Abstract
Purpose
LRRK2 (leucine-rich repeat kinase 2) has recently been proven to be a promising drug target for Parkinson’s disease (PD) due to an apparent enhanced activity caused by mutations associated with familial PD. To date, there have been no reports in which a LRRK2 inhibitor has been radiolabeled and used for in in vitro or in vivo studies of LRRK2. In the present study, we radiolabeled the LRRK2 ligand, LRRK-IN-1, for the purposes of performing in vitro (IC50, K d , B max, autoradiography) and in vivo (biodistribution, and blocking experiments) evaluations in rodents and human striatum tissues.
Procedures
[3H]LRRK2-IN-1 was prepared with high radiochemical purity (>99 %) and a specific activity of 41 Ci/mmol via tritium/hydrogen (T/H) exchange using Crabtree’s catalyst. For IC50, K d , and B max determination, LRRK2-IN-1 was used as a competing drug for nonspecific binding assessment. The specific binding of the tracer was further evaluated via an in vivo blocking study in mice with a potent LRRK2 inhibitor, Pf-06447475.
Results
In vitro binding studies demonstrated a saturable binding site for [3H]LRRK2-IN-1 in rat kidney, rat brain striatum and human brain striatum with K d of 26 ± 3 and 43 ± 8, 48 ± 2 nM, respectively. In rat, the density of LRRK2 binding sites (B max) was higher in kidney (6.4 ± 0.04 pmol/mg) than in brain (2.5 ± 0.03 pmol/mg), however, in human brain striatum, the B max was 0.73 ± 0.01 pmol/mg protein. Autoradiography imaging in striatum of rat and human brain tissues gave results consistent with binding studies. In in vivo biodistribution and blocking studies in mice, co-administration with Pf-06447475 (10 mg/kg) reduced the uptake of [3H]LRRK2-IN-1 (%ID/g) by 50–60% in the kidney or brain.
Conclusion
The high LRRK2 brain density observed in our study suggests the feasibility for positron emission tomography imaging of LRRK2 (a potential target) with radioligands of higher affinity and specificity.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
LRRK2 (leucine-rich repeat kinase 2) enzyme is a type of ROCO-serine/threonine specific kinase proteins. It shows of two distinct enzymatic functions, i.e., GTPase and kinase. The enzymatic core of LRRK2 comprises of two main proportions which include Roc-COR domain (Ras-like guanine nucleotide binding domain-C-terminal of Roc) and catalytic kinase domain (Fig. 1) [1]. Additionally, it has four other domains: armadillo repeats (ARM), ankyrin repeats (ANK), leucine-rich repeats (LRR), and WD40 domain (hydrophilic domain) (Fig. 1) [2]. The genetic mutations G2019S (Gly2019Ser) and I2020T in the catalytic kinase domain lead to either increased or slightly decreased kinase activity, respectively [3–8]. However, the mutations in G2019S account as a major factor for late-onset Parkinson’s disease (PD) [9–15]. In addition, several other mutations in LRRK2 such as in LRR (I112V), Roc (N1437H, R1441C/G/H), and COR (Y1699C) are also associated with genetic causes of PD [12, 13]. Since mutation-induced increases in LRRK2 activity are associated with PD, inhibiting the activity of this enzyme has been investigated as a therapeutic approach to delaying the onset of this disease. Potential mechanisms for inhibition of LRRK2 activity include obstruction of GTP binding (Roc, essential for protein kinase activity), dimerization inhibition (COR), blockage of protein-protein interactions (ARM, ANK, and WD), stabilization of 14-3-3 interaction (LRR), and direct kinase inhibition (kinase) (Fig. 1) [16–22].
To date, many LRRK2 kinase inhibitors have been developed and evaluated in vitro [23–32]. Most of these kinase inhibitors compete with ATP for ATP-binding site by targeting either by active (DFG-in) or inactive conformation (DFG-out) [26]. LRRK2-IN-1 was one of the first inhibitors and has shown potent inhibition of both wild-type (WT) and G2019S mutant enzyme [22, 23]. By structure-activity relationship study [33, 34], it was evident that the pyrimidyl moiety along with the benzodiazepine (benzene ring without substituent) in LRRK2-IN-1 is the effective motif for binding in the ATP pocket (Fig. 2) [24, 32]. These inhibitors cause dephosphorylation in LRRK2 kinase (Ser 910/Ser 935) [35, 36].
