Abstract
Thraustochytrium sp. 26185 is a marine protist that can produce a large amount of docosahexaenoic acid (DHA, 22:6n-3), an ω3 very long chain polyunsaturated fatty acid (VLCPUFA) of nutritional importance. However, the mechanism of how this fatty acid is synthesized and assembled into the storage lipid triacylglycerol is unclear. Here we report sequencing of the whole genome and genomic analysis of genes involved in the biosynthesis and assembly of the fatty acids in this species. Genome sequencing produced a total of 2,418,734,139 bp clean sequences with about 62 fold genome coverage. Annotation of the genome sequences revealed 10,797 coding genes. Among them, 10,216 genes could be assigned into 25 KOG classes where 451 genes were specifically assigned to the group of lipid transport and metabolism. Detailed analysis of these genes revealed co-existence of both aerobic pathway and anaerobic pathways for the biosynthesis of DHA in this species. However, in the aerobic pathway, a key gene encoding stearate Δ9 desaturase introducing the first double bond to long chain saturated fatty acid 18:0 was missing from the genome. Genomic survey of genes involved in the acyl trafficking among glycerolipids showed that, unlike plants, this protist did not possess phosphatidylcholine:diacylglycerol cholinephosphotransferase, an important enzyme in bridging two types of glycerolipids, diacylglycerols (DAG) and phosphatidylcholines (PtdCho). These results shed new insight on the biosynthesis and assembly of VLCPUFA in the Thraustochytrium.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Very long chain polyunsaturated fatty acids (VLCPUFA) such as arachidonic acid (ARA, 20:4n-6), eicosapentaenoic acid (EPA, 20:5n-3) and docosahexaenoic acid (DHA, 22:6n-3) are crucial for human health and well-being. Dietary supplementation of these fatty acids have shown that they can improve performance of eyes, brain and immune systems, and provide protection against chronic diseases such as cardiovascular disorders, inflammatory diseases and metabolic syndrome [1]. Many clinical and animal studies demonstrate that VLCPUFA improves neural development, decrease inflammation, promote angiogenesis, and prevent atherothrombosis through the lipid mediators derived from VLCPUFA by a variety of enzymes, such as cyclooxygenases, lipoxygenases and cytochrome P450 enzymes [2–5].
De novo biosynthesis of VLCPUFA is believed to occur only in microorganisms and go through two distinct pathways [6]. The aerobic pathway involves alternating elongation and desaturation steps to introduce double bonds and to extend the acyl chain. Desaturations are catalyzed by a variety of desaturases including front-end and methyl-end desaturases and require molecular oxygen as a cofactor for the oxidation. Elongations are catalyzed by an elongase complex of four catalytic enzymes with substrate specificity primarily defined by the condensing enzyme. For instance, in the aerobic pathway of DHA synthesis, saturated stearic acid (18:0) produced primarily by a fatty acid synthase (FAS) goes through a series of desaturation and elongation steps by various desaturases and elongases such as ∆9 desaturase, ∆12 desaturase, ∆15 desaturase, ∆6 desaturase, ∆6 elongase, ∆5 desaturase, ∆5 elongase and ∆4 desaturase [6]. On the other hand, the anaerobic pathway is catalyzed by a polyketide synthase (PKS)-like mega-enzyme called polyunsaturated fatty acid (PUFA) synthase using acetate as the precursor to synthesize VLCPUFA directly. The double bonds are introduced during the extending process of acyl chains without molecular oxygen involvement [6].
Thraustochytrium sp. 26185 is a marine protist that can produce a high level of VLCPUFA such as DHA and docosapentaenoic acid (DPA, 22:5n-6) in storage lipid triacylglycerols (TAG) [7, 8]. This source of VLCPUFA has been attempted for use in functional food and animal feeds. However, the biosynthesis and assembly of VLCPUFA in glycerolipids is not well understood. In 2001, the first ∆4 desaturase involved in the biosynthesis of DHA was identified and characterized from this species, providing unambiguous evidence that DHA could be synthesized through the final ∆4 desaturation step in the aerobic pathway [9]. Almost at the same time, a PKS-like PUFA synthase was found for the biosynthesis of DHA in Schizochytrium sp. 20888, a closely related species to Thraustochytrium sp. 26185 [10–12]. This result has raised questions about if an anaerobic pathway also exists for VLCPUFA synthesis in Thraustochytrium sp. 26185, and if it does, which pathway is more important for the DHA biosynthesis. To answer these questions, we attempted to sequence the entire genome of Thraustochytrium sp. 26185. The genome sequence de novo assembly and annotation revealed the genome size of Thraustochytrium sp. 26185 at about 38.6 Mb with a GC content of 63 %, and 10,797 coding genes. Genomic analysis of these genes showed that both aerobic and anaerobic pathways for the biosynthesis of VLCPUFA indeed co-exist in this species. However, in the aerobic pathway, a gene encoding stearate ∆9 desaturase for introducing the first double bond to long chain fatty acid 18:0 synthesized by Type I fatty acid synthase, was missing from the genome, implying the aerobic pathway might be incomplete and the alternative anaerobic pathway might be mainly responsible for the biosynthesis of VLCPUFA. Additionally, genome sequence analysis of genes involved in acyl trafficking among glycerolipids showed that the fatty acids could be acylated into the glycerol backbone to synthesize both neutral lipids and polar lipids. However, a gene encoding phosphatidylcholine:diacylglycerol cholinephosphotransferase (PDCT), a critical enzyme involved in the acyl trafficking between phosphatidylcholine (PtdCho) and diacylglycerol (DAG), was missing from the genome, implying that DAG, an immediate precursor for the biosynthesis of TAG might not derive from the catalytic process, as seen in plants. These results shed new insights into the biosynthesis and assembly of VLCPUFA in the Thraustochytrium, which is instrumental for not only genetic improvement of the production of VLCPUFA in the species, but also for the transgenic production of these fatty acids in heterologous systems, particularly oil seed crops.
