Abstract
Agriculture accounts for ~ 70% of all water use and the world population is increasing annually; soon more people will need to be fed, while also using less water. The use of plant-associated bacteria (PAB) is an eco-friendly alternative that can increase crop water use efficiency. This work aimed to study the effect of some PAB on increasing soybean tolerance to drought stress, the mechanisms of the drought tolerance process, and the effect of the PAB on promoting plant growth and on the biocontrol of Sclerotinia sclerotiorum. PAB were isolated from soybean rhizosphere and S. sclerotiorum sclerotia. The strains identified as UFGS1 (Bacillus subtilis), UFGS2 (Bacillus thuringiensis), UFGRB2 and UFGRB3 (Bacillus cereus) were selected on their ability to grow in media with reduced water activity. Soybean plants were inoculated with the PAB and evaluated for growth promotion, physiological and molecular parameters, after drought stress. Under drought stress, UFGS2 and UFGRB2 sustained potential quantum efficiency of PSII (Fv/Fm), while a decrease was found in the control plants. Moreover, UFGS2 and UFGRB3 maintained the photosynthetic rates in non-stressed conditions compared to the control. UFGS2-treated plants showed a higher stomatal conductance and higher transpiration than the control, after drought stress. Some PAB-treated plants also had other beneficial phenotypes, such as increases in fresh and dried biomass relative to the control. Differential gene expression analysis of genes involved in plant stress pathways shows changes in expression in PAB-treated plants. Results from this study suggest that PAB can mitigate drought stress in soybean and may improve water efficiency under certain conditions.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Cultivable land and productivity have been globally reduced as a result of industrialization and abiotic stresses (Gamalero et al. 2009). Drought, for instance, plays a major role in reducing plant growth parameters and also negatively interferes with plants’ physiology by causing losses in CO2 accumulation and then reducing photosynthetic rate or the photochemical efficiency (Muller et al. 2011; Sadeghipour and Abbasi 2012).
To resist drought-stress effects, plants use different mechanisms, which are generally regulated by genes, such as ethylene response factors (Gmereb), dehydration-responsive binding protein (Gmdreb1a), galactinol synthase (Gmgols), and Δ1-pyrroline-5-carboxylase synthetase (Gmp5cs). These genes have been reported to be the main drought-associated genes that are directly or indirectly related to the plant’s response to water stress tolerance (Chen et al. 2016; Shinozaki and Yamaguchi-Shinozaki 2007; Stolf-Moreira et al. 2010).
In some cases, in addition to abiotic stresses, plants from their early stages have to cope with biotic stresses, such as plant pathogens emerging from over-wintering structures, among which is the sclerotium produced by Sclerotinia sclerotiorum (Lib.) de Bary. S. sclerotiorum is an ubiquitous causal agent of disease in more than 500 plant species from 75 families, including soybean (Glycine max (L.) Merr.), one of the most important crops in the world, on which S. sclerotiorum causes white mold (Kamal et al. 2016). S. sclerotiorum sclerotium is a hard dark structure comprised of a mass of hyphal threads, capable of remaining dormant without host interaction for years (Geraldine et al. 2013).To make matters worse, with the exception of the highly toxic fumigant chemicals, no fungicide is effective in preventing these sclerotia from germinating and then infecting their hosts (Hu et al. 2014).
On the other hand, plant-associated bacteria (PAB) are among the environmentally friendly tools which can be used as biocontrol agents against plant pathogens, and some strains may offer additional benefits to the plants, such as making them more tolerant to abiotic stresses and promoting growth (Geraldine et al. 2013; Martins et al. 2015a, 2014). Among PAB, endospore-forming species, such as those belonging to the Bacillus genus, have the advantage relative to other species do to their ability to survive under harsh environments (Timmusk et al. 2013).
In nature, PAB can be found aboveground in the phyllosphere on tissues, such as buds, flowers and leaves, and also within plant tissues (endophytes). However, PAB are most often found belowground in the rhizosphere (Berendsen et al. 2012). The bacterial cells measured colonizing each gram of the root is around 109–1011, which often not only surpasses the number of plant host cells but also the number of people existing on Earth (Berg et al. 2016).
In addition to being found in high populations in plants, beneficial microbials can also be naturally present on pathogen survival structures (e.g., sclerotia), in which case they are named hyperparasites (Heydari and Pessarakli 2010; Martins et al. 2015b). Researchers have been screening hyperparasites in an attempt to isolate microbials with a higher chance of succeeding against a specific pathogen. Here, we hypothesized that bacteria isolated from soybean rhizosphere and from S. sclerotiorum sclerotium and screened under low water activity could increase soybean tolerance to drought stress in addition to inhibiting the sclerotium germination.
The aims of this study were to screen the PAB under low water activity medium for potential bacterium resistant to drought stress; to evaluate the contribution of the selected PAB to increasing soybean tolerance to drought stress and to study the evolved mechanism(s), to test the PAB as a plant growth promoter, and to evaluate the antagonistic activity of the PAB against S. sclerotiorum sclerotium.
Materials and methods
Plant-associated bacteria isolation
The PAB were isolated from Sclerotinia sclerotiorum sclerotia as well as from rhizosphere of asymptomatic soybean plants in an agricultural area located in the Federal University of Goiás (UFG), in Goiânia, Goiás, Brazil (altitude 716 m; 16° 35′ 39″ S; 49° 17′ 16″ W). The soil characterized as Rhodic Hapludox and presented the following characteristics: pH(H2O): 4.9, sum of bases (S value) 4.2 cmolc dm, organic matter 18.0 g kg−1, and clay content 470 g kg−1.
