Abstract
We studied the diversity of arbuscular mycorrhizal fungi (AMF) in semiarid sandy areas of the Great Hungarian Plain representing the westernmost of the Eurasian steppe belt. AM fungal spores were extracted (i) directly from soils collected around 30 Juniperus communis trees of three sampling sites and (ii) from pot trap cultures established with soils from those samples. Altogether spores of 31 AMF species belonging to 15 genera were identified from the field and pot-culture samples. Three taxa could be identified only on the genus level. During the study, one fungus showed a unique combination of morphological characters. Molecular phylogenetic analyses of SSU, ITS, and LSU regions of the nrDNA positioned the fungus in the genus Diversipora and confirmed its novelty when compared with all Diversipora spp. species sequenced to date. This fungus is here described as D. jakucsiae sp. nov.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Arbuscular mycorrhiza (AM) is formed by fungi belonging to the phylum Glomeromycota (Schwarzott et al. 2001) and by plants from almost all main land-plant groups (Wang and Qiu 2006). Although AMF have a worldwide distribution, occur in all climatic regions, and have a long evolutionary history, the species described to date number only ca. 270 (Schüßler and Walker 2010; Schüßler 2013). Molecular diversity screening techniques usually showed more phylotypes than morphotypes revealed by extraction of AMF spores from soil (Helgason et al. 1998; 1999); nevertheless, recent studies report better differentiation of AMF communities by morphological spore identification than by molecular analyses (Oehl et al. 2004; Hijri et al. 2006; Wetzel et al. 2014). Bioassay pot cultures established with mixtures of rhizosphere soil and roots and in planta diversity molecular studies proved that some species of AMF existing inside roots produce extraradical spores only seasonally, rarely, or not at all in the field, so such fungi could go undetected in surveys based only on spores (Błaszkowski 2012; Stutz and Morton 1996). Molecular diversity examinations targeting roots to study in planta diversity of AMF in field studies is the only known way to connect a fungus with its host plant. Nevertheless, time-consuming spore-based investigations are still necessary, because morphologically characterized isolates are needed to describe novel taxa, and collections of AM fungal isolates have great value as potential inocula for both scientific and applied purposes.
Mycorrhizae have special importance in abiotically stressed environments such as arid and semiarid regions and nutrient-limited soils: sandy areas, dunes, etc. Although knowledge on the mycorrhizal fungal communities of different regions of the world has been accumulating, (semi)arid and sandy habitats are still underrepresented. This geographic bias is true for AMF studies (Morton et al. 1995), especially when time-consuming spore-based investigations are considered.
The semiarid sandy areas of interfluvia of the Danube and Tiscia, the two main rivers of Hungary, are the westernmost representatives of the Eurasian steppes with woodland patches. These sandy open grasslands with semi-desert features are among the botanically most intensively studied habitat types in the Carpathian Basin (Biró et al. 2013; Fekete et al. 2002; Kovács-Láng et al. 2000; 2008). As past mycorrhizal status studies showed, characteristic plant communities in the regions are dominated by AM plant species (Kovács and Bagi 2001; Kovács and Szigetvári 2002). Unlike our knowledge of, e.g., the mushrooms and lichens of these habitats (Hollós 1913; Babos 1999; Nagy 2004; Révay and Nagy 2005; Nagy and Gorliczai 2007; Knapp et al. 2012), we had only a limited knowledge of their AM fungal species. Early workers reported AMF species with macroscopic sporocarps (Hollós 1911; Szemere 1965). Takács and Bratek (2006) then reported the first spore-based study of AMF of these sandy areas. They identified six species and Bratek et al. (2013) added two more to the list. The first molecular study of AM fungal diversity in plant roots from the region identified several AM fungal lineages colonizing roots of the sporophyte of the fern Botrychium virginianum (Kovács et al. 2007).
As a part of a complex diversity research project on non-pathogenic root-colonizing fungi of these sandy areas, we studied which AM fungi can be detected from the soil spore bank. We extracted AM fungal spores (i) directly from soils originating from the sampling sites and (ii) from pot trap cultures established with soils from the sampling sites. During the study, one fungus showed a unique combination of morphological characters; subsequent molecular phylogenetic analyses positioned it in the genus Diversispora and confirmed its novelty when compared with all Diversispora spp. species sequenced to date. Below we describe the species as D. jakucsiae sp. nov.
Materials and methods
Sampling sites, sample collection
Soil samples were collected from three sandy sites with characteristic plant communities of the interfluvia of the Danube and Tiscia Rivers near the villages Bugac, Fülöpháza, and Tatárszentgyörgy, Hungary. Detailed descriptions, including photos, of the three sampling sites were published previously (Knapp et al. 2012). The plant communities and soil characteristics of all sites are similar. In August 2007, soil samples were collected around Juniperus communis L. trees at each site; those 30 trees were the targets of our long-term root sampling as well. Four to six soil cores 15–20 cm deep by 5 cm diameter were collected around each tree and mixed thoroughly to mask the differences usually existing around trees standing in open sites. Those 30 soil samples were transferred to the laboratory and air-dried in paper bags at room temperature in a ventilator chamber.
Pot cultures and isolation of spores
Soil samples collected from the field were partitioned and used two ways for screening AMF spores: directly and from pot trap cultures. The latter were established by adding 0.1 volume of soil sample to the composite material (see in Błaszkowski et al. 2013) into which Plantago lanceolata L. was planted as host plant. We followed the method detailed previously by Błaszkowski et al. (2006; Błaszkowski 2012). The trap cultures were harvested 6 months after the establishment.
