Abstract
The growing use of silver nanoparticles (AgNPs) is likely to result in increased environmental contamination. Although AgNPs have been reported to affect microbial communities in a range of ecosystems, there is still a lack of information concerning the effect of low concentrations of AgNPs on soil microbial community structures and functional groups involved in biogeochemical cycling. In this study, the concentration-dependent effects of AgNPs and silver micron particles (AgMPs) on bacterial and fungal community structures in an agricultural pastureland soil were examined in a microcosm-based experiment using enzyme analysis, molecular fingerprinting, qPCR and amplicon sequencing. Soil enzyme processes were impacted by Ag contamination, with soil dehydrogenase activity reduced by 1 mg kg−1 of AgNPs and AgMPs. Soil urease activity was less susceptible, but was inhibited by ≥ 10 mg kg−1 AgNPs. The significant (P ≤ 0.001) decrease in copy numbers of the amoA gene by 10 mg kg−1 AgNPs indicated that archaea ammonia oxidisers may be more sensitive to AgNP contamination than bacteria. Amplicon sequencing revealed the bacterial phyla Acidobacteria and Verrucomicrobia to be highly sensitive to AgNP contamination. A broad reduction in the relative abundance of Acidobacterial genera was observed, with the exception of the genus Geothrix which increased in response to AgNP and AgMP amendment. Broad tolerance to Ag was observed among the Bacteriodetes, with higher relative abundance of most genera observed in the presence of AgNPs and AgMPs. The proteobacterial genus Dyella was highly tolerant to AgNPs and AgMPs and relative abundance of this genus increased with Ag concentration. Soil fungal community structure responded to both AgNPs and AgMPs, but the nanoparticle had an impact at a lower concentration. This study demonstrates that pastureland soil microbial communities are highly sensitive to AgNP amendment and key functional processes may be disrupted by relatively low levels of contamination.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Soil microbial communities play key roles in providing functional services in both natural and agricultural terrestrial ecosystems (Trivedi et al. 2016). The soil microbial community contributes greatly to soil organic matter formation (Kallenbach et al. 2016) and nutrient biogeochemical cycling processes (Ma et al. 2016) both of which are indispensable for maintaining healthy productive soils. Soil microbial biogeochemical processes are generally performed by a wide range of different species, many of which share common properties (Banerjee et al. 2016), although certain groups are known to contribute to a greater extent than others. Bacteria and archaea contribute to the degradation of organic matter, although fungi are regarded as the primary decomposers of leaf matter in terrestrial soils (Kuramae et al. 2016). Nevertheless, some key soil processes, such as nitrification, have lower levels of functional redundancy within the community structure and are undertaken by specific functional groups of microbes such as ammoni-oxidising bacteria (AOB) and archaea (AOA) (Banning et al. 2015). Anthropogenic disturbances of soil microbial community processes such as nitrogen cycling have been shown to compromise ecosystem functioning and subsequently result in reduced productivity within the soil system (Calderón et al. 2016).
The production and release of nanoparticle emissions into natural soil ecosystems is regarded as a growing environmental problem (Simonin and Richaume 2015). Anthropogenic AgNPs in particular are seen as one of the most potentially hazardous environmental NPs given their antimicrobial activity (León-Silva et al. 2016). A recent estimate of global AgNP production indicated that between 220 and 312 t were being produced annually (Seltenrich 2013), while the market for antimicrobial AgNPs has since been predicted to further grow from 0.79 to 2.54 billion $ (USD) from 2014 to 2022 (Pachapur et al. 2016). Although the potential extent and routes of exposure by which nanomaterials will come to pollute natural environments is currently poorly understood, NPs have been predicted to reach soil systems through a number of routes including the application of organic wastes such as sludge to soils (Dale et al. 2015). Forms of AgNPs entering environmental systems are predicted to be highly varied. AgNPs which reach soils through application of bio solids are predicted to undergo transformation under the anaerobic conditions into Ag+, AgCl and AgS-NPs prior to reaching soil systems (Hashimoto et al. 2017; Sekine et al. 2015). Direct application of AgNPs into soil systems has found AgNPs to largely persist in aerobic environments: however, in anaerobic soils, AgNPs undergo transformation into AgS-NPs (Hashimoto et al. 2017). The majority of AgNPs which are incubated in anaerobic conditions have been found to bind sulphur forming sulphide bridges transforming into AgS-NPs (Levard et al. 2011; Pradas del Real et al. 2016).
To date, several studies have shown AgNPs affect certain aspects of the soil microbiome. Hansch and Emmerling (2010) demonstrated that AgNP contamination of soil reduced the total microbial biomass over a 120-day period, although total soil respiration inversely increased over the period. Production of certain soil exo-enzymes linked to the soil microbial community was shown to be decreased by AgNP contamination by Shin et al. (2012). The ability of soil ectomycorrhizal fungi to colonise pine roots has been shown to be reduced in soil contaminated with AgNP concentrations ≥ 350 mg kg−1 (Sweet and Singleton 2015). Kumar et al. (2011, 2014) found that bacterial and fungal communities in arctic soil were altered in the presence of relatively high AgNP concentrations (660–66,000 mg kg−1), and furthermore, abundance of key nitrifying bacteria (Rhizobium) appeared to be slightly reduced. The study of Doolette et al. (2016) found AgNP and silver sulphide nanoparticles (Ag2S-NP) reduced soil nitrate production by 50% at concentrations of 42 and 619 mg kg−1, respectively, although AOB abundance was increased or remained unchanged up to these concentrations, while AOA were not assessed. Abundance of the AOA has been shown to be reduced to a greater extent than AOB in pastureland soils at an AgNP concentration of 50 mg kg−1, although the effect on nitrification rates was not assessed (McGee et al. 2017). The Doolette et al. (2016) study also indicated that up to 20% of soil bacterial OTUs may be affected at AgNP concentrations as low as 1.4 mg kg−1, though the composition of the bacterial community and the specific changes were not described.
