Abstract
Genetic analysis and fine mapping of a resistance gene against brown planthopper (BPH) biotype 2 in rice was performed using two F2 populations derived from two crosses between a resistant indica cultivar (cv.), AS20-1, and two susceptible japonica cvs., Aichi Asahi and Lijiangxintuanheigu. Insect resistance was evaluated using F1 plants and the two F2 populations. The results showed that a single recessive gene, tentatively designated as bph19(t), conditioned the resistance in AS20-1. A linkage analysis, mainly employing microsatellite markers, was carried out in the two F2 populations through bulked segregant analysis and recessive class analysis (RCA), in combination with bioinformatics analysis (BIA). The resistance gene locus bph19(t) was finely mapped to a region of about 1.0 cM on the short arm of chromosome 3, flanked by markers RM6308 and RM3134, where one known marker RM1022, and four new markers, b1, b2, b3 and b4, developed in the present study were co-segregating with the locus. To physically map this locus, the bph19(t)-linked markers were landed on bacterial artificial chromosome or P1 artificial chromosome clones of the reference cv., Nipponbare, released by the International Rice Genome Sequencing Project. Sequence information of these clones was used to construct a physical map of the bph19(t) locus, in silico, by BIA. The bph19(t) locus was physically defined to an interval of about 60 kb. The detailed genetic and physical maps of the bph19(t) locus will facilitate marker-assisted gene pyramiding and cloning.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The brown planthopper (Nilaparvata lugens Stål, BPH), with characteristics of migration, paroxysm and rampancy, is one of the most destructive insect pests in the rice-producing areas, especially for shift to short-stature and heavy-tilling rice cultivars (cvs.) (Dyck and Thomas 1979). The severity of damage and the frequency of outbreak have increased since 1960s, because of the use of greater quantities of nitrogenous fertilizers and insecticides (Pathak 1972; Sogawa 1982, Sogawa et al. 2003; Holt et al. 1996). In China, this insect has caused yield losses of over 500,000 tons each year (Zhu et al. 2004b). Using resistant cvs, which is friendly to the environment than chemical pesticides, has proven to be one of the most efficient ways to control this pest (Pataki 1969; Sogawa 1982).
It is well known that the native indica cvs. expresses a much higher level of resistance than that of another subspecies japonica cvs. Up to now, about nine major BPH resistance genes were identified in the diversified indica cvs. Athwal et al. (1971) identified the first dominant gene Bph1 and the first recessive gene bph2 in cvs. Mudgo and Karsamba Red ASD7, respectively, through genetic segregation analysis. Later, Hirabayashi and Ogawa (1995) identified the Bph1 in cv. IR28 on the long arm of chromosome 12 through restriction fragment-length polymorphism (RFLP) analysis. As for the bph2 gene, Murata et al. (1998) and Murai et al. (2001) also assigned bph2 on the long arm of chromosome 12 through RFLP and amplified fragment-length polymorphism analyses. Lakshminarayana and Khush (1977) identified the second dominant gene Bph3 and the second recessive gene bph4 in cvs. Rathu Heenati and Babawee, respectively, through genetic segregation analysis. Later, Huang (2003) mapped the Bph3 gene to the long arm of chromosome 4 through simple sequence repeat (SSR) and position-specific microsatellite (PSM) analysis. Kawaguchi et al. (2001) mapped the bph4 gene on the short arm of chromosome 6 through linkage analysis with SSR and RFLP markers. Khush et al. (1985), and Kabir and Khush (1988) reported that three new resistant genes, bph5, Bph6 and bph7, were found in rice cvs. ARC10550, Swarnalata and T12, respectively, through genetic segregation analysis. Nemoto et al. (1989) reported that three cvs., Col. 5 Thailand, Col. 11 Thailand and Chin saba, carry the same resistant gene bph8(t), and Bph9(t) in other three cvs., Balamawee, Kaharamana and Pokkali, through allelism tests. Murata et al. (2001) later identified Bph9 on the long arm of chromosome 12, through RFLP and random-amplified polymorphic DNA (RAPD) analyses.
The rice wild species have been considered to be the most important resistant resources other than the native indica cvs for the BPH. Up to date, about nine BPH resistance genes have been identified in the various wild species. Ishii et al. (1994) first identified a resistance gene, Bph10, in an introgression line IR65482-4-136-2-2 derived from the wild species Oryza australiensis, on the long arm of chromosome 12, through RFLP analysis. Yang et al. (2002) identified the resistance gene Bph12(t) on the short arm of chromosome 4 in cv. B14, which is derived from O. latifolia, through SSR and RFLP analysis. Liu et al. (2001) identified a major dominant gene, Bph13(t) on the long arm of chromosome 2 in an O. eichingeri-derived line, acc105159, through RFLP and SSR analyses. Renganayaki et al. (2002) identified a resistance gene, which was also designated as Bph13(t), in IR54745-2-21-12-17-6, a line with O. officinalis-derived resistance to BPH biotype 4, on the short arm of chromosome 3, through RAPD analysis. Huang et al. (2001) reported that B5, a highly resistant line that derived its resistant genes from O. officinalis, carries two major resistance genes, Qbp1 and Qbp2 (later named Bph14 and Bph15, respectively) on the long arm of chromosome 3 and the short arm of chromosome 4, respectively, through linkage analysis and quantitative trait loci (QTL) analysis with RFLP markers. Jena et al. (2005) mapped the resistance gene Bph18(t) on the subterminal region of the long arm of chromosome 12 in an O. australiensis-derived line, IR65482-7-216-1-2, through SSR and sequence-tagged site (STS) analysis, in combination with statistical analysis. Hirabayashi et al. (1998) also found two recessive genes, bph11(t) and bph12(t) on the long arm of chromosome 3 and the middle arm of chromosome 4, respectively, in introgression lines of O. officinalis, through RFLP analysis.
