Abstract
Brain atlases play a key role in modern neuroimaging analysis of brain structure and function. We review available atlas databases for humans and animals and illustrate common state-of-the-art workflows in neuroimaging research based on image registration. Advances in noninvasive imaging methods, 3D ex vivo microscopy, and image processing are summarized which will eventually close the current resolution gap between brain atlases based on conventional 2D histology and those based on 3D in vivo imaging.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
In neuroimaging, atlasing refers to registering acquired higher dimensional brain data, e.g., 4D time series or multimodal images, to a 3D reference space called “atlas” that is parcellated and labeled based on known functional and/or anatomical features. Anatomical parcellation of the brain and atlas creation is one of the oldest neuroscience modalities initiated in modern neuroscience with phrenology in the nineteenth century (Fig. 1) [2]. Numerous different atlases are currently available for a number of species, and this battery is constantly growing as applications in experimental neurological research progress to better understand structural and functional relationships of the healthy and diseased brain. During the last decades, with the advancement of non-invasive neuroimaging and computational technology, atlasing is rejuvenated and many ongoing large projects and initiatives such as the Human Connectome Project (https://www.humanconnectome.org), the Allen Brain Initiative (http://www.brain-map.org), and Brain/MINDS (http://brainminds.jp/en/) put concerted efforts in mapping the brains of different species to great detail. To this end, very recently, atlases started to serve more specific, tailored applications. For example, methodology to generate high-resolution mouse vascular atlases that are very important in imaging techniques such as magnetic resonance imaging (MRI), positron emission tomography (PET), X-ray computed tomography (CT), functional ultrasound (fUS), and others have impacted the neuroimaging field. To date, the vast majority of neuroimaging researchers use brain atlases since these provide possibilities for automated, more systematic and novel analyses of imaging data. However, there seems little consensus and guidance on the choice of atlas and/or software to align imaging data with it, especially in basic and preclinical research. In this review, we therefore would like to provide an overview of available atlas resources for both animal species and humans. However, due to the fact that many laboratories use own atlas adaptations, we cannot guarantee completeness. A brief insight into methodology to register imaging data to an atlas and examples of research applications will be given as well as a discussion of remaining methodological and conceptual challenges from the point of view of the neuroimaging community.
Phrenological chart of the faculties from the late nineteenth century. Adapted from [1].
Histological Atlases
Brain atlases are of utmost importance for any neuroimaging approach as soon as a certain structure, a region of interest (ROI), has to be anatomically identified. Historically, anatomical brain atlases have been derived from two-dimensional histological sections, which were labeled by experts, e.g., the very well-known and highly appreciated atlases for mouse and rat by Paxinos et al. [3, 4] or the human Talairach atlas [5]. Other frequently used databases exist for mice and rats [6, 7], dogs [8, 9], cats [10], or monkeys [11, 12]. These histological atlases provide microscopic, i.e., cellular resolution, which was and still is of importance to sufficiently delineate the different substructures in the brain. At this cellular level, histochemistry provided specific markers like Nissl staining or ACh-esterase staining serving as basic information for brain parcellation. This approach goes back to the times of Brodmann’s parcellation of the human brain [13]. Despite advances of modern neuroimaging like non-invasive 3D datasets, 2D histological atlases remain the gold standard for the broad community of neuroscience researchers. This results from the fact that immunohistochemical and histological staining techniques provide molecular insight, e.g., antibody as well as genetically defined (mRNA) markers at the (sub-) cellular level. Therefore, there is a need to register 2D images into a “real world” 3D reference system. However, this remains a challenge (see below). In the neuroscience community and the public, the most perceived initiative to encompass all of this information in a single, publicly available database, is the Allen Brain Institute, which provides 3D cellular, genetical, and connectivity information of the mouse but also human brain (http://atlas.brain-map.org/). Most importantly, compared to atlases based on single or few specimens, the Allen brain atlas (ABA) of the mouse brain used modern nonlinear image registration/warping techniques to build an anatomical atlas from > 1500 specimens resulting in a group template that takes into account the interindividual differences within a species.
Neuroimaging Atlases and the Resolution Gap
Despite technical advances, all histological atlases suffer to a different degree from cutting and preparation/staining artifacts, and most importantly, the full three-dimensional context is lost although several approaches have been worked out to cope with some of these constraints, e.g., z-stacking of 2D slices [14, 15], the block-face method [16,17,18,19], or serial two photon microscopy [20, 21]. Furthermore, the role of fast 3D light sheet microscopic imaging (LSM) of cleared tissue will certainly grow to improve the quality of histological atlas databases [22]. Despite these advances of ex vivo techniques, methodologically inherent in 3D imaging techniques like MRI, SPECT, PET, or CT is the 3D context, the preserved original in vivo geometry, and the high if not even artifact-free data quality. For example, the size and shape of ventricles might substantially change from the in vivo to the ex vivo condition and obviously even more during any further preparation step. Such critical issues must be taken more into account in the future. Moreover, the noninvasive methods allow for repetitive anatomical but also functional studies on the very same specimen. Unfortunately, all these imaging methods only have less spatial resolution than histological techniques. However, technical improvements particular in the MRI field lead to much higher resolutions in the order of tens of microns [23, 24] which coined the term MRI microscopy [25, 26]. Based on these resolutions and improved imaging contrasts, MRI-based full 3D brain atlases became available not only particularly for mice and rats [27,28,29,30,31,32,33,34,35,36] but also for other species like gerbils [37], dogs [38, 39], cats [40], and monkeys [41,42,43,44]. Of note, after 3D imaging, the brain undistorted in the skull, e.g., by MRI microscopy, classical histology, and staining procedures can be applied adding dimensions, i.e., new information. Therefore, multimodal whole brain atlases were generated combining the best from the two worlds like the Waxholm datasets of the rat [33] and the mouse [34] as only two examples and multiple efforts for human brain, reviewed in [45]. Very valuable information resources do exist on the internet providing various collections of brain atlases:
Atlas Registration
At this point, another methodological progress is of great importance: non-affine registration or warping approaches for building probabilistic atlases that take the variance across a group of individual specimens from the same species into account. Probabilistic atlases, due to averaging across many individual subjects (after appropriate registration), provide much higher SNR in the datasets and thus more accurate delineation of structures, and it is possible to constantly add individual data to the cohort underlying the database [45]. This provides a mechanism to regularly improve the representativeness of the atlas for the species of interest [46]. Generating an atlas from a huge number of individuals (> 1500 in case of ABA) provides an advancement on its own render new anatomical details possible as the individual barrels of mouse barrel field cortex which cannot be seen on single subjects (extended Fig. 8 in [6]).