Materials and Methods
Thraustochytrium Cultivation
The strain, Thraustochytrium sp. 26185, was purchased from the American Type Culture Collection (ATCC 26185). It was cultured at room temperature in a BY+ medium containing 0.1 % (wt/vol) yeast extract, 0.1 % (wt/vol) peptone, and 0.5 % (wt/vol) d-glucose in seawater. Seawater was prepared by dissolving 40 g of sea salts (Sigma-Aldrich) in 1 L of distilled water. The cells were harvested by centrifugation at 2500×g for 20 min, washed twice with 50 mL of seawater, and used for genomic DNA extraction.
DNA Preparation and Library Construction
Genomic DNA was extracted from Thraustochytrium sp. 26185 using E.Z.N.A HP Fungal DNA kit (Omega, bio-tek) according to the manufacturer’s instructions. Sequencing libraries were prepared according to the Illumina protocol (Illumina, San Diego, CA, USA). A total of 5 μg of genomic DNA was fragmented using a Branson sonicator and the ends of DNA fragments were repaired according to previously published method [13]. After being end-blunted, ligated with Hiseq adaptors and amplified by polymerase chain reaction (PCR), the products were purified using agarose gel and cloned into a TOPO plasmid to construct DNA libraries for sequencing. The 300 bp library was used for Illumina Hiseq paired-end sequencing, and the 3 kb library was used for Illumina Hiseq mate-pair sequencing. Sequencing of both libraries was performed on a Hiseq 2000 sequencer.
De Novo Assembly
Processing and assembly of the sequenced data was carried out using a de novo method as described previously [14–17]. Raw data were filtered out of adaptors, low quality and duplicated reads via FastQC. Reads with quality score <20 (one error in 100) and length <25 bp were all reckoned as low quality reads. The de novo sequence assembly was performed by SOAPdenovo assembler (http://soap.genomics.org.cn/soapdenovo.html) [18–20]. Contigs were generated by assembling clean paired-end data and scaffolds were then generated after adding clean mate-pair data.
Gene Annotation and Analysis
The draft genome sequence was annotated according to the standard method of Integrated Microbial Genomes EXPERT Review (IMG-ER) platform [15, 21]. The sequences of coding region were predicted by Augustus program (http://bioinf.uni-greifswald.de/augustus/) and were confirmed by BLASTP search [22] with an E value threshold of 1E−05 against non-redundant protein database [23]. Gene ontology (GO) annotation of each predicted protein sequence was performed using Blast2GO program (v.2.4.2) with default parameters [24, 25]. The KOG and KEGG pathway annotation was also performed by BLAST search against Eukaryotic Cluster of Orthologous Groups database (http://www.ncbi.nlm.nih.gov/COG/) and Kyoto Encyclopedia of Genes and Genomes database (http://www.genome.jp/kegg/), respectively.
Results
Genome Sequencing and De Novo Assembly
To obtain a genome-wide survey of genes associated with the biosynthesis and assembly of VLCPUFA in Thraustochytrium sp. 26185, the whole genome was sequenced using Illumina sequencing platform. After cleaning and quality checks, 2 × 16,399,848 and 2 × 9,797,372 raw paired-end and mate-pair reads with the length of 101 bp were generated. The Poisson distributions of GC content and sequencing depth, as well as centralized plots of the distributions of sequencing reads with their GC content and depth (Fig. 1) indicated that no external DNA contaminations and sequencing bias were observed and the libraries generated for sequencing were of high quality.
After filtering out low quality data, a total of 2,418,734,139 bp clean sequences were obtained and assembled into 8,130 contigs with 5449 large contigs (>1000 bp) and the largest contig reached 88,386 bp in length. These contigs were then assembled into 2250 scaffolds with the largest scaffold being 819,459 bp in length. The calculated genome size of Thraustochytrium sp. 26185 was about 38.6 Mb with a GC content of 62.9 % and an N rate (sequencing gaps) of 3.3 % (Table 1).
