Abstract
Euglena gracilis, a unicellular phytoflagellate, can accumulate a large amount of medium-chain wax esters under anaerobic growth conditions. Here we report the identification and characterization of two genes involved in the biosynthesis of wax esters in E. gracilis. The first gene encodes a fatty acyl-CoA reductase (EgFAR) involved in the conversion of fatty acyl-CoAs to fatty alcohols and the second gene codes for a wax synthase (EgWS) catalyzing esterification of fatty acyl-CoAs and fatty alcohols, yielding wax esters. When expressed in yeast (Saccharomyces cerevisiae), EgFAR converted myristic acid (14:0) and palmitic acid (16:0) to their corresponding alcohols (14:0Alc and 16:0Alc) with myristic acid as the preferred substrate. EgWS utilized a broad range of fatty acyl-CoAs and fatty alcohols as substrates with the preference towards myristic acid and palmitoleyl alcohol. The wax biosynthetic pathway was reconstituted by co-expressing EgFAR and EgWS in yeast. When myristic acid was fed to the yeast, myristyl myristate (14:0–14:0), myristyl palmitoleate (14:0–16:1), myristyl palmitate (14:0–16:0) and palmityl myristate (16:0–14:0) were produced. These results indicate EgFAR and EgWS are likely the two enzymes involved in the biosynthesis of medium-chain wax esters in E. gracilis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Euglena gracilis is a unicellular phytoflagellate protist that can grow photoautotrophically in a minimal medium as well as heterotrophically in an organic carbon-rich medium. Under light or aerobic conditions, Euglena accumulates polysaccharides (mainly β-1,3 glucan, known as paramylon) and wax esters as energy reserves [1, 2]. Once the cell culture is switched from light to dark or from aerobic to anaerobic conditions, the polysaccharide reserve is converted to wax esters in the cytosol and the accumulated wax esters can reach up to 62% of the total lipid content [3]. Wax esters produced in Euglena are in a range of the 20- to 36-carbon chains comprised of saturated fatty acids and alcohols of 12–18 carbon chains with myristyl myristate (14:0–14:0) as the major species [4]. Under aerobic growth, Euglena mainly accumulates even-numbered wax esters, however an anaerobic growth condition promotes the accumulation of odd-numbered wax esters in some Euglena strains [3–6]. In addition to light and oxygen, other environmental factors, such as nutrients and temperature can also affect the production and composition of storage wax esters in Euglena cells. The production of wax esters is more efficient when the culture medium contains organic carbon source [7] or when the culture is shifted from low to high temperature (15 °C to 33 °C) [8]. Supplementation of the medium with unsaturated fatty acids (oleic acid, linoleic acid, linolenic acid) and unusual fatty acid (ricinoleic acid) leads to the accumulation of these fatty acids in both acyl and alcohol moieties of wax esters [6].
The wax ester biosynthetic pathway consists of two successive steps: conversion of fatty acyl-CoA to fatty alcohol and esterification of fatty acyl-CoA and fatty alcohol [9]. The enzymes responsible for these reactions are fatty acyl-CoA reductase (FAR) and acyl-CoA:fatty alcohol acyltransferase or wax synthase (WS), respectively. The genes encoding FARs have been identified from a variety of living organisms, including mouse [10], human [10], silkmoth [11], bean borer moth [12], jojoba [13], Arabidopsis [14, 15] and wheat [16]. BmFAR, a fatty acyl-CoA reductase from silkmoth (Bombyx mori), is responsible for production of sex pheromone bombykol, (E,Z)-10,12–hexadecadien-1-ol [11], whereas OsFARXIII, another fatty acyl-CoA reductase from bean borer moth (Ostrinia scapulalis), catalyzes the production of (Z)-11-tetradecenol, which can be further converted to acetate or aldehyde pheromones [12]. ScFAR, a fatty acyl-CoA reductase from jojoba (Simmondsia chinensis), is responsible for producing storage wax esters in developing seeds [13]. Arabidopsis fatty acyl-CoA reductase AtCER4 (At4g33790) is involved in the synthesis of cuticular wax lipids [15]. TaTAA1a, TaTAA1b and TaTAA1c, three orthologs of the jojoba ScFAR, were isolated from wheat [16]; they are involved in producing the lipid component in the outer pollen wall.
Wax synthases have also been isolated and characterized from a wide range of living organisms. Some show only wax synthase activity such as mouse WS, human WS [17], jojoba ScWS [18] and petunia PhWS1 [19], while the other exhibit both wax synthase and acyl-CoA:diacylglycerol acyltransferase (DGAT) activities, including Acinetobacter WS/DGAT [20] and Arabidopsis WSD1 [21]. Mouse and human WSs (MmWS and HsWS) are highly expressed in sebaceous-rich tissues, such as preputial glands and eyelids [17]. Jojoba ScWS was isolated from developing seeds and is involved in the synthesis of liquid waxes that accumulate in seeds [18]. Arabidopsis AtWSD1 (At5g37300) is involved in synthesis of stem epicuticular wax esters as shown by a severe reduction of the wax ester content (44 and 46 carbons) in Arabidopsis wsd1 mutants [21]. Petunia PhWS1 is highly expressed in petals and involved in the production of very long chain fatty acid esters of methyl, isoamyl and short to medium straight chain alcohols (4–12 carbons) [19].