The aim of the present study was to investigate the effect of relatively low concentrations, 1–50 mg kg−1, of AgNPs and AgMPs on enzyme activities, bacterial and fungal community structures and the abundance of the ammonia oxidation gene in both the bacteria and archaea in soil. Though the majority of AgNPs are now predicted to be transformed into AgS-NP prior to reaching soil systems, this study investigated the effects of pristine AgNPs. The toxic mechanism of both AgNP and AgS-NP is considered the release of Ag+ into the environment; thus, the effects on the susceptible soil microbial community can be considered to be somewhat similar. The concentration effects of AgNPs on soil systems have received little attention to date. Both AgNPs and AgMPs were investigated to provide a contrast between the effects of different-sized Ag particles, to determine if there were specific effects related to the nano-scale. Although there have been several studies exploring the effects of AgNPs on individual components of soil systems, these studies have often employed relatively high concentrations. In addition, these studies frequently do not assess the response of both soil activities and changing microbial community structures, particularly at contamination levels likely to be found in the environment. Crucially, there is scant knowledge of which particular bacterial taxa respond to AgNPs and/or the composition of the bacterial community post contamination. To date, the response of the AOA to AgNPs has also received little attention, and given their integral role in soil nitrification, this deserves further investigation. By undertaking this study, we set out to elucidate the response of several components of the soil microbial system by utilising a broad range of techniques. Molecular techniques such as of next-generation sequencing (NGS) on an Illumina MiSeq platform and automated ribosomal intergenic spacer analysis (ARISA) were utilised to profile the effects of AgNPs and AgMPs on soil bacterial and fungal community structures. The abundance of bacterial and archaeal amoA genes in soil was determined using quantitative PCR (qPCR). Soil dehydrogenase and urease enzyme activities were also used as a general indication of activity in soil.
Materials and methods
Silver particles
The AgNPs and AgMPs investigated in this study were obtained from commercial suppliers and were chosen based on the physicochemical properties stated by the manufacturers. A spherical silver nanoparticle (AgNP) preparation (stock# 45405) was obtained from Sky Spring Nanomaterials Inc. (Houston, Texas, USA) which had a specified average particle size (APS) of 20–30 nm. A silver micron particle (AgMP) preparation (stock# 41597) was obtained from Alfa Aesar (Hard Hill, Massachusetts, USA) with a specified particle size range of between 0.5 and 1 μm.
Physicochemical characterisation of particles
Physicochemical characterisation of the particles was performed for zeta potential, size and shape using the protocols detailed in McGee et al. (2017). Briefly, particle shape and size were determined using a Tecnai G2 20 TWIN electron microscope (FEI Company, Oregon, USA) through imaging using an accelerating voltage of 200 kV on air-dried particle suspensions (1 mg ml−1) on Formvar carbon-coated 200-mesh copper grids (Electron Microscopy Sciences, Hatfield, PA, USA).
Particles were characterised for size and zeta potential using a Zetasizer (Malvern, Worcestershire, UK) at pH levels of 4, 5, 6, 7 and 8.
Microcosm set-up
Soil used in the microcosm was harvested at a depth of 5–15 cm from an Irish pastureland, and details of sampling, soil analysis and basic microcosm set up are detailed in McGee et al. (2017). Briefly, AgNPs and AgMPs suspensions were prepared in 0.1 M NaCl and added to 50 g of fresh weight soil to give final concentrations of 1, 5, 10, 25 and 50 mg kg−1 of fresh weight homogenised soil. The final concentration of AgNPs/AgMPs in soil was based on the fresh weight of soil prior to amendment with the particle suspensions. Controls were amended with 0.1 M NaCl suspensions. Microcosm pots were incubated in the dark at 22 °C for a period of 30 days during which time soil moisture content was maintained as described previously. Three replicate pots were destructively sampled for each treatment on days 0, 3, 6, 10, 20 and soil enzyme levels measured immediately. Five grams of soil from each replicate was also stored at − 20 °C for subsequent molecular analysis.
Soil dehydrogenase and urease activities
Soil dehydrogenase and urease activities were determined using modified methods of Thalmann (1968) and Kandeler and Gerber (1988), respectively, as described in McGee et al. (2017).
Nucleic acid extraction and purification
DNA was extracted using the method developed by Griffiths et al. (2000), and then purified in a two-step process using ethanol precipitation and a High Pure PCR Product Purification Kit (Roche Cat 11732676001). The concentration and quality of the DNA extracts were determined using spectrophotometry (ND-1000 Spectrophotometer, NanoDrop, USA).
High-throughput sequencing of 16S gene amplicons
The bacterial community was characterised on day 30 of the microcosm experiment using MiSeq sequencing. PCR and preparation of DNA amplicons for next-generation sequencing of the 16S rRNA gene was performed using a modified protocol of Kozich et al. (2013) as described in McGee et al. (2017). Quality control and sequencing was carried out by the Centre for Genomic Research, University of Liverpool, UK, using an Illumina MiSeq system.
ARISA profiling of soil fungal community structure
Fungal community structures were examined using automated ribosomal intergenic spacer analysis (ARISA) as reported in McGee et al. (2017). ITS amplicons lengths and abundances were determined using an ABI3130xl genetic analyser (Applied Biosystems, CA, USA) on sampling days 0, 6 and 30.
Quantitative analysis of bacterial and archaeal amoA gene copy number
Quantification of bacterial amoA and archaeal amoA gene copy numbers was carried out following a modified protocol described by Abell et al. (2010) as described in McGee et al. (2017). Reactions were performed using an ABI ViiATM real time PCR system.
Data analysis
The data generated from MiSeq sequencing was processed using Mothur (Schloss et al. 2009) and datasets subsequently analysed using PRIMER version 6.1.9 with the PERMANOVA add-on version 1.0.1 (Primer-E Ltd., UK). After merging the paired-end reads, sequence lengths > 275 bp were removed in a quality control filtering step prior to clustering. A 97% threshold was used in clustering against a reference alignment which had a set homoploymer length of 8. To reduce the noise in the sequences, pre-clustering allowed up to 2 differences between sequences. The resulting sequences were screened for chimeras using VSEARCH. A Bayesian classifier was then used to classify those sequences against the Ribosomal Database Project 16S rRNA gene training set, and archaeal, eukaryotic, mitochondria and chloroplast 16S fragments were removed.
ARISA datasets were transformed to the fourth root and resemblance matrices were constructed using the Bray-Curtis dissimilarity measure in the Primer software. The Kruskal fit scheme 1 was used to rank dissimilarities to construct nMDS plots. Comparisons between factors and terms were performed with PERMANOVA main tests and pair-wise comparisons. Where the number of unique permutations was not sufficient to give meaningful values, Monte-Carlo simulations were used. One-way analysis of variance (ANOVA) and post-hoc pdiff tests were performed in Microsoft Excel 2010 for examining enzyme activities.