To isolate the bacteria, six sclerotia (~ 100 mg), which were collected from the field described above, were immersed in tubes containing 10 mL of sterile distilled water, vortexed three times for 5 s each time and serially diluted until 10−4. A volume of 100 μL of each dilution was spread-plated on nutrient agar (NA) medium (3.0 g L−1 meat extract, 3.5 g L−1 meat peptone, 5.0 g L−1 NaCl, 20.0 g L−1 agar) (n = 3) (Martins et al. 2013). Similarly, approximately 100 mg of soybean root samples were immersed in tubes containing 10 mL of sterile distilled water, vortexed three times for 5 s each time and serially diluted until 10−4. A volume of 100 μL of each dilution was spread-plated on NA medium. Plates from each dilution were incubated at 28 °C and analyzed for the colonies after 24 h. Bacteria were preserved in peptone glycerol at − 80 °C for long-term storage.
Screening test
To test the bacterial growth under low water activity, the isolated bacteria with four replicates each were grown in 5-mL Eppendorf tubes filled with NA-broth medium altered with 2.5% (w/v) of glycerol and arranged in a completely randomized design. As a control, the same ten bacteria were grown in NA-broth medium. The bacteria were cultivated for 24 h on a shaker at 150 rpm at room temperature (28 °C) and then measured for their growth by checking the optical density at 620 nm for each tube using a spectrophotometer.
Bacterial DNA extraction and identification
The bacteria were selected based on the previous test and cultivated as described previously. Then, 1.5 mL of bacterial suspension of each sample was collected and transferred to 2-mL tubes. The samples were centrifuged at 10000 rpm for 8 min at 25 °C. The supernatant was discarded and the pellet washed with autoclaved milli-Q water followed by one more centrifugation step. The supernatant was discarded and the pellet was used for DNA extraction. The DNA extraction was performed by a modified CTAB method (Boiteux et al. 1999). In each sample, 600 μL of extraction buffer (0.1 M EDTA; 0.1 M Tris pH 8.0; 1.4 M NaCl; 2% CTAB; 0.2% β-mercaptoethanol) were added and incubated at 65 °C for 10 min. A volume of 600 μL of chloroform: isoamyl alcohol (24:1) was added and vortexed for 30 s, followed by centrifugation for 5 min at 12000 rpm. The supernatant was transferred into new tubes and 0.6× of the isopropanol volume was added and incubated for 10 min at room temperature (28 °C). The samples were centrifuged for 13 min at 12000 rpm to form the pellet. The supernatant was carefully discarded and the pellet washed with 300 μL of 70% ethanol (− 20 °C). The samples were centrifuged at 12000 rpm for 5 min. The supernatant was discarded and the pellet dried at room temperature (28 °C). The pellet was resuspended in 30 μL of milli-Q water, and the DNA was quantified in a NanoDrop Lite spectrophotometer (Thermo Scientific).
For the bacterial identification, 16S PCR was done with primers 27F (5′-AGAGTTTGATCCTGGCTCAG-3′) and 1492R (5′-GGTTACCTTGTTACGACTT-3′) using amplitaq gold PCR master mix under the following condition: initial denaturation 95 °C for 10 min, followed by 35 cycles of 95 °C for 30 s, 55 °C for 30 s, and 72 °C for 1 min, and a final extension at 72 °C for 7 min. The resultant amplicon was prepared for sequencing using a Nextera XT library preparation kit following the manufacturer’s suggested protocols. The prepared libraries were sequenced using a MiSeq DNA sequencer with the MiSeq V3 2 × 300 sequencing kit. The resulting reads were quality trimmed to the Q30 confidence level. The draft genome was assembled using CLCbio Genomics Workbench 10.1 (Qiagen Inc., Cambridge, MA) using default parameters. The sequences were deposited in NCBI Genbank under accession numbers MF526966–MF526969. The species identifications were determined using EzBioCloud 16S database (Yoon et al. 2017). Phylogenetic analysis was performed in MEGA 7 using the Maximum Likelihood method with a HKY85 model and a nearest-neighbor interchange heuristic search method was used. The level of bootstrap support was calculated from 1500 replicates.
Biological seed treatment with the selected PAB
The in vivo tests were carried out at the Federal University of Goiás (UFG), in Goiânia, Goiás, Brazil, at the greenhouse (altitude 749 m; 16° 36′ 12″ S; 49° 15′ 35″ W) under the following conditions: maximum temperature 31 °C, minimum temperature 12 °C, average relative humidity 38%, and light intensity of 860 μmol m−2 s−1.
Selected PAB were streaked on NA plates and cultured at 28 °C for 48 h, then transferred to the NA-broth medium and cultivated for 48 h in a shaker at 150 rpm at 28 °C. The cell concentration was determined in a Neubauer hemocytometer chamber and adjusted to 1 × 105 CFU mL−1 for the seed treatment.
