Abstract
Wastewater treatment plants are hot spots for antibiotic-resistant bacteria (ARB) and antibiotic resistance genes (ARGs). However, limited studies have been conducted to compare the reductions of ARB and ARGs by various biological treatment processes. The study explored the reductions of heterotrophic bacteria resistant to six groups of antibiotics (vancomycin, gentamicin, erythromycin, cephalexin, tetracycline, and sulfadiazine) and corresponding resistance genes (vanA, aacC1, ereA, ampC, tetA, and sulI) by five bench-scale biological reactors. Results demonstrated that membrane bioreactor (MBR) and sequencing batch reactor (SBR) significantly reduced ARB abundances in the ranges of 2.80∼3.54 log and 2.70∼3.13 log, respectively, followed by activated sludge (AS). Biological filter (BF) and anaerobic (upflow anaerobic sludge blanket, UASB) techniques led to relatively low reductions. In contrast, ARGs were not equally reduced as ARB. AS and SBR also showed significant potentials on ARGs reduction, whilst MBR and UASB could not reduce ARGs effectively. Redundancy analysis implied that the purification of wastewater quality parameters (COD, NH4 +-N, and turbidity) performed a positive correlation to ARB and ARGs reductions.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Wastewater treatment plants (WWTPs) are considered as important reservoirs of antibiotic-resistant bacteria (ARB) and antibiotic resistance genes (ARGs) (Rizzo et al. 2013; Pruden 2014). Wastewater provides a favorable environment, including high contents of microbial biomass, relatively abundant nutrients and antibiotic agents, for both the ARB survival and the ARGs transfer (Guardabassi et al. 2002; Rizzo et al. 2013). Bacteria including many pathogens (e.g., enterobacteria, enterococci, Staphylococcus aureus, and Aeromonas) resistant to nearly all clinically relevant antibiotics have been reported in WWTPs worldwide (Guardabassi et al. 2002; Castiglioni et al. 2008; Borjesson et al. 2009; Novo and Manaia 2010; Reinthaler et al. 2010). More than 140 ARGs have also been detected in WWTPs up to 2009 (Szczepanowski et al. 2009; Zhang et al. 2009). Because WWTPs are not specially designed for the removal of ARB and ARGs (Pruden et al. 2013), their excess sludge biomass and effluents still contain large amounts of the contaminants (Reinthaler et al. 2003; Borjesson et al. 2009; Novo and Manaia 2010), which transfer to the subsequent environments and impose great potential threats on public health.
Controlling antibiotic resistance in WWTPs is of great concerns of microbiologists and wastewater engineers (Rizzo et al. 2013). Wastewater disinfection, as a pathogen inactivation process, is considered as an important barrier to limit the release of ARB and ARGs into the environment (Guo et al. 2013b). However, the general observation is that the disinfection adopted in WWTPs shows limited efficacy on antibiotic resistance reduction. Some kinds of ARB are reported to express tolerance to chlorination (Huang et al. 2011; Yuan et al. 2015a), causing them difficult to be removed. Ultraviolet (UV) disinfection is also frequently reported for a weak reduction of ARB or ARGs (Munir et al. 2011; Guo et al. 2013a; Zhang et al. 2015). The limited reduction by chlorination and UV disinfection possibly partly attributes to the excessive abundances of ARB or ARGs. Increasing the dose usually contribute to the significant improvement of reduction efficiency (Engemann et al. 2006; Macauley et al. 2006; Zhang et al. 2015). One lab-scale study noted that doubling the typical UV dose applied in WWTPs was required to destroy vanA, mecA, tetA, and ampC (McKinney and Pruden 2012). Apparently, a biological process with the effective reduction of ARB and ARGs is essential for the subsequent disinfection.
Numerous studies have explored the effects of biological treatment on the reduction of antibiotic resistance level (Sharma et al. 2016). Christgen and colleagues used metagenomics approaches to contrast the fate of ARGs in anaerobic, aerobic, and anaerobic–aerobic sequence (AAS) bioreactors treating domestic wastewater (Christgen et al. 2015); they found that AAS and aerobic reactors were better than anaerobic units in reducing ARGs abundances. Result of Munir et al. (2011) indicated that membrane bioreactor displayed significantly higher removals of ARGs and ARB than conventional treatment plants. Chen and Zhang (2013) investigated the removal efficiencies of 6 ARGs (tetM, tetO, tetQ, tetW, sulI, and sulII) by three advanced treatment processes and found that constructed wetlands showed higher reductions (1∼3 log) than biological aerated filter (0.6∼1.2 log).