Results
Particle and soil physicochemical characteristics
Physicochemical characterisation of the AgNP has been previously reported (McGee et al. 2017). Briefly, TEM imaging found the AgNPs to be spherical in shape, while DLS found average particle sizes (APS) of the AgNPs to be 19 ± 1 nm. The AgMP sizes were initially characterised using DLS and TEM to confirm that they were MPs. The measured APS of the AgMP particles (829 ± 61 nm) was within the range specified by the manufacturer. Based on TEM analysis, AgMP shapes were mostly spherical and met the assumptions of DLS. The AgMPs were found to have negative surface charges between pH 5 and pH 6.
The soil had a pH of 5.15 and a moisture content of 28.8%. It had an organic matter content of 7.5% and N, P and K levels of 0.5, 0.1 and 0.4% (on a dry weight basis), respectively.
Effect of particles on soil dehydrogenase and urease activities
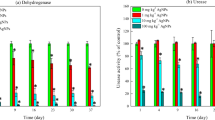
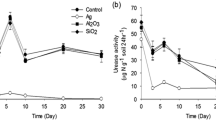
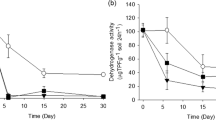
Broadly, ANOVA indicated that AgNP and AgMP concentrations of 1–50 mg kg−1 significantly (P ≤ 0.0001) reduced soil dehydrogenase activity from day 3–30 of the experiment (Fig. 1). However, similar concentrations of AgNPs reduced soil dehydrogenase activity to a greater extent than AgMPs. Dehydrogenase activity declined significantly (P ≤ 0.0001) immediately after soil was amended with AgNPs at concentrations ≥ 25 mg kg−1 and AgMPs at 50 mg kg−1 compared to unamended control soil. By day 20–30, dehydrogenase activity was barely detectable in soil amended with ≥ 25 mg kg−1 of AgNP and 50 mg kg−1 AgMP. Dehydrogenase activity increased between day 0–10 in the controls before decreasing towards the termination of the experiment, this is often a property of microcosm experiments after soil has been aerated after mixing.
Enzyme activities in AgNP- and AgMP-amended soils and unamended soils. Dehydrogenase activity in unamended soil (○) and soils amended with 1(●), 5(▼), 10(▲), 25(■) and 50(♦) mg kg−1 of AgNPs (a) and AgMPs (b). Urease activity in unamended soil (○) and soils amended with 10(▲), 25(■) and 50(♦) mg kg−1 of AgNPs (c) and AgMPs (d). Microcosm pots were sampled destructively on days 0, 3, 6, 10, 20 and 30 and activity was measured. Each marker is a mean of three measurements with error bars depicting SD
Urease activity in soil amended with ≥ 10 mg kg−1 AgNP was significantly (P ≤ 0.05) lower than control soil on days 3, 6 and 10. However, after day 20, no significant reductions in activity were detected, though urease activity was found to significantly increase compared to the control on day 30 (P ≤ 0.01). In contrast, only 50 mg kg−1 of AgMP was found to significantly reduce soil urease activity between days 6 and 10 (P ≤ 0.05) with no other significant reductions observed for lesser concentrations of AgMP. Soil urease activity in AgNP-and AgMP-amended soils recovered by day 30. Urease activity in the controls and treatments was exceptionally high on day 0, most likely a property of the soil being aerated after mixing.
Effect of Ag particles on soil bacterial community structure
Amending soil with AgNPs and AgMPs impacted the composition of the bacterial community. At phylum level, all concentrations of AgNPs tested reduced the relative abundance of the Acidobacteria and Verrucomicrobia compared to the unamended soil, while the relative abundance of the Proteobacteria and Bacteriodetes increased (Fig. 2a). The relative abundance of the Planctomycetes was only reduced by concentrations of 25 and 50 mg kg−1 AgNPs and no effect was observed at 10 mg kg−1. AgMPs only affected the relative abundance of similar bacterial phyla at 25 and 50 mg kg−1. Concentrations of 25 and 50 mg kg−1 AgMPs reduced the relative abundance of both the Acidobacteria and Verrucomicrobia, while increasing the relative abundance of the Proteobacteria and Bacteriodetes (Fig. 2b). The relative abundance of many phyla such as the Actinobacteria, Chloroflexi, Firmicutes and unclassified bacteria remained relatively stable in the presence of AgNPs and AgMPs.
Percentage contribution of bacterial phyla contributing > 1% of total sequences in unamended soil ( ) and soil amended with 10 (
) and soil amended with 10 ( ), 25(
), 25( ) and 50(
) and 50( ) mg kg−1 AgNP (a) and AgMP (b) on day 30. Each measurement represents the mean of 3 replicate samples and bars indicate SD. Significant differences between the relative abundance of phyla in unamended soil and AgNP- and AgMP-amended soils are indicated by the presence of A or B above the bar charts
) mg kg−1 AgNP (a) and AgMP (b) on day 30. Each measurement represents the mean of 3 replicate samples and bars indicate SD. Significant differences between the relative abundance of phyla in unamended soil and AgNP- and AgMP-amended soils are indicated by the presence of A or B above the bar charts
The unamended soil was dominated by three genera (Bacillus, Sporosarcina and unclassified 1) of Firmicutes whose sequences comprised 39.9% of total sequences detected (Table 1). No genera from the Firmicutes were affected by either AgNPs (Table 1) or AgMPs (Table 2), apart from one, the unclassified 6 whose relative abundance decreased in a concentration-dependent manner in AgNP-amended soil. Two genera of Proteobacteria, Dyella and Rhodanobacter increased in both AgNP- and AgMP-amended soil. The relative abundance of Dyella increased significantly (P ≤ 0.0001) from 0.2 to 5.3, 15.4 and 15.9% in soil amended with 10, 25 and 50 mg kg−1 AgNPs, respectively. In AgMP-amended soil, the relative abundance of Dyella increased slightly at 10 mg kg−1 from 0.2 to 0.6%, although at the higher concentrations of 25 and 50 mg kg−1, relative abundance increased significantly to 3.6 and 8.9% (P ≤ 0.0001). The relative abundance of Rhodanobacter was low in unamended soil (0.7%) but increased significantly (P ≤ 0.001) at every concentration of AgNP and AgMP amendment.