Single-species cultures were also established and grown as given in Błaszkowski et al. (2012). Briefly, the cultures (six of the 10 established) were successfully established from ca. 10 spores extracted from trap cultures and then stored in water at 4 °C for 10 days. After removal of soils debris, spores of uniform morphology were collected in a pipette, placed at the bottom of a hole ca. 1 cm wide and 4 cm deep, formed in a wetted growing substrate filling 8 cm polystyrene pots (250 cm3), buried in the growing substrate, and finally seeded with ca. 20 seeds of P. lanceolata. The growing substrate of the cultures was autoclaved commercially available coarse-grained sand (grains 1.0–10.0 mm diameter, 80.50 %; grains 0.1–1.0 mm diameter, 17.28 %; grains < 0.1 mm diameter, 2.22 %) mixed (5:1, v/v) with clinopthilolite (Zeocem, Bystré, Slovakia) of grains 2.5–5 mm. Clinopthilolite is a crystaline hydrated alumosilicate of alkali metals and alkaline earth metals having, e.g., high ion exchange capability and selectivity, as well as reversible hydration and dehydration. The sand-clinopthilolite mixture had a pH of 7.3. The cultures were kept in transparent plastic bags, 15 cm wide and 22 cm high as suggested by Walker and Vestberg (1994). The cultures were watered with tap water once or twice a week and harvested for the first time after 4 months when spores were extracted for study. To obtain additional data, later spores were still repeatedly extracted from the same cultures.
Spores were extracted from field samples, trap and single-species cultures by the modified wet sieving and decanting method (Gerdemann and Nicholson 1963). The samples were mixed with tap water using a glass rod; after a few seconds the mixture was poured through sieves with pore sizes 500 μm and 40 μm. Spores from the 40 μm sieve were washed into Petri dishes, then picked from the surface with a needle or sharpened wooden stick and used for (i) identification, (ii) preparation of reference slides, (iii) establishment of single-species pot cultures, or iv) DNA extraction. To reveal mycorrhizal root structures, root fragments located ca. 1–5 cm below the upper level of the growing medium were cut off with a scalpel and stained as described in Błaszkowski (2012).
Microscopy
Characteristics of intact and crushed spores and their wall structure were determined by examining them in water, lactic acid, polyvinyl alcohol/lactic acid/glycerol (PVLG; Omar et al. 1979), and a mixture of PVLG and Melzer’s reagent (1:1, v/v). For species description, at least 100 spores differing in colour and size were studied as described previously (Błaszkowski et al. 2006; Błaszkowski 2012). An Olympus BX 50 compound microscope equipped with Nomarski DIC optics was used for morphological investigation, and photo documentation was carried out with a Sony 3CCD color video camera coupled to the microscope.
Spore structure terminology suggested by Stürmer and Morton (1997) and Walker (1983) was followed. Spore colour was examined under a dissecting microscope on fresh specimens immersed in water and illuminated with light of the same colour and intensity as colour samples given in Methuen Handbook of Colour (Kornerup and Wanscher 1983); colour names are from the same book. Nomenclature of fungi and the authors of fungal names are those presented in the Index Fungorum. Voucher specimens mounted in PVLG and a mixture of PVLG and Melzer’s reagent (1:1, v/v) were deposited in the Department of Ecology and Protection of Environment (DEPE), West Pomeranian University of Technology, Szczecin, Poland, and in the herbarium at Oregon State University (OSC) in Corvallis, Oregon, USA; reference slides of the other AMF taxa detected were also deposited in DEPE.
Molecular methods and phylogenetic analyses
DNA was extracted from three to nine spores as described by Błaszkowski et al. (2013). The extracts were used as template in polymerase chain reactions (PCR). We amplified four segments of the nrDNA. A part of the SSU was amplified with AML1–AML2 primers (Lee et al. 2008). The ITS region was amplified with the primer system designed by Redecker (2000), whereas the partial LSU segment was gained by use of the primers 28G1 and 28G2 (da Silva et al. 2006). To obtain the long SSU–ITS–LSU nrDNA segment we used the primer system designed by Krüger et al. (2009). The PCRs were performed as the original papers recommend and as we described previously (Błaszkowski et al. 2013). Only high fidelity polymerases were used (either HighFidelity Enzyme Mix /Fermentas/ or Phusion /Fermentas/). Amplicons were cloned as described in Błaszkowski et al. (2010). Plasmids containing appropriate amplicons were sequenced by universal primers at LGC Genomics (Berlin, Germany). The SSU–ITS–LSU nrDNA sequences were also gained from three species originally described as Glomus arenarium Błaszk., Tadych & Madej, G. gibbosum Błaszk., and G. insculptum Błaszk. (Błaszkowski 1997; Błaszkowski et al. 2001; 2004a, b). Schüßler and Walker (2010, 2011) placed these fungi among species of uncertain generic position because of the lack of molecular evidence, and Oehl et al. (2011a) transferred them to the genus Diversispora based on morphology of their spores. Spores of these species were extracted from the same single-species cultures from which their holotypes were designated. The sequences were deposited in GenBank (KJ850181–KJ850204).
Pilot analyses were run to test the generic position of the new taxon and the species of uncertain generic affiliation. For final analyses, sequences gained in the present study and collected from public databases were aligned with the online MAFFT v. 7 (Katoh and Standley 2013) by the auto option. The indel positions were coded by means of the simple indel coding algorithm (Simmons et al. 2001) as implemented in GapCoder (Young and Healy 2003), which converts all indels with different starting and/or end positions to a matrix of binary presence/absence characters. Indels showing a complete overlap with a longer indel were coded as unknown characters. Leading and trailing gaps of the alignments were scored as missing data. This binary character set was added to the nucleotide alignment knowing that these characters reinforce the robustness of phylogenetic analyses (Nagy et al. 2012). Bayesian (BI) analyses were performed with MrBayes 3.1.2 (Huelsenbeck and Ronquist 2001; Ronquist and Huelsenbeck 2003) by use of SSU–ITS–LSU sequences plus indel characters divided into four partitions. The GTR + G and two-parameter Markov (Mk2 Lewis) models were applied for the nucleotide partitions and indel matrices, respectively. Four Markov chains were run for 10,000,000 generations, sampling every 1,000 steps, and with a burn in at 3,000 sampled trees. The convergence of the MCMC Bayesian phylogenetic inference was checked by AWTY online (Wilgenbusch et al. 2004).