Genome Sequence Annotation
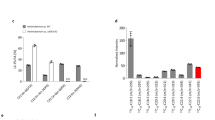
To annotate the genome sequences, the assembled contigs and scaffolds were analyzed by Augustus program with default parameters. In total, 10,797 predicted coding genes were detected, of which 8360 had homologous hits by BLAST searches. From Gene Ontology (GO) annotation, 3178 genes were assigned into 46 groups in three main GO categories: cellular component, molecular function and biological process (Fig. 2). Assignments of these genes to the biological process made up the majority, followed by cellular component and molecular function. To better describe the genome composition, the annotated sequences were BLAST searched against the Eukaryotic Clusters of Orthologous Groups (KOG) database. In total, 10,216 genes were assigned into 25 KOG classifications (Fig. 3). Among the KOG groups, the cluster of signal transduction mechanisms (13.4 %) represented the largest group, followed by the groups of general function prediction (12.9 %), posttranslational modification, protein turnover, chaperones (8.6 %), and transcription (6.4 %). To our interest, there were 451 genes assigned into the group of lipid transport and metabolism. In order to better classify the annotated genes, we BLAST searched the genes against KEGG pathway database. In total, 2634 genes were mapped into a different pathway with 135 in lipid metabolisms (Table 2).
Genomic Analysis of Genes Involved in VLCPUFA Biosynthesis
In Thraustochytrium sp. 26185, like other fungi, the primary fatty acid biosynthesis from acetic acid to long chain saturated fatty acids (16:0 and 18:0) was catalyzed by Type I fatty acid synthase (FAS), a large multifunctional polypeptide with 4053 amino acids containing four catalytic domains for biochemical reactions: condensation, ketoacyl reduction, hydroxyacyl dehydration and enoyl reduction. In addition, it also possessed an acyl carrier protein (ACP) domain for binding phosphopantetheine prosthetic group to carry an acyl chain. Moreover, a malonyl-CoA ACP transacylase (MAT) domain for catalyzing the conversion of malonyl-CoA to malonyl-ACP was also found in the enzyme. The FAS was highly homologous to FAS from Schizochytrium sp. with about 90 % identity at the amino acid level. Furthermore, a coding region for acetyl-CoA carboxylase (ACC) of 2326 amino acids with biotin carboxylase and biotin-carboxyl-transferase domains was also identified in this species. This enzyme converted acetyl-CoA into malonyl-CoA, which was the rate limiting step of fatty acid synthesis. Additionally, two separate genes encoding MAT were also detected in the genome (Table 3).
The biosynthesis of VLCPUFA in the aerobic pathway involves a series of desaturation and elongation reactions (Table 3). In the Thraustochytrium, four putative methyl-end desaturase genes were detected. The functions of two of them were confirmed by heterologous expression in yeast (unpublished data). One encoded a Δ12 desaturase converting oleic acid (18:1-9) to linoleic acid (18:2-9, 12) while the other coded for an ω3 desaturase for converting very long chain ω6 fatty acids to their corresponding ω3 counterparts. In the category of front-end desaturases, the genome possessed three putative Δ6 desaturases, one Δ5 desaturase and one Δ4 desaturase to sequentially introduce three double bonds towards the carboxyl end. Among them, the gene encoding Δ5 desaturase and Δ4 desaturase were previously characterized [9]. Catalytic function of one putative Δ6 desaturase was recently confirmed by heterologous expression in yeast (unpublished data). All these front-end desaturases had a cytochrome b5-like domain at their N-termini. However, no gene encoding any Δ9 desaturase to convert stearic acid (18:0) to oleic acid (18:1-9) was identified in the genome.
In addition, three condensing enzymes (ELO type) were found to be possibly responsible for elongating polyunsaturated fatty acids such as Δ9 desaturated fatty acids linoleic acid (LA, 18:2-9,12) and α-linolenic acid (ALA, 18:3-9,12,15), or Δ6 desaturated fatty acids γ-linoleic acid (GLA, 18:3-6,9,12) and stearidonic acid (SDA, 18:4-6,9,12,15) or Δ5 desaturated fatty acids arachidonic acid (ARA, 20:4-5,8,11,14) and EPA (20:5-5,8,11,14,17) (Table 3) [26]. One of these elongases was previously characterized in both yeast and plant [27]. The elongation of fatty acids involves four biochemical reactions for ketoacyl-CoA condensation, ketoacyl-CoA reduction, hydroxyacyl-CoA dehydration and enoyl-CoA reduction. Accordingly, one gene was found to encode ketoacyl-CoA reduction, three genes were identified to encode hydroxyacyl-CoA dehydration and one gene was detected to encode enoyl-CoA reduction in the genome.
