Abstract
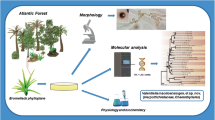
The systematic position of three yeast strains isolated from a plant cell culture, a piece of termite nest, or as a foliar endophyte of Coffea arabica, respectively, is evaluated using morphological, physiological, and phylogenetical characteristics. In culture, all three isolates produced white, pale orange to pink colored colonies of cylindrical cells with monopolar budding and pseudohyphae. Standard phenotypic, biochemical, physiological characterization, and phylogenetic analyses of the combined 26S rRNA gene (D1/D2 domains) and ITS region sequences showed the conspecificity of these isolates and suggest their placement within the Exobasidiales (Ustilaginomycotina) as a sister lineage of the sampled and sequenced Graphiola species. Here, we describe this species as Graphiola fimbriata sp. nov. MycoBank MB 825077 (holotype: PC1T; ex-type cultures: IBRC-M 30158T = CBS 13945T = DSM 104832T). This is the first species described in the genus Graphiola for which only the asexual, saprobic developmental phase is known. The description of the genus Graphiola is therefore emended to allow species known only from a saprobic state.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Fungi with yeast and yeast-like morphology are present within all three main lineages of Basidiomycota, namely the subphyla Agaricomycotina, Pucciniomycotina, and Ustilaginomycotina (Sampaio 2004; Boekhout et al. 2011; Kurtzman and Boekhout 2017). Yeast and filamentous taxa are intermixed as revealed by recent phylogenetic studies (Liu et al. 2015a, 2015b; Wang et al. 2015a, 2015b, 2015c). With a few exceptions, a transition from filamentous to yeast stage is associated with the change from a parasitic to saprobic life style (Begerow et al. 2017). The subphylum Ustilaginomycotina comprises a highly diverse assemblage of fungi, including teliosporic plant pathogens, non-teliosporic plant pathogens, anamorphic plant pathogens, as well as endophytic, lipophilic, saprobic, and zoophilic yeasts (Begerow et al. 2014). The orders Malasseziales, Monilielalles, and Violaceomycetales include exclusively yeast species of the genera Malassezia, Moniliella, and Violaceomyces, respectively (Wang et al. 2014; Albu et al. 2015), while the remaining yeasts are scattered among the remaining orders of Ustilaginomycotina, with the exception of the members of Doassansiales, Tilletiales, and Uleiellales for which no yeasts have been discovered so far.
The anamorphic saprobic members of Ustilaginomycotina have previously been classified mainly in the polyphyletic genera Pseudozyma and Tilletiopsis (Begerow et al. 2000; Sampaio 2004; Kurtzman et al. 2011, 2015) and later in several monophyletic genera Acaromyces, Farysizyma, Fereydounia, Jaminaea, Meira, and Sympodiomycopsis (Sugiyama et al. 1991; Boekhout et al. 2003; Inácio et al. 2008; Sipiczki and Kajdacsi 2009; Nasr et al. 2014). Sexual and asexual morphs in Ustilaginomycotina were recently grouped together in order to unify the taxonomy of plant parasites and species known only from their yeast states (Piątek et al. 2015; Wang et al. 2015a; Kijpornyongpan and Aime 2016). As the result, several monophyletic genera (Dirkmeia, Golubevia, Kalmanozyma, and Robbauera) were erected to accommodate species previously classified in the genera Pseudozyma and Tilletiopsis (Wang et al. 2015a). The current classification scheme recommends that each novel lineage not linked with a teleomorphic genus or recognized anamorphic genus should be assigned to a new genus. The subphylum Ustilaginomycotina comprises the least number of yeast and yeast-like species (71) compared to the two other subphyla of Basidiomycota (Wang et al. 2015a). However, studies that involved environmental sequencing suggest a much greater, unexplored diversity of fungi in this group (e.g., Richards et al. 2012; Nasanit et al. 2015; Dunthorn et al. 2017; Jimu et al. 2017).
In the course of independent studies two yeast strains have been isolated. The first culture (PC1) was isolated in the Iranian Biological Resource Center (IBRC) from a contaminated plant culture, and the second isolate (SNB-CN72) was obtained from a nest of the termite species Nasutitermes corniger harvested in Rémire-Montjoly, French Guiana. Phylogenetic analyses of the ITS region and the D1/D2 domains of the LSU rRNA gene placed the strains within Graphiolaceae (Exobasidiomycetes, Exobasidiales) close to plant parasites of the genus Graphiola and revealed relatedness to a third yeast isolate (IBL 03150) reported by Posada et al. (2007) and deposited in the CBS yeast collection of the Westerdijk Fungal Biodiversity Institute, Utrecht, The Netherlands. Species in the plant pathogenic genus Graphiola are traditionally circumscribed based on their host spectrum and symptoms, and nucleotide sequence data for these fungi are rare (Wang et al. 2015a).
The aim of the study was to characterize the yeast-like isolates and determine their phylogenetic placement, incorporating morphological, physiological, and molecular data available for members of the genus Graphiola.