Maximum likelihood (ML) phylogenetic analyses were carried out with the raxmlGUI (Silvestro and Michalak 2012) implementation of RAxML (Stamatakis 2014) with GTRGAMMA for DNA and default set for binary (indel) characters. Rapid bootstrap analysis with 1,000 replicates was used to test the support of the branches. Phylogenetic trees were visualized and edited in MEGA6 (Tamura et al. 2013).
Results
Study of the spores
Altogether, spores of 31 AMF species were identified from the field and trap cultures. Three taxa could be identified only at the genus level. Based on spore morphological characters, the 31 AMF species represented 15 genera (Table 1). Septoglomus constrictum (Trappe) Sieverd., G.A. Silva & Oehl was most frequently identified and found in all field-collected soil samples and trap cultures (Table 1). Two other frequent species were G. microcarpum Tul. & C. Tul. and Diversispora arenaria (Błaszk., Tadych & Madej) Oehl, G.A. da Silva & Sieverd., extracted only from field samples and trap cultures, respectively (Table 1). Two trap cultures from the Bugac site contained spores of an unknown species, described below as a sp. nov. (Table 1).
Phylogenetic analyses
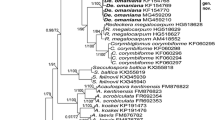
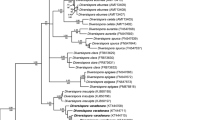
The sequences gained from the isolated AMF assumed to be a new species unambiguously grouped into the genus Diversispora (Fig. 1). When analyzed, the SSU–ITS–LSU sequences separated unambiguously from other taxa and branched in a sister group relation with the fungus Diversispora sp. EE1 analyzed by Thiéry et al. (2012) (Fig. 1). Their clade formed a well supported monophyletic group with D. arenaria and Tricispora nevadensis (Palenz., Ferrol, Azcón-Aguilar & Oehl) Oehl, Palenz., G.A. Silva & Sieverd. (Fig. 1).
50% majority rule consensus phylogram inferred from the Bayesian analysis of nrDNA SSU–ITS–LSU sequences of the new species together with other known AMF species. Corymbiglomus corymbiforme served as outgroup. Numbers above branches are posterior probabilities, below the branches denote bootstrap values from 1,000 replications, shown as percentages. Values below 0.90 and 70 %, respectively, are not shown. Bar indicates 0.5 expected change per site per branch
The results obtained using our ITS and LSU sequences gained separately among the other sequences shown in Fig. 1 did not differ from the results of the analyses of the SSU–ITS–LSU sequences mentioned above (data not shown). The SSU sequences of the putative new species also showed it to belong to Diversispora, but that region is not suitable for species level comparison within this genus. The sequences showed high similarities with the Diversispora VTX00054 of the MaarjAM database (Öpik et al. 2010; 2014).
In addition, the phylogenetic analyses of our SSU–ITS–LSU sequences confirmed Oehl et al.’s (2011a) conclusions drawn from studies of spore morphology that D. arenaria, D. gibbosa (Błaszk.) Błaszk. & Kovács, and D. insculpta (Błaszk.) Oehl, G.A. Silva & Sieverd. belong in the genus Diversispora (Fig. 1).
Taxonomy
Diversispora jakucsiae Błaszk., Balázs & Kovács, sp. nov. Figs. 2–12
Spores of Diversispora jakucsiae. 2. Intact spores with subtending hyphae (sh). 3–6. Spore wall layers (swl) 1–3; on 4. swl1 separated from swl2 due to vigorous crushing of the spore. 7. One-layered hyaline subtending hypha (sh) with septum (s). 8. One-layered hyaline subtending hypha (sh) with septum (s) and spore wall layers (swl) 2 and 3. 9. Spore wall layers (swl) 1–3 and subtending hyphal wall layers (shwl) 1 and 2 continuous with swl1 and 2; note the thick, coloured shwl2 and the septum (s) of the subtending hypha continuous with some innermost laminae of swl2. Mounting: 2: in lactic acid. 3: in PVLG. 4–7 and 9: in PVLG + Melzer’s reagent. 2–9: Differential interference microscopy. Bars: 2: 50 μm, 3–9: 10 μm
Spores and mycorrhizal structures of Diversispora jakucsiae (10–12) and spores of D. arenaria (13–17). 10. Subtending hyphal wall layers (shwl) 1 and 2. 11. Subtending hypha (sh) and spore wall layers (swl) 2 and 3; note swl3 forms septum (s) in the lumen of sh. 12. Mycorrhizal structures of D. jakucsiae in P. lanceolata roots stained in 0.1 % trypan blue: arbuscule (a) and intraradical hyphae (ih) stained faintly. 13–17. Spores of D. arenaria. 13. Intact spores. 14–16. Spore wall layers 1–3; note the deteriorating swl1. 17. Hyaline subtending hypha (sh) with septum (s) and spore wall layers (swl) 2 and 3; swl1 is completely sloughed. Mounting: 13: in lactic acid. 11, 12, 15, 17: in PVLG. 10, 13, 16: in PVLG + Melzer’s reagent. 10–17: Differential interference microscopy. Bars: 13: 100 μm, 10–12 and 14–17: 10 μm
MycoBank MB 808855
Diagnosis: Differs from Diversispora arenaria in the spore wall structure, the phenotypic characters of spore wall layers, and its phylogenetic position.
Type: Poland, Szczecin, under bioassay pot-cultured Plantago lanceolata, 10 May 2009, (Original soil collected: Hungary, Bugac puszta August 2007), Błaszkowski, J., 3222 [Holotype, Department of Ecology and Protection of Environment (DEPE)]; Błaszkowski, J., 3221, 3223–3243 (Istotypes, DEPE) and two slides at OSC.