Even more importantly, registration of high quality, in particular the nonlinear transformation techniques, allows for a new branch of anatomical analyses, namely voxel-wise group statistics, which were introduced under the term “voxel-based morphometry” (VBM) [47]. In the meantime, this research area has diversified widely [48, 49]. These analyses led to fundamentally new insights into neurobiological research exemplified here in the context of learning and memory with the hallmark papers of Maguire et al. [50, 51]. The authors showed that London taxi drivers have an above average hippocampal volume for performing their task. Thereafter, VBM was used to assess different pathologies of the human brain like in the context of chronic pain, e.g., migraine or comparing across different chronic diseases [52,53,54,55]. This success story was translated to mice, e.g., paralleling the taxi driver finding in mice which learned different spatial tasks [56]. VBM also serves as an important tool in preclinical research, especially in rodent studies of brain pathologies that come along with morphometric changes, e.g., due to atrophy in Huntington’s disease [57], edema-induced tissue swelling [58], genic modification [59], and neurodegeneration [60, 61].
A typical workflow on how atlas registration is used for neuroimaging research is illustrated in Fig. 2. An accurate image registration technique is the main prerequisite to the validity of such a workflow. Nowadays, the most prevalent approach for medical imaging is the use of intrinsic image registration to an average template based on intensity-based similarity measures such as Mutual Information, followed by affine and an optional elastic transformation (reviewed in [62,63,64]). Fortunately, several image registration tools combining these steps for neuroimaging data are available nowadays such as FSL, SPM, FreeSurfer, ANTs, AFNI, and Vinci (for a comprehensive list and links for download see for example www.nitrc.org). However, most of these tools were developed and optimized for human neuroimaging data and often fail when processing animal data. Several projects have therefore generated dedicated databases and registration or segmentation pipelines, e.g., for the mouse [58, 65, 66] or ovine [67] brain.
Typical workflow for the use of atlases in neuroimaging studies. Using image registration techniques, individual imaging data are aligned with a group average and an atlas in a standard coordinate space allowing region- and voxel-wise group analyses. The inverse transformation of the image registration allows mapping the results back in standard space on the original data. Example images are taken from MRI acquisitions in mouse models of neurological diseases and from the Allen mouse brain atlas. Adapted from [58].
One caveat of atlas registration is that several preprocessing steps (e.g., brain-extraction) and tuning of a large set of registration, regularization, and interpolation parameters is necessary. This leaves many degrees of freedom to individual researchers. Although variability in atlases and protocols is still high between research groups [68], the human MRI field could be a good example on how to tackle this problem. To this end, the Montreal Neurological Institute (MNI) human atlas is generally considered a standard and detailed data processing pipelines for image segmentation, registration, and voxel-based statistics have been published for large cohort studies such as the Human Connectome Project in order to make data processing more coherent [69, 70]. For animal studies, large population imaging studies are in their infancy and common agreements are scarce. Efforts to improve this, e.g., publication of detailed parameter settings when using image registration software, are highly encouraged. A good practice example is given by the elastix wiki parameter database which allows users to upload their settings when using the software along the publication of their results (http://elastix.isi.uu.nl/).
Role of Atlases for Functional Imaging
In addition, and somewhat parallel to structural analyses, there is he usage of brain atlases in the context of functional imaging studies. Here, activation biomarkers assessed by 2-deoxy-d-[14C]glucose autoradiography [71,72,73], 2-deoxy-2-[18F]fluoro-d-glucose PET, SPECT [70], or functional MRI blood-oxygenation-level-dependent (fMRI BOLD) activation are intended to be analyzed in a brain structure-specific manner and compared between groups of subjects. To obtain a “match” between functional data, in general at a (much) lower resolution, and higher resolved brain atlases, different registration approaches are applied (for review see [74]). Here, the obtained quality of such a match between atlas and the single subject data space is of great impact for the final results. Automated functional analysis pipelines, which are used in a “black-box” manner, starting from the raw data as input and provide (significant) statistical differences at the output should be much more scrutinized or better falsified than is currently the case. In addition, the complexity and consequently the constraints of the statistical analysis have to be taken into account avoiding false positive results [75, 76].
Structural-Functional Relationship
The possibility to obtain atlas-based multiregional functional activation parameters, like BOLD, tracer accumulation in Manganese-enhanced MRI, PET, SPECT, etc. as well as anatomical data (connectivity, brain region volume), and correlation/regression with behavioral/functional testing, allows for more modern, higher dimensional data analyses. For example, voxel-wise analysis of lesion location over groups in atlas space allows mapping of regions whose damage best explain a functional deficit after stroke [77]. More and more, the dynamic and complex interaction mechanisms between multiple brain regions are taken into account using a combination of statistics used to analyze functional data and graph theory [78,79,80,81]. Vice versa, a better understanding of the relationship between brain structure and function has recently been used to feed multimodal neuroimaging data [82], or neuroimaging fused with genetic data [83] into personalized models of a simulated brain that can be used to predict the brain’s function in silico. Another new endeavor is to provide multimodal anatomical-functional parcellation schemes, e.g., incorporate functional data like resting state fMRI data to obtain parcellations of brain structures at a much higher level fusing anatomical/histological and functional information [84, 85].
