Abstract
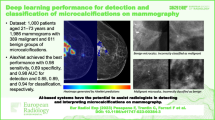
Early detection of breast cancer is clinically important to reduce the mortality rate. In this study, a new computer-aided detection (CAD) and classification system is introduced to classify two types of mammogram tumors (i.e., mass and calcification) as either benign or malignant. In this CAD system, the tumor-like regions (TLRs) are identified using the automated optimal Otsu thresholding method. Afterward, deep convolutional neural networks (CNNs) process the extracted TLRs to extract relevant mammogram features, investigating AlexNet and ResNet-50 architectures. The normalized extracted CNN features are further input to a support vector machine classifier to decode the classes of mammogram structures (i.e., Benign Calcification, Benign Mass, Malignant Calcification, and Malignant Mass nodules). The experimental results are tested on 2800 mammogram images from the Curated Breast Imaging Subset of Digital Database of Screening Mammography, a publicly available dataset. The accuracy of the proposed CAD system, to classify the ROI into one of the four classes, achieves 0.91 using AlexNet and 0.84 using ResNet-50 models, using fivefold cross-validation. Comparison results with the related methods confirm the advantages of the proposed CAD system.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
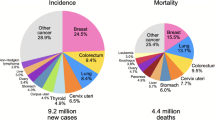
According to the World Health Organization (WHO), breast cancer is the most globally common cancer in women [1]. Each year, around 2.1 million women are diagnosed with breast cancer, resulting in the highest number of women cancer-related deaths. A recent report in 2018 estimates around 627,000 women cancer-related deaths, accounting for around 15% of all women cancer deaths [1, 2]. Low-energy X-rays mammography is considered the standard screening tool to identify the breast cancer abnormalities. Mammogram screening has shown to reduce breast cancer mortality by approximately 20% in high-resource settings [3]. Early detection is clinically important to reduce the cancer mortality rate [4, 5]. However, the early detection of breast cancer on screening mammography is challenging due to the small sizes of potential nodules with respect to the entire breast [4]). Therefore, computer-aided detection (CAD) systems for breast cancer are clinically important to reduce the workload and improve the detection accuracy of radiologists and experts [5].
Recently, convolutional neural networks (CNNs) show impressive performance in the field of pattern recognition, classification, object detection [6, 7], and more specifically in the field of breast cancer detection [8,9,10,11,12,13,14,15,16]. For example, Lévy and Jain [8] used three different network architectures (a CNN model, AlexNet, and GoogLeNet) to classify benign and malignant breast masses, using areas around the breast masses, cropped from the DDSM dataset, achieving accuracies of 60%, 89%, and 92.9%, on the three tested models, respectively. Castro et al. [10] trained a CNN architecture for mass classification into benign or malignant and validated it on three available full mammogram datasets (INbreast, CBIS-DDSM, and a Breast Cancer Digital Repository (BCRP)). The reported sensitivities on the tested datasets are 80%, 80%, and 60%, respectively. To classify breast masses as benign or malignant, Tsochatzidis et al. [12] investigated different deep CNNs (AlexNet, VGG-16, VGG-19, ResNet-50, ResNet-101, ResNet-152, GoogLeNet, Inception-BN-v2), showing that the fine-tuning of the pre-trained ResNet-101 has achieved the best accuracies on DDSM and Region of Interest (ROI) CBIS-DDSM databases of 78.5% and 75.3%, respectively. Chun-minga et al. [13] classified mammography images of a dataset, collected from DDSM and CBIS-DDSM databases, into five classes, i.e., benign calcification (BC), benign mass (BM), malignant calcification (MC), and malignant mass (MM) and normal. They fused two deep CNN networks, achieving an accuracy of 91%. Ragab et al. [14] used AlexNet and SVM classification to classify breast masses in as benign or malignant with an accuracy of 87.2% on preprocessed ROI CBIS-DDSM. Falconí et al. [15] applied a preprocessing step followed by feature extraction using ResNet50 and MobileNet models to classify benign and malignant ROI CBIS-DDSM breast masses, achieving an accuracy of 78.4% and 74.3%, for the two models, respectively. Ansar et al. [16] used the MobileNet model to classify breast masses into benign or malignant, achieving an accuracy of 74.5% on preprocessed ROI CBIS-DDSM data. Although different methods have achieved considerable success to detect and classify breast cancer, there is still a need to investigate how to further improve the classification accuracy. This paper focuses to develop an approach for the early detection of the breast cancer, using the ROI CBIS-DDSM [17] (see Fig. 1), to overcome the following limitations of the existing work:
-
Building new CNN models require huge amount of data for training, to avoid overfitting. In addition, these methods are computationally expensive, especially in the training phase.