Materials and methods
Sample collection and isolation
A plant cell culture of an unidentified plant species mixed with a yeast strain was obtained from the plant bank section of the IBRC in 2013 (strain designation: PC1). Another strain was isolated from a piece of N. corniger nest collected in Rémire-Montjoly (Cayenne, French Guiana) in July 2011 (strain designation: SNB-CN72; Nirma et al. 2013, 2015). A third strain was obtained from the CBS yeast collection of the Westerdijk Fungal Biodiversity Institute, Utrecht, The Netherlands (original strain designation: IBL 03150). It was isolated as foliar endophyte of Coffea arabica (Posada et al. 2007). Designations and GenBank accession numbers of the yeast strains used in this study are given in Table 1 .
Cultures were maintained on YPG agar medium (0.5% yeast extract, 1% peptone, 2% glucose, 2% agar, w/v) at 25 °C during experiments.
To expand the ITS sequence sampling for molecular phylogenetic analyses, a specimen of the plant parasitic genus Graphiola was additionally used and newly sequenced: Graphiola phoenicis (Moug. ex Fr.) Poit. on Phoenix reclinata Jacq., South Africa, Kwa Zulu-Natal, Durban, Yellowwood Park, Kenneth Stainbank Nature Reserve, 14 Feb. 2012, leg. A.R. Wood 883, KR-M-0042315.
Morphological examination
Colony morphology images were taken using a stereo microscope coupled with the Nikon zoom digital camera. In addition, scanning electron microscopy was performed on the isolate using a VEGA3-TESCAN SEM instrument (Van Wyk and Wingfield 1994). Briefly, the cells were fixed in 3.0%, 0.1-M, pH 7.0 sodium phosphate-buffered glutaraldehyde for 3 h at room temperature, followed by 1 h fixation in 2% osmium tetroxide. The cells were dehydrated by increasing ethanol concentrations (30, 50, 70, 90, and 96%) for 30 min and two 30-min washes in 100% ethanol. The standard characterization of the yeast isolates was performed according to methods described earlier (Barnett et al. 2000; Kurtzman et al. 2011). Assimilation of carbon and nitrogen sources was carried out on solid and in liquid media, respectively.
DNA extraction, PCR, and sequencing
Nuclear DNA was extracted by the method of Hanna and Xiao (2006). For the G. phoenicis specimen, the genomic DNA was isolated directly from the herbarium specimen. The 5′-end (D1/D2 region) of the nuclear large subunit ribosomal DNA (LSU) and the ITS 1 and ITS 2 regions of the nuclear rDNA including the 5.8S rDNA (ITS) were amplified and sequenced using primer pairs NL1 (5′GCATATCAATAAGCGGAGGAAAAG-3′) and NL4 (5′-GGTCCGTGTTTCAAGACGG3′) (Kurtzman and Robnett 1998) and ITS1 (5′-TCCGTAGGTGAACCTGCGG-3′) and ITS4 (5′-TCCTCCGCTTATTGATATGC-3′) (White et al. 1990), respectively. DNA sequences determined for this study were deposited in GenBank, accession numbers are given in Figs. 1 and 2 and in Table 1. Additional sequences of Graphiola cylindrica and G. phoenicis were obtained from GenBank and the public catalog of the NITE Biological Resource Center collection (NBRC), Japan.
Bayesian inference of phylogenetic relationships within the sampled Exobasidiales: Markov chain Monte Carlo analysis of an alignment of LSU base sequences using the GTR + I + G model of DNA substitution with gamma distributed substitution rates and estimation of invariant sites, random starting trees, and default starting parameters of the DNA substitution model. A 50% majority-rule consensus tree is shown computed from 75,000 trees that were sampled after the process had become stationary. The topology was rooted with the ustilaginomycetous species U. ficariae and U. hordei. Numbers on branches before slashes are estimates for a posteriori probabilities; numbers on branches after slashes are ML bootstrap support values. Branch lengths were averaged over the sampled trees. They are scaled in terms of expected numbers of nucleotide substitutions per site. The taxonomical concept applied corresponds to Begerow et al. (2014)
Bayesian inference of phylogenetic relationships within the sampled Exobasidiales: Markov chain Monte Carlo analysis of an alignment of ITS + LSU base sequences using the GTR + I + G model of DNA substitution with gamma distributed substitution rates and estimation of invariant sites, random starting trees, and default starting parameters of the DNA substitution model. A 75% majority-rule consensus tree is shown computed from 75,000 trees that were sampled after the process had become stationary. The topology was rooted with the cryptobasidiaceous species Acaromyces ingoldii and Clinoconidium sp. Numbers on branches before slashes are estimates for a posteriori probabilities; numbers on branches after slashes are ML bootstrap support values. Branch lengths were averaged over the sampled trees. They are scaled in terms of expected numbers of nucleotide substitutions per site. The taxonomical concept applied corresponds to Begerow et al. (2014)
Phylogenetic analyses
To elucidate the phylogenetic position of the isolated strains, their sequences were analyzed within a LSU and a concatenated ITS + LSU dataset. Since preliminary analyses and Blast searches (Altschul et al. 1997) revealed an affinity of the strains to the order Exobasidiales (class Exobasidiomycetes), the LSU dataset was reduced to members of the Exobasidiales and some representatives of the Ustilaginomycetes were used as an outgroup. Isolates and clones, for which only ITS sequences were available, were analyzed in the LSU + ITS dataset. If present in GenBank, sequences of the respective type species were used. Additionally sequences of all available brachybasidiaceous species and all available sequences clustering within the Graphiolaceae including sequences from the Biological Resource Center, NITE, Japan were added.