Etymology: We name this species in honor of Dr. Erzsébet Jakucs, who worked for almost four decades at the Eötvös Loránd University and established and taught several mycological courses for many students. She strongly influenced modern ectomycorrhiza research in Hungary and diligently publicized mycology, recently as president of the Hungarian Mycological Society.
Sporocarps unknown. Spores formed singly in soil and develop blastically at the tip of sporogenous hyphae continuous with extraradical hyphae (Fig. 2). Spores reddish yellow (4A7) to brownish red (8C8), globose to subglobose, (85–)110(−145) μm diam, rarely ovoid, 105–120 × 110–150 μm, with one subtending hypha (Figs. 2, 7–11). Spore wall composed of three layers (layers 1–3, Figs. 3–6, 8, 9, 11). Layer 1, forming the spore surface, permanent, smooth, hyaline to yellowish white (4A2), (0.8–)1.0(−1.3) μm thick (Figs. 3–6, 9). Layer 2, laminate, smooth, reddish yellow (4A7) to brownish red (8C8), (4.2)9.2(14.5) μm thick, frequently swelling slightly in spores crushed in PVLG (Figs. 3–11). Layer 3, flexible to semi-flexible, likely hyaline to lighter than layer 2, (0.6–)1.0(−1.3) μm thick, usually easily separating from the lower surface of layer 2 in crushed spores (Figs. 3–6, 8, 9, 11). None of the spore wall layers reacts in Melzer’s reagent (Figs. 4, 6, 9). Subtending hypha hyaline to orange (5B8), straight or recurved, cylindrical to funnel-shaped, sometimes constricted at the spore base, (5.0–)8.6(−13.5) μm wide at the spore base (Figs. 2, 7–11). Wall of subtending hypha hyaline to orange (5B88), (1.3–)2.9(−4.4) μm thick at the spore base, composed of either one hyaline layer, when spore wall layer 2 begins its development at the spore base (Figs. 8, 9), or two layers, when the coloured spore wall layer 2 starts its development below the spore base (Figs. 9, 10); layer 2 extends up to 13.6 μm below the spore base and sometimes thickens towards the centre of the subtending hyphal lumen near the spore base, making the lumen narrow. Pore (0.6–)2.3(−6.8) μm diam, occluded by a straight or slightly curved septum, 0.8–1.4 μm thick, continuous with some innermost laminae of spore wall layer 2 (Figs. 7–9) and spore wall layer 3 (Fig. 11), which usually tightly adheres to the lower surface of the group of the innermost laminae of spore wall layer 2 that forms the septum in the subtending hypha and thereby this layer is difficult to see; septum positioned at or up to 2.4 μm below the spore base. Spore contents of a hyaline oily substance. Germination unknown.
Mycorrhizal associations. In the field, D. jakucsiae occurred in soils collected around the roots of J. communis. In single-species cultures with P. lanceolata as host plant, D. jakucsiae formed mycorrhizae with arbuscules and intra- and extraradical hyphae (Fig. 12). No vesicles were found. Arbuscules were infrequent and widely dispersed along the root fragments examined. Intraradical hyphae grew parallel to the longitudinal root axis, were straight to slightly curved, (2.0–)4.8(−8.3) μm wide. They frequently formed ellipsoidal coils, 15.5–37.0 × 33.3–106.0 μm when seen in a plan view. Extraradical hyphae occurred very rarely and were (1.6–)2.4(−3.8) μm wide. In 0.1 % trypan blue, arbuscules stained violet white (16A2) to pastel violet (16A4), and intra- and extraradical hyphae and coils violet white (15A2) to lilac (16B5, Fig. 12).
Notes: The most distinctive morphological characters of D. jakucsiae are its reddish yellow to brownish red spores with a three-layered spore wall, of which layer 1, forming the spore surface, is persistent and smooth, the laminate layer 2 frequently swells slightly in PVLG, and therefore, is thicker than in spores crushed in water, and layer 3 is flexible to semi-flexible, likely colorless or lighter than spore wall layer 2, and closes the subtending hyphal pore along with some innermost laminae of spore wall layer 2 (Figs. 2–6, 9, 11). The spore subtending hyphal wall of D. jakucsiae, which may be hyaline, when one-layered (Figs. 7, 8), or coloured, when the coloured spore wall layer 2 begins its development below the spore base and forms subtending hyphal wall layer 2 (Figs. 9, 10), also takes notice. However, the morphological variability of the subtending hyphal wall is within the range of variability of subtending hyphae of Diversispora spp. (Oehl et al. 2011a, b). In addition the species is unique in its molecular phylogeny (Fig. 1).
The expansion of the laminate spore wall layer 2 of D. jakucsiae in PVLG was not given in descriptions of any other Diversispora spp. and those of species of other genera of the Glomeromycota with glomoid spores known to date. We also did not find this phenomenon in any Diversispora spp. kept in our cultures and showed in Fig. 1, except for D. clara. Błaszkowski (2012) and Franke and Morton (1994) stated similar swelling of the structural laminate spore wall layer of Fuscutata heterogama Oehl, F.A. Souza, L.C. Maia & Sieverd. [formerly Scutellospora heterogama (T.H. Nicolson & Gerd.) C. Walker & F.E. Sanders], but only in young spores. With age this layer transformed from an amorphous structure into a compact laminate layer. In D. jakucsiae the laminate spore wall layer 2 swell more frequently in darker-coloured, likely older spores.
