Abstract
The study of the soil microbial community represents an important step in better understanding the environmental context. Therefore, biological characterisation and physicochemical integration are keys when defining contaminated sites. Fungi play a fundamental role in the soil, by providing and supporting ecological services for ecosystems and human wellbeing. In this research, 52 soil fungal taxa were isolated from in situ pilot reactors installed to a contaminated site in Czech Republic with a high concentration of hexachlorocyclohexane (HCH). Among the identified isolates, 12 strains were selected to evaluate their tolerance to different isomers of HCH by using specific indices (Rt:Rc; T.I.) and to test their potential in xenobiotic biotransformation. Most of the selected taxa was not significantly affected by exposure to HCH, underlining the elevated tolerance of all the tested fungal taxa, and different metabolic intermediates of HCH dechlorination were observed. The oxidative stress responses to HCH for two selected species, Penicillium simplicissimum and Trichoderma harzianum, were investigated in order to explore their toxic responses and to evaluate their potential functioning in bioremediation of contaminated environments. This research suggests that the isolated fungal species may provide opportunities for new eco-friendly, integrated and cost-effective solutions for environmental management and remediation, considering their efficient adaptation to stressful conditions.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Over the last 30 years, organochlorine pesticides have been widely used for agriculture and public health purposes (Giri et al. 2014). Ecotoxicological data indicate that the reiterated use of pesticides has contributed to undesirable environmental effects on animals, plants, microorganisms and risks to human health (Sherif and Elhussein 2011; Hussain et al. 2015; Maqbool et al. 2016). Consequently, the problems associated with contaminated sites assume increasing prominence in many countries (Vidali 2001; Gurung et al. 2018).
Specifically, during the last few decades, scientific studies of persistent organic pollutants (POPs), like hexachlorocyclohexane (HCH), have revealed that these chemicals cause problems on a local as well as a global scale (Scheringer 2004; Nadal et al. 2015; Ashraf 2017). HCH exists in isomers, and one of these, γ-HCH or lindane, was widely used for agricultural purposes (Mrema et al. 2013). It has been proven that HCH may cause serious damages to ecosystems and human health in the short and long term (Saez et al. 2017). Worldwide HCH waste production has been estimated to range between 4.8 and 7.2 million tonnes (Vijgen et al. 2011). In 2009, the list of chemicals established by the Stockholm Convention in 2001, was extended to include some of the isomers which were used during this study, i.e. α-HCH, β-HCH and γ-HCH (Vijgen et al. 2011). Due to its low solubility in water (0.2–8 mg/L), HCH has a strong affinity with the suspended particulate matter in water (Hu et al. 2010) which can increase its stability and resistance to biodegradation. Consequently, it tends to bioaccumulate along the food chain (Borgå et al. 2001; Caicedo et al. 2011; Saez et al. 2017).
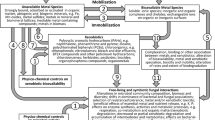
HCH bioremediation is especially necessary at industrial post-production or waste dumping sites, because decontamination through natural attenuation by microorganisms is slow and problematic (Phillips et al. 2005). In this regard, fungi can play an important role in the decontamination of a compromised area (Gadd 2001; Singh 2006; Harms et al. 2011; Anastasi et al. 2013; Deshmukh et al. 2016). To resolve these problems, approaches that link remediation to the possibility of reusing contaminated sites may provide feasible solutions (Tu 1994; Nawab et al. 2003; Komínková et al. 2018). Bioremediation mediated by microorganisms (e.g. fungi and bacteria) is an environmentally friendly technology which is able to reduce hazardous substances, and several biotechnological applications can be leveraged (Salam et al. 2013; Ceci et al. 2015a, b; Spina et al. 2018). Fungi can tolerate extreme and very restrictive environmental conditions, such as high concentrated mixtures of toxic substances, and are characterised by unique attributes such as robust morphology, diverse metabolic capacity and powerful and low specific enzymes that make them an effective tool for biodegradation of toxic organic chemicals and the most vigorous agents for the decomposition of waste matter (Harms et al. 2011; Maqbool et al. 2016; Morillo and Villaverde 2017). Compared to other microorganisms, they benefit from extended mycelial networks, and they are also able to use xenobiotic compounds as a source of carbon and energy (Gadd 2001; Singh 2006; Roze et al. 2011; Kulshreshtha et al. 2014; Czaplicki et al. 2016; Maqbool et al. 2016). In addition, fungal bioremediation or mycoremediation offers an environmentally oriented and sustainable solution in comparison with less efficient and less cost-effective traditional physical and chemical methods (Gadd 2001; Singh 2006; Ceci et al. 2015a, b; Maqbool et al. 2016). Several reports on filamentous fungi as HCH degraders are reported in literature (Ceci et al. 2015a; Guillén-Jiménez et al. 2012; Phillips et al. 2005; Taşeli 2006).
In order to maximise the application of saprotrophic soil microfungi in the bioremediation of contaminated environments, it is crucial to focus on the site’s indigenous fungi when assessing them in a remediation project (D’Annibale et al. 2006; Czaplicki et al. 2016; Godoy et al. 2016; Awasthi et al. 2017). In this research, the study of fungal assemblages in field reactors provides useful information on the fungal biodiversity and functionality; the bioremediation processes involved, such as biosorption, biostimulation and bioaugmentation; and the efficiency of biological treatment of the wastewaters. The investigated reactors represent different treatment systems for reduction of the environmental toxicity of HCH into surface waters and nearby ecosystems, and the investigation on functionality could be useful for shedding further light on the time dynamics and the relationships between fungal ecophysiology responses and the physicochemical features of the reactors.
The aims of this research were (1) to characterise the fungal assemblages in the reactors in order to study the biodiversity, the ecology and the bioremediation potential of isolated fungi; (2) to develop a better understanding of the fungal role in bioremediation processes by analysing the expected metabolites in the HCH biodegradation pathway; and (3) to investigate the oxidative stress responses of two selected species of microfungi in order to evaluate the stress-induced responses through the production of ROS and antioxidant enzymes such as superoxide dismutase (SOD), catalase (CAT) and glutathione S-transferase (GST) due to the presence of HCH.
Materials and methods
Contaminated site and sampling description
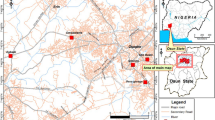
The analysed matrices samples were taken from the contaminated site of Hájek in western Bohemia (the Czech Republic). In the 1960s, a large amount of waste from the production of γ-HCH was placed into a dump at this site. In recent studies, seepage water was collected by drainage pipes which flow into an open channel discharging into a nearby lake. In the channel, the sum of HCH concentrations exceeded the appropriate limit (0.02 μg/L) (Wacławek et al. 2016). Several variants of treatment systems for limiting the leakage of HCH into surface waters were tested at the site. Specifically, four of the analysed matrices, known as N-A, N-B, N-C and N-D (Fig. 1), came from HCH degradation reactors which provide an artificial aerobic wetland with soil, gravel and emerged plants (N-A); a wetland with green mulch, ground grain, bark and wood chips (N-B); a barrier of zero valent iron for chemical reduction (N-C); and a sorption with peat, bark and wood chips (N-D). The fifth analysed sample comes from a natural wood wetland soil (WW) (Fig. 1). Lastly, it was carried out an analysis from the fifth sample (WW) amended with a mixture of different HCH isomers at the final concentration of 4 mg/L (WW-HCH) in order to select tolerant species with a potential for bioremediation and verify the occurrence of indigenous fungal species in the investigated reactors.
Isolation and identification of potential HCH degrading fungal strains
The strains were obtained through fungal isolation using dilution plate method (soil/water ratio of 1:1000) according to Persiani et al. (2008). Aliquots from each matrix suspension (0.1 mL) were plated in five Petri dishes with a soil extract agar (culture medium) prepared using soil from the sampling area (Maggi et al. 2005). After the incubation period (7 days at 25 °C) on the soil extract agar, the number of colonies was counted and expressed as colony-forming units (CFUs). All the species were identified through conventional taxonomic keys based on macro- and microscopic characteristics after growing on a culture medium suitable for this purpose (Malloch and Cain 1971; von Arx and Müller 1975; Ellis 1976; Pitt 1979; Ramirez and Martinez 1982; Gams and Dingley 2006; Domsch et al. 2007; Samuels and Hebbar 2015). All the isolates were transferred as pure cultures and stored at 4 °C in the culture collection of the Fungal Biodiversity Laboratory (FBL, Sapienza, University of Rome).
Genetic identification of selected fungal strains
The identification of the fungal strains was confirmed by sequencing the internal transcribed spacer (ITS) 1, the 5.8S gene and the ITS2 regions of the ribosomal DNA (White et al. 1990), in conjunction with other protein-coding genes. The b-tubulin gene sequence was used as secondary barcode marker in the case of Penicillium strain (Glass and Donaldson 1995), while the translation elongation factor 1-alpha (tef1) with intron 4, in combination with intron 5 regions, has been used for the molecular identification of the Trichoderma strain (Jaklitsch et al. 2005).
Fungal DNA was extracted from fresh mycelium using the DNeasy PowerSoil Kit, which has proved effective in DNA cleaning even in the presence of pigments, oils and other fungal metabolites. Both the purity and quantity of DNA were checked by agarose gel electrophoresis, determined with the NanoDrop 8000 spectrophotometer (Thermo Scientific, Waltham, MA, USA), and the DNA concentration was adjusted to 10 ng μl−1.
