Abstract
Rice (Oryza sativa) is sensitive to salt stresses and cannot survive under low salt conditions, such as 50 mM NaCl. In an attempt to improve salt tolerance of rice, we introduced katE, a catalase gene of Escherichia coli, into japonica rice cultivar, Nipponbare. The resultant transgenic rice plants constitutively expressing katE were able to grow for more than 14 days in the presence of 250 mM NaCl, and were able to form flower and produce seeds in the presence of 100 mM NaCl. Catalase activity in the transgenic rice plants was 1.5- to 2.5-fold higher than non-transgenic rice plants. Our results clearly indicate that simple genetic modification of rice to express E. coli-derived catalase can efficiently increase its tolerance against salt stresses. The transformant presented here is one of the most salt-tolerant rice plants created by molecular breeding so far.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plants are exposed to many types of environmental stresses such as salt, drought, and freezing. Among these stresses, salinity is one of the major factors that limit crop production world-wide (Epstein et al. 1980; Yancey et al. 1982; Zhu 1999). Salt tolerance of plants is a complex trait that is controlled by a number of salt stress-responsive genes (Zhu 2000; Wang et al. 2003, 2004; Ueda et al. 2002). The detrimental effects of salt on plants are consequences of both water deficit and interference with biochemical processes, which are brought about by high osmoticum and excess sodium ion, respectively. Organisms that thrive in hyper-saline environments possess various mechanisms to avoid these effects, e.g., osmotic adjustment, exclusion of sodium ion, and sequestration of sodium ion in the vacuole. The ability to accumulate organic compatible solutes is essential for osmoregulation in plants under high salinity (Bohnert et al. 1995; Hasegawa et al. 2000; Zhu 2002). Plants also use inorganic ions to adjust osmotic balance and to distribute these ions to keep sodium ion away from the cytoplasm (Serrano 1996; Blumwald et al. 2000). Many stress-responsible genes, which are controlled by specific transcription factors, function in increasing salt tolerance in plants (Nakashima et al. 2000; Narusaka et al. 2003).
Sodium ion influx into plant cells results in oxidative stress. Sodium ion suppresses CO2 fixation, which causes the over-reduction of the photosynthetic electron transport chain and the production of reactive oxygen species (ROS), such as singlet oxygen (1O2*), super oxide anion (O −2 ), hydrogen peroxide (H2O2), and hydroxyl radical (HO•).
In photosynthetic organisms, H2O2 is derived from proton and O −2 generated by Mehler reaction. H2O2 is most stable among ROS. However, when present together with Fe2+, H2O2 produces HO•, the most harmful ROS that causes serious damage to plant cells. Therefore, the decomposition of H2O2 is pivotal for protecting the cells from the oxidative damage.
Catalase is one of the major endogenous antioxidants, and catalyzes H2O2 decomposition. Like the other ROS scavenging enzymes, catalase is up-regulated at the transcriptional level upon exposure to high concentrations of NaCl (Wang et al. 2004). In cyanobacteria, introduction of a catalase gene of Escherichia coli, katE, was found to reduce ROS production under salt stresses and confer salt tolerance. The result indicates that the quenching of H2O2 was an important factor for salt tolerance (Kaku et al. 2000).
Here, we report the phenotypes of transgenic japonica rice plants into which the katE gene had been introduced. The results indicate the potential of katE in improving salt tolerance of rice by genetic modification.
Materials and methods
Plant material and construction of plasmid vector pIG121/Hm/katE
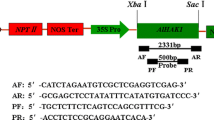
Oryza sativa japonica cultivar Nipponbare was used throughout the present study. The katE DNA was prepared from pAMkatE72 (Ossowski et al. 1991) by partial digestion with XbaI and SacI. The resulting katE-containing fragment was ligated into the XbaI and SacI double-digested site of a binary vector, pIG121/Hm (Akama et al. 1992) to construct pIG121-Hm/katE (Fig. 1). The pIG121-Hm/katE plasmid was introduced into Agrobacterium tumefaciens strain EHA101 by electroporation (Hood et al. 1986).
