Abstract
Mutations in the leucine-rich repeat kinase 2 (LRRK2) are the most common known cause of autosomal dominant Parkinson’s disease (PD), accounting for approximately 1% of “sporadic” and 4% of familial cases. These mutations either lead directly to an increased kinase activity (G2019S and I2020T are in the kinase activation loop) or to a reduced GTPase activity (R1441C/G and Y1699C), that in turn positively regulate kinase activity. The physiological substrate of the LRRK2 kinase has yet to be definitively identified, yet autophosphorylation is emerging as a relatively robust measure of its activity. LRRK2 has been implicated in a number of diverse cellular processes such as vesicular trafficking, microtubule dynamics, protein translation control, inflammation, and immune function, all of which have been linked to PD. LRRK2 is a large, multi-domain protein; a thorough understanding of the protein domain organization and identification of interacting partners is important to determine the underlying mechanism of LRRK2. Substantial recent effort has been directed towards identifying potent LRRK2 kinase inhibitors, from the repurposed kinase inhibitors to the first through third generation of LRRK2-focused kinase inhibitors, from a range of chemotypes, which are now providing researchers with new tools to better interrogate LRRK2 function.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
Parkinson’s disease (PD) is the most common movement disorder and the second most common neurodegenerative disorder. The etiology of PD is complex but the most common phenotype is the loss of dopaminergic neurons of the substantia nigra (SN) leading to the clinical symptoms of bradykinesia, resting tremors, rigidity, and postural instability. Historically, at a pathological level, PD is characterized by inclusions containing the synaptic protein α-synuclein (α-syn) in the cell bodies and processes of surviving neurons (known as Lewy bodies and Lewy neurites, respectively). However, more recent observations describe not only a wider distribution of α-syn pathology, but also accumulation of tau and Aβ aggregates, suggesting a more complex pathology [1]. In addition, inflammation, including reactive gliosis, is observed in the striatum and substantia nigra of PD patients [2, 3].
The penetrance of the disease increases with age with 1% of the population over 65 being affected, rising to 5% by age 85 [4]. The past 17 years has led to considerable progress in both the identification of mutations that cause disease and in the mapping of common variants that alter risk for PD [5]. It is now clearly established that many, if not all, forms of Parkinson’s disease (PD) contain a genetic component [5], and mutations in the leucine-rich repeat kinase 2 (LRRK2) are the most common known cause of autosomal dominant PD, accounting for approximately 1% of “sporadic” and 4% of familial cases [5–7]. In specific populations, notably Ashkenazi Jews and North African Berber Arabs, the prevalence of LRRK2 mutations can be as high as 40% [8]. Of the LRRK2 mutations, the most common is G2019S with penetrance ranging from ~30 to 70% by the age of 80 [8–10]. This mutation occurs in the kinase domain of the protein, increasing the kinase activity, and consequently, a great deal of effort has been directed towards identifying potent LRRK2 kinase inhibitors, with a biopharmaceutical profile congruent with clinical evaluation, as novel therapeutics for PD. This chapter describes the current state of the art in developing appropriate kinase inhibitors.
2 LRRK2 Biology and Pharmacology
2.1 LRRK2 Genetics and Human Biology
The LRRK2 gene has 51 exons, with multiple potential splice sites, which encodes a large, 2,527 amino acid protein containing two predicted enzymatic domains (GTPase and kinase) and multiple protein–protein interaction domains (Fig. 1). See below for a more detailed account of LRRK2 structural biology (Sect. 3). A number of novel variants have been identified in this gene in PD patients, but only seven of these (N1437H, R1441C, R1441G, S1761R, Y1699C, G2019S, and I2020T) can be considered as definitively disease causing, on the basis of co-segregation with disease in families, and an absence in controls [5, 11–13]. These mutations either lead to an increased kinase activity (G2019S and I2020T) or to a reduced GTPase activity (R1441C/G and Y1699C), which in turn regulates kinase activity [14–16] (vide infra).
The G2019S mutation is relatively frequent in some populations from Southern Europe and in certain populations, such as the Ashkenazi Jews and North African Berber Arabs, where the prevalence can be as high as 40% [8]. However, there is an incomplete but age-related penetrance, ranging from ~30 to 70% by 80 years old in different studies, that has been estimated for carriers of the G2019S mutation, and the associated range of PD onset age is broad, including patients with early and late disease onset [8–10]. Dopaminergic neuronal loss and gliosis in the substantia nigra are the common pathological features in patients with LRRK2 mutations, and classical Lewy bodies are found in the majority of them. However, in some cases α-synuclein-positive inclusions are not observed, and only tau-positive or ubiquitin-positive inclusions are seen [7]. Overall, the clinical characteristics of patients with LRRK2 mutations, particularly those with the more common G2019S mutation, are very similar to those with sporadic PD [8, 17].
The incomplete penetrance clearly indicates that other factors are involved in the pathogenesis (Fig. 1); these may include other genetic contributions, both known (e.g., α-synuclein, Tau, RAB7L1, GAK) [5, 18] and yet to be defined, and/or “environmental” factors such as inflammation or oxidative/nitrative/unfolded and/or misfolded protein stress [19–21], many of which are actively being investigated. For example, LRRK2 kinase levels are increased by inflammatory mediators, such as IFNγ and LPS, in vitro [22, 23] and in vivo [23], and two recent reports suggest that LRRK2 levels are increased in sporadic PD brains [24, 25], potentially regulated by microRNA-205 [24]. LRRK2 has been shown to exist as a dimeric protein [26–29] with this form having greater kinase activity [28, 30]. These data have led to a proposed model in which LRRK2 cycles from a cytoplasmic, low-activity monomer to a higher-activity plasma membrane-associated dimer driven by, for example, LPS [28, 31, 32]. However, such biochemical experiments, investigating these potentially more active conformations of LRRK2, have not yet been performed in postmortem patient brains. Furthermore, to date, there is no direct evidence of increased LRRK2 kinase activity in postmortem patient brains from either G2019S mutation carriers or patients with sporadic disease. This is partly due to the challenges of immunoprecipitating sufficient active LRRK2 for in vitro phosphorylation assays, compounded by the potential effects of postmortem delay on the activity [33]. The lack of a validated substrate (vide infra) and the reliance on surrogate substrate, e.g., LRRKtide and NICtide [34, 35], also hampered such efforts. Despite this we, and others, are currently investigating these assays, and other end-points, to provide a critical link, which is currently missing, between LRRK2 and sporadic disease.
2.2 LRRK2 Substrates
The physiological substrate of the LRRK2 kinase has yet to be definitively identified. A number of potential substrates, other than LRRK2 (see next section), have been proposed (Table 1). However, full validation of these, including a clear demonstration of an increase in human PD patient cells and tissues and reduction in phosphorylation with multiple, structurally diverse inhibitors and by knocking out LRRK2, remains to be demonstrated.
2.3 Effects of Mutations on Kinase and GTPase Activity
The G2019S mutation, in the activation loop, has been consistently shown to result in increased kinase activity of LRRK2 [57, 58]. However, the functional consequences of the other mutations reported in the literature are conflicting. The other kinase domain mutation (I2020T) has been reported to either increase [59] or decrease kinase activity [34, 60]. Similarly, mutations in the GTPase domain have been demonstrated to increase kinase activity [57, 61], whereas in other studies, they have had little or no effect [34, 58]. More recent data have shown that the R1441C/G and Y1699C mutations reduced the GTPase activity that in turn regulates kinase activity [14, 15]. The majority of studies have used either recombinantly expressed and purified LRRK2 [58] or immunoprecipitated LRRK2 from recombinant mammalian expression systems [34, 57, 61] and investigated autophosphorylation or the use of surrogate substrates (myelin basic protein, LRRKtide, NICtide), all which are relatively poor substrates under those assay conditions.
LRRK2’s autophosphorylation has been characterized [30, 62, 63], and given the caveats of the validation of the exogenous substrate, perhaps this end-point is currently the most relevant. This is supported by recent data that demonstrated an increase in phosphorylation at S1292 by the pathological mutations [16]. Importantly, and in contrast to most of the previous results using purified enzyme, the magnitude of increase in phosphorylation of S1292 by, for example, G2019S is over 10-fold, when compared to wild-type LRRK2 [16] versus the 2–3-fold typically observed with the isolated enzyme [58]. This is clearly suggesting that the microenvironment of LRRK2 within the cell, including accessory proteins, interaction partners, and membrane association, is critical for the optimal kinase activity and that such a measure of kinase activity is a potentially valuable biomarker if it can be applied to patient samples. This will be a challenge as the stoichiometry of phosphorylation at S1292 is very low (<1%) ([16], unpublished observations), which may also reflect the tight control of LRRK2’s activity in the cellular context.
There are additional phosphorylation sites on LRRK2, including S910 and S935 [64], which have been shown to interact with 14-3-3 proteins and regulate the subcellular localization of LRRK2 [65]. While not autophosphorylation sites, they are clearly sensitive to LRRK2 kinase inhibitors, potentially due to conformational changes [66] exposing these residues to phosphatases, and have been used extensively in both cellular and in vivo studies (vide infra).
2.4 Role of LRRK2 in Normal and Pathological Biological Pathways
The understanding of LRRK2’s role in normal and pathological biological pathways is still at a relatively early stage, with a number of key questions that remain to be answered. For example, the endogenous substrate for the kinase domain is not known (vide supra and Table 1), and despite recent progress identifying interaction partners [67, 68], the full understanding of LRRK2’s function and interactome in different cell types remains an area of active research. The signaling pathways, through which LRRK2 elicits its actions and others that it potentially modulates, are emerging [69, 70] and its role in PD pathophysiology, while not definitive, is also developing.
LRRK2 has been implicated in a number of diverse cellular processes [19, 71, 72] including autophagy, vesicular trafficking, microtubule dynamics, neurite outgrowth, endosomal/synaptic dysregulation, protein translation control, mitochondrial pathology/ROS, WNT and MAPK/MEK/ERK/EIF2 signaling pathways, inflammation, and immune function, all of which have been linked to PD.