All PCR reactions were performed in a Veriti 96-Well Thermal Cycler (Applied Biosystems) using the GoTaq® DNA Polymerase (Promega). The reactions were carried out in a 25-μL volume containing 1× Reaction Buffer, 1.5 mM MgCl2, 0.2 mM each dNTP, 1.0 μM upstream primer, 1.0 μM downstream primer, 1.25u GoTaq® DNA polymerase, 1 μg bovine serum albumin and < 0.3 μg/25 μl of the DNA template.
The internal transcribed spacer (ITS) region of the rRNA (~ 450–800 bp) was amplified using the primer pair ITS1-ITS4 (White et al. 1990), the beta-tubulin (tub2/BenA) (~ 500 bp) using the primer pair Bt2a-Bt2b (Glass and Donaldson 1995) and the translation elongation factor 1-alpha (tef1) (~ 1300 bp) using the primer pair EF1-728F- TEF1LLErev (Jaklitsch et al. 2005).
The thermocycling programmes were as follows:
-
1)
ITS: 5-min denaturation at 94 °C, followed by 30 cycles of 1-min denaturation at 94 °C, 1-min annealing at 60 °C and 1-min extension at 72 °C. Ten minutes at 72 °C were used as a final extension step.
-
2)
tub2: 5-min denaturation at 95 °C, followed by 35 cycles of 30-s denaturation at 95 °C, 30-s annealing at 58 °C and 1-min extension at 72 °C. Seven minutes at 72 °C were used as a final extension step.
-
3)
tef1 (touchdown cycle): 2-min denaturation at 94 °C, 1 amplification cycle with annealing at 66 °C, subsequently incrementally reduced by 1 °C per cycle over the next 9 cycles. 36 amplification cycles were then performed, each consisting of 30-s denaturation (94 °C), 30-s annealing (56 °C), 1-min extension (72 °C), concluding with 0-min incubation (72 °C) as the final extension step, followed by a quick cooling to room temperature.
PCR products were checked by agarose gel electrophoresis, purified with QIAquick PCR Purification Kit (Qiagen) and sent for Sanger sequencing to NHM, Molecular Laboratory, UK.
The forward and reverse electropherograms obtained for each fungal isolate were verified visually and aligned using CLUSTALW (version 2.0) to obtain consensus sequences that were then compared using the BLAST search programme (Altschul et al. 1997) with the NCBI database (Karsch-Mizrachi et al. 2012). Comparative ITS sequence analyses were performed using online public databases: the National Centre for Biotechnology Information (NCBI), the UNITE database (Kõljalg et al. 2005; Abarenkov et al. 2010) and the TrichOKey v. 1.0 (barcode sequence identification programme for Trichoderma species with the web interface located on www.isth.info) (Druzhinina et al. 2005). The Trichoderma tef1 sequence was used in TrichoBLAST v. 1.0 and the tef1 4th intron, tef1 5th intron and tef1 6th exon searched with BLAST within the Multiloci Database of Phylogenetic Markers using this component of the TrichoKeys database (http://isth.info/).
Chemicals and analytical reagents
The four HCH isomers (α-, β-, γ- and δ-HCH, 98.2% purity) were purchased from Sigma-Aldrich (Seelze, Germany). Fungal culture media were amended 1 mg/L for each isomer using a stock solution of α-, β-, γ- and δ-HCH with a final concentration of 4 mg/L (α/β/γ/δ = 1:1:1:1) prepared in toluene (> 99.9% purity, ROMIL Ltd, Cambridge, UK). Ethyl acetate was purchased from ROMIL Ltd (Cambridge, UK) with a chemical purity > 99.9%. Antioxidant enzyme assay kits were purchased from Sigma-Aldrich (Seelze, Germany).
Screening of fungal isolates for HCH toxicity tolerance
In order to test the tolerance, the pure cultures were maintained on a Malt Extract Agar (MEA) for 7 days at 25 °C. The medium was prepared according to the following composition (g/L in distilled water): malt extract, 20; peptone, 1; glucose, 20; agar, 20. Pure fungal isolates were further screened for their growth tolerance on treatment plates supplemented with the mixture of four HCH isomers, α-HCH, β-HCH, γ-HCH and δ-HCH (1:1:1:1 ratio) at the final concentration of 4 mg/L. Petri dishes were inoculated at the centre and cultures were incubated at 25 °C in the dark. Mycelial diametric growth was daily recorded for 7 days. All the experiments were carried out in triplicate. Prior to inoculation, an 84-mm-diameter sterile cellophane membrane was placed aseptically on the surface of the agar in each Petri dish to separate the fungi from the medium, allow the passage of nutrients and metabolites between the medium and the colony and provide a convenient means of recovering the mycelium (Ceci et al. 2015b). Strain inoculation was carried out using a 5-mm-diameter mycelium core from the active growth colony margin using a cork borer. After 7 days, the fungal colonies were removed from the medium by peeling the biomass from the membranes using a sterile razor blade. The mycelium was oven-dried at 100 °C until reaching a constant weight for at least 2 days. Fungal tolerance to the HCH isomers was evaluated using two indices: (1) the tolerance index (Rt:Rc) defined as the ratio of the colony extension rates in the presence of HCHs (Rt) to the control extension rate (Rc); (2) the tolerance index (TI) based on the dry weights of the fungal biomass (DW) as follows (Ceci et al. 2015b, 2018):
After the membrane and mycelium were removed, the surface pH of the culture media was measured at 20-mm intervals across the diameter of the Petri dish using a conical tip FC 202D pH electrode (Hanna Instruments, Woonsocket, RI, USA) and a portable pH metre, HI 99161 (Hanna Instruments, Woonsocket, RI, USA). The pH measurements were used to show the pH profile for treatments, while the average surface pH values were used to calculate the differences (ΔpH) between the control and the test.
Chemical analysis
The agar and the membranes were collected and analysed to detect the formation of metabolites according to Ceci et al. (2015a, 2018). Specimens were posed in glass tubes with 15 mL of ethyl acetate and sonicated for 30 min. After centrifugation at 2000 rpm, the ethyl acetate solutions were recovered in vials and 10 mL was analysed (Ceci et al. 2015a, 2018). To improve the metabolite detection, a concentration procedure was performed in which 0.5 mL of ethyl acetate solution from each sample was collected, evaporated and recovered in 100 μL of toluene solution. The formation of metabolites was analysed by gas chromatography-mass spectrometry (GC-MS). Operating conditions are reported in Ceci et al. (2015a). A Hewlett-Packard 6890 gas chromatograph with a 5973A mass selective detector (Agilent Technologies, Palo Alto, CA, USA) was used. The qualitative analyses concerned the formation of intermediate metabolites from the biotransformation process, such as dechlorination of HCH (e.g. pentachlorocyclohexene, tetrachlorocyclohexene) or intermediates of HCH reductive dechlorination and hydroxylation, as reported in other bioremediation studies with bacteria and fungi (Phillips et al. 2005; Guillén-Jiménez et al. 2012; Singh 2017).
Oxidative stress analysis: ROS determination and antioxidant enzyme activities
The isolates P. simplicissimum and T. harzianum out of the 12 evaluated for the fungal tolerance to HCH were selected for monitoring the changes in reactive oxygen species (ROS) production and activities of the following antioxidant enzymes in the presence of HCH: superoxide dismutase (SOD), catalase (CAT), glutathione S-transferase (GST). The two taxa were selected by considering sporulation capacity, diametric growth and biomass production. After being grown on MEA at 25 °C for 7 days, a spore suspension was prepared at a concentration of 2 × 108 spores/mL by using a counting Thoma chamber. The spores were removed from the culture dishes with the aid of a sterile loop by adding sterile distilled water amended with Tween 20. The resulted suspension was filtrated by sterile gauzes several times to remove residues of hyphae or conidiophores and 5 ml of the final spore suspension was inoculated in 75 ml Czapek-Dox medium (sucrose 30 g/L; NaNO3 3 g/L; KH2PO4 1 g/L; MgSO4·7H2O 0.5 g/L; KCl 0.5 g/L; FeSO4 H2O 0.01 g/L, distilled water 1 L) adjusted to pH 5.6 with 1 M HCl, before autoclaving. After 24 h, the mycelium was harvested by centrifugation and resuspended in a fresh Czapek-Dox medium with or without the addition of the oxidative stress agent (HCH isomeric mixture at a final concentration of 4 mg/L). After further growth, the mycelium was harvested by filtration, washed in sterile distilled H2O and then in cold 50 mM potassium buffer (pH 7.8) and resuspended in the same buffer according to Angelova et al. (2005). The cell suspension was disrupted using the Tissue Lyser LT (Qiagen) and after centrifugation (12.000g for 20 min at 4 °C), the supernatant was collected to determinate ROS production and enzymatic activities.
ROS production was determined using oxidative stress cell-permeant 2′,7′dichlorodihydrofluorescein diacetate dye (H2DCFDA, Sigma-Aldrich) which is oxidised to a fluorescence dye 2,7-dichlorofluorescein (DCF) with excitation/emission wavelengths of 350 nm/600 nm (Galdiero et al. 2016; Russo et al. 2017). CAT, SOD and GST activities were determined using commercial assays kits (Sigma-Aldrich) following the manufacturer’s protocols. CAT activity was determined by measuring the decrease of H2O2 at 240 nm with a spectrophotometer (Hach lange DR5000) at different times (30, 60, 90, 120 s). SOD and GST activities were measured by a microplate reader (BioTek™ Synergy™ H4 Hybrid). SOD activity was expressed by determining the decrease in the colour development of the samples at 440 nm. GST activity was calculated by measuring the variations in absorbance recorded at 340 nm. Protein content was measured using the protocol reported by Bradford (1976) with bovine serum albumin as a standard. The protein content was used to normalise the results of the enzymatic activities. All the tests were performed in triplicate.