Seeds of the soybean cultivar TMG 132, known to be susceptible to drought, were initially disinfected in alcohol (70% ethanol) for 30 s and sodium hypochlorite (5% active chloride) for 2 min and subsequently washed thoroughly with sterile water and air-dried in a flow cabinet for 8 h. Disinfected seeds were soaked for 30 min in the bacterial suspension (2 mL L−1 seed), resistance inductor acibenzolar-S-methyl (ASM at 10 mg L−1) or water (control). Treated seeds were sown in pots of 3 L containing a mixture of autoclaved soil and sand (2:1), with three seeds per pot. Plants were kept under greenhouse conditions as described previously and watered to field capacity. Five replicates of each treatment arranged in a completely randomized design were used for the growth promotion. Three replicates of each treatment arranged in a completely randomized design were used for the physiological parameters. Both experiments were conducted twice.
Plant phenotype evaluation after drought stress
At 23 days after planting (dap), plants were subjected to drought by withholding irrigation, as has been used in recent works (Cohen et al. 2010; Staudinger et al. 2016).
At the first, second, and third days after the irrigation suspension (hereafter DAIS), plants were assessed for the potential quantum efficiency of photosystem II (PSII) (given by the Fv/Fm ratio) by using the fluorometer model Hansatech PEA MK2, Kings Lynn, England. Leaves of the second node (from the plant apex) of five plants chosen randomly for each treatment were assessed. The leaf blades were initially submitted to 20 min in the darkness before the measurements, through the use of clips connected to the fluorometer chamber. Pulse saturated light of 3000 μmol m−2 s−1 for 5 s was used. Fv/Fm ratio was measured between 8 a.m. and 9 a.m. Moreover, at the first and second DAIS photosynthetic rate (A, μmol m−2 s−1), stomatal conductance (gs, mol H2O m−2 s−1), and transpiration rate (E, mmol m−2 s−1) were measured between 9 a.m. and 11 a.m. on the second fully expanded leaf from the base of the five plants of each treatment. For the measurements, a portable LI-6400XTR infrared gas analyzer (LI-COR Biosciences, Lincoln, NE, USA) was used and adjusted to a constant chamber temperature of 24 °C. An attached LED light source was used to provide 1000 μmol photons m−2 s−1. Leaves were placed in a 6-cm2 chamber and data were recorded at 1 min after exposition. At the third DAIS, the plants were collected and checked for growth promotion parameters: percentage of seedling emergence, fresh and dry weights, and plant height. Roots were thoroughly washed in tap water, and both shoots and roots were wrapped and oven-dried at 70 °C for 72 h to a constant weight. The experiment was repeated. However, right before the plants were collected, three pots of each treatment were chosen randomly, and three leaves that had reached the fully expanded leaf stage were excised, immediately wrapped in aluminum foil, dipped in liquid nitrogen, and then kept in the freezer at − 80 °C until the total RNA extraction for the qRT-PCR.
qRT-PCR of soybean drought stress-related genes
From the stored leaf samples at − 80 °C four leaf discs of 100 mg of each treatment were placed in 1.5-mL tubes and grounded in liquid nitrogen to extract the total RNA. The extraction process was performed using TRIzol reagent (Invitrogen) following the manufacturer instructions. For the cDNA synthesis, the enzyme M-MLV (Invitrogen) was used following the manufacturer instructions. The primers used in the qRT-PCR were selected for the amplifications of the genes Gmp5cs, Gmgols, Gmdreb1a, and Gmereb, encoding for Δ1-pyrroline-5-carboxylase synthetase, galactinol synthase, dehydration-responsive protein, and ethylene response factor, respectively (Table 1). The Gm18SrRNA was used as a normalization gene, since its expression level is not affected by abiotic stresses (Stolf-Moreira et al. 2010). The reactions were performed in three technical replicates using iQ5 real time PCR system (Bio-Rad), and the conditions were 10 min at 95 °C (1 cycle), 15 s at 95 °C followed by 1 min at 60 °C (40 cycles) and a melting curve ramping from 60 to 95 °C with an increasing temperature of 0.5 °C every 10 s (1 cycle). Data were collected continuously. Each reaction contained cDNA template, forward and reverse primers (500 nM each), and 10 μL of MAXIMA® SYBR-green PCR Master mix (Fermentas). Gene expression levels were estimated according to 2−ΔΔCT method (Livak; 2001). The method was validated through verification of primer efficiency using cDNA serial dilutions (1:10).
Biocontrol of S. sclerotiorum sclerotia on semi-selective medium
The strain used in this work was Sclerotinia sclerotiorum BRM 29673, which is preserved by the Brazilian Agricultural Research Corporation (EMBRAPA). A mycelial disc of a pure culture of the pathogen was transferred to the center of Petri dishes with 20 mL of PDA medium. The plates were incubated at 20 °C for 7 days, which was the period of time necessary for the sclerotia to be produced. PDA medium was amended with 150 ppm of bromophenol blue to be named NEON (Peres et al. 2002). Then, sclerotia from BRM 29673 were treated with UFGS1, UFGS2, UFGRB2, and UFGRB3. The bacterial cell concentration was determined in a Neubauer chamber and adjusted to 1 × 105 CFU mL−1. Water and thiophanate-methyl (2 mL/L) were used as negative and positive controls, respectively. Then, three S. sclerotiorum sclerotia were transferred to each 9-mm Petri dish containing 20 mL of the NEON medium and were then incubated at 20 °C for 5 days. Plates were observed for color change and assessed for S. sclerotiorum mycelial growth. The experiment was performed twice with four replicates for treatment in each experiment.