GenBank accession numbers of the sequences used for both the LSU and ITS + LSU dataset (Begerow et al. 1997, 2001, 2002; Guo et al. 2001; Boekhout et al. 2003; Castlebury et al. 2005; Stoll et al. 2005; Yasuda et al. 2005, 2006; Posada et al. 2007; Tanaka et al. 2008; Piepenbring et al. 2010, 2012; Yuan et al. 2011; Jusino et al. 2015; Urbina et al. 2016) are given in Figs. 1 and 2.
Sequence alignment was obtained independently for both the LSU dataset and the ITS and LSU part of the ITS + LSU dataset using MAFFT 7.313 (Katoh and Standley 2013) using the L-INS-i option. To obtain reproducible results, manipulation of the alignments by hand as well as manual exclusion of ambiguous sites were avoided as suggested by Gatesy et al. (1993) and Giribet and Wheeler (1999), respectively. Instead, highly divergent portions of the alignments were omitted using GBlocks 0.91b (Castresana 2000) with the following options for the LSU dataset: “Minimum Number of Sequences for a Conserved Position”: 20, “Minimum Number of Sequences for a Flank Position”: 20, “Maximum Number of Contiguous Non-conserved Positions”: 8, “Minimum Length of a Block”: 5, and “Allowed Gap Positions” to “With half,” for the ITS part of the ITS + LSU dataset: 11/11/8/5/”With half” and for the LSU part of the ITS + LSU dataset: 12/12/8/5/”With half.” After alignment the ITS and LSU part of the ITS + LSU dataset were concatenated. The resulting alignments [LSU dataset: new number of positions: 591 (32% of the original 1801 positions), number of variable sites: 279; ITS + LSU dataset: new number of positions: 1121 (34% of the original 3264 positions), number of variable sites: 468] were used for phylogenetic analyses using a maximum likelihood (ML) and a Bayesian approach (BA) following Vasighzadeh et al. (2014).
For the LSU dataset, trees were rooted with the ustilaginomycetous species Urocystis ficariae and Ustilago hordei, for the ITS + LSU dataset trees were rooted with the brachybasidiaceous species Dicellomyces gloeosporus and Meira geulakonigii.
Metabolite profiling analysis was carried out for the three yeast strains and the results are presented as supplementary data.
Results
Morphological examination
Morphology of the yeast strains IBL 03150, PC1, and SNB-CN72 showed no significant differences; details are included in the species description.
Metabolite profiling with UHPLC-MS was carried out for the three yeast strains and the results are provided in supplementary data. In the absence of metabolic data from closely related species, the analysis of UHPLC-MS profiles was restricted to the tree strains. We observed that the culture medium impacted more significantly on the profiles than the nature of the strain (Supplementary data). Hierarchical clustering analyses (HCA) conducted on the 3 (N) × 29 (X) matrix of merged profiles showed that strains PC1 and IBL 03150 are more alike than SNB-CN72 (Supplementary data).
Phylogenetic analyses
The different runs of the BA that were performed and the ML analyses yielded consistent topologies. To illustrate the results, the consensus trees of one run of the BA of the LSU and the concatenated ITS + LSU dataset are presented (Figs. 1 and 2). Using the ustilaginomycetous species as outgroup in the LSU analysis, the clades in the phylogenetic tree were congruent to the families discussed in Begerow et al. (2014). In all analyses, the sequences of the yeast isolates clustered within the Exobasidiales together with the sequence MF334501 from an uncultured fungus clone forming the sister lineage of the sampled Graphiola spp. of which the G. phoenicis cluster included several sequences of yeast isolates and uncultured fungus clones, respectively. The clade comprising G. cylindrica, G. geonomae, G. phoenicis, and the yeast isolates received good statistical support (BA: 100%, ML: 93%). The yeast isolates branched first in the Graphiola clade and were placed sister to G. cylindrica, G. geonomae, and G. phoenicis. Combined ITS + LSU analyses (Fig. 2) revealed the same groups and relations including more sequences from Graphiola spp. as well as from the studied yeast isolates and uncultured fungus clones. The three strains shared identical LSU sequences and showed five variable positions (two substitutions and three indels) in the ITS region.