Data treatment
All data obtained were presented as the mean ± standard error (SE) and tested for statistical significance using one-way analysis of variance (ANOVA) followed by the Tukey test by XLSTAT (version 2018.1.49386) software (Addinsoft 2007-Pro, Paris, France). The critical value for statistical significance was p < 0.05.
Results
Fungal characterisation
On the basis of the investigations carried out on the five samples studied, a total of 52 fungal taxa was detected. The most represented taxa were Penicillium with 13 species, Trichoderma with five, Sphaeropsidales with eight and Aspergillus, Gaeumannomyces and Talaromyces with two, while the remaining taxa were present as single species (data not shown). N-D was clearly the first in terms of species richness with 22 species, while WW (19 species) was the second followed by WW-HCH (14 species), N-A (six species), N-B (four species) and N-C (zero species). The relationship between species richness and genus richness recorded from all the matrices is shown in Fig. 2. Although the highest number of species was present in N-D, the total number of genera was relatively low; in fact, in this matrix, there was evidence of the highest amount of species belonging to the genus Penicillium which counts for 10 species among the 13 isolated from the overall matrices. By comparing the number of isolates of the natural wetland WW with the amended ones (WW-HCH), a slight decrease (from 19 to 14) was observed, but it was still possible to highlight several taxa as, e.g. Harzia acremonioides, Metarhizium sp., Sphaeropsidales sp., Talaromyces sp. and Trichoderma polysporum common to both treatments (WW; WW-HCH). Instead, by comparing the number of isolates of the natural wetland WW with the other matrices (N-A, N-B, N-C, N-D), it was possible to underline two common species only with N-D: Aspergillus fumigatus and Penicillium simplicissimum.
A total of 332 colony-forming units (CFUs) was isolated from all the matrices (Fig. 3). The matrices with the highest number of CFUs were N-D (56%), followed by WW (27%), WW-HCH (9%) and N-B (6%), while the matrix with the lowest number of colonies was N-A (2%). Lastly, no species were observed in N-C (Fig. 3). The 332 screened CFUs were represented by different genera: Aspergillus (16%), Cadophora (13%), Metarhizium (5%), Penicillium (18%), Trichoderma (8%), Talaromyces (7%), Sporothrix (5%), Gaeumannomyces (4%) and Zopfiella (4%). Sphaeropsidales (8%) and Agonomycetales (formerly Mycelia sterilia) (8%), followed by other genera with a percentage not exceeding 1% (e.g. Acremonium, Cladosporium, Harzia, Humicola, Mucor, Westerdykella, Phialocephala) also occurred. Of the 52 isolated taxa, 12 strains were selected to evaluate their tolerance and to test their potential in biotransformation of HCH (Fig. 4). Of these 12 strains, two species, P. simplicissimum and T. harzianum, were selected to investigate the oxidative stress responses in the presence of a high HCH concentration.
Distribution of the 332 colony-forming units (CFUs) isolated from five matrices (WW, WW-HCH, N-A, N-B, N-C, N-D).The largest number of colonies was isolated from N-D (184 CFUs, 56%), followed by WW with 91 CFUs, 27%; WW-HCH with 30 CFUs, 9%; N-A with 7 CFUs, 2%; 20 CFUs, 6% for N-B; and 0 CFUs, 0% for N-C
Selected taxa for fungal tolerance evaluation growth on MEA for 7 days at 25 °C. The pure cultures are maintained in fungal collection of Fungal Biodiversity Laboratory (FBL). a Cadophora fastigiata Lagerb. & Melin FBL 557; b Harzia acremonioides (Harz) Costantin FBL 571; c Metarhizium sp. FBL 568; d Mucor hiemalis f. hiemalis Wehmer FBL 572; e Penicillium simplicissimum K.M. Zaleski FBL 573; f Phialocephala dimorphospora W.B. Kendr FBL 636; g Sphaeropsidales sp. FBL 570; h Talaromyces sp. FBL 574; k Trichoderma sp. FBL 575; i Trichoderma harzianum J. Webster & Rifai FBL 576; j Westerdykella sp. FBL 569; l Zopfiella latipes (N. Lundq.) Malloch & Cain FBL 567
Evaluation of fungal growth rate and tolerance
The selected taxa for fungal tolerance evaluation are reported in Fig. 4. Exposure to HCHs did not significantly reduce the diametric growth of most of the selected taxa, underlining the elevated tolerance of all the tested fungal taxa. Only H. acremonoides and P. simplicissimum showed lower values of fungal extension than the control, and ANOVA followed by the Tukey test showed statistically significant differences (p < 0.05). A marked difference in the levels of HCH tolerance was revealed in three species, namely M. hiemalis f. hiemalis, T. harzianum and Trichoderma sp. These taxa showed very high tolerance and exhibited strong growth and, as “hyper-tolerant”, reached the maximum radial growth of 86 mm after 7 days. These patterns reflect the tolerance development of the fungi, expressed as the tolerance index (Rt:Rc) (Table 1). None of the colony extensions were significantly reduced (Rt:Rc > 0.75) by exposure to HCHs. The lowest value occurred in P. simplicissimum (Rt:Rc = 0.77) reflecting a relatively moderate diametric growth, while the highest value occurred in Sphaeropsidales sp. (Rt:Rc = 1.02) despite a low growth.
The degree of tolerance was also measured using the tolerance index (TI). The tolerance values of these selected fungi after growth for 7 days is presented in Fig. 5. Dry weights of the recovered fungal biomass (dw) were not strongly reduced (TI > 50%) by exposure to HCHs. The lowest value occurred in Z. latipes (TI = 58%), while the highest TI value occurred in Westerdykella sp. (TI = 123%). Nevertheless, multiple comparisons using the Tukey test revealed no significant differences between the average values of dry weight.
Medium pH profile under experimental conditions
The average surface pH values of the media are reported in Table 2. The addition of HCH to MEA did not significantly change the medium pH by comparing the treatments (5.21 ± 0.02) with the control conditions (5.30 ± 0.03). Medium pH values increased (alkalinisation) after fungal growth in the case of all the taxa, except for C. fastigiata, Sphaeropsidales sp., T. harzianum and Westerdykella sp. Despite these results, multiple comparisons using the Tukey test revealed no significant differences. ΔpH values are reported in Fig. 6.
GC-MS analysis: identification of metabolites and degradation products of HCH
Different metabolic intermediates of HCH dechlorination were observed in different fungal species, and different isomers of pentachlorocyclohexene (PCCH), tetrachlorocyclohexene (TCCH), trichlorobenzene (TCB), dichlorobenzene (DCB) and chlorobenzene (CB) were detected (Table 3). The concentration procedure significantly improved the number of intermediates found. Without this procedure, only PCCH was detected in the solid Czapek-Dox medium experiments for C. fastigiata, H. acremonioides, Metarhizium sp., M. hiemalis f. hiemalis, P. simplicissimum, P. dimorphospora, Sphaeropsidales sp., Talaromyces sp., Trichoderma sp., T. harzianum, Westerdikella sp. and Z. latipes, while DCB and CB were found in all the tested taxa. After the concentration, all the intermediates were detected in P. dimorphospora and Sphaeropsidales sp., while in C. fastigiata, H. acremonioides, Metarhizium sp., P. dimorphospora, Sphaeropsidales sp., Talaromyces sp. and Westerdikella sp., PCCH and TCCH were also observed (Table 3).
Genetic analyses
The pairwise comparison of fungal ITS and Btub2 sequences with those available in the public online databases partially confirmed the identity at the species level of Penicillium simplicissimum (Oudem.) Thom (Btub2 100% similarity with KR709181.1 P. simplicissimum from culture collection MUT, Italy; ITS 99.65% similarity with sequence MH856014.1 in GenBank corresponding to P. simplicissimum from culture collection CBS 280.39). The Penicillium strain, however, showed high similarity according to both ITS and BTub2 also with P. ochrochloron Biourge (ITS 99.83% similarity with AJ509865.1), and P. pulvillorum Turfitt (Btub2 99.37% similarity with GU981670.1 from culture-collection CBS:280.39).
The pairwise comparison of fungal ITS and Tef1 sequences confirmed the identity at the species level of Trichoderma harzianum Rifai (Kunze) (Tef1 98.21% similarity with sequence MF125298.1 in GenBank; TrichoKey confirmed the identification as T. harzianum on the basis of Tef1 sequence that contains region 1 (9 nt), region 2 (158 nt) and region 3 (196 nt) genetic markers). The Trichoderma ITS1, 5.8S ribosomal RNA gene, ITS2 and partial LSU sequence and large subunit ribosomal RNA gene showed 99.84% similarity with GenBank sequence MH651391.1 corresponding to T. citrinoviride Bisset, and 99.84% similarity also with sequence MH651377.1, corresponding to a T. harzianum isolate. The pairwise comparison of Trichoderma ITS in UNITE database confirmed a best match with Trichoderma harzianum (KC403944).