Consequently, a total of 18 BPH resistance genes have been identified and about half of these resistant genes were derived from the wild species. It is noteworthy that nine resistance genes assigned Bph1 to Bph9 were identified by classical genetic approach in the 1970s and 1980s, of which Bph1, bph2, Bph3, bph4 and Bph9 were further identified by molecular genetic approach in the 1990s and 2000s. However, the other nine resistance genes assigned Bp10 to Bph18 were identified by molecular genetic approach, only, from the 1990s. This is partially due to the fact that allelism test is a tedious task, especially in the case that the donor cvs. of resistance genes cannot be exchanged, freely, and that the number of known resistance genes are strikingly increased. On the other hand, the target gene may be more reliable to differentiate from other BPH gene(s) by comparing the relative map positions determined.
On the other hand, genetic mapping studies of these 18 resistance genes have led to rice and the BPH becoming ideal model systems for the study of interactions between plants and sucking herbivorous insects (Yang et al. 2004). However, the only example of a cloned resistance gene to a pest is the Mi gene, and none of the BPH resistance genes have been cloned in crop plants (Vos et al. 1998). Therefore, cloning BPH resistance genes and elucidating its resistance mechanism in rice become very necessary. As we know, map-based cloning is an effective method to get the target genes. However, this way is often tedious and time-consuming, particularly in the process of construction and screening of artificial chromosome library, such as bacterial artificial chromosome (BAC) or P1 artificial chromosome (PAC, Peters et al. 2003; Chen et al. 2005; Liu et al. 2005). Fortunately, the availability and utilization of the sequence information for the rice whole-genome of two subspecies, i.e. the japonica reference cv. Nipponbare released by the International Rice Genome Sequencing Project (IRGSP) (http://rgp.dna.affrc.go.jp) and the indica reference cv. 93-11 released by Beijing Genomics Institute (BGI) (http://www.genomics.org.cn), have made map-based cloning in rice much more efficient. That is, bioinformatics analysis (BIA) of the reference sequences using various software tools has enabled map-based cloning as a routine work in the laboratories related. Doubtlessly, sequence-based map of the target gene is a crucial step for both marker-assisted gene pyramiding and cloning (Schuler 1998; Pan et al. 2003; Gu et al. 2004; Yang et al. 2004).
In the present study, we analyzed the inheritance pattern of a recessive resistance gene against BPH in an indica cv. AS20-1, which is tentatively named bph19(t), and finely mapped to a 60-kb interval, on rice chromosome 3, using PCR-based markers including SSR, STS and CAPS (cleaved amplified polymorphic sequence) markers.
Materials and methods
Plant materials
An indica rice (Oryza sativa L.) cv. AS20-1, which is resistant to the BPH biotype 2, and two susceptible japonica cvs., Aichi Asahi and Lijiangxintuanheigu (LTH), were used as the parental cvs. in this study. Two F2 populations, F2-1 and F2-2, derived from crosses AS20-1/LTH and AS20-1/Aichi Asahi, which consist of 1,330 and 1,403 plants, respectively, were used as a mapping population. For determining the inheritance pattern of the BPH resistance in cv. AS20-1, 13 F1 plants derived from the cross of AS20-1/LTH were also tested. To confirm the success of infestation and objectively evaluate the progeny plants, the parental cvs. and the indica cv. Taichung Native 1 (TN1), which is highly susceptible to all biotypes of the BPH, were used as resistant and susceptible controls.
Insects
The BPH colony used for the research was biotype 2, which was identified by using a set of international testers (Li et al. 1999), and kindly provided by Dr. Huang FK at Guangxi Academy of Agricultural Science, China. The biotype 2 was recognized as a predominant biotype in most of the rice fields in China since 1990s (Wang et al. 1999). The insects were fed on TN1 for about 5 weeks to produce a sufficiently large population, and second-to-third-instar nymphs were selected for infestation. The rest were maintained with alternative plants of TN1 for subsequent infestation.