As indicated our phylogenetic analyses, of the described species of AMF, D. jakucsiae is closest to D. arenaria (Fig. 1) (Błaszkowski et al. 2001; Oehl et al. 2011a). Both species form single spores (not in clusters or sporocarps) that are similar in size and have a three-layered spore wall, of which none reacts in Melzer’s reagent. However, D. arenaria may produce markedly darker spores, despite their walls may be up to 1.8-fold thinner (Fig. 13; Błaszkowski 2012). In addition spore wall layers 1 and 2 of D. arenaria slough with age (Figs. 14–17; vs. permanent in D. jakucsiae, Figs. 3–6, 9), and spore wall layer 3 of the former species is laminate (Figs. 14–17; vs. flexible to semi-flexible, Figs. 3–6, 8, 9, 11). The spore subtending hypha of D. arenaria is more regular in shape (cylindrical to flared, Fig. 17; vs. cylindrical to funnel-shaped in D. jakucsiae, Figs. 2, 7–11), and at the spore base it is up to 1.7-fold wider and has an up to 2.6-fold thicker wall.
Of the other known Diversispora spp., morphologically D. jakucsiae spores most resemble D. aurantia spores, which also arise singly and are similar in size and the structure of spore wall (Błaszkowski et al. 2004a, b; Błaszkowski 2012). However, mature and even old D. aurantia spores never are brownish red as those of D. jakucsiae (Figs. 2, 5, 6, 10, 11). The phenotypic and histochemical characters of spore wall layers 1–3 of both species are almost identical, except for two features. First, in intact spores mounted in PVLG spore wall layer 1 of D. aurantia frequently balloons and then separates from laminate layer 2 (vs. it always remains adherent to layer 2 in D. jakucsiae spores mounted in PVLG, Figs. 3–6, 9). Second, the laminate spore wall layer 2 of D. aurantia is ca. 1.5-fold thinner than that of D. jakucsiae. In addition the spore subtending hypha of D. aurantia is cylindrical to slightly flared (vs. cylindrical to funnel-shaped in D. jakucsiae, Figs. 2, 7–10) and may be up to 1.6-fold narrower at the spore base.
Discussion
Our study of the AMF spores of semiarid sandy areas in Hungary explored by us resulted in identification of 31 species. There was a kind of tendency in some species whether they were detected in field or trap culture or both. Beside this, the prevalence of the different species differed considerably. However, we should bear in mind that our sampling was not quantitative, and as spores were studied from one sampling, the seasonality of the spore production and inoculum potential might have influenced the results. We could identify ca. 30 AMF taxa by their spores; for the reasons listed above, this species number is surely an underestimation of the AMF present in the area. Takács and Bratek (2006) identified six species of AMF from spores found in the Carpathian Basin, and two additional AM species were reported later (Bratek et al. 2013). Of them, Corymbiglomus corymbiforme Błaszk. & Chwat, G. microcarpum, S. constrictum, and Racocetra persica (Koske & C. Walker) Oehl, F.A. Souza & Sieverd. occurred in Fülöpháza, where we also sampled those species. Both in our study and that of Takács and Bratek (2006) the most common and abundant spores were formed by S. constrictum. We also found C. corymbiforme in Fülöpháza, and G. microcarpum occurred in all three regions explored by us. Interestingly, Scutellospora dipurpurescens J.B. Morton & Koske was found in both study sites but neither we nor Takács and Bratek (2006) reported the species from Fülöpháza. All together Takács and Bratek (2006) reported six AMF species from the sandy grassland region; here we found all and identified 22 additional species.
Schüßler and Walker (2010, 2011) pointed the possible wrong generic assignation of several Glomus species that could not be revised. Here we could gain appropriate sequence information to support the generic position of G. arenarium, G. gibbosum and G. insculptum (Błaszkowski 1997; Błaszkowski et al. 2001; 2004a, b) revised by Oehl et al. (2011a), who transferred them to the genus Diversispora based on morphological studies.
Thiéry et al. (2012) characterized the genetic diversity of a Diversispora sp. strain “EE1” which was found in an inoculum sample of uncertain origin. The distinct lineage of strain EE1 was as a sister group to the new D. jakucsiae. However, not only the phylogenetic analyses, but also certain morphological features also separate the two AMF. Nevertheless, based on the available sequence data and the photo documentation of the spores (unpublished results kindly sent us by Thiery and Öpik), one cannot exclude the possibility that D. jakucsiae and EE1 are conspecific. These two taxa formed a clade with D. arenaria and Tricispora nevadensis. However, T. nevadensis, originally described as Entrophospora nevadensis Palenz., N. Ferrol, Azcón-Aguilar & Oehl (Palenzuela et al. 2010) was defined as forming spores inside a neck of the sporiferous saccule (so called entrophosporoid spores), and not at the tip of sporogenous hyphae (glomoid spores) as all Diversispora spp. known to date do it. Redecker et al. (2013) concluded that the morphology of T. nevadensis was likely erroneously defined; the formation of spores of the species from sporiferous saccules was not proven and the spores could have come from contaminant AMF. Further studies of specimens of T. nevadensis grown in single-species cultures are needed to clarify the doubts.
The SSU region where the NS31–AM1 segment is localized cannot distinguish Diversispora species (Gamper et al. 2009; Stockinger et al. 2010; Schüßler et al. 2011). However, this region is still the most frequent segment used in environmental AMF diversity studies and underlies the VTX-system of the MaarjAM database (Öpik et al. 2010; 2014). When the SSU sequence of D. jakucsiae was compared with the MaarjAM database it was identical with the Diversispora sp. EE1 and unambiguously fit into “VTX00054”. Because of the poor resolution of this sequence in the genus Diversispora it is not surprising that VTX00054 contains the sequences of several described species and environmental samples with diverse geographic origin (Palenzuela et al. 2008; Gamper et al. 2009; Alguacil et al. 2011, 2012; Liu et al. 2011; Oehl et al. 2011a, b; Schüßler et al. 2011; Thiéry et al. 2012; Torrecillas et al. 2012).