-
Training with the whole details of the ROI image represents an overhead and may lead to reduce the detection accuracy.
-
There is a need to design advanced methods and develop new ideas to improve the diagnosis accuracy, especially for early cases, where the nodule size is small with respect to the image size.
Typical samples for the four classes of the ROI CBIS-DDSM database [17]. From left to right: Benign Calcification (BC), Benign Mass (BM), Malignant Calcification (MC), and Malignant Mass (MM) images. Nodules are outlined using white circles
The main features/contributions of this work are as follows:
-
Unlike the traditional CNN, which work on the original mammogram or cropped regions, the proposed CAD system applies an automated optimal dynamic thresholding method to extract the tumor-like regions (TLRs), eliminating the need for processing the whole ROI image for detecting breast nodules and enabling the CNN model to focus on the fine TLRs details, improving the chance of the early detection of the small size nodules.
-
Performance is evaluated on the challenging standard CBIS-DDSM dataset [17], achieving superior performance over the competing methods.
2 Materials and methods
The proposed deep learning system, see Fig. 2, is designed based on three processing stages. In the first sage, the TLRs are identified based on an automated thresholding method. In the second stage, a transfer deep learning is performed by investigating two architectures, i.e., ResNet-50 and AlexNet. Finally, a classification stage is used to classify TLR images based on the normalized deep extracted features.
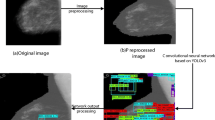
Diagram of the proposed CAD system for breast cancer detection. First, tumor-like regions (TLRs) are identified from the input raw image by Otsu thresholding. Second, TLRs are processed using a CNN model to form a feature descriptor. Finally, classification is performed to produce the breast classes (BC, BM, MC, and MM)
2.1 Collected database (CBIS-DDSM)
To test the proposed CAD system, the ROI CBIS-DDSM [17] images are utilized, composed of 3549 ROI mammogram images (1852 calcification (1132 BC and 720 MC) and 1697 mass (913 BM and 784 MM) images), see Fig. 1. CBIS-DDSM is an updated and standardized version of the Digital Database for Screening Mammography (DDSM) [18]. Image type has been converted from DICOM format to PNG format. Figure 1 shows typical samples of CBIS-DDSM database of the four classes (i.e., BC, BM, MC, and MM).
AlexNet [27] architecture
ResNet-50 [28] architecture
2.2 Extracting tumor-like regions (TLRs)
The first stage in the proposed CAD system aims at extracting TLRs to enable the CNN model to focus on the fine TLRs details, improving the chance of the early detection of the small size nodules and eliminating the need for processing the unnecessary details of the original CBIS-DDSM image. TLRs is a grey-level image, where the regions with similar grey levels as the tumors are extracted. To achieve this goal, a Gaussian smoothing is firstly applied to the original ROI image, \(\mathrm {IM_{ORG}}\), in order to enable the proposed Otsu method to focus on the global image features. Let the Gaussian kernel be G. The image is convolved with G [19, 20]:
where * denotes the convolution operator and \(\mathrm {IM_{Smooth}}\) is the smoothed image. The Gaussian kernel G is defined as follows:
where (x, y) is the pixel’s Cartesian coordinates and \(\sigma \) denotes the standard deviation of the Gaussian kernel. Afterward, the Otsu method, an optimum global thresholding method, proposed by Otsu [21], is used to extract TLRs, utilizing the zeros and the first-order cumulative moments of the gray-level histogram. Otsu thresholding is optimum in the sense that it maximizes the between class variance, a well-known measure used in statistical discriminant analysis [22, 23]. Assume that the grey level histogram of the image, \(\mathrm {IM_{Smooth}}\), is composed of L pins. Let p(i) be the probability of an intensity grey level i,\(i\) \(\in \){0,L-1}. Let T be an arbitrary random threshold. Let the weights \(\omega _{0}\) and \(\omega _{1}\) define the probabilities of the two classes, separated by the threshold T, as follows: \({\omega _{0} (T) = \sum ^{T-1}_{i=0}{p(i)}}\) and \({\omega _{1} (T) = \sum ^{i=T}_{L-1} {p(i)}} \). The means of the two classes are:
The between class variance, \(\sigma _{b}^2\) , can be expressed as follows [22, 23]: \({\sigma _{b}^2(T) = \omega _{0}\omega _{1} (\mu _{0}- \mu _{1})^2}\). To estimate the TLRs, an algorithm of six steps is followed. First, set initial random values for the class probabilities and means, i.e., \(\omega _{0}\),\(\omega _{1}\),\(\mu _{0}\), and \(\mu _{1}.\) Second, for all possible thresholds T=1,2, ..., L-1: Update \(\omega _{0},\omega _{1},\mu _{0}, \mathrm {and~} \mu _{1}\) accordingly. Third, compute \(\sigma _{b}^2(T)\) for each threshold, T. Forth, determine the desired optimal global threshold, \(T\mathrm {_{OPT}}\), that globally maximizes the between class variance to correspond to the maximum \(\sigma _{b}^2(T)\) for all \(\mathrm {T}\) values. Fifth, compute \(\mathrm {IM_{BTLRs}}\), where all pixels less than \(T\mathrm {_{OPT}}\) are classified as background (binary zero), and the remaining pixels are TLRs (binary one), (see Fig. 3). Finally, Output the grey-level TLRs image \(\mathrm {IM_{TLRs}}\), by the pixel-by-pixel multiplication of \(\mathrm {IM_{BTLRs}}\) with the original image \(\mathrm {IM _{ORG}}\) as follows: \(\mathrm {IM_{TLRs}}\)=\(\mathrm {IM _{ORG}}\) * \(\mathrm {IM_{BTLRs}}\). Figure 3 illustrates the step-by-step algorithm of the proposed method for extracting TLRs. Note that the TLRs are not the tumor class, but it contains the regions of the grey levels that are similar to the tumor grey levels (bright pixels).
2.3 Utilized CNN models
A convolutional neural network (CNN) consists of two types of layers; convolutional layers to extract lower- and higher-order image features, and fully connected layers to perform the classification [24,25,26]. AlexNet [27] and ResNet [28] CNN models are from the most commonly used popular architectures, that achieve a remarkable success in medical applications (i.e., lung cancer detection [29] and face recognition [30]), and in particular breast cancer detection [8, 11, 12, 14]. Therefore, they have been adopted in the proposed system.
AlexNet has five convolution layers, three pooling layers, and three fully connected layers [27, 31] (see Table 1). The AlexNet CNN architecture is shown in Fig. 4 The layers of Conv. Layer 1 to Conv. Layer 5, in Fig. 4, are the convolution layers. Each neuron in the convolution layers computes a dot product between their weights and the local region that is connected to the input volume [32]. Each convolutional layer is followed by a pooling layer in order to perform a down sampling operation along the spatial dimensions to reduce the amount of the computations and improve the robustness [32]. Additionally, the fully connected layers are FC6, FC7 and FC8, as shown in Fig. 4. The role of the fully connected layers are to process the extracted convolutional layers’ features and output the most relevant feature vector for classification [32].
ResNet is a short name for Residual Network. The basic idea of a ResNet model is to skip blocks of convolutional layers by using shortcut connections (see Fig. 5) [12]. ResNet-50 is based on a residual learning framework, where layers within a network are reformulated to learn a residual mapping rather than the desired unknown mapping between the inputs and outputs. Such a network is easier to optimize and consequently enables training of deeper networks, which correspondingly leads to an overall improvement in the network capacity and performance [12].
In order to reduce the training overhead, the weights of the convolutional layers of the pretrained models are transferred without training, and only the fully connected layers are trained using the CBIS-DDSM data. To apply transfer learning of the CNN models, the last fully connected layer of the pre-trained models (FC8 layer in AlexNet or FC1000 layer in ResNet50) is replaced with a shallow classifier (SVM). The vectors of activities of the FC7 layer in AlexNet or the flatten layer (just before FC1000) in Res-Net50, represent the feature descriptor of the input CBIS-DDSM image. Features are further normalized between 0 and 1 to the input of the SVM classifier.
2.4 Classification
To classify the mammogram images, a SVM shallow classifier is used. SVM is a supervised machine learning algorithm that sorts data in categories. The idea of SVM is to formulate a computationally efficient way of learning by separating hyper planes in a high dimensional feature space [33, 34]. The input of the classifier is the vectors of activities (FC7 in AlexNet or flatten layer in ResNet-50) and the output is the ROI class. The SVM classifier is chosen using binary nonlinear kernel to reflect the variability of the classes. For the case of four classes, We used one-versus-all SVM, in which we fuse four binary trained classifiers, each processes a class, where the data from the class is treated as positive, and the data from all the other classes is treated as negative. The performance is evaluated for either the ResNet-50 or AlexNet, each along with the SVM classifier based on a standard performance evaluation metric, i.e., the classification accuracy, in order to select the best model/classifier.