Plant transformation and culture condition
Rice transformation was conducted based on the methods appeared in the publications (Hiei et al. 1994; Toki 1997). Callus was induced from rice seeds on N6 medium (Chu et al. 1975) supplemented with 2 mg/l 2,4-D. Two weeks after callus induction, the callus was infected with A. tumefaciens EHA101 carrying pIG121/Hm/katE. After 3 days of co-cultivation, the callus was sterilized with 500 mg/l carbenicillin and cultured on N6 selection medium containing 50 mg/l hygromycin and 100 mg/l carbenicillin for 2 weeks. After the selection of transformed cells, shoot regeneration was induced by culture on MS medium (Murashige and Skoog 1962) supplemented with 1 mg/l NAA, 2 mg/l kinetin, 50 mg/l hygromycin, and 100 mg/l carbenicillin. Regenerated shoots were transferred to hormone-free MS medium for the induction of root formation. Regenerants were then acclimatized in pots with soil and submerged in water. Progeny was obtained from those transgenic plants by self-mating.
PCR analysis
Genomic DNA was extracted from young leaf tissues of T0 transgenic plants, T1 transgenic plants and non-transformed control plants. PCR was performed in a reaction mixture containing 25 ng genomic DNA, 100 μM dNTP mix, 0.2 μM forward and reverse primers, and 1 unit of Taq polymerase (TaKaRa). The PCR temperature profile consisted of 30 cycles of 94°C for 30 s (denaturation), 50°C for 40 s (annealing), and 72°C for 50 s (extension). The sequences of the PCR primers were as follows: katE-U, 5′-ATGTCGCAACAAACGA-3′; katE-L, 5′-TCAGGCAGGAATTTG-3′.
Southern blot analysis
Total genomic DNA extracted from T1 transgenic plant and non-transformed control plant was digested with KpnI and EcoRI and electrophoresed on 0.8% agarose gel. Separated DNA fragments were transferred to a nylon membrane (Hybond-N+, Amersham), and subjected to hybridization with 32P-labeled katE as a probe according to the standard method of Southern blotting. Hybridization signals were detected by autoradiography.
RNA isolation and RT-PCR analysis
Total RNA was extracted from T1 transgenic plants and non-transformed control plants by the SDS-phenol method (Ozeki et al. 1990) and first-strand cDNA was synthesized using reverse transcriptase (Superscript II; GIBCO BRL) and oligo-dT primer. PCR was carried out with the katE-U and katE-L primers as described above.
Enzyme extraction and measurement of catalase activity
Enzyme extraction and measurement of catalase were conducted according to the published protocols (Ngo and Lenhoff 1980; McClung 1997; Aebi 1984). Leaf tissues of T1 transgenic plants and non-transformed control plants were ground in liquid nitrogen into a fine powder with a pestle and mortar, and homogenized in 50 mM sodium phosphate buffer (pH 7.0) containing 0.1% (w/v) sodium metabisulphite and 0.1% (w/v) Triton X-100.
Polyvinylpolypyrrolidone (1/50 weight) was then added to the homogenate, which was filtered through one layer muslin and centrifuged at 16,000 g for 20 min at 4°C. Glycerol was added to the supernatant to give a final concentration of 10% (v/v), and the mixture was frozen in liquid nitrogen and stored at −80°C until use. The total protein content in enzyme extracts was determined by the Lawry method.
The activity of catalase in the extract was assayed in 50 mM sodium phosphate buffer (pH 7.3) containing 5 mM H2O2. The decrease in absorbance was monitored at 240 nm at 25°C for 1 min using a spectrophotometer (Shimadzu, UV160A).
Evaluation of salt tolerance
Salt tolerance was evaluated both at the vegetative stage and at the reproductive stage. In the vegetative stage, T0 transgenic rice plants, T1 transgenic rice plants and wild-type rice plants were soaked in 0, 50, 100, 150, 200, 250, 300, 400, 500, or 600 mM (sea water level) NaCl solution. Growth was monitored every 5 days by measuring the length of leaves and the area of surviving green part in leaves. NaCl solution was replaced every 5 days. For each condition, five plants were used.
For salt tolerance evaluation at the reproductive stage, flowering T1 transgenic rice plants and wild-type rice plants grown under normal growth condition were soaked in 250 mM NaCl solution for 14 days and scored for the number of seeds produced.
Results and discussion
In the present study, we transformed rice, Oryza sativa japonica cultivar Nipponbare, with the katE gene that had been derived from E. coli and designed to be expressed under the control of the 35S RNA promoter of cauliflower mosaic virus. The resultant transgenic rice plants were evaluated for katE expression, catalase activity, growth, and salt tolerance.