Evaluation of the BPH resistance
The evaluation of BPH resistance was conducted, using the standard seedbox screening technique (IRRI 1988) with some modifications. At the 3.5- to 4-leaf stage, half a leaf of each seedling was clipped for DNA extraction after removing the weak plants. On the next day, the tray was transferred into an iron sheet case (80×55×8 cm3), water, 2- to 3-cm deep, being flowed into the tray, for keeping humidity high and ants off the seedlings. Then, the plants were infested with the selected nymphs at a density of six to seven nymphs per seedling as evenly as possible. After infestation, each tray was covered with a nylon-net cage (size: 100 meshes/cm2; 65×45×50 cm3), immediately. To objectively evaluate the F2 plants, all the plants in a tray were rated individually every 2 days from the fourth day of infestation. The scoring system proposed by the International Rice Research Institute (IRRI 1988) with some modifications was used to rate each seedling: 0 = no visible damage; 1 = partial yellowing of first leaf; 3 = first and second leaves partially yellow; 5 = pronounced yellowing or some stunting; 7 = mostly wilting, the plant was still alive; 9 = the plant completely wilted or died. Since the damage of severities of plants is dynamic, the score of each seedling rated on a day when more than 90% of the susceptible parents died, was recognized as its reaction. The mean value of damage severity scores for parental and F1 plants were used as their reaction. For data analysis, plants with a scale of 0–5 and 7–9 were designated as resistant and susceptible, respectively. Segregation in the two F2 populations was tested by Chi-square analysis for goodness of fit to the appropriate ratio.
Initial localization of the BPH resistance gene locus
Genomic DNA, from the clipped leaves of F2 plants as well as the parental plants, was extracted, using the CTAB method (Murray and Thompson 1980).
For SSR analysis, amplification reactions were performed with the Programmable Thermal Controller PTC 100TM (MJ Research Inc., Watertown, MA, USA) or the GeneAmp® PCR System 9700 (PE Applied Biosystems, Foster City, CA, USA). Primers of the SSR markers (Temnykh et al. 2000) were synthesized by Shanghai Sangon Biotechnology Co., Ltd. (Shanghai, China). The PCR procedures described by Temnykh et al. (2000) were followed, with the exception that amplified PCR products were separated on 6.0% polyacrylamide denaturing gels and visualized by means of silver staining (Zhu et al. 2004a).
The bulked segregant analysis (BSA, Michelmore et al. 1991) was used to identify the candidate SSR markers linked to the BPH resistance gene. According to the results of the BPH resistance evaluation, two contrasting bulks were prepared, each containing DNA from 10 extremely resistant (score: 0 or 1) or susceptible (score: 9) F2 plants (both were selected from the F2-1 population). Based on the results of the segregation analysis, candidate markers were validated by means of the recessive class analysis (RCA, Pan et al. 2003), using 497 extremely resistant individuals of the F2-1 and F2-2 populations (Fig. 1, and also see subsequently). The recombination frequency between a marker and the resistance gene was calculated using the following formula: c = N r/2N, in which N is the total number of the resistant individuals tested, N r is the number of recombination events occurred at the respective locus (Pan et al. 2003; Gu et al. 2004). The recombination frequency was transformed into centimorgans according to the Kosambi function (Kosambi 1944).
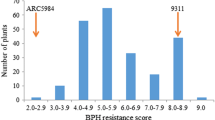
Distribution of the brown planthopper (BPH) resistance scores in two F2 populations of AS20-1/Aichi Asahi and AS20-1/Lijiangxintuanheigu (LTH). The arrows with number 1, 2, 3 and 4 represent the average severities of resistant cultivar (cv.) AS20-1, F1, and susceptible cvs. Aichi Asahi and LTH, respectively
Fine mapping of the BPH resistance gene locus
The BPH resistance gene locus was finely mapped by BIA. On the one hand, known SSR markers, covering the BPH resistance gene region, were adopted from the public database released by the International Rice Microsatellite Initiative (IRMI, http://www.gramene.org/microsat). On the other hand, additional PCR-based markers were subsequently developed according to the sequence information of the reference cvs., Nipponbare and 93–11. For SSR marker, the putative simple-sequence repeat motifs were searched and identified using Simple Sequence Repeat Identification Tool (SSRIT, http://www.gramene.org/db/searches/ssrtool). For STS markers, the putative sequences, which are highly divergent between the two reference genomes, were targeted. All PCR primers were designed using Primer Premier 5.0 (http://www.premierbiosoft.com) and were synthesized by SBS Genetech Co., Ltd. (Beijing, China). The amplification products of the monomorphic STS marker developed were digested with the appropriate restriction enzyme, and converted into CAPS markers, if possible.
Physical mapping of the BPH resistance gene locus
Physical map of the BPH resistance gene locus was constructed, in silico, via BIA. Molecular markers linked with the BPH resistance gene locus were landed on the BAC or PAC clones of the reference cv. Nipponbare, released by IRGSP, using sequence homology search tool, BLASTN (http://www.ncbi.nlm.nih.gov/BLAST). Sequences of these clones were downloaded and aligned using sequence olignment tool, Pairwise BLAST (http://www.ncbi.nlm.nih.gov/blast /bl2seq/bl2.html) for constructing the BAC/PAC contigs spanning the BPH resistance gene locus.
Results
Genetic analysis of the BPH resistance and construction of a mapping population
When more than 90% of susceptible parental plants died, the average severity scores of the three parents, AS20-1, Aichi Asahi and LTH, were 3.88, 8.56 and 8.89, respectively. The average severity score of the 13 F1 plants was 8.43, which resembled to the susceptible parents Aichi Asahi and LTH, indicating that the BPH resistance in AS20-1 was under recessive gene(s) control.