In E. gracilis, the activities of both FAR and WS were found in the microsomal fraction of cultures grown in the dark [22]. The biochemical assays showed that FAR used 14:0, 16:0 and 18:0 as substrates and required NADH as a cofactor [22, 23]. Although the biosynthesis of wax esters in Euglena has been extensively studied, the genes encoding these enzymes have not been described. Here we report the identification and characterization of two genes, EgFAR and EgWS, involved in the biosynthesis of wax esters in E. gracilis. Heterologous expression of these genes in yeast revealed the unique properties of the enzymes—preferential utilization of medium chain fatty acyl and alcohol as substrates for synthesis of wax esters.
Experimental Procedures
Isolation of Putative Euglena Fatty Acyl-CoA Reductase (EgFAR) and Euglena Wax Synthase (EgWS)
Euglena gracilis was grown aerobically in a TSY medium (0.1 g/l sodium acetate, 0.1 g/l beef extract, 0.2 g/l pepto-tryptone, 0.2 g/l yeast extract, 0.2 µg vitamin B12, 1 µg/l biotin, 100 µg/l thiamine-HCl, 0.1 µg/l niacinamide, pH 7.0 [24]) at 25 °C for 16 h (120 μE m−2 s−1)/8 h (light/dark) regime. The Euglena cells can accumulate up to 28% wax ester under these conditions. Total RNA was extracted from the E. gracilis culture using TRIZOL® reagent (Invitrogen). The full length cDNAs of EgFAR and EgWS were obtained using Marathon™ cDNA Amplifcation kit under conditions detailed by the supplier (Clontech). The gene-specific primers used for 5′- and 3′-RACE of EgFAR are ACR1 (5′-GGCTGGTTGGAGTTGACGTAGCA-3′) and ACF1 (5′-GCCATGAACGATTTCTACGCGGG-3′) primers, respectively. The gene-specific primers used for 5′- and 3′-RACE of EgWS are WSR2 (5′-CTCCGGGTGACCTTTCGGC-3′) and WSF2 primers (5′-CCAGCCCTACTTTTCCACATCTCTGAG-3′), respectively.
DNA Sequencing and Analysis
All synthesis and sequencing work was performed by the DNA Technologies Unit at the Plant Biotechnology Institute, National Research Council of Canada. Nucleotide sequence and amino acid sequence comparisons were conducted using Lasergene7 (DNASTAR).
Amino Acid Sequence Alignment and Phylogenetic Analysis
Phylogenetic analysis of functionally characterized FARs and WSs were performed as previously described [25].
Construction of Yeast Expression Vectors Harboring EgFAR, EgWS and EgFAR-EgWS
The EgWS open reading frame (ORF) was amplified using primers WSFLF (5′-TTCGCGATGGATTTTTTGGGG-3′) and WSFLR (5′-GCCACCCAAGGCACTTGGCCT-3′) and cloned into pYES2.1/V5-His/lacZ vector (Invitrogen), yielding plasmid pPT504. The EgFAR ORF was amplified using primers PT0037 (5′-GATCGGATCCATGAACGATTTCTACGCG-3′)-PT0041 (5′-ATCAGCTAGCCTATCACAGCATGGCCCGC-3′) and digested with BamHI and NheI. The amplified fragment was subsequently cloned into the corresponding restriction sites of a yeast expression vector pESC-URA (Stratagene), yielding pPT515. The amplification was performed using 2.5 units of Platinum® Pfx DNA polymerase (Invitrogen) in the presence of 5% (v/v) DMSO. PCR conditions for EgFAR were 35 cycles of 94 °C for 15 s, 55 °C for 30 s, 68 °C for 1 min with the final extension at 68 °C for 5 min. PCR conditions for EgWS were identical to EgFAR, except that the extension time was 30 s instead of 1 min. For the co-expression study, the EgWS ORF was amplified from pPT504 using primers PT0049 (5′-GATCATCGATATGGATTTCTTAGGTTTTCCTGAC-3′)-PT0050 (5′-CTGTAGATCTCTATCAGACAGACAGACCTAGC-3′). The amplified fragment was digested with ClaI and BamHI and subsequently cloned into the corresponding sites of pPT515, yielding pPT516.
Functional Analysis of EgFAR and EgWS in Yeast
pPT515 (EgFAR) and pPT504 (EgWS) plasmids were transformed into yeast (Saccharomyces cerevisiae) strain H1246 (MATα; are1-Δ::HIS3 are2-Δ::LEU2 dga1-Δ::KanMX4 lro1-Δ::TRP1 ADE2) [26] using S.c. EasyComp™ transformation kit (Invitrogen). Yeast strains transformed with pPT515 (EgFAR), pPT504 (EgWS), pPT516 (EgFAR + EgWS), pESC-URA or pYES2.1 plasmids were grown at 30 °C for 2 days in 10 ml of the synthetic dropout medium containing 0.17% (w/v) yeast nitrogen base, 0.5% (w/v) ammonium sulfate, 2% (w/v) glucose and 0.06% (w/v) dropout supplement lacking uracil (DOB + GLU-URA). After two washes with 10 ml of sterile distilled water, the expression of transgene in yeast were induced by culturing the yeast at 20 °C for 4 days for pPT515 (EgFAR) and 30 °C for 2 days for pPT504 (EgWS) in 10 ml of the synthetic dropout medium containing 2% (w/v) galactose (DOB + GAL-URA) with or without substrate supplementation in the presence of 0.1% (v/v) tergitol (Nonidet P-40). Two-hundred fifty micromolar of fatty acid was used as a substrate for the pPT515 (EgFAR) expression, whereas 250 μM of fatty acid and 250 μM of fatty alcohol were used as substrates for the pPT504 (EgWS) expression. After induction, the cultures were washed once with 10 ml of 1% (v/v) tergitol and once with 10 ml of distilled water, and then subjected to fatty acid analysis. For the yeast feeding experiment, fatty acid and fatty alcohol substrates were initially prepared as stock solutions in ethanol at the concentration of 500 mM and the appropriate amount of the stock was then diluted in 10% tergitol at the final concentration of 50 mM used for feeding the yeast.