ITS sequences for Penicillium simplicissimum FBL 573 and Trichoderma harzianum FBL 576 were deposited in GenBank with the accession numbers MK789684 and MK793273, respectively; P. simplicissimum beta-tubulin gene and T. harzianum TEF1 region were also deposited in GenBank with the accession numbers MK840446 and MK840447, respectively.
Oxidative stress analysis
The results showed that high HCH levels promoted ROS production (Fig. 7a). In both species, it was possible to highlight a similar trend with an increase of about 42% of ROS compared to the control.
ROS production by DCF fluorescence (a) and antioxidant enzyme activities (b–d) in P. simplicissimum and T. harzianum in the control and treatment exposed to HCH isomers, respectively. SOD (b), GST (c) and CAT (d) activities were normalised to protein content and expressed as unit/milligram. The data were expressed as a mean ± SE of independent biological replicates. Asterisk denotes a significant difference between the treatment and the control (*p < 0.05; **p < 0.005; ***p < 0.0001).
The three analysed antioxidant enzyme activities, namely SOD, CAT and GST, are shown in Fig. 7b–d. GST and CAT activities increased compared to the control in both species. SOD activity increased only in P. simplicissimum, while no increase was observed in T. harzianum compared to the control (Fig. 7b).
Discussion
The isolated species from the study area mainly belonged to the phylum Ascomycota. It has been reported in several studies that this phylum is the most common in contaminated soils, being the most diverse group of fungi which can colonise most ecological niches (Harms et al. 2011; Marco-Urrea et al. 2015; Godoy et al. 2016). By comparing the results from a natural wood wetland (WW) with the amended ones (WW-HCH), only a slight decrease in terms of the richness of species was reported; as such indigenous fungi were found to be HCH tolerant, they may play a role in increasing the efficiency of bioremediation at contaminated sites. The high species richness observed by comparing the results from WW with those of the other matrices (N-A, N-B, N-C and N-D) may also be related to the presence of a limited number of common species. This resulted in a considerable heterogeneity, probably due to the diversity of organic matter in the reactors. It is known that successful mycoremediation requires the selection of fungi that tolerate the contamination and are well adapted to the environmental conditions of the site (D’Annibale et al. 2006; Marco-Urrea et al. 2015; Czaplicki et al. 2016; Godoy et al. 2016). Indeed, several researchers suggested that, in this context, the indigenous fungal isolates are more tolerant and can be an excellent tool for biotransformation and biodegradation of contaminants, being linked to long-term exposure with a consequent adaptation to a high selective pressure (Godoy et al. 2016; Awasthi et al. 2017; Gururajan and Belur 2018). According to Spina et al. (2018), in order to accelerate the natural attenuation processes at contaminated sites, it is beneficial to lean towards solutions inspired by nature. In this way, natural wetlands have been used as biological filters to cope with different environmental and water quality issues (Sheoran and Sheoran 2006). These systems rely on different basic processes including intrinsic contaminant biodegradation by indigenous microorganisms in association with plants (Yu et al. 2005; Sheoran and Sheoran 2006; Fernandes and Guiomar 2018). On the other hand, no fungi from the N-C reactor samples were isolated, raising questions about the effects of this treatment system on fungal biodiversity and functionality. This result may be explained by the high Fe concentration and very reductive and anaerobic conditions, given that this reactor consists of a barrier with zero valent iron for chemical reduction of HCH isomers. The impact of these barriers on fungal assemblages should be carefully assessed for the possible toxic effects involved in the context of a wider integration of biological and physicochemical approaches. The microbiological characterisation of the semipermeable barriers has only been previously studied regarding bacteria (Gu et al. 1999; Scherer et al. 2000). Methanogens, sulphate reducers and iron-reducing bacteria were found within and downgradient of the barrier, while iron oxidising bacteria could exploit the increased availability of ferrous iron near the upgradient face of the barrier. A wide variety of geochemical transitions, chemical redox and precipitation reactions can be stimulated, associated with the concentration of iron chemical species, dissolved oxygen and CO2 consumption, and cathodic hydrogen production (Scherer et al. 2000). Consequently, the fungal occurrence in the context of Fe0 barriers has received little or no attention in the literature. In the N-A, N-B and N-D reactors, fungal species richness resulted in a high number of isolated taxa. These reactors provided different physicochemical conditions, from aerobic (N-A) to relatively anaerobic (N-B), and different organic matrices for xenobiotic biosorption, biostimulation and biotransformation. Despite the relative anaerobic conditions in N-B, taxonomic and functional fungal biodiversity was not substantially affected as it was in the N-C reactor. Biosorption of xenobiotic compounds on organic matrices and microbial biomass may be the first step in the biodegradation process of HCH isomers in these reactors. Soil, gravel and emerged plants in N-A, green mulch, ground grain, bark and wood chips in N-B, and peat, bark and wood chips in N-D, may have created favourable conditions for bioremediation. Biostimulation of microbial activity by different organic matrices is considered a powerful remediation strategy to accelerate the degradation of xenobiotics in the soils through the appropriate utilisation of organic amendments and nutrients (Kanissery and Sims 2011; Adams et al. 2015). Moreover, organic amendment can synergistically affect the adsorption, movement and biodegradation of pesticides (Diez 2010). Pesticide biotransformations may occur via metabolism or co-metabolism with specific organic substrates in a multi-step process in which the synergistic actions of plants and microorganisms can improve the bioremediation process (Hoagland et al. 2000; du Jardin 2015). With regard to the experimental conditions of the studied reactors which simulated constructed wetlands, several studies are reported in the literature supporting the functional roles played by fungi (Groudeva et al. 2001; Giraud et al. 2001).
In the tolerance tests, all the fungi grew without significant inhibition in terms of growth rate, biomass production, and high percentage tolerance index values (Fig. 5 and Table 1). This high tolerance may be explained by detoxification resulting from defence mechanisms implemented by the fungi, but less information has been produced verifying the correlation between the tolerance and detoxification strategies of fungi (Chen et al. 2015). The criteria for selecting the two taxa (P. simplicissimum and T. harzianum) to assess the oxidative stress responses are reported in the “Materials and methods” section. The genera Trichoderma and Penicillium have been described in several studies as potential bioremediators for environmental remediation. Tripathi et al. (2013) reported the genus Trichoderma as being tolerant to a range of recalcitrant pollutants including toxic metals, pesticides and polyaromatic hydrocarbons. Argumedo-Delira et al. (2012) reported tolerance and growth of 11 Trichoderma strains to crude oil, naphthalene, phenanthrene and benzo[a]pyrene. Moreover, studies have been performed on Penicillium spp. acting as fungal strains degrading hazardous chemicals (Taşeli 2006; Leitão 2009; Zhao et al. 2010; Gonçalves 2012; Alvarenga et al. 2014; Ceci et al. 2015a).
Results of oxidative stress responses showed an increase in ROS production in both of the tested taxa (Fig. 7a). In fact, fungi can adapt to an increase in ROS production by upregulating the activity of their antioxidant enzymes, particularly CAT and SOD, which represent a line of defence against stress oxidative conditions (Bai et al. 2003). The importance of antioxidant enzymes in the response to oxidative stress in filamentous fungi is reported in several other studies (Emri et al. 1997; Kawasaki and Aguirre 2001; Kreiner et al. 2002; Verdin et al. 2004; Angelova et al. 2005; Li et al. 2008; Chakraborty et al. 2013; Montibus et al. 2015). These enzymes have physiological functions under non-stressed conditions, but their activities may increase under oxidative stress, and they can also be used as biomarkers (Srivastava and Thakur 2006; Galdiero et al. 2016, 2017). Chakraborty et al. (2013) demonstrated that the good tolerance and growth of Aspergillus niger in the presence of elevated concentrations of Pb(II) was related to the increased activities of antioxidant enzymes. Nevertheless, it may be difficult to predict the behaviour of fungal cultures exposed to stress conditions, underlining the fact that the responses may be different and highly specific (Bai et al. 2003). Addition of definite concentrations of exogenous oxidative stressors provides a simulation of oxidative stress in microorganisms and these studies must be interpreted with care (Bai et al. 2003). Results showed that SOD activity significantly increased under HCH stress conditions compared to the control in P. simplicissimum, while SOD activity did not significantly increase in T. harzianum. CAT and GST increased their activity in both of the tested taxa (Fig. 7b–d). Similar results have been reported by other authors; e.g. Kreiner et al. (2002) reported that the effects of menadione in A. niger resulted in an increase in CAT activity but did not increase specific SOD activity. This supposed that the SOD levels at the start of menadione stress were high enough to dismutate the generated superoxide radicals resulting in an immediate increase of specific CAT activity as primary response, and later, after adding higher amounts of MD to the culture, SOD activity also started to increase. On the other hand, the decline in activity of antioxidant enzymes may correspond to complete detoxification according to Li et al. (2008). In this sense, further investigation is necessary to deepen the knowledge of enzyme activity, despite the fact that the number of studies of oxidative stress have increased in the last few years in different fields like medicine, toxicology and environmental science (Lushchak 2011).