Due to the clinical significance of LRRK2 in PD, the availability of positron emission tomography (PET) imaging agents will provide a non-invasive means both to examine the role of LRRK2 in PD and to determine target engagement of new LRRK2 inhibitors. Despite research efforts invested to date, there is no radiotracer available for in vivo imaging of LRRK2 using PET. Similarly, there has been no report of a labeled LRRK2 inhibitor for in vitro studies. In the present study, we synthesized and evaluated the binding affinity of the H-3 labeled LRRK2 inhibitor, LRRK2-IN-1, in in vitro competition assays and saturation studies and autoradiography (ARG) in rat kidney, rat brain, and in human brain tissues. The specific binding of the tracer to LRRK2 was further evaluated in biodistribution studies in mice, both alone and with blockade with a potent LRRK2 kinase inhibitor, Pf-06447475 (3-[4-(morpholin-4-yl)-7H–pyrrolo[2,3-d]pyrimidin-5-yl] benzonitrile, IC50 = 3 nM) [30, 31].
Materials and Methods
General
All chemicals and solvents were of biological grade with high purity (≥98 %). LRRK2-IN-1 (reference standard) and Pf-06447475 were purchased from Tocris. [3H]LRRK2-IN-1 was prepared by Novandi (collaborator, synthesis procedures are included). The HPLC (Dionex GP40; UV detector: Knauer K-2500; column: Phenomenex Luna, 150 × 4.7 mm, 5 μ; scintillation counter as a detector to differentiate radioactive and non-radioactive fractions: Ludlum) was used to check the purity of radioligands. For measurements of H-3 levels in samples, a liquid scintillation counter (Packard, Tri carb, 2900) was used whereas for binding assays, filters were counted using a Wallac liquid scintillation and luminescence counter (Microbeta 1450-021 Trilux, Perkin Elmer). For protein analyses, absorbance was measured using spectrometer plate reader (ELX-808, Bio-Tek). 96-well plates were incubated in micro-plate shaker (VWR). For ARG studies, films were scanned in phosphor imager (Fujifilm, FLA-7000). All animals (mice and rats) were purchased from Charles River (Charles River Laboratories International, Inc., USA). Human brain striatum (healthy brain) was purchased from Analytical Biological Services Inc. (Wilmington, DE, USA). The sectioning of tissues was performed using a Lieca Microm HM 560 cryostat. All animal procedures were performed in accordance with the National Institutes of Health Animal Care Guidelines.
Radiochemistry
Synthesis of [3H]LRRK2-IN-1
[3H]LRRK2-IN-1 was labeled by tritium/hydrogen (T/H) exchange with an organoiridium catalyst (Crabtree’s catalyst) in dichloromethane (DCM). Purification was performed on reversed phase HPLC which gave [3H]LRRK2-IN-1 with a radiochemical purity of 99 % and with a specific activity of 41 Ci/mmol (1.51 TBq/mmol). The concentration of a radiotracer stock solution for further experiments was 33.7 MBq/ml in ethanol (EtOH).
HPLC Analysis
To verify the identity and radiochemical purity of [3H]LRRK2-IN-1, 20 μ of a mixture of standard and radiolabeled LRRK2-IN-1 were co-injected into HPLC. The HPLC analysis was performed using gradient elution by acetonitrile (MeCN; 20–60 %) with 0.1 % of ammonium formate (pH: 6) using a reverse phase C18 analytical column (4.6 mm × 250 mm, Phenomenex) at a flow rate of 1 ml/min for 20 min. The fractions were collected throughout analysis and were subsequently measured with liquid scintillation counter (LSC). Radioactivity co-eluted with the UV peak with a retention time of 7 min.
Biological Experiments
Buffer Preparation
For binding and autoradiographic assays, the Tris-buffer composition was as follows: 50 mM Tris, 5 mM MgCl2, 0.5 mM EDTA, and pH 7.4.
Preparation of Membranes
The samples of tissues (rat kidney, rat brain striatum, and human brain striatum) (rats: Sprague Dawley, males, avg. weight: 250 g, avg. age: 10 weeks) were prepared for competition assays (IC50) and saturation studies (K d , B max). The frozen tissues (1 g, −80 °C) were homogenized in 12 ml of ice-cold Tris-buffer containing protease inhibitor cocktail (0.1 ml; Sigma) using a polytron and were centrifuged at 800×g for 3–4 min. Afterwards, supernatants were centrifuged at 20,000×g for 15 min. The supernatants were discarded and the pellets were re-suspended in Tris-buffer (without protease) and centrifuged again. After a third spin, the pellets were re-suspended in cryomedium (10 % sucrose in Tris-buffer; ca. 500 μl). The samples for human brain striatum (protein: 2.65 ng/ml) were prepared in a similar way. An aliquot from the final resuspensions was analyzed for protein using a BCA protein kit (Pierce) together with bovine serum albumin (BSA) protein standards and measured for absorbance using a spectrometer.