2.5 Animal Models to Understand LRRK2 Function
Studies in Drosophila melanogaster [73] and Caenorhabditis elegans [74] have provided important insights into LRRK2 toxicity in these species. The LRRK2 transgenic mouse models created to date do not completely recapitulate the hallmarks of PD (i.e., dopaminergic neuronal loss, α-synuclein accumulation, the development of Lewy bodies, and behavioral phenotype) [75–77], and with the exception of pharmacodynamic end-points, measuring reduction of LRRK2 phosphorylation at S935 and S1292 has not been used, to date, for any long-term LRRK2 inhibitor studies [16]. In contrast, viral overexpression of LRRK2 does result in dopaminergic neuronal loss [78, 79], but these models need to be fully validated with the more selective, brain-penetrant compounds described herein.
Toxins such as 6-hydroxydopamine (6-OHDA) or 1-methyl-4-phenyl-1,2,3,6-tetrahydropyridine (MPTP) have been effective in acutely inducing DA cell loss [80]. However, the LRRK2 knockout mouse displays the same sensitivity to MPTP as wild-type mice [81] suggesting that LRRK2 may not be involved in the events downstream of the mitochondrial toxicity in the mouse brain. There are no published reports on 6-OHDA in in vivo models, with LRRK2 overexpression or with endogenous LRRK2. The use of other toxins, such as LPS, may provide a more relevant model as LRRK2 kinase levels are increased by LPS in vitro [22, 23] and in vivo [23].
The lack of validated mammalian preclinical models of LRRK2 has been an impediment to the development of LRRK2 kinase inhibitors. This has also been hampered by the lack of suitable tool compounds that possess the appropriate pharmacokinetic properties and safety profiles for long-term dosing studies that are likely to be required.
2.6 LRRK2 and Potential Safety Concerns
LRRK2 is widely expressed, with highest expression in kidney, lung, and peripheral blood monocytes and lower levels in the brain [82]. LRRK2 functions in the peripheral tissues are also not well understood, and transgenic animals, knock-ins and knockouts, have been generated in an attempt to gain greater insight into LRRK2 biology. In all cases, the animals are viable and exhibit normal life spans. Initial studies with LRRK2 knockout (KO) mice exhibited phenotypes which showed a lack of hypersensitivity to MPTP, caused impairment of protein degradation pathways, resulted in accumulation of α-synuclein in kidney, not brain, and resulted in apoptotic cell death [76, 80, 83]. A single report with conditional LRRK2 G2019S overexpression in rats showed impaired dopamine reuptake and improved locomotor activity without loss of SN dopaminergic neurons [84]. LRRK2 KO rats are now commercially available [85]. While these animals appear to not have functional impairment, they do exhibit two distinct peripheral phenotypes, similar to the LRRK2 KO mice: (1) the kidneys present with a dark color that show inclusion bodies upon microscopic examination, and (2) type II pneumocytes of the lungs show changes in lamellar body morphology with altered surfactant secretion, measured in vitro [86–89]. The changes in type 2 pneumocyte vacuolation/lamellar body enlargement have recently been observed with 7- and 29-day NHP toxicology studies [90]. It remains to be determined whether KO phenotypes are a consequence of a total lack of LRRK2 from the embryonic stage to adult and if the drug-mediated phenotypes are chemotype or mechanism based.
3 LRRK2 Structural Biology
3.1 LRRK2 Protein Domain Structure
The LRRK2 gene encodes a 286 kDa protein comprising 2,527 amino acids [6, 91]. LRRK2 protein belongs to the Roco family of proteins whose members contain multiple domains including a tandem Roc (Ras of complex proteins)–COR (C-terminal of Roc) motif [92, 93]. As with the other members of the Roco family, LRRK2 contains multiple domains (Fig. 1): the enzymatic Roc–COR and kinase domains, which are flanked by putative protein–protein interaction domains, including the N-terminal armadillo (ARM) repeats, ankyrin repeats (AR or ANK) and leucine-rich repeats (LRR), and the C-terminal WD40 repeats [94, 95]. Currently, there is no unanimous agreement on the identity and number of armadillo, ankyrin, leucine-rich, and WD40 repeats in the LRRK2 protein [94]. A thorough understanding of the protein domain organization and identification of interacting partners is important to understand the function(s) of LRRK2.
3.1.1 Protein–Protein Interaction Domains
No X-ray crystal structures have yet been reported for any of the LRRK2 putative protein–protein interaction domains; therefore, its domain identification relies on bioinformatic analysis of the primary sequence to predict domain composition and boundaries.
The N-terminal LRRK2-specific repeats characterized by Marín [94] have been predicted to adopt the folds of 13 armadillo-type repeats from residues E49 to K657 [95]. Each ARM repeat is made up of ~42 amino acids that form three α-helices; tandem ARM repeats form a super helical structure that is involved in protein–protein interactions in other ARM repeat proteins [96]. This domain is predicted to be the largest LRRK2 domain containing ~600 amino acids.
Seven ANKs have been predicted to comprise the subsequent LRRK2 domain, made up of residues F676–S901 [95, 97]. Each AR is made up of 33 amino acids that form two antiparallel α-helices followed by a β-hairpin or a long loop, assembling to form a curved structure [98]. AR proteins are involved in a wide range of functions and typically mediate protein–protein interactions [99].
The next domain in sequence is the LRR domain, from where LRRK2 gets its name. As many as 14 LRRs have been predicted from residues I984 to R1320 [95, 100], though fewer have also been suggested [92, 94, 97, 101]. Each LRR is typically made up of 20–30 amino acids (~24 in LRRK2) and contains an unusually high number of leucine residues that form a β-strand followed by an α-helix or extended chain [102]. Tandem LRRs assemble into a horseshoe- or solenoid-type shape whose curvature depends on the specifics of the secondary structure. The framework of LRR domains makes them well suited for protein–protein interactions, and the concave surface of the LRR structure is usually, though not always, the site of macromolecular recognition or dimerization [103].
The C-terminal domain has been predicted to contain one [94], two [92], or seven [95, 97] WD40 repeats; the seven repeats are predicted to occur between residues N2164 and K2515 [95]. Each WD40 repeat contains a four-stranded, antiparallel β-sheet made up of 40 residues [97]. They usually occur in multiples of six or seven and assemble to form a propeller-like structure that serves as a rigid scaffold for protein–protein interactions [104].
3.1.2 Enzymatic Domains
The Roc domain of LRRK2 is a Ras-like GTPase domain that is followed immediately by the COR domain as in all members of the ROCO protein family [93]. These domains approximately span residues M1335–N1510 and F1511–E1878, respectively [97]. Ras family GTPases act as molecular switches between an active GTP-bound state and an inactive GDP-bound state [105, 106]. Evolutionary analysis indicates that not all ROCO proteins contain a kinase domain; as such, it has been suggested that GTPase activity is the primary function of ROCO proteins and those with kinase domains evolved independently to regulate that function [92, 93, 95, 107].
Two X-ray crystal structures have been solved providing the ability to form hypotheses about the LRRK2 Roc and COR domains and their functions based on structural information. One is a structure of the human LRRK2 Roc domain bound to GDP-Mg2+, which reveals a domain-swapped homodimer [108]. This structure suggests the Roc dimer acts as one functional unit with each nucleotide binding site formed by contributions from both monomers. The other relevant X-ray crystal structure contains both the Roc and COR domains from the bacteria Chlorobium tepidum and does not exhibit the domain swapping configuration [109]. This structure suggests that the COR domain is responsible for dimerization, that the Roc domain is stabilized by the COR domain, and that the Roc–COR domain is stabilized by nucleotide binding. Much discussion of the validity and relevance of these X-ray crystal structures has taken place [105, 107, 110–114]. Regardless, these structures both have implications for LRRK2 GTPase function that have spurred important discussions, and results from hypotheses generated based on these structures will only improve our understanding of the LRRK2 protein.
The kinase domain has the most straightforward therapeutic potential and thus has garnered the most interest of all of the domains in the LRRK2 protein. Characterized as a Ser/Thr kinase in the TKL group of kinases, it has the highest homology to LRRK1 and the receptor-interacting protein (RIP) kinases [94, 115], which are sensors of intracellular and extracellular stresses [116]. The LRRK2 kinase domain spans residues Q1879–V2138. Protein kinases catalyze the transfer of a phosphoryl group from ATP to serine, threonine, or tyrosine residues in protein substrates and mediate most cellular signaling processes [117]. The physiological substrate of the LRRK2 kinase has yet to be definitively identified (vide supra). The X-ray crystal structure of the kinase domain from the slime mold Dictyostelium discoideum Roco4 protein may provide insight into the structure of the LRRK2 kinase domain [118]. With a kinase domain that is ~30% identical and ~50% similar to that of LRRK2, as well as a similar domain structure that contains LRRs, Roc–COR, kinase, and WD40 repeat domains, the D. discoideum Roco4 kinase domain may be a suitable crystallography surrogate to help build an understanding of LRRK2 kinase structure and mechanism.
3.2 LRRK2 Pathogenic PD Mutations
To date, the proven pathogenic PD mutations (Fig. 1) segregate to the enzymatic Roc–COR and kinase domains of LRRK2 [13, 119, 120], though there are a number of variants that have been identified as PD risk factors that are distributed throughout the entire protein [121, 122]. As such, these mutations have the potential to disrupt not only LRRK2 enzymatic functions but also protein–protein interactions. In the absence of LRRK2 X-ray crystal structures, homology models [123, 124] have been valuable tools to help understand the position of pathogenic mutations and the effect(s) they may have on the function of the LRRK2 protein.
3.2.1 Roc–COR Pathogenic Mutations
The R1441C/G/H and N1437H pathogenic mutations are located in the Roc domain and the Y1699C and S1761R mutations are located in the COR domain. The two current models for the Roc domain suggest different roles for these residues (Fig. 2) [125]. In the domain-swapped homodimer X-ray crystal structure of the human LRRK2 Roc domain proposed by Cookson and coworkers [108], R1441 and N1437 are situated such that they stabilize the Roc–Roc dimer interface through interactions with the other monomer. R1441 forms HB interactions with the backbone carbonyl of F1401 and the side chain of T1404 and forms a cation-π interaction with the side chain of F1401, while N1437 forms a HB interaction with the side chain of R1398.