Experimental design and statistical analysis
The completely randomized design was used for the in vitro and in vivo tests. Data with five and four replications were submitted to analysis of variance (ANOVA) respectively for in vitro and in vivo tests. Duncan’s multiple range, Dunnett’s, Scott-Knott’s, and Student’s t tests (P < 0.05) were applied for significant means when necessary. For all analyses, the assumption of normality was checked by the Shapiro-Wilk and Kolmogorov-Smirnov tests prior to analysis, and no transformation was necessary. SigmaPlot® version 11, Sisvar (Build 72) and GraphPad Prism 7.0 were used for statistical analyses and artworks (Ferreira 2011).
Results
Five bacteria from the sclerotia and five from soybean rhizosphere were isolated and used for the in vitro screening test for growth in reduced water activity. Under the medium with 2.5% of glycerol, most of the isolated bacteria had a decrease in their growth compared to the control (Fig. 1). However, four strains identified as Bacillus subtilis UFGS1 (Universidade Federal de Goiás sclerotia 1), Bacillus thuringiensis UFGS2 (Universidade Federal de Goiás sclerotia 2), Bacillus cereus UFGRB2 (Universidade Federal de Goiás rhizobacteria 2), and Bacillus cereus UFGRB3 (Universidade Federal de Goiás rhizobacteria 3) did not have their growth reduced under lower water activity compared to the control (P = 0.1757, P = 0.7646, P = 0.7309, P = 0.9451, respectively) (Fig. 1, Supplementary 1).
Effect of the addition of glycerol (2.5%) in the nutrient agar (NA) medium on the bacterial growth, assessed at 48 h. *Significant at the 0.05 probability level by Student’s t test. ***Significant at the 0.001 probability level by Student’s t test. ns = not significant. Error bars represent ± SE. (Means of four replicates for each treatment with glycerol and without glycerol)
Although the treatments did not differ among themselves regarding plant height, there was an increase by 2-fold in the number of seedlings when seeds were treated with UFGRB2 compared to the control (water) (P = 0.002) (Fig. 2).
Effect of seed treatment with plant-associated bacteria (PAB), acibenzolar-S-methyl (ASM) or water on the emergence and height of soybean seedling cultivar TMG 132, assessed at the third day after the irrigation suspension (DAIS). **Significant at the 0.01 probability level by Student’s t test compared to the water (control). ns = not significant compared to the water (control). Error bars represent ± SE. The experiment was repeated. (Means of two experiments of five replicates each)
When soybean seedlings were assessed regarding potential quantum efficiency of photosystem II (Fv/Fm) at 1, 2, and 3 days after the irrigation suspension (DAIS), there was a decrease at 3 DAIS, except in UFGS2 (P = 0.323) and UFGRB2 (P = 0.250). These two plant-associated bacteria sustained the plant photochemical efficiency at the same level in all the assessments under drought stress (Fig. 3).
Effect of seed treatment with plant-associated bacteria, acibenzolar-S-methyl (ASM) or water on the potential quantum efficiency of photosystem II (Fv/Fm) of soybean plant cultivar TMG 132, assessed at 1, 2, and 3 days after the irrigation suspension (DAIS). Bars with the same letter are similar at the 5% level according to Duncan’s multiple range tests. ns = not significant. Error bars represent ± SE. The experiment was repeated. (Means of two experiments of three replicates each). Fv/Fm* = optimal quantum yield
Likewise, when soybean seedlings were assessed for photosynthetic rate, stomatal conductance, and transpiration at 1 and 2 days after the irrigation suspension (DAIS), UFGRB3 maintained the photosynthetic rates in higher levels when compared to the control (water) at both 1 and 2 DAIS (P = 0.0130 and P = 0.0179, respectively) (Fig. 4a). In addition, UFGS2 sustained photosynthetic rates at 2 DAIS when compared to the control water (P = 0.0059) (Fig. 4a). On the other hand, for the UFGS1-treated plants there was a decrease in the photosynthetic rates at 1 DAIS (P = 0.0014) (Fig. 4a). However, at 2 DAIS, the UFGS1 treatment did not differ from the water control (P = 0.5426) (Fig. 4a).
Effect of seed treatment with plant-associated bacteria, acibenzolar-S-methyl or water on the photosynthetic rate (a), stomatal conductance (b), and transpiration rate (c) of soybean plants cultivar TMG 132 assessed at 1 and 2 days after the irrigation suspension (DAIS). The plants on the bottom are 24 days old, and the pictures were taken at 2 DAIS. *Significant at the 0.05 probability level by Student’s t test compared to the water (control). **Significant at the 0.01 probability level by Student’s t test compared to the water (control). ***Significant at the 0.001 probability level by Student’s t test compared to the water (control). ns = not significant water (control). Error bars represent ± SE. The experiment was repeated. (Means of two experiments of three replicates each)
Regarding stomatal conductance, there was a decline at 1 DAIS for UFGS1 and UFGS2 treatments (P = 0.0006 and P = 0.0104, respectively) (Fig. 4b). However, at 2 DAIS UFGS2 showed an increase in the stomatal conductance compared to the water control (P = 0.0028) (Fig. 4b).
A decrease in transpiration was found at 1 DAIS for the UFGS1, UFGS2, and UFGRB2 treatments (P = 0.0001, P = 0.0079, and P = 0.0274, respectively) (Fig. 4c). In contrast, at 2 DAIS UFGS2 presented a higher transpiration compared to the water control (P = 0.0077) (Fig. 4c).