Drug Preparation
For competition and autoradiography assays, the [3H]LRRK2-IN-1 solution (in EtOH) was evaporated to dryness under argon and dissolved in the final assay buffer. The final concentration of [3H]LRRK2-IN-1 used for competition and autoradiography assays was 10 nM. For saturation assays, the final [3H]LRRK2-IN-1 concentration ranged from 5 to 140 nM. For non-specific binding, LRRK2-IN-1 standard drug solutions were prepared by dissolving the drug in DMSO (2 mg/ml, stock solution) and then diluting further using assay buffer to the final working concentration. Final assay concentrations were 0 to 3000 nM for binding assays and 0 to 1000 nM for autoradiographic assays. Non-specific binding was defined by using 10 μM (LRRK2-IN-1, per well) for competition assays, and as 1 μM (LRRK2-IN-1, per slide) for autoradiographic assays.
Filtration Binding
One hundred fifty microliters of tissue samples (90–110 μg protein/well), 50 μL of LRRK2-IN-1 (various concentrations), and 50 μl of radioligand ([3H]LRRK2-IN-1) were added to each well of a 96-well microtiter plate. The plate was incubated at 30 °C with continuous rotation for 1 h. The incubation was stopped by vacuum filtration onto 0.3 % PEI pre-soaked GF/C filters using a 96-well FilterMate™ harvester, followed by four washes with ice-cold wash buffer. Filters were then dried for 30 min under warm air flow. The filter was sealed in polyethylene, scintillation cocktail (Betaplate Scint; PerkinElmer) added, and the radioactivity counted in a LSC (Wallac).
In Vitro Autoradiography
In vitro ARG studies were performed with rat brain striatum, cortex, kidney, and human brain striatum tissues. For harvesting tissues, rats (Sprague Dawley, avg. weight: 250 g, avg. age: 10 weeks) were sacrificed and tissues were taken out and frozen immediately with cold hexane (−80 °C) in dry ice. Afterwards, the tissues were sectioned at a 20 μm thickness using a cryostat and stored at −80 °C until further use.
For autoradiography, the slides were allowed to warm to room temperature (RT) and then, pre-incubated in assay buffer for 10 min. Following this step, the slides were placed in a humidified box and 1 ml of a solution of the unlabeled compound (for non-specific) and radiotracer (total) in buffer loaded onto each section. The sections were incubated in this solution for 30 min at RT. The slides were then washed in ice-cold buffer (two changes) for 6 min and finally dried under a stream of warm air for 10 min. Once dried, the slides were placed on a tritium-sensitive phosphor screen in a light-proof cassette together with autoradiographic standards. The latter were prepared by spotting known quantities of radioactivity (measured using LSC) onto spare tissue sections. All slides were then exposed to the screens for 3–4 days, after which the screens were scanned in a phosphor imager. In addition, slides were subsequently stained by cresyl violet Nissl staining using a standard procedure to correlate the radioactivity images with histology.
The regions of interest (ROI) were drawn around tissue specimen and mean radioactivity values (in PSL units) were calculated using ImageJ. The PSL/mm3 values were converted to DPM/mm3 by reference to the autoradiographic standards.
In Vivo Biodistribution and Blocking Studies
For biodistibution study, albino BALB/c mice (female, avg. weight: 35 g; age: 2–3 months; n = 4 for each group) were used. [3H]LRRK2-IN-1 (ca. 1 μCi, 14 ng) together with different doses of Pf-06447475 (0, 1, 3, 10 mg/kg) were co-administered via tail vein injection. Mice were sacrificed at 45 min post-injection and tissues (kidney, spleen, lung, heart, liver, muscle, and brain) were collected into pre-weighed LSC-vials. The blood samples were also collected in plasma separator tubes and were centrifuged for plasma separation. One milliliter of 0.2 M NaOH was added and the vials were placed on shaker at 50 °C for a day to digest the tissue. Three hundred microliters of 1 M HCl were then added, followed by 100 μl of Tris-buffer (pH: 7.4). For decolorization, 150 μl H2O2 were added in each vial. All LSC-vials were vortexed and left at RT for 30 min. Finally, leekofluor was added to each vial and the vials were placed in a LSC for counting.
Data Analysis
For binding experiments, specific binding (CPM) was obtained by subtracting the non-specific binding from total binding. In hot saturation assays, the specific binding was converted from CPM to fmol/well from the specific activity of the tracer. The specific binding values for the competition and saturation binding assays were plotted and analyzed with GraphPrism 7.0 using the variable slope nonlinear regression to obtain IC50, K d , and B max. The P values (P < 0.05) for significance were calculated by one-way ANOVA (Dunnett’s test) using the statistical routines in GraphPrism 7.0 (Table 1). All the data has been expressed as ±SEM.
Results
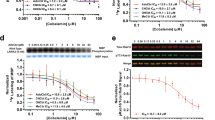
In the radioligand binding assays, specific binding of [3H]LRRK2-IN-1 was observed both in the rat tissues (kidney and brain) and in the post-mortem human tissues (brain). The binding was displaced by unlabeled LRRK2-IN-1 with an IC50 values of 40 ± 4 in rat kidney, 65 ± 3 in rat brain striatum, and 73 ± 6 nM in human brain striatum tissues (Fig. 3a, b, c).