Developing Brain and Exotic Species
Brain development is a highly complex process in terms of 3D relationships during development. In order to visualize these relationships, developmental atlases like emap (https://www.emouseatlas.org/emap/home.html) and ABA mouse and human were introduced. Further mention should be made of the fact that more and more high-resolution atlases are being made available for single brain structures/areas like the cerebral cortex [86] or the rat hippocampus [86]. Of note, brain atlases are becoming available for more and more species such as insects (drosophila, honey bee, desert locust) [87,88,89], birds [90,91,92,93,94], fishes [95,96,97,98,99,100], and opossum rat [101], using state of the art ex vivo and in vivo imaging technologies (see, e.g., https://scalablebrainatlas.incf.org/ for an overview for the mammalian species [102]). It is noteworthy that most of these atlases are freely accessible to the scientific community. Finally and very recently, additional data entities led to an enormous boost in dimensionality of digital brain atlases to name only a few topics: connectivity assessed by MR diffusion tensor imaging [103], connectivity assessed by viral trace injections in the ABA [6], physiological activity in response to visual stimuli in the Allen brain observatory project (http://observatory.brain-map.org/visualcoding/), or OMICS approaches like ViBrism [104].
Vasculature
Next to the general mouse whole brain anatomical atlases, more specific atlases have been developed focusing on the brain’s vasculature describing vascular spatial location, vascular distribution patterns, and regional vascular density [105,106,107]. The cerebral vasculature has a key supporting role by supplying the brain with oxygen and nutrients, and removing brain waste metabolites. By doing so, it is maintaining the homeostasis energy metabolism of the brain. Structural vascular abnormalities can have severe impact on normal brain functioning and have been observed in various neuropathological diseases such as stroke, Alzheimer’s disease, Huntington’s disease, multiple sclerosis, and brain tumors [108]. Hence, in-depth knowledge about the cerebral vascular architecture and its alterations in disease are quintessential. Furthermore, spatial information of cerebral vasculature is very useful for improving processing in techniques depending on blood flow or oxygenation such as (resting state) fMRI, fUS, and intrinsic optical signal imaging.
In recent years, advancements in preclinical imaging techniques increased the possibilities for visualization of the cerebral vasculature at global brain scale thus allowing the development of full brain vascular atlases. Ex vivo histology-based automatic sectioning imaging techniques such as knife-edge scanning microscopy (KESM), micro-optical sectioning tomography (MOST), and serial two photon tomography allow whole brain vascular imaging at submicron resolution [20, 109,110,111]. However, acquisition of these datasets is very time consuming due to the embedding procedures and extensive imaging time. Nonetheless, these techniques allow precise investigation of the whole brain cerebral vascular architecture at the capillary level (< 10 μm). More recently, other ex vivo histological techniques have been developed allowing whole brain vascular imaging based on brain clearing techniques such as Clarity and iDisco in combination with LSM for vascular imaging [112,113,114,115]. These techniques rely upon imaging an almost transparent whole brain combined with vasculature fluorescent staining. Examples are lectin-based staining or vascular casting [113, 116]. In contrast to automatic sectioning techniques, the fast imaging time of LSM allows high throughput vascular imaging up to a resolution of ~ 5 μm. Furthermore, LSM also provides additional auto-fluorescence background images which can later on be used to co-register the whole brain vasculature to anatomical atlases, e.g., the ABA [15, 117].
In contrast to ex vivo imaging techniques, non-invasive in vivo imaging techniques have the clear advantage to assess the development of whole brain macro-vasculature and venous sinuses within the same subject over time. CT and transcranial functional ultrasound can be used to visualize the whole brain macro-vasculature with resolutions down to 19 and 50 μm, respectively [118,119,120]. Although these techniques are able to acquire high-quality whole brain vasculature, they lack anatomical information that can help to spatially localize vessels. Thus, combining these techniques with MRI for soft tissues can provide additional information that can help area identification and alignment to other atlases. Furthermore, MRI can also acquire whole brain vasculature by either intrinsic vascular signal using time of flight imaging sequence (TOF) or contrast agents, allowing combined acquisition of anatomical and vascular information up to the resolution of 20 μm (Fig. 3) [121,122,123].
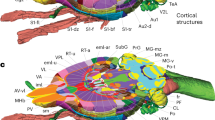
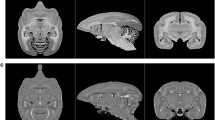
Generation of a vascular atlas of the mouse brain. In vivo 3D structural MR image (a) and time of flight angiography (b) of large vessels with high blood flow. Vascular casting followed by light sheet microscopic imaging adds an angiogram of all vessels including small vessels and veins (c). Autofluorescence images acquired in the same orientation and the structural MR image can be registered to the Allen brain atlas (d) in order to build up a probabilistic vascular atlas database in standard space.
Conclusion
Given the great success of 3D brain atlases and their analysis framework through the flagship project Allen brain atlas already allows in silico studies on the anatomy and function of the brain. This is very effective and cost-saving, reduces the number of laboratory animals (3R principle), and allows to address completely new research questions. Advances in ex vivo and in vivo imaging techniques will further close the gap between histological and neuroimaging atlases which will add more and more atlas dimensions and will help to better understand the complex interplay between the brain’s electrical activity, cellular/molecular composition, structural connectivity, vascular supply, and gene expression.