However, most current studies are conducted in various WWTPs with different raw sewage and abiotic environments (Chen and Zhang 2013). The reduction of antibiotic resistance by various biological treatment processes with identical conditions is rarely reported. This possibly weakens the reliability of the comparison, since wastewater qualities and abiotic conditions are frequently reported to affect the ARB/ARGs fates during biological treatment processes (Mc Mahon et al. 2007; Novo et al. 2013; Yang et al. 2014; Yuan et al. 2014). On the other hand, the fate of ARGs in WWTPs was usually inconsistent with their ARB (Gao et al. 2012; Huang et al. 2014; Huang et al. 2015). The single exploration of ARB or ARGs might both make biases, since numerous genes usually encode resistance to the same antibiotic, and only a tiny minority (<1 %) of the whole bacterial community is cultivable (Amann et al. 1995), making it necessary to combine the exploration of both ARB and ARGs.
Over the past decade, a steep increase in the number of WWTPs was observed in China (Chen and Zhang 2013). Most extensively adopted techniques in Chinese WWTPs achieve an effective purification of a majority of wastewater pollutants, while ARB and ARGs, as emerging contaminants, call for more strict requirements on the treatment processes of emerging WWTPs (Pruden 2014). To provide theoretical support on the selection and optimization of biological processes in WWTPs concerning the control of antibiotic resistance, the objective of this study was to explore the reductions of six groups of ARB and corresponding ARGs by various biological treatment processes in comparable abiotic conditions. Heterotrophic bacterial resistant to vancomycin (VAN), gentamicin (GEN), erythromycin (ERY), cephalexin (CEP), tetracycline (TC), and sulfadiazine (SD), as well as corresponding ARGs (vanA, aacC1, ereA, ampC, tetA, and sulI) were investigated as representative contaminants with five bench-scale bioreactors, including aerobic activated sludge (AS), biological filter (BF), aerobic sequencing batch reactor (SBR), membrane biological reactor (MBR), and upflow anaerobic sludge blanket (UASB).
Materials and methods
Bench-scale reactor design and operation
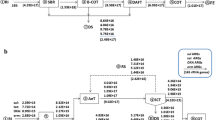
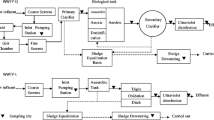
The five reactors were all made of Plexiglass®. The schematic diagram of the five reactors is presented in Fig. 1. The working volume of UASB system was 6 L, while the others were at 4 L. In AS, SBR, MBR and UASB system, a magnetic stirrer (84-1A, Sile apparatus, Shanghai) was used in each reactor for liquid agitation at a constant rate (100 rpm). An aquarium air pump (CX-0088, Chuangxing Electric, Guangdong, China) was set in AS, SBR, and MBR systems, to keep the dissolved oxygen concentration at 3∼4 mg/L. In the AS system, the reaction mixture was separated by a micro secondary sedimentation tank, with the effluent discharged and the biomass partly refluxed into the reactor (Fig. 1a). SBR operation was consisted of four stages in each cycle: filling, reaction, settling and decantation. A microcomputer control switch was used in each system to control the inflow pump, effluent electromagnetic valve and air pump, with all equipment running automatically (Fig. 1b). A flat sheet microfiltration membrane (inorganic ceramic, 30 cm × 20 cm, pore size 1 μm) was used in the MBR system (Fig. 1c). The transmembrane pressure of MBR was 0.2 MPa. Three liters of pall rings (propene polymer) was used as fillings within the BF system (Fig. 1d). The UASB reactor employed a three-phase separator to separate the treated effluent, biomass and biogas, and a water seal was established to keep an anaerobic environment (Fig. 1e). The amount of CH4 was 0.2∼0.4 m3/kg COD. The schematic diagram of the five reactors is presented in Fig. 1.
All reactors were seeded with a mixture of return-activated sludge and the effluent of the primary clarifier (considered as raw sewage) collected from a WWTP in Shanghai. The reactors were then fed with “fresh” sewage continuously by peristaltic pumps (BT300-2J/YZII25, Longer Pump) and operated according to the schedule listed in Table S1. When all reactors achieved a steady purification of turbidity and chemical oxygen demand (COD) for at least 3 weeks, it was assumed that stable operational conditions were achieved. Then, the reactors were operated for 12 weeks. The hydraulic loading rate of each reactor was changed moderately for 3 stages (every 4 weeks), so as to simulate its fluctuation in actual WWTPs (Table S1). All reactors were placed within the same room with a temperature of 25 ± 2 °C.
The reactors were sampled twice a week for physical and chemical analysis according to national standard methods, to ensure that they were running stably. Regular analyses included pH, turbidity, COD, ammonia nitrogen (NH4 +-N), total nitrogen (TN), and total phosphorus (TP). Purification efficiencies of COD, NH4 +-N, TN, and TP by the five reactors are listed in Table 1.