In hot saturation assays using [3H]LRRK2-IN-1, B max values (pmol/mg protein) of 6.4 ± 0.04, 2.5 ± 0.03, 0.73 ± 0.01, and K d values (nM) of 26 ± 3, 43 ± 8, 48 ± 2 were obtained for rat kidney, rat striatum, and human caudate-putamen (striatum), respectively (Fig. 4a, b, c). The fraction of non-specific binding, as a percentage of total binding, was 10 % for rat kidney, 35 % for rat brain striatum, and 70 % for human brain striatum.
In addition to above assays, autoradiographic imaging with [3H]LRRK2-IN-1 was performed in rat kidney and brain striatum and in human brain striatum tissues. For better resolution of images, different slides, e.g., gelatin subbed, regular, and silanized were examined in order to minimize the interactions between radiotracer and slide surface. The best results with lowest background binding of the tracer to the slide were obtained with silanized slides. The images of the tissue sections are shown in Fig. 5a. Strong binding of [3H]LRRK2-IN-1 was observed in the kidney tissue sections that was both regionally heterogeneous and was displaced by the unlabeled LRRK2-IN-1, indicating specificity for LRRK2. In the rat brain sections, cut at the level of the striatum, the specific binding was lower than kidney (Fig. 5b). In the brain sections, there was evidence of a slightly higher level of binding in the rat brain striatum than in cortex. However, this difference did not reach significance. As in the rat brain, human caudate-putamen (striatum) similarly showed modest levels of [3H]LRRK2-IN-1 binding that was displaced by the unlabeled compound.
a Autoradiographic images of [3H]LRRK2-IN-1 binding in rat kidney, rat brain striatum, and human brain striatum. The same sections are shown with both a colored scale (left) and a gray-scale (right). The corresponding cresyl violet-stained sections are shown on the far right; b mean values (DPM/mm3) for the specific binding of [3H]LRRK2-IN-1 in rat kidney (RK), rat brain striatum (RBS), rat brain cortex (RBC), and human brain striatum (HBS, caudate-putamen) tissues.
The biodistribution of [3H]LRRK2-IN-1 was performed in mice with tail vein administration of the tracer. The following rank order of tracer uptake (%ID/g) was observed: kidney (28.5 ± 0.8) > liver (13 ± 0.9) > lung (4.7 ± 0.5) > spleen (3.4 ± 0.3) ≅ heart (3.2 ± 1) > muscle (1.8 ± 0) > plasma (1.2 ± 0.1) > brain (0.39 ± 0.1) (Fig. 6). To determine if uptake represented the binding to LRRK2, blocking studies were performed in which mice were co-administered the LRRK2 inhibitor, Pf-06447475 (Pf), together with the [3H]LRRK2-IN-1 at doses of 1, 3, and 10 mg/kg. In the presence of the blocker, [3H]LRRK2-IN-1 uptake showed a dose-dependent inhibition in all tissues. The inhibition was the highest at 10 mg/kg Pf-06447475 by reducing the uptake of [3H]LRRK2-IN-1 by 54 % in kidney and 62 % in brain (Table 1). In other peripheral tissues: lung (78 %) > heart (72 %) > muscle (62 %) > kidney (54 %) > spleen (40 %) > liver (34 %).
Based on Dunnette’s test, 10 mg/kg was significantly different from vehicle (except for heart) and the levels of significance are: kidney (0.0001, ***), spleen (0.0209, *), brain (0.0238, *), muscle (0.0001, ****), plasma (0.006, **), heart (0.1235, ns), lung (0.0001, ****), and liver (0.0007, ***).
Discussion
Literature reports indicate that LRRK2 related mutations in G2019S are correlated with familial late-onset Parkinson’s. This suggests that targeting LRRK2 enzyme with kinase inhibitors would be a promising approach for clinical diagnostics [3–15]. A PET ligand for imaging LRRK2 in vivo would be of much value in enabling the brain LRRK2 occupancy of these novel inhibitors to be quantified and correlating this occupancy with dose or plasma levels of drug. Thus, an in vivo imaging biomarker would be critical both for drug development studies and as an outcome measure in clinical trials.
Among the list of LRRK2 ligands, LRRK2-IN-1 stands as the first one which has been reported as a potent and selective kinase inhibitor. It has shown very high affinity with both WT (IC50 = 13 nM) and mutant G2019S type (IC50 = 6 nM) LRRK2 [24]. However, to date, there has been no report in which a LRRK2 inhibitor has been radiolabeled for use as a research tool for in vitro or in vivo studies of LRRK2. In present work, [3H]LRRK2-IN-1 was synthesized and used for binding assays, autoradiographic imaging in rats and human tissues, and for in vivo biodistribution and blocking studies in mice.