To further confirm the inheritance pattern of the BPH resistance, the two F2 populations were separately tested with the same biotype of insects. In the F2-1 population, the damage severity scores of the 1,330 individuals ranged from 0 to 9, and performed a bimodal distribution (Fig. 1). The resistant and susceptible plants well fitted a 1:3 ratio (347R: 983S, χ 2=0.79, P>0.05), suggesting that a recessive gene confers the BPH resistance in cv. AS20-1 (Fig. 1). In the F2-2 population, a total of 1,403 individuals, also produced the similar result (1R: 3S = 371: 1032, χ 2=1.48, P>0.05), which further confirmed the BPH resistance in AS20-1 conditioned by a single recessive gene (Fig. 1). Since the resistance gene in cv. AS20-1 showed essentially the identical performance in the two F2 populations, the mapping population for the resistance gene was constructed by pooling the two F2 populations.
Identification of SSR makers linked with the BPH resistance gene locus
To efficiently identify the candidate SSR markers linked with the BPH resistance, the BSA method was employed. A total of 142 SSR markers, selected from 12 chromosomes with an interval of ≈15 centimorgans (cM), were tested. Among the selected markers, only three markers, RM231, RM545 and RM7, were polymorphic between both two parents and bulks. These three markers are positioned in a contiguous region on the short arm of chromosome 3 (Fig. 2) and, therefore, considered as candidate markers for the resistance gene. For confirming the candidate markers, the plants consisting of the resistant and susceptible bulks were tested, individually. The results showed that these three markers were linked with the resistance gene (data not shown). Additional 4 SSR markers, RM517, RM546, RM36 and RM218, around these three markers were selected to survey for polymorphism, of which two markers i.e. RM517 and RM218 were also polymorphic between both two parents and bulks.
a Genetic map of the bph19(t) locus and three other BPH resistance genes/QTLs on the short arm of chromosome 3. The locations of three genes/QTLs other than bph19(t) identified in the present research were estimated based on the data from a, Renganayaki et al. 2002; b, Xu et al. 2002; and c, Alam and Cohen 1998. CEN: centromere. b Finely genetic map of the bph19(t) locus. The distance (cM) between markers is indicated to the left; markers used in this phase of the study are indicated to the right; d: new markers used in the present paper
The candidate markers identified by BSA were used for linkage analysis in the mapping population consisting of 497 extremely resistant plants by the RCA. The results showed that most of the recombinant plants identified at the RM231, RM545 and RM517 loci were distinguished from those of RM218 and RM7 loci (data not shown). It, therefore, revealed that the resistance gene locus was flanked by these markers on both sides. RM231, RM545 and RM517 were linked to the BPH resistance locus on telomere side with genetic distances of 11.7, 7.1 and 2.0 cM, respectively, while RM218 and RM7 were found to be 3.8 and 6.5 cM, respectively, on the centromere side (Fig. 2a). According to this result, the BPH resistance locus was roughly mapped within a 5.8-cM region flanked by markers RM517 and RM218 (Fig. 2a), and was therefore tentatively designated as bph19(t). A total of 58 recombination events, 20 detected at RM517 locus and 38 at RM218 locus, were subsequently used to finely map the bph19(t) locus.
Fine mapping of the bph19(t) locus
In order to fine map the bph19(t) locus, additional SSR markers developed in the region defined by RM517 and RM218 were adopted from IRMI database. Fortunately, 9 out of the 10 SSR markers selected showed polymorphism between the resistant and susceptible parents. These nine markers were used to test the 58 recombination events. The results showed that 14, 12, 12, 10, 2 and 2 recombination events occurred at the RM517 locus were detected at RM3872, RM5925, RM2326, RM3766, RM1002 and RM6308 loci, respectively. That is, these marker loci were located on the telomere side with genetic distances of 1.4, 1.2, 1.2, 1.0, 0.2 and 0.2 cM, respectively (Fig. 2b). In contrast, 28, 8 and 0 recombination event(s) occurred at the RM218 locus were detected at RM5639, RM3134 and RM1022 loci, respectively, suggesting that these three markers were located on the centromere side of the resistance gene with genetic distances of 2.8, 0.8 and 0.0 cM, respectively (Fig. 2b). Hence, the bph19(t) locus was mapped in a smaller 1.0-cM region, delimited by RM6308 and RM3134, and co-segregated with RM1022. Notably, the orders of two pairs of SSR markers, RM218 and RM7, and RM2326 and RM3872, disagreed with those in the previous maps (McCouch et al. 2001, 2002). It may be because of the inversions taken place in those small chromosome regions in the donor cv. AS20-1.
For map-based cloning of the bph19(t) gene conveniently in the future, additional four markers were created in the RM6308-RM3134 interval based on the sequence information from the reference cv. Nipponbare. Of the newly designed markers, two (SSR marker b1 and STS marker b3) were polymorphic between the parents, and another two STS markers b2 and b4 were monomorphic on the parents. The monomorphic STS markers were then converted into CAPS markers, successively (Table 1). Thus, these four polymorphic markers were also tested with the 58 recombination events. The results showed that all the four markers co-segregated with the bph19(t) locus (Fig. 2b). According to the previous results, a high-resolution genetic map was constructed in which the bph19(t) locus was finally localized to the 1.0-cM interval flanked by markers RM6308 and RM3134, and completely co-segregated with five PCR-based markers, RM1022, b1, b3, b2 and b4 (Fig. 2b).