References
(1879) The people’s cyclopedia of universal knowledge: with numerous appendixes invaluable for reference in all departments of industrial life, the whole brought down to the year 1883. Phillips & Hunt
Simpson D (2005) Phrenology and the neurosciences: contributions of F.J. Gall and J.G. Spurzheim. ANZ J Surg 75:475–482
Paxinos G, Franklin KBJ (2004) The mouse brain in stereotaxic coordinates. Elsevier Acad. Press, Amsterdam
Paxinos G, Watson C (2018) The rat brain in stereotaxic coordinates. Elsevier Acad. Press, Amsterdam
Talairach J, Tournoux P (1988) Co-planar stereotaxic atlas of the human Brain.3-dimensional proportional system: an approach to cerebral imaging. Thieme Medical Publishers, Stuttgart
Oh SW, Harris JA, Ng L, Winslow B, Cain N, Mihalas S, Wang Q, Lau C, Kuan L, Henry AM, Mortrud MT, Ouellette B, Nguyen TN, Sorensen SA, Slaughterbeck CR, Wakeman W, Li Y, Feng D, Ho A, Nicholas E, Hirokawa KE, Bohn P, Joines KM, Peng H, Hawrylycz MJ, Phillips JW, Hohmann JG, Wohnoutka P, Gerfen CR, Koch C, Bernard A, Dang C, Jones AR, Zeng H (2014) A mesoscale connectome of the mouse brain. Nature 508:207–214
Sunkin SM, Ng L, Lau C, Dolbeare T, Gilbert TL, Thompson CL, Hawrylycz M, Dang C (2012) Allen brain atlas: an integrated spatio-temporal portal for exploring the central nervous system. Nucleic Acids Res 41:D996–D1008
Adrianov OS, Mering TA, Mering TA (2010) Atlas of the canine brain. NPP Books, Arlington
Palazzi X (2011) The beagle brain in stereotaxic coordinates. Springer, New York
Snider RS, Niemer WT (1961) A stereotaxic atlas of the cat brain. University of Chicago Press, Chicago
Paxinos G, Huang X-F, Toga AW (2000) The rhesus monkey brain in stereotaxic coordinates. Academic Press, Sat Lake City
Palazzi X, Bordier N (2008) The marmoset brain in stereotaxic coordinates. The marmoset brain in stereotaxic coordinates. Springer, New York, pp 1–59
Brodmann K (1909) Vergleichende Lokalisationslehre der Grosshirnrinde in ihren Prinzipien dargestellt auf Grund des Zellenbaues. J.A. Barth, Leipzig
Hess A, Lohmann K, Gundelfinger ED, Scheich H (1998) A new method for reliable and efficient reconstruction of 3-dimensional images from autoradiographs of brain sections. J Neurosci Methods 84:77–86
Simonetti AW, Elezi VA, Farion R, Malandain G, Segebarth C, Rémy C, Barbier EL (2006) A low temperature embedding and section registration strategy for 3D image reconstruction of the rat brain from autoradiographic sections. J Neurosci Methods 158:242–250
Roy D, Steyer GJ, Gargesha M, Stone ME, Wilson DL (2009) 3D cryo-imaging: a very high-resolution view of the whole mouse. Anat Rec (Hoboken) 292:342–351
Wilson D, Roy D, Steyer G et al (2008) Whole mouse cryo-imaging. Proc SPIE Int Soc Opt Eng 6916:69161I–69161I9
Tajika Y, Murakami T, Iijima K, Gotoh H, Takahashi-Ikezawa M, Ueno H, Yoshimoto Y, Yorifuji H (2017) A novel imaging method for correlating 2D light microscopic data and 3D volume data based on block-face imaging. Sci Rep 7:3645
Kimura J, Hirano Y, Takemoto S, Nambo Y, Ishinazaka T, Himeno R, Mishima T, Tsumagari S, Yokota H (2005) Three-dimensional reconstruction of the equine ovary. Anat Histol Embryol 34:48–51
Amato SP, Pan F, Schwartz J, Ragan TM (2016) Whole brain imaging with serial two-photon tomography. Front Neuroanat 10:31
Ragan T, Kadiri LR, Venkataraju KU, Bahlmann K, Sutin J, Taranda J, Arganda-Carreras I, Kim Y, Seung HS, Osten P (2012) Serial two-photon tomography for automated ex vivo mouse brain imaging. Nat Methods 9:255–258
Richardson DS, Lichtman JW (2015) Clarifying tissue clearing. Cell 162:246–257
Kim WH, Adluru N, Chung MK, Okonkwo OC, Johnson SC, B. Bendlin B, Singh V (2015) Multi-resolution statistical analysis of brain connectivity graphs in preclinical Alzheimer’s disease. Neuroimage 118:103–117
Kovačević N, Henderson JT, Chan E, Lifshitz N, Bishop J, Evans AC, Henkelman RM, Chen XJ (2005) A three-dimensional MRI atlas of the mouse brain with estimates of the average and variability. Cereb Cortex 15:639–645
Badea A, Johnson GA (2012) Magnetic resonance microscopy. Anal Cell Pathol (Amst) 35:205–227
Johnson GA, Benveniste H, Black RD, Hedlund LW, Maronpot RR, Smith BR (1993) Histology by magnetic resonance microscopy. Magn Reson Q 9:1–30
Dorr AE, Lerch JP, Spring S, Kabani N, Henkelman RM (2008) High resolution three-dimensional brain atlas using an average magnetic resonance image of 40 adult C57Bl/6J mice. Neuroimage 42:60–69
Ma Y, Hof PR, Grant SC, Blackband SJ, Bennett R, Slatest L, McGuigan MD, Benveniste H (2005) A three-dimensional digital atlas database of the adult C57BL/6J mouse brain by magnetic resonance microscopy. Neuroscience 135:1203–1215