Confirmation of the presence of the katE transgene
We obtained 50 regenerated candidates of T0 transgenic rice plants by the induction of shoot regeneration following hygromycin selection of callus transformed with katE. From 46 plants out of the 50, PCR analysis detected the presence of the katE gene (Fig. 2). PCR analysis also detected the presence of the katE gene in T1 transgenic rice plants (data not shown). To estimate the copy number of the katE gene in the T1 transgenic rice plants, we carried out Southern blot analysis using the full-length katE coding sequence as a probe. Under high-stringency condition, only a single band was detected in the KpnI and EcoRI double-digested genome of the transgenic plant (Fig. 3). This indicates that a single copy of katE gene was introduced in the genome since the katE gene has no internal restriction site for KpnI and EcoRI.
Southern blot analysis for the detection of the katE gene. Total genomic DNA samples prepared from the T1 transgenic plant (a) and the wild type (b) were digested with KpnI (Kp) and EcoRI (Ec), separated by electrophoresis, and then subjected to Southern hybridization. Arrows represent katE-containing fragments
Expression of the katE transgene
To determine whether the katE transgene was actively transcribed in rice, we carried out RT-PCR analysis with the katE-specific primers. From all the T1 transgenic plants tested, we detected DNA fragments derived from the transcript of the katE gene (Fig. 4), which indicated a significant level of expression of the katE mRNA.
Catalase activities of the katE-expressing transgenic plants
For investigating the influence of the katE transgene on total catalase activity, we measured the catalase activities of the extracts extracted from T1 transgenic rice plants. All the transgenic lines showed approximately 1.5- to 2.5-fold higher activity of catalase than the wild-type rice plants (Fig. 5).
Growth and fertility of the katE-expressing transgenic plants
Vegetative growth, flowering, and seed formation and maturation of the katE-expressing transgenic plants were similar to those of the wild type under a standard growth condition (data not shown). However, germination rate was lower in seeds produced by T1 transgenic plants (57.9% on average) than in the wild-type seeds (86.7% on average). Grain yield in the T1 transgenic plants was similar to that of the wild-type rice.
Salt tolerance of the katE-expressing transgenic plants
Salt tolerance was compared among the katE-expressing T0 transgenic plants, T1 transgenic plants and the wild-type plants at various concentrations of NaCl. While wild-type rice could not survive for more than 5 days in the presence of low concentrations of NaCl, such as 50 mM, the transgenic plants were not affected significantly in vegetative growth by 100 mM NaCl (Fig. 6; Table 1). At the level of 100 mM NaCl, the T1 transgenic plants grew without suffering noticeable damage for more than 3 months to form seeds. Seed production of the T1 transgenic plants under this condition was about 10% of that of the unstressed transgenic plants. The T0 transgenic plants and T1 transgenic plants could grow almost normally for 14 days even in the presence 250 mM NaCl (Fig. 7), but thereafter chlorosis began gradually (Table 1). Transgenic plants could not grow for more than 5 days at 300 mM (Table 1). When the T1 transgenic plants at the reproductive stage, which had been grown under the standard condition without salt stress, were soaked in 250 mM NaCl solution, they survived for more than 14 days and produced mature seeds (Fig. 8a). Such treatment caused severe damage in the wild type (Fig. 8b).
Comparison of salt tolerance among transgenic rice plants reported
Many efforts have been made to produce salt tolerant rice by genetic manipulation of proteins related to stress protection, such as glutamine synthase (GS), choline oxidase (COD), calcium-dependent protein kinase (CDPK), Na+/H+ antiporter, and others (Hoshida et al. 2000; Sakamoto et al. 1998; Saijo et al. 2000; Ohta et al. 2002; Xu et al. 1996; Tanaka et al. 1999). Transformed rice plants overexpressing GS2, a gene encoding chloroplastic GS, could survive for 2 weeks at 150 mM NaCl (Hoshida et al. 2000). A bacterial gene encoding COD, an enzyme that is involved in the synthesis of compatible solute, glycinebetaine was introduced into rice and the resultant transgenic rice could survive for 1 week at 150 mM NaCl (Sakamoto et al. 1998). Overexpression of OsCDPK7, a gene encoding a stress-responsive CDPK, enabled rice plants to survive for 3 days at 200 mM NaCl (Saijo et al. 2000). AgNHX1, a halophyte gene encoding Na+/H+ antiporter was introduced into rice and the transgenic plants were tolerant against transient exposure to 300 mM NaCl for 3 days (Ohta et al. 2002). In none of these cases, however, has tolerance to long-term treatment with high concentrations of salt been reported so far. In this respect, the katE-expressing transgenic rice plant reported here, which could survive for more than 2 weeks at 250 mM NaCl, and could grow for more than 3 months and produce seeds at 100 mM, is of much note.