In Vitro Assays of EgFAR and EgWS
To investigate substrate specificity of EgFAR and EgWS in vitro, the microsomal fractions of yeast expressing EgFAR and EgWS were prepared as previously described [27, 28]. For the EgFAR assay, the enzyme reaction (500 μl) contained 200 μg of microsomal proteins, 0.3 M sucrose, 0.1 M MOPS (pH 6.5) or 0.1 M Tris–HCl, (pH 7.4), 1 mM EDTA, 2.5 mM DTT, 5 mM MgCl2, 1 mM PMSF, 100 μM acyl-CoA, 2.5 mM NADH, 2.5 mM NADPH. The reaction was carried out at 30 °C with gentle shaking for 30 min to 2 h. For the EgWS assay, the same amount of protein and the reaction buffer (pH 7.4) were used. One-hundred micromolar of acyl-CoA and 100 μM fatty alcohol were added as substrates. The reaction was carried out at 30 °C with gentle shaking for 2 h. The enzyme reactions were terminated by addition of 100 μl of 6 M HCl. After adding 1 ml of phosphate buffered saline and 1 ml of 0.9% (w/v) NaCl solution, lipids were extracted twice with 2 ml hexane. The known amount of methyl eicosanoate (20:0-ME) was added as the internal standard. The solvent was subsequently removed under a nitrogen stream and the lipid samples were resuspended in 20 μl of hexane, which was derivatized with bis(trimethylsilyl)-acetamide (TMS) and analyzed by GC for the EgFAR assay or was directly analyzed by GC for the EgWS assay (see below). Acyl-CoA stock solutions were prepared in 10 mM sodium acetate buffer (pH5.2)/ethanol (1:1, v/v) at a concentration of 50 mM.
Co-Expression of EgFAR and EgWS in Yeast
To reconstitute the Euglena wax biosynthesis pathway in yeast, the plasmid pPT516 harboring the two-gene cassette (EgFAR and EgWS) was transformed into the yeast strain H1246. The transformants were grown in the DOB + GLU-URA medium at 30 °C for 2 days and the expression of EgFAR and EgWS was induced in the DOB + GAL-URA medium supplemented with 500 μM 14:0 in the presence of 0.1% (v/v) tergitol at 20 °C for 4 days. The yeast cells were harvested and analyzed for fatty alcohol and wax monoester production as described above.
Analysis of Fatty Acids, Fatty Alcohols and Wax Esters of Yeast Transformants
Fatty acids of yeast cells were transmethylated with 2 ml of methanol/HCl (3 M) at 80 °C for 2 h and the reaction was terminated by adding 1 ml of 0.9% NaCl solution. Total fatty acid methyl esters (FAMEs) and fatty alcohols were then extracted twice with 2 ml of hexane and the hexane phase was transferred to a new tube, evaporated under a nitrogen stream and resuspended in 200 μl of hexane. Fifty microliters of samples were derivatized with 50 μl of TMS/pyridine (1:1, v/v) at 80 °C for 30 min and the derivative was analyzed by gas chromatography (GC). For wax ester analysis, the total lipid was extracted by homogenizing yeast cells in the presence of 6 ml of chloroform/methanol (2:1, v/v). The organic phase was dried under a stream of nitrogen gas, resuspended in 50 μl of hexane and analyzed by GC, GC–mass spectrophotometry (MS) and thin layer chromatography (TLC).
GC, GC–MS and TLC Analysis
Fatty acids (10:0, 12:0, 14:0, 16:0, 16:1n-9, 18:0, and 18:1n-9), fatty alcohols (10:0Alc, 12:0Alc, 14:0Alc, 16:0Alc, 16:1n-9Alc, 18:0Alc and 18:1n-9Alc), wax esters (14:0–12:0, 14:0–14:0, 14:0–16:0, 14:0–16:1n-9, 14:0–18:0, 14:0–18:1n-9, 12:0–14:0, 16:0–14:0, 16:1n-9–14:0, 18:0–14:0, 18:1n-9–14:0) and TLC reference standard (cholesterol, cholesteryl oleate, triolein, oleic acid, methyl oleate) with 99% purity were purchased from Nuchek-Prep, Inc. For GC analysis, one-microliter samples were analyzed on an Agilent 6890 N GC equipped with a DB-5 column (10 m × 0.25 mm) (J&W Scientific). The following temperature programs were employed: 70 °C for 1 min, then 10 °C/min to 300 °C, and 300 °C for 10 min with H2 as the carrier gas. For MS analysis, the mass selective detector was run under standard electron impact conditions (70 eV), scanning an effective m/z range of 40–700 at 2.26 scans/s. Identities of FAMEs, fatty alcohols-TMS derivatives and wax esters were identified by comparing their retention times with those of the standards and confirmed by GC–MS on the basis of their fragmentation patterns. For TLC analysis, total lipid samples and standards were spotted on 60-Å silica gel SIL G-25 plates (Macherey–Nagel) and resolved in hexane/diethylether/acetic acid (90:7.5:1, v/v/v) [21]. The plate was air-dried and the lipid metabolites were detected by primuline spraying (0.05% w/v; Sigma) and the pictures were taken under UV light at 254 nm.