Detection of heterotrophic antibiotic-resistant bacteria
After four-week operation of each stage, influent and effluent samples of all reactors were sampled and analyzed for the antibiotic resistance of total heterotrophic bacteria. Culture-dependent spread-plating technique was adopted according to our previous publications (Yuan et al. 2014; Yuan et al. 2015b). Briefly, 1 mL of each homogenized sample was removed, serially diluted, and then plated on nutrient agar (Beef extract 3 g/L, peptone 10 g/L, NaCl 5 g/L, and agar 15 g/L, pH: 7.2 ± 0.2). All the plates were spiked with a certain antibiotic. Concentrations of the six antibiotics in the agar were: VAN 32 mg/L; GEN 16 mg/L; ERY 8 mg/L; CEP 16 mg/L; TC 16 mg/L; and SD 512 mg/L. The antibiotic concentrations were defined as the maximum value of all minimum inhibitory concentrations (MICs) for the pathogen resistance to an antibiotic listed in the CLSI (Clinical and Laboratory Standards Institute) documentation (CLSI 2011). All samples were processed by the standard count technique. Each sample was also incubated in nutrient agar with no antibiotic added to determine total heterotrophic bacterial count (HPC) levels.
DNA extraction and quantitative real-time PCRs
Total genome DNA was extracted from each sample after the analysis of the bacterial resistance. Twenty milliliters (raw sewage) or 200 mL (treated effluent) of each sample was filtered through a 0.22 μm cellulose acetate fiber filter. The filters were cut into small pieces and added directly to DNA extraction tubes. The extractions were conducted using the FastDNA Spin Kit for Soil (MP Biomedicals, CA, USA) according to the manufacturer’s protocol. The extraction yield and the quality of the DNA were verified by agarose gel electrophoresis and spectrophotometry (NanoDrop 8000, NanDrop Technologies, USA).
Six ARGs (vanA, aacC1, ereA, ampC, tetA, and sulI) and 16S rRNA were quantitatively detected using SYBR Green II qPCR. These genes are all prevalent ones according to previous publications and our preliminary experiment result (Szczepanowski et al. 2009; Yuan et al. 2014). In brief, the development and validation of ARGs primers were described previously (Table S2). The genes were cloned to plasmids to generate qPCR standard curves to determine the abundances. A six-point calibration curve with the gene abundance of 105∼1010 copies/L was generated for each ARG using 10-fold serial dilution of the plasmids. Based on the calibration curves, the Ct value of a test sample was used to calculate the abundance of each ARG. The detection and quantification limits were 103 and 105 copies/L, respectively.
The qPCRs were conducted in 8 trip tubes with a final volume of 20 μL, containing 10 μL Power SYBR® Green PCR Master Mix (Tiangen Biotech, Beijing), plus 0.3 μL of each primer (10 μM) and 1 μL of the template DNA. Thermal cycling and fluorescence detection were conducted on an ABI 7500 (Applied Biosystems, USA), using the following protocol: 95 °C for 15 min, followed by 40 cycles of 95 °C for 10 s, annealing at defined temperatures (Table S2) for 20 s and 72 °C for 32 s. Each reaction was run in triplicate. The amplification efficiency for ARGs was 88∼105 %. A total of 1 × 106 copies of the plasmids carrying each gene were added in serial dilutions of sample DNA to check for real-time PCR inhibition. The final concentration of template DNA in the reaction volume was controlled at <0.25 ng/μL for all primers to avoid amplification suppression. The specificity of the qPCR products was further checked by melting curves and agarose gel electrophoresis.
Data analysis
The reduction of each ARB and ARG by various biological reactors was determined as follows:
where C 0 and C i represent ARB (CFU/mL) or ARG (copies/L) abundance in the raw sewage and effluent of each reactor, respectively.
A bivariate correlation analysis was conducted between the reductions of six types of ARB and corresponding ARGs (Pearson correlation with SPSS 19.0). The Student’s t test was used to assess statistically significant differences (p < 0.05) among the values of antibiotic resistance abundance or reduction (performed with SPSS 19.0). A p value ≤0.05 was applied to reject the null hypothesis that the ARB (or ARGs) abundances or reductions were not different between different samples.
Redundancy analysis (RDA, software package CANOCO version 4.5) was used according to our previous study (Yuan et al. 2014), to assess the ARB or ARG reduction as a function of purification efficiencies of wastewater quality (COD, turbidity, NH4 +–N, HPC, and 16S rRNA). In brief, the raw data for multivariate analyses comprised species data (ARB and ARGs reductions) and other environmental variables (purification efficiencies of wastewater quality). All variables were included in the ordination. The significance of the relationship between species data and the environmental data was tested by Monte Carlo permutations test (n = 199).