In the unamended control soil, the Acidobacterial community consisted of 17 genera which accounted for 15.4% of the total community. In response to 10 mg kg−1 AgNP amendment, relative abundance decreased significantly to 9.5% (P < 0.05), while concentrations of 25 and 50 mg kg−1 both caused a decrease to 5.6% (P < 0.01). The decrease in relative abundance of the phylum Acidobacteria in response to AgNP and AgMP amendment was reflected in a general decrease in most genera from this phylum, particularly Gp 1, 2 and 6. However, the genus Geothrix behaved differently, with increased relative abundance observed at 10–50 mg kg−1 concentrations of AgNPs and AgMPs. Interestingly, AgMPs did not affect the overall relative abundance of the Acidobacteria at 10 mg kg−1. However, this may reflect the fact that the relative abundance of Geothrix increased from 0.5% in the unamended soil to 5.4% in soil amended with 10 mg kg−1 AgMP. The relative abundance of most other Acidobacterial genera decreased slightly at this concentration of AgMPs. The decrease in the relative abundance of the phylum Verrucomicrobia in AgNP-amended soil at all concentrations tested could be attributed to significant decreases in the genus Spartobacteria (P < 0.05). Similarly, AgMPs induced significant decreases in Spartobacterial relative abundance at 25 and 50 mg kg−1 (P < 0.05).
Effect of particles on soil fungal community structure
The response of soil fungal community structures to the addition of AgNPs and AgMPs was examined using F-ARISA and found to differ in terms of concentration and time. n-MDS plots indicated that fungal community structures in unamended soil amended were significantly different from those in soils amended with concentrations of both 25 and 50 mg kg−1 AgNPs and AgMPs after 30 days of exposure (Fig. 3). PERMANOVA analysis, however, revealed that fungal communities in AgNP-amended soil were significantly different to those from unamended controls on day 30 at concentrations of 10, 25 and 50 mg kg−1 (P ≤ 0.05, 0.01 and 0.01, respectively) (Table 2). AgMP-amended communities were found to differ significantly from the unamended soil by day 30 at concentrations of 25 and 50 mg kg−1 (P ≤ 0.01).
nMDS plots of fungal community structures derived from ITS fragments in unamended soil and AgNP- and AgMP-amended soils. Plots a, b and c represent the response of the fungal community in the pastureland soil to 10 (triangles), 25 (squares) and 50 (diamonds) mg kg−1 Ag NP and MPs, respectively. Communities in unamended soil represented by ○, while those in soil amended with AgNPs are represented by closed symbols (▲, ■ and ♦) while AgMPs are represented by the corresponding open symbols (Δ, □ and ◊). Numbers beside symbol refer to sampling day
A total of 57 unique fungal ribotypes were detected in the pastureland soil. Simper analysis indicated that the ribotypes that were below the limit of detection (LOD) in unamended soils rose to detectable levels in AgNP-and AgMP-amended soil and this contributed to the differences in community structures. In unamended soil, the fungal community appeared to be dominated by 2 ribotypes (738 and 739 bp), with the relative abundance of most ribotypes below 1%. Addition of AgNPs and AgMPs caused a decline in the relative abundance of the two dominant ribotypes, and an increase in the relative abundance of many other ribotypes and detection of novel ribotypes above limits of detection (LODs).
Quantitative analysis of bacterial and archaeal amoA gene copy number
The ammonia-oxidising potential of control, AgNP- and AgMP-amended soils was assessed by quantifying the copy numbers of bacterial and archaeal amoA genes on day 30 of the microcosm experiment (Fig. 4). AgNPs and AgMPs significantly decreased archaeal amoA gene copy numbers at all concentrations tested (P ≤ 0.001). AgNPs also reduced bacterial amoA copy numbers significantly, but only at a concentration of 50 mg kg−1 (P < 0.05). Although copy numbers of bacterial amoA appeared lower in the presence of 10 and 25 mg kg−1 AgNPs, these differences were significant. AgMPs had no effect on bacterial amoA at any concentration tested.
Discussion
Currently, there is a lack of research data indicating the lower thresholds at which NPs can potentially disrupt the composition and functioning of terrestrial ecosystems. A limited number of dose-dependent studies have been conducted on the effects of AgNPs in soil systems, to date. In a previous study, we demonstrated that amending a pastureland soil with a concentration of 50 mg kg−1 AgNPs significantly affected microbial community structures and a key step in the nitrogen cycle to a greater extent than silica and aluminium oxide NP (McGee et al. 2017). However, it was unclear if lower concentrations would also have an impact soil microbial community structures and functional processes. To address this knowledge gap, we performed a concentration-dependent study to (1) confirm the findings of our previous study, (2) to determine if the effects of lower concentrations (dose-dependent effects) of NPs could be detected and (3) to compare and contrast the effects of AgNPs concentrations to similar levels of AgMPs.
Particle physicochemical characteristics
Characterisation of the AgMPs revealed that the particles corresponded to the manufacturer’s specifications and were mostly spherical. Similar to the AgNP, the AgMP was found to have a negative surface charge in the pH range of 5–6, values commonly found in soil. The distinctly differing size ranges but similar chemical properties of the AgNPs and AgMPs made these particles ideal for a comparative study.
Soil enzyme activities
A distinct contrast was observed between the effects of AgNPs and AgMPs on soil enzyme activities. AgNPs were found to cause more pronounced reductions of soil dehydrogenase and urease enzyme activities than similar concentrations of AgMPs. This was particularly evident in soil urease activity, where AgNP caused a much more marked reduction in activity than AgMP. However, soil urease activity displayed resilience and recovered prior to the termination of the experiment at all concentrations tested. In contrast to urease activity, dehydrogenase activity was found to be reduced by AgNP and AgMP in a nearly dose-dependent manner for the duration of the experiment across the concentrations range tested. These results indicate that low concentrations of AgNP contamination (≤ 1 mg kg−1) in soil can reduce the functional activity of the soil community, although concentrations of ≥ 5 mg of AgNPs were required to consistently reduce soil dehydrogenase activity significantly (P < 0.05) over the full duration of the microcosm experiment. These findings support the results of our previous study which found similar effects of 50 mg kg−1 of AgNP on soil dehydrogenase activity and were in accordance with the study conducted by Shin et al. (2012), which also found AgNPs reduced soil dehydrogenase and urease activities. However, in their soil system, Shin et al. (2012) found urease activity to be more susceptible to inhibition than dehydrogenase activity. The differences between these two studies could possibly be due to a range of biotic and abiotic factors influencing Ag contaminants, such as the differing composition of the soil microbial communities present or the physicochemical properties of the soils (Xian et al. 2015).