How did the katE transgene confer such a strong salt tolerance to rice? The total catalase activity of the transgenic rice was approximately 1.5- to 2.5-fold higher than that of the non-transgenic rice. The simplest explanation is that reinforcement of the ROS scavenging system by the increased catalase activity resulted in a remarkable enhancement of salt tolerance. In addition to the total activity, the subcellular localization of catalase should be taken into consideration. In higher plants, catalase is known to exist in the microbody, mitochondria, and chloroplast, but not in the cytosol. Catalase produced from the katE transgene may reside in the cytosol due to the lack of any specific targeting signals, which might also modulate the ROS scavenging system. This is the quite important and very interesting issue to be addressed, and is under investigation.
References
Aebi H (1984) Catalase in vitro. Methods Enzymol 105:121–126
Akama K, Shiraishi H, Ohta S, Nakamura K, Okada K, Shimura Y (1992) Efficient transformation of Arabidopsis thaliana: comparison of the efficiencies with various organs, plant ecotypes and Agrobacterium strains. Plant Cell Rep 12:7–11
Blumwald E, Aharon GS, Apse MP (2000) Sodium transport in plant cell. Biochem Biophys Acta 1465:140–151
Bohnert HJ, Nelson DE, Jensen RG (1995) Adaptations to environmental stresses. Plant Cell 7:1099–1111
Chu CC, Wang CC, Sun CS, Chen H, Yin KC, Chu YC, Bi FY (1975) Establishment of an efficient medium for anther culture of rice through comparative experiments on the nitrogen saurces. Sci Sin 18:659–668
Epstein E, Norlyn JD, Rush DW, Kingsbury RW, Kelley DB (1980) Cunningham GA, Wrona AF, Saline culture of crops: a genetic approach. Science 210:399–404
Hasegawa PM, Bressan RA, Zhu JK, Bohnert HJ (2000) Plant cellular and molecular responses to high salinity. Annu Rev Plant Physiol Plant Mol Biol 51:463–499
Hiei Y, Komari T, Kumashiro T (1994) Efficient transformation of rice (Oryza sativa L.) mediated by Agrobacterium tumrefaciens and sequence analysis of the boundaries of the T-DNA. Plant J 6:271–281
Hood EE, Helmer GL, Freiley RT, Chilton MD (1986) The hypervirulence of Agrobacterium tumefaciens A281 is encoded in a region of pTiBo542 outside of T-DNA. J Bacteriol 168:1291–1301
Hoshida H, Tanaka Y, Hibino T, Hayashi Y, Tanaka A, Takabe T, Takabe T (2000) Enhanced tolerance to salt stress in transgenic rice that overexpresses chloroplast glutamine synthase. Plant Mol Biol 43:103–111
Kaku N, Hibino T, Meng Y, Tanaka Y, Araki E, Takabe T, Takabe T (2000) Effects of overexpression of Escherichia coli katE and bet genes on the tolerance for salt stress in a freshwater cyanobacterium Synechococcus sp. PCC 7942. Plant Sci 159:281–288
McClung CR (1997) Regulation of catalase in Arabidopsis. Free Radical Biol Med 23:489–496
Murashige T, Skoog F (1962) A revised medium for rapid growth and bioassays with tobacco tissue. Physiol Plant 15:473–497
Nakashima K, Shinwari ZK, Sakuma Y, Seki M, Miura S, Shinozaki K, Yamaguchi-Shinozaki K (2000) Organization and expression of two arabidopsis DREB2 genes encoding DRE-binding proteins involved in dehydration- and high-salinity-responsive gene expression. Plant Mol Biol 42:657–665
Narusaka Y, Nakashima K, Shinwari ZK, Sakuma Y, Furihata T, Abe H, Narusaka M, Shinozaki K, Yamaguchi-Shinozaki K (2003) Interaction between two cis-acting elements, ABRE and DRE, in ABA-dependant expression of arabidopsis RD29A gene in response to dehydration and high-salinity stresses. Plant J 34:137–148