Taxonomy
GraphiolaPoit., Ann. Sci. Nat. (Paris): 473 (1824) emend. “S. Nasr, M. Lutz, D. Stien & A. Yurkov”
According to the current genus concept of Graphiola (Piepenbring et al. 2012), the genus comprises plant pathogens on palms (Arecaceae). Tubaki and Yokoyama (1971) and Oberwinkler et al. (1982) obtained and studied G. phoenicis in culture. They provided also the diagnosis of this species, including morphological and physiological properties observed in culture. Emendation of the diagnosis of the genus Graphiola is proposed to allow classification of asexual, known only from a saprobic state yeast species in the genus.
Sexual reproduction is observed in some species. Fungi are dimorphic with a filamentous sexual form parasitizing on plants and free-living saprobic yeast states. On malt extract, colonies are white, pale orange to pink in color. Budding cells are present in culture. Colonies are white, pale orange to pink in color. Ballistoconidia are not produced. Pseudohyphae may be present in cultures. Fermentation is absent. DBB reaction is positive. Urease activity is positive.
Graphiola fimbriata S. Nasr, M. Lutz, D. Stien & A. Yurkov, sp. nov. (Fig. 3) MycoBank MB 825077
Etymology: Referring to the margin of the colonies.
Description: After 3 days on YPG agar at 25 °C, the cells are cylindrical, 1–2.5 × 4–8.5 μm (n = 20), and occur single or in small chains. Budding is monopolar. After 5 days, the colony is white, convex, and the margin is filiform. Pellicles are formed in liquid media. After 1 week at 25 °C on MEA and Cornmeal agar, in the slide culture undifferentiated pseudohyphae are formed. Fermentation of D-glucose, D-galactose, sucrose, maltose, lactose, raffinose and α-α-trehalose, starch, melibiose, cellobiose, and D-xylose is negative. The following compounds are assimilated: glucose, sucrose, maltose, trehalose, melezitose, D-xylose, raffinose, L-arabinose, D-mannitol, myo-inositol, D-ribose, and arbutin; assimilation of salicin, glycerol, and cellobiose is weak. No growth occurs on galactose, lactose, L-rhamnose, melibiose, inulin, D-arabinose, soluble starch, ethanol, methanol, DL-lactate, succinate, citrate, and n-hexadecane. Sodium nitrate, potassium nitrate, L-lysine, ethylamine hydrochloride (variable), and cadaverine dihydrochloride (variable) are assimilated. No growth occurs on glucosamine, imidazole, creatine, or creatinine. Growth in vitamin-free medium is positive. Growth at 15, 25, 30, or 34 °C is positive but not at 4, 37, and 40 °C. Growth occurs on YM agar supplemented with 5% (w/v) NaCl, 10% (w/v) NaCl, but not on YM agar with 16% (w/v) NaCl. Starch-like compounds are not produced. No growth occurs on media supplemented with 0.01 and 0.1% cycloheximide and 1% acetic acid. The diazonium blue B reaction is weakly positive. Urease activity is positive.
Molecular characteristics: nucleotide sequences of ITS and LSU (D1/D2 domains) rDNA sequences are deposited in GenBank (Table 1.)
Deposits: holotype, PC1T (=IBRC-M 30158T) isolated as a contaminant of an unidentified plant culture, preserved in a metabolically inactive state at the Iranian Biological Resource Centre, Teheran (holotype), Iran. Ex-type cultures are deposited in the CBS yeast collection of the Westerdijk Fungal Biodiversity Institute, Utrecht, The Netherlands (=CBS 13945T) and in the German Collection of Microorganisms and Cell Cultures, Braunschweig, Germany (DSM 104832T).
Strain examined: PC1T, SNB-CN72 (=DSM 104833), IBL 03150 (for further data see Material and Methods)
Discussion
Morphological and physiological characteristics of the three yeast isolates examined in this study are uniform, differences in ITS and LSU sequences are low all that suggesting assignment to a single species. Molecular phylogenetic analyses reveal the three yeast isolates within the Graphiolaceae as a sister lineage of G. cylindrica, G. geonomae, and G. phoenicis. Thus, we propose the new species G. fimbriata to accommodate the three yeast strains. Based on sequencing results for ITS and D1/D2 regions and the phylogenetic analyses of the same loci, the novel species differs from Graphiola species that were included in the analysis (Figs. 1 and 2).
We also emend the description of the genus Graphiola to allow the inclusion of species known from saprobic state only (see “Taxonomy” section).
The ecology and distribution range of the novel species is unknown. All described Graphiola species are plant pathogens on palms (Arecaceae) during their sexual developmental stage, and a few species were reported to produce yeasts for asexual, saprotrophic growth, and multiplication. We hypothesize that the novel species is probably associated with plants, as also suggested by the origin of the studied isolates and closely related culture of a species of Exobasidiomycetidae AUMC 10262 (KX011608) from the palm weevil Rhynchophorus ferrugineus that infests palms, which is a primary host of Graphiola. Although the novel species is known from its asexual state, it is most likely that it has a host like all other known Graphiola species. Widespread transcontinental transfer of plants results in a global dissemination of plant pathogens and crop pests (Bebber 2015; Hurley et al. 2016). Among them, fungal pathogens currently lead the global invasion of agriculture, despite their more restricted host range (Bebber et al. 2014; Wingfield et al. 2017). In our study, a new species of the plant pathogenic genus Graphiola was identified over a broad geographic range that includes the Old World (Iran and Egypt), tropical Asia, and the Americas (French Guiana and USA). It is very likely that the ongoing transport of plants and pests promoted dissemination of G. fimbriata between continents, localities, and habitats.