Roc–COR structures with positions of pathogenic mutations and interacting residues. (a) Self-homology model of the domain-swapped LRRK2 Roc–Roc homodimer (PDB ID: 2ZEJ). Orange: GDP; Green: monomer A; Gray: monomer B; Yellow: interacting residues. (b) Homology model of LRRK2 Roc–COR dimer based on the C. tepidum structure (PDB ID: 3DPU). Cyan: Roc monomer A; Pink: COR monomer A; Light blue: Roc monomer B; Light pink: COR monomer B; Yellow: interacting residues
Alternatively, in the X-ray crystal structure of the Roc–COR dimer from C. tepidum proposed by Wittinghofer and coworkers [109], the residues equivalent to R1441, N1437, and Y1699 (Y558C. tepidum, H554C. tepidum, and Y804C. tepidum, respectively) are clustered together at the interface of the Roc and COR domains, involved in interactions within and between the Roc and COR domains. Based on the Y558C. tepidum interactions, it is hypothesized that the aliphatic side chain of R1441 is involved in structurally important hydrophobic interactions with the COR domain. A homology model based on this structure suggests that R1441 may also form HB interactions with the side chains of E1400 and S1403 in the Roc domain and that N1437 forms HB interaction with the side chains of E1399 in the Roc domain and Y1699 in the COR domain [126] (Fig. 2a). Based on the Y804C. tepidum interactions, Y1699 may form a hydrogen bond with N1437 (H554C. tepidum). A homology model based on this structure suggests Y1699 may also form a HB with the N1437 side chain or the G1397 backbone carbonyl and a CH-π interaction with the Cα of N1437 [126] (Fig. 2b).
Neither model places these residues close enough to the binding site to interact with GTP or GDP, so the pathogenic mutations do not appear to disrupt nucleotide binding directly. In both models, mutation of R1441 to Cys, Gly, or His and N1437 mutation to His would disrupt HB and van der Waals (VDW) interactions, and in the C. tepidum model, mutation of Y1699 to Cys would disrupt a HB between the Roc and COR domains. Therefore, evidence from both models suggests that these residues play an important role in dimer formation or interactions between domains, affecting LRRK2 protein function (vide supra). In addition, these three residues form interactions with residues in the Switch II region of the Roc domain [109], a conserved region in Ras proteins that is important for GTPase function [127, 128]. This suggests that mutation of these residues could also affect LRRK2 GTPase function by altering interactions with the Switch II region.
The S1761 residue in the COR domain corresponds to S852C. tepidum in the C. tepidum structure. This residue is farther away from the Roc–COR interface and Switch II than the other pathogenic mutations and is situated on the loop between β9 and β10. As such, it may be involved in protein–protein interactions with other LRRK2 domains or other scaffolding proteins. Mutation to Arg could affect these protein–protein interactions or the flexibility of the loop.
3.2.2 Kinase Domain Pathogenic Mutations
The G2019S and I2020T pathogenic mutations are located in the kinase domain. In the absence of an X-ray crystal structure of the LRRK2 kinase domain, a number of homology models have been reported and used to understand the position of these residues and the implications of the pathogenic mutations. There is no consensus on a “best” template to build LRRK2 kinase domain homology models due to the fact that the kinases for which X-ray crystal structures have been published have at most ~30% sequence identity to LRRK2. The most common templates have been from B-Raf [94, 101, 129–131], JAK-2 [132–138], MLK1 [132, 136–139], ROCO4 [131, 140], and TAK1 [132, 141, 142] kinases, chosen based on overall kinase sequence identity, ATP-binding site identity, and/or crossover of inhibitor activity. Consensus from these alignments is that G2019 is the Gly in the conserved DFG motif (DYG in LRRK2) and I2020 is the subsequent residue in the activation loop (Fig. 3) [125].
Modeling of the G2019S mutation suggests that the change from a flexible Gly residue to a Ser causes the activation loop to be less flexible due to an increase in HB interactions with D2017, E1920, or other nearby residues, increasing the population of conformations in the active state [114, 131, 136, 141]. The X-ray crystal structure of Roco4 with the mutation that corresponds to LRRK2 G2019S, G1179SRoco4, was solved to 2.04 Å resolution and reveals a HB interaction between G1179SRoco4 and R1077Roco4 in the αC-helix, which corresponds to Q1919 in LRRK2 [118]. This construct was shown to have increased kinase activity compared to WT. In addition, the double mutants G1179SRoco4/R1077ARoco4 in Roco4 and G2019S/Q1919A in LRRK2, in which the Ser is unable to make a HB interaction to the αC-helix to stabilize the active conformation, have nearly wild-type activity. Metadynamic simulations combined with kinetics studies suggested that the energy barrier to achieve the inactive DYG-out conformation in LRRK2 is much higher for the G2019S mutant than for WT [131]. Together, this evidence suggests that an additional hydrogen bond in the G2019S mutant stabilizes the active kinase conformation, causing an increase in kinase activity.
Modeling of the I2020T mutation, on the other hand, does not suggest a clear hypothesis for its pathogenicity. The X-ray crystal structure of Roco4 kinase with the corresponding mutation, L1180T Roco4, was solved to 2.3 Å resolution and shows the side chain of the Thr interacting with solvent [118]. It was suggested that the effect of the mutation may only be evident upon LRRK2 dimerization or interactions with other domains, which cannot be discerned from the X-ray crystal structure of the kinase domain alone. Alternatively, molecular dynamics simulations suggest the Thr mutation may form a HB interaction with the backbone carbonyl of D2017, which is involved in substrate binding [60, 114]. Metadynamic simulations combined with kinetic studies suggested that the I2020T mutant stabilized the active DYG-in conformation in LRRK2 compared to WT [60].
3.3 Using LRRK2 Structure to Guide Kinase Inhibitor Design
Most of the LRRK2 kinase inhibitors that have been published to date are ATP-competitive inhibitors that contain hinge-binding motifs common to many kinase inhibitors (vide infra). Through the use of structure–activity relationships (SAR), X-ray crystal structures, and homology models, specific residues in the LRRK2 ATP-binding pocket have been targeted for inhibitor interactions to improve potency and selectivity for LRRK2 kinase (Fig. 4). Hydrophobic interactions with A2016 have been shown to be important for potency and selectivity of some LRRK2 kinase inhibitors [35, 140, 143, 144]. Selectivity of these compounds is especially improved over kinases that have a more polar residue at that position – 28% of kinases have a Ser or Thr at that position. Mutation of A2016 to Thr abolishes activity in some inhibitors [35, 140, 143], but not others [145–147], which is indicative of the inhibitor binding modes.
The non-conserved residues L1949, S1954, R1957, and F1883 were identified as potential selectivity handles in the LRRK2 kinase domain, due to their proximity to the ATP-binding pocket for a series of diaminopyrimidine compounds [132–135]. L1949 near the hinge region was targeted for selectivity since it is a smaller residue than Phe or Tyr, which is found in this position in approximately 60% of the kinome. Inhibitors with a nitrogen lone pair or substituents or that extend into this region can be accommodated with the Leu residue, but sterically clash with the larger Phe or Tyr. This strategy was used to obtain selectivity for LRRK2 over the JAK family of kinases and JAK2 was found to be a useful surrogate for predicting general kinome selectivity. Interactions with S1954 were targeted to achieve both general kinase selectivity and to overcome specific selectivity issues with the kinase TTK. Approximately 55% of the kinome has a larger group at this position, and approximately 50% has a negatively charged Asp or Glu residue in this position. Strategies to mitigate crossover to TTK, which has an Asp residue in this position, have included the introduction of small groups that would cause unfavorable steric and/or electrostatic interactions with the Asp side chain [132, 133, 135]. Selectivity of a series of 7-aryl-substituted quinoline derivatives may also be due to the placement of polar groups in the vicinity of the unconserved S1954 and R1957 residues [138]. Interactions with the unconserved R1957 and H1998 LRRK2 residues may also contribute to selectivity in a series of indolinone compounds [144]. Optimizing interactions with H1998 and the catalytic K1906 in this series of compounds was hypothesized to improve kinase selectivity, especially over RET kinase.
Alternative strategies used to obtain selectivity in kinases are through Type II inhibitors, which stabilize an inactive DFG-out conformation, or Type III inhibitors, which are not ATP-competitive. Only a handful of LRRK2 inhibitors have been reported that fall into these categories [131, 148]. These inhibitors were shown to be more potent against WT LRRK2 kinase compared to G2019S, supporting the hypothesis that the pathogenic G2019S mutation stabilizes the active conformation of LRRK2 [60, 114, 131, 136, 141]. This suggests a Type II kinase inhibitor may not be the optimal approach for the treatment of PD due to the LRRK2 G2019S mutation. The X-ray crystal structure of Roco4 kinase domain co-crystallized with the inhibitor H1152 revealed two binding sites for this inhibitor: one in the expected ATP-binding site and the other close to the αC-helix [118]. The implications of this second binding site for LRRK2 are unclear, but suggest not completely ruling out the possibility of developing Type III inhibitors for LRRK2 kinase.
4 Medicinal Chemistry: LRRK2 Kinase Inhibition
Historically, kinase inhibitors have been a key target for oncology indications. The critical role of kinases in cell cycle or apoptotic signaling pathways made them a logical target. From a drug discovery perspective for non-oncology indications, pharmacologically promiscuous kinase inhibitors can potentially give rise to safety concerns due to their varied roles in signaling pathways. These safety issues are particularly acute for the treatment of chronic diseases that require long-term treatment, typically experienced with neurodegenerative disorders. Recently, kinase inhibitors have been developed for non-oncology indications. Xeljanz® (tofacitinib), a JAK3 inhibitor, is a good example and was approved in 2012 for moderate to severe rheumatoid arthritis (http://www.xeljanz.com/).
For a central nervous system (CNS) indication, in addition to the off-target selectivity, one needs to contend with the blood–brain barrier (BBB) and designing compounds that have access to the central compartment. The medicinal chemistry challenge is readily apparent when one analyzes the current marketed kinase chemical space relative to the historical CNS chemical space. From a kinase inhibitor perspective, the potential pharmaceutical agent must contend with equilibria of active, inactive, and intermediate conformations (e.g., DFG-in vs. DFG-out). Most of the initially marketed kinases targeted the DFG-out conformation which generates a much larger active site pocket, thus giving rise to compounds with MW, logP, or PSA parameters outside the range of typical CNS space. Comparison of these physicochemical parameters is presented in Table 2. These data were generated from an analysis of marketed kinases relative to the marketed CNS drugs and a set of preclinical CNS candidates from the Pfizer pipeline. Additionally, employing the CNS multiparameter optimization (MPO) score [149], as a guide to the likelihood the compound may be CNS drug like, clearly shows the current chemical matter is outside this space. Furthermore, as a consequence to the high concentration of ATP present in the cell, any potential kinase inhibitor will need to have high potency to be able to compete at a mass balance level.