Results
Isolation of Two cDNAs Encoding Putative Fatty acyl-CoA Reductase and Wax Synthase from E. gracilis
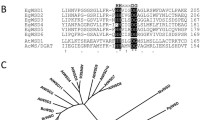
The partial cDNA sequences of Euglena putative fatty acyl-CoA reductase EgFAR and wax synthase EgWS were obtained by a homology search of the Euglena EST database using jojoba ScFAR and ScWS as query sequences and the full-length putative EgFAR and EgWS cDNAs were obtained by 5′- and 3′-RACE using Euglena cDNAs as the template. Sequence analysis indicated putative EgFAR encodes a polypeptide of 514 amino acids in length with the predicted molecular mass of 56.5 kDa. The deduced protein EgFAR contains a Rossmann-fold NAD/NADP binding domain (NABD; 289 amino acids) [29] linked with a Male Sterile 2 domain (MS2, 97 amino acids) [30] at the carboxyl end (Fig. 1a). A conserved motif (I/V/F)-X-(I/L/V)-T-G-X-T-G-F-L-(G/A), found in other fatty acyl-CoA reductases, was also observed in the NABD domain of EgFAR (Fig. 1b). The hydropathy analysis indicated that the putative EgFAR contains five high hydrophobic regions; one located at the N-terminus and the rest present in the central portion of the protein. Phylogenetic analysis of the putative EgFAR and related sequences reveals that EgFAR is more closely related to plant fatty acyl-CoA reductases, including Arabidopsis AtCER4 [15], wheat TaTAA1 [16] and jojoba ScFAR [31], whereas the insect and mammalian fatty acyl-CoA reductases [10] form a distant group (Fig. 2a). Sequence analysis of the putative EgWS indicated that it encodes a polypeptide of 368 amino acids with the predicted molecular mass of 41.2 kDa. The deduced protein EgWS contains six distinct hydrophobic regions. Phylogenetic analysis of EgWS and related sequences reveals that Euglena EgWS clusters with jojoba ScWS [18], which is distantly related to the rest members of wax synthases, including the mammalian WSs [17], Acinetobacter AcWS/DGAT [20], Arabidopsis AtWSD1 [21] and petunia PhWS1 [19] (Fig. 2b).
A schematic diagram illustrating the functional domains (a) and a conserved motif (b) of EgFAR and other related sequences. The numbers indicate amino acid positions. The accession numbers of these sequences are indicated in Fig. 2
Phylogenetic analysis of EgFAR (a) and EgWS (b) and their related sequences. The GenBank accession numbers of the sequences are as follows: mouse MmFAR1, BC007178; mouse MmFAR2, BC055759; human HsFAR1, AY600449; human HsFAR2, BC022267; silkmoth BmFAR, AB104896; borer moth OsFARXIII, EU817405; wheat FAR (TaTAA1a), AJ459249; jojoba ScFAR, AF149917; Arabidopsis AtCER4, AY070065-At4g33790; mouse MmWS, AY611032; human HsWS, AY605053; Arabidopsis AtWSD1, NM_123089-AT5G37300; Acinetobacter AcWS/DGAT, AF529086; jojoba ScWS, AF149919; petunia PhWS1, DQ093641
Functional Analysis of EgFAR and EgWS in Yeast
To determine the function of putative EgFAR and EgWS, the full-length cDNAs were cloned into the yeast expression vectors separately and the recombinant plasmids were transformed into the Saccharomyces cerevisiae quadruple mutant strain H1246 [26], in which four acyltransferase genes involved in the triacylglycerol (TAG) and sterol ester biosynthesis were disrupted. This strain appeared to have diminished capacity for wax ester biosynthesis (Fig. 3b), thus can serve as a good host system to examine the function of genes involved in the biosynthesis of wax esters.
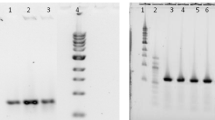
Activity of EgFAR (a) and EgWS (b, c) in yeast. a GC analysis of fatty acid methyl esters and fatty alcohol TMS ethers prepared from the yeast transformed with the pESC-URA vector control or EgFAR. b TLC analysis of the total lipid of the yeast transformed with the pYES2.1 vector control (lanes 5 and 6) or EgWS (lanes 7 and 8) in the absence (lanes 5 and 7) and presence (lane 6 and 8) of substrates (14:0 and 14:0Alc). Lane 1–4, 9 are standards: DAG diacylglycerol, TAG triacylglycerol (c) GC analysis of the total lipid prepared from yeast transformed with EgWS or the pYES2.1 vector control and fed with 14:0 and 14:0Alc substrates
The activity of the putative Euglena fatty acyl-CoA reductase was investigated by feeding the yeast strain carrying EgFAR with a variety of probable fatty acid substrates. Analysis of total fatty alcohols in the cells indicated that the transformant expressing EgFAR produced new peaks only when myristic acid (14:0) and palmitic acid (16:0) were used as substrates, compared with the vector control (Fig. 3a). The chemical structures of these new peaks were determined as myristyl alcohol (14:0Alc) and palmityl alcohol (16:0Alc) by GC–MS. The conversion efficiencies of the two alcohols were 34 ± 4 and 24 ± 3%, respectively. No other alcohol products were detected when the yeast strain was fed with other saturated fatty acids (10:0, 12:0 and 18:0) or unsaturated fatty acids (16:1n-9, 18:1n-9 and 18:2n-6). A similar result was obtained when EgFAR was expressed in yeast strain INVSc.