With respect to the relative density and expression of LRRK2 enzyme in different brain regions, many studies in rodents have reported that brain striatum and cortex have the highest expression of mRNA-LRRK2 [37–41], we thus focused on rat brain striatum tissue in comparison with human caudate-putamen (striatum) tissue. In addition to brain tissues, the peripheral tissues have also been reported to express the enzyme, with especially high levels in the kidney [40, 42–47]. Our results also showed a significantly higher specific binding for the tritiated LRRK2 tracer in the rat kidney. However, moderate levels of specific binding were also observed in the rat brain striatum tissue. The IC50 values in these tissues for displacement of [3H]LRRK2-IN-1 by unlabeled LRRK2-IN-1 were similar or slightly higher to the literature reported value IC50 for LRRK2-IN-1 (13 nM) inhibition of enzyme activity for WT (wild-type) LRRK2.
The saturation analysis in the present study indicated a high density of binding sites for LRRK2 in the kidney and lower, although still substantial, density of binding sites in both the rat brain striatum and in post-mortem human caudate-putamen (striatum). Based on the results from our current study, the non-specific binding fraction in human striatum tissue is lower than that in rat brain striatum which is consistent with the lower B max value in the human tissue. This lower B max may be due to either the species difference or the confounding effects of post-mortem delays, which is one of the limitations of the study.
The value of 0.73 pmol/mg protein for B max obtained in the human striatum in the current study is relatively high compared to that observed for common brain receptors that have been successfully imaged using PET. For example, in human brain, common G-protein coupled receptors such as D2 dopamine receptors (B max = 0.2–0.4 pmol/mg protein in striatum), mu opoid receptors (B max = 0.055 pmol/mg protein in caudate-putamen), serotonin receptors (5HT1A, B max = 0.15 pmol/mg protein in cortex), and nicotinic acetylcholine receptors, or nAChRs (α4β2, B max = 0.05 pmol/mg in striatum) have been the potential targets for neuroimaging by PET since decades [48–53].
To qualify as a tracer suitable for PET, a radioligand must show high affinity (K d ) to the target together with a low level of non-specific binding. Additionally, for the target, a sufficient density (B max) is necessary for PET imaging. For PET imaging, a suitable radiotracer should possess a binding potential (B max/K d ) greater than 10 to give a strong signal. The present study indicates that the density of LRRK2 enzyme (B max) in human caudate-putamen (striatum) is sufficient for PET imaging. Using the estimated value of 73 nM for B max of LRRK2 in the brain, and K d = 48 nM for of LRRK2-IN-1, the binding potential of LRRK2-IN-1 in human brain striatum was low (1.5) compared to the normally accepted value for an ideal PET tracer (>10). This suggests that despite the high density of binding sites for LRRK2 in the brain, LRRK2-IN-1 itself may not be suitable as a tracer for PET imaging.
Notably, the ratios specific binding values (DPM) of HBS/RK (0.14) and RBS/RK (0.30) (human brain striatum: HBS, rat brain striatum: RBS, rat kidney: RK) from in vitro ARG studies were similar to the ratios of B max values (HBS/RK: 0.11, RBS/RK: 0.39; target density). Hence, the results were consistent between the homogenate binding assays and the autoradiographic imaging with respect to the relative levels of radiotracer binding between tissues.
Though the binding studies were conducted using rat tissue, the in vivo biodistribution studies were performed using mice. This was due to the relative ease of injecting mice via a tail vein for radiotracer administration, combined with considerations of reduced radiotracer consumption stemming from their smaller size. In the mice in vivo biodistribution studies, a high uptake was observed in kidney but a low uptake in brain striatum. The former is consistent with the high enzyme activity of LRRK2 reported in the literature in the kidney [40, 42–47]. Similarly, the rank order of uptake in other peripheral tissues such as spleen, lung, and heart approximately paralleled that of literature reports of expression levels of LRRK2 mRNA for enzyme activity [40, 42, 43]. An exception was the [3H]LRRK2-IN-1 accumulation in the liver, which was higher than may have been predicted from relative expression levels of LRRK2 mRNA reported in the literature [37, 40]. This likely stems from metabolism of the tracer in this organ. In the presence of the blocking agent (Pf-06447475), the uptake of [3H]LRRK2-IN-1 in the tissues was blocked either partially or more strongly in lung and heart, strongly suggesting the in vivo specific binding of the tracer to LRRK2.
The development of potential radioligands for neuroimaging of brain LRRK2 with PET depends on the capability of radioligands to cross the blood-brain barrier (BBB). It has been suggested that the ratio of the uptake in brain to plasma can provide an initial guidance for brain permeability. If the ratio less than 0.1, the tracer is considered unable to cross the BBB. When this ratio lies between 0.3–0.5, partial access from the BBB would be possible, while crossing of the BBB would be achieved readily if the ratio is greater than 1 [51]. In our work, a low uptake of [3H]LRRK2-IN-1 was observed in brain which was apparent both from the percentage %ID/g values and from the brain to plasma radioactivity ratio (avg. values derived from four-dose blocking studies). The ratio of [3H]LRR2-IN-1 in brain versus blood was 0.33 with saline alone, and 0.28 with 1 mg/kg, 0.22 with 3 mg/kg, and 0.23 with 10 mg/kg of Pf-06447475. These results suggest a poor brain permeability of [3H]LRRK2-IN-1, which is consistent with reports in the literature [24]. In addition to poor BBB permeability of [3H]LRRK2-IN-1, the low brain uptake could have also potentially resulted from a fast peripheral metabolism of the tracer.