Schwarz AJ, Danckaert A, Reese T, Gozzi A, Paxinos G, Watson C, Merlo-Pich EV, Bifone A (2006) A stereotaxic MRI template set for the rat brain with tissue class distribution maps and co-registered anatomical atlas: application to pharmacological MRI. Neuroimage 32:538–550
Schweinhardt P, Fransson P, Olson L, Spenger C, Andersson JLR (2003) A template for spatial normalisation of MR images of the rat brain. J Neurosci Methods 129:105–113
MacKenzie-Graham A, Lee E-F, Dinov ID, Bota M, Shattuck DW, Ruffins S, Yuan H, Konstantinidis F, Pitiot A, Ding Y, Hu G, Jacobs RE, Toga AW (2004) A multimodal, multidimensional atlas of the C57BL/6J mouse brain. J Anat 204:93–102
Scholz J, LaLiberté C, van Eede M, Lerch JP, Henkelman M (2016) Variability of brain anatomy for three common mouse strains. Neuroimage 142:656–662
Papp EA, Leergaard TB, Calabrese E, Johnson GA, Bjaalie JG (2014) Waxholm space atlas of the Sprague Dawley rat brain. Neuroimage 97:374–386
Johnson GA, Badea A, Brandenburg J, Cofer G, Fubara B, Liu S, Nissanov J (2010) Waxholm space: an image-based reference for coordinating mouse brain research. Neuroimage 53:365–372
Hjornevik T, Leergaard TB, Darine D et al (2007) Three-dimensional atlas system for mouse and rat brain imaging data. Front Neuroinform 1:4
Nie B, Chen K, Zhao S, Liu J, Gu X, Yao Q, Hui J, Zhang Z, Teng G, Zhao C, Shan B (2013) A rat brain MRI template with digital stereotaxic atlas of fine anatomical delineations in paxinos space and its automated application in voxel-wise analysis. Hum Brain Mapp 34:1306–1318
Radtke-Schuller S, Schuller G, Angenstein F, Grosser OS, Goldschmidt J, Budinger E (2016) Brain atlas of the Mongolian gerbil (Meriones unguiculatus) in CT/MRI-aided stereotaxic coordinates. Brain Struct Funct 221:1–272
Kaufman HH, Cohen G, Glass TF, Huchton JD, Pruessner JL, Ostrow PT, Andia-Waltenbaugh AM, Dujovny M (1981) CT atlas of the dog brain. J Comput Assist Tomogr 5:529–537
Datta R, Lee J, Duda J, Avants BB, Vite CH, Tseng B, Gee JC, Aguirre GD, Aguirre GK (2012) A digital atlas of the dog brain. PLoS One 7:e52140
Stolzberg D, Wong C, Butler BE, Lomber SG (2017) Catlas: an magnetic resonance imaging-based three-dimensional cortical atlas and tissue probability maps for the domestic cat (Felis catus). J Comp Neurol 525:3190–3206
Liu C, Ye FQ, Yen CC-C, Newman JD, Glen D, Leopold DA, Silva AC (2018) A digital 3D atlas of the marmoset brain based on multi-modal MRI. Neuroimage 169:106–116
Woodward A, Hashikawa T, Maeda M, Kaneko T, Hikishima K, Iriki A, Okano H, Yamaguchi Y (2018) The brain/MINDS 3D digital marmoset brain atlas. Sci Data 5:180009
Ghosh P, O’Dell M, Narasimhan PT et al (1994) Mouse Lemur microscopic MRI brain atlas. Neuroimage 1:345–349
Hutchinson EB, Schwerin SC, Radomski KL, Sadeghi N, Jenkins J, Komlosh ME, Irfanoglu MO, Juliano SL, Pierpaoli C (2017) Population based MRI and DTI templates of the adult ferret brain and tools for voxelwise analysis. Neuroimage 152:575–589
Toga AW, Thompson PM, Mori S, Amunts K, Zilles K (2006) Towards multimodal atlases of the human brain. Nat Rev Neurosci 7:952–966
Mazziotta JC, Toga AW, Evans A, Fox P, Lancaster J (1995) A probabilistic atlas of the human brain: theory and rationale for its development. The international consortium for brain mapping (ICBM). Neuroimage 2:89–101
Ashburner J, Friston KJ (2000) Voxel-based morphometry—the methods. Neuroimage 11:805–821
Whitwell JL (2009) Voxel-based morphometry: an automated technique for assessing structural changes in the brain. J Neurosci 29:9661–9664
Abbott DF, Pell GS, Pardoe HR, Jackson GD (2012) Selecting appropriate voxel-based methods for neuroimaging studies. Neuroimage 59:885–886
Maguire EA, Gadian DG, Johnsrude IS, Good CD, Ashburner J, Frackowiak RSJ, Frith CD (2000) Navigation-related structural change in the hippocampi of taxi drivers. Proc Natl Acad Sci U S A 97:4398–4403
Maguire EA, Woollett K, Spiers HJ (2006) London taxi drivers and bus drivers: a structural MRI and neuropsychological analysis. Hippocampus 16:1091–1101
May A (2008) Chronic pain may change the structure of the brain. Pain 137:7–15
Matharu MS, Good CD, May A, Bahra A, Goadsby PJ (2003) No change in the structure of the brain in migraine: a voxel-based morphometric study. Eur J Neurol 10:53–57
Rodriguez-Raecke R, Niemeier A, Ihle K, Ruether W, May A (2013) Structural brain changes in chronic pain reflect probably neither damage nor atrophy. PLoS One 8:e54475
Baliki MN, Schnitzer TJ, Bauer WR, Apkarian AV (2011) Brain morphological signatures for chronic pain. PLoS One 6:e26010