Physical mapping of the bph19(t) locus
According to the primer sequences and genomic sequence information of the reference cv. Nipponbare, the bph19(t)-linked SSR, STS and CAPS markers were landed on the 19 Nipponbare BAC clones by the BLASTN analysis. Sequences of the 19 clones were downloaded and aligned through Pairwise BLAST analysis. The results showed that two BAC clones, OSJNBa0049C20 and OJ1081D05, which were anchored by two closely linked markers RM6308 and RM3134, respectively, are overlapped. Thus, a physical map of the bph19(t) locus was generated as shown in Fig. 3 (partial data not shown). The physical distance between RM6308 and RM3134 is estimated to be ≈60 kb in length based on the Nipponbare sequence.
Physical map of the bph19(t) locus. The numerals in parentheses indicate the recombination events occurred at the corresponding marker loci, and the numbers below the long line are relative physical distances in kilobytes estimated based on the positions of markers linked to the bph19(t) locus. The short horizontal lines represent bacterial artificial chromosome clones of cv. Nipponbare, which contain the bph19(t)-linked markers and were released by International Rice Genome Sequencing Project (IRGSP). The vertical line denotes the position of the respective markers
Discussion
In the present paper, a powerful strategy, which consists of BSA, RCA and BIA approaches, was employed to construct both genetic and physical maps of the rice BPH resistance gene, bph19(t). Firstly, BSA provides a rapid, technically simple means for identifying candidate markers in the target region (Michelmore et al. 1991). Secondly, RCA enables to confirm the candidate markers for mapping the target gene on its chromosome using the homo-recessive progenies (Pan et al. 2003; Zhu et al. 2004a). Third, BIA allows us to adopt and/or develop molecular markers in the target region, and to construct a physical map spanning the target gene locus, in silico (Gu et al. 2004; Chen et al. 2005; Liu et al. 2005). As a result, the bph19(t) locus was genetically defined to a chromosomal interval of 1.0 cM in length with five co-segregated markers, and physically mapped to a DNA fragment of 60-kb in length.
It is of interest that plant resistance genes, including major (qualitative) and minor (quantitative, QTL) genes, tend to be clustered in the particular regions of chromosomes (Beavis et al. 1991; Wang et al. 1994; Xu et al. 2002; Ramalingam et al. 2003; Yang et al. 2003). Aside from the bph19(t) gene, one major gene Bph13(t) and two QTLs were mapped on the short arm of rice chromosome 3 (Fig. 2a). The Bph13(t) gene was located near the bph19(t) gene (Renganayaki et al. 2002), but it is clearly different from the bph19(t) gene for the following reasons: (i) Bph13(t) is a dominant gene, while bph19(t) is a recessive gene based on the evaluation of BPH resistance in our two populations; (ii) According to the report of Renganayaki et al. (2002), Bph13(t) resides in the region flanked by RG100 and RG191, while bph19(t) was mapped in the region flanked by RM517 and RM218 in the present study (Fig. 2a). One QTL accounted for only 3.8% of the phenotypic variance was identified to a marker interval, RG944-RG348, which is farther than the Bph13(t) locus from the bph19(t) locus (Fig. 2a; Xu et al. 2002). Another QTL predominantly associated with a single resistance mechanism, antixenosis, was also identified to a marker interval, RG191-RZ678, which covers the bph19(t) location (Fig. 2a; Alam and Cohen 1998). Currently, four hypotheses could explain their genetic relationships: (i) they are different loci but tightly linked to each other; (ii) they are different alleles at the same locus; (iii) they are the same allele but show differential reactions to different biotypes of the BPH; and (iv) they are the same gene but expressed different reactions in the different background (Wang et al. 1994; Monna et al. 2002; Ramalingam et al. 2003). One way to test the hypothesis above is to isolate the single QTL line and then construct its high-resolution map (Paterson et al. 1990; Inukai et al. 1996). Since this QTL has not been named and finely mapped, the resistance gene identified in the present study was reasonably designated as bph19(t).
As for disease resistance genes, recessive resistance genes generally do not cluster with other resistant genes in rice (Ronald 1997, 1998; Richter and Ronald 2000). But this may not be the case for BPH resistance genes. The most striking example is that a recessive BPH resistance gene bph2, with other three dominant genes, Bph1, Bph9 and Bph10, were mapped to a 25-cM interval on the long arm of chromosome 12 (Murai et al. 2001; Murata et al. 2001; Sharma et al. 2002). One more interesting phenomenon was observed by the same research group. That is, the recessive gene bph2 behaved as a dominant gene (Murata et al. 1998; Murai et al. 2001). A possible explanation might be that recessive resistance genes behave as dominant genes under different genetic backgrounds and with different pathotypes (races) or biotypes (Murai et al. 2001). Another possible reason is that gene conversion or unequal crossing-over events occurred at the bph2 locus during meiosis, and resulted in the instability of bph2, which is similar to the rust resistance gene Rp1 (Sudupak et al. 1993; Richter and Ronald 2000). A similar case is on the short arm of chromosome 3, where the recessive gene, bph19(t), the dominant gene, Bph13(t), as well as two QTLs were located (Fig. 2a). It seems that the bph19(t) locus is unstable, due to inversions taken place in the two marker pairs, RM218-RM7 and RM2326-RM3872 on the both sides. Although allelism of the BPH resistance genes resided at the clusters, including the bph19(t) cluster, are still not well understood, fine mapping and molecular isolation of these BPH genes will shed light on this issue.