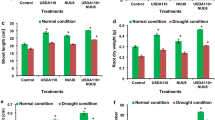
The bacterial strains UFGS2, UFGRB3, and UFGRB2 increased the dry shoot weight by 14, 31, and 41%, respectively (P = 0.0455), in addition to increasing the dry root weight by 23, 50, and 45%, respectively, when compared to the control (water) (P = 0.0217) (Fig. 5). Furthermore, the dry root weight was 25% higher for the ASM (P = 0.0217). Additionally, there was an increase in the fresh shoot weight by 29, 64, and 28% when seeds were treated with ASM and with the bacteria UFGS2 and UFGRB3, respectively (P = 0.0197). Moreover, fresh root weight was 29 and 62% higher for the treatments UFGRB3 and UFGS2, respectively (P = 0.0132).
Effect of seed treatment with plant-associated bacteria (PAB), acibenzolar-S-methyl (ASM) or water on the fresh and dry weight of soybean seedlings, assessed at 26 days after planting and 3 days after the irrigation suspension (DAIS). Bars with the same letter are similar at the 5% level according to Scott-Knott’s tests. ns = not significant. Error bars represent ± SE. The experiment was repeated. (Means of two experiments of ten replicates each)
Figure 6 shows the expression of the drought-related genes Gmp5cs, Gmgols, Gmdreb1a, and Gmereb analyzed 26 days after seed treatment with the plant-associated bacteria (PAB) and 3 DAIS.
Expression analysis of the drought-associated genes: Δ1-pyrroline-5-carboxylase synthetase (Gmp5cs), galactinol synthase (Gmgols), dehydration-responsive binding protein (Gmdreb1a) and ethylene response factors (Gmereb), in leaf samples of plants grown 26 days after seed treatment with plant-associated bacteria (PAB), acibenzolar-S-methyl (ASM) or water, and 3 days after the irrigation suspension (DAIS). *p < 0.05, **p < 0.01, and ***p < 0.001 are significant compared to the water (control). ns = not significant compared to the water (control). Expression values were compared by one-way ANOVA followed by Dunnett’s test. Error bars represent ± SE. The experiment was repeated. (Means of two experiments of three replicates each)
None of the PAB treatments differed from the control (water) regarding Gmp5cs expression. However, there was an upregulation for both Gmgols and Gmdreb1a in the presence of UFGRB2 by 1.7- and 5-fold, respectively. Moreover, UFGS1 upregulated Gmdreb1a by 2-fold. On the other hand, Gmereb was downregulated by ASM, UFGS2, and UFGRB3 by 1-, 1-, and 0.9-fold, respectively.
When the antagonistic activity of the four PAB was tested against Sclerotinia sclerotiorum sclerotia, it was found that one of the tested PAB strains, Bacillus cereus UFGRB3 inhibited by 4.24-fold the germination of Sclerotinia sclerotiorum sclerotia when compared to the water control (P < 0.001) (Fig. 7).
Effect of sclerotium treatment with plant-associated bacteria, thiophanate-methyl (2 mL/L), or water on the Sclerotinia sclerotiorum strain BRM 29673 sclerotium germination, assessed at 5 days after planting. Bars with the same letter are similar at the 5% level according to Scott-Knott’s tests. Error bars represent ± SE. The experiment was repeated. (Means of two experiments of four replicates each) *TM = thiophanate-methyl
Discussion
Even though there has been significant research on improving the ability of crop plants to tolerate drought stress through genetic modification (Nakaya and Isobe 2012; Yu et al. 2012), there are still significant hurdles to overcome. This may be associated with the complexity of networks involved in drought tolerance and the likely yield penalty associated with breeding (Krannich et al. 2015). In this study, we presented some plant-associated bacteria (PAB) as an eco-friendly alternative to cope with drought stress. PAB isolated from soybean rhizosphere and from Sclerotinia sclerotiorum sclerotia demonstrated they could increase the water use efficiency in soybean seedlings stressed with low water availability. Some plant physiological responses, such as potential quantum of photosystem II (Fv/Fm), photosynthetic rate, stomatal conductance, and transpiration were shown to have an important role in the process of minimizing drought effects in soybean (Figs. 3 and 4a–c). In a non-invasive way, chlorophyll fluorescence provides useful data for estimating the potential quantum efficiencies of PSII photochemistry, and photosynthesis data can link them to carbon assimilation (Genty et al. 1989). These parameters may be used as a stress indicator by comparing the metabolic and energetic balance of photosynthesis in plants under drought stress, as shown by Rahbarian et al. (2011) and Mishra et al. (2012). For UFGS2 and UFGRB2 treatments, even at the third day after irrigation suspension, no decrease in the Fv/Fm was found, suggesting integrity of the plant photochemical apparatus. The high Fv/Fm ratio associated with the decrease in gs (stomatal conductance) and E (transpiration rate) at the first day after irrigation suspension seems to have contributed to the turgor maintenance of UFGS2-treated plants, as shown in Fig. 4, which was taken at 2 DAIS. In addition, UFGS2-treated plants presented higher dry biomass, which indicates an effective photoassimilate increase, as expected, owing to the higher A (photosynthetic rate) at 2 DAIS, in relation to the control. Likewise, plants treated with UFGRB2 showed a higher dry biomass and a decrease in E (transpiration rate) at the first day after irrigation suspension, in addition to maintaining a high Fv/Fm ratio on the third day after irrigation suspension. However, no variation in A (photosynthetic rate) was found. Similarly, the increase in the dry biomass of UFGRB3-treated plants was also related to high photosynthetic rates, when compared to the control plants.