Lerch JP, Yiu AP, Martinez-Canabal A, Pekar T, Bohbot VD, Frankland PW, Henkelman RM, Josselyn SA, Sled JG (2011) Maze training in mice induces MRI-detectable brain shape changes specific to the type of learning. Neuroimage 54:2086–2095
Sawiak SJ, Wood NI, Williams GB, Morton AJ, Carpenter TA (2013) Voxel-based morphometry with templates and validation in a mouse model of Huntington’s disease. Magn Reson Imaging 31:1522–1531
Koch S, Mueller S, Foddis M et al (2017) Atlas registration for edema-corrected MRI lesion volume in mouse stroke models. J Cereb Blood Flow Metab. https://doi.org/10.1177/0271678X1772663
Badea A, Nicholls PJ, Johnson GA, Wetsel WC (2007) Neuroanatomical phenotypes in the Reeler mouse. Neuroimage 34:1363–1374
Badea A, Johnson GA, Jankowsky JL (2010) Remote sites of structural atrophy predict later amyloid formation in a mouse model of Alzheimer’s disease. Neuroimage 50:416–427
Badea A, Kane L, Anderson RJ, Qi Y, Foster M, Cofer GP, Medvitz N, Buckley AF, Badea AK, Wetsel WC, Colton CA (2016) The fornix provides multiple biomarkers to characterize circuit disruption in a mouse model of Alzheimer’s disease. Neuroimage 142:498–511
Zitová B, Flusser J (2003) Image registration methods: a survey. Image Vis Comput 21:977–1000
Viergever MA, Maintz JBA, Klein S, Murphy K, Staring M, Pluim JPW (2016) A survey of medical image registration—under review. Med Image Anal 33:140–144
Maintz JB, Viergever MA (1998) A survey of medical image registration. Med Image Anal 2:1–36
Sawiak SJ, Wood NI, Williams GB, et al (2009) SPMMouse: a new toolbox for SPM in the animal brain. Proc Int’l Soc Mag Res Med. p 1086
Hikishima K, Komaki Y, Seki F, Ohnishi Y, Okano HJ, Okano H (2017) In vivo microscopic voxel-based morphometry with a brain template to characterize strain-specific structures in the mouse brain. Sci Rep 7:85
Nitzsche B, Frey S, Collins LD et al (2015) A stereotaxic, population-averaged T1w ovine brain atlas including cerebral morphology and tissue volumes. Front Neuroanat 9:69
Dickie DA, Shenkin SD, Anblagan D et al (2017) Whole brain magnetic resonance image atlases: a systematic review of existing atlases and caveats for use in population imaging. Front Neuroinform 11:1
Glasser MF, Smith SM, Marcus DS, Andersson JLR, Auerbach EJ, Behrens TEJ, Coalson TS, Harms MP, Jenkinson M, Moeller S, Robinson EC, Sotiropoulos SN, Xu J, Yacoub E, Ugurbil K, van Essen DC (2016) The human connectome project’s neuroimaging approach. Nat Neurosci 19:1175–1187
Van Essen DC, Smith SM, Barch DM et al (2013) The WU-Minn human connectome project: an overview. Neuroimage 80:62–79
Hess A, Stiller D, Kaulisch T, Heil P, Scheich H (2000) New insights into the hemodynamic blood oxygenation level-dependent response through combination of functional magnetic resonance imaging and optical recording in gerbil barrel cortex. J Neurosci 20:3328–3338
Ohl FW, Scheich H (1997) Orderly cortical representation of vowels based on formant interaction. Proc Natl Acad Sci U S A 94:9440–9444
Sokoloff L, Reivich M, Kennedy C, Rosiers MHD, Patlak CS, Pettigrew KD, Sakurada O, Shinohara M (1977) The [14C]deoxyglucose method for the measurement of local cerebral glucose utilization: theory, procedure, and normal values in the conscious and anesthetized albino rat. J Neurochem 28:897–916
Gholipour A, Kehtarnavaz N, Briggs R, Devous M, Gopinath K (2007) Brain functional localization: a survey of image registration techniques. IEEE Trans Med Imaging 26:427–451
Eklund A, Nichols TE, Knutsson H (2016) Cluster failure: why fMRI inferences for spatial extent have inflated false-positive rates. Proc Natl Acad Sci U S A 113:7900–7905
Vul E, Harris C, Winkielman P, Pashler H (2009) Puzzlingly high correlations in fMRI studies of emotion, personality, and social cognition. Perspect Psychol Sci 4:274–290
Bates E, Wilson SM, Saygin AP, Dick F, Sereno MI, Knight RT, Dronkers NF (2003) Voxel-based lesion-symptom mapping. Nat Neurosci 6:448–450
Hess A, Axmann R, Rech J, Finzel S, Heindl C, Kreitz S, Sergeeva M, Saake M, Garcia M, Kollias G, Straub RH, Sporns O, Doerfler A, Brune K, Schett G (2011) Blockade of TNF-α rapidly inhibits pain responses in the central nervous system. Proc Natl Acad Sci 108:3731–3736
Gu S, Pasqualetti F, Cieslak M, Telesford QK, Yu AB, Kahn AE, Medaglia JD, Vettel JM, Miller MB, Grafton ST, Bassett DS (2015) Controllability of structural brain networks. Nat Commun 6:8414
Sporns O (2013) Structure and function of complex brain networks. Dialogues Clin Neurosci 15:247–262
Bullmore E, Sporns O (2009) Complex brain networks: graph theoretical analysis of structural and functional systems. Nat Rev Neurosci 10:186–198