Taken together, these results indicate that an aerobic pathway exists for the biosynthesis of VLCPUFA in Thraustochytrium sp. 26185. However, the pathway might not be complete since a Δ9 desaturase for the introduction of the first double bond into stearic acid was not present in the genome. The Δ9 desaturated monounsaturated acid, oleic acid (18:1-9), is the key precursor for the VLCPUFA synthesis in the aerobic pathway.
In an anaerobic pathway for the biosynthesis of VLCPUFA, three huge genes without introns encoding three subunits (A, B and C) of a PUFA synthase were found in Thraustochytrium sp. 26185 (Table 3). Subunit A (TsPfs-A) was 2813 amino acid long composed of one 3-ketoacyl-ACP synthase domain (KS1) for condensing malonyl-ACP with the existing acyl chain to form 3-ketoacyl-ACP, one malonyl-CoA:ACP acyltransferase domain (MAT) for transferring malonyl-CoA to ACP domains, 8 acyl carrier protein (ACP) domains, one ketoacyl-ACP reductase domain (KR) for reducing keto group of acyl chains and one hydroxyacyl-ACP dehydrogenase (DH1) domain for removing a water molecule and generating a double bond. Subunit B was 2049 amino acids in length comprising two KS domains (KS2 and KS3), one acyl transferase domain (AT), and one enoyl-ACP reductase domain (ER1) for reducing double bonds. Two KS domains (KS2 and KS3) in Subunit B were located side by side. KS2 had sequence similarity to E. coli FabB while KS3 had sequence similarity to PKS chain length factors. This arrangement might be vital for retaining double bonds during the VLCPUFA synthesis. Subunit C was 1497 amino acids in length consisting of two dehydratase domains (DH2 and DH3) for introducing double bonds, and one ER domain (ER2). Two DH domains (DH2 and DH3) in Subunit C resided also side by side and both shared sequence similarity to E. coli FabA. Overall, the PUFA synthase in Thraustochytrium sp. 26185 was highly homologous to those from other VLCPUFA-producing microbes. In addition, a discrete phosphopantetheinyl transferase (TsPPTase) with 206 amino acids required for attaching phosphopantetheine prosthetic group to the ACP domains was also identified in the genome.
Genomic Analysis of Genes Involved in the Assembly of VLCPUFA
Acyl-CoA synthetase (ACS) is a member of the ligase family that activates free fatty acids to their acyl-CoA forms, playing an important role in enabling free fatty acids to be incorporated into various glycerolipids. In the Thraustochytrium, five genes were identified encoding long chain acyl-CoA synthetase (Table 4). They all shared similar structure such as an AMP-binding domain “INYTSGTTGAPK” which was commonly found in members of the AMP-binding proteins such as acyl-CoA synthetase [28].
Once having been activated into the acyl-CoA form, newly synthesized fatty acids could be assembled into neutral lipids such as TAG, wax esters and sterol esters or polar phospholipids such as phosphatidylethanolamine (PtdEtn) and PtdCho using various acyltransferases. For instance, in the neutral glycerolipid biosynthesis pathway, a fatty acid was first acylated onto sn-1 position of glycerol-3-phosphate (G3P) by G3P acyltransferase (GPAT) to form lysophosphatidic acid (LysoPtdOH), which was further acylated by LysoPtdOH-acyltransferase (LPAT) to form phosphatidic acid (PtdOH). DAG was then formed via the dephosphorylation of PtdOH by phosphatidic acid phosphatase (PAP). Finally, TAG was synthesized by incorporation of an acyl chain to DAG catalyzed by DAG acyltransferase (DGAT) [29]. Structurally, these acyl transferases belonged to a membrane-bound O-acyltransferase (MBOAT) superfamily with the conserved motifs catalyzing acylation of an acyl-CoA into various acceptors such as G3P, LysoPtdOH, LysoPtdCho or alcohol [30]. In Thraustochytrium sp. 26185, five genes encoded GPAT and four genes encoded LPAT in the superfamily. In addition, six genes were identified as DGAT or possibly wax synthase (WS) from the superfamily. There was also one gene found to encode PAP (Table 5).
In Thraustochytrium sp. 26185, two main phospholipids of biological membranes, PtdEtn and PtdCho, could be synthesized by two pathways, backbone-activating pathway (CDP-DAG) and head-activating (de novo) pathway. In the backbone-activating pathway, CDP-DAG was firstly synthesized from PtdOH and CTP by CDP-DAG synthase (CDS). Two genes from this species were assumed to encode CDS as they both were identified as a member of CTP transferase family and shared 42 and 45 % similarity with CDS from Zea mays and Budvicia aquatica, respectively. Phosphatidylserine (PtdSer) could be synthesized from CDP-DAG by phosphatidylserine synthase (PSS). One gene from this species was identified to encode PSS, with 534 amino acids long sharing 37 % sequence identity with the PSS from Dictyostelium discoideum. PtdSer could be decarboxylated by phosphatidylserine decarboxylase (PSD) to produce PtdEtn. Two genes were identified to encode PSD with 318 and 421 amino acids, respectively. On the other hand, PtdEtn could go through three sequential methylation reactions catalyzed by PtdEtn methyltransferase (PEMT) to produce PtdCho. One gene in this species was identified to encode PEMT with 864 amino acids. The protein had two PEMT domains, a common feature in PEMT (Table 4).