References
Alguacil MM, Torrecillas E, Caravaca F, Fernandez DA, Azcon R, Roldan A (2011) The application of an organic amendment modifies AMF communities colonizing native and non-native seedlings grown in a heavy metal polluted soil. Soil Biol Biochem 43:1498–1508. doi:10.1016/j.soilbio.2011.03.026
Alguacil MM, Torrecillas E, Roldan A, Diaz G, Torres MP (2012) Perennial plant species from semiarid gypsum soils support higher AMF diversity in roots than the annual Bromus rubens. Soil Biol Biochem 49:132–138. doi:10.1016/j.soilbio.2012.02.024
Babos M (1999) Higher fungi (Basidiomycotina) of the Kiskunság National Park and its environs. In: Lőkös L, Rajczy M (eds) The flora of the Kiskunság National Park, II. Cryptogams. MTM, Budapest, pp 199–298
Biró M, Szitár K, Horváth F, Bagi I, Molnár Z (2013) Detection of long-term landscape changes and trajectories in a Pannonian sand region: comparing land-cover and habitat-based approaches at two spatial scales. Community Ecol 14:219–230. doi:10.1556/ComEc.14.2013.2.12
Błaszkowski J (1997) Glomus gibbosum, a new species from Poland. Mycologia 89:339–345
Błaszkowski J, Tadych M, Madej T (2001) Glomus arenarium, a new species in Glomales (Zygomycetes). Acta Soc Bot Pol 70:97–101
Błaszkowski J, Adamska I, Czerniawska B (2004a) Glomus insculptum, a new arbuscular mycorrhizal species from Poland. Mycotaxon 89:225–234
Błaszkowski J, Blanke V, Renker C, Buscot F (2004b) Glomus aurantium and G. xanthium, new species in Glomeromycota. Mycotaxon 90:447–467
Błaszkowski J, Renker C, Buscot F (2006) Glomus drummondii and G. walkeri, two new species of arbuscular mycorrhizal fungi (Glomeromycota). Mycol Res 110:555–566. doi:10.1016/j.mycres.2006.02.006
Błaszkowski J, Kovács GM, Balázs KT, Orłowska E, Sadravi M, Wubet T, Buscot F (2010) Glomus africanum and G. iranicum, two new species of arbuscular mycorrhizal fungi (Glomeromycota). Mycologia 102:1450–1462. doi:10.3852/09-302
Błaszkowski J (2012) Glomeromycota. W. Szafer Institute of Botany, Polish Academy of Sciences, Kraków
Błaszkowski J, Kovács GM, Gáspár BK, Balázs TK, Buscot F, Ryszka P (2012) The arbuscular mycorrhizal Paraglomus majewskii sp. nov. represents a new distinct basal lineage in Paraglomeraceae (Glomeromycota). Mycologia 104:148–156. doi:10.3852/10-430
Błaszkowski J, Chwat G, Kovács GM, Gáspár BK, Ryszka P, Orłowska E, Pagano MC, Araújo FS, Wubet T, Buscot F (2013) Septoglomus fuscum and S. furcatum, two new species of arbuscular mycorrhizal fungi (Glomeromycota). Mycologia 105:670–680. doi:10.3852/12-127
Bratek Z, Merényi Z, Varga T (2013) Changes of hypogeous funga in the Carpathian-Pannonian region in the past centuries. Acta Mycol 48:33–39. doi:10.5586/am.2013.005
da Silva GA, Lumini E, Maia LC, Bonfante P, Bianciotto V (2006) Phylogenetic analysis of Glomeromycota by partial LSU rDNA sequences. Mycorrhiza 16:183–189. doi:10.1007/s00572-005-0030-9
Fekete G, Molnár Z, Kun A, Botta-Dukát Z (2002) On the structure of the Hungarian forest steppe: grasslands on sand. Acta Zool Hung 48:137–150
Franke M, Morton JB (1994) Ontogenetic comparisons of arbuscular mycorrhizal fungi Scutellospora heterogama and Scutellospora pellucida: revision of taxonomic character concepts, species descriptions, and phylogenetic hypotheses. Can J Bot 72:122–134
Gamper HA, Walker C, Schüßler A (2009) Diversispora celata sp. nov: molecular ecology and phylotaxonomy of an inconspicuous arbuscular mycorrhizal fungus. New Phytol 182:495–506. doi:10.1111/j.1469-8137.2008.02750
Gerdemann JW, Nicholson TH (1963) Spores of mycorrhizal Endogone species extracted from soil by wet sieving and decanting. Trans Brit Mycol Soc 46:235–244. doi:10.1016/S0007-1536(63)80079-0
Helgason T, Daniell TJ, Husband R, Fitter AH, Young JPW (1998) Ploughing up the wood-wide web? Nature 394:431. doi:10.1038/28764
Helgason T, Fitter AH, Young JPW (1999) Molecular diversity of arbuscular mycorrhizal fungi colonizing Hyacinthoides non-scripta (bluebell) in a seminatural woodland. Mol Ecol 8:659–666. doi:10.1046/j.1365-294x.1999.00604.x
Hijri I, Sykorova Z, Oehl F, Ineichen K, Mäder P, Wiemken A, Redecker D (2006) Communities of arbuscular mycorrhizal fungi in arable soils are not necessarily low in diversity. Mol Ecol 15:2277–2289. doi:10.1111/j.1365-294X.2006.02921.x
Hollós L (1911) Magyarország földalatti gombái, szarvasgombaféléi (Fungi hypogaei Hungariae). Budapest
Hollós L (1913) Kecskemét vidékének gombái. (Fungi of Kecskemét and its country-side) Budapest
Huelsenbeck JP, Ronquist F (2001) MrBayes: Bayesian inference of phylogeny. Bioinformatics 17:754–755. doi:10.1093/bioinformatics/17.8.754
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30:772–780. doi:10.1093/molbev/mst010
Knapp DG, Pintye A, Kovács GM (2012) The dark side is not fastidious – Dark septate endophytic fungi of native and invasive plants of semiarid sandy areas. PLoS ONE 7:e32570. doi:10.1371/journal.pone.0032570
