther (trinucleate stage) with a wide range of molecular weight and pI values. Pollen allergens, ATP synthase, actin, profilin, and β-expansin proteins were highly abundant in mature anthers, reflecting anther maturation, pollen germination, and pollen tube elongation functions. The proteomes of cold-exposed and control anthers were compared and, in total, 441 differentially expressed proteins were detected. GO enrichment and Mapman analysis of the differentially expressed proteins suggested that the cold tolerance of Dianxi 4 resulted from increased levels of proteins associated with cellular components that limited damage from cold stress. Cold exposure was associated with reduced levels of starch metabolism proteins. Among the proteins that exhibited increased expression after cold exposure, C2 domain proteins and GRPs were identified as candidate signaling factors for mediation of the cold tolerance response. Protein expression changes may not directly contribute to cold tolerance. Nevertheless, understanding protein expression patterns in cold-tolerant rice varieties can contribute to our understanding of tolerance responses and can highlight candidate genes for further study.
References
Bocca SN, Magioli CS, Mangeon A, Margis R, Junqueira RM, Jorge VC, Martins GS (2005) Survey of glycine-rich proteins (GRPs) in the Eucalyptus expressed sequence tag database (ForEST). Genet Mol Biol 28:608–624
Castro AJ, Clément C (2007) Sucrose and starch catabolism in the anther of Lilium during its development: a comparative study among the anther wall, locular fluid and microspore/pollen fractions. Planta 225:1573–1582
Chinnusamy V, Zhu J, Zhu JK (2007) Cold stress regulation of gene expression in plants. Trends Plant Sci 12:444–451
Dai S, Chen T, Chong K, Xue Y, Liu S, Wang T (2007) Proteomics identification of differentially expressed proteins associated with pollen germination and tube growth reveals characteristics of germinated oryza sativa pollen. Mol Cell Proteom 6:207–230
Dat J, Vandenabeele S, Vranová E, Van Montagu M, Inzé D, Van Breusegem F (2000) Dual action of the active oxygen species during plant stress responses. Cell Mol Life Sci 57:779–795
Dixon RA, Paiva NL (1995) Stress-induced phenylpropanoid metabolism. Plant Cell 7:1085–1097
Du Z, Zhou X, Ling Y, Zhang Z, Su Z (2010) agriGO: a GO analysis toolkit for the agricultural community. Nucl Acids Res 38:W64–W70
Flores Fusaro A, Sachetto-Martins G (2007) Blooming time for plant glycine-rich proteins. Plant Signal Behav 2:386–387
Fusaro A, Mangeon A, Junqueira RM, Rocha CAB, Coutinho TC, Margis R, Sachetto-Martins G (2001) Classification, expression pattern and comparative analysis of sugarcane expressed sequences tags (ESTs) encoding glycine-rich proteins (GRPs). Genet Mol Biol 24:263–273
Griffin NM, Yu J, Long F, Oh P, Shore S, Li Y, Koziol AJ, Schnitzer JE (2010) Label-free, normalized quantification of complex mass spectrometry data for proteomic analysis. Nat Biotech 28:83–89
Hayase H, Satake T, Nishiyama I, ITO N (1969) Male sterility caused by cooling treatment at the meiotic stage in rice plants: II. The most sensitive stage to cooling and the fertilizing ability of pistils. Jpn J Crop Sci 38:706–711
Imin N, Kerim T, Rolfe BG, Weinman JJ (2004) Effect of early cold stress on the maturation of rice anthers. Proteomics 4:1873–1882
Kerim T, Imin N, Weinman JJ, Rolfe BG (2003) Proteome analysis of male gametophyte development in rice anthers. Proteomics 3:738–751
Khan AA, Downing RD (1968) Cytokinin reversal of abscisic acid inhibition of growth and α-amylase synthesis in barley seed. Physiol Plant 21:1301–1307
Kim CY, Koo YD, Jin JB, Moon BC, Kang CH, Kim ST, Park BO, Lee SY, Kim ML, Hwang I, Kang KY, Bahk JD, Lee SY, Cho MJ (2003) Rice C2-domain proteins are induced and translocated to the plasma membrane in response to a fungal elicitor. Biochemistry 42:11625–11633
Kim YO, Kim JS, Kang H (2005) Cold-inducible zinc finger-containing glycine-rich RNA-binding protein contributes to the enhancement of freezing tolerance in Arabidopsis thaliana. Plant J 42:890–900
Kim M, Kim H, Lee W, Lee Y, Kwon SW, Lee J (2015) Quantitative shotgun proteomics analysis of rice anther proteins after exposure to high temperature. Int J Genomics 2015:9
Kovar DR, Drøbak BK, Staiger CJ (2000) Maize profilin isoforms are functionally distinct. Plant Cell 12:583–598
Lapik YR, Kaufman LS (2003) The arabidopsis cupin domain protein AtPirin1 Interacts with the G protein α-subunit GPA1 and regulates seed germination and early seedling development. Plant Cell 15:1578–1590
Lee J, Garrett WM, Cooper B (2007) Shotgun proteomic analysis of Arabidopsis thaliana leaves. J Sep Sci 30:2225–2230