In the head-activating pathway for PtdCho and PtdEtn biosynthesis, ethanolamine and choline was firstly activated by their kinases, ethanolamine kinase (EK) and choline kinase (CK), to form phosphoethanolamine and phosphocholine, respectively. Afterwards, they were incorporated with a CMP group by cytidylyltransferases (PECT and PCCT) to form CDP-ethanolamine and CDP-choline, respectively. The final step was catalyzed by ethanolamine phosphotransferase (EPT) or choline phosphotransferase (CPT) to convert CDP-ethanolamine and CDP-choline into PtdEtn and PtdCho, respectively. In Thraustochytrium sp. 26185, one gene was identified each encoding EK, CK, PECT and PCCT, respectively. Both ATP binding site and active site were found in EK and CK sequences [31], while “HXGH” and “KMSKS” motifs were found in PECT and PCCT sequences, which were highly conserved among the cytidylyltransferase family [32, 33]. There were two genes identified as either EPT or CPT in the genome with 385 and 423 amino acids long, respectively, and both shared 32 % sequence identity with those from Tetrahymena thermophile and Chrysochromulina sp., respectively (Table 4).
Lands cycle [34] plays an important role in the biosynthesis of glycerophospholipid in plants by using a lyso-phosphatidylcholine (LysoPtdCho) acyltransferase (LPCAT) for incorporation of freshly synthesized fatty acids to PtdCho [35]. In Thraustochytrium sp. 26185, there were two LPCAT of 556 and 612 amino acids for acylation of the sn-2 position of LysoPtdCho to form PtdCho. Both shared high amino acid identity (70 and 42 %) with LPCAT from Aurantiochytrium limacinum, with a conserved acyl-acceptor pocket and a calcium binding site.
In plants, acyl switching between PtdCho and DAG could be catalyzed by phosphatidylcholine: diacylglycerol cholinephosphotransferase (PDCT), whereby DAG with acyl chains modified on PtdCho could be assembled into TAG by DGAT [36]. Surprisingly, PDCT was not found in Thraustochytrium sp. 26185.
Discussion
Thraustochytrium sp. 26185 has long been known to accumulate a large amount of VLCPUFA in its storage lipids. The goal of this research was to survey the metabolic pathways for the biosynthesis and assembly of VLCPUFA in this species through genome sequencing and genomic analysis of genes involved in the processes. Genome sequencing produced a total of 2,418,734,139 bp clean sequences with more than 62 fold genome coverage. Annotation of the genome sequence revealed 10,797 coding genes. Among them, 10,216 genes could be assigned into 25 KOG classes where 451 genes were specifically assigned to the group of lipid transport and metabolism.
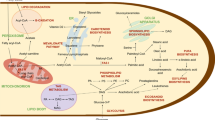
Analysis of these genes revealed that two pathways for the biosynthesis of VLCPUFA co-exist in this species. The aerobic pathway was more complex involving many different kinds of enzymes, such as fatty acid synthase for the synthesis of long chain saturated fatty acids (16:0 and 18:0), desaturases and elongases for introducing various double bonds towards both front and methyl ends and for elongating fatty acids with various chain lengths (Fig. 4). On the other hand, the anaerobic pathway is a de novo system with fewer chemical reactions using a single PUFA enzyme complex of three subunits each with multiple catalytic domains for the biosynthesis. Detailed analysis of genes in the aerobic pathway showed a key gene involved in the introduction of the first double bond into stearic acid synthesized by Type I FAS is missing from the genome, indicating this pathway is incomplete. This observation was recently confirmed by our in vivo feeding experiment (unpublished data). Co-existence of both aerobic and anaerobic pathways had been observed in Thraustochytrids [37, 38]. In Schizochytrium sp., a Δ12 desaturase activity is missing from the aerobic pathway, while in Thraustochytrium aureum, the aerobic pathway is intact. Similar to Thraustochytrium sp. 26185, the anaerobic pathway is believed to be primarily responsible for the biosynthesis of VLCPUFA in these two species. It is likely that the aerobic pathway is a progenitor system, while the anaerobic pathway is newly acquired, and after acquisition of this efficient system, the progenitor pathway becomes relic in the species. During evolution, different components in the aerobic pathway can be lost or retained in different species.