Kornerup A, Wanscher JH (1983) Methuen handbook of colour, 3rd edn. Eyre Methuen, London
Kovács GM, Bagi I (2001) Mycorrhizal status of a mixed deciduous forest from the Great Hungarian Plain with special emphasis on the potential mycorrhizal partners of Terfezia terfezioides (Matt.) Trappe. Phyton 41:161–168
Kovács GM, Szigetvári C (2002) Mycorrhizae and other root-associated fungal structures of the plants of a sandy grassland ont the Great Hungarian Plain. Phyton 42:211–223
Kovács GM, Balázs T, Pénzes Z (2007) Molecular study of the arbuscular mycorrhizal fungi colonizing the sporophyte of the eusporangiate rattlesnake fern (Botrychium virginianum, Ophioglossaceae). Mycorrhiza 17:597–605. doi:10.1007/s00572-007-0137-2
Kovács-Láng E, Kröel-Dulay G, Kertész M, Fekete G, Mika J et al (2000) Changes in the composition of sand grasslands along a climatic gradient in Hungary and implications for climate change. Phytocoenologia 30:385–407. doi:10.1127/phyto/30/2000/385
Kovács-Láng E, Molnár E, Kröel-Dulay G, Barabás S (2008) The KISKUN LTER: Long-term ecological research in the Kiskunság, Hungary. Hungarian Academy of Sciences, Institue of Ecology and Botany, Vácrátót
Krüger M, Stockinger H, Krüger C, Schüßler A (2009) DNA-based species level detection of Glomeromycota: one PCR primer set for all arbuscular mycorrhizal fungi. New Phytol 183:212–223. doi:10.1111/j.1469-8137.2009.02835.x
Liu Y, He J, Shi G, An L, Öpik M, Feng H (2011) Diverse communities of arbuscular mycorrhizal fungi inhabit sites with very high altitude in Tibet Plateau. FEMS Microbiol Ecol 78:355–365. doi:10.1111/j.1574-6941.2011.01163.x
Lee J, Lee S, Young JPW (2008) Improved PCR primers for the detection and identification of arbuscular mycorrhizal fungi. FEMS Microbiol Ecol 65:339–349. doi:10.1111/j.1574-6941.2008.00531.x
Morton JB, Bentivenga SP, Bever JD (1995) Discovery, measurement, and interpretation of diversity in symbiotic endomycorrhizal fungi. Can J Bot 73S:25–32. doi:10.1139/b95-221
Nagy L (2004) Fungisztikai vizsgálatok az Alföldön 1997 és 2003 között. Mikol Közlem Clusiana 43:15–46
Nagy L, Gorliczai Z (2007) Újabb adatok az Alföld gombavilágához. Mikol Közlem Clusiana 46:211–256
Nagy LG, Kocsube S, Csana Z, Kovács GM, Petkovits T, Vágvölgyi C, Papp T (2012) Re-mind the gap! insertion – deletion data reveal neglected phylogenetic potential of the nuclear ribosomal internal transcribed spacer (ITS) of fungi. PLoS ONE 7:e49794. doi:10.1371/journal.pone.0049794
Oehl F, Sieverding E, Mäder P, Dubois D, Ineichen K, Boller T, Wiemken A (2004) Impact of long-term conventional and organic farming on the diversity of arbuscular mycorrhizal fungi. Oecologia 138:574–583. doi:10.1007/s00442-003-1458-2
Oehl F, da Silva GA, Goto BT, Sieverding E (2011a) Glomeromycota: three new genera, and glomoid species reorganized. Mycotaxon 116:75–120. doi:10.5248/116.75
Oehl F, da Silva GA, Sanchez-Castro I, Goto BT, Maia LC, Vieira HEE, Barea JM, Sieverding E, Palenzuela J (2011b) Revision of Glomeromycetes with entrophosporoid and glomoid spore formation, with two genera nova. Mycotaxon 117:297–316. doi:10.5248/117.297
Omar MB, Bollan L, Heather WA (1979) A permanent mounting medium for fungi. Bull Br Mycol Soc 13:31–32
Öpik M, Vanatoa A, Vanatoa E, Moora M, Davison J, Kalwij JM, Reier U, Zobel M (2010) The online database MaarjAM reveals global and ecosystemic distribution patterns in arbuscular mycorrhizal fungi (Glomeromycota). New Phytol 188:223–241. doi:10.1111/j.1469-8137.2010.03334.x
Öpik M, Davison J, Moora M, Zobel M (2014) DNA-based detection and identification of Glomeromycota: the virtual taxonomy of environmental sequences. Botany 92:135–147. doi:10.1139/cjb-2013-0110
Palenzuela J, Barea JM, Ferror N, Azcón-Aguilar C, Oehl F (2010) Entrophospora nevadensis, a new arbuscular mycorrhizal fungus from Sierra Nevada national Park (southeastern Spain). Mycologia 102(3):624–632
Palenzuela J, Ferrol N, Boller T, Azcon-Aguilar C, Oehl F (2008) Otospora bareai, a new fungal species in the Glomeromycetes from a dolomitic shrub land in Sierra de Baza National Park (Granada, Spain). Mycologia 100:296–305. doi:10.3852/mycologia.100.2.296
Redecker D (2000) Specific PCR primers to identify arbuscular mycorrhizal fungi within colonized roots. Mycorrhiza 10:73–80. doi:10.1007/s005720000061
Redecker D, Schüβler A, Stockinger H, Stürmer SL, Morton JB, Walker C (2013) An evidence-based consensus for the classification of arbuscular mycorrhizal fungi (Glomeromycota). Mycorrhiza 23:515–531. doi:10.1007/s00572-013-0486-y
Révay Á, Nagy L (2005) Myxomycetes data from the Danube-Tisza interfluve and some other parts of Hungary. Stud Bot Hung 36:117–121
Ronquist F, Huelsenbeck JP (2003) MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19:1572–1574. doi:10.1093/bioinformatics/btg180