In the bioremediation of halogenated pollutants, dehalogenation is the first important reaction for the removal of halogen atoms (Camacho-Pérez et al. 2012; Singh 2017). The toxicity and the persistence in the atmosphere of a xenobiotic compound are generally related to the degree of halogenation and the stability of the carbon-halogen bond (Singh 2017). HCH and its isomers have six chlorine atoms and their removal is a very significant step for the reduction of both the biodegradation recalcitrance and the risk of forming toxic intermediates during subsequent biotransformation steps (Nagata et al. 2007; Camacho-Pérez et al. 2012; Singh 2017). In this study, dechlorination intermediates were observed in the tested fungi. The first step of the biotransformation process, resulting in the formation of pentachlorocyclohexene (PCCH), was observed for all the tested species, except for M. hiemalis f. hiemalis and Trichoderma sp. (Table 3). For these species, PCCH may be totally transformed or occur in concentrations below the detection limits. This intermediate was reported in different studies on HCH dehalogenation under aerobic conditions with bacteria and fungi (Phillips et al. 2005; Camacho-Pérez et al. 2012; Guillén-Jiménez et al. 2012; Ceci et al. 2015a, 2018). Referring to non-white-rot fungi, PCCH was reported in different investigations. PCCH isomers were observed in the degradation of lindane and other HCH isomers by Fusarium verticillioides AT-100, Candida VITJzN04, Rhodotorula sp. VITJzN03 and Penicillium griseofulvum FBL 500 (Guillén-Jiménez et al. 2012; Salam et al. 2013; Salam and Das 2014; Ceci et al. 2015a, 2018). The presence of tetrachlorocyclohexene (TCCH) in seven species was confirmed by concentrating the samples and enhancing the resolution of qualitative analysis (Table 3). This metabolite has been generally observed for several bacterial species and in fungi (Phillips et al. 2005; Singh 2017; Ceci et al. 2018). In white-rot fungi, γ-tetrachlorocyclohexene (γ-TCCH) has been reported as the initial metabolite of lindane biodegradation (Phillips et al. 2005). In Trametes hirsutus, Phanerochaete chrysosporium, Cyathus bulleri and P. sordida, TCCH has been found after lindane dechlorination (Mougin et al. 1996; Singh and Kuhad 1999, 2000; Phillips et al. 2005). In non-white-rot fungi, 1,3,4,6-tetrachloro-1,4-cyclohexadiene (1,4-TCCHdiene) and TCCH have been observed in Rhodotorula sp. VITJzN03 and P. griseofulvum FBL 500, respectively (Salam et al. 2013; Ceci et al. 2018). Through a final series of dehydrochlorination reactions, mixed isomers of chlorinated benzene intermediates may result from HCH biotransformation, namely trichlorobenzene (TCB), dichlorobenzene (DCB) and monochlorobenzene (CB). TCB was detected only for P. dimorphospora and Sphaeropsidales sp., while DCB and CB were observed for all the tested species (Table 3). As these taxa could likely possess different enzymes with specific metabolic and physiologic peculiarities, their complete pathways of HCH biodegradation may be very different, resulting for instance in different dechlorination intermediates, as was observed in this research. For instance, P. dimorphospora and Sphaeropsidales sp. showed all the intermediates of dechlorination from PCCH to CB: for these species, a similar pathway of HCH biodegradation to that observed previously for Rhodotorula sp. VITJzN03 and P. griseofulvum FBL 500 can be hypothesised (Salam et al. 2013; Ceci et al. 2018). Lastly, in this research, other metabolites belonging to the successful steps of HCH biodegradation were not detected. This may be due to (1) interactions between the growth mycelia, fungal enzymes, growth medium and pollutants; (2) the biodegradation kinetics involved; and (3) the low detection limits of the transient products and their different chemical-physical features (volatility and polarity) (Ulčnik et al. 2012; Guillén-Jiménez et al. 2012). Furthermore, the bioremediation potential of the tested taxa needs to be considered since several case studies are reported in literature for the biodegradation of different organic pollutants. Synergistic rhizosphere degradation of γ-HCH (lindane) through the combinatorial plant-fungal action among Megathyrsus maximus and some species belonging to Talaromyces, Aspergillus and Yarrowia genera has been observed and different intermediates of HCH biodegradation (PCCH, TCB, DCB, CB) have been detected (Asemoloye et al. 2017). Phialophora fastigiata (Basionym: Cadophora fastigiata) has been reported to degrade wood preservatives based on imidazolium compounds and quaternary ammonium compounds, scots pine and beech wood exposed used for testing of wood preservatives, and pentachlorophenol (Benoit-Guyod et al. 1994; Zabielska-Matejuk and Czaczyk 2006; Råberg et al. 2013). Metarhizium cylindrosporae has been reported to degrade the chlorinated herbicide acetochlor under stirred culture media (Erguven 2018), while biodegradation of ametryn, a representative of a class of s-triazine herbicides, by the entomopathogenic cosmopolite fungus Metarhizium brunneum has been observed (Szewczyk et al. 2018). Mucor hiemalis has been observed to be able to degrade diclofenac and endosulfan (Martens 1976; Esterhuizen-Londt et al. 2017). P. simplicissimum has been reported to transform recalcitrant pollutants, such as polyethylene and halophenols (Marr et al. 1996; Leitão 2009). P. dimorphospora and two Trichoderma species have been isolated from creosote-treated crosstie wastes resulting in them being resistant to polycyclic aromatic hydrocarbons (Kim et al. 2010). T. harzianum has been reported to degrade different organochlorine pesticides (Katayama and Matsumura 1993). A Zopfiella species, Z. karachiensis, has successfully been tested for the biodegradation of polyester polyurethane along with other endophytic fungi (Russell et al. 2011). In this study, the medium pH increased in most of studied taxa, resulting in a medium alkalinisation to neutral condition (Fig. 6 and Table 2). On the contrary, the HCH dehydrochlorination resulted in a decrease in the culture pH value in Pandorea species, probably due to the production of acidic metabolites during the metabolism of HCH (Siddique et al. 2003; Guillén-Jiménez et al. 2012). In fungi, acidic pH could lead to toxic effects due to the production of intermediates, such as benzoates, during HCH biotransformation (Guillén-Jiménez et al. 2012; Ceci et al. 2015a, 2018). Hence, fungal modulation of medium pH may be fundamental in HCH biotransformation. It is worth mentioning that fungal modulation can also regulate a plethora of enzymes, such as those implicated in fungal pathogenicity (Alkan et al. 2013). In Paecilomyces marquandii, a neutral pH of the medium has proven to be favourable both for alachlor biodegradation and for the reduction of ROS production during biodegradation (Słaba et al. 2015). The efficiency of fungal decolourisation and the enzymatic process involved is highly pH-dependent with the optimal pH range for biodegradation being from 3 to 6 (Prasad 2017).
Conclusion
The use of green technologies and ecosystem management has become a necessity for environmental remediation due to the worldwide existence of millions of sites contaminated with organic pollutants. It should be kept in mind that in terms of sustainability, bioremediation is able to provide an effective reduction of hazardous chemicals. In this research, fungal assemblages of a site historically contaminated with high concentrations of HCHs were isolated and characterised from different in situ pilot reactors. Based on the experimental results, most of the tested fungi exhibited elevated tolerance and a good response to oxidative stress caused by HCH isomers. The selected saprotrophic fungal species were able to biotransform isomers of HCH mixtures at high concentrations. In addition, the data presented in this study underline the relevant role that indigenous fungal species may play in the overall remedial process. Fungal bioremediation as a complex process requires monitoring in planned periods to assess the efficiency of xenobiotic biodegradation and its implementation.