Genomic analysis of genes involved in the assembly of glycerolipids showed that freshly-synthesized VLCPUFA would be effectively incorporated into various phospholipids through both CDP-DAG pathway and head-activating pathway, as all the genes were found in the genome (Fig. 5). In addition, the flux of freshly-synthesized fatty acids can go through several pathways into storage glycerolipids. They can be esterified onto the sn-1 and sn-2 positions of PtdCho through acyl-editing, or be used to acylate G3P for the de novo synthesis of DAG, or be used for sn-3 acylation of DAG to generate TAG directly [39]. However, the relative flux efficiency of nascent VLCPUFA through these pathways is unknown in the Thraustochytrium. Acyl-editing of PtdCho is believed to be important in the acyl trafficking in plants where LPCAT plays an important role to shuffling freshly-synthesized saturated and monounsaturated fatty acids from chloroplast to PtdCho for further desaturation and other modification (hydroxylation and epoxidation for example) [40, 41]. In plants, de novo synthesized DAG starting from G3P is mainly used to synthesize PtdCho while PtdCho-derived DAG is the main precursor for the TAG synthesis generated either by PDCT or possibly reverse catalytic activity of CPT [39]. This mechanism warrants that fatty acids on TAG have already been modified on PtdCho. Intriguingly, the candidate gene for phospatidylcholine:diacylglycerol cholinetransferase (PDCT) to produce DAG from PtdCho was not detected in the genome sequence of Thraustochytrium sp. 26185. This implies that, unlike plants, this species might not use PtdCho-derived DAG generated by PDCT for the TAG biosynthesis. Instead, DAG precursor for the TAG biosynthesis might be either the de novo one from the Kennedy pathway or PtdCho-derived one generated by reverse activity of CPT, although the latter activity has not been absolutely confirmed in any species.
Abbreviations
- ACC:
-
Acyl-CoA carboxylase
- ACP:
-
Acyl carrier protein
- ACS:
-
Acyl-CoA synthetase
- ALA:
-
α-Linolenic acid
- ARA:
-
Arachidonic acid
- AT:
-
Acyl transferase
- CDS:
-
CDP-DAG synthase
- CK:
-
Choline kinase
- CPT:
-
Choline phosphotransferase
- DAG:
-
Diacylglycerol
- DGAT:
-
Diacylglycerol acyltransferase
- DH:
-
Dehydrase
- DHA:
-
Docosahexaenoic acid
- DPA:
-
Docosapentaenoic acid
- EK:
-
Ethanolamine kinase
- EPT:
-
Ethanolamine phosphotransferase
- ER:
-
Enoyl reductase
- FAS:
-
Fatty acid synthase
- G3P:
-
Glycerol-3-phosphate
- GLA:
-
γ-Linolenic acid
- GO:
-
Gene ontology
- GPAT:
-
Glycerol-3-phosphate acyltransferase
- KEGG:
-
Kyoto encyclopedia of genes and genomes
- KOG:
-
Eukaryotic orthologous groups
- KR:
-
Ketoacyl reductase
- KS:
-
Ketoacyl synthase
- LA:
-
Linoleic acid
- LysoPtdOH:
-
Lysophosphatidic acid
- LPAT:
-
Lysophosphatidic acid acyltransferase
- LysoPtdCho:
-
Lyso-phosphatidylcholine
- LPCAT:
-
Lyso-phosphatidylcholine acyltransferase
- MAT:
-
Malonyl-CoA ACP transacylase
- MBOAT:
-
Membrane-bound O-acyltransferase
- PAP:
-
Phosphatidic acid phosphatase
- PCCT:
-
Phosphocholine cytidylyltransferase
- PDCT:
-
Phosphatidylcholine:diacylglycerol cholinephosphotransferase
- PEMT:
-
Phosphatidylethanolamine methyltransferase
- PKS:
-
Polyketide synthase
- PPTase:
-
Phosphopantetheinyl transferase
- PSD:
-
Phosphatidylserine decarboxylase
- PSS:
-
Phosphatidylserine synthase
- PtdCho:
-
Phosphatidylcholine
- PtdEtn:
-
Phosphatidylethanolamine
- PtdOH:
-
Phosphatidic acid
- PtdSer:
-
Phosphatidylserine
- SDA:
-
Stearidonic acid
- TAG:
-
Triacylglycerol
- VLCPUFA:
-
Very long chain polyunsaturated fatty acid
- WS:
-
Wax synthase
References
Abedi E, Sahari MA (2014) Long-chain polyunsaturated fatty acid sources and evaluation of their nutritional and functional properties. Food Sci Nutr 2(5):443–463
Robinson LE, Mazurak VC (2013) N-3 polyunsaturated fatty acids: relationship to inflammation in healthy adults and adults exhibiting features of metabolic syndrome. Lipids 48(4):319–332