Schüßler A, Walker C (2010) The Glomeromycota: a species list with new families and new genera. + Corrigendum (2011) Schüßler A, Walker C Gloucester. Published in libraries at The Royal Botanic Garden Edinburgh, The Royal Botanic Garden Kew, Botanische Staatssammlung Munich, and Oregon State University
Schüßler A, Krüger M, Walker C (2011) Revealing natural relationships among arbuscular mycorrhizal fungi: culture line BEG47 represents Diversispora epigaea not Glomus versiforme. PLoS ONE 6:e23333. doi:10.1371/journal.pone.0023333
Schüßler A (2013) Phylogeny and taxonomy of Glomeromycota. http://schuessler.userweb.mwn.de/amphylo/. Accessed 15 June 2013
Schwarzott D, Walker C, Schüßler A (2001) Glomus, the largest genus of the arbuscular mycorrhizal fungi (Glomales) is nonmonophyletic. Mol Phylogenet Evol 21:190–197. doi:10.1006/mpev.2001.1007
Silvestro D, Michalak I (2012) raxmlGUI: a graphical front-end for RAxML. Org Divers Evol 12:335–337. doi:10.1007/s13127-011-0056-0
Simmons MP, Ochoterena H, Carr TG (2001) Incorporation, relative homoplasy, and effect of gap characters in sequence-based phylogenetic analyses. Syst Biol 50:454–462. doi:10.1080/106351501300318049
Stamatakis A (2014) RAxML Version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinforma (Online Early). doi:10.1093/bioinformatics/btu033
Stockinger H, Krüger M, Schüßler A (2010) DNA barcoding of arbuscular mycorrhizal fungi. New Phytol 187:461–474. doi:10.1111/j.1469-8137.2010.03262.x
Stutz JC, Morton JB (1996) Successive pot cultures reveal high species richness of arbuscular mycorrhizal fungi in arid ecosystems. Can J Bot 74:1883–1889. doi:10.1139/b96-225
Stürmer SL, Morton JB (1997) Developmental patterns defining morphological characters in spores of four species in Glomus. Mycologia 89:72–81. doi:10.2307/3761174
Szemere L (1965) Die unterirdischen Pilze des Karpatenbeckens. Akadémiai Kiadó, Budapest
Takács T, Bratek Z (2006) Description of AM fungi species from semiarid open sandy grasslands in Hungary. Acta Bot Hung 48:179–188. doi:10.1556/ABot. 48.2006.1-2.16
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: Molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729. doi:10.1093/molbev/mst197
Thiéry O, Moora M, Vasar M, Zobel M, Öpik M (2012) Inter- and intrasporal nuclear ribosomal gene sequence variation within one isolate of arbuscular mycorrhizal fungus, Diversispora sp. Symbiosis 58:135–147. doi:10.1007/s13199-012-0212-0
Torrecillas E, Alguacil MM, Roldan A (2012) Host preferences of arbuscular mycorrhizal fungi colonizing annual herbaceous plant species in semiarid mediterranean prairies. Appl Environ Microbiol 78:6180–6186. doi:10.1128/AEM. 01287-12
Walker C (1983) Taxonomic concepts in the Endogonaceae: spore wall characteristics in species descriptions. Mycotaxon 18:443–455
Walker C, Vestberg M (1994) A simple and inexpensive method for producing and maintaining closed pot cultures of arbuscular mycorrhizal fungi. Agric Sci Finl 3:233–240
Wang B, Qiu YL (2006) Phylogenetic distribution and evolution of mycorrhizas in land plants. Mycorrhiza 16:299–363. doi:10.1007/s00572-005-0033-6
Wetzel K, da Silva GA, Matczinski U, Oehl F, Fester T (2014) Superior differentiation of arbuscular mycorrhizal fungal communities from till and no-till plots by morphological spore identification when compared to T-RFLP. Soil Biol Biochem 72:88–96. doi:10.1016/j.soilbio.2014.01.033
Wilgenbusch JC, Warren DL, Swofford DL (2004) AWTY: A system for graphical exploration of MCMC convergence in Bayesian phylogenetic inference. Available from http://ceb.csit.fsu.edu/awty [accessed 7 January 2014]
Young ND, Healy J (2003) GapCoder automates the use of indel characters in phylogenetic analysis. BMC Bioinforma 4:6. doi:10.1186/1471-2105-4-6
Acknowledgments
The authors are grateful to Drs. Odile Thiéry and Maarja Öpik for kindly sharing unpublished results about Diversispora sp. EE1. Fritz Oehl’s and James M. Trappe’s valuable comments on the manuscript are highly appreciated. This study was supported by the Hungarian Scientific Research Fund (OTKA K72776) and Polish National Centre of Science, grants no. 2012/05/B/NZ8/00498 and 2012/07/N/NZ8/02363. The study was partly conducted during T. K. Balázs’ research stay supported by a research scholarship of FEMS at Prof. Janusz Blaszkowski’s laboratory. G. M. Kovács is supported by a János Bolyai Research Scholarship of the Hungarian Academy of Sciences.
Author information
Authors and Affiliations
Corresponding author
Additional information
Tímea K. Balázs and Janusz Błaszkowski equally contributed.
Rights and permissions
About this article
Cite this article
Balázs, T.K., Błaszkowski, J., Chwat, G. et al. Spore-based study of arbuscular mycorrhizal fungi of semiarid sandy areas in Hungary, with Diversispora jakucsiae sp. nov.. Mycol Progress 14, 1021 (2015). https://doi.org/10.1007/s11557-014-1021-z
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11557-014-1021-z