Abstract
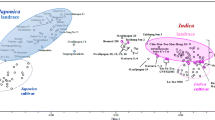
Preserving many kinds of rice resources and rich variations, Guizhou Province is one of the districts with the highest genetic diversity of cultivated rice (Oryza sativa L.) in China. In the current research, genetic diversity and structure of 537 accessions of cultivated rice from Guizhou were studied using 36 microsatellite markers and 39 phenotypic characters. The results showed that the model-based genetic structure was the same as genetic-distance-based one using SSRs but somewhat different from the documented classification (mainly based on phenotype) of two subspecies. The accessions being classified into indica by phenotype but japonica by genetic structure were much more than that being classified into japonica by phenotype but indica by genetic structure. Like Ding Ying’s taxonomic system of cultivated rice, the subspecific differentiation was the most distinct differentiation within cultivated rice. But the differentiation within indica or japonica population was different: japonica presented clearer differentiation between soil-watery ecotypes than indica, and indica presented clearer differentiation between seasonal ecotypes than japonica. Cultivated rices in Guizhou revealed high genetic diversity at both DNA and phenotypic levels. Possessing the highest genetic diversity and all the necessary conditions as a center of genetic diversity, region Southwestern of Guizhou was suggested as the center of genetic diversity of O. sativa L. from Guizhou.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Zhang Z X. Rice Germplasm resources of Guizhou. In: Ying C S. ed. Rice Germplasm Resources of China (in Chinese). Beijing: China Agriculture Press, 1993. 201–222
Chen H C, Jin T Y, You J M, et al. Analysis and evaluation on quality characters of rice germplasm resources in Guizhou. Guizhou Agric Sci (in Chinese), 1998, 26(3): 527
Chen H C, Zhang Z X, Ruan R C, et al. Evaluation and identification of tolerance to cold and resistance to drought on Guizhou HE (Oryza sativa L.) germplasm resources. Guizhou Agric Sci (in Chinese), 1999, 27(6): 38–40
Wei X H, Zhang L H, Luo L J, et al. Multilocational evaluation of the elite rice germplasm collected in Guizhou. J Southwest Agric Univ (in Chinese), 1997, (19): 334–337
Jin T Y, Zhang Z X, Chen H C, et al. Screening and observation on poly-embryonic seedings in rice germplyasm resources of Guizhou. Guizhou Agric Sci (in Chinese), 1999, 27(6): 44–46
Ruan R C, Chen H C, Yang Y S, et al. Screening of rice (Oryza sativa L.) germplasm with restoring genes to CMS from locavarieties collected in Guizhou. Guizhou Agric Sci (in Chinese), 1999, 27(6): 34–37
Ruan R C, Chen H C, You J M, et al. Assessment, conversation and utilization on elite rice genetic resources in Guizhou. Acta Bot Yunnan (in Chinese), 2001, (Suppl. XIII): 11–17
Yu X Q, Jiang X H, Wu H B, et al. SSR genetic diversity of cold-tolerant native rice varieties in Guizhou. Southwest China J Agric Sci (in Chinese), 2005, 18(1): 1–4
Ruan R C, Chen H C, Zhang Z X, et al. Progress and prospects of research and utilization on genetic diversity of rice germplasm resources in Guizhou. Acta Bot Yunnan (in Chinese), 2000, (Suppl XII): 134–138
Wu K S, Tanksley S D. Abundance, polymorphism and genetic mapping of microsatellites in rice. Mol Gen Genet, 1993, 241: 225–235
Yang G P, Saghai Maroof M A, Xu C G, et al. Comparative analysis of microsatellite DNA polymorphism in landraces and cultivars rice. Theor Appl Genet, 1994, 245: 187–194
Chen X, Temnykh S, Xu Y, et al. Development of a microsatellite framework map providing genome-wide coverage in rice (Oryza sativa L.). Theor Appl Genet, 1997, 95: 553–567
Zhao X P, Kochert G. Characterization and genetic mapping of a short, highly repeated, interspersed DNA sequence from rice (Oryza sativa L). Theor Appl Genet, 1992, 231: 353–359
Svetlana T, Genevieve D C, Angelika L, et al. Computational and experimental analysis of microsatellites in rice (Oryza sativa L): Frequency, length variation, transposon associations, and genetic marker potential. Genome Res, 2001, 11: 1441–1452
Davierwala A P, Chowdari K V, Kumar S, et al. Use of three different marker systems to estimate genetic diversity of Indian elite rice varieties. Genetica, 2000, 108: 269–284