It cannot be precluded that G. fimbriata represents the anamorphic stage of a palm pathogen of which no sequence data are available. Of the 12 Graphiola species described (Piepenbring et al. 2012), only three are represented by sequence data in GenBank. Moreover species diversity seems underestimated in the genus Graphiola. Assuming that its host specificity developed to the same degree shown for other plant pathogens (Piątek et al. 2013; Savchenko et al. 2014; Vasighzadeh et al. 2014; Choi and Thines 2015; Scholler et al. 2016), the number of 12 Graphiola species on 38 palm species from 21 plant genera (Farr and Rossman n.d.; Piepenbring et al. 2012) is lower than in other known plant-parasite systems. Intensive sampling and molecular analyses were used successfully in other groups to link anamorph and teleomorph stages (Sampaio 2004; Boekhout et al. 2006; Inácio et al. 2008; Wang et al. 2015a; Kruse et al. 2017; Piątek et al. 2017). However, both sexual and asexual species are yet strongly under-sampled in many lineages (Liu et al. 2015a; Wang et al. 2015a; Wang et al. 2015b). Although these studies unified systematics of plant parasites and yeast taxa, the results strongly suggest that the available genetic data is insufficient to resolve anamorph-teleomorph relationships in sister lineages (e.g., Boekhout et al. 2006; Kruse et al. 2017).
The lack of sequence data of sexual species also complicates biodiversity assessments and new species discovery. With this study, we provide the first overview on Graphiola diversity from sequence data available in public databases. We propose the new species G. fimbriata to accommodate yeasts from several localities and habitats. Our results show that sequences related to Graphiola species are rapidly accumulating in public databases as taxonomically unassigned isolates and clones. We agree that taxonomic redundancy cannot be ruled out in the genus Graphiola considering that sequence data are available only for few species. Rediscovery and description of already known species as new may happen when stable and informative morphological characters for species differentiation are limited, both in plant material and in culture. Thus, future studies should be addressed on resampling and sequencing of new and already existing species in order to better understand genetic diversity and taxonomy in this group of fungi.
There is little known about physiological properties of dimorphic plant parasites, whereas fungi traditionally recognized as “yeasts” were intensively studied in this respect. Under-sampling and the lack of genetic (including housekeeping genes) and physiological data do not allow delimitation of many genera (e.g., Wang et al. 2015a). In this study, we decided not to propose a new genus for the novel fungus, but to accommodate it in the genus Graphiola in order to reduce the taxonomic complexity in this group. Neither phylogenetic analyses nor the analysis of physiological data revealed a basis to distinguish the new species from other Graphiola species. Therefore, we modified the description of the genus Graphiola to include information about its asexual form considering own results and previous reports by Tubaki and Yokoyama (1971) and Oberwinkler et al. (1982). Despite limited genetic data, our study identified several fungal isolates and metabarcoding clones as G. fimbriata and G. phoenicis (Figs. 1 and 2). These fungi were found in Indomalayan and Neotropical realms (e.g., Takashima et al. 2012; Urbina et al. 2016), which are the regions where palms are widely distributed (Kissling et al. 2012).
References
Albu S, Toome M, Aime MC (2015) Violaceomyces palustris gen. et sp. nov. and a new monotypic lineage, Violaceomycetales ord. nov. in Ustilaginomycetes. Mycologia 107:1193–1204
Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Barnett JA, Payne RW, Yarrow D (2000) Yeasts: characteristics and identification, 3rd edn. Cambridge University Press, Cambridge
Bebber DP (2015) Range-expanding pests and pathogens in a warming world. Annu Rev Phytopathol 53:335–356
Bebber DP, Holmes T, Gurr SJ (2014) The global spread of crop pests and pathogens. Glob Ecol Biogeogr 23:1398–1407
Begerow D, Bauer R, Oberwinkler F (1997) Phylogenetic studies on nuclear LSU rDNA sequences of smut fungi and related taxa. Can J Bot 75:2045–2056