In contrast to enzymatic assessments of soil microbial community activities, a study conducted by Hansch and Emmerling (2010) measuring soil respiration rates indicated that amending soil with AgNPs could increase soil community level activity in a concentration-dependent manner. However, the study of Frenk et al. (2013) found that while amending soil with metal oxide NPs increased measured soil respiration, soil enzymatic processes were inversely reduced. To resolve this anomaly, their study investigated NP effects on sterilised soil and found that metal oxide NP interactions with the physical components of the soil were the cause of increased CO2 levels and that the perceived increases in soil respiration were not a result of biological processes (Frenk et al. 2013). Their work supports the findings that AgNPs negatively affect soil community activity and highlight that enzymatic assays may be a more reliable method for assaying the effects of NP on soil activity responses than measurements of soil respiration.
Effect of AgNPs and AgMPs on soil bacterial community
Amplicon sequencing of the bacterial community in the unamended soils revealed that the natural pastureland soil microbiome was composed predominantly of Acidobacteria, Actinobacteria, Firmicutes, Proteobacteria and Verrucomicrobia. The natural community composition was comparable to other studies on natural pastureland soils (Naethe et al. 2012; Will et al. 2010). AgNP and AgMP amendment of soil was not found to affect the relative abundance of the dominant bacterial phylum, the Firmicutes. Other changes detected in the composition of the microbial community in response to contamination with AgNPs were in accordance with other studies (Kumar et al. 2011, 2014; McGee et al. 2017). However, this study was unique in demonstrating dose-dependent effects of comparatively low AgNP concentrations (10 mg kg−1) on bacterial pastureland soil communities.
Regardless of the type of Ag particle investigated (NP or MP), a significant (P ≤ 0.05) increase in the abundance of the Bacteriodetes was observed at all concentrations tested. In the unamended soil, 3 genera of Bacteriodetes were detected, representing approximately 0.17% of the total community abundance. In Ag-amended soils, 17 unique Bacteriodetes genera were detected and accounted for between 1.53 and 2.01% of all bacterial sequences across all treatments (AgNPs and AgMPs). No distinguishing trends were determined between the effects of AgNPs and AgMPs on the Bacteriodetes. Our previous study (McGee et al. 2017) found the increase in Bacteriodetes to be largely due to the genus Ohtaekwangia; however, with the expanded dataset gained from this study, it appears that there may be a broad tolerance to Ag across the phylum.
In total, 111 unique genera of Proteobacteria were detected making this the most diverse phylum in this soil system. The relative abundance of the Proteobacteria was found to increase in response to Ag amendment, although no relationship was found between increasing Proteobacterial relative abundance and diversity. The increase in Proteobacterial relative abundance was rather driven by two genera of the family Xanthomonadaceae, Dyella and Rhodanobacter. However, in AgMP-amended soil, relative abundance of Dyella was found to increase to a much lesser extent than at similar concentrations AgNP. In contrast, relative abundance of Rhodanobacter increased to a greater extent in AgMP-amended soils compared to AgNP-amended soil. This is not the first study to report increases in Rhodanobacter in AgNP-contaminated environments. The study of Samarajeewa et al. (2017) isolated high levels of a Rhodanobacter strain from soil amended with between 49 and 287 mg kg−1 AgNPs. Our previous study found the increase in Proteobacterial abundance to be largely due to the genus Dyella. This study has confirmed that in this soil system, a representative of this genus appeared to have a substantial tolerance for Ag contamination. The genus Dyella was first isolated and characterised in 2005 (Xie and Yokota 2005) and still very little is known about its functional role in ecosystems. Recently, a strain of an unclassified Dyella species was found to be capable of causing neonatal bacteremia (Hakima et al. 2017), although cases of pathogenicity in humans appear to be rare. Studies on the 10 currently described species indicate potential roles in mineral weathering and a high capacity for degrading certain environmental pollutants (Bao et al. 2014; Kong et al. 2013). Characterisation of Dyella genomes has highlighted the presence of several copper resistance cus genes. Copper resistance mechanisms encoded by genes in the cus operon are known to also confer silver resistance (Behlau et al. 2011) and it is plausible that the increases in relative abundance of Dyella observed in this study may reflect their ability to resist the toxic effect of Ag through copper resistance mechanisms.
The genus Rhodanobacter is closely related to the genus Dyella (Kostka et al. 2012) and currently contains 13 recognised species, most of which have been isolated from soil (Madhaiyan et al. 2014). Denitrification potential has been shown to be sporadically present throughout several species (Kostka et al. 2012), while an isolate of R. xiangquanii has been shown to be capable of degrading the herbicide lindane indicating potential roles for bioremediation (Zhang et al. 2016). A study of Rhodanobacter genomes indicated that high levels of duplication and transfer of mercuric and Co2+/Zn2+/Cd2+ resistance operons take place in metal-contaminated environments within this genus (Hemme et al. 2016). Additionally, the study performed by Zhang et al. (2011) found the genus to be highly abundant in copper mine tailings indicating high levels of copper resistance. Though no studies have highlighted the presence of Ag resistance genes in Rhodanobacter species, reviewing the current literature indicates that this genus can tolerate 2+ charged metals which possibly explains its ability to tolerate Ag in this soil environment.
Decreases in relative abundance were observed in the Acidobacteria and Verrucomicrobia in response to both AgNP and AgMP amendment. The Verrucomicrobia are widely found in soil systems and based on 16S transcripts generally account for between 1 and 10% of the total bacterial community (Sangwan et al. 2004). Of the 13 genera of Verrucomicrobia detected in this soil system, the genus Spartobacteria was found to dominate accounting for 85.3% of the total Verrucomicrobial abundance. AgNP amendment of pastureland soil significantly decreased the relative abundance of Spartobacteria at all concentrations tested (P < 0.05), while AgMPs had an impact at 25 and 50 mg kg−1, indicating that the Spartobacteria are highly susceptible to Ag contamination. Metagenomic studies have shown that Spartobacteria is generally the most abundant Verrucomicrobial genus present in terrestrial and aquatic habitats (Bergmann et al. 2011). A study of the Spartobacteria by Herlemann et al. (2014) characterised a putative genome of an aquatic phylotype of Spartobacteria and found a diversity of glycoside hydrolases present, in addition to a positive association between abundances of Spartobacteria and cyanobacteria populations, indicating a role in the marine carbon cycle. However, currently little is known the ecological role of Spartobacteria in terrestrial habitats due to a lack in cultivatable and characterised isolates.