Leyva A, Jarillo JA, Salinas J, Martinez-Zapate JM (1995) Low Temperature induces the accumulation of phenylalanine ammonia-lyase and chalcone synthase mRNAs of Arabidopsis thaliana in a light-dependent manner. Plant Physiol 108:39–46
Long R, Yang Q, Kang J, Zhang T, Wang H, Li M, Zhang Z (2013) Overexpression of a novel salt stress-induced glycine-rich protein gene from alfalfa causes salt and ABA sensitivity in Arabidopsis. Plant Cell Rep 32:1289–1298
Mangeon A, Junqueira RM, Sachetto-Martins G (2010) Functional diversity of the plant glycine-rich proteins superfamily. Plant Signal Behav 5:99–104
McCurdy DW, Kovar DR, Staiger CJ (2001) Actin and actin-binding proteins in higher plants. Protoplasma 215:89–104
Mittler R (2002) Oxidative stress, antioxidants and stress tolerance. Trends Plant Sci 7:405–410
Nalefski EA, Falke JJ (1996) The C2 domain calcium-binding motif: structural and functional diversity. Protein Sci 5:2375–2390
Nishiyama I (1970) Male sterility caused by cooling treatment at the young microspore stage in rice plants: VII. Electron microscopical observations on tapetal cells dilated by the cooling treatment. Jpn J Crop Sci 39:480–486
Nishiyama I (1995) Damage due to extreme temperatures. Sci Rice Plant 2:769–812
Oliver SN, Dennis ES, Dolferus R (2007) ABA regulates apoplastic sugar transport and is a potential signal for cold-induced pollen sterility in rice. Plant Cell Physiol 48:1319–1330
Paoletti AC, Parmely TJ, Tomomori-Sato C, Sato S, Zhu D, Conaway RC, Conaway JW, Florens L, Washburn MP (2006) Quantitative proteomic analysis of distinct mammalian Mediator complexes using normalized spectral abundance factors. Proc Natl Acad Sci 103:18928–18933
Rivero RM, Ruiz JM, Garcıa PC, Lopez-Lefebre LR, Sánchez E, Romero L (2001) Resistance to cold and heat stress: accumulation of phenolic compounds in tomato and watermelon plants. Plant Sci 160:315–321
Sachetto-Martins G, Franco LO, de Oliveira DE (2000) Plant glycine-rich proteins: a family or just proteins with a common motif? Biochimica et Biophysica Acta (BBA). Gene Struct Expr 1492:1–14
Satake T, Hayase H (1970) Male sterility caused by cooling treatment at the young micro-spore stage in rice plants. 5. Estimations of pollen developmental stage and the most sensitive stage to coolness. Nihon Sakumotsugaku Kai Kiji. Proc Crop Sci Soc Jpn 39:468–473
Schutzki RE, Cregg B (2007) Abiotic plant disorders: symptoms, signs and solutions: a diagnostic guide to problem solving. Michigan State University
Songnuan W (2013) Wind-pollination and the roles of pollen allergenic proteins. Asian Pac J Allergy Immunol 31:261
Takaha T, Yanase M, Okada S, Smith SM (1993) Disproportionating enzyme (4-alpha-glucanotransferase; EC 2.4.1.25) of potato. Purification, molecular cloning, and potential role in starch metabolism. J Biol Chem 268:1391–1396
Thimm O, Bläsing O, Gibon Y, Nagel A, Meyer S, Krüger P, Selbig J, Müller LA, Rhee SY, Stitt M (2004) mapman: a user-driven tool to display genomics data sets onto diagrams of metabolic pathways and other biological processes. Plant J 37:914–939
Wang W, Vinocur B, Shoseyov O, Altman A (2004) Role of plant heat-shock proteins and molecular chaperones in the abiotic stress response. Trends Plant Sci 9:244–252
Wilkins MR, Appel RD, Van Eyk JE, Chung M, Görg A, Hecker M, Huber LA, Langen H, Link AJ, Paik YK, Patterson SD, Pennington SR, Rabilloud T, Simpson RJ, Weiss W, Dunn MJ (2006) Guidelines for the next 10 years of proteomics. Proteomics 6:4–8
Yamaguchi T, Nakayama K, Hayashi T, Yazaki J, Kishimoto N, Kikuchi S, Koike S (2004) cDNA microarray analysis of rice anther genes under chilling stress at the microsporogenesis stage revealed two genes with DNA transposon castaway in the 5′-flanking region. Biosci Biotechnol Biochem 68:1315–1323
Yokotani N, Ichikawa T, Kondou Y, Maeda S, Iwabuchi M, Mori M, Hirochika H, Matsui M, Oda K (2009) Overexpression of a rice gene encoding a small C2 domain protein OsSMCP1 increases tolerance to abiotic and biotic stresses in transgenic Arabidopsis. Plant Mol Biol 71:391
Zhu J-K (2016) Abiotic stress signaling and responses in plants. Cell 167:313–324
Zybailov B, Mosley AL, Sardiu ME, Coleman MK, Florens L, Washburn MP (2006) Statistical analysis of membrane proteome expression changes in saccharomyces cerevisiae. J Proteom Res 5:2339–2347
Acknowledgements
This work was supported by the National Institute of Crop Science of the Rural Development Administration, Korea [Grant number PJ01099903].
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Lee, J., Lee, Y., Kim, M. et al. Quantitative shotgun proteomic analysis of cold-stressed mature rice anthers. Plant Biotechnol Rep 11, 417–427 (2017). https://doi.org/10.1007/s11816-017-0459-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11816-017-0459-2