Almost all kinase inhibitors in general bind to the ATP-binding pocket and thus competitively inhibit these enzymes [24]. Similarly, [3H]LRRK2-IN-1 binds to the same site as ATP does [24]. The estimates for intracellular concentration of ATP in cells are generally between 1 and 10 mM [54]. This is considerably higher than the Km of LRRK2 for ATP which is in the mid-micromolar range [55], suggesting that the ATP site will be mostly saturated by ATP under normal conditions. As such, the binding of a tracer level of [3H]LRRK2-IN-1 will be lowered by competition with endogenous ATP. Nonetheless, despite the high intracellular levels of ATP we were able to observe binding of [3H]LRRK2-IN-1 in vivo, especially in the periphery, suggesting that the ATP-binding site is not fully saturated by ATP in vivo. Although, the endogenous ATP will largely out-compete binding of a reversible inhibitor to the ATP-binding site on the enzyme, it could be speculated that an irreversible inhibitor would be less susceptible to such competition and thus ultimately be more effective as strategy for a PET tracer for this target [56, 57].
Conclusion
In conclusion, our studies suggest that LRRK2 binding sites are present at high levels in both peripheral tissues (kidney) and brain and this enzyme may hence, serve as a suitable target for PET studies. Additionally, our studies indicated that [3H]LRRK2-IN-1 is able to specifically label LRRK2 binding sites in vitro and thus can be used as a research tool for radioligand binding assays and autoradiographic imaging in tissue sections. Furthermore, our in vivo mice biodistribution studies have provided evidence that [3H]LRRK2-IN-1 is able to bind to and label the LRRK2 enzyme both peripherally and in the brain. However, its low level of brain uptake together with its poor B max/K d ratio suggests that this tracer itself may not be suitable as a PET ligand for brain imaging of LRRK2.
Change history
01 May 2017
An erratum to this article has been published.
References
Bosgraaf L, van Haastert PJ (2003) Roc, a Ras/GTPase domain in complex proteins. Biochim Biophys Acta 1643:5–10
Gilsbach BK, Kortholt A (2014) Structural biology of the LRRK2 GTPase and kinase domains: implications for regulation. Front Mol Neurosci 7:32
West AB, Moore DJ, Biskup S et al (2005) Parkinson’s disease-associated mutations in leucine-rich repeat kinase 2 augment kinase activity. Proc Natl Acad Sci U S A 102:16842–16847
Smith WW, Pei Z, Jiang H et al (2006) Kinase activity of mutant LRRK2 mediates neuronal toxicity. Nat Neurosci 9:1231–1233
Bonifati V (2006) The LRRK2-G2019S mutation: opening a novel era in Parkinson’s disease genetics. Eur J Hum Genet 14:1061–1062
Luzón-Toro B, Rubio de la Torre E, Delgado A (2007) Mechanistic insight into the dominant mode of the Parkinson’s disease-associated G2019S LRRK2 mutation. Hum Mol Genet 16:2031–2039
Jaleel M, Nichols RJ, Deak M et al (2007) LRRK2 phosphorylates moesin at threonine-558: characterization of how Parkinson’s disease mutants affect kinase activity. Biochem J 405:307–317
Anand VS, Reichling LJ, Lipinski K et al (2009) Investigation of leucine-rich repeat kinase 2: enzymological properties and novel assays. FEBS J 276:466–478
Paisan-Ruiz C, Jain S, Evans EW et al (2004) Cloning of the gene containing mutations that cause PARK8-linked Parkinson’s disease. Neuron 44:595–600
Zimprich A, Biskup S, Leitner P et al (2004) Mutations in LRRK2 cause autosomal-dominant parkinsonism with pleomorphic pathology. Neuron 44:601–607
Adams JR, van Netten H, Schulzer M et al (2005) PET in LRRK2 mutations: comparison to sporadic Parkinson’s disease and evidence for presymptomatic compensation. Brain 128(Pt 12):2777–2785
Guo L, Gandhi PN, Wang W et al (2007) The Parkinson’s disease-associated protein, leucine-rich repeat kinase 2 (LRRK2), is an authentic GTPase that stimulates kinase activity. Exp Cell Res 313:3658–3670
Cookson MR, Bandmann O (2010) Parkinson’s disease: insights from pathways. Hum Mol Genet 19:R21–R27
Cookson MR (2010) The role of leucine-rich repeat kinase 2 (LRRK2) in Parkinson’s disease. Nat Rev Neurosci 11:791–797