The Acidobacteria were similarly seen to decrease in relative abundance in response to AgNP and AgMP amendment. The decrease in Acidobacterial abundance was due to a broad decrease in most Acidobacterial genera, with the exception of Geothrix, the abundance of Geothrix increased in response to AgNP and AgMP amendment. Geothrix fermentans is currently the only classified species of the genus Geothrix, this species is the sole member of the Acidobacteria known to be capable of reducing iron (Mehta-Kolte and Bond 2012). The high level of redox capabilities associated with this species may explain its ability to tolerate Ag contamination and even increase in abundance. Our previous study which investigated 50 mg kg−1 of AgNP on this pastureland soil detected a slight increase in abundance of Geothrix from 0.4% in the unamended soil to 1% in the presence of AgNP but the increases detected at the lower concentrations in the current study were considerably greater, revealing the subtle response of this genus to even low concentrations of the particles. The phylum Acidobacteria is a relatively newly identified phylum the ecological role of which is still poorly understood owing to difficulties in cultivation, currently only 8 of the 26 subdivisions have representatives in culture (Kielak et al. 2016). However, current indications are that Acidobacteria may possibly live oligotrophic lifestyles and that certain members appear to play roles in the nitrogen cycle (Kielak et al. 2016).
Effect of AgNP and AgMP on soil fungal community structure
Similar to the bacteria, pastureland soil fungal community structures were affected by both AgNP and AgMP amendment. The detection of novel fungal ribotypes in Ag-amended soil appeared to be a driving factor in changing community structures. The study of MacDonald et al. (2008) also reported the appearance of novel fungal ribotypes when examining metal-contaminated soils. Although the response of soil fungal communities to NPs has received relatively less attention than bacterial communities, several studies have reported an effect of these particles on fungi. AgNP concentrations of ≥ 350 mg kg−1 were shown to inhibit or prevent ectomycorrhizal colonisation of pine roots (Sweet and Singleton 2015) and the sensitivity of arctic soil fungal communities to AgNP concentrations of 660 and 66,000 mg kg−1 was reported by Kumar et al. (2011, 2014). The current study indicates that soil fungal communities respond to much lower concentrations of AgNPs than has been previously reported and highlights the sensitivity of fungal communities to Ag contamination. However, further studies are still required to investigate the taxonomic changes in the fungal community and to determine if these changes result in negative effects on the functional roles that of the fungal communities perform in soil ecosystems.
Ammonia-oxidising functional groups
Ammonia oxidation is a key step in the soil nitrogen cycle which is performed by a functional group composed of ammonia-oxidising archaea (AOA) and bacteria (AOB). These functional organisms have been shown in in vitro studies to be sensitive to AgNPs (Shahrokh et al. 2014). In accordance with the in vitro studies, our current investigation found AOB to be sensitive to Ag contamination with the abundance of the amoA gene copies decreasing in response to AgNP contamination ≥ 10 mg kg−1, albeit only significantly at 50 mg kg−1 (P < 0.05). Similar concentrations of AgMPs did not have an effect on AOB gene copy numbers. A study by Doolette et al. (2016) found a disparity between AOB gene copy number and soil nitrification in response to Ag contamination. Their study indicated that AOB gene copy numbers either increased or remained static at AgNP concentrations up to 40–50 mg kg−1, while soil nitrification was inhibited by AgNP concentrations ≥ 10 mg kg−1. This anomaly could possibly be explained by the contribution of the AOA which were not quantified. In both the current and our previous study, AOA were found to be more sensitive to AgNP contamination than AOB. AgNPs and AgMPs significantly decreased copy numbers of the AOA-associated gene at all concentrations tested (P < 0.001), with relatively little difference observed between similar levels of AgNPs and AgMPs. This indicates that AOA are highly sensitive to low levels of Ag contamination. While copy numbers of the AOA gene were lower than those of the AOB in this soil system, AOA have been found to be generally higher in abundance than AOB across a broad range of soil systems and possibly contribute more to soil nitrification (Leininger et al. 2006; Zeglin et al. 2011).
Conclusions
Soil microbial communities were found to be highly sensitive to AgNP and AgMP contamination, but the effect of AgNPs and AgMPs differed as concentration decreased. AgNP and AgMPs effects on soil dehydrogenase activity were found to be dose-dependent across the concentration range tested in this study. Relatively low concentrations (1 mg kg−1) of AgNPs significantly reduced biological activity in soil and 10 mg kg−1 impacted both soil bacterial and fungal community structures to a greater degree than that observed with a similar concentration of AgMPs. In addition, ammonia-oxidising microbes, in particular the archaea appeared to highly sensitive to AgNP amendment. Given the importance of bacteria, fungi and archaea in nutrient cycling in soil, these results highlight the potential compromising effects AgNPs may have on natural ecosystems through disruption of key biogeochemical processes. In order to fully understand the potential impacts of AgNPs, further studies exploring AgNPs and the potential transformed products of these NPs are required to determine how these substances may negatively affect microbial functional groups and biological activity in natural ecosystems.