By sustaining the plant physiological activities at normal levels under low water availability, PAB prevented the plant from using unnecessary metabolic energy, which could be applied to other important metabolic processes, such as growth and reproduction. As shown by Niu et al. (2011), Arabidopsis thaliana primed with a plant growth-promoting rhizobacteria, Bacillus cereus AR156, invested less energy in activating cellular defense responses to biotic stress.
Among the genes that were analyzed in this study, dehydration-responsive binding protein (Gmdreb1a) was the one that showed the strongest expression in response to drought stress in the presence of the PAB (Fig. 6). Chen et al. (2006) and Stolf-Moreira et al. (2010) have shown that Gmdreb1a was also upregulated in wild and drought-sensitive soybean genotypes, respectively. In this work, we have shown that the bacteria UFGS1 and UFGRB2 increased Gmdreb1a expression of a drought-sensitive cultivar (TMG 132) when compared to the plant’s natural response to drought stress.
In most cases, when applied to plants, some PAB can offer additional benefits (Hu et al. 2011; Martins et al. 2015a). Here, we have shown that in addition to enhancing soybean water-use efficiency, some PAB strains promoted plant growth by increasing both fresh and dry plant biomass as well as seedling emergence (Figs. 1 and 5). Since agriculture accounts for 70% of all water use and the world population has continued to increase over the years, in order to avoid a possible water gap, we will have to feed more people with less water. A higher fresh biomass conferred by the PAB, as shown in this work, suggests that the microbiolized plants were able to hold more water in their tissues and made the plants more tolerant to drought (Fig. 5). Furthermore, no disease symptoms or signs of phytotoxicity in soybean plants were found with the use of the PAB. Another benefit promoted by the PAB in this study was that Bacillus cereus UFGRB3 showed in vitro germination inhibition of Sclerotinia sclerotiorum sclerotia, one of the most important pathogens in soybean (Fig. 7). The control of S. sclerotiorum sclerotia currently relies mostly on the application of chemicals, through the use of highly toxic fumigants, due to the difficulty of managing the overwinter fungal structure. However, the misuse of pesticides has led to serious human health problems, the proliferation of pesticide-resistant pathogens, and contamination of the environment (Firoz et al. 2016; Medeiros et al. 2012; Turdi and Yang 2016). Moreover, depending on the inoculum density in an area, it might require the grower to abandon the infested area, to insert crop-free periods, or to grow a less- or non-economic crop. On the other hand, the use of free-living plant-associated bacteria has been highlighted as a promising broad-spectrum means to improve plant growth and health, while being a relatively simple and low-cost alternative strategy (Kim et al. 2013; Martins et al. 2014; Timmusk and Behers 2012).
One of the forms of applying beneficial microbes to plants and that was used in this work is via seed treatment, which reduces costs due to the lower volume of the product applied and also causes minimum interference with biological equilibrium, when compared to the spray method (Prasad et al. 2016). Seed coating also helps plants ward off threats in priming by making them more adaptable to biotic and abiotic stresses (Mahmood et al. 2016). In addition, it increases the chance of having a higher plant stand, as was shown in this work (Fig. 1), and possibly a higher yield.
Although none of the tested PAB have shown a simultaneously positive result for all the analyzed variables, a better outcome might be possible through the application of a mixture of strains, a tendency that has been shown in the last decades (Jetiyanon 2007; Wang et al. 2012). In order to use this strategy in further studies, a compatibility test will be performed prior to the in vivo test, so as to select the best bacterial combinations.
Although it is unrealistic to think that solely the use of microbials will solve all agriculture problems, its use has the potential in the integrated management system to cut down the amount of chemicals used and mitigate drought due to lack of rain at some point, once most soybean plantations are not irrigated.