2.5 Performance evaluation
To test the proposed system, we used six standard metrics to evaluate the system performance, i.e., Accuracy (ACC), Sensitivity (SEN), Specificity (SPE), Positive Predictive Value (PPV), Negative Predictive Value (NPV), and F1 score (FSC) defined as follows [35]:
3 Experimental results and discussions
This section illustrates, in details, the experimental setup, comparison results, and related discussions.
3.1 Experimentation setting
To evaluate the proposed system, CBIS-DDSM ROI dataset is used. To train the classifier, data is randomly divided into training set (70%) and testing set (30%). To train the deep learning model (AlexNet and ResNet-50), the Bayesian optimizer is used to minimize the binary cross entropy function, using a learning rate of \(10^{(-4)}\). During training, the data is shuffled using a mini-patch size of 128. The maximum number of epochs is set to 20.
3.1.1 Threshold cases
To verify the advantages of using the automated adaptive Otsu thresholding to identify TLRs, different values of fixed thresholds are investigated and the performance results of the proposed system have been carried and compared with values of fixed threshold cases. Assuming that the ROI grey levels are normalized between 0 and 1, Case I uses original ROI dataset. Case II to case VII use ROI with thresholds T of 0.60 to 0.80 with a step of 0.05.
3.1.2 Data balancing
The experiments are run through two setting (a) imbalance data that contains the whole data (3549 images of imbalanced number of images per each class), and (b) balanced data which contains 2800 images (700 image per each class) The proposed system adopted balancing the data to avoid the bias toward the classes of large numbers.
3.2 Comparison results
3.2.1 Experiments using the whole imbalanced data
Two investigated deep models (i.e., AlexNet and ResNet-50) have been compared, using the different values of thresholds, for breast nodule diagnosis, i.e., classification of breast masses into benign and malignant (BM or MM), and classification of breast calcification into benign and malignant (BC or MC). Table 2 overviews the achieved comparison results. Otsu’s thresholding, besides being automated and adaptive to each input TLR, achieves the best performance for both the investigated ResNet-50, and AlexNet models, as shown in Table 2. In addition, Table 2 demonstrates that when classification breast mass the AlexNet model achieves slightly better results than the ReseNet-50 model. This may be due to the large number of layers of the Resenet50 (50 layers) compared to the AlexNet model (five convolutional and two fully connected layers), that lead to data overfitting in the case of ResNet-50 model, taking into account limited size of the standard ROI CBIS-DDSM data. Figure 6 shows visual comparison results of the proposed method using the whole data. As shown in Fig. 6, the proposed method shows superior results over the competing fixed thresholding methods. In addition to the ability of classifying breast calcification into benign or malignant (BC or MC), see Table 2, the proposed system is tested to detect the malignancy in case of calcification nodules, which is more challenging, since the nodule shape in case of the calcification nodules is complex and distributed. In this experiments, ResNet-50 achieves slightly better results, due to the complex task of the nodule diagnosis, in case of the calcification nodules.
Furthermore, the proposed system is tested to classify the ROI data into one of the four classes, i.e., BC, MC, BM, or MM. As shown in Table 2, the proposed system achieves accuracies of 0.81 and 0.90 for using ResNet-50 and AlexNet models, respectively, in case of using the whole imbalanced data. As expected, the accuracy of the 4-class classifier is significantly below that of a 2-class classifier. This is due to the added challenge of differentiating between benign and malignant, in additional to distinguishing between mass and calcification.
3.2.2 Experiments using the balanced data
To remove the bias toward the classes of large number, we adopted balancing the data, such that each class contains 700 images (a total of 2800 images selected randomly). As expected, using balanced data further improve the performance (see Table 2). Figure 7 shows the confusion matrix for classifying the four classes using Alexnet and balanced data using our proposed method. As shown in the figure, the total accuracy of all classes reaches 93.2%, which outperforms the case of using imbalanced data, as expected. In order to investigate the sensitivity of the proposed method to the selected training data, fivefold cross-validation is applied, achieving relatively low standard deviation, which indicates that our proposed method is insensitive to the training data selection (see Table 3).