References
Abarenkov K, Henrik Nilsson R, Larsson K-H, Alexander IJ, Eberhardt U, Erland S, Høiland K, Kjøller R, Larsson E, Pennanen T, Sen R, Taylor AFS, Tedersoo L, Ursing BM, Vrålstad T, Liimatainen K, Peintner U, Kõljalg U (2010) The UNITE database for molecular identification of fungi - recent updates and future perspectives: letters. New Phytol 186:281–285. https://doi.org/10.1111/j.1469-8137.2009.03160.x
Adams GO, Fufeyin PT, Okoro SE, Ehinomen I (2015) Bioremediation, biostimulation and bioaugmention: a review. Int J Environ Bioremediat Biodegrad 3:28–39. https://doi.org/10.12691/ijebb-3-1-5
Alkan N, Espeso EA, Prusky D (2013) Virulence regulation of phytopathogenic fungi by pH. Antioxid Redox Signal 19:1012–1025. https://doi.org/10.1089/ars.2012.5062
Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997). Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Research, 25:3389-3402. https://doi.org/10.1093/nar/25.17.3389
Alvarenga N, Birolli WG, Seleghim MHR, Porto ALM (2014) Biodegradation of methyl parathion by whole cells of marine-derived fungi Aspergillus sydowii and Penicillium decaturense. Chemosphere 117:47–52. https://doi.org/10.1016/j.chemosphere.2014.05.069
Anastasi A, Tigini V, Varese GC (2013) The bioremediation potential of different ecophysiological groups of fungi. In: Goltapeh EM, Danesh YR, Varma A (eds) Fungi as bioremediators, Soil Biology, vol 32. Springer, Berlin, pp 29–49. https://doi.org/10.1007/978-3-642-33811-3_2
Angelova MB, Pashova SB, Spasova BK, Vassilev SV, Slokoska LS (2005) Oxidative stress response of filamentous fungi induced by hydrogen peroxide and paraquat. Mycol Res 109:150–158. https://doi.org/10.1017/S0953756204001352
Argumedo-Delira R, Alarcón A, Ferrera-Cerrato R, Almaraz JJ, Peña-Cabriales JJ (2012) Tolerance and growth of 11 Trichoderma strains to crude oil, naphthalene, phenanthrene and benzo[a]pyrene. J Environ Manag 95:S291–S299. https://doi.org/10.1016/j.jenvman.2010.08.011
Asemoloye MD, Ahmad R, Jonathan SG (2017) Synergistic rhizosphere degradation of γ-hexachlorocyclohexane (lindane) through the combinatorial plant-fungal action. PLoS One 12:e0183373. https://doi.org/10.1371/journal.pone.0183373
Ashraf MA (2017) Persistent organic pollutants (POPs): a global issue, a global challenge. Environ Sci Pollut Res 24:4223–4227. https://doi.org/10.1007/s11356-015-5225-9
Awasthi AK, Pandey AK, Khan J (2017) A preliminary report of indigenous fungal isolates from contaminated municipal solid waste site in India. Environ Sci Pollut Res 24:8880–8888. https://doi.org/10.1007/s11356-017-8472-0
Bai Z, Harvey LM, McNeil B (2003) Oxidative stress in submerged cultures of fungi. Crit Rev Biotechnol 23:267–302. https://doi.org/10.1080/07388550390449294
Benoit-Guyod J-L, Seigle-Murandi F, Steiman R, Sage L, Toe A (1994) Biodegradation of pentachlorophenol by micromycetes. III. Deuteromycetes. Environ Toxicol Water Qual 9:33–44. https://doi.org/10.1002/tox.2530090106
Borgå K, Gabrielsen G, Skaare J (2001) Biomagnification of organochlorines along a Barents Sea food chain. Environ Pollut 113:187–198. https://doi.org/10.1016/S0269-7491(00)00171-8
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Caicedo P, Schröder A, Ulrich N, Schröter U, Paschke A, Schüürmann G, Ahumada I, Richter P (2011) Determination of lindane leachability in soil–biosolid systems and its bioavailability in wheat plants. Chemosphere 84:397–402. https://doi.org/10.1016/j.chemosphere.2011.03.070
Camacho-Pérez B, Ríos-Leal E, Rinderknecht-Seijas N, Poggi-Varaldo HM (2012) Enzymes involved in the biodegradation of hexachlorocyclohexane: a mini review. J Environ Manag 95:S306–S318. https://doi.org/10.1016/j.jenvman.2011.06.047
Ceci A, Pierro L, Riccardi C, Pinzari F, Maggi O, Persiani AM, Gadd GM, Petrangeli Papini M (2015a) Biotransformation of β-hexachlorocyclohexane by the saprotrophic soil fungus Penicillium griseofulvum. Chemosphere 137:101–107. https://doi.org/10.1016/j.chemosphere.2015.05.074
Ceci A, Rhee YJ, Kierans M, Hillier S, Pendlowski H, Gray N, Persiani AM, Gadd GM (2015b) Transformation of vanadinite [Pb5(VO4)3Cl] by fungi: fungal biotransformation of vanadinite. Environ Microbiol 17:2018–2034. https://doi.org/10.1111/1462-2920.12612
Ceci A, Pinzari F, Riccardi C, Maggi O, Pierro L, Petrangeli Papini M, Gadd GM, Persiani AM (2018) Metabolic synergies in the biotransformation of organic and metallic toxic compounds by a saprotrophic soil fungus. Appl Microbiol Biotechnol 102:1019–1033. https://doi.org/10.1007/s00253-017-8614-9
Chakraborty S, Mukherjee A, Das TK (2013) Biochemical characterization of a lead-tolerant strain of Aspergillus foetidus: an implication of bioremediation of lead from liquid media. Int Biodeterior Biodegrad 84:134–142. https://doi.org/10.1016/j.ibiod.2012.05.031
Chen R, Zhou Z, Liu Y, Jiang J, Li Q, Song H, Pei D, Xu H (2015) Mycoremediation potential and tolerance responses of Oudemansiella radicata in cadmium-pyrene co-contaminated soil. J Soils Sediments 15:1083–1093. https://doi.org/10.1007/s11368-015-1093-7
Czaplicki LM, Cooper E, Ferguson PL, Stapleton HM, Vilgalys R, Gunsch CK (2016) A new perspective on sustainable soil remediation-case study suggests novel fungal genera could facilitate in situ biodegradation of hazardous contaminants: a new perspective on sustainable soil remediation. Remediat J 26:59–72. https://doi.org/10.1002/rem.21458
D’Annibale A, Rosetto F, Leonardi V et al (2006) Role of autochthonous filamentous fungi in bioremediation of a soil historically contaminated with aromatic hydrocarbons. Appl Environ Microbiol 72:28–36. https://doi.org/10.1128/AEM.72.1.28-36.2006
Deshmukh R, Khardenavis AA, Purohit HJ (2016) Diverse metabolic capacities of fungi for bioremediation. Indian J Microbiol 56:247–264. https://doi.org/10.1007/s12088-016-0584-6
Diez M (2010) Biological aspects involved in the degradation of organic pollutants. J Soil Sci Plant Nutr 10:244–267
Domsch KH, Gams W, Anderson T-H (2007) Compendium of soil fungi, 2. ed., taxonomically rev. IHW-Verl, Eching
Druzhinina IS, Kopchinskiy AG, Komoń M, Bissett J, Szakacs G, Kubicek CP (2005) An oligonucleotide barcode for species identification in Trichoderma and Hypocrea. Fungal Genet Biol 42:813–828. https://doi.org/10.1016/j.fgb.2005.06.007
du Jardin P (2015) Plant biostimulants: definition, concept, main categories and regulation. Sci Hortic 196:3–14. https://doi.org/10.1016/j.scienta.2015.09.021
Ellis MB (1976) More dematiaceous hyphomycetes. Commonwealth Mycological Institute, Kew
Emri T, Pócsi I, Szentirmai A (1997) Glutathione metabolism and protection against oxidative stress caused by peroxides in Penicillium chrysogenum. Free Radic Biol Med 23:809–814. https://doi.org/10.1016/S0891-5849(97)00065-8
Erguven GO (2018) Comparison of some soil fungi in bioremediation of herbicide acetochlor under agitated culture media. Bull Environ Contam Toxicol 100:570–575. https://doi.org/10.1007/s00128-018-2280-1
Esterhuizen-Londt M, Hendel A-L, Pflugmacher S (2017) Mycoremediation of diclofenac using Mucor hiemalis. Toxicol Environ Chem 99:795–808. https://doi.org/10.1080/02772248.2017.1296444
Fernandes JP, Guiomar N (2018) Nature-based solutions: the need to increase the knowledge on their potentialities and limits. Land Degrad Dev 29:1925–1939. https://doi.org/10.1002/ldr.2935
Gadd GM (2001) Fungi in bioremediation. Cambridge Univ. Press, Cambridge
Galdiero E, Siciliano A, Maselli V, Gesuele R, Guida M, Fulgione D, Galdiero S, Lombardi L, Falanga A (2016) An integrated study on antimicrobial activity and ecotoxicity of quantum dots and quantum dots coated with the antimicrobial peptide indolicidin. Int J Nanomedicine 11:4199–4211. https://doi.org/10.2147/IJN.S107752
Galdiero E, Falanga A, Siciliano A, Maselli V, Guida M, Carotenuto R, Tussellino M, Lombardi L, Benvenuto G, Galdiero S (2017) Daphnia magna and Xenopus laevis as in vivo models to probe toxicity and uptake of quantum dots functionalized with gH625. Int J Nanomedicine 12:2717–2731. https://doi.org/10.2147/IJN.S127226
Gams W, Dingley JM (2006) Hypocrea and Trichoderma studies marking the 90th birthday of Joan M. Dingley. CBS, Centraalbureau voor Schimmelcultures, Utrecht
Giraud F, Guiraud P, Kadri M, Blake G, Steiman R (2001) Biodegradation of anthracene and fluoranthene by fungi isolated from an experimental constructed wetland for wastewater treatment. Water Res 35:4126–4136. https://doi.org/10.1016/S0043-1354(01)00137-3
Giri K, Rawat AP, Rawat M, Rai JPN (2014) Biodegradation of hexachlorocyclohexane by two species of Bacillus isolated from contaminated soil. Chem Ecol 30:97–109. https://doi.org/10.1080/02757540.2013.844795