References
Abell, GCJ, Revill, AT, Smith, C, Bissett, AP, Volkman, JK and Robert, SS (2010). Archaeal ammonia oxidizers and nirS-type denitrifiers dominate sediment nitrifying and denitrifying populations in a subtropical macrotidal estuary. The ISME Journal 4:286–300
Banerjee S, Kirkby CA, Schmutter D, Bissett A, Kirkegaard JA, Richardson AE (2016) Network analysis reveals functional redundancy and keystone taxa amongst bacterial and fungal communities during organic matter decomposition in an arable soil. Soil Biol Biochem 97:188–198
Banning NC, Maccarone LD, Fisk LM, Murphy DV (2015) Ammonia-oxidising bacteria not archaea dominate nitrification activity in semi-arid agricultural soil. Sci Rep 5:11146
Bao Y, Kwok AHY, He L, Jiang J, Huang Z, Leung FC, Sheng X (2014) Complete genome sequence of Dyella jiangningensis strain SBZ3-12, isolated from the surfaces of weathered rock. Genome Announc 2(3):e00416–e00414
Behlau F, Canteros BI, Minsavage GV, Jones JB, Graham JH (2011) Molecular characterization of copper resistance genes from Xanthomonas citri subsp. citri and Xanthomonas alfalfae subsp. Citrumelonis. Appl Environ Microb 77(12):4089–4096
Bergmann GT, Bates ST, Eilers KG, Lauber CL, Caporaso JG, Walters WA, Knight R, Fierer N (2011) The under-recognized dominance of Verrucomicrobia in soil bacterial communities. Soil Biol Biochem 43:1450–1455
Calderón K, Spor A, Breuil MC, Bru D, Bizouard F, Violle C, Barnard RL, Philippot L (2016) Effectiveness of ecological rescue for altered soil microbial communities and functions. ISME J 11:272–283
Dale AL, Casman EA, Lowry GV, Lead JR, Viparelli E, Baalousha M (2015) Modeling nanomaterial environmental fate in aquatic systems. Environ Sci Technol 49:2587−2593
Doolette CL, Gupta VVSR, Lu Y, Payne JL, Batstone DJ, Kirby JK, Navarro DA, Mclaughlin M (2016) Quantifying the sensitivity of soil microbial communities to silver sulfide nanoparticles using metagenome sequencing. PLoS One 11(8):e0161979
Frenk S, Ben-Moshe T, Dror I, Berkowitz B, Minz D (2013) Effect of metal oxide nanoparticles on microbial community structure and function in two different soil types. PLoS One 8(12):e84441
Griffiths RI, Whiteley AS, O'Donnell AG, Bailey MJ (2000) Rapid method for coextraction of DNA and RNA from natural environments for analysis of ribosomal DNA- and rRNA-based microbial community composition. Appl Environ Microbiol 66:5488–5491
Hakima N, Bidet P, Lopez M, Rioualen S, Carol A, Bonacorsi S (2017) First case of neonatal bacteremia due to Dyella genus. Diagn Microbiol Infect Dis 87:199–201
Hansch M, Emmerling C (2010) Effects of nanoparticles on the microbiota and enzyme activity in soil. J Plant Nutr 1:1–5
Hashimoto Y, Takeuchi S, Mitsunobu S, Ok Y-S (2017) Chemical speciation of silver (Ag) in soils under aerobic and anaerobic conditions: Ag nanoparticles vs. ionic Ag. J Hazard Mater 322:318–324
Hemme CL, Green SJ, Rishishwar L, Prakash O, Pettenato A, Chakraborty R, Deutschbauer AM, Van Nostrand JD, Wu L, He Z, Jordan IK, Hazen TC, Arkin AP, Kostka JE, Zhou J (2016) Lateral gene transfer in a heavy metal-contaminated-groundwater microbial community. MBio 7(2):e02234–e02215
Herlemann DPR, Lundin D, Labrenz M, Jürgens K, Zheng Z, Aspeborg H, Andersson AF (2014) Metagenomic de novo assembly of an aquatic representative of the verrucomicrobial class Spartobacteria. MBio 4(3):e00569–e00512
Kallenbach CM, Fre SD, Grandy SA (2016) Direct evidence for microbial-derived soil organic matter formation and its ecophysiological controls. Nat Commun 7:13630
Kandeler E, Gerber H (1988) Short-term assay of soil urease activity using colorimetric determination of ammonium. Biol Fert Soils 6(1):68–72
Kielak AM, Barreto CC, Kowalchuk GA, van Veen JA, Kuramae EE (2016) The ecology of Acidobacteria: moving beyond genes and genomes. Front Microbiol 7:744
Kong C, Wang L, Li P, Qu Y, Tang H, Wang J, Zhou H, Ma Q, Zhou J, Xu P (2013) Genome sequence of Dyella ginsengisoli strain LA-4, an efficient degrader of aromatic compounds. Genome Announc 1(6):e00961–e00913
Kostka JE, Green SJ, Rishishwar L, Prakash O, Katz LS, Mariño-Ramírez L, Jordan IK, Munk C, Ivanova N, Mikhailova N, Watson DB, Brown SD, Palumbo AV, Brooks SC (2012) Genome sequences for six Rhodanobacter strains, isolated from soils and the terrestrial subsurface, with variable denitrification capabilities. J Bacteriol 194(16):4461–4462
Kozich JJ, Westcott SL, Baxter NT, Highlander SK, Schloss PD (2013) Development of a dualindex sequencing strategy and curation pipeline for analyzing amplicon sequence data on the MiSeq Illumina sequencing platform. Appl Environ Microbiol 79(17):5112–5120
Kumar N, Shah V, Walker VK (2011) Perturbation of an arctic soil microbial community by metal nanoparticles. J Hazard Mater 190:816–822
Kumar N, Palmer GR, Shah V, Walker VK (2014) The effect of silver nanoparticles on seasonal change in arctic tundra bacterial and fungal assemblages. PLoS One 9(6):e99953
Kuramae EE, Hillekens RHE, de Hollander M, van der Heijden MGA, van den Berg M, van Straalen NM, Kowalchuk GA (2016) Structural and functional variation in soil fungal communities associated with litter bags containing maize leaf. FEMS Microbiol Ecol 84:519–531
Leininger S, Urich T, Schloter M, Schwark L, Qi J, Nicol GW, Prosser JI, Schuster SC, Schleper C (2006) Archaea predominate among ammonia-oxidizing prokaryotes in soils. Nat Lett 442:806–809
León-Silva S, Fernández-Luqueño F, López-Valdez F (2016) Silver nanoparticles (AgNP) in the environment: a review of potential risks on human and environmental health. Water Air Soil Pollut 227:306
Levard C, Reinsch BC, Michel FM, Oumahi C, Lowry GV, Brown GE Jr (2011) Sulfidation processes of PVP-coated silver nanoparticles in aqueous solution: impact on dissolution rate. Environ Sci Technol 45:5260–5266
Ma W, Jiang S, Assemien F, Qin M, Ma B, Xie Z, Liu Y, Feng H, Du G, Ma X, Le Roux X (2016) Response of microbial functional groups involved in soil N cycle to N, P and NP fertilization in Tibetan alpine meadows. Soil Biol Biochem 101:195–206
MacDonald CA, Campbell CD, Bacon JR, Singh BK (2008) Multiple profiling of soil microbial communities identities potential genetic markers of metal-enriched sewage sludge. FEMS Microbiol Ecol 65:555–564