Table 3 shows that the proposed system achieves a better results to the most similar related work [13]. Note that our experiments do not consider the normal (N) class, since the utilized standard ROI CBIS-DDSM does not contain any normal record. To demonstrate the advantage of the proposed system for breast cancer identification, Table 3 compares the results of the proposed system with the related state-of the-art methods. Table 3 shows the superior of the proposed system for classifying breast masses into benign and malignant. This is due to that the proposed automated thresholding improves the detection rate, since the deep model focuses to detect tumors within only the TLR regions.
3.3 Limitations
Although the proposed method has achieved promising results comparing with the competing methods, it fails to correctly classify all the images. Last column in Fig. 6 shows an example of incorrect classification. In the future, we will try adding more features and/or different CNN models to improve further the classification accuracy.
4 Conclusion
In this work, a CAD system for early detect breast cancer based on deep learning is proposed. Unlike related work, the utilized CNN models extract features from the TLRs based on automated adaptive Otsu thresholding, in order to improve the training capabilities of the deep learning model. A SVM classifier is used to classify mammogram images into the nodule classes, i.e., BC, MC, BM, and MM. Experiments results on the ROI CBIS-DDSM data confirm the superior of the proposed method over other related works. In the future, other databases will be investigated to test the robustness of the proposed system. In addition, other feature models will be tested in order to improve the performance.
References
WHO. World health organization. https://www.who.int/cancer/prevention/diagnosis-screening/breast-cancer/en/ (2020). Accessed 7 Nov 2020
DeSantis, C.E., Miller, K.D., GodingSauer, A., Jemal, A., Siegel, R.L.: Cancer statistics for African Americans. CA Cancer J. Clin. 69(3), 211 (2019)
Chen, H.L., Yang, B., Wang, G., Wang, S.J., Liu, J., Liu, D.Y.: Support vector machine based diagnostic system for breast cancer using swarm intelligence. J. Med. Syst. 36(4), 2505 (2012)
Shen, L., Margolies, L.R., Rothstein, J.H., Fluder, E., McBride, R., Sieh, W.: Deep learning to improve breast cancer detection on screening mammography. Sci. Rep. 9(1), 1 (2019)
Hamidinekoo, A., Denton, E., Rampun, A., Honnor, K., Zwiggelaar, R.: Deep learning to improve breast cancer detection on screening mammography. Med. Image Anal. 47, 45 (2018)
Chan, H.P., Samala, R.K., Hadjiiski, L.M., Zhou, C.: Deep Learning in Mammography and Breast Histology, An Overview and Future Trends, Medical Image Analysis, pp. 3–21. Springer, Berlin (2020)
Altaf, F., Islam, S.M., Akhtar, N., Janjua, N.K.: Going deep in medical image analysis: concepts, methods, challenges, and future directions. IEEE Access 7, 99540 (2019)
Lévy, D., Jain, A.: Breast mass classification from mammograms using deep convolutional neural networks, arXiv preprint arXiv:1612.00542 (2016)
Yi, D., Sawyer, R.L., CohnIII, D., Dunnmon, J., Lam, C., Xiao, X., Rubin, D.: Optimizing and visualizing deep learning for benign/malignant classification in breast tumors, arXiv preprint arXiv:1705.06362 (2017)
Castro, E., Cardoso, J.S., Pereira, J.C.: Elastic deformations for data augmentation in breast cancer mass detection. In: 2018 IEEE EMBS International Conference on Biomedical and Health Informatics (BHI), pp. 230–234. IEEE (2018)
Xi, P., Shu, C., Goubran, R.: Abnormality detection in mammography using deep convolutional neural networks. In: 2018 IEEE International Symposium on Medical Measurements and Applications (MeMeA), pp. 1–6. IEEE (2018)
Tsochatzidis, L., Costaridou, L., Pratikakis, I.: Deep learning for breast cancer diagnosis from mammograms—a comparative study. J. Imaging 5(3), 37 (2019)
Chun-ming, T., Xiao-mei, C., Xiang, Y., Fan, Y., et al.: Five classification of mammography images based on deep cooperation convolutional neural network. Am. Sci. Res. J. Eng. Technol. Sci. ASRJETS 57(1), 10 (2019)