On the other hand, only one natural insect resistance gene, Mi, has been cloned, so far. It belongs to the nucleotide-binding site-leucine-rich repeat (NBS-LRR) class of resistance genes, and resembles to that of most disease resistance genes (Vos et al. 1998). In our work, putative open reading frames (ORFs) were predicted in the 60-kb interval based on the Nipponbare sequences, using gene annotation systems (http://ricegaas.dna.affrc.go.jp; http://www.softberry.com/berry.phtml). The results show that there are ten ORFs, but none possesses the NBS-LRR structure (data not shown). Therefore, it is probably that the BPH resistance gene bph19(t) encodes a novel form of insect resistance. Recent research indicates that plant responses to phloem-feeding insects, such as aphids, have similarities to those by pathogens, because of the similar effects insect stylets and fungal hyphae have on their hosts during insect feeding and pathogen infection (Fidantsef et al. 1999; Walling 2000). The related knowledge may be helpful to clarify the resistance mechanism in rice against BPH, because it also belongs to the phloem-feeding insect. We have undertaken the isolation of bph19(t), which will certainly discover the real resistance mechanism in rice against BPH.
References
Alam SN, Cohen MB (1998) Detection and analysis of QTLs for resistance to the brown planthopper, Nilaparvata lugens, in a doubled-haploid rice population. Theor Appl Genet 97:1370–1379
Athwal DS, Pathak MD, Bacalangco EH, Pura CD (1971) Genetics of resistance to brown planthoppers and green leafhoppers in Oryza sativa L. Crop Sci 11:747–750
Beavis WD, Grant D, Albersen M, Fincher R (1991) Quantitative trait loci for plant height in four maize populations and their associations with qualitative genetic loci. Theor Appl Genet 83:141–145
Chen S, Wang L, Que ZQ, Pan RQ, Pan QH (2005) Genetic and physical mapping of Pi37(t), a new gene conferring resistance to rice blast in the famous cultivar st. no. 1. Theor Appl Genet 111:1563–1570
Dyck VA, Thomas B (1979) The brown planthopper problem. In: International Rice Research Institute (eds) Brown planthopper: threat to rice production in Asia. IRRI, Los Baños, Philippines, pp 3–17
Fidantsef AL, Stout MJ, Thaler JS, Duffey SS, Bostock RM (1999) Signal interactions in pathogen and insect attack: expression of lipoxygenase, proteinase inhibitor II, and pathogenesis-related protein P4 in the tomato, Lycopersicon esculentum. Physiol Mol Plant Pathol 54:97–114
Gu K, Tian D, Yang F, Wu L, Sreekala C, Wang D, Wang GL, Yin Z (2004) High-resolution genetic mapping of Xa27(t), a new bacterial blight resistance gene in rice, Oryza sativa L. Theor Appl Genet 108:800–807
Hirabayashi H, Ogawa T (1995) RFLP mapping of Bph-1 (brown planthopper resistance gene) in rice. Breed Sci 45:369–371
Hirabayashi H, Angeles ER, Kaji R, Ogawa T, Brar DS, Khush GS (1998) Identification of brown planthopper resistance gene derived from O. officinalis using molecular markers in rice (in Japanese). Breed Sci Suppl 48:82
Hirabayashi H, Kaji R, Angeles ER, Ogawa T, Brar DS, Khush GS (1999) RFLP analysis of a new gene for resistance to brown planthopper derived from O. officinalis on rice chromosome 4 (in Japanese). Breed Sci Suppl 49:48
Holt J, Chancellor TCB, Reynolds DR, Tiongco ER (1996) Risk assessment for rice planthopper and tungro disease outbreaks. Crop Protection 15(4):359–368
Huang CF (2003) Development of position specific microsatellite markers and molecular mapping of insect resistant genes in rice (Oryza sativa L.) (in Chinese). MS Thesis, South China Agricultural University
Huang Z, He G, Shu L, Li X, Zhang Q (2001) Identification and mapping of two brown planthopper resistance genes in rice. Theor Appl Genet 102:929–934
Inukai T, Zeigler RS, Sarkarung S, Bronson M, Dung LV, Kinoshita T, Nelson RJ (1996) Development of pre-isogenic lines for rice blast-resistance by marker-aided selecting from inbred population. Theor Appl Genet 93:560–567
IRRI (1988) Standard evaluation system for rice. International Rice Research Institute, Manila
Ishii T, Brar DS, Multani DS, Khush GS (1994) Molecular tagging of genes for brown planthopper resistance and earliness introgressed from Oryza australiensis into cultivated rice, O. sativa. Genome 37:217–221