For a CNS indication, such as PD, clearly having free drug in the brain at the site of action is an absolute requirement. Moreover, to minimize the potential for off-target toxicology, the requisite concentration of free drug in the brain should be achieved with minimal exposure in the periphery. Combining all these attributes generates a CNS kinase design strategy that incorporates the selection of chemical matter that has low MW and is polar and neutral with high brain availability. These compounds additionally will need to have high ligand binding efficiencies (LE), lipophilic efficiencies (LipE), and low efficacious concentration (Ceff), all the while having high kinome selectivity.
The current public domain LRRK2 kinase inhibitor chemical matter will be summarized with an eye to how they compare to the design strategy articulated (vide supra).
4.1 LRRK2 Patent Space Analysis
As a target, LRRK2 was first identified in 2004 and its connection to PD was confirmed by genome-wide associate studies (GWAS) in 2010 (For a recent PD genetics review: [150]). This short-time frame coupled with the lack of its (patho)physiological role(s) (vide supra) has presented a challenge to the medicinal chemist as witnessed by a relatively small compound set compared to targets with a more rich medicinal chemistry history. Conversely, the highly conserved nature of the ATP-binding site of kinases does give rise to the great potential for crossing-over of chemical scaffolds from inhibitors designed for alternate kinases. Indeed, the first set of LRRK2 kinase inhibitors investigated were compounds repurposed from other kinase programs (vide infra). A handful of LRRK2 selective kinase inhibitors have been published in the primary literature; however, they are typically specific examples from a greater set of compounds claimed in the LRRK2 patent literature. The Markush structures for this set of LRRK2 kinase inhibitors are presented in Fig. 5. In general terms, the LRRK2 ATP-binding site can accommodate known core scaffolds that incorporate 1-, 2-, and potentially 3-point hinge-binding motifs. Addressing the safety, CNS design features, and chemical novelty will arise from how these scaffolds are adorned with requisite substituents.
The attributes that the medicinal chemist has the greatest design control over are the physicochemical properties. All subsequently measured compound properties are embedded once the structure has been fixed. Figure 6 provides an overview of the LRRK2 patented chemical matter as it relates to physicochemical space. The box plots provide a summary of each property showing the data distribution with the average (red line), the CNS drug set average (green line), and the marketed kinase set (black dashed line). MW shows a normal distribution with the average between those for the kinase and CNS drug sets. This tendency towards the CNS drug set is not observed for PSA with the LRRK2 and kinase sets being essentially the same. Measures of lipophilicity (clogP and logD) again show a normal distribution with the average closer to the CNS drug set. HB donor count (HBD) for the LRRK2 compounds will be skewed to the kinase set due to the required hinge interactions, and this was observed. The distribution for pK a appears to be bimodal for the LRRK2 set, but the average tends closer to the CNS drug set.
Physicochemical analysis of LRRK2 patent chemical matter. Each box overlays the corresponding distribution for the descriptor on its box plots. The dots represent outliers and the lines denote LRRK2 data set average (red), CNS drug set average (green), and marketed kinase drugs average (black dashed)
Figure 7 provides an overview of the probability of in vivo toxicology findings (tox plot) [151]. The upper left quadrant (clogP > 3 and PSA < 75) was determined to have the greatest likelihood of in vivo safety findings. Conversely, the lower right quadrant (clogP < 3 and PSA > 75) was found to have the lowest probability of a safety finding. The pie charts are scaled to the number of compounds in each quadrant, and Fig. 7 is color coded for CNS MPO desirability. Clearly, most of the compounds reside in good CNS space (MPO > 4), but a significant number of these compounds reside in physicochemical space expected to give rise to safety findings.
Figure 8 illustrates the breakdown of chemical matter by the patent assignee. Again, most organizations are working in chemical space that is congruent with CNS drugs, and in several instances, the companies have leveraged existing chemotypes from marketed kinases (Fig. 5). John Hopkins University has patented the use of existing kinase inhibitors, e.g., staurosporine, damnacanthal, SP600125, 5-iodotubercidin, GW5074, and indirubin-3′-monoxime as LRRK2 inhibitors, whereas Tautatis focused on modified versions of staurosporine analogues. Novartis, Southern Methodist University, and Zenobia focused on the oxindole framework as exemplified by sunitinib (Sutent®). The diaminopyrimidine core can be found in patents from Cellzome, Dana Farber, and Roche (Genentech). The Medical Research Council alone and in collaboration with Genentech, DCAM Pharma, Merck, Arrien, Origenis, Pfizer, and Southern Research Institute have all generated variations on a 6,5-fused bicyclic framework. Ipsen/Oncodesign extended the use of this scaffold by exemplifying macrocyclic variants of this core. GSK has disclosed a novel LRRK2 kinase inhibitor scaffold in the form of an aryl ether amide.
Comparing this set of compounds, as a whole, to the data for CNS drugs and marketed kinases (vide supra) provides potential insight as to the position in chemical space these compounds have relative to the space currently defined by the CNS drug and marketed kinase sets. All these compounds are ATP-competitive inhibitors presumably targeting the DYG-in conformation as the equilibrium distribution between the DYG-in and DYG-out (inactive conformation) appears to be skewed towards the active conformation by 5–6 kcal/mol [131]. Despite the lack of a LRRK2 crystal structure, one can leverage the knowledge from the kinase literature and predict that these compounds will interact with the hinge motif in a 1- or 2-point manner. Thus, to drive selectivity and ultimately safety, the substitution pattern from the core is crucial to accomplish this requisite attribute. From homology models, Genentech has identified two critical residues necessary to drive LRRK2 selectivity. These include the hinge residue Leu1949 and the activation loop residue Arg1957. In the case of the diaminopyrimidines, Genentech was able to increase selectivity to off-targets (e.g., MST2 and JAK1) with substituents on the pendant aniline or aminopyrazole moieties.
4.2 LRRK2 Chemical Matter Overview
It has been suggested that the ability to study a protein’s function is enhanced when small molecule tools for the target exist [152]. Ideally a “compound toolbox” would be available, covering different chemical scaffolds and physicochemical and ADMET properties, and these compounds would be well characterized in terms of on- and off-target effects. With distinct scaffolds, overlap of off-target effects, for example, between the compounds is likely to be reduced, and consistent findings across the compounds would therefore be indicative of activity at the target in question.
The 28 patent applications of small molecules (Fig. 5), to date, represent roughly 10 unique chemical scaffolds. Through the use of SAR, X-ray crystal structures (surrogate crystallography), and homology models, specific residues in the LRRK2 ATP-binding pocket have been targeted for inhibitor interactions to improve potency and selectivity for LRRK2 kinase. Initial activities focused on LRRK2 potency and selectivity. As the compounds achieved these goals, focus shifted to address ADME and safety issues that were revealed as these compounds were used to expand LRRK2 biology. The diversity of the current chemical matter provides a toolbox of compounds that should enable the detailed elaboration of LRRK2 kinase function.
4.2.1 Repurposed Kinase Inhibitors
Early work in the field of LRRK2 kinase inhibitor development sought to compliment the range of biochemical tools that were becoming available [153] with small molecule kinase inhibitors. By screening panels of commercial compounds, groups began to identify known kinase inhibitors that also displayed activity against LRRK2 [35, 78, 154]. Initially identified compounds, using LanthaScreen detection and a surrogate peptide substrate (LRRKtide), included 7–9 and 11–12. Staurosporine, 10, was one of the most potent LRRK2 inhibitors reported (wt IC50 = 6 nM); however, it is also highly promiscuous across the kinome [155]. Capitalizing on this observation, 4 illustrates how modifications to the bis-indole core were pursued to potentially address the selectivity issue. Sunitinib, 34, is a potent LRRK2 inhibitor (wt IC50 = 37 nM), but also displays similar promiscuity. Related indolidinones (1, 17, and 27) have been disclosed. Screening of a number of Rho-kinase inhibitors led to the identification of 35 [35]. Although it has a number of off-target effects, selectivity is much improved over 10 and 34, although potency against LRRK2 is also reduced (wt IC50 = 244 nM).

Although these early inhibitors are structurally different, they have in common potent activity across a range of kinases, and thus it is difficult to separate effects at LRRK2 from off-target effects, especially when examined in isolation. These compounds proved useful for initial study of the function of LRRK2; however, it was clear that the field could benefit from more selective compounds.

4.2.2 First-Generation LRRK2-Focused Kinase Inhibitors
In 2011, the first LRRK2-optimized kinase inhibitor LRRK2-IN-1, 36, was reported (derived from 3) [143]. Due to its high potency against both wild-type and mutant LRRK2 (IC50 values of 13 and 6 nM, respectively, measured in cell-free enzymatic assays), and its improved selectivity across a range of kinases, it was clearly a vast improvement on previous compounds and was initially employed throughout the field as a selective LRRK2 inhibitor. While 36 is potent against both LRRK2 WT and G2019S mutant enzymes, it has been demonstrated to display moderate potency in a whole cell assay, 200–600 nM [40, 146]. Further, high doses are required to observe any in vivo effects due to its low permeability and high levels of plasma protein binding. Additionally, no CNS effects were observed as 36 had low brain availability (unbound Cbrain/unbound Cplasma). Although 36 demonstrated improved selectivity in comparison to previous LRRK2 inhibitors, it has some significant off-target potency, most notably displaying equipotent activity against ERK5 (BMK1, MAPK7), DCAMKL, PLK1, and PLK4. This off-target activity confounded some of the biological assays in which the effects of 36 are attributed to LRRK2, in particular when assessing effects on neurite outgrowth and inflammation end-points [40]; consequently, use of 36 has diminished.
4.2.3 Second-Generation LRRK2-Focused Kinase Inhibitors
As intensive research into the function and dysfunction of LRRK2 and its mutants continued, a number of other tool compounds and inhibitor series have been published from both academic and industrial laboratories. The most potent of these derived from 2, in both enzyme (5–7 nM in WT and G2019S, respectively) and cell-based assays (attenuation of G2019S and R1441C-induced neuronal injury and death in a concentration-dependent manner with an EC50 of <10 nM), was CZC-54252 (37) reported in 2011 [156]. Unfortunately, 37 had disappointing selectivity across the kinome. Although CZC-54252 may prove a useful tool for cell-based assays, its poor CNS MPO score (2.12) suggested it was outside of CNS space which will limit its application in in vivo models (confirmed with a reported brain availability of 0.05).