The activity of the putative EgWS was investigated by feeding the yeast strain expressing EgWS with 14:0 and 14:0Alc substrates. The total neutral lipids including triacylglycerols and wax esters in the transformant were first analyzed by TLC. In contrast to the vector control, the yeast expressing EgWS produced wax esters in presence of 14:0 and 14:0Alc that were detected on the TLC plate (Fig. 3b). Like the control, the transformant expressing EgWS did not produce any triacylglycerols, indicating EgWS possess wax synthase activity, but not diacylglycerol acyltransferase activity as seen in bacterial WS/DGAT [32]. Wax composition analysis by GC and GC–MS indicated that the major components of wax esters produced in the transformant were myristyl myristate (14:0–14:0), myristyl palmitate (14:0–16:0) and myristyl pamitoleate (14:0–16:1) (Fig. 3c). It was also noted that the vector control could produce a trace amount of myristyl myristate (14:0–14:0) when the yeast was fed with substrates (Fig. 3b, c).
To examine the fatty acid substrate specificity of EgWS, the yeast strain was fed with 14:0Alc in combination with a range of fatty acids including 10:0, 12:0, 14:0, 16:0, 16:1n-9,18:0, and 18:1n-9. The wax analysis showed that EgWS could incorporate 12:0, 14:0, 16:0 and 16:1–9 fatty acids into wax esters, but it could not use 10:0, 18:0 or 18:1n-9 as the substrate. Quantitative analysis of the ratio of products versus substrates indicated that the highest conversion efficiency of fatty acids was observed on 14:0, which was followed by 12:0, 16:0 and 16:1n-9 (Table 1). In order to examine fatty alcohol substrate specificity, the yeast strain was fed with 14:0, the most preferred acyl-CoA substrate for EgWS in combination with a range of fatty alcohols including 10:0Alc, 12:0Alc, 14:0Alc, 16:0Alc, 16:1n-9Alc, 18:0Alc and 18:1n-9Alc. Wax analysis indicated that EgWS could utilize 12:0Alc, 14:0Alc, 16:0Alc and 16:1n-9Alc fatty alcohols, but not 10:0Alc, 18:0Alc and 18:1n-9Alc as the substrates. The preferred fatty alcohol was 16:1n-9Alc, which was followed by 14:0Alc, 12:0Alc and 16:0Alc (Table 1).
To study the substrate specificity of EgFAR and EgWS in vitro, the microsomal fractions of yeast expressing EgFAR or EgWS were incubated with a series of fatty acids or fatty acid and alcohol combinations. However, only small activity of wax ester synthesis was detected in the in vitro assay of EgWS, which was basically in agreement with the result from the in vivo experiment. Furthermore, we could not detect any fatty acyl-CoA reductase activity in the in vitro assay of EgFAR with all possible substrates. The reason why the in vitro EgWS and EgFAR activities in yeast were low is unknown. It is noteworthy that so far only three reports have described the in vitro fatty acyl-CoA reductase and wax synthase activities in heterologous systems, including mammalian FARs and WSs in the HEK293 human cell line [10, 17] and Acinetobacter WS/DGAT in E. coli [20]. The difficulty in the yeast in vitro assays of EgFARs and EgWSs might be related to the nature of this type of enzymes or simply the low activities of the two enzymes in yeast.
Reconstitution of the Euglena Wax Biosynthetic Pathway in Yeast
To reconstitute the Euglena wax biosynthetic pathway in yeast, EgFAR and EgWS were cloned into a yeast expression vector, pESC-URA, in which the expression of EgFAR and EgWS were driven by GAL1 and GAL10 inducible promoters, respectively. The recombinant plasmid was then transformed into the yeast strain H1246 and the transformants were grown in the selective medium supplemented with 14:0, a common preferred substrate for both EgFAR and EgWS. Wax ester analysis indicated that the co-expressing yeast, like the EgFAR-expressing yeast, could convert 14:0 and 16:0 to 14:0Alc and 16:0Alc, respectively (Fig. 4). In addition, the co-expressing yeast produced three new wax ester peaks which were not detected in either EgFAR-expressing yeast or vector control yeast. Identities of these peaks were determined by comparing their retention times and mass spectra to those of standards. Peak 1 had the identical retention time and mass spectrum to myristyl myristate (14:0–14:0) which accounted for ~23% of total wax esters (Fig. 5a) while peak 2 had the identical retention time and mass spectrum to myristyl palmitoleate (14:0–16:1) (Fig. 5b) which accounted for ~22% of total wax esters. Peak 3 appeared to contain two wax ester products, myristyl palmitate (14:0–16:0) and palmityl myristate (16:0–14:0) together they accounted for ~55% of total wax esters. GC and GC–MS analysis showed that it had the same retention time as myristyl palmitate (14:0–16:0) and palmityl myristate (16:0–14:0) and possessed the mass spectrum (Fig. 5c) with diagnostic ions representing the mass spectra of both myristyl palmitate (14:0–16:0) (Fig. 5d) and palmityl myristate (16:0–14:0) (Fig. 5e). Based on the abundance of the diagnostic fragments and the relative response factors of 14:0–16:0 (m/z 257) and 16:0–14:0 (m/z 229), peak 3 contained ~58% of 14:0–16:0 and ~41% of 16:0–14:0. Collectively, these results indicated that EgWS in the co-expressing yeast could esterify 14:0Alc and 16:0Alc produced by EgFAR with 14:0, 16:0 and 16:1n-9 fatty acids, producing medium chain wax esters, myristyl myristate (14:0–14:0), myristyl palmitate (14:0–16:0), myristyl palmitoleate (14:0–16:1) and palmityl myristate (16:0–14:0).