Guichardant M, Calzada C, Bernoud-Hubac N, Lagarde M, Véricel E (2015) Omega-3 polyunsaturated fatty acids and oxygenated metabolism in atherothrombosis. Biochimica et Biophysica Acta (BBA) Mol Cell Biol Lipids 1851(4):485–495
Wang W, Zhu J, Lyu F, Panigrahy D, Ferrara KW, Hammock B, Zhang G (2014) ω-3 Polyunsaturated fatty acids-derived lipid metabolites on angiogenesis, inflammation and cancer. Prostaglandins Other Lipid Mediat 113:13–20
Janssen CI, Kiliaan AJ (2014) Long-chain polyunsaturated fatty acids (LCPUFA) from genesis to senescence: the influence of LCPUFA on neural development, aging, and neurodegeneration. Prog Lipid Res 53:1–17
Qiu X (2003) Biosynthesis of docosahexaenoic acid (DHA, 22:6-4, 7,10,13,16,19): two distinct pathways. Prostaglandins Leukot Essent Fatty Acids 68(2):181
Lewis TE, Nichols PD, McMeekin TA (1999) The biotechnological potential of thraustochytrids. Mar Biotechnol 1:580–587
Weete JD, Kim H, Gandhi SR, Wang Y, Dute R (1997) Lipids and ultrastructure of Thraustochytrium sp. ATCC 26185. Lipids 32:839–845
Qiu X, Hong H, MacKenzie SL (2001) Identification of a Delta 4 fatty acid desaturase from Thraustochytrium sp. involved in the biosynthesis of docosahexanoic acid by heterologous expression in Saccharomyces cerevisiae and Brassica juncea. J Biol Chem 276(34):31561–31566
Metz JG, Roessler P, Facciotti D, Levering C (2001) Production of polyunsaturated fatty acids by polyketide synthases in both prokaryotes and eukaryotes. Science 293:4
Metz JG, Flatt JH, Kuner JM (2006) The genes for the enzymes of the polyunsaturated fatty acid polyketide synthase of Schizochytrium and their use in the manufacture of polyunsaturated fatty acids. International Patent Publication WO. 2006135866
Metz JG, Kuner J, Rosenzweig B, Lippmeier JC, Roessler P, Zirkle R (2009) Biochemical characterization of polyunsaturated fatty acid synthesis in Schizochytrium: release of the products as free fatty acids. Plant Physiol Biochem PPB/Societe Francaise de Physiologie Vegetale 47(6):472–478
Yang Y, Zhao H, Barrero RA, Zhang B, Sun G, Wilson IW, Xie F, Walker KD, Parks JW, Bruce R (2014) Genome sequencing and analysis of the paclitaxel-producing endophytic fungus Penicillium aurantiogriseum NRRL 62431. BMC Genom 15(1):69
DiGuistini S, Liao NY, Platt D, Robertson G, Seidel M, Chan SK, Docking TR, Birol I, Holt RA, Hirst M (2009) De novo genome sequence assembly of a filamentous fungus using Sanger, 454 and Illumina sequence data. Genome Biol 10(9):R94
Nielsen M, Schreiber L, Finster K, Schramm A (2015) Draft genome sequence of Bacillus azotoformans MEV2011, a (Co-) denitrifying strain unable to grow with oxygen. Stand Genom Sci 9(1):23
Soliai MM, Meyer SE, Udall JA, Elzinga DE, Hermansen RA, Bodily PM, Hart AA, Coleman CE (2014) De novo genome assembly of the fungal plant pathogen Pyrenophora semeniperda. PloS One 9(1):e87045
Ellwood SR, Liu Z, Syme RA, Lai Z, Hane JK, Keiper F, Moffat CS, Oliver RP, Friesen TL (2010) A first genome assembly of the barley fungal pathogen Pyrenophora teres f. teres. Genome Biol 11(11):R109
Wang Z, Fang B, Chen J, Zhang X, Luo Z, Huang L, Chen X, Li Y (2010) De novo assembly and characterization of root transcriptome using Illumina paired-end sequencing and development of cSSR markers in sweet potato (Ipomoea batatas). BMC Genom 11(1):726
Luo R, Liu B, Xie Y, Li Z, Huang W, Yuan J, He G, Chen Y, Pan Q, Liu Y (2012) SOAPdenovo2: an empirically improved memory-efficient short-read de novo assembler. Gigascience 1(1):18
Ter-Hovhannisyan V, Lomsadze A, Chernoff YO, Borodovsky M (2008) Gene prediction in novel fungal genomes using an ab initio algorithm with unsupervised training. Genome Res 18(12):1979–1990
Markowitz VM, Mavromatis K, Ivanova NN, Chen I-MA, Chu K, Kyrpides NC (2009) IMG ER: a system for microbial genome annotation expert review and curation. Bioinformatics 25(17):2271–2278
Stanke M, Steinkamp R, Waack S, Morgenstern B (2004) AUGUSTUS: a web server for gene finding in eukaryotes. Nucleic Acids Res 32(suppl 2):W309–W312
Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25(17):3389–3402