References
Berendsen RL, Pieterse CM, Bakker PA (2012) The rhizosphere microbiome and plant health. Trends Plant Sci 17:478–486. https://doi.org/10.1016/j.tplants.2012.04.001
Berg G, Rybakova D, Grube M, Köberl M (2016) The plant microbiome explored: implications for experimental botany. J Exp Bot 67:995–1002. https://doi.org/10.1093/jxb/erv466
Boiteux LS, Fonseca MEN, Simon PW (1999) Effects of plant tissue and DNA purification method on randomly amplified polymorphic DNA-based genetic fingerprinting analysis in carrot. J Am Soc Hort Sci 124:32–38
Chen Y, Chen P, de los Reyes B (2006) Differential responses of the cultivated and wild species of soybean to dehydration stress. Crop Sci 46:2041–2046
Chen W, Yao Q, Patil GB, Agarwal G, Deshmukh RK, Lin L, Wang B, Wang Y, Prince SJ, Song L, Xu D, An YC, Valliyodan B, Varshney RK, Nguyen HT (2016) Identification and comparative analysis of differential gene expression in soybean leaf tissue under drought and flooding stress revealed by RNA-seq. Front Plant Sci. https://doi.org/10.3389/fpls.2016.01044
Cohen D, Bogeat-Triboulot MB, Tisserant E, Balzergue S, Martin-Magniette ML, Lelandais G, Ningre N, Renou JP, Tamby JP, le Thiec D, Hummel I (2010) Comparative transcriptomics of drought responses in Populus: a meta-analysis of genome-wide expression profiling in mature leaves and root apices across two genotypes. BMC Genomics 11:630. https://doi.org/10.1186/1471-2164-11-630
Ferreira DF (2011) Program SISVAR a Computer statistic analysis system. Ciência e Agrotecnologia 35:1039–1042. https://doi.org/10.1590/S1413-70542011000600001
Firoz MJ, Xiao X, Zhu FX, Fu YP, Jiang DH, Schnabel G, Luo CX (2016) Exploring mechanisms of resistance to dimethachlone in Sclerotinia sclerotiorum. Pest Manag Sci 72:770–779. https://doi.org/10.1002/ps.4051
Gamalero E, Berta G, Glick BR (2009) The use of microorganisms to facilitate the growth of plants in saline soils. In: Khan MS, Zaidi A, Musarrat J (eds) Microbial strategies for crop improvement. Springer, Berlin, pp 1–22
Genty B, Briantais J, Baker NR (1989) The relationship between the quantum yield of photosynthetic electron transport and quenching of chlorophyll fluorescence. Biochimica et Biphysica Acta 990:87–92. https://doi.org/10.1016/S0304-4165(89)80016-9
Geraldine AM, Lopes FAC, Carvalho DDC, Barbosa ET, Rodrigues AR, Brandão RS, Ulhoa CJ, Lobo Junior M (2013) Cell wall-degrading enzymes and parasitism of sclerotia are key factors on field biocontrol of white mold by Trichoderma spp. Biol Control 67:308–316. https://doi.org/10.1016/j.biocontrol.2013.09.013
Heydari A, Pessarakli M (2010) A review on biological control of fungal plant pathogens using microbial antagonists. J Biol Sci 10:273–290
Hu X, Roberts DP, Maul JE, Emche SE, Liao X, Guo X, Liu Y, McKenna LF, Buyer JS, Liu S (2011) Formulations of the endophytic bacterium Bacillus subtilis Tu-100 suppress Sclerotinia sclerotiorum on oilseed rape and improve plant vigor in field trials conducted at separate locations. Can J Microbiol 57:539–546. https://doi.org/10.1139/W11-041
Hu X, Roberts DP, Xie L, Maul JE, Yu C, Li Y, Jiang M, Liao X, Che Z, Liao X (2014) Formulations of Bacillus subtilis BY-2 suppress Sclerotinia sclerotiorum on oilseed rape in the field. Biol Control 70:54–64. https://doi.org/10.1016/j.biocontrol.2013.12.005
Jetiyanon K (2007) Defensive-related enzyme response in plants treated with a mixture of Bacillus strains (IN937a and IN937b) against different pathogens. Biol Control 42:178–185. https://doi.org/10.1016/j.biocontrol.2007.05.008
Kamal MM, Savocchia S, Lindbeck KD, Ash GJ (2016) Biology and biocontrol of Sclerotinia sclerotiorum (Lib.) de Bary in oilseed Brassicas. Australas Plant Pathol 45:1–14. https://doi.org/10.1007/s13313-015-0391-2
Kim YC, Glick B, Bashan Y, Ryu CM (2013) Enhancement of plant drought tolerance by microbes. In: Aroca R (ed) Plant responses to drought stress. Springer Verlag, Berlin, pp 383–413
Krannich CT, Maletzki L, Kurowsky C, Horn R (2015) Network candidate genes in breeding for drought tolerant crops. Int J Mol Sci 16:16378–16400. https://doi.org/10.3390/ijms160716378
Mahmood A, Turgay OC, Farooq M, Hayat R (2016) Seed biopriming with plant growth promoting rhizobacteria: a review. FEMS Microbiol Ecol 92. https://doi.org/10.1093/femsec/fiw112
Martins SJ, Medeiros FHV, Souza RM, Resende MLV, Ribeiro Junior PM (2013) Biological control of bacterial wilt of common bean by plant growth-promoting rhizobacteria. Biol Control 66:65–71
Martins SJ, Medeiros FHV, Souza RM, Vilela LAF (2014) Is the curtobacterium-wilt biocontrol temperature-dependent? Acta Sci Agron 36:409–415. https://doi.org/10.4025/actasciagron.v36i4.18018
Martins SJ, Medeiros FHV, de Souza RM, Faria AF, Cancellier EL, Silveira HRO, Rezende MLV, Guilherme LRG (2015a) Common bean growth and health promoted by rhizobacteria and the contribution of magnesium to the observed responses. Appl Soil Ecol 87:49–55. https://doi.org/10.1016/j.apsoil.2014.11.005
Martins SJ, Soares AC, Medeiros FHV, Santos DBC, Pozza EA (2015b) Contribution of host and environmental factors to the hyperparasitism of coffee rust under field conditions. Australas Plant Pathol 44:605–610. https://doi.org/10.1007/s13313-015-0375-2