Jena KK, Jeung JU, Lee JH, Choi HC, Brar DS (2005) High-resolution mapping of a new brown planthopper (BPH) resistance gene, Bph18(t), and marker-assisted selection for BPH resistance in rice (Oryza sativa L.) Theor Appl Genet, DOI 10. 1007/s00122-005-1027-8
Kabir MA, Khush GS (1988) Genetic analysis of resistance to brown planthopper in rice, Oryza sativa L. Plant Breed 100:54–58
Kawaguchi M, Murata K, Ishii T, Takumi S, Mori N, Nakamura C (2001) Assignment of a brown planthopper (Nilaparvata lugens Stål) resistance gene bph4 to the rice chromosome 6. Breed Sci 51:13–18
Khush GS, Rezaul-Karim ANM, Angeles ER (1985) Genetics of resistance of rice cultivar ARC10550 to Bangladesh brown planthopper biotype. J Genet 64:121–125
Kosambi DD (1944) The estimation of map distances from recombination values. Ann Eugen 12:172–175
Lakshminarayana A, Khush GS (1977) New genes for resistance to the brown planthopper in rice. Crop Sci 17:96–100
Li Q, Luo SY, Wei SM (1999) Mensuration of the brown planthopper biotypes and the relationship of the migration. Entomol Knowl 36:257–260
Liu GQ, Yan HH, Fu Q, Qian Q, Zhang ZT, Zhai WX, Zhu LH (2001) Mapping of a new gene for brown planthopper resistance in cultivated rice introgressed from Oryza eichingeri. Chin Sci Bull 46:1459–1462
Liu XQ, Wang L, Chen S, Lin F, Pan QH (2005) Genetic and physical mapping of Pi36(t), a novel rice blast resistance gene located on rice chromosome 8. Mol Gen Genomics 274:394–401
McCouch SR, Temnykh S, Lukashova A, Coburn J, Declerck G, Cartinhour S, Harrington S, Thomson M, Septiningsih E, Semon M, Moncada P, Li JM (2001) Microsatellite markers in rice: abundance, diversity, and applications. In: Khush GS, Brar DS, Hardy B (eds) Rice Genetics IV. Science Publishers, Enfield, pp 117–135
McCouch SR, Teytelman L, Xu YB, Lobos KB, Clare K, Walton M, Fu BY, Maghirang R, Li ZK, Xing YZ, Zhang QF, Kono I, Yano M, Fjellstrom R, Declerck G, Schneider D, Cartinhour S, Ware D, Stein L (2002) Development and mapping of 2240 new SSR markers for rice (Oryza sativa L.). DNA Res 9:199–207
Michelmore RW, Paran I, Kesseli RV (1991) Identification of markers linked to disease-resistance genes by bulked segregant analysis: A rapid method to detect markers in specific genomic regions by using segregating populations. Proc Natl Acad Sci USA 88:9828–9832
Monna L, Lin HX, Kojima S, Sasaki T, Yano M (2002) Genetic dissection of a genomic region for quantitative trait locus, Hd3, into two loci, Hd3a and Hd3b, controlling heading date in rice. Theor Appl Genet 104:772–778
Murai H, Hashimoto Z, Sharma PN, Shimizu T, Murata K, Takumi S, Mori N, Kawasaki S, Nakamura C (2001) Construction of a high-resolution linkage map of a rice brown planthopper (Nilaparvata lugens Stål) resistance gene bph2. Theor Appl Genet 103:526–532
Murata K, Fujiwara M, Kaneda C, Takumi S, Mori N, Nakamura C (1998) RFLP mapping of a brown planthopper (Nilaparvata lugens Stål) resistance gene bph2 of indica rice introgressed into a japonica breeding line ‘Norin-PL4’. Genes Genet Syst 73:359–364
Murata K, Fujiwara M, Murai H, Takumi S, Mori N, Nakamura C (2001) Mapping of a brown planthopper (Nilaparvata lugens Stål) resistance gene Bph9 on the long arm of rice chromosome 12. Cereal Res Commun 29:245–250
Murray MG, Thompson WK (1980) Rapid isolation of high-molecular-weight plant DNA. Nucleic Acids Res 8:4321–4325
Nemoto H, Ikeda R, Kaneda C (1989) New genes for resistance to brown planthopper, Nilaparvata lugens Stål, in rice. Jpn J Breed 39:23–28
Pan QH, Hu ZD, Takatoshi T, Wang L (2003) Fine mapping of the blast resistance gene Pi15, linked to Pii, on rice chromosome 9. Acta Bot Sin 45:871–877
Paterson AH, Deverna JW, Lanini B, Tanksley SD (1990) Fine mapping of quantitative trait loci using selected overlapping recombinant chromosomes, in an interspecies cross of tomato. Genetics 124:736–742
Pataki MD (1969) Resistance to Nephotettix impicticeps and N. lugens in varieties of rice. Nature 223:502–504
Pathak MD (1972) Resistance to insect pests in rice varieties. In: International Rice Research Institute (ed) Rice breeding. IRRI, Los Baños, Philippines, pp 325–341
Peters JL, Cnudde F, Gerats T (2003) Forward genetics and map-based cloning approaches. Trends Plant Sci 8:484–491
Ramalingam J, Vera-Cruz CM, Kukreja K, Chittoor JM, Wu JL, Lee SW, Baraoidan M, George ML, Cohen MB, Hulbert SH, Leach JE, Leung H (2003) Candidate defense genes from rice, barley, and maize and their association with qualitative and quantitative resistance in rice. Mol Plant Microbe Interact 16:14–24