Begerow D, Bauer R, Boekhout T (2000) Phylogenetic placements of ustilaginomycetous anamorphs as deduced from nuclear LSU rDNA sequences. Mycol Res 104:53–60
Begerow D, Bauer R, Oberwinkler F (2001) Muribasidiospora: Microstromatales or Exobasidiales? Mycol Res 105:798–810
Begerow D, Bauer R, Oberwinkler F (2002) The Exobasidiales: an evolutionary hypothesis. Mycol Prog 1:187–199
Begerow D, Schäfer AM, Kellner R, Yurkov A, Kemler M, Oberwinkler F, Bauer R (2014) Ustilaginomycotina. In: McLaughlin DJ, Spatafora JW (eds) The Mycota, systematics and evolution, vol vol. 7A. Springer, Berlin, pp 299–330
Begerow D, Kemler M, Feige A, Yurkov A (2017) Parasitism in yeasts. In: Buzzini P, Lachance MA, Yurkov A (eds) Yeasts in natural ecosystems: ecology. Springer, Cham, pp 179–210
Boekhout T, Theelen B, Houbraken J, Robert V, Scorzetti G, Gafni A, Gerson U, Sztejnberg A (2003) Novel anamorphic mite-associated fungi belonging to the Ustilaginomycetes: Meira geulakonigii gen. nov., sp. nov., Meira argovae sp. nov. and Acaromyces ingoldii gen. nov., sp. nov. Int J Syst Evol Microbiol 53:1655–1664
Boekhout T, Gildemacher P, Theelen B, Müller WH, Heijne B, Lutz M (2006) Extensive colonization of apples by smut anamorphs causes a new postharvest disorder. FEMS Yeast Res 6:63–76
Boekhout T, Fonseca Á, Sampaio JP (2011) Discussion of teleomorphic and anamorphic basidiomycetous yeasts. In: Kurtzman CP, Fell JW, Boekhout T (eds) The yeasts, a taxonomic study, vol 2011, 5th edn. Elsevier, Amsterdam, pp 1339–1372
Castlebury LA, Carris LM, Vánky K (2005) Phylogenetic analysis of Tilletia and allied genera in order Tilletiales (Ustilaginomycetes; Exobasidiomycetidae) based on large subunit nuclear rDNA sequences. Mycologia 97:888–900
Castresana J (2000) Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol Biol Evol 17:540–552
Choi Y-J, Thines M (2015) Host jumps and radiation, not co-divergence drives diversification of obligate pathogens. A case study in downy mildews and Asteraceae. PLoS One 10:e0133655
Dunthorn M, Kauserud H, Bass D, Mayor J, Mahé F (2017) Yeasts dominate soil fungal communities in three lowland neotropical rainforests. Environ Microbiol Rep 9:668–675
Farr DF, Rossman AY (n.d.) Fungal databases. U.S. National Fungus Collections, ARS, USDA Retrieved April 18, 2018, from https://nt.ars-grin.gov/fungaldatabases/
Gatesy J, DeSalle R, Wheeler W (1993) Alignment-ambiguous nucleotide sites and the exclusion of systematic data. Mol Phylogenet Evol 2:152–157
Giribet G, Wheeler WC (1999) On gaps. Mol Phylogenet Evol 13:132–143
Guo LD, Hyde KD, Liew EC (2001) Detection and taxonomic placement of endophytic fungi within frond tissues of Livistona chinensis based on rDNA sequences. Mol Phylogenet Evol 20:1–13
Hanna M, Xiao W (2006) Isolation of nucleic acids. In: Xiao W (ed) Yeast protocols (methods in molecular biology), vol 313. Humana Press, New York, pp 15–20
Hurley BP, Garnas J, Wingfield MJ, Branco M, Richardson DM, Slippers B (2016) Increasing numbers and intercontinental spread of invasive insects on eucalypts. Biol Invasions 18:921–933
Inácio J, Landell MF, Valente P, Wang PH, Wang YT, Yang SH, Manson JS, Lachance MA, Rosa CA, Fonseca Á (2008) Farysizyma gen. nov., an anamorphic genus in the Ustilaginales to accommodate three novel epiphytic basidiomycetous yeast species from America, Europe and Asia. FEMS Yeast Res 8:499–508
Jimu L, Kemler M, Mujuru L, Mwenje E (2017) Illumina DNA metabarcoding of Eucalyptus plantation soil reveals the presence of mycorrhizal and pathogenic fungi. Forestry 19:1–8
Jusino MA, Lindner DL, Banik MT, Walters JR (2015) Heart rot hotel: fungal communities in red-cockaded woodpecker excavations. Fungal Ecol 14:33–43
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30:772–780
Kijpornyongpan T, Aime MC (2016) Rare or rarely detected? Ceraceosorus guamensis sp. nov.: a second described species of Ceraceosorales and the potential for underdetection of rare lineages with common sampling techniques. Antonie Van Leeuwenhoek 109:1127–1139
Kissling WD, Eiserhardt WL, Baker WJ, Borchsenius F, Couvreur TL, Balslev H, Svenning JC (2012) Cenozoic imprints on the phylogenetic structure of palm species assemblages worldwide. Proc Natl Acad Sci U S A 109:7379–7384