Medeiros FHV, Martins SJ, Zucchi TD, Melo IS, Batista LR, Machado JC (2012) Biological control of mycotoxin-producing molds. Ciência e Agrotecnologia 36:483–497. https://doi.org/10.1590/S1413-70542012000500001
Mishra KB, Iannacone R, Petrozza A, Mishra A, Armentano N, la Vecchia G, Trtílek M, Cellini F, Nedbal L (2012) Engineered drought tolerance in tomato plants is reflected in chlorophyll fluorescence emission. Plant Sci 182:79–86. https://doi.org/10.1016/j.plantsci.2011
Muller B, Pantin F, Génard M, Turc O, Freixes S, Piques M, Gibon Y (2011) Water deficits uncouple growth from photosynthesis, increase C content, and modify the relationships between C and growth in sink organs. J Exp Bot 62:1715–1729. https://doi.org/10.1093/jxb/erq438
Nakaya A, Isobe SN (2012) Will genomic selection be a practical method for plant breeding? Ann Bot 110:13003–13016. https://doi.org/10.1093/aob/mcs109
Niu DD, Liu HX, Jiang CH, Wang YP, Wang QY, Jin HL, Guo JH (2011) The plant growth-promoting rhizobacterium Bacillus cereus AR156 induces systemic resistance in Arabidopsis thaliana by simultaneously activating salicylate–and jasmonate/ethylene-dependent signaling pathways. Mol Plant-Microbe Interact 24:533–542. https://doi.org/10.1094/MPMI-09-10-0213
Peres AP, Nasser LCB, Machado JC (2002) Use of semi-seletive media for detection of Sclerotinia sclerotiorum on bean and soybean seeds. Fitopatol Bras 27:123–127. https://doi.org/10.1590/S0100-41582002000200001
Prasad SR, Kamble UR, Sripathy KV, Bhaskar KU, Singh DP (2016) Seed bio-priming for biotic and abiotic stress management. In: Singh DP, Singh HB, Prabha R (eds) Microbial inoculants in sustainable agricultural productivity. Springer, Berlin, pp 211–228
Rahbarian R, Khavari-Nejad R, Ganjeali A, Bagheri A, Najafi F (2011) Drought stress effects on photosynthesis, chlorophyll fluorescence and water relations in tolerant and susceptible chickpea (Cicer arietinum L.) genotypes. Acta Biologica Cracoviensia 53:47–56
Sadeghipour O, Abbasi S (2012) Soybean response to drought and seed inoculation. World Appl Sci J 17:55–60
Shinozaki A, Yamaguchi-Shinozaki K (2007) Gene networks involved in drought stress response and tolerance. J Exp Bot 58:221–227. https://doi.org/10.1093/jxb/erl164
Staudinger C, Mehmeti-Tershani V, Gil-Quintana E, Gonzalez EM, Hofhansl F, Bachmann G (2016) Evidence for a rhizobia-induced drought stress response strategy in Medicago truncatula. J Proteome 136:202–213. https://doi.org/10.1016/j.jprot.2016.01.006
Stolf-Moreira R, Medri ME, Neumaier N, Lemos NG, Pimenta JA, Tobita S, Brogin RL, Marcelino-Guimaraes FC, Oliveira MCN, Farias JRB, Abdelnoor RV, Nepomuceno AL (2010) Soybean physiology and gene expression during drought. Genet Mol Res 9:1946–1956. https://doi.org/10.4238/vol9-4gmr851
Timmusk S, Behers L (2012) Rhizobacterial application for sustainable water management on the areas of limited water resources. Irrig Drain Syst Eng. https://doi.org/10.4172/2168-9768.1000e111
Timmusk S, Timmusk K, Behers L (2013) Rhizobacterial plant drought stress tolerance enhancement: towards sustainable water resource management and food security. J Food Security 1:6–10. https://doi.org/10.12691/jfs-1-1-2
Turdi M, Yang L (2016) Trace elements contamination and human health risk assessment in drinking water from the agricultural and pastoral areas of Bay County, Xinjiang, China. Int J Environ Res Public Health 13. https://doi.org/10.3390/ijerph13100938
Wang CJ, Yang W, Wang C, Gu C, Niu DD, Liu H-X (2012) Induction of drought tolerance in cucumber plants by a consortium of three plant growth- promoting rhizobacterium strains. PLoS One 7:e52565. https://doi.org/10.1371/journal.pone.0052565
Yoon S, Ha S, Kwon S, Lim J, Kim Y, Seo H, Chun J (2017) Introducing EzBioCloud: a taxonomically united database of 16S rRNA gene sequences and whole-genome assemblies. Int J Syst Evol Microbiol 67:1613–1617. https://doi.org/10.1099/ijsem.0.001755
Yu S, Liao F, Wang F, Wen W, Li J, Mei H, Luo L (2012) Identification of rice transcription factors associated with drought tolerance using ecotilling method. PLoS One 7:e30765. https://doi.org/10.1371/journal.pone.0030765
Acknowledgements
The authors express their appreciation to the Brazilian Agricultural Research Corporation (EMBRAPA) for kindly providing the Sclerotinia sclerotiorum strain BRM 29673 to be used in this work. We also thank Caroline Elizabeth Martins for providing an English review of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible editor: Diane Purchase
Electronic supplementary material
Supplementary 1
Maximum Likelihood tree of 16S rRNA of subject strains. The tree was produced using the HKY85 model and complete deletion of missing data. The level of bootstrap support was calculated from 1500 replicates. The bar is equal to 0.01 substitutions per bp. (PNG 110 kb)
Rights and permissions
About this article
Cite this article
Martins, S.J., Rocha, G.A., de Melo, H.C. et al. Plant-associated bacteria mitigate drought stress in soybean. Environ Sci Pollut Res 25, 13676–13686 (2018). https://doi.org/10.1007/s11356-018-1610-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-018-1610-5