Co-expression of EgFAR and EgWS in yeast. GC analysis of the total lipid showing wax ester production prepared from yeast transformed with pESC-URA (empty vector), EgFAR and EgFAR-EgWS and fed with 14:0 substrate. Peak 1 myristyl-myristate (14:0–14:0), Peak 2 myristyl-palmitoleate (14:0–16:1), Peak 3 myristyl palmitate (14:0–16:0) and palmityl myristate (16:0–14:0). Selected regions of the chromatograms of control, EgFAR and EgFAR-EgWS expressing yeast are magnified in boxes A, B and C, respectively
GC/MS analysis of wax ester peak 1 a, peak 2 b and peak 3 c in Fig. 4, myristyl palmitate (14:0–16:0) standard (d) and palmityl myristate (16:0–14:0) standard e. Asterisks indicate diagnostic ions of the wax esters
Discussion
The biosynthesis of wax esters comprises two consecutive catalytic steps, reduction of fatty acyl-CoA to fatty alcohol and subsequent esterification of fatty acyl-CoA and fatty alcohol. The reduction of fatty acid to fatty alcohol occurs through aldehyde intermediate and is catalyzed by either one or two enzyme reactions. In the two-enzyme reaction, acyl-CoA reductase first converts the fatty acyl-CoA to aldehyde and then the aldehyde reductase catalyzes the reduction of the aldehyde to alcohol. Evidence supporting the two-step reaction came from the early identification of aldehyde reductase from Brassica oleracea [33] and the recent isolation of the acyl-CoA reductase gene from Acinetobacter, catalyzing the production of fatty aldehyde [34]. However, support for the existence of the two-enzyme reduction is scarce in eukaryotes. In fact, a single enzyme reaction for the fatty acid reduction appears to occur widely in nature where the FAR catalyzes reduction of fatty acyl-CoA to fatty alcohol directly, although aldehyde intermediates could be detected by indirect trapping assays [23]. This type of fatty acyl-CoA reductases has been identified from jojoba cotyledon, pea leaves [23, 35, 36] and many others [37–39], including the one we describe here from Euglena.
Although E. gracilis has long been known to accumulate a large amount of medium-chain wax esters under dark and anaerobic growth conditions, the genes involved in the biosynthesis of these wax esters have yet to be identified. In this study, we report the isolation and characterization of two cDNAs, EgFAR and EgWS, from E. gracilis encoding two enzymes involved in the biosynthesis of medium chain wax esters. The pairwise sequence comparison of EgFAR and previously identified FARs revealed that EgFAR shares low amino acid identity with other FARs (23–27.4%). Functional analysis of EgFAR in yeast indicated that it could effectively convert 14:0 and 16:0 fatty acids to their corresponding alcohols. Compared with other biochemically characterized FARs, EgFAR possesses a narrower substrate range, only using saturated fatty acids with 14 and 16 carbon chains as substrates with the preferred fatty acid being 14:0 when expressed in yeast.
The pairwise sequence comparison of EgWS and related wax synthases revealed that EgWS also shares low amino acid identity to its related enzymes (in the range of 13–27%). It was noted that EgWS does not contain an N-terminal domain with the proposed active site (HHXXXD) found in Acinetobacter AcWS/DGAT [20], Arabidopsis AtWSD1 [21] and petunia PhWS1 [19]. This domain was believed to be essential for their acyl-CoA acyltransferase activities for the synthesis of wax esters and TAGs [20]. It will be interesting to know how EgWS functions without this domain. To date, three wax synthases including EgWS we report here have been functionally characterized in yeast. AtWSD1 mainly synthesizes wax esters with 16:0 fatty acid and 18:0Alc, 24:0Alc and 28:0Alc alcohols, and PhWS1 produces wax esters with very long chain fatty acids and methyl, isoamyl short to medium chain alcohols (4–12 carbons), whereas EgWS prefers 14 carbon chain fatty acid as the substrate.
Wax esters consisting of medium chain saturated fatty alcohols are sporadically found in animals and microorganisms [40–43]. E. gracilis can accumulate up to 28 or 62% of the total lipid as wax esters when grown in aerobic or anaerobic conditions [3] and the major molecular species of the wax esters accumulated is myristyl myristate [4, 6]. Functional expression of EgFAR and EgWS separately in yeast revealed that both enzymes could use 14 and 16 fatty acid substrates and the co-expression of EgFAR and EgWS in yeast resulted in production of medium-chain wax esters. These results are consistent with the composition of wax esters naturally present in Euglena, suggesting an important role of EgFAR and EgWS in synthesizing the wax esters in Euglena. EgFAR and EgWS were identified by the homology search of an EST database using jojoba ScFAR and ScWS as query sequences and only one EST of each EgFAR and EgWS was found in the database. However, it could not be excluded that there are additional genes involved in the biosynthesis of medium-chain wax esters in Euglena that were not present in the EST database. It should also be noted that Euglena can accumulate unsaturated fatty acid (e.g. 18:1n-9) wax esters when the culture medium is supplemented with such fatty acid [6]. Although we have not observed any activity of EgFAR and EgWS towards 18:1n-9 when they were expressed in yeast, this should not exclude the possibility that the difference of the cellular micro-environment between the native host and yeast could have effect on the substrate specificity of the enzymes. Alternatively, there might be additional FARs and WSs present in Euglena that can utilize unsaturated fatty acids as substrates.