Ngo TT, Lenhoff HM (1980) A sensitive and versatile chromogenic assay for peroxidase and peroxidase-coupled reactions. Anal Biochem 105:389–397
Ohta M, Hayashi Y, Nakashima A, Hamada A, Tanaka A, Nakamura T, Hayakawa T (2002) Introduction of a Na+/H+ antiporter gene from Atriplex gmelini confers salt tolerance to rice. FEBS Lett 532:279–282
Ossowski I, Mulvey MR, Leco PA, Borys A, Loewen PC (1991) Nucleotide sequence of Escherichia coli katE, which encodes catalase HPII. J Bacteriol 173:514–520
Ozeki Y, Matui K, Sakuta M, Matsuoka M, Ohashi Y, Kano-Murakami Y, Yamamoto N, Tanaka Y (1990) Differential regulation of phenylalanine ammonia-lyase genes during anthocyanin synthesis and by transfer effect in carrot cell suspension cultures. Physiol Plant 80:379–387
Saijo Y, Hata S, Kyozuka J, Shimamoto K, Izui K (2000) Over-expression of a single Ca+–dependant protein kinase confers both cold and salt/drought tolerance on rice plants. Plant J 23:319–327
Sakamoto A, Murata A, Murata N (1998) Metabolic engineering of rice leading to biosynthesis of glycinebetaine and tolerance to salt and cold. Plant Mol Biol 38:1011–1019
Serrano R (1996) Salt tolerance in plants and microorganisms: toxicity targets and defense responses. Int Rev Cytol 165:1–52
Tanaka Y, Hibino T, Hayashi Y, Tanaka A, Kishitani S, Takabe T, Yokota S, Takabe T (1999) Salt tolerance of transgenic rice overexpressing yeast mitochondrial Mn-SOD in chloroplasts, Plant Sci 143:131–138
Toki S (1997) Rapid and efficient Agrobacterium-mediated transformation in rice, Plant Mol Biol Rep 15:10–21
Ueda A, Shi W, Nakamura T, Takabe T (2002) Analysis of salt-inducible genes in barley roots by differential display. J Plant Res 115:119–130
Wang H, Miyazaki S, Kawai K, Deyholos M, Galbraith DW, Bohnert HJ (2003) Temporal progression of gene expression responses to salt shock in maize roots. Plant Mol Biol 52:873–891
Wang ZL, Li PH, Fredricksen M, Gong ZZ, Kim CS, Zhang C, Bohnert HJ, Zhu JK, Bressan RA, Hasegawa PM, Zhao YX, Zhang H (2004) Expressed sequence tags from Thellungiella halophila, a new model to study plant salt-tolerance. Plant Sci 166:609–616
Xu D, Duan X, Wang B, Hong B, Ho THD, Wu R (1996) Expression of a late embryogenesis abundant gene, HVA1, from barley confers tolerance to water deficit and salt stress in transgenic rice. Plant Physiol 110:249–257
Yancey PH, Clark ME, Hand SC, Bowlus RD, Somero GM (1982) Living with water stress: evolution of osmolyte system. Science 217:1214–1222
Zhu JK (1999) Understanding and improving plant tolerance to salinity stress. AgBiotechnet 1:1–5
Zhu JK (2000) Genetic analysis of plant salt tolerance using Arabidopsis. Plant Physiol 124:941–948
Zhu JK (2002) Salt and drought stress signal transduction in plants. Annu Rev Plant Biol 53:247–273
Acknowledgments
We thank Mr. Masateru Yamanaka and Mr. Masataka Asada, Department of Agriculture, Tokyo University of Agriculture for technical advice and valuable comments. We also thank Dr. Munetaka Sugiyama, Graduate School of Science, The University of Tokyo for critical reading of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Nagamiya, K., Motohashi, T., Nakao, K. et al. Enhancement of salt tolerance in transgenic rice expressing an Escherichia coli catalase gene, katE. Plant Biotechnol Rep 1, 49–55 (2007). https://doi.org/10.1007/s11816-007-0007-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11816-007-0007-6