The diaminopyrimidine core has been extensively examined by several groups. In addition to compounds derived from 2 and 3 (vide infra), variations including analogues from 6, 13, 16, 21, and 31 have been reported. An early example is exemplified by 38 (CNS MPO = 3.97), but more advanced analogues successfully balanced good physicochemical properties with high degrees of LRRK2 selectivity [157, 158]. Focusing on diaminopyrimidine HTS hits, due to high ligand efficiency and reasonable PK properties, optimization of the hits using a LRRK2 homology model to identify both key interactions for LRRK2 potency and sites that may offer enhanced kinase selectivity yielded 39 (MPO = 5.37) with improved kinase selectivity over their initial hit, suitable in vivo exposures, and similar cellular activity (IC50 = 389 nM) to 36. While disclosed in a patent the previous year (WO2012062783; see Fig. 5, above), 39 was subsequently described as HG-10-102-01 and reported to exhibit in vivo mouse brain activity [132].

Having identified TTK (MPS1) as the major off-target interaction of this series, and significant safety concern, continued optimization by Genentech scientists, using structure-based drug design initially arrived at 40 (CNS MPO = 4.60), a compound having excellent selectivity for LRRK2 [145]. The in vivo activity of this compound and related analogues was shown using dephosphorylation of S1292 as a measure of LRRK2 kinase activity. Although 40 is an excellent tool compound for probing the effect of LRRK2 inhibition in vivo, it has some major liabilities, thus further optimization was needed for progression towards a potential clinical candidate. As the diaminopyrimidine forms the key hinge interactions with LRRK2, optimization was focused around the side chain. The aniline motif, which could lead to idiosyncratic toxicity findings, was replaced with an aminopyrazole, and side chains to enhance aqueous solubility were incorporated [133–135]. This optimization lead to GNE-0877, 41 (CNS MPO = 5.45), which has been progressed to preclinical safety studies (see Sect. 2.6, above).
4.2.4 Third-Generation LRRK2-Focused Kinase Inhibitors
Moving away from diaminopyrimidines, several other chemotypes have been reported. A series defined by 5 and 18, while likely to be ATP-competitive, do not contain a common hinge-binding motif [135]. This attribute may be responsible for the excellent kinase selectivity. GSK2578215A, 42, is a representative analogue from this series and showed reasonable potency (IC50 = 47 nM). However, it has poor rodent PK, with a relatively low oral bioavailability (%F = 12) and half life of 1.1 h [159]. The lack of LRRK2 inhibition observed in the brain (potentially predicted by its CNS MPO score of 3.82), in comparison to kidney and spleen, suggested low levels of unbound compound in brain potentially limiting the use of this series for in vivo models.
Quinoline derivatives, 26 and 28, are known single-point hinge binders and have been found to be good LRRK2 kinase inhibitors [139, 147]. Although the cinnoline variants [141] were potent LRRK2 inhibitors (wt IC50 = 7 nM), they were found to be promiscuous in a small kinase panel. By focusing the core to quinoline [138], e.g., 43, kinase selectivity was significantly improved and in vivo activity was demonstrated (CNS MPO = 5.05).

Pyrazolopyri(mi)dines, 15, 19, 23–25, and 30, have been well represented in the patent literature. For analogues, such as 44 (CNS MPO = 5.83), potency is comparable to 36 with overall good ADME [159]. Alternate 6,5-fused ring systems, 14, 20, and 22, have been reported; however, the pyrrolopyrimidine 33 appears to have the best alignment of properties. Specifically, 45 (CNS MPO = 5.83) is one of the most active LRRK2 inhibitors in vivo (brain free drug wt IC50 = 15 nM) along with high kinase selectivity [160].

One final variation of the 6,5-fused ring system is illustrated by 32. These compounds are also postulated to be single-point hinge binders, interacting through one of the triazole nitrogens. A representative compound from this series, 46 (CNS MPO = 4.88), displayed excellent in vitro potency (wt IC50 = 31 nM; G2019S IC50 = 8 nM) against LRRK2, but this did not translate to cellular assays resulting in a 100-fold right shift [137]. As witnessed with other scaffolds (vide supra), compounds from this chemotype would make good tool compounds for probing LRRK2 in vitro biology.
4.2.5 Pharmacological Profiles for Key LRRK2-Focused Kinase Inhibitors
As the relatively young LRRK2 field of research has evolved, there have been frequent advances in the biochemical reagents available, as the biology has developed. In part, this has been facilitated by the improvement of the chemical tools (vide supra). However, the literature data documenting the pharmacological profiles of the tool compounds is often not consistent in terms of the assays or conditions used in reporting their potencies. Thus, Table 3 presents a head-to-head comparison of several of the top compounds, from our internal assessments, with the intention of providing LRRK2 researchers with additional data to better enable selecting of the best compound(s) for their studies.
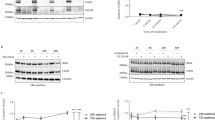
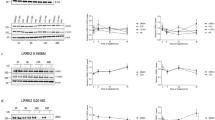
The LRRK2 inhibitors described below have been characterized in a number of in vitro and in vivo models. Activity against LRRK2 wild-type and G2019S mutant was assayed using truncated enzyme from Invitrogen in a LanthaScreen format at 50 μM and 1 mM ATP concentrations [58]. The whole cell potency is measured in HEK293 cells transfected with WT LRRK2, using the antibody against pS935 [64]. The RRCK (Ralph Russ Canine Kidney) assay is a measure of passive permeability, using a canine kidney cell line which expresses low levels of efflux transporters [161]. MDR1 is a measure of efflux by human P-gp transporter and is used to assess the likelihood of brain impairment [162]. HLM and RLM are an assessment of metabolism in human and rat liver microsomes, respectively. Several safety assays are performed; “dof” is a measure of the ability of compounds to displace [3H]-labeled dofetilide from the hERG K+-channel, important for cardiac safety in clinical candidates [163]. General cytotoxicity is measured using the THLE assay [164]. Effects on cell health are followed up with a panel of assays performed in HepG2 cells, measuring mitochondrial function [165]. Kinome selectivity is reported from a Dundee Panel [166] using recombinant enzymes at the ATP KM or ActivX data [167], which used lysed PBMC cells to assay selectivity in a more native system.
5 Summary, Conclusions, and Outlook
The answers to some key questions are important for a more thorough understanding of LRRK2 structure, function, and the development of inhibitors, including how the Roc domain regulates the kinase domain, what the role of the COR domain is, and how the pathogenic mutations alter the function of the LRRK2 protein. X-ray crystal structures of the LRRK2 enzymatic domains, including both WT and the pathogenic mutations, could contribute to an understanding of these questions and would be a key accomplishment to help drive inhibitor design.
The strength of research in the field of LRRK2, both academic and industrial, has generated a wealth of chemical tools for the study of LRRK2. Where the initially reported compounds had activity across multiple kinase targets, new inhibitors have a high degree of kinome selectivity along with improved physicochemical properties, providing opportunities to study inhibition of LRRK2 kinase activity from the isolated enzyme through to in vivo assays. Ultimately, this collection of inhibitors should allow researchers the flexibility to design experiments to probe LRRK2 function, knowing that they have the right tools for the job.
Abbreviations
- 6-OHDA:
-
6-Hydroxydopamine
- ANK/AR:
-
Ankyrin repeats
- ARM:
-
Armadillo
- BA:
-
Brain availability
- BBB:
-
Blood brain barrier
- BI:
-
Brain impairment
- CNS:
-
Central nervous system
- COR:
-
C-terminal of Roc
- GPCR:
-
G-protein coupled receptor
- GWAS:
-
Genome-wide association study
- HB:
-
hydrogen bond
- HBD:
-
hydrogen bond donor count
- hERG:
-
Human ether-a-go-go-related gene
- HLM:
-
Human liver microsome
- KO:
-
Knockout
- LE:
-
Ligand efficiency
- LipE:
-
Lipophilic ligand efficiency
- logD:
-
Natural logarithm of the distribution coefficient
- logP:
-
Natural logarithm of the water/octanol partition coefficient
- LPS:
-
Lipopolysaccharide
- LRR:
-
Leucine-rich repeats
- LRRK2:
-
Leucine-rich repeat kinase 2
- MDR1:
-
Multidrug resistance
- MPO:
-
Multiparameter optimization
- MPTP:
-
1-Methyl-4-phenyl-1,2,3,6-tetrahydropyridine
- MW:
-
Molecular weight
- NHP:
-
Nonhuman primate
- PBMC:
-
Peripheral blood mononuclear cell
- PD:
-
Parkinson’s disease
- PSA:
-
Polar surface area
- RLM:
-
Rat liver microsome
- Roc:
-
Ras of complex proteins
- ROS:
-
Reactive oxygen species
- RRCK:
-
Ralph Russ Canine Kidney
- SAR:
-
Structure–activity relationship
- SN:
-
Substantia nigra
- THLE:
-
Transformed human liver epithelial
- VDW:
-
Van der Waals
- WD40:
-
WD40 repeats
- α-syn:
-
α-Synuclein
References
Antony PM, Diederich NJ, Krüger R, Balling R (2013) The hallmarks of Parkinson’s disease. FEBS J 280:5981
Deleidi M, Gasser T (2013) The role of inflammation in sporadic and familial Parkinson’s disease. Cell Mol Life Sci 70:4259
Pradhan S, Andreasson K (2013) Commentary: progressive inflammation as a contributing factor to early development of Parkinson’s disease. Exp Neurol 241:148
Reeve A, Simcox E, Turnbull D (2014) Ageing and Parkinson’s disease: why is advancing age the biggest risk factor? Ageing Res Rev 14:19
Singleton AB, Farrer MJ, Bonifati V (2013) The genetics of Parkinson’s disease: progress and therapeutic implications. Mov Disord 28:14
Paisan-Ruiz C, Jain S, Evans EW et al (2004) Cloning of the gene containing mutations that cause PARK8-linked Parkinson’s disease. Neuron 44:595
Zimprich A, Biskup S, Leitner P et al (2004) Mutations in LRRK2 cause autosomal-dominant parkinsonism with pleomorphic pathology. Neuron 44:601
Healy DG, Falchi M, O’Sullivan S, Bonifati V, Durr A, Bressman S, Brice A, Aasly J, Zabetian CP, Goldwurm S, Ferreira JJ, Tolosa E, Kay DM, Klein C, Williams DR, Marras C, Lang AE, Wszolek ZW, Bericiano J, Schapira AHV, Lynch T, Bhatia KP, Gasser T, Lees AJ, Wood NW (2008) Phenotype, genotype, and worldwide genetic penetrance of LRRK2-associated Parkinson's disease: a case–control study. Lancet Neurol 7:583
Goldwurm S, Zini M, Mariani L et al (2007) Evaluation of LRRK2 G2019S penetrance: relevance for genetic counseling in Parkinson disease. Neurology 68:1141
Latourelle JC, Sun M, Lew MF et al (2008) The Gly2019Ser mutation in LRRK2 is not fully penetrant in familial Parkinson’s disease: the GenePD study. BMC Med 6:32
Gasser T (2009) Molecular pathogenesis of Parkinson disease: insights from genetic studies. Expert Rev Mol Med 11:e22
Aasly JO, Vilariño-Güell C, Dachsel JC, Webber PJ, West AB, Haugarvoll K, Johansen KK, Toft M, Nutt JG, Payami H, Kachergus JM, Lincoln SJ, Felic A, Wider C, Soto-Ortolaza AI, Cobb SA, White LR, Ross OA, Farrer MJ (2010) Novel pathogenic LRRK2 p. Asn1437His substitution in familial Parkinson’s disease. Mov Disord 25:2156
Lorenzo-Betancor O, Samaranch L, Ezquerra M, Tolosa E, Lorenzo E, Irigoyen J, Gaig C, Paster MA, Soto-Ortolaza AI, Ross OA, Rodriguez-Oroz MC, Valldeoriola F, Marti MJ, Luquin MR, Perez-Tur J, Burguera JA, Obeso JA, Pastor P (2012) LRRK2 haplotype-sharing analysis in Parkinson's disease reveals a novel p.S1761R mutation. Mov Disord 27:146
Taymans JM, Vancraenenbroeck R, Ollikainen P, Beilina A, Lobbestael E, De Maeyer M, Baekelandt V, Cookson MR (2011) LRRK2 kinase activity is dependent on LRRK2 GTP binding capacity but independent of LRRK2 GTP binding. PLoS One 6:e23207
Biosa A, Trancikova A, Civiero L, Glauser L, Bubacco L, Greggio E, Moore DJ (2013) GTPase activity regulates kinase activity and cellular phenotypes of Parkinson’s disease-associated LRRK2. Hum Mol Genet 22:1140
Sheng Z, Zhang S, Bustos D, Kleinheinz T, Le Pichon CE, Dominguez SL, Solanoy HO, Drummond J, Zhang X, Ding X, Cai F, Song Q, Li X, Yue Z, van der Brug MP, Burdick DJ, Gunzner-Toste J, Chen H, Liu X, Estrada AA, Sweeney ZK, Scearce-Levie K, Moffat JG, Kirkpatrick DS, Zhu H (2012) Ser1292 autophosphorylation is an indicator of LRRK2 kinase activity and contributes to the cellular effects of PD mutations. Sci Transl Med 4:164ra161
Haugarvoll K, Rademakers R, Kachergus JM, Nuytemans K, Ross OA, Gibson JM, Tan EK, Gaig C, Tolosa E, Goldwurm S, Guidi M, Riboldazzi G, Brown L, Walter U, Benecke R, Berg D, Gasser T, Theuns J, Pals P, Cras P, De Deyn PP, Engelborghs S, Pickut B, Uitti RJ, Foroud T, Nichols WC, Hagenah J, Klein C, Samii A, Zabetian CP, Bonifati V, Van Broeckhoven C, Farrer MJ, Wszolek ZK (2008) LRRK2 R1441C parkinsonism is clinically similar to sporadic Parkinson disease. Neurology 70:1456
Hyun CH, Yoon CY, Lee HJ, Lee SJ (2013) LRRK2 as a potential genetic modifier of synucleinopathies: interlacing the two major genetic factors of Parkinson’s disease. Exp Neurobiol 22:249
Russo I, Bubacco L, Greggio E (2014) LRRK2 and neuroinflammation: partners in crime in Parkinson’s disease? J Neuroinflammation 11:52
Zuo L, Motherwell MS (2013) The impact of reactive oxygen species and genetic mitochondrial mutations in Parkinson's disease. Gene 532:18
Manzoni C, Lewis PA (2013) Dysfunction of the autophagy/lysosomal degradation pathway is a shared feature of the genetic synucleinopathies. FASEB J 27:3424
Gardet A, Benita Y, Li C, Sands BE, Ballester I, Stevens C, Korzenik JR, Rioux JD, Daly MJ, Xavier RJ, Podolsky DK (2010) LRRK2 is involved in the IFN-gamma response and host response to pathogens. J Immunol 185:5577
Moehle MS, Webber PJ, Tse T, Sukar N, Standaert DG, DeSilva TM, Cowell RM, West AB (2012) LRRK2 inhibition attenuates microglial inflammatory responses. J Neurosci 32:1602
Cho HJ, Liu G, Jin SM, Parisiadou L, Xie C, Yu J, Sun L, Ma B, Ding J, Vancraenenbroeck R, Lobbestael E, Baekelandt V, Taymans JM, He P, Troncoso JC, Shen Y, Cai H (2013) MicroRNA-205 regulates the expression of Parkinson’s disease-related leucine-rich repeat kinase 2 protein. Hum Mol Genet 22:608
Guerreiro PS, Huang Y, Gysbers A, Cheng D, Gai WP, Outeiro TF, Halliday GM (2013) LRRK2 interactions with α-synuclein in Parkinson's disease brains and in cell models. J Mol Med 91:513
Greggio E, Zambrano I, Kaganovich A, Beilina A, Taymans JM, Daniëls V, Lewis P, Jain S, Ding J, Syed A, Thomas KJ, Baekelandt V, Cookson MR (2008) The Parkinson disease-associated leucine-rich repeat kinase 2 (LRRK2) is a dimer that undergoes intramolecular autophosphorylation. J Biol Chem 283:16906
Sen S, Webber PJ, West AB (2009) Dependence of leucine-rich repeat kinase 2 (LRRK2) kinase activity on dimerization. J Biol Chem 284:36346
Berger Z, Smith KA, Lavoie MJ (2010) Membrane localization of LRRK2 is associated with increased formation of the highly active LRRK2 dimer and changes in its phosphorylation. Biochemistry 49:5511
Civiero L, Vancraenenbroeck R, Belluzzi E, Beilina A, Lobbestael E, Reyniers L, Gao F, Micetic I, De Maeyer M, Bubacco L, Baekelandt V, Cookson MR, Greggio E, Taymans JM (2012) Biochemical characterization of highly purified leucine-rich repeat kinases 1 and 2 demonstrates formation of homodimers. PLoS One 7:e43472
Webber PJ, Smith AD, Sen S, Renfrow MB, Mobley JA, West AB (2011) Autophosphorylation in the leucine-rich repeat kinase 2 (LRRK2) GTPase domain modifies kinase and GTP-binding activities. J Mol Biol 412:94
James NG, Digman MA, Gratton E, Barylko B, Ding X, Albanesi JP, Goldberg MS, Jameson DM (2012) Number and brightness analysis of LRRK2 oligomerization in live cells. Biophys J 102:L41
Schapansky J, Nardozzi JD, Felizia F, Lavoie MJ (2014) Membrane recruitment of endogenous LRRK2 precedes its potent regulation of autophagy. Hum Mol Genet. doi:10.1093/hmg/ddu138
Davies P, Hinkle KM, Sukar NN, Sepulveda B, Mesias R, Serrano G, Alessi DR, Beach TG, Benson DL, White CL, Cowell RM, Das SS, West AB, Melrose HL (2013) Comprehensive characterization and optimization of leucine rich repeat kinase 2 (LRRK2) monoclonal antibodies. Biochem J 453:101
Jaleel M, Nichols RJ, Deak M, Campbell DG, Gillardon F, Knebel A, Alessi DR (2007) LRRK2 phosphorylates moesin at threonine-558: characterization of how Parkinson’s disease mutants affect kinase activity. Biochem J 405:307
Nichols RJ, Dzamko N, Hutti JE, Cantley LC, Deak M, Moran J, Bamborough P, Reith AD, Alessi DR (2009) Substrate specificity and inhibitors of LRRK2, a protein kinase mutated in Parkinson’s disease. Biochem J 424:47
Imai Y, Gehrke S, Wang HQ, Takahashi R, Hasegawa K, Oota E, Lu B (2008) Phosphorylation of 4E-BP by LRRK2 affects the maintenance of dopaminergic neurons in Drosophila. EMBO J 27:2432
Kumar A, Greggio E, Beilina A, Kaganovich A, Chan D, Taymans JM, Wolozin B, Cookson MR (2010) The Parkinson’s disease associated LRRK2 exhibits weaker in vitro phosphorylation of 4E-BP compared to autophosphorylation. PLoS One 5:e8730
Trancikova A, Mamais A, Webber PJ, Stafa K, Tsika E, Glauser L, West AB, Bandopadhyay R, Moore DJ (2012) Phosphorylation of 4E-BP1 in the mammalian brain is not altered by LRRK2 expression or pathogenic mutations. PLoS One 7:e47784