Glass NL, Donaldson GC (1995) Development of primer sets designed for use with the PCR to amplify conserved genes from filamentous ascomycetes. Appl Environ Microbiol 61:1323–1330
Godoy P, Reina R, Calderón A, Wittich RM, García-Romera I, Aranda E (2016) Exploring the potential of fungi isolated from PAH-polluted soil as a source of xenobiotics-degrading fungi. Environ Sci Pollut Res 23:20985–20996. https://doi.org/10.1007/s11356-016-7257-1
Gonçalves MS (2012) Isolation of filamentous fungi present in swine wastewater that are resistant and with the ability to remove atrazine. Afr J Biotechnol 11:11074–11077. https://doi.org/10.5897/AJB11.4018
Groudeva VI, Groudev SN, Doycheva AS (2001) Bioremediation of waters contaminated with crude oil and toxic heavy metals. Int J Miner Process 62:293–299. https://doi.org/10.1016/S0301-7516(00)00060-0
Gu B, Phelps TJ, Liang L, Dickey MJ, Roh Y, Kinsall BL, Palumbo AV, Jacobs GK (1999) Biogeochemical dynamics in zero-valent iron columns: implications for permeable reactive barriers. Environ Sci Technol 33:2170–2177. https://doi.org/10.1021/es981077e
Guillén-Jiménez F de M, Cristiani-Urbina E, Cancino-Díaz JC et al (2012) Lindane biodegradation by the Fusarium verticillioides AT-100 strain, isolated from Agave tequilana leaves: kinetic study and identification of metabolites. Int Biodeterior Biodegrad 74:36–47. https://doi.org/10.1016/j.ibiod.2012.04.020
Gurung B, Race M, Fabbricino M, Komínková D, Libralato G, Siciliano A, Guida M (2018) Assessment of metal pollution in the Lambro Creek (Italy). Ecotoxicol Environ Saf 148:754–762. https://doi.org/10.1016/j.ecoenv.2017.11.041
Gururajan K, Belur PD (2018) Screening and selection of indigenous metal tolerant fungal isolates for heavy metal removal. Environ Technol Innov 9:91–99. https://doi.org/10.1016/j.eti.2017.11.001
Harms H, Schlosser D, Wick LY (2011) Untapped potential: exploiting fungi in bioremediation of hazardous chemicals. Nat Rev Microbiol 9:177–192. https://doi.org/10.1038/nrmicro2519
Hoagland RE, Zablotowicz RM, Hall JC (2000) Pesticide metabolism in plants and microorganisms: an overview. In: Hall JC, Hoagland RE, Zablotowicz RM (eds) Pesticide biotransformation in plants and microorganisms: similarities and divergences, ACS Symposium Series, vol 777. American Chemical Society, Washington, DC, pp 2–27. https://doi.org/10.1021/bk-2001-0777.ch001
Hu W, Wang T, Khim JS, Luo W, Jiao W, Lu Y, Naile JE, Chen C, Zhang X, Giesy JP (2010) HCH and DDT in sediments from marine and adjacent riverine areas of North Bohai Sea, China. Arch Environ Contam Toxicol 59:71–79. https://doi.org/10.1007/s00244-009-9455-z
Hussain S, Arshad M, Springael D, SøRensen SR, Bending GD, Devers-Lamrani M, Maqbool Z, Martin-Laurent F (2015) Abiotic and biotic processes governing the fate of phenylurea herbicides in soils: a review. Crit Rev Environ Sci Technol 45:1947–1998. https://doi.org/10.1080/10643389.2014.1001141
Jaklitsch WM, Komon M, Kubicek CP, Druzhinina IS (2005) Hypocrea voglmayrii sp. nov. from the Austrian Alps represents a new phylogenetic clade in Hypocrea/Trichoderma. Mycologia 97:1365–1378. https://doi.org/10.1080/15572536.2006.11832743
Kanissery RG, Sims GK (2011) Biostimulation for the enhanced degradation of herbicides in soil. Appl Environ Soil Sci 2011:1–11. https://doi.org/10.1155/2011/843450
Karsch-Mizrachi I, Nakamura Y, Cochrane G, on behalf of the International Nucleotide Sequence Database Collaboration (2012) The international nucleotide sequence database collaboration. Nucleic Acids Res 40:D33–D37. https://doi.org/10.1093/nar/gkr1006
Katayama A, Matsumura F (1993) Degradation of organochlorine pesticides, particularly endosulfan, by Trichoderma harzianum. Environ Toxicol Chem 12:1059–1065. https://doi.org/10.1002/etc.5620120612
Kawasaki L, Aguirre J (2001) Multiple catalase genes are differentially regulated in Aspergillus nidulans. J Bacteriol 183:1434–1440. https://doi.org/10.1128/JB.183.4.1434-1440.2001
Kim M-J, Lee H, Choi Y-S, Kim GH, Huh NY, Lee S, Lim YW, Lee SS, Kim JJ (2010) Diversity of fungi in creosote-treated crosstie wastes and their resistance to polycyclic aromatic hydrocarbons. Antonie Van Leeuwenhoek 97:377–387. https://doi.org/10.1007/s10482-010-9416-6
Kõljalg U, Larsson K-H, Abarenkov K, Nilsson RH, Alexander IJ, Eberhardt U, Erland S, Høiland K, Kjøller R, Larsson E, Pennanen T, Sen R, Taylor AF, Tedersoo L, Vrålstad T, Ursing BM (2005) UNITE: a database providing web-based methods for the molecular identification of ectomycorrhizal fungi: methods. New Phytol 166:1063–1068. https://doi.org/10.1111/j.1469-8137.2005.01376.x
Komínková D, Fabbricino M, Gurung B, Race M, Tritto C, Ponzo A (2018) Sequential application of soil washing and phytoremediation in the land of fires. J Environ Manag 206:1081–1089. https://doi.org/10.1016/j.jenvman.2017.11.080
Kreiner M, Harvey LM, McNeil B (2002) Oxidative stress response of a recombinant Aspergillus niger to exogenous menadione and H2O2 addition. Enzym Microb Technol 30:346–353. https://doi.org/10.1016/S0141-0229(01)00517-8
Kulshreshtha S, Mathur N, Bhatnagar P (2014) Mushroom as a product and their role in mycoremediation. AMB Express 4:29. https://doi.org/10.1186/s13568-014-0029-8
Leitão AL (2009) Potential of Penicillium species in the bioremediation field. Int J Environ Res Public Health 6:1393–1417. https://doi.org/10.3390/ijerph6041393
Li Q, McNeil B, Harvey LM (2008) Adaptive response to oxidative stress in the filamentous fungus Aspergillus niger B1-D. Free Radic Biol Med 44:394–402. https://doi.org/10.1016/j.freeradbiomed.2007.09.019
Lushchak VI (2011) Adaptive response to oxidative stress: bacteria, fungi, plants and animals. Comp Biochem Physiol Part C Toxicol Pharmacol 153:175–190. https://doi.org/10.1016/j.cbpc.2010.10.004
Maggi O, Persiani AM, Casado MA, Pineda FD (2005) Effects of elevation, slope position and livestock exclusion on microfungi isolated from soils of Mediterranean grasslands. Mycologia 97:984–995. https://doi.org/10.3852/mycologia.97.5.984
Malloch D, Cain RF (1971) New cleistothecial Sordariaceae and a new family, Coniochaetaceae. Can J Bot 49:869–880. https://doi.org/10.1139/b71-127
Maqbool Z, Hussain S, Imran M, Mahmood F, Shahzad T, Ahmed Z, Azeem F, Muzammil S (2016) Perspectives of using fungi as bioresource for bioremediation of pesticides in the environment: a critical review. Environ Sci Pollut Res 23:16904–16925. https://doi.org/10.1007/s11356-016-7003-8
Marco-Urrea E, García-Romera I, Aranda E (2015) Potential of non-ligninolytic fungi in bioremediation of chlorinated and polycyclic aromatic hydrocarbons. New Biotechnol 32:620–628. https://doi.org/10.1016/j.nbt.2015.01.005
Marr J, Kremer S, Sterner O, Anke H (1996) Transformation and mineralization of halophenols by Penicillium simplicissimum SK9117. Biodegradation 7:165–171. https://doi.org/10.1007/BF00114628
Martens R (1976) Degradation of [8,9,-14C]endosulfan by soil microorganisms. Appl Environ Microbiol 31:853–858
Montibus M, Pinson-Gadais L, Richard-Forget F, Barreau C, Ponts N (2015) Coupling of transcriptional response to oxidative stress and secondary metabolism regulation in filamentous fungi. Crit Rev Microbiol 41:295–308. https://doi.org/10.3109/1040841X.2013.829416
Morillo E, Villaverde J (2017) Advanced technologies for the remediation of pesticide-contaminated soils. Sci Total Environ 586:576–597. https://doi.org/10.1016/j.scitotenv.2017.02.020
Mougin C, Pericaud C, Malosse C, Laugero C, Asther M (1996) Biotransformation of the insecticide lindane by the white rot basidiomycete Phanerochaete chrysosporium. Pestic Sci 47:51–59. https://doi.org/10.1002/(SICI)1096-9063(199605)47:1<51::AID-PS391>3.0.CO;2-V
Mrema EJ, Rubino FM, Brambilla G, Moretto A, Tsatsakis AM, Colosio C (2013) Persistent organochlorinated pesticides and mechanisms of their toxicity. Toxicology 307:74–88. https://doi.org/10.1016/j.tox.2012.11.015
Nadal M, Marquès M, Mari M, Domingo JL (2015) Climate change and environmental concentrations of POPs: a review. Environ Res 143:177–185. https://doi.org/10.1016/j.envres.2015.10.012
Nagata Y, Endo R, Ito M et al (2007) Aerobic degradation of lindane (γ-hexachlorocyclohexane) in bacteria and its biochemical and molecular basis. Appl Microbiol Biotechnol 76:752. https://doi.org/10.1007/s00253-007-1066-x
Nawab A, Aleem A, Malik A (2003) Determination of organochlorine pesticides in agricultural soil with special reference to gamma-HCH degradation by Pseudomonas strains. Bioresour Technol 88:41–46