Results and discussion
Comparison of ARB reduction by various biological treatment processes
The log reductions of six kinds of ARB through various biological treatment processes are presented in Fig. 2. The AS system contributed to significant reductions on all targeted ARB (p < 0.05), ranged 1.76∼2.06 log. The value was consistent with previous studies, with 1.5∼2 log removal efficiencies (Guardabassi et al. 2002; Blanch et al. 2003; Reinthaler et al. 2003). By comparison, SBR system generated higher reductions of 2.70∼3.13 log. Huang et al. (2014) reported that SBR caused TC-resistant heterotrophic bacteria abundance decreased from (5.5∼42.1) × 105 CFU/mL to (0.2∼1.1) × 105 CFU/mL. SBR was rarely reported before for its reduction of ARB possibly because it had not been as frequently adopted as AS in WWTPs.
Log reductions of six kinds of ARB through various biological treatment processes. C i and C 0 represent ARB concentrations in the effluent of each reactor and the raw sewage, respectively. VAN, GEN, ERY, CEP, TC, and SD represent the total heterotrophic bacteria resistant to vancomycin, gentamicin, erythromycin, cephalexin, tetracycline, and sulfadiazine, respectively. Error bars indicate the deviation of each ARB reduction by each reactor of various hydraulic loading rates
MBR resulted in the highest reduction on ARB abundances among the five reactors, ranging between 2.80∼3.54 log. Apparently the excellent entrapment ability of the microfiltration membrane held back the leakage of most bacteria and suspended solid. The study of Munir et al. (2011) noted that MBR utility provided higher treatment efficiency (ranging from 2.57∼7.06 log for ARGs and ARB) than other conventional treatment techniques (Munir et al. 2011). Cheng and colleagues treated antibiotic wastewater through MBR; the treatment efficiencies were 92.5 % for BOD, 96.0 % for COD, 81.5 % for suspended solids, and more than 99.9 % for antibiotics (Cheng et al. 2015). The present study showed that MBR was more effective on ARB reductions than other treatment processes, which possibly arose from the fact that most ARB were transferred into biosolids as our previous study indicated (Yuan et al. 2014). MBR could easily achieve a much longer biomass retention than other reactors by membrane entrapment (Smith et al. 2013).
BF contributed to 0.87∼1.23 log reductions of tested ARB, significantly lower than that of AS, SBR and MBR (p < 0.05). The poor efficiency of the biological filter was also observed by Novo and Manaia (2010); they reported that normal trickling filter reduced abundances of antibiotic-resistant heterotrophs, enterobacteria and enterococci very slightly. The form of biofilter treatment process possibly influenced the ARB reduction level. Novo and Manaia (2010) found that submerged aerated filter (SAF) was more efficient on ARB removal than the trickling filter.
Significant reductions of ARB by the anaerobic reactor (UASB) were also observed, but with rather low levels ranging between 0.95∼1.16 log. Oxygen condition was reported to greatly affect ARB prevalence (generally with a negative correlation) in biological treatment system (Yuan et al. 2014). Low oxygen conditions were considered relevant to the development of bacterial resistance to antibiotics (HØiby et al. 2010).
Comparison of ARGs reduction by various biological treatment processes
The log reductions of six kinds of ARGs by various biological treatment processes are explored in Fig. 3. AS and SBR contributed to significant reductions of targeted ARGs (p < 0.05), with values ranging between 2.36∼4.24 log and 1.66∼3.56 log, respectively. The potentials were similar with some previous reports (e.g., Munir et al. 2011; Al-Jassim et al. 2015;). In AS and SBR, effective sedimentation and precipitation of biomass probably caused most bacteria and intracellular ARGs transferred into biosolids. The variation of antibiotic-resistant microbial community after activated sludge treatment possibly also resulted in the reductions of ARB and ARGs prevalence, as previously reported (Al-Jassim et al. 2015; Guo et al. 2015). BF also contributed to a noticeable reduction of targeted ARGs abundances, but with a low level ranging from 0.58∼1.18.