Conesa A, Götz S, García-Gómez JM, Terol J, Talón M, Robles M (2005) Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21(18):3674–3676
Götz S, García-Gómez JM, Terol J, Williams TD, Nagaraj SH, Nueda MJ, Robles M, Talón M, Dopazo J, Conesa A (2008) High-throughput functional annotation and data mining with the Blast2GO suite. Nucleic Acids Res 36(10):3420–3435
Damude HG, McGonigle B, Zhu QQ, Xue Z (2011) Delta-9-elongases and their use in making polyunsaturated fatty acids. Google Patents
Wu G, Truksa M, Datla N, Vrinten P, Bauer J, Zank T, Cirpus P, Heinz E, Qiu X (2005) Stepwise engineering to produce high yields of very long-chain polyunsaturated fatty acids in plants. Nat Biotechnol 23(8):1013–1017
He X, Chen GQ, Kang ST, McKeon TA (2007) Ricinus communis contains an acyl-CoA synthetase that preferentially activates ricinoleate to its CoA thioester. Lipids 42(10):931–938
Coleman R (2004) Enzymes of triacylglycerol synthesis and their regulation. Prog Lipid Res 43(2):134–176
Lewin TM, Wang P, Coleman RA (1999) Analysis of amino acid motifs diagnostic for the sn-glycerol-3-phosphate acyltransferase reaction. Biochemistry 38(18):5764–5771
Malito E, Sekulic N, Too WCS, Konrad M, Lavie A (2006) Elucidation of human choline kinase crystal structures in complex with the products ADP or phosphocholine. J Mol Biol 364(2):136–151
Lee J, Johnson J, Ding Z, Paetzel M, Cornell RB (2009) Crystal structure of a mammalian CTP: phosphocholine cytidylyltransferase catalytic domain reveals novel active site residues within a highly conserved nucleotidyltransferase fold. J Biol Chem 284(48):33535–33548
Saridakis V, Christendat D, Kimber MS, Dharamsi A, Edwards AM, Pai EF (2001) Insights into ligand binding and catalysis of a central step in NAD+ synthesis: structures of Methanobacterium thermoautotrophicum adenylyl transferase complexes. J Biol Chem 276(10):7225–7232
Lands WE (2000) Stories about acyl chains. Biochimica et Biophysica Acta (BBA) Mol Cell Biol Lipids 1483(1):1–14
Bates PD, Fatihi A, Snapp AR, Carlsson AS, Browse J, Lu C (2012) Acyl editing and headgroup exchange are the major mechanisms that direct polyunsaturated fatty acid flux into triacylglycerols. Plant Physiol 160(3):1530–1539
Lu C, Xin Z, Ren Z, Miquel M (2009) An enzyme regulating triacylglycerol composition is encoded by the ROD1 gene of Arabidopsis. Proc Natl Acad Sci 106(44):18837–18842
Matsuda T, Sakaguchi K, Hamaguchi R, Kobayashi T, Abe E, Hama Y, Hayashi M, Honda D, Okita Y, Sugimoto S (2012) Analysis of Δ12-fatty acid desaturase function revealed that two distinct pathways are active for the synthesis of PUFAs in T. aureum ATCC 34304. J Lipid Res 53(6):1210–1222
Lippmeier JC, Crawford KS, Owen CB, Rivas AA, Metz JG, Apt KE (2009) Characterization of both polyunsaturated fatty acid biosynthetic pathways in Schizochytrium sp. Lipids 44(7):621–630
Bates PD, Durrett TP, Ohlrogge JB, Pollard M (2009) Analysis of acyl fluxes through multiple pathways of triacylglycerol synthesis in developing soybean embryos. Plant Physiol 150(1):55–72. doi:10.1104/pp.109.137737
Bates PD, Ohlrogge JB, Pollard M (2007) Incorporation of newly synthesized fatty acids into cytosolic glycerolipids in pea leaves occurs via acyl editing. J Biol Chem 282(43):31206–31216. doi:10.1074/jbc.M705447200
Bates PD, Stymne S, Ohlrogge J (2013) Biochemical pathways in seed oil synthesis. Curr Opin Plant Biol 16(3):358–364. doi:10.1016/j.pbi.2013.02.015
Acknowledgments
This research was supported by Natural Sciences and Engineering Research Council of Canada. We also wish to acknowledge Jack Chen for technical assistance in the genome sequencing.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflicts of interest.
About this article
Cite this article
Zhao, X., Dauenpen, M., Qu, C. et al. Genomic Analysis of Genes Involved in the Biosynthesis of Very Long Chain Polyunsaturated Fatty Acids in Thraustochytrium sp. 26185. Lipids 51, 1065–1075 (2016). https://doi.org/10.1007/s11745-016-4181-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11745-016-4181-6