Madhaiyan M, Poonguzhali S, Saravanan VS, Kwon SW (2014) Rhodanobacter glycinis sp. nov., a yellow pigmented gammaproteobacterium isolated from the rhizoplane of field-grown soybean. Int J Sys Evol Microbiol 64:2023–2028
McGee CF, Storey S, Clipson N, Doyle E (2017) Soil microbial community responses to contamination with silver, aluminium oxide and silicon dioxide nanoparticles. Ecotoxicology 26(3):449–458
Mehta-Kolte MG, Bond DR (2012) Geothrix fermentans secretes two different redox-active compounds to utilize electron acceptors across a wide range of redox potentials. Appl Environ Microbiol 78(19):6987–6995
Naethe A, Foesel BU, Naegele V, Wüst PK, Weinert J, Bonkowski M, Alt F, Oelmann Y, Polle A, Lohaus G, Gockel S, Hemp A, Kalko EKV, Linsenmair KE, Pfeiffer S, Renner S, Schöning I, Weisser WW, Wells K, Fischer M, Overmann J, Friedrich MW (2012) Environmental factors affect acidobacterial communities below the subgroup level in grassland and forest soils. Appl Environ Microbiol 78(20):7398–7406
Pachapur VL, Larios AD, Cledón M, Brar SK, Verma M, Surampalli RY (2016) Behavior and characterization of titanium dioxide and silver nanoparticles in soils. Sci Total Environ 563–564:933–943
Pradas del Real AE, Castillo-Michel H, Kaegi R, Sinnet B, Magnin V, Findling N, Villanova J, Carriere M, Santaella C, Fernandez-Martinez A, Levard C, Sarret G (2016) Fate of Ag-NPs in sewage sludge after application on agricultural soils. Environ Sci Technol 50:1759–1768
Samarajeewa AD, Velicogna JR, Princz JI, Subasinghe RM, Scroggins RP, Beaudette LA (2017) Effect of silver nano-particles on soil microbial growth, activity and community diversity in a sandy loam soil. Environ Pollut 220(Pt A):504–513
Sangwan P, Chen X, Hugenholtz P, Janssen PH (2004) Chthoniobacter flavus gen. nov., sp. nov., the first pure-culture representative of subdivision two, Spartobacteria classis nov., of the phylum Verrucomicrobia. Appl Environ Microbiol 70(10):5875–5881
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB, Lesniewski RA, Oakley BB, Parks DH, Robinson CJ, Sahl JW, Stres B, Thallinger GG, van Horn DJ, Weber CF (2009) Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75(23):7537–7541
Sekine R, Brunetti G, Donner E, Khaksar M, Vasilev K, Jämting ÅK, Scheckel KG, Kappen P, Zhang H, Lombi E (2015) Speciation and lability of Ag-, AgCl-, and Ag2S-nanoparticles in soil determined by X-ray absorption spectroscopy and diffusive gradients in thin films. Environ Sci Technol 49:897–905
Seltenrich N (2013) Nanosilver: weighing the risks and benefits. Environ Health Perspect 121:221–225
Shahrokh S, Hosseinkhani B, Emtiazi G (2014) The impact of silver nanoparticles on bacterial aerobic nitrate reduction process. J Biopro Biotechnol 4(3):152
Shin YJ, Kwak JI, An YJ (2012) Evidence for the inhibitory effects of silver nanoparticles on the activities of soil exoenzymes. Chemosphere 88(4):524–529
Simonin M, Richaume A (2015) Impact of engineered nanoparticles on the activity, abundance, and diversity of soil microbial communities: a review. Environ Sci Pollut R 22(18):3710–13723
Sweet MJ, Singleton I (2015) Soil contamination with silver nanoparticles reduces Bishop pine growth and ectomycorrhizal diversity on pine roots. J Nanopart Res 17:448
Thalmann A (1968) Zur methodik der bestimmung der dehydrogenaseaktivität im boden mittles triphenyltetrazoliumchlorid (TTC). Landwitsch Forch 21:249–258
Trivedi P, Delgado-Baquerizo M, Anderson IC, Singh BK (2016) Response of soil properties and microbial communities to agriculture: implications for primary productivity and soil health indicators. Front Plant Sci 7:990
Will C, Thurmer A, Wollherr A, Nacke H, Herold N, Schrumpf M, Gutknecht J, Wubet T, Buscot F, Daniel R (2010) Horizon-specific bacterial community composition of German grassland soils, as revealed by pyrosequencing-based analysis of 16S rRNA genes. Appl Environ Microbiol 76(20):6751–6759
Xian Y, Wang M, Chen W (2015) Quantitative assessment on soil enzyme activities of heavy metal contaminated soils with various soil properties. Chemosphere 139:604–608
Xie CH, Yokota A (2005) Dyella japonica gen. nov., sp. nov., a γ-proteobacterium isolated from soil. Inter J Sys Evol Microbiol 55:753–756
Zeglin LH, Taylor AE, Myrold DD, Bottomley PJ (2011) Bacterial and archaeal amoA gene distribution covaries with soil nitrification properties across a range of land uses. Environ Microbiol Rep 3(6):717–726
Zhang J, Zheng JW, Hang BJ, Ni YY, He J, Li SP (2011) Rhodanobacter xiangquanii sp. nov., a novel anilofos-degrading bacterium isolated from a wastewater treating system. Current Microbiology 62:645–649
Zhang X, Niu J, Liang Y, Liu X, Yin H (2016) Metagenome-scale analysis yields insights into the structure and function of microbial communities in a copper bioleaching heap. BMC Genet 17:21
Acknowledgments
The authors would also like to thank Maria Benson, Alexandre De Menezes, Ciara Murphy, Kate Randall and Bas Boots for their valuable technical assistance during the study.
Funding
This work was funded by the Irish Research Council (IRC) as part of a Graduate Research Education Program (GREP) structured Ph.D. program in Sustainable Development.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Additional information
Responsible editor: Robert Duran
Electronic supplementary material
ESM 1
(DOC 29 kb)
Rights and permissions
About this article
Cite this article
McGee, C.F., Storey, S., Clipson, N. et al. Concentration-dependent responses of soil bacterial, fungal and nitrifying communities to silver nano and micron particles. Environ Sci Pollut Res 25, 18693–18704 (2018). https://doi.org/10.1007/s11356-018-2087-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-018-2087-y







 ) and soil amended with 10 (
) and soil amended with 10 ( ), 25(
), 25( ) and 50(
) and 50( ) mg kg−1 of AgNP and AgMP on day 30. Each marker is a mean of three measurements with error bars depicting SD
) mg kg−1 of AgNP and AgMP on day 30. Each marker is a mean of three measurements with error bars depicting SD