Log reductions of six kinds of ARGs through various biological treatment processes. C 0 and C i represent ARG concentrations in the raw sewage and treated effluent of each reactor, respectively. vanA, aacC1, ereA, ampC, tetA, and sulI represent the gene encoding resistance to vancomycin, gentamicin, erythromycin, cephalexin, tetracycline, and sulfadiazine, respectively. Error bars indicate the deviation of each ARB reduction by each reactor of various hydraulic loading rates
In contrast, MBR led to little reduction of targeted ARGs abundances. Actually, most ARGs abundances increased in wastewater treated by MBR. It was not expected since MBR was the most effective process on ARB reduction, implying that the intracellular ARGs shall be retained by the membrane. The discrepancy between the traditional cultivation and qPCR method that only a tiny minority of the whole bacterial community is cultivable might only account for the difference to a small degree. MBR led to a rather effective removal of all cultivable bacteria, implied that non-cultivable bacteria was unlikely to persist in a large prevalence. It was supposed that extracellular DNA possibly persisted in MBR treated effluent (e.g., released by donors during horizontal transfer), which could leak from the membrane pores while most bacteria were entrapped at the moment. The extracellular DNA probably arose from the leakage of intracellular DNA from bacteria lysis by aeration and stir. A direct evidence was that 16S rRNA abundance in MBR-treated effluent was more prevalent compared to other reactors (Fig. S1). Higher biomass concentration (expressed by MLSS, 3.5∼5 g/L) than AS (1.5∼2.3 g/L) and SBR (1.6∼2.5 g/L) implied that more intracellular DNA be leaked. In addition, intercellular DNA transfer would also contribute to the prevalent ARGs. MBR had been reported for its property of facilitating the horizontal transfer of ARG (RP4) owning to the high sludge bacterial density, abundant nutrition, as well as the presence of biofilms (Yang et al. 2013). It should be noted that an effective sedimentation process might be important for the reduction of extracellular DNA. MBR could not prevent the leakage of dissociative extracellular DNA with the absence of effective sedimentation, compared to AS and SBR. The extracellular DNA was noticed for the ability of conferring resistance genotypes with the absence of live donor cells (Dodd 2012). A further investigation of the extracellular ARGs status in MBR effluent would help to reveal the result more explicitly. On the other hand, the membrane pore size might also affect the MBR reduction level (Breazeal et al. 2013). Another WWTP adopting MBR process with the mean pore size of 0.1∼0.4 μm sharply declined the abundances of ARGs (tetG, tetW, tetX, sul1) and integron-integrase genes (intI1) (Du et al. 2014). A smaller pore size might improve the reduction of antibiotic resistance, which is worth further study.
Similar to MBR, UASB exhibited no effective reduction of targeted ARGs with their prevalence significantly improved in the treated effluent (p < 0.05). Anaerobic condition seemed to be favorable for the persistence and development of antibiotic resistance, as Threedeach et al. (2012) indicated. Huang and colleagues reported a similar result; they found that the relative abundance of eight tetR genes changed slightly with no significant difference after anaerobic treatment (Huang et al. 2015). Results of Tao et al. (2014) noted the highest level of five ARGs existed in the anaerobic treatment tank compared to the activated sludge unit and the effluent. It should be pointed out, however, that the result did not imply the limitation of anaerobic technique application. Anaerobic technique was generally applied as a pre-treatment process in WWTPs. Anaerobic and aerobic sequence bioreactor was reported to reduce ARB or ARGs abundances significantly (Christgen et al. 2015). Another study of Chen et al. (2015) reported that the integrated constructed wetlands, which provided alternate aerobic and anaerobic environments, effectively reduced sul R, tet R, and erm R genes by over 99 %. Further optimization of the combination of anaerobic and aerobic process in antibiotic resistance reduction is interesting for a future study.
Correlation between wastewater quality parameter purifications and ARB/ARGs reductions
It was speculated above that the reductions of ARB/ARGs of biological treatment processes might be related to the purifications of conventional wastewater quality parameters, such as turbidity. The correlation between purification efficiencies of wastewater quality and ARB/ARGs was therefore explored by the redundancy analysis (RDA) (Fig. 4). All ARB reductions were observed to be tightly connected to HPC reduction (Fig. 4a). Similar correlation was also noticed between the reductions of six ARGs and 16S rRNA (Fig. 4b). The data indicated that various ARB or ARGs were nearly equally affected during the biological treatment with bacteria or gene showing no obvious tolerance. The positive correlations between the reductions of ARB and HPC, or ARGs and 16S rRNA, were also reported in other WWTPs (Chen and Zhang 2013; Wang et al. 2015).
Redundancy analysis of log reductions of six types of ARB (a) and corresponding ARGs (b) in function of purification efficiencies of wastewater quality by various biological treatment processes, including COD, turbidity, NH4 +-N, HPC, and 16S rRNA. a relations between purification efficiencies of ARB and wastewater quality; b relations between purification efficiencies of ARGs and wastewater quality. Log ARB/ARGs removals are represented by blue hollow arrows; wastewater quality purification efficiencies are represented by red solid arrows
The result implied that the detection of HPC and 16S rRNA would help to explore most ARB and ARGs status during biological treatment processes, which possibly simplified the exploration of antibiotic resistance reduction during wastewater treatment. However, no significant correlation (p > 0.05) between the reductions of ARB and corresponding ARGs was detected (Table S3). The independence probably stemmed from the discrepancy between the traditional cultivation and qPCR method, as well as the persistence of extracellular ARGs, as mentioned above. The result stressed the necessity of comprehensive investigation of both ARB and ARGs. On the other hand, the complex composition of wastewater made the significant correlation between ARB and ARGs usually difficult explore, as previously reported (Gao et al. 2012; Huang et al. 2015).