Kruse J, Doehlemann G, Kemen E, Thines M (2017) Asexual and sexual morphs of Moesziomyces revisited. IMA Fungus 8:117–129
Kurtzman CP, Boekhout T (2017) Yeasts as distinct life forms of Fungi. In: Buzzini P, Lachance MA, Yurkov A (eds) Yeasts in natural ecosystems: ecology. Springer, Cham, pp 1–37
Kurtzman CP, Robnett CJ (1998) Identification and phylogeny of ascomycetous yeasts from analysis of nuclear large subunit (26S) ribosomal DNA partial sequences. Antonie Van Leeuwenhoek 73:331–371
Kurtzman CP, Fell JW, Boekhout T (2011) The yeasts, a taxonomic study, 5th edn. Elsevier Science Publication, Amsterdam
Kurtzman CP, Mateo RQ, Kolecka A, Theelen B, Robert V, Boekhout T (2015) Advances in yeast systematics and phylogeny and their use as predictors of biotechnologically important metabolic pathways. FEMS Yeast Res 15:fov050
Liu XZ, Wang QM, Göker M, Groenewald M, Kachalkin AV, Lumbsch HT, Millanes AM, Wedin M, Yurkov AM, Boekhout T, Bai FY (2015a) Towards an integrated phylogenetic classification of the Tremellomycetes. Stud Mycol 81:85–147
Liu XZ, Wang QM, Theelen B, Groenewald M, Bai FY, Boekhout T (2015b) Phylogeny of tremellomycetous yeasts and related dimorphic and filamentous basidiomycetes reconstructed from multiple gene sequence analyses. Stud Mycol 81:1–26
Nasanit R, Tangwong-O-Thai A, Tantirungkij M, Limtong S (2015) The assessment of epiphytic yeast diversity in sugarcane phyllosphere in Thailand by culture-independent method. Fungal Biol 119:1145–1157
Nasr S, Soudi MR, Shahzadeh Fazeli SA, Nguyen HDT, Lutz M, Piątek P (2014) Expanding evolutionary diversity in the Ustilaginomycotina: Fereydouniaceae fam. nov. and Fereydounia gen. nov., the first urocystidalean yeast lineage. Mycol Prog 13:1012
Nirma C, Eparvier V, Stien D (2013) Antifungal agents from Pseudallescheria boydii SNB-CN73 isolated from a Nasutitermes sp. termite. J Nat Prod 76:988–991
Nirma C, Eparvier V, Stien D (2015) Antibacterial ilicicolinic acids C and D and ilicicolinal from Neonectria discophora SNB-CN63 isolated from a termite nest. J Nat Prod 78:159–162
Oberwinkler F, Bandoni RJ, Blanz P, Deml G, Kisimova-Horovitz L (1982) Graphiolales: Basidiomycetes parasitic on palms. Pl Syst Evol 140:251–277
Piątek M, Lutz M, Chater AO (2013) Cryptic diversity in the Antherospora vaillantii complex on Muscari species. IMA Fungus 4:5–19
Piątek M, Lutz M, Yorou NS (2015) A molecular phylogenetic framework for Anthracocystis (Ustilaginales), including five new combinations (inter alia for the asexual Pseudozyma flocculosa), and description of Anthracocystis grodzinskae sp. nov. Mycol Prog 14:88
Piątek M, Lutz M, Sousa FMP, Santos ARO, Félix CR, Landell MF, Gomes FCO, Rosa CA (2017) Pattersoniomyces tillandsiae gen. et comb. nov.: linking sexual and asexual morphs of the only known smut fungus associated with Bromeliaceae. Org Divers Evol 17:531–543
Piepenbring M, Espinoza J, Saldaña L, Cáceres O (2010) New records, host plants, morphological and molecular data of Exobasidiales (Basidiomycota) from Panama. Nova Hedwigia 91:231–242
Piepenbring M, Nold F, Trampe T, Kirschner R (2012) Revision of the genus Graphiola (Exobasidiales, Basidiomycota). Nova Hedwigia 94:67–96
Posada F, Aime MC, Peterson SW, Rehner SA, Vega FE (2007) Inoculation of coffee plants with the fungal entomopathogen Beauveria bassiana (Ascomycota: Hypocreales). Mycol Res 111:748–757
Richards TA, Jones MD, Leonard G, Bass D (2012) Marine fungi: their ecology and molecular diversity. Annu Rev Mar Sci 4:495–522
Sampaio JP (2004) Diversity, phylogeny and classification of basidiomycetous yeasts. In: Agerer R, Blanz P, Piepenbring M (eds) Frontiers in basidiomycote mycology. IHW-Verlag, Eching, pp 49–80
Savchenko KG, Carris LM, Castlebury LA, Heluta VP, Wasser SP, Nevo E (2014) Revision of Entyloma (Entylomatales, Exobasidiomycetes) on Eryngium. Mycologia 106:797–810
Scholler M, Schmidt A, Siahaan SAS, Takamatsu S, Braun U (2016) A taxonomic and phylogenetic study of the Golovinomyces biocellatus complex (Erysiphales, Ascomycota) using asexual state morphology and rDNA sequence data. Mycol Prog 15:56
Sipiczki M, Kajdacsi E (2009) Jaminaea angkorensis gen. nov., sp. nov., a novel anamorphic fungus containing an S943 nuclear small-subunit rRNA group IB intron represents a basal branch of Microstromatales. Int J Syst Evol Microbiol 59:914–920