Daechsel JC, Farrer MJ (2010) LRRK2 and Parkinson disease. Arch Neurol 67:542–547
Lewis PA, Greggio E, Beilina A et al (2007) The R1441C mutation of LRRK2 disrupts GTP hydrolysis. Biochem Biophys Res Commun 357:668–671
Sen S, Webber PJ, West AB (2009) Dependence of leucine-rich repeat kinase 2 (LRRK2) kinase activity on dimerization. J Biol Chem 284:36346–36356
Berger Z, Smith KA, LaVoie MJ (2010) Membrane localization of LRRK2 is associated with increased formation of the highly active LRRK2 dimer and changes in its phosphorylation. Biochemist 49:5511–5523
Nichols RJ, Dzamko N, Morrice NA et al (2010) 14-3-3 binding to LRRK2 is disrupted by multiple Parkinson’s disease-associated mutations and regulates cytoplasmic localization. Biochem J 430:393–404
Civiero L, Vancraenenbroeck R, Belluzzi E et al (2012) Biochemical characterization of highly purified leucine-rich repeat kinases 1 and 2 demonstrates formation of homodimers. PLoS One 7:e43472
James NG, Digman MA, Gratton E et al (2012) Number and brightness analysis of LRRK2 oligomerization in live cells. Biophys J 102:L41–L43
Bahnassawy L, Nicklas S, Palm T et al (2013) The Parkinson’s disease-associated LRRK2 mutation R1441G inhibits neuronal differentiation of neural stem cells. Stem Cells Dev 22:2487–2496
Zhang J, Yang PL, Gray NS (2009) Targeting cancer with small molecule kinase inhibitors. Nat Rev Cancer 9:28–39
Deng X, Dzamko N, Prescott A et al (2011) Characterization of a selective inhibitor of the Parkinson’s disease kinase LRRK2. Nat Chem Biol 7:203–205
Reith AD, Bamborough P, Jandu K et al (2012) GSK2578215A; a potent and highly selective 2-arylmethyloxy-5-substitutent-N-arylbenzamide LRRK2 kinase inhibitor. Bioorg Med Chem Lett 22:5625–5629
Estrada AA, Liu X, Baker-Glenn C et al (2012) Discovery of highly potent, selective, and brain-penetrable leucine-rich repeat kinase 2 (LRRK2) small molecule inhibitors. J Med Chem 55:9416–9433
Kavanagh ME, Doddareddy MR, Kassiou M (2013) The development of CNS-active LRRK2 inhibitors using property-directed optimisation. Bioorg Med Chem Lett 23:3690–3696
Davies P, Hinkle KM, Sukar NN et al (2013) Comprehensive characterization and optimization of anti-LRRK2 (leucine-rich repeat kinase 2) monoclonal antibodies. Biochem J 453:101–113
Estrada AA, Chan BK, Baker-Glenn C et al (2014) Discovery of highly potent, selective, and brain-penetrant aminopyrazole leucine-rich repeat kinase 2 (LRRK2) small molecule inhibitors. J Med Chem 57:921–936
Henderson JL, Kormos BL, Hayward MM et al (2015) Discovery and preclinical profiling of 3-[4-(morpholin-4-yl)-7H-pyrrolo[2, 3-d]pyrimidin-5-yl] benzonitrile (Pf-06447475), a highly potent, selective, brain penetrant, and in vivo active LRRK2 kinase inhibitor. J Med Chem 58:419–432
Daher JP, Abdelmotilib HA, Hu X et al (2015) Leucine-rich repeat kinase 2 (LRRK2) pharmacological inhibition abates α-synuclein gene-induced neurodegeneration. J Biol Chem 290:19433–19444
Koshibu K, van Asperen J, Gerets H et al (2015) Alternative to LRRK2-IN-1 for pharmacological studies of Parkinson’s disease. Pharmacology 96:240–247
Liu Z, Galemmo RA Jr, Fraser KB et al (2014) Unique functional and structural properties of the LRRK2 protein ATP-binding pocket. J Biol Chem 289:32937–32951
Gray NS, Waller D, Choi HG et al (2014) Pyrimido-diazepinone compounds and methods of treating disorders. Patent: WO 2014145909:A3
Dzamko N, Deak M, Hentati F et al (2010) Inhibition of LRRK2 kinase activity leads to dephosphorylation of Ser(910)/Ser(935), disruption of 14-3-3 binding and altered cytoplasmic localization. Biochem J 430:405–413
Taylor SS, Kornev AP (2011) Protein kinases: evolution of dynamic regulatory proteins. Trends Biochem Sci 36:65–77
Giesert F, Hofmann A, Buerger A et al (2013) Expression analysis of Lrrk1, Lrrk2 and Lrrk2 splice variants in mice. PLoS One 8:e63778
West AB, Cowell RM, Daher JP et al (2014) Differential LRRK2 expression in the cortex, striatum, and substantia nigra in transgenic and nontransgenic rodents. J Comp Neurol 522:2465–2480