Ritter P, Schirner M, McIntosh AR, Jirsa VK (2013) The virtual brain integrates computational modeling and multimodal neuroimaging. Brain Connect 3:121–145
Ganglberger F, Kaczanowska J, Penninger JM, Hess A, Bühler K, Haubensak W (2018) Predicting functional neuroanatomical maps from fusing brain networks with genetic information. Neuroimage 170:113–120
Grandjean J, Zerbi V, Balsters JH, Wenderoth N, Rudin M (2017) Structural basis of large-scale functional connectivity in the mouse. J Neurosci 37:8092–8101
Glasser MF, Coalson TS, Robinson EC, Hacker CD, Harwell J, Yacoub E, Ugurbil K, Andersson J, Beckmann CF, Jenkinson M, Smith SM, van Essen DC (2016) A multi-modal parcellation of human cerebral cortex. Nature 536:171–178
Amunts K, Zilles K (2015) Architectonic mapping of the human brain beyond Brodmann. Neuron 88:1086–1107
Peng H, Chung P, Long F, Qu L, Jenett A, Seeds AM, Myers EW, Simpson JH (2011) BrainAligner: 3D registration atlases of Drosophila brains. Nat Methods 8:493–498
Kurylas AE, Rohlfing T, Krofczik S, Jenett A, Homberg U (2008) Standardized atlas of the brain of the desert locust, Schistocerca gregaria. Cell Tissue Res 333:125–145
Brandt R, Rohlfing T, Rybak J, Krofczik S, Maye A, Westerhoff M, Hege HC, Menzel R (2005) Three-dimensional average-shape atlas of the honeybee brain and its applications. J Comp Neurol 492:1–19
Poirier C, Vellema M, Verhoye M, van Meir V, Wild JM, Balthazart J, van der Linden A (2008) A three-dimensional MRI atlas of the zebra finch brain in stereotaxic coordinates. Neuroimage 41:1–6
De Groof G, George I, Touj S et al (2016) A three-dimensional digital atlas of the starling brain. Brain Struct Funct 221:1899–1909
Vellema M, Verschueren J, Van Meir V, Van der Linden A (2011) A customizable 3-dimensional digital atlas of the canary brain in multiple modalities. Neuroimage 57:352–361
Karten HJ, Brzozowska-Prechtl A, Lovell PV, Tang DD, Mello CV, Wang H, Mitra PP (2013) Digital atlas of the zebra finch (Taeniopygia guttata) brain: a high-resolution photo atlas. J Comp Neurol 521:3702–3715
Güntürkün O, Verhoye M, De Groof G, Van der Linden A (2013) A 3-dimensional digital atlas of the ascending sensory and the descending motor systems in the pigeon brain. Brain Struct Funct 218:269–281
Simões JM, Teles MC, Oliveira RF, van der Linden A, Verhoye M (2012) A three-dimensional stereotaxic MRI brain atlas of the cichlid fish Oreochromis mossambicus. PLoS One 7:e44086
Wulliman MF, Rupp B, Reichert H (2012) Neuroanatomy of the zebrafish brain: a topological atlas. Birkhäuser, Basel
Anken RH, Bourrat F (1998) Brain atlas of the medakafish: Oryzias latipes. Institute National de la Recherche Agronomique, Versailles
Randlett O, Wee CL, Naumann EA, Nnaemeka O, Schoppik D, Fitzgerald JE, Portugues R, Lacoste AMB, Riegler C, Engert F, Schier AF (2015) Whole-brain activity mapping onto a zebrafish brain atlas. Nat Methods 12:1039–1046
D’angelo L (2013) Brain atlas of an emerging Teleostean model: Nothobranchius furzeri. Anat Rec 296:681–691
Maler L, Sas E, Johnston S, Ellis W (1991) An atlas of the brain of the electric fish Apteronotus leptorhynchus. J Chem Neuroanat 4:1–38
Majka P, Chlodzinska N, Turlejski K et al (2017) A three-dimensional stereotaxic atlas of the gray short-tailed opossum (Monodelphis domestica) brain. Brain Struct Funct 223:1779–1795
Bakker R, Tiesinga P, Kötter R (2015) The scalable brain atlas: instant web-based access to public brain atlases and related content. Neuroinformatics 13:353–366
Calabrese E, Badea A, Cofer G, Qi Y, Johnson GA (2015) A diffusion MRI tractography connectome of the mouse brain and comparison with neuronal tracer data. Cereb Cortex 25:4628–4637
Okamura-Oho Y, Shimokawa K, Takemoto S, Hirakiyama A, Nakamura S, Tsujimura Y, Nishimura M, Kasukawa T, Masumoto KH, Nikaido I, Shigeyoshi Y, Ueda HR, Song G, Gee J, Himeno R, Yokota H (2012) Transcriptome tomography for brain analysis in the web-accessible anatomical space. PLoS One 7:e45373
Dorr A, Sled JG, Kabani N (2007) Three-dimensional cerebral vasculature of the CBA mouse brain: a magnetic resonance imaging and micro computed tomography study. Neuroimage 35:1409–1423
Xiong B, Li A, Lou Y, Chen S, Long B, Peng J, Yang Z, Xu T, Yang X, Li X, Jiang T, Luo Q, Gong H (2017) Precise cerebral vascular atlas in stereotaxic coordinates of whole mouse brain. Front Neuroanat 11:128
Joshi R, Yanasak N (2011) Magnetic resonance angiography study of a normal mouse brain for creating a three-dimensional cerebral vasculature atlas and software for labeling vessels. 2011 IEEE International Conference on Bioinformatics and Biomedicine Workshops (BIBMW). pp 966–968