Persiani AM, Maggi O, Montalvo J, Casado MA, Pineda FD (2008) Mediterranean grassland soil fungi: patterns of biodiversity, functional redundancy and soil carbon storage. Plant Biosyst 142:111–119. https://doi.org/10.1080/11263500701872713
Phillips TM, Seech AG, Lee H, Trevors JT (2005) Biodegradation of hexachlorocyclohexane (HCH) by microorganisms. Biodegradation 16:363–392. https://doi.org/10.1007/s10532-004-2413-6
Pitt JI (1979) The genus Penicillium and its teleomorphic states Eupenicillium and Talaromyces. Academic Press, London
Prasad R (ed) (2017) Mycoremediation and environmental sustainability. Springer International Publishing, Cham
Råberg U, Terziev N, Daniel G (2013) Degradation of Scots pine and beech wood exposed in four test fields used for testing of wood preservatives. Int Biodeterior Biodegrad 79:20–27. https://doi.org/10.1016/j.ibiod.2012.12.010
Ramirez C, Martinez AT (1982) Manual and atlas of the Penicillia. Elsevier Biomedical Press ; Sole distributors for the USA and Canada, Elsevier/North-Holland, Amsterdam ; New York : New York, N.Y
Roze LV, Chanda A, Linz JE (2011) Compartmentalization and molecular traffic in secondary metabolism: a new understanding of established cellular processes. Fungal Genet Biol 48:35–48. https://doi.org/10.1016/j.fgb.2010.05.006
Russell JR, Huang J, Anand P, Kucera K, Sandoval AG, Dantzler KW, Hickman DS, Jee J, Kimovec FM, Koppstein D, Marks DH, Mittermiller PA, Núñez SJ, Santiago M, Townes MA, Vishnevetsky M, Williams NE, Vargas MPN, Boulanger LA, Bascom-Slack C, Strobel SA (2011) Biodegradation of polyester polyurethane by endophytic fungi. Appl Environ Microbiol 77:6076–6084. https://doi.org/10.1128/AEM.00521-11
Russo D, Siciliano A, Guida M, Galdiero E, Amoresano A, Andreozzi R, Reis NM, Li Puma G, Marotta R (2017) Photodegradation and ecotoxicology of acyclovir in water under UV254 and UV254/H2O2 processes. Water Res 122:591–602. https://doi.org/10.1016/j.watres.2017.06.020
Saez JM, Alvarez A, Fuentes MS, Amoroso MJ, Benimeli CS (2017) An overview on microbial degradation of lindane. In: Singh SN (ed) Microbe-induced degradation of pesticides. Springer International Publishing, Cham, pp 191–212
Salam JA, Das N (2014) Lindane degradation by Candida VITJzN04, a newly isolated yeast strain from contaminated soil: kinetic study, enzyme analysis and biodegradation pathway. World J Microbiol Biotechnol 30:1301–1313. https://doi.org/10.1007/s11274-013-1551-6
Salam JA, Lakshmi V, Das D, Das N (2013) Biodegradation of lindane using a novel yeast strain, Rhodotorula sp. VITJzN03 isolated from agricultural soil. World J Microbiol Biotechnol 29:475–487. https://doi.org/10.1007/s11274-012-1201-4
Samuels GJ, Hebbar PK (2015) Trichoderma: identification and agricultural applications. APS Press, St. Paul, Minn
Scherer MM, Richter S, Valentine RL, Alvarez PJJ (2000) Chemistry and microbiology of permeable reactive barriers for in situ groundwater clean up. Crit Rev Microbiol 26:221–264. https://doi.org/10.1080/10408410091154237
Scheringer M (2004) Persistent organic pollutants (POPs) in the focus of science and politics. Environ Sci Pollut Res 11:1–2. https://doi.org/10.1007/BF02980278
Sheoran AS, Sheoran V (2006) Heavy metal removal mechanism of acid mine drainage in wetlands: acritical review. Miner Eng 19:105–116. https://doi.org/10.1016/j.mineng.2005.08.006
Sherif A, Elhussein A (2011) Biodegradation of fungicide Thiram (TMTD) in soil under laboratory conditions. Am J Biotechnol Mol Sci 1:57–68. https://doi.org/10.5251/ajbms.2011.1.2.57.68
Siddique T, Okeke BC, Arshad M, Frankenberger WT (2003) Biodegradation kinetics of endosulfan by Fusarium ventricosum and a Pandoraea species. J Agric Food Chem 51:8015–8019. https://doi.org/10.1021/jf030503z
Singh H (2006) Mycoremediation: fungal bioremediation. Wiley, Hoboken
Singh SN (2017) Microbe-induced degradation of pesticides. Springer International Publishing, Cham
Singh BK, Kuhad RC (1999) Biodegradation of lindane (γ-hexachlorocyclohexane) by the white-rot fungus Trametes hirsutus. Lett Appl Microbiol 28:238–241. https://doi.org/10.1046/j.1365-2672.1999.00508.x
Singh BK, Kuhad RC (2000) Degradation of insecticide lindane (γ-HCH) by white-rot fungi Cyathus bulleri and Phanerochaete sordida. Pest Manag Sci 56:142–146. https://doi.org/10.1002/1526-4998(200002)56:2<142::AID-PS104>3.0.CO;2-I
Słaba M, Różalska S, Bernat P, Szewczyk R, Piątek MA, Długoński J (2015) Efficient alachlor degradation by the filamentous fungus Paecilomyces marquandii with simultaneous oxidative stress reduction. Bioresour Technol 197:404–409. https://doi.org/10.1016/j.biortech.2015.08.045
Spina F, Cecchi G, Landinez-Torres A, Pecoraro L, Russo F, Wu B, Cai L, Liu XZ, Tosi S, Varese GC, Zotti M, Persiani AM (2018) Fungi as a toolbox for sustainable bioremediation of pesticides in soil and water. Plant Biosyst 152:474–488. https://doi.org/10.1080/11263504.2018.1445130
Srivastava S, Thakur IS (2006) Evaluation of bioremediation and detoxification potentiality of Aspergillus niger for removal of hexavalent chromium in soil microcosm. Soil Biol Biochem 38:1904–1911. https://doi.org/10.1016/j.soilbio.2005.12.016
Szewczyk R, Kuśmierska A, Bernat P (2018) Ametryn removal by Metarhizium brunneum: biodegradation pathway proposal and metabolic background revealed. Chemosphere 190:174–183. https://doi.org/10.1016/j.chemosphere.2017.10.011
Taşeli BK (2006) Dehalogenation of lindane by Penicillium camemberti. Bull Environ Contam Toxicol 77:882–887. https://doi.org/10.1007/s00128-006-1226-1
Tripathi P, Singh PC, Mishra A, Chauhan PS, Dwivedi S, Bais RT, Tripathi RD (2013) Trichoderma: a potential bioremediator for environmental clean up. Clean Techn Environ Policy 15:541–550. https://doi.org/10.1007/s10098-012-0553-7
Tu CM (1994) Effects of herbicides and fumigants on microbial activities in soil. Bull Environ Contam Toxicol 53:12–17
Ulčnik A, Kralj Cigić I, Zupančič-Kralj L et al (2012) Bioremediation of lindane by wood-decaying fungi. Drv Ind 63:271–276
Verdin A, Sahraoui AL-H, Durand R (2004) Degradation of benzo[a]pyrene by mitosporic fungi and extracellular oxidative enzymes. Int Biodeterior Biodegrad 53:65–70. https://doi.org/10.1016/j.ibiod.2003.12.001
Vidali M (2001) Bioremediation. An overview. Pure Appl Chem 73:1163–1172. https://doi.org/10.1351/pac200173071163
Vijgen J, Abhilash PC, Li Y et al (2011) Hexachlorocyclohexane (HCH) as new Stockholm Convention POPs—a global perspective on the management of lindane and its waste isomers. Environ Sci Pollut Res 18:152–162. https://doi.org/10.1007/s11356-010-0417-9
von Arx JA, Müller E (1975) A re-evaluation of the bitunicate ascomycetes with keys to families and genera. Stud Mycol 9:1–159
Wacławek S, Antoš V, Hrabák P, Černík M, Elliott D (2016) Remediation of hexachlorocyclohexanes by electrochemically activated persulfates. Environ Sci Pollut Res 23:765–773. https://doi.org/10.1007/s11356-015-5312-y
White TJ, Bruns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ (eds) PCR Protocols: a guide to methods and applications. Academic Press, New York, pp 315–322
Yu KSH, Wong AHY, Yau KWY, Wong YS, Tam NFY (2005) Natural attenuation, biostimulation and bioaugmentation on biodegradation of polycyclic aromatic hydrocarbons (PAHs) in mangrove sediments. Mar Pollut Bull 51:1071–1077. https://doi.org/10.1016/j.marpolbul.2005.06.006
Zabielska-Matejuk J, Czaczyk K (2006) Biodegradation of new quaternary ammonium compounds in treated wood by mould fungi. Wood Sci Technol 40:461–475. https://doi.org/10.1007/s00226-005-0065-2
Zhao R, Bao H, Liu Y (2010) Isolation and characterization of Penicillium oxalicum ZHJ6 for biodegradation of methamidophos. Agric Sci China 9:695–703. https://doi.org/10.1016/S1671-2927(09)60145-0
Acknowledgements
We greatly thank Dr. Flavia Pinzari for her precious support in the genetic analysis and bioinformatics. M. Cernik thanks the Ministry of Education, Youth and Sports of the Czech Republic and the European Union - European Structural and Investment Funds in the frames of Operational Programme Research, Development and Education - project Hybrid Materials for Hierarchical Structures (HyHi, Reg. No. CZ.02.1.01/0.0/0.0/16_019/0000843).
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible editor: Philippe Garrigues
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Russo, F., Ceci, A., Maggi, O. et al. Understanding fungal potential in the mitigation of contaminated areas in the Czech Republic: tolerance, biotransformation of hexachlorocyclohexane (HCH) and oxidative stress analysis. Environ Sci Pollut Res 26, 24445–24461 (2019). https://doi.org/10.1007/s11356-019-05679-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-019-05679-w