Purification efficiencies of COD, NH4 +-N, and turbidity all exhibited positive correlations on reductions of ARB (Fig. 4a) or ARGs (Fig. 4b). Most bacteria were considered to be adhered to suspended solids or colloids in wastewater, causing the reductions of ARB/ARGs positively related to COD or turbidity to some extent. DNA-colloid interactions were also reported previously; Breazeal et al. (2013) found a significant correlation between ARGs (vanA and bla TEM) reductions and corresponding colloidal fractions.
With the crescent attention on the emerging contaminants, developments of more advanced treatment techniques are greatly proposed. The partial correlation between wastewater qualities and ARB/ARGs implied a biological treatment process with effective sedimentation performance was important for antibiotic resistance reduction. SBR process performed an effective potential on the reductions of ARB/ARGs abundances. The investigation of membrane characteristics was essential for MBR to achieve an efficient performance. The forms of biofilm reactor configurations and the combination with aerobic treatment for anaerobic treatment were also highly interesting to optimize so as to promote their capacities of controlling antibiotic resistance. On the other hand, none of the reactors could completely remove antibiotic resistance, with (8.5∼2.1 × 105) CFU/mL of ARB and (1.0∼6.0 × 107) copies/L of ARGs persisting in treated effluents. Disinfection process was essential for the further reduction of antibiotic resistance to an acceptable level. In addition, the exploration of novel treatment processes is of great interest to effectively control antibiotic resistance in final effluents.
Conclusions
The study compared the reductions of six groups of ARB and corresponding ARGs in municipal wastewater by five biological treatment reactors. MBR and SBR reduced ARB abundances significantly with values ranging between 2.80∼3.54 log and 2.70∼3.13 log, respectively, followed by AS system of 1.76∼2.06 log. BF and UASB led to lower reduction levels, with values of 0.87∼1.23 log and 0.95∼1.16 log, respectively. However, ARGs were not equally affected by various biological processes. AS and SBR also demonstrated significant potentials on ARGs reductions, with values ranged 2.36∼4.24 log and 1.66∼3.56 log. BF resulted in an obvious reduction on ARGs abundances, but with a low value ranged 0.58∼1.18. In contrast, MBR and UASB could not reduce ARGs effectively, which caused ARGs abundances increased in treated effluents.
Redundancy analysis showed HPC purification level was strongly correlated to all ARB reductions. Similar correlation was also detected between the purifications of 16S rRNA and six ARGs. However, the correlation between the reductions of ARB and corresponding ARGs was not significant (p > 0.05). The purifications of COD, NH4 +-N, and turbidity all exhibited positive correlations to ARB or ARG reductions. Biological processes with effective purifications of conventional wastewater quality parameters would partially contribute to the promotion of antibiotic resistance reduction.
References
Al-Jassim N, Ansari MI, Harb M, Hong PY (2015) Removal of bacterial contaminants and antibiotic resistance genes by conventional wastewater treatment processes in Saudi Arabia: is the treated wastewater safe to reuse for agricultural irrigation? Water Res 73:277–290
Amann RI, Ludwig W, Schleifer K (1995) Phylogenetic identification and in situ detection of individual microbial cells without cultivation. Microbiol Rev 59:143–169
Blanch AR, Caplin JL, Iversen A, Kuhn I, Manero A, Taylor HD, Vilanova X (2003) Comparison of enterococcal populations related to urban and hospital wastewater in various climatic and geographic European regions. J Appl Microbiol 94:994–1002
Borjesson S, Melin S, Matussek A, Lindgren PE (2009) A seasonal study of the mecA gene and Staphylococcus aureus including methicillin-resistant S. aureus in a municipal wastewater treatment plant. Water Res 43:925–932
Breazeal MV, Novak JT, Vikesland PJ, Pruden A (2013) Effect of wastewater colloids on membrane removal of antibiotic resistance genes. Water Res 47:130–140
Castiglioni S, Pomati F, Miller K, Burns BP, Zuccato E, Calamari D, Neilan BA (2008) Novel homologs of the multiple resistance regulator marA in antibiotic-contaminated environments. Water Res 42:4271–4280