Stoll M, Begerow D, Oberwinkler F (2005) Molecular phylogeny of Ustilago, Sporisorium, and related taxa based on combined analyses of rDNA sequences. Mycol Res 109:342–356
Sugiyama J, Tokuoka K, Suh SO, Hirata A, Komagata K (1991) Sympodiomycopsis: a new yeast-like anamorph genus with basidiomycetous nature from orchid nectar. Antonie Van Leeuwenhoek 59:95–108
Takashima M, Sugita T, Van BH, Nakamura M, Endoh R, Ohkuma M (2012) Taxonomic richness of yeasts in Japan within subtropical and cool temperate areas. PLoS One 7:e50784
Tanaka E, Shimizu K, Imanishi Y, Yasuda F, Tanaka C (2008) Isolation of basidiomycetous anamorphic yeast-like fungus Meira argovae found on Japanese bamboo. Mycoscience 49:329–333
Tubaki K, Yokoyama T (1971) Cultural aspects of Graphiola phoenicis. Mycopathol Mycol Appl 43:49–60
Urbina H, Scofield DG, Cafaro M, Rosling A (2016) DNA-metabarcoding uncovers the diversity of soil-inhabiting fungi in the tropical island of Puerto Rico. Mycoscience 57:217–227
Van Wyk PWJ, Wingfield MJ (1994) Ultrastructure of ascus arrangement and ascospore development in Ophiostoma seticolle. Mycologia 86:607–614
Vasighzadeh A, Zafari D, Selçuk F, Hüseyin E, Kurşat M, Lutz M, Piątek M (2014) Discovery of Thecaphora schwarzmaniana on Rheum ribes in Iran and Turkey: implications for the diversity and phylogeny of leaf smuts on rhubarbs. Mycol Prog 13:881–892
Wang QM, Theelen B, Groenewald M, Bai FY, Boekhout T (2014) Moniliellomycetes and Malasseziomycetes, two new classes in Ustilaginomycotina. Persoonia 33:41–47
Wang QM, Begerow D, Groenewald M, Liu XZ, Theelen B, Bai FY, Boekhout T (2015a) Multigene phylogeny and taxonomic revision of yeasts and related fungi in the Ustilaginomycotina. Stud Mycol 81:55–83
Wang QM, Groenewald M, Takashima M, Theelen B, Han PJ, Liu XZ, Boekhout T, Bai FY (2015b) Phylogeny of yeasts and related filamentous fungi within Pucciniomycotina determined from multigene sequence analyses. Stud Mycol 81:27–53
Wang QM, Yurkov AM, Göker M, Lumbsch HT, Leavitt SD, Groenewald M, Theelen B, Liu XZ, Boekhout T, Bai FY (2015c) Phylogenetic classification of yeasts and related taxa within Pucciniomycotina. Stud Mycol 81:149–189
White TJ, Bruns TD, Lee S, Taylor JW (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ (eds) PCR protocols: a guide to methods and applications. Academic Press, San Diego, pp 315–322
Wingfield MJ, Slippers B, Wingfield BD, Barnes I (2017) The unified framework for biological invasions: a forest fungal pathogen perspective. Biol Invasions 19:3201–3214
Yasuda F, Yamagishi D, Akamatsu H, Izawa H, Kodama M, Otani H (2005) Fruit stain of Japanese pear caused by basidiomycetous, yeast-like fungi, Acaromyces ingoldii and Meira sp. Jpn J Phytopathol 71:156–165
Yasuda F, Yamagishi D, Akamatsu H, Izawa H, Kodama M, Otani H (2006) Meira nashicola sp. nov., a novel basidiomycetous, anamorphic yeastlike fungus isolated from Japanese pear fruit with reddish stain. Mycoscience 47:36–40
Yuan ZL, Rao LB, Chen YC, Zhang CL, Wu YG (2011) From pattern to process: species and functional diversity in fungal endophytes of Abies beshanzuensis. Fungal Biol 115:197–213
Acknowledgements
The authors are grateful to C. Nirma for isolating SNB-CN72 and to R. Constantino for identification of the host termite. Authors are grateful to Prof. M. Catherine Aime (Purdue University, USA) and Dr. Marizeth Groenewald (Westerdijk Fungal Biodiversity Institute, The Netherlands) for granting access to the strain IBL 03150.
Funding
This work has benefited from an “Investissement d’Avenir” grant managed by the Agence Nationale de la Recherche (CEBA, ref. ANR-10-LABX-0025).
The authors gratefully acknowledge financial support from the Iranian Biological Resource Centre (IBRC), ACECR.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
The authors declare that there are no conflicts of interest.
Additional information
Section Editor: Dominik Begerow
Electronic supplementary material
ESM 1
(DOCX 23 kb)
Rights and permissions
About this article
Cite this article
Nasr, S., Lutz, M., Amoozegar, M.A. et al. Graphiola fimbriata: the first species of Graphiolaceae (Exobasidiales, Basidiomycota) described only based on its yeast stage. Mycol Progress 18, 359–368 (2019). https://doi.org/10.1007/s11557-018-1450-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11557-018-1450-1