Renganayaki K, Fritz AK, Sadasivam S, Pammi S, Harrington SE, McCouch SR, Kumar SM, Reddy AS (2002) Mapping and progress toward map-based cloning of brown planthopper biotype-4 resistance gene introgressed from Oryza officinalis into cultivated rice, O. sativa. Crop Sci 42:2112–2117
Richter TE, Ronald PC (2000) The evolution of disease resistance genes. Plant Mol Biol 42:195–204
Ronald PC (1997) The molecular basis of disease resistance in rice. Plant Mol Biol 35:179–186
Ronald PC (1998) Resistance gene evolution. Curr Opin Plant Biol 1:294–298
Schuler GD (1998) Electronic PCR: bridging the gap between genome mapping and genome sequencing. Trends Biotechnol 16:456–459
Sharma PN, Ketipearachchi Y, Murata K, Torii A, Takumi S, Mori N, Nakamura C (2002) RFLP/AFLP mapping of a brown planthopper (Nilaparvata lugens Stål) resistance gene Bph1 in rice. Euphytica 129:109–117
Sogawa K (1982) The rice brown planthopper: feeding physiology and host plant interactions. Annu Rev Entomol 27:49–73
Sogawa K, Liu GJ, Shen JH (2003) A review on the hyper-susceptibility of Chinese hybrid rice to insect pests. Chin J Rice Sci 17:23–30
Sudupak MA, Bennetzen JL, Hulbert SH (1993) Unequal exchange and meiotic instability of the Rp1 region disease resistance genes in maize. Genetics 133:119–125
Temnykh S, Park WD, Ayres N, Cartinhour S, Hauck N, Lipovich L, Cho YG, Ishii T, McCouch SR (2000) Mapping and genome organization of microsatellite sequences in rice (Oryza sativa L.). Theor Appl Genet 100:697–712
Vos P, Simons G, Jesse T, Wijbrandi J, Heinen L, Hogers R, Frijters A, Groenendijk J, Diergaarde P, Reijans M, Fierens-Onstenk J, Both MD, Peleman J, Liharska T, Hontelez J, Zabeau M (1998) The tomato Mi-1 gene confers resistance to both root-knot nematodes and potato aphids. Nat Biotechnol 16:1365–1369
Wang GL, Mackill DJ, Bonman JM, McCouch SR, Champoux MC, Nelson RJ (1994) RFLP mapping of genes conferring complete and partial resistance to blast in a durably resistant rice cultivar. Genetics 136:1421–1434
Wang GR, Lai FX, Fu Q, Zhang ZT, Guo LF (1999) Virulent shift in populations of Nilaparvata lugens (Homptera: Delphacidae) (in Chinese). Chin J Rice Sci 13:229–232
Walling LL (2000) The myriad plant responses to herbivores. J Plant Growth Regul 19:195–216
Xu XF, Mei HW, Luo LJ, Cheng XN, Li ZK (2002) RFLP-facilitated investigation of the quantitative resistance of rice to brown planthopper (Nilaparvata lugens). Theor Appl Genet 104:248–253
Yang HY, Ren X, Weng QM, Zhu LL, He GC (2002) Molecular mapping and genetic analysis of a rice brown planthopper (Nilaparvata lugens Stål) resistance gene. Hereditas 136:39–43
Yang Z, Sun X, Wang S, Zhang Q (2003) Genetic and physical mapping of a new gene for bacterial blight resistance in rice. Theor Appl Genet 106:1467–1472
Yang HY, You AQ, Yang ZF, Zhang FT, He RF, Zhu LL, He GC (2004) High-resolution genetic mapping at the Bph15 locus for brown planthopper resistance in rice (Oryza sativa L.). Theor Appl Genet 110:182–191
Zhu ML, Wang L, Pan QH (2004a) Identification and characterization of a new blast resistance gene located on rice chromosome 1 through linkage and differential analyses. Phytopathology 94:515–519
Zhu LL, Zhu CL, Weng QM, Huang Z, He GC (2004b) Research progress on brown planthopper resistance genes in rice (in Chinese). Hubei Agric Sci 1:19–24
Acknowledgements
We are grateful to F.-K. Huang at Guangxi Academy of Agricultural Sciences, China, for kindly providing the insect source and advice on BPH bioassay, and R.-Y. He for critical reading of the manuscript. This research was supported by grants from the National Basic Research Program of China (2006CB100200-G), the National Special Program for Functional Genomics and Biologic Chips (2002AA2E1002), the Guangdong Provincial Natural Science Foundation (000575; 039254), and the Special Project for the Distinguished University Professor from the Department of Education of Guangdong Province, China.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by R. Hagemann
Rights and permissions
About this article
Cite this article
Chen, J.W., Wang, L., Pang, X.F. et al. Genetic analysis and fine mapping of a rice brown planthopper (Nilaparvata lugens Stål) resistance gene bph19(t). Mol Genet Genomics 275, 321–329 (2006). https://doi.org/10.1007/s00438-005-0088-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-005-0088-2