Qing H, Wong W, McGeer EG, McGeer PL (2009) Lrrk2 phosphorylates alpha synuclein at serine 129: Parkinson disease implications. Biochem Biophys Res Commun 387:149
Luerman GC, Nguyen C, Samaroo H, Loos P, Xi H, Hurtado-Lorenzo A, Needle E, Stephen Noell G, Galatsis P, Dunlop J, Geoghegan KF, Hirst WD (2014) Phosphoproteomic evaluation of pharmacological inhibition of leucine-rich repeat kinase 2 reveals significant off-target effects of LRRK-2-IN-1. J Neurochem 128:561
Ohta E, Kawakami F, Kubo M, Obata F (2011) LRRK2 directly phosphorylates Akt1 as a possible physiological substrate: impairment of the kinase activity by Parkinson’s disease-associated mutations. FEBS Lett 585:2165
Stafa K, Trancikova A, Webber PJ, Glauser L, West AB, Moore DJ (2012) GTPase activity and neuronal toxicity of Parkinson's disease-associated LRRK2 is regulated by ArfGAP1. PLoS Genet 8:e1002526
Xiong Y, Yuan C, Chen R, Dawson TM, Dawson VL (2012) ArfGAP1 is a GTPase activating protein for LRRK2: reciprocal regulation of ArfGAP1 by LRRK2. J Neurosci 32:3877
Gillardon F (2009) Leucine-rich repeat kinase 2 phosphorylates brain tubulin-beta isoforms and modulates microtubule stability–a point of convergence in Parkinsonian neurodegeneration? J Neurochem 110:1514
Matta S, Van Kolen K, da Cunha R, van den Bogaart G, Mandemakers W, Miskiewicz K, De Bock PJ, Morais VA, Vilain S, Haddad D, Delbroek L, Swerts J, Chávez-Gutiérrez L, Esposito G, Daneels G, Karran E, Holt M, Gevaert K, Moechars DW, De Strooper B, Verstreken P (2012) LRRK2 controls an EndoA phosphorylation cycle in synaptic endocytosis. Neuron 75:1008
Parisiadou L, Xie C, Cho HJ, Lin X, Gu XL, Long CX, Lobbestael E, Baekelandt V, Taymans JM, Sun L, Cai H (2009) Phosphorylation of ezrin/radixin/moesin proteins by LRRK2 promotes the rearrangement of actin cytoskeleton in neuronal morphogenesis. J Neurosci 29:13971
Kanao T, Venderova K, Park DS, Unterman T, Lu B, Imai Y (2010) Activation of FoxO by LRRK2 induces expression of proapoptotic proteins and alters survival of postmitotic dopaminergic neuron in Drosophila. Hum Mol Genet 19:3747
Pungaliya PP, Bai Y, Lipinski K, Anand VS, Sen S, Brown EL, Bates B, Reinhart PH, West AB, Hirst WD, Braithwaite SP (2010) Identification and characterization of a leucine-rich repeat kinase 2 (LRRK2) consensus phosphorylation motif. PLoS One 5:e13672
Zach S, Felk S, Gillardon F (2010) Signal transduction protein array analysis links LRRK2 to Ste20 kinases and PKC zeta that modulate neuronal plasticity. PLoS One 5:e13191
Gloeckner CJ, Schumacher A, Boldt K, Ueffing M (2009) The Parkinson disease-associated protein kinase LRRK2 exhibits MAPKKK activity and phosphorylates MKK3/6 and MKK4/7, in vitro. J Neurochem 109:959
Chen CY, Weng YH, Chien KY, Lin KJ, Yeh TH, Cheng YP, Lu CS, Wang HL (2012) (G2019S) LRRK2 activates MKK4-JNK pathway and causes degeneration of SN dopaminergic neurons in a transgenic mouse model of PD. Cell Death Differ 19:1623
Hsu CH, Chan D, Greggio E, Saha S, Guillily MD, Ferree A, Raghavan K, Shen GC, Segal L, Ryu H, Cookson MR, Wolozin B (2010) MKK6 binds and regulates expression of Parkinson’s disease-related protein LRRK2. J Neurochem 112:1593
Martin I, Kim JW, Lee BD, Kang HC, Xu JC, Jia H, Stankowski J, Kim MS, Zhong J, Kumar M, Andrabi SA, Xiong Y, Dickson DW, Wszolek ZK, Pandey A, Dawson TM, Dawson VL (2014) Ribosomal protein s15 phosphorylation mediates LRRK2 neurodegeneration in Parkinson’s disease. Cell 157:472
Yun HJ, Park J, Ho DH, Kim H, Kim CH, Oh H, Ga I, Seo H, Chang S, Son I, Seol W (2013) LRRK2 phosphorylates Snapin and inhibits interaction of Snapin with SNAP-25. Exp Mol Med 45:e36
Kawakami F, Yabata T, Ohta E, Maekawa T, Shimada N, Suzuki M, Maruyama H, Ichikawa T, Obata F (2012) LRRK2 phosphorylates tubulin-associated tau but not the free molecule: LRRK2-mediated regulation of the tau-tubulin association and neurite outgrowth. PLoS One 1:e30834
Bailey RM, Covy JP, Melrose HL, Rousseau L, Watkinson R, Knight J, Miles S, Farrer MJ, Dickson DW, Giasson BI, Lewis J (2013) LRRK2 phosphorylates novel tau epitopes and promotes tauopathy. Acta Neuropathol 126:809
West AB, Moore DJ, Biskup S, Bugayenko A, Smith WW, Ross CA, Dawson VL, Dawson TM (2005) Parkinson’s disease-associated mutations in leucine-rich repeat kinase 2 augment kinase activity. Proc Natl Acad Sci U S A 102:16842
Anand VS, Reichling LJ, Lipinski K, Stochaj W, Duan W, Kelleher K, Pungaliya P, Brown EL, Reinhart PH, Somberg R, Hirst WD, Riddle SM, Braithwaite SP (2009) Investigation of leucine-rich repeat kinase 2: enzymological properties and novel assays. FEBS J 276:466
Gloeckner CJ, Kinkl N, Schumacher A, Braun RJ, O'Neill E, Meitinger T, Kolch W, Prokisch H, Ueffing M (2006) The Parkinson disease causing LRRK2 mutation I2020T is associated with increased kinase activity. Hum Mol Genet 15:223
Ray S, Bender S, Kang S, Lin R, Glicksman MA, Liu M (2014) The Parkinson disease-linked LRRK2 protein mutation I2020T stabilizes an active state conformation leading to increased kinase activity. J Biol Chem 289:13042
West AB, Moore DJ, Choi C, Andrabi SA, Li X, Dikeman D, Biskup S, Zhang Z, Lim KL, Dawson VL, Dawson TM (2007) Parkinson’s disease-associated mutations in LRRK2 link enhanced GTP-binding and kinase activities to neuronal toxicity. Hum Mol Gene 16:223
Gloeckner CJ, Boldt K, von Zweydorf F, Helm S, Wiesent L, Sarioglu H, Ueffing M (2010) Phosphopeptide analysis reveals two discrete clusters of phosphorylation in the N-terminus and the Roc domain of the Parkinson-disease associated protein kinase LRRK2. J Proteome Res 9:173
Kamikawaji S, Ito G, Sano T, Iwatsubo T (2013) Differential effects of familial parkinson mutations in LRRK2 revealed by a systematic analysis of autophosphorylation. Biochemistry 52:6052
Dzamko N, Deak M, Hentati F, Reith AD, Prescott AR, Alessi DR, Nichols RJ (2010) Inhibition of LRRK2 kinase activity leads to dephosphorylation of Ser(910)/Ser(935), disruption of 14-3-3 binding and altered cytoplasmic localization. Biochem J 430:405
Nichols RJ, Dzamko N, Morrice NA, Campbell DG, Deak M, Ordureau A, Macartney T, Tong Y, Shen J, Prescott AR, Alessi DR (2010) 14-3-3 binding to LRRK2 is disrupted by multiple Parkinson's disease-associated mutations and regulates cytoplasmic localization. Biochem J 430:393
Gillardon F, Kremmer E, Froehlich T, Ueffing M, Hengerer B, Gloeckner CJ (2013) ATP-competitive LRRK2 inhibitors interfere with monoclonal antibody binding to the kinase domain of LRRK2 under native conditions. A method to directly monitor the active conformation of LRRK2? J Neurosci Methods 214:62
Beilina A, Rudenko IN, Kaganovich A, Civiero L, Chau H, Kalia SK, Kalia LV, Lobbestael E, Chia R, Ndukwe K, Ding J, Nalls MA; International Parkinson’s Disease Genomics Consortium; North American Brain Expression Consortium, Olszewski M, Hauser DN, Kumaran R, Lozano AM, Baekelandt V, Greene LE, Taymans JM, Greggio E, Cookson MR. (2014) Unbiased screen for interactors of leucine-rich repeat kinase 2 supports a common pathway for sporadic and familial Parkinson disease. Proc Natl Acad Sci U S A 111:2626
Piccoli G, Onofri F, Cirnaru MD, Kaiser CJ, Jagtap P, Kastenmüller A, Pischedda F, Marte A, von Zweydorf F, Vogt A, Giesert F, Pan L, Antonucci F, Kiel C, Zhang M, Weinkauf S, Sattler M, Sala C, Matteoli M, Ueffing M, Gloeckner CJ (2014) Leucine-rich repeat kinase 2 binds to neuronal vesicles through protein interactions mediated by its C-terminal WD40 domain. Mol Cell Biol 34:2147
Berwick DC, Harvey K (2011) LRRK2 signaling pathways: the key to unlocking neurodegeneration? Trends Cell Biol 21:257
Verma M, Steer EK, Chu CT (2013) ERKed by LRRK2: a cell biological perspective on hereditary and sporadic Parkinson’s disease. Biochim Biophys Acta. doi:10.1016/j.bbadis.2013.11.005
Rideout HJ, Stefanis L (2014) The neurobiology of LRRK2 and its role in the pathogenesis of Parkinson’s disease. Neurochem Res 39:576
Cookson MR (2010) The role of leucine-rich repeat kinase 2 (LRRK2) in Parkinson’s disease. Nat Rev Neurosci 12:791
Hindle SJ, Elliott CJ (2013) Spread of neuronal degeneration in a dopaminergic, Lrrk-G2019S model of Parkinson disease. Autophagy 9:936
Yao C, Johnson WM, Gao Y, Wang W, Zhang J. Deak M, Alessi DR, Zhu X, Mieyal JJ, Roder H, Wilson-Delfosse AL, Chen SG (2013) Kinase inhibitors arrest neurodegeneration in cell and C. elegans models of LRRK2 toxicity. Hum. Mol Genet 22:328
Li X, Patel JC, Wang J, Avshalumov MV, Nicholson C, Buxbaum JD, Elder GA, Rice ME, Yue Z (2010) Enhanced striatal dopamine transmission and motor performance with LRRK2 overexpression in mice is eliminated by familial Parkinson's disease mutation G2019S. J Neurosci 30:1788
Li Y, Liu W, Oo TF, Wang L, Tang Y, Jackson-Lewis V, Zhou C, Geghman K, Bogdanov M, Przedborski S, Beal MF, Burke RE, Li C (2009) Mutant LRRK2(R1441G) BAC transgenic mice recapitulate cardinal features of Parkinson’s disease. Nat Neurosci 12:826