Medium- and long-chain wax esters have been widely used in food, pharmaceuticals, textiles, perfumes and flavoring. They can also be used in the production of fatty sulfate salts and alcohol ethoxylates in the detergent industry. The current supply of wax from natural sources is limited due to the high production cost and cannot meet the growing demand for its widespread uses. Metabolic engineering of oilseed plants to produce wax esters has been viewed as an attractive alternative to provide cost-effective sources for biological wax. The first attempt of wax metabolic engineering in plants was undertaken by expressing jojoba ScFAR and ScWS, along with Lunaria annua β-ketoacyl-CoA synthase (a component of fatty acid elongase) resulting in the production of very long chain wax esters in Brassica napus seeds [18]. Here, we describe two genes EgFAR and EgWS from Euglena encoding fatty acyl-CoA reductase and wax synthase that have a substrate preference towards medium chain substrates. It will be interesting to see how these genes perform in the production of industrially important medium-chain wax esters in oil seed crops.
Abbreviations
- CoA:
-
Coenzyme A
- FAMEs:
-
Fatty acid methyl esters
- FAR:
-
Fatty acyl-CoA reductase
- RACE:
-
Rapid amplification of cDNA ends
- WS:
-
Wax synthase
References
Inui H, Miyatake K, Nakano Y, Kitaoka S (1982) Wax ester fermentation in Euglena gracilis. FEBS Lett 150:89–93
Rosenberg A (1963) A comparison of lipid patterns in photosynthesizing and non-photosynthesizing cells of Euglena gracilis. Biochemistry 2:1148–1154
Tucci S, Vacula R, Krajcovic J, Proksch P, Martin W (2009) Variability of wax ester fermentation in natural and bleached Euglena gracilis strains in response to oxygen and the elongase Inhibitor Flufenacet. J Eukaryot Microbiol 57:63–69
Koritala S (1989) Microbiological synthesis of wax esters by Euglena gracilis. J Am Oil Chem Soc 66:133–134
Schneider T, Betz A (1985) Waxmonoester fermentation in Euglena gracilis T. Factors favouring the synthesis of odd-numbered fatty acids and alcohols. Planta 166:67–73
Tani Y, Okumura M, Li S (1987) Liquid wax ester production by Euglena gracilis. Agric Biol Chem 51:225–230
Regnault A, Chervin D, Chammai A, Pitona F, Calvayraca R, Mazliakb P (1995) Lipid composition of Euglena gracilis in relation to carbon–nitrogen balance. Phytochemistry 40:725–733
Kawabata A, Inui H, Miyatake K, Nakano Y, Kitaoka S (1990) Production and composition of Euglena wax esters at high temperature. Agric Biol Chem 54:811–812
Samuels L, Kunst L, Jetter R (2008) Sealing plant surfaces: cuticular wax formation by epidermal cells. Annu Rev Plant Biol 59:683–707
Cheng JB, Russell DW (2004) Mammalian wax biosynthesis. I. Identification of two fatty acyl-coenzyme A reductases with different substrate specificities and tissue distributions. J Biol Chem 279:37789–37797
Moto K, Yoshiga T, Yamamoto M, Takahashi S, Okano K, Ando T, Nakata T, Matsumoto S (2003) Pheromone gland-specific fatty-acyl reductase of the silkmoth, Bombyx mori. Proc Natl Acad Sci USA 100:9156–9161
Antony B, Fujii T, Moto K, Matsumoto S, Fukuzawa M, Nakano R, Tatsuki S, Ishikawa Y (2009) Pheromone-gland-specific fatty-acyl reductase in the adzuki bean borer, Ostrinia scapulalis (Lepidoptera: Crambidae). Insect Biochem Mol Biol 39:90–95
Miwa T (1971) Jojoba oil wax esters and derived fatty acids and alcohols: gas chromatographic analyses. J Am Oil Chem Soc 48:259–264
Doan TT, Carlsson AS, Hamberg M, Bulow L, Stymne S, Olsson P (2008) Functional expression of five Arabidopsis fatty acyl-CoA reductase genes in Escherichia coli. J Plant Physiol 166:787–796
Rowland O, Zheng H, Hepworth SR, Lam P, Jetter R, Kunst L (2006) CER4 encodes an alcohol-forming fatty acyl-coenzyme A reductase involved in cuticular wax production in Arabidopsis. Plant Physiol 142:866–877
Wang A, Xia Q, Xie W, Dumonceaux T, Zou J, Datla R, Selvaraj G (2002) Male gametophyte development in bread wheat (Triticum aestivum L.): molecular, cellular, and biochemical analyses of a sporophytic contribution to pollen wall ontogeny. Plant J 30:613–623
Cheng JB, Russell DW (2004) Mammalian wax biosynthesis. II. Expression cloning of wax synthase cDNAs encoding a member of the acyltransferase enzyme family. J Biol Chem 279:37798–37807
Lardizabal KD, Metz JG, Sakamoto T, Hutton WC, Pollard MR, Lassner MW (2000) Purification of a jojoba embryo wax synthase, cloning of its cDNA, and production of high levels of wax in seeds of transgenic Arabidopsis. Plant Physiol 122:645–655
King A, Nam JW, Han J, Hilliard J, Jaworski JG (2007) Cuticular wax biosynthesis in petunia petals: cloning and characterization of an alcohol-acyltransferase that synthesizes wax-esters. Planta 226:381–394
Kalscheuer R, Steinbuchel A (2003) A novel bifunctional wax ester synthase/acyl-CoA: diacylglycerol acyltransferase mediates wax ester and triacylglycerol biosynthesis in Acinetobacter calcoaceticus ADP1. J Biol Chem 278:8075–8082