Chen H, Zhang M (2013) Effects of advanced treatment systems on the removal of antibiotic resistance genes in wastewater treatment plants from Hangzhou, China. Environ Sci Technol 47:8157–8163
Chen J, Liu YS, Su HC, Ying GG, Liu F, Liu SS, He LY, Chen ZF, Yang YQ, Chen FR (2015) Removal of antibiotics and antibiotic resistance genes in rural wastewater by an integrated constructed wetland. Environ Sci Pollut Res 22: 1794–1803
Cheng SF, Lee YC, Kuo CY, Wu TN (2015) A case study of antibiotic wastewater treatment by using a membrane biological reactor system. Int Biodeterior Biodegrad 102:398–401
Christgen B, Yang Y, Ahammad SZ, Li B, Rodriquez DC, Zhang T, Graham DW (2015) Metagenomics shows that low-energy anaerobic–aerobic treatment reactors reduce antibiotic resistance gene levels from domestic wastewater. Environ Sci Technol 49:2577–2584
Clinical and Laboratory Standards Institute (CLSI) (2011) Performance standards for antimicrobial susceptibility testing: twenty-first informational supplement, 31
Dodd MC (2012) Potential impacts of disinfection processes on elimination and deactivation of antibiotic resistance genes during water and wastewater treatment. J Environ Monit 14:1754–1771
Du J, Geng J, Ren H, Ding L, Xu K, Zhang Y (2014) Variation of antibiotic resistance genes in municipal wastewater treatment plant with A2O-MBR system. Environ Sci Pollut Res 22:3715–3726
Engemann CA, Adams L, Knapp CW, Graham DW (2006) Disappearance of oxytetracycline resistance genes in aquatic systems. FEMS Microbiol Lett 263:176–182
Gao P, Munir M, Xagoraraki I (2012) Correlation of tetracycline and sulfonamide antibiotics with corresponding resistance genes and resistant bacteria in a conventional municipal wastewater treatment plant. Sci Total Environ 421-422:173–183
Guardabassi L, Lo Fo Wong DMA, Dalsgaard A (2002) The effects of tertiary wastewater treatment on the prevalence of antimicrobial resistant bacteria. Water Res 36:1955–1964
Guo MT, Yuan QB, Yang J (2013a) Microbial selectivity of UV treatment on antibiotic-resistant heterotrophic bacteria in secondary effluents of a municipal wastewater treatment plant. Water Res 47:6388–6394
Guo MT, Yuan QB, Yang J (2013b) Ultraviolet reduction of erythromycin and tetracycline resistant heterotrophic bacteria and their resistance genes in municipal wastewater. Chemosphere 93:2864–2868
Guo MT, Yuan QB, Yang J (2015) Insights into the amplification of bacterial resistance to erythromycin in activated sludge. Chemosphere 136:79–85
HØiby N, Bjarnsholt T, Givskov M, Molin S, Oana C (2010) Antibiotic resistance of bacterial biofilm. Int J Antimicrob Agents 25:322–332
Huang JJ, Hu HY, Tang F, Li Y, Lu SQ, Lu Y (2011) Inactivation and reactivation of antibiotic-resistant bacteria by chlorination in secondary effluents of a municipal wastewater treatment plant. Water Res 45:2775–2781
Huang MH, Zhang W, Zheng Y, Zhang W (2014) Correlation among extracellular polymeric substances, tetracycline resistant bacteria and tetracycline resistance genes under trace tetracycline. Chemosphere 117:658–662
Huang MH, Zhang W, Liu C, Hu HY (2015) Fate of trace tetracycline with resistant bacteria and resistance genes in an improved AAO wastewater treatment plant. Process Saf Environ Prot 93:68–74
Macauley JJ, Qiang Z, Adams CD, Surampalli R, Mormile MR (2006) Disinfection of swine wastewater using chlorine, ultraviolet light and ozone. Water Res 40:2017–2026
Mc Mahon MA, Blair IS, Moore JE, Mc Dowell DA (2007) The rate of horizontal transmission of antibiotic resistance plasmids is increased in food preservation-stressed bacteria. J Appl Microbiol 103:1883–1888
McKinney CW, Pruden A (2012) Ultraviolet disinfection of antibiotic resistant bacteria and their antibiotic resistance genes in water and wastewater. Environ Sci Technol 46:13393–13400
Munir M, Wong K, Xagoraraki I (2011) Release of antibiotic resistant bacteria and genes in the effluent and biosolids of five wastewater utilities in Michigan. Water Res 45:681–693
Novo A, Manaia CM (2010) Factors influencing antibiotic resistance burden in municipal wastewater treatment plants. Appl Microbiol Biotechnol 87:1157–1166
Novo A, Andre S, Viana P, Nunes OC, Manaia CM (2013) Antibiotic resistance, antimicrobial residues and bacterial community composition in urban wastewater. Water Res 47:1875–1887
Pruden A (2014) Balancing water sustainability and public health goals in the face of growing concerns about antibiotic resistance. Environ Sci Technol 48:5–14