Chen C-C V, Chen Y-C, Hsiao H-Y et al (2013) Neurovascular abnormalities in brain disorders: highlights with angiogenesis and magnetic resonance imaging studies. J Biomed Sci 20:47
Mayerich D, Kwon J, Sung C, Abbott L, Keyser J, Choe Y (2011) Fast macro-scale transmission imaging of microvascular networks using KESM. Biomed Opt Express 2:2888–2896
Wu J, He Y, Yang Z, Guo C, Luo Q, Zhou W, Chen S, Li A, Xiong B, Jiang T, Gong H (2014) 3D BrainCV: simultaneous visualization and analysis of cells and capillaries in a whole mouse brain with one-micron voxel resolution. Neuroimage 87:199–208
Xue S, Gong H, Jiang T, Luo W, Meng Y, Liu Q, Chen S, Li A (2014) Indian-ink perfusion based method for reconstructing continuous vascular networks in whole mouse brain. PLoS One 9:e88067
Liebmann T, Renier N, Bettayeb K, Greengard P, Tessier-Lavigne M, Flajolet M (2016) Three-dimensional study of Alzheimer’s disease hallmarks using the iDISCO clearing method. Cell Rep 16:1138–1152
Lugo-Hernandez E, Squire A, Hagemann N, Brenzel A, Sardari M, Schlechter J, Sanchez-Mendoza EH, Gunzer M, Faissner A, Hermann DM (2017) 3D visualization and quantification of microvessels in the whole ischemic mouse brain using solvent-based clearing and light sheet microscopy. J Cereb Blood Flow Metab 37:3355–3367
Zhang L-Y, Lin P, Pan J, Ma Y, Wei Z, Lu J, Wang L, Song Y, Wang Y, Zhang Z, Jin K, Wang Q, Yang G-Y (2018) Clarity for high-resolution imaging and quantification of vasculature in the whole mouse brain. Aging Dis 9:262–272
Di Giovanna AP, Tibo A, Silvestri L et al (2017) Whole-brain vasculature reconstruction at the single capillary level. bioRxiv
Robertson RT, Levine ST, Haynes SM, Gutierrez P, Baratta JL, Tan Z, Longmuir KJ (2015) Use of labeled tomato lectin for imaging vasculature structures. Histochem Cell Biol 143:225–234
Renier N, Adams EL, Kirst C, Wu Z, Azevedo R, Kohl J, Autry AE, Kadiri L, Umadevi Venkataraju K, Zhou Y, Wang VX, Tang CY, Olsen O, Dulac C, Osten P, Tessier-Lavigne M (2016) Mapping of brain activity by automated volume analysis of immediate early genes. Cell 165:1789–1802
Deffieux T, Demene C, Pernot M, Tanter M (2018) Functional ultrasound neuroimaging: a review of the preclinical and clinical state of the art. Curr Opin Neurobiol 50:128–135
Tiran E, Ferrier J, Deffieux T, Gennisson JL, Pezet S, Lenkei Z, Tanter M (2017) Transcranial functional ultrasound imaging in freely moving awake mice and anesthetized young rats without contrast agent. Ultrasound Med Biol 43:1679–1689
Starosolski Z, Villamizar CA, Rendon D, Paldino MJ, Milewicz DM, Ghaghada KB, Annapragada AV (2015) Ultra high-resolution in vivo computed tomography imaging of mouse cerebrovasculature using a long circulating blood pool contrast agent. Sci Rep 5:10178
Pastor G, Jiménez-González M, Plaza-García S, Beraza M, Padro D, Ramos-Cabrer P, Reese T (2017) A general protocol of ultra-high resolution MR angiography to image the cerebro-vasculature in 6 different rats strains at high field. J Neurosci Methods 289:75–84
Howles GP, Ghaghada KB, Qi Y, Mukundan S Jr, Johnson GA (2009) High-resolution magnetic resonance angiography in the mouse using a nanoparticle blood-pool contrast agent. Magn Reson Med 62:1447–1456
Klohs J, Baltes C, Princz-Kranz F, Ratering D, Nitsch RM, Knuesel I, Rudin M (2012) Contrast-enhanced magnetic resonance microangiography reveals remodeling of the cerebral microvasculature in transgenic ArcAβ mice. J Neurosci 32:1705–1713
Acknowledgements
The writing of this review was initiated by the members of the Molecular Neuroimaging Study Group of the European Society for Molecular Imaging (ESMI). We gratefully thank the ESMI for their support and the possibility of establishing the study group as a platform for scientific exchange within the society and beyond.
Funding
This work is supported by the Deutsche Forschungsgemeinschaft (DFG Cluster of Excellence NeuroCURE, Exc 257 to P.B-S.), the German Federal Ministry of Education and Research (BMBF; 01EO0801, Center for Stroke Research Berlin to P.B-S. and BMBF NeuroRad (02NUK034D to A.H.), BMBF NeuroImpa (01EC1403C) to A.H.), INCF Digital Atlasing program to A.H., the Research Foundation - Flanders (FWO G048917N to R.H. and G.A.K.), and Flagship ERA-NET (FLAG-ERA) FUSIMICE (grant agreement G.0D7651N to R.H. and G.A.K.).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Hess, A., Hinz, R., Keliris, G.A. et al. On the Usage of Brain Atlases in Neuroimaging Research. Mol Imaging Biol 20, 742–749 (2018). https://doi.org/10.1007/s11307-018-1259-y
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11307-018-1259-y