Zhou H F, Xie Z W, Ge S. Microsatellite analysis of genetic diversity and population genetic structure of a wild rice (Oryza rufipogon Griff) in China. Theor Appl Genet, 2003, 107: 332–339
Olufowote J O, Xu Y, Chen X, et al. Comparative evaluation of within-cultivar variation of rice (Oryza sativa L) using microsatellite and RFLP markers. Genome, 1997, 40: 370–378
Garland S H, Lewin L, Abedinia M, et al. The use of microsatellite polymorphisms for the identification of Australian breeding lines of rice (Oryza sativa L). Euphytica, 1999, 108: 53–63
Liu X C, Wu J L. SSR heterogenic patterns of parents for marking and predicting heterosis in rice breeding. Mol Breed, 1998, 4: 263–268
Li Z C. Studies on sampling strategy for core ccollection of Chinese landrace rice and genetic diversity of phenotypes and isozymes (in Chinese). Dissertation for the Doctoral Degree. Beijing: China Agricultural University, 2001. 22–39
SaghiMaroof M A, Soliman K M, Jorgensen A R, et al. Ribisinal DNA space length polymorphism in barley: Mendelian inheritance, chromosomal location and population dynamics. Proc Natl Acad Sci USA, 1984, 81: 8014–8018
Panaud O, Chen X L, McCouch S R. Development of microsatellite markers and characterization of simple sequence length polymorphism (SSLP) in rice (Oryza sativa L). Mol Gen Genet, 1996, 252: 597–607
Sheldon A L. Equitability indices: Dependence on the species count. Ecology, 1969, 50: 466–467
Nei M. Analysis of diversity in subdivided populations. Proc Natl Acad Sci USA, 1973, 70: 3321–3323
Pritchard J K, Stephens M, Donnelly P. Inference of population structure using multilocus genotype data. Genetics, 2000, 155: 945–959
Falush D, Stephens M, Pritchard J K. Inference of population structure using multilocus genotype data: Linked loci and correlated allele frequencies. Genetics, 2003, 164: 1567–1587
Liu K, Muse S. PowerMarker: New Genetic Data Analysis Software, Version 2.7 (http://www.powermarker.net), 2004
Nei, M, Tajima F A. Tateno. Accuracy of estimated phylc-genetic trees from molecular data. J Mol Evol, 1983, 19: 153–170
Takezaki N, Nei M. Genetic distances and reconstruction of phylogenetic trees from microsatellite DNA. Genetics, 1996, 144: 389–399
Vigouroux Y, Matsuoka Y, Doebley J. Directional evolutionary for microsatellite size in maize. Mol Biol Evol, 2003, 20: 1480–1483
Weir B S, Cockerham C C. Estimating F-statistics for the analysis of population structure. Evolution, 1984, 38: 1358–1370
Yang C D. Regionalization of rice cropping in Guizhou. In: Min S K, Wu X Z. Regionalization of Rice Cropping in China (in Chinese). Hangzhou: Zhejiang Science Technology Press, 1989. 99–103
Glaszman J C. Isozymes and classification of Asian rice varieties. Theor Appl Genet, 1987, 74: 21–30
Oka H I. Origin of Cultivated Rice. Tokyo: Jnp Sci Soc Press, 1988
Tang S X, Jiang Y Z, Wei X H, et al. Genetic diversity of isozymes of cultivated rice in China. Acta Agron Sin (in Chinese), 2002, 28(3): 203–207
Ting Y. The origin and evolution of cultivated rice in China. Acta Agron Sin (in Chinese), 1957, 8(3): 243–260
Cheng K S. A statistical evaluation of classification of rice cultivars into hsien and keng subspecies. Rice Genet Newsl, 1985, 2: 46–48
Garris A J, Tai T H, Coburn J, et al. Genetic structure and diversity in Oryza sativa L. Genetics. 2005, 169: 1631–1638
Semon M, Nielsen R, Jones P M, et al. The population structure of African Cultivated rice Oryza glaberrima (Steud.): Evidence for elevated levels of linkage disequilibrium caused by admixture with O. sativa and ecological adaptation. Genetics. 2005, 169: 1639–1647
Lu H, Redus M A, Coburn J R, et al. Population structure and breeding patterns of 145 U.S. rice cultivars based on SSR marker analysis. Crop Sci, 2005, 45: 66–76
Author information
Authors and Affiliations
Corresponding author
Additional information
Contributed equally to this work
Supported by the National Basic Research Program of China (“973” Program) (Grant No. 2004CB117201) and the Key Technologies R&D Programme of China (Grant No. 2004BA525B02-05)
About this article
Cite this article
Zhang, D., Zhang, H., Wei, X. et al. Genetic structure and diversity of Oryza sativa L. in Guizhou, China. CHINESE SCI BULL 52, 343–351 (2007). https://doi.org/10.1007/s11434-007-0063-x
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/s11434-007-0063-x