Pruden A, Larsson DG, Amezquita A, Collignon P, Brandt KK, Graham DW, Lazorchak JM, Suzuki S, Silley P, Snape JR, Topp E, Zhang T, Zhu YG (2013) Management options for reducing the release of antibiotics and antibiotic resistance genes to the environment. Environ Health Perspect 121:878–885
Reinthaler FF, Posch J, Feierl G, Wüst G, Haas D, Ruckenbauer G, Mascher F, Marth E (2003) Antibiotic resistance of E. coli in sewage and sludge. Water Res 37:1685–1690
Reinthaler FF, Feierl G, Galler H, Haas D, Leitner E, Mascher F, Melkes A, Posch J, Winter I, Zarfel G, Marth E (2010) ESBL-producing E. coli in Austrian sewage sludge. Water Res 44:1981–1985
Rizzo L, Manaia C, Merlin C, Schwartz T, Dagot C, Ploy MC, Michael I, Fatta-Kassinos D (2013) Urban wastewater treatment plants as hotspots for antibiotic resistant bacteria and genes spread into the environment: a review. Sci Total Environ 447C:345–360
Sharma VK, Johnson N, Cizmas L, McDonald TJ, Kim H (2016) A review of the influence of treatment strategies on antibiotic resistant bacteria and antibiotic resistance genes. Chemosphere 150:702–714
Smith AL, Skerlos SJ, Raskin L (2013) Psychrophilic anaerobic membrane bioreactor treatment of domestic wastewater. Water Res 47:1655–1665
Szczepanowski R, Linke B, Krahn I, Gartemann KH, Gutzkow T, Eichler W, Puhler A, Schluter A (2009) Detection of 140 clinically relevant antibiotic-resistance genes in the plasmid metagenome of wastewater treatment plant bacteria showing reduced susceptibility to selected antibiotics. Microbiology 155:2306–2319
Tao CW, Hsu BM, Ji WT, Hsu TK, Kao PM, Hsu CP, Shen SM, Shen TY, Wan TJ, Huang YL (2014) Evaluation of five antibiotic resistance genes in wastewater treatment systems of swine farms by real-time PCR. Sci Total Environ 496:116–121
Threedeach S, Chiemchaisri W, Watanabe T, Chiemchaisri C, Honda R, Yamamoto K (2012) Antibiotic resistance of Escherichia coli in leachates from municipal solid waste landfills: comparison between semi-aerobic and anaerobic operations. Bioresour Technol 113:253–258
Wang J, Mao D, Mu Q, Luo Y (2015) Fate and proliferation of typical antibiotic resistance genes in five full-scale pharmaceutical wastewater treatment plants. Sci Total Environ 526:366–373
Yang D, Wang J, Qiu Z, Jin M, Shen Z, Chen Z, Wang X, Zhang B, Li JW (2013) Horizontal transfer of antibiotic resistance genes in a membrane bioreactor. J Biotechnol 167:441–447
Yang CW, Chang YT, Chao WL, Shiung II, Lin HS, Chen H, Ho SH, Lu MJ, Lee PH, Fan SN (2014) An investigation of total bacterial communities, culturable antibiotic-resistant bacterial communities and integrons in the river water environments of Taipei city. J Hazard Mater 277:159–168
Yuan QB, Guo MT, Yang J (2014) Monitoring and assessing the impact of wastewater treatment on release of both antibiotic-resistant bacteria and their typical genes in a Chinese municipal wastewater treatment plant. Environ Sci Process Impacts 16:1930–1937
Yuan QB, Guo MT, Yang J (2015a) Fate of antibiotic resistant bacteria and genes during wastewater chlorination: implication for antibiotic resistance control. PLoS One 10:e0119403
Yuan QB, Guo MT, Yang J (2015b) The sludge loading rate regulates the growth and release of heterotrophic bacteria resistant to six types of antibiotics in wastewater activated sludge. Environ Sci Process Impacts 17:206–212
Zhang XX, Zhang T, Fang HH (2009) Antibiotic resistance genes in water environment. Appl Microbiol Biotechnol 82:397–414
Zhang Y, Zhuang Y, Geng J, Ren H, Zhang Y, Ding L, Xu K (2015) Inactivation of antibiotic resistance genes in municipal wastewater effluent by chlorination and sequential UV/chlorination disinfection. Sci Total Environ 512-513:125–132
Acknowledgment
This project was funded by the National Natural Science Foundation of China (51308399) and the Shanghai Natural Science Foundation (13ZR1443300). The authors would like to thank the engineers of the WWTP for their assistance in obtaining the wastewater samples.
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible editor: Gerald Thouand
Electronic supplementary material
ESM 1
(DOCX 63.6 kb)
Rights and permissions
About this article
Cite this article
Yuan, QB., Guo, MT., Wei, WJ. et al. Reductions of bacterial antibiotic resistance through five biological treatment processes treated municipal wastewater. Environ Sci Pollut Res 23, 19495–19503 (2016). https://doi.org/10.1007/s11356-016-7048-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-016-7048-8