Li F, Wu X, Lam P, Bird D, Zheng H, Samuels L, Jetter R, Kunst L (2008) Identification of the wax ester synthase/acyl-coenzyme A: diacylglycerol acyltransferase WSD1 required for stem wax ester biosynthesis in Arabidopsis. Plant Physiol 148:97–107
Khan AA, Kolattukudy PE (1973) Control of synthesis and distribution of acyl moieties in etiolated Euglena gracilis. Biochemistry 12:1939–1948
Kolattukudy PE (1970) Reduction of fatty acids to alcohols by cell-free preparations of Euglena gracilis. Biochemistry 9:1095–1102
Schlösser UG (1994) SAG—Sammlung von Algenkulturen at the University of Göttingen, catalogue of strains. Bot Acta 107:113–186
Meesapyodsuk D, Reed DW, Covello PS, Qiu X (2007) Primary structure, regioselectivity, and evolution of the membrane-bound fatty acid desaturases of Claviceps purpurea. J Biol Chem 282:20191–20199
Sandager L, Gustavsson MH, Stahl U, Dahlqvist A, Wiberg E, Banas A, Lenman M, Ronne H, Stymne S (2002) Storage lipid synthesis is non-essential in yeast. J Biol Chem 277:6478–6482
Katavic V, Mietkiewska E, Barton DL, Giblin EM, Reed DW, Taylor DC (2002) Restoring enzyme activity in nonfunctional low erucic acid Brassica napus fatty acid elongase 1 by a single amino acid substitution. Eur J Biochem 269:5625–5631
Li R, Reed DW, Liu E, Nowak J, Pelcher LE, Page JE, Covello PS (2006) Functional genomic analysis of alkaloid biosynthesis in Hyoscyamus niger reveals a cytochrome P450 involved in littorine rearrangement. Chem Biol 13:513–520
Rossmann MG, Moras D, Olsen KW (1974) Chemical and biological evolution of nucleotide-binding protein. Nature 250:194–199
Aarts MG, Hodge R, Kalantidis K, Florack D, Wilson ZA, Mulligan BJ, Stiekema WJ, Scott R, Pereira A (1997) The Arabidopsis MALE STERILITY 2 protein shares similarity with reductases in elongation/condensation complexes. Plant J 12:615–623
Metz JG, Pollard MR, Anderson L, Hayes TR, Lassner MW (2000) Purification of a jojoba embryo fatty acyl-coenzyme A reductase and expression of its cDNA in high erucic acid rapeseed. Plant Physiol 122:635–644
Kalscheuer R, Luftmann H, Steinbuchel A (2004) Synthesis of novel lipids in Saccharomyces cerevisiae by heterologous expression of an unspecific bacterial acyltransferase. Appl Environ Microbiol 70:7119–7125
Kolattukudy PE (1971) Enzymatic synthesis of fatty alcohols in Brassica oleracea. Arch Biochem Biophys 142:701–709
Reiser S, Somerville C (1997) Isolation of mutants of Acinetobacter calcoaceticus deficient in wax ester synthesis and complementation of one mutation with a gene encoding a fatty acyl coenzyme A reductase. J Bacteriol 179:2969–2975
Vioque J, Kolattukudy PE (1997) Resolution and purification of an aldehyde-generating and an alcohol-generating fatty acyl-CoA reductase from pea leaves (Pisum sativum L.). Arch Biochem Biophys 340:64–72
Pollard MR, McKeon T, Gupta LM, Stumpf PK (1979) Studies on biosynthesis of waxes by developing jojoba seed. II. The demonstration of wax biosynthesis by cell-free homogenates. Lipids 14:651–662
Moore C, Snyder F (1982) Properties of microsomal acyl coenzyme A reductase in mouse preputial glands. Arch Biochem Biophys 214:489–499
Wang X, Kolattukudy PE (1995) Solubilization, purification and characterization of fatty acyl-CoA reductase from duck uropygial gland. Biochem Biophys Res Commun 208:210–215
Johnson RC, Gilbertson JR (1972) Isolation, characterization, and partial purification of a fatty acyl coenzyme A reductase from bovine cardiac muscle. J Biol Chem 247:6991–6998
Falk-Petersen S, Hagen W, Kattner G, Clarke A, Sargent JR (2000) Lipids, trophic relationships, and biodiversity in Arctic and Antarctic krill. Can J Fish Aquat Sci 57:178–191
Kattner G, Hagen W (1998) Lipid metabolism of the Antarctic euphausid Euphausia crystallorophias and its ecological implications. Mar Ecol Prog Ser 170:203–213
Kolattukudy PE, Bohnet S, Rogers L (1987) Diesters of 3-hydroxy fatty acids produced by the uropygial glands of female mallards uniquely during the mating season. J Lipid Res 28:582–588
Rijpstra WI, Reneerkens J, Piersma T, Damste JS (2007) Structural identification of the β-hydroxy fatty acid-based diester preen gland waxes of shorebirds. J Nat Prod 70:1804–1807
Acknowledgments
We are very grateful to Patricia Vrinten for technical assistance in gene isolation. We thank Devin Polichuk, Darwin Reed and Valerie Catinot for technical help and discussions. This work is part of ICON, a European Community Seventh Framework Programme.
Author information
Authors and Affiliations
Corresponding author
About this article
Cite this article
Teerawanichpan, P., Qiu, X. Fatty Acyl-CoA Reductase and Wax Synthase from Euglena gracilis in the Biosynthesis of Medium-Chain Wax Esters. Lipids 45, 263–273 (2010). https://doi.org/10.1007/s11745-010-3395-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11745-010-3395-2