Ragab, D.A., Sharkas, M., Marshall, S., Ren, J.: Breast cancer detection using deep convolutional neural networks and support vector machines. PeerJ 7, e6201 (2019)
Falconí, L.G., Pérez, M., Aguilar, W.G.: Transfer learning in breast mammogram abnormalities classification with mobilenet and nasnet. In: 2019 International Conference on Systems, Signals and Image Processing (IWSSIP), pp. 109–114. IEEE (2019)
Ansar, W., Shahid, A.R., Raza, B., Dar, A.H.: Breast cancer detection and localization using MobileNet based transfer learning for mammograms. In: International Symposium on Intelligent Computing Systems, pp. 11–21. Springer (2020)
Lee, R.S., Gimenez, F., Hoogi, A., Miyake, K.K., Gorovoy, M., Rubin, D.L.: A curated mammography data set for use in computer-aided detection and diagnosis research. Sci. Data 4(1), 1 (2017)
Bowyer, K., Kopans, D., Kegelmeyer, W., Moore, R., Sallam, M., Chang, K., Woods, K.: The digital database for screening mammography. In: Third International Workshop on Digital Mammography, vol. 58, p. 27 (1996)
Al-Najdawi, N., Biltawi, M., Tedmori, S.: Mammogram image visual enhancement, mass segmentation and classification. Appl. Soft Comput. 35, 175 (2015)
Lukac, R.: Single-Sensor Imaging: Single-Sensor Imaging: Methods and Applications for Digital Cameras. CRC Press, Boca Raton (2018)
Otsu, N.: A threshold selection method from gray-level histograms. IEEE Trans. Syst. Man Cybern. 9(1), 62 (1979)
Deepa, S., Subbiah, B.V.: Efficient ROI segmentation of digital mammogram images using Otsu’s N thresholding method. Natl. J. Adv. Comput. Manag. 4, 1 (2013)
Swetha, T., Bindu, C.H.: Detection of breast cancer with hybrid image segmentation and Otsu’s thresholding. In: 2015 International Conference on Computing and Network Communications (CoCoNet), pp. 565–570. IEEE (2015)
LeCun, Y., Kavukcuoglu, K., Farabet, C.: Convolutional networks and applications in vision. In: Proceedings of 2010 IEEE International Symposium on Circuits and Systems, pp. 253–256. IEEE (2010)
Anagnostis, A., Asiminari, G., Papageorgiou, E., Bochtis, D.: A convolutional neural networks based method for anthracnose infected walnut tree leaves identification. Appl. Sci. 10(2), 469 (2020)
Truong, T.D., Pham, H.T.T.: Breast cancer histopathological image classification utilizing convolutional neural network. In: International Conference on the Development of Biomedical Engineering in Vietnam, pp. 531–536. Springer (2018)
Krizhevsky, A., Sutskever, I., Hinton, G.E.: Imagenet classification with deep convolutional neural networks. Adv. Neural Inf. Process. Syst. 25, 1097 (2012)
He, K., Zhang, X., Ren, S., Sun, J.: Deep residual learning for image recognition (2015), arXiv preprint arXiv:1512.03385 (2016)
Elnakib, A., Amer, H.M., Abou-Chadi, F.E.: Early lung cancer detection using deep learning optimization. Int. J. Online Biomed. Eng. iJOE 16(06), 82 (2020)
Moustafa, A.A., Elnakib, A., Areed, N.F.: Age-invariant face recognition based on deep features analysis. Signal Image Video Process. 14, 1027 (2020)
Wu, Z., Shen, C., Van DenHengel, A.: Wider or deeper: revisiting the resnet model for visual recognition. Pattern Recogn. 90, 119 (2019)
Suzuki, S., Zhang, X., Homma, N., Ichiji, K., Sugita, N., Kawasumi, Y., Ishibashi, T., Yoshizawa, M.: Mass detection using deep convolutional neural network for mammographic computer-aided diagnosis. In: 2016 55th Annual Conference of the Society of Instrument and Control Engineers of Japan (SICE), pp. 1382–1386 IEEE (2016)
Gunn, S.R., et al.: Support vector machines for classification and regression. ISIS Tech. Rep. 14(1), 5 (1998)
Pisner, D.A., Schnyer, D.M.: Support Vector Machine, Machine Learning, pp. 101–121. Elsevier, Amsterdam (2020)
Zhu, W., Zeng, N., Wang, N., et al.: Sensitivity, specificity, accuracy, associated confidence interval and ROC analysis with practical SAS implementations. NESUG Proc. Health Care Life Sci. Baltim. Md. 19, 67 (2010)
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Hekal, A.A., Elnakib, A. & Moustafa, H.ED. Automated early breast cancer detection and classification system. SIViP 15, 1497–1505 (2021). https://doi.org/10.1007/s11760-021-01882-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11760-021-01882-w