Dorval V, Mandemakers W, Jolivette F et al (2014) Gene and MicroRNA transcriptome analysis of Parkinson’s related LRRK2 mouse models. PLoS One 9:e85510
Maekawa T, Kubo M, Yokoyama I et al (2010) Age-dependent and cell-population-restricted LRRK2 expression in normal mouse spleen. Biochem Biophys Res Commun 392:431–435
Taymans JM, Van den Haute C, Baekelandt V (2006) Distribution of PINK1 and LRRK2 in rat and mouse brain. J Neurochem 98:951–961
Miklossy J, Arai T, Guo JP et al (2006) LRRK2 expression in normal and pathologic human brain and in human cell lines. J Neuropathol Exp Neurol 65:953–963
Westerlund M, Belin AC, Anvret A et al (2008) Developmental regulation of leucine-rich repeat kinase 1 and 2 expression in the brain and other rodent and human organs: implications for Parkinson’s disease. Neuroscience 152:429–436
Herzig MC, Kolly C, Persohn E et al (2010) LRRK2 protein levels are determined by kinase function and are crucial for kidney and lung homeostasis in mice. Hum Mol Genet 20:4209–4223
Baptista MA, Dave KD, Frasier MA et al (2012) Loss of leucine-rich repeat kinase 2 (LRRK2) in rats leads to progressive abnormal phenotypes in peripheral organs. PLoS One 8:e80705
Tong Y, Giaime E, Yamaguchi H et al (2012) Loss of leucine-rich repeat kinase 2 causes age-dependent bi-phasic alterations of the autophagy pathway. Mol Neurodegener 7:2
Fell MJ, Mirescu C, Basu K et al (2015) MLi-2, a potent, selective, and centrally active compound for exploring the therapeutic potential and safety of LRRK2 kinase inhibition. J Pharmacol Exp Ther 355:397–409
Joyce JN, Sapp DW, Marshall JF (1986) Human striatal dopamine receptors are organized in compartments. Proc Nat Acad Sci USA 83:8002–8006
Hall H, Sedvall G, Magnusson O et al (1994) Distribution of D1- and D2-dopamine receptors, and dopamine and its metabolites in the human brain. Neuropsychopharmacol 11:245–256
Quirion R, Pilapil C, Allaoua H et al (1995) Autoradiographic distribution of multiple opioid, sigma and phencyclidine receptor binding sites in the human brain. In: Biegon A, Volkow ND (eds) ISBN: 084937653XSites of drug action in the human brain, pp 117–141
Kulkarni AD, Patel HM, Surana SJ et al (2016) Brain-blood ratio: implications in brain drug delivery. Expert Opin Drug Deliv 13:85–92
Convents A, De Keyser J, De Backer JP et al (1989) [3H] rauwolscine labels alpha 2-adrenoceptors and 5-HT1A receptors in human cerebral cortex. Eur J Pharmacol 159(3):307–310
Durany N, Zoechling R, Boissl KW et al (2000) Human post-mortem striatal α4β2 nicotinic acetylcholine receptor density in schizophrenia and Parkinson's syndrome. Neurosci Lett 287:109–112
Traut TW (1994) Physiological concentrations of purines and pyrimidines. Mol Cell Biochem 140:1–22
Thermofisher Tools and protocols on kinase basic module—theory: https://tools.thermofisher.com/content/sfs/manuals/LRRK2_G2019S_LanthaScreen_Activity.pdf
Hicks JW, VanBrocklin HF, Wilson AA et al (2010) Radiolabeled small molecule protein kinase inhibitors for imaging with PET or SPECT. Molecules 15:8260–8278
Sun J, Cai L, Zhang K et al (2014) A pilot study on EGFR-targeted molecular imaging of PET/CT with 11C-PD153035 in human gliomas. Clin Nucl Med 39:e20–e26
Acknowledgements
The work is supported by the Center for Advanced Imaging Innovation and Research (CAI2R, www.cai2r.net) at New York University School of Medicine is supported by NIH/NIBIB grant number P41 EB017183. We would like to acknowledge the expert assistance of Mr. Yianni Piyis (METIS Laboratories) for help with the sectioning.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Additional information
An erratum to this article is available at https://doi.org/10.1007/s11307-017-1086-6.
Rights and permissions
About this article
Cite this article
Malik, N., Gifford, A.N., Sandell, J. et al. Synthesis and In Vitro and In Vivo Evaluation of [3H]LRRK2-IN-1 as a Novel Radioligand for LRRK2. Mol Imaging Biol 19, 837–845 (2017). https://doi.org/10.1007/s11307-017-1070-1
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11307-017-1070-1