Ramonet D, Daher JP, Lin BM, Stafa K, Kim J, Banerjee R, Westerlund M, Pletnikova O, Glauser L, Yang L, Liu Y, Swing DA, Beal MF, Troncoso JC, McCaffery JM, Jenkins NA, Copeland NG, Galter D, Thomas B, Lee MK, Dawson TM, Dawson VL, Moore DJ (2011) Dopaminergic neuronal loss, reduced neurite complexity and autophagic abnormalities in transgenic mice expressing G2019S mutant LRRK2. PLoS One 6:e18568
Lee BD, Shin JH, VanKampen J, Petrucelli L, West AB, Ko HS, Lee YI, Maguire-Zeiss KA, Bowers WJ, Federoff HJ, Dawson VL, Dawson TM (2010) Inhibitors of leucine-rich repeat kinase-2 protect against models of Parkinson’s disease. Nat Med 16:998
Dusonchet J, Kochubey O, Stafa K, Young SM Jr, Zufferey R, Moore DJ, Schneider BL, Aebischer P (2011) A rat model of progressive nigral neurodegeneration induced by the Parkinson's disease-associated G2019S mutation in LRRK2. J Neurosci 31:07
Tieu K (2011) A guide to neurotoxic animal models of Parkinson's disease. Cold Spring Harb Perspect Med 1:a009316
Andres-Mateos E, Mejias R, Sasaki M, Li X, Lin BM, Biskup S, Zhang L, Banerjee R, Thomas B, Yang L, Liu G, Beal MF, Huso DL, Dawson TM, Dawson VL (2009) Unexpected lack of hypersensitivity in LRRK2 knock-out mice to MPTP (1-methyl-4-phenyl-1,2,3,6-tetrahydropyridine). J Neurosci 29:15846
Biskup S, Moore DJ, Rea A, Lorenz-Deperieux B, Coombes CE, Dawson VL, Dawson TM, West AB (2007) Dynamic and redundant regulation of LRRK2 and LRRK1 expression. BMC Neurosci 8:102
Tong Y, Yamaguchi H, Giaime E, Boyle S, Kopan R, Kelleher RJ, Shen J. (2010) Loss of leucine-rich repeat kinase 2 causes impairment of protein degradation pathways, accumulation of alpha-synuclein, and apoptotic cell death in aged mice. Proc Natl Acad Sci U S A. 107: 9879
Zhou H, Huang C, Tong J, Hong WC, Liu YJ, Xia XG (2011) Temporal expression of mutant LRRK2 in adult rats impairs dopamine reuptake. Int J Biol Sci 7:753
LEH-LRRK2tm1sage from SAGE Labs, product number: TGRL4620. http://www.sageresearchlabs.com/files/Parkinsons%20Disease%20KORat%20Flyer_1.pdf. Accessed 4 August 2014
Herzig MC, Kolly C, Persohn E, Theil D, Schweizer T, Hafner T, Stemmelen C, Troxler TJ, Schmid P, Danner S, Schnell CR, Mueller M, Kinzel B, Grevot A, Bolognani F, Stirn M, Kuhn RR, Kaupmann K, van der Putten PH, Rovelli G, Shimshek DR (2011) LRRK2 protein levels are determined by kinase function and are crucial for kidney and lung homeostasis in mice. Hum Mol Genet 20:4209
Ness D, Ren Z, Gardai S, Sharpnack D, Johnson VJ, Brennan RJ, Brigham EF, Olaharski AJ (2013) Leucine-rich repeat kinase 2 (LRRK2)-deficient rats exhibit renal tubule injury and perturbations in metabolic and immunological homeostasis. PLoS One 8:e66164
Miklavc P, Ehinger K, Thompson KE, Hobi N, Shimshek DR, Frick M (2010) Surfactant secretion in LRRK2 knock-out rats: changes in lamellar body morphology and rate of exocytosis. PLoS One 9:e84926
Baptista MA, Dave KD, Frasier MA, Sherer TB, Greeley M, Beck MJ, Varsho JS, Parker GA, Moore C, Churchill MJ, Meshul CK, Fiske BK (2013) Loss of leucine-rich repeat kinase 2 (LRRK2) in rats leads to progressive abnormal phenotypes in peripheral organs. PLoS One 8:e80705
Fuji, R (2013) Nonclinical safety studies of selective LRRK2 kinase inhibitors. Presented at Seventh annual Parkinson’s disease therapeutics conference, The New York Academy of Sciences, October 24, 2013
The UniProt Consortium (2013) Update on activities at the Universal Protein Resource (UniProt) in 2013. Nucl Acids Res 41:D43
Bosgraaf L, Van Haastert PJM (2003) Roc, a Ras/GTPase domain in complex proteins. Biochim Biophys Acta 1643:5
Marin I, van Egmond WN, van Haastert PJM (2008) The Roco protein family: a functional perspective. Faseb J 22:3103
Marin I (2006) The Parkinson disease gene LRRK2: evolutionary and structural insights. Mol Biol Evol 23:2423
Mills RD, Mulhern TD, Cheng HC, Culvenor JG (2012) Analysis of LRRK2 accessory repeat domains: prediction of repeat length, number and sites of Parkinson’s disease mutations. Biochem Soc Trans 40:1086
Tewari R, Bailes E, Bunting KA, Coates JC (2010) Armadillo-repeat protein functions: questions for little creatures. Trends Cell Biol 20:470
Mata IF, Wedemeyer WJ, Farrer MJ, Taylor JP, Gallo KA (2006) LRRK2 in Parkinson’s disease: protein domains and functional insights. Trends Neurosci 29:286
Mosavi LK, Cammett TJ, Desrosiers DC, Zy P (2004) The ankyrin repeat as molecular architecture for protein recognition. Protein Sci 13:1435
Li J, Mahajan A, Tsai MD (2006) Ankyrin repeat: a unique motif mediating protein–protein interactions. Biochemistry 45:15168
Vancraenenbroeck R, Lobbestael E, Weeks SD, Strelkov SV, Baekelandt V, Taymans JM, DeMaeyer M (2012) Expression, purification and preliminary biochemical and structural characterization of the leucine rich repeat namesake domain of leucine rich repeat kinase 2. Biochim Biophys Acta 1824:450
Guo L, Wang W, Chen SG (2006) Leucine-rich repeat kinase 2: relevance to Parkinson’s disease. Int J Biochem Cell Biol 38:1469
Kobe B, Kajava AV (2001) The leucine-rich repeat as a protein recognition motif. Curr Opin Struct Biol 11:725
Bella J, Hindle KL, McEwan PA, Lovell SC (2008) The leucine-rich repeat structure. Cell Mol Life Sci 65:2307
Xu C, Min J (2011) Structure and function of WD40 domain proteins. Protein Cell 2:202
Taymans JM (2012) The GTPase function of LRRK2. Biochem Soc Trans 40:1063
Xiong Y, Dawson VL, Dawson TM (2012) LRRK2 GTPase dysfunction in the pathogenesis of Parkinson’s disease. Biochem Soc Trans 40:1074
Tsika E, Moore D (2013) Contribution of GTPase activity to LRRK2-associated Parkinson disease. Small GTPases 4:164
Deng J, Lewis PA, Greggio E, Sluch E, Beilina A, Cookson MR (2008) Structure of the ROC domain from the Parkinson’s disease-associated leucine-rich repeat kinase 2 reveals a dimeric GTPase. Proc Natl Acad Sci U S A 105:1499
Gotthardt K, Weyand M, Kortholt A, VanHaastert PJM, Wittinghofer A (2008) Structure of the Roc-COR domain tandem of C. tepidum, a prokaryotic homologue of the human LRRK2 Parkinson kinase. EMBO J 27:2239
Gotthardt K, Weyand M, Kortholt A, VanHaastert PJ, Wittinghofer A (2008) Structure of the Roc-COR domain tandem of C. tepidum, a prokaryotic homologue of the human LRRK2 Parkinson kinase. EMBO J 27:2352
Gasper R, Meyer S. Gotthardt K, Sirajuddin M, Wittinghofer A (2009) It takes two to tango: regulation of G proteins by dimerization. Nat Rev Mol Cell Biol 10:423
Greggio E, Cookson MR (2009) Leucine-rich repeat kinase 2 mutations and Parkinson’s disease: three questions. ASN Neuro 1:13
Li Y, Dunn L, Greggi E, Krumm B, Jackson GS, Cookson MR, Lewis MR, Deng J (2009) The R1441C mutation alters the folding properties of the ROC domain of LRRK2. Biochim Biophys Acta 1792:1194
Ray S, Liu M (2012) Current understanding of LRRK2 in Parkinson’s disease: biochemical and structural features and inhibitor design. Future Med Chem 4:1701
Manning G, Whyte DB, Martinez R, Hunter T, Sudarsanam S (2002) The protein kinase complement of the human genome. Science 298:1912
Zhang D, Lin J, Han J (2010) Receptor-interacting protein (RIP) kinase family. Cell Mol Immunol 7:243
Adams JA (2001) Kinetic and catalytic mechanisms of protein kinases. Chem Rev 101:2271
Gilsbach BK, Ho FY, Vetter IR, vanHaastert PJM, Wittinghofer A, Kortholt A (2012) Roco kinase structures give insights into the mechanism of Parkinson disease-related leucine-rich-repeat kinase 2 mutations. Proc Natl Acad Sci U S A 109:10322
Aasly JO, Toft M, Fernandez-Mata I, Kacherqus J, Hulihan M, White LR, Farrer M (2005) Clinical features of LRRK2-associated Parkinson’s disease in central Norway. Ann Neurol 57:762
Bardien S, Lesage S, Brice A, Carr J (2011) Genetic characteristics of leucine-rich repeat kinase 2 (LRRK2) associated Parkinson's disease. Parkinsonism Relat Disord 17:501
Paisan-Ruiz C (2009) LRRK2 gene variation and its contribution to Parkinson disease. Hum Mutat 30:1153
Paisan-Ruiz C, Lewis PA, Singleton AB (2013) LRRK2: cause, risk, and mechanism. J Parkinson’s Dis 3:85
Cavasotto CN, Phatak SS (2009) Homology modeling in drug discovery: current trends and applications. Drug Discov Today 14:676
Krieger E, Nabuurs SB, Vriend G (2005) Homology modeling. In: Bourne PE, Weissig H (eds) Structural bioinformatics, vol 44. Wiley, Hoboken. doi:10.1002/0471721204.ch25
LRRK2 homology models were constructed with MOE (Molecular Operating Environment (MOE), 2013.08; Chemical Computing Group Inc., 1010 Sherbooke St. West, Suite #910, Montreal, QC, Canada, H3A 2R7, 2013) using the AMBER99 force field (Wang JM, Cieplak P, Kollman PA. (2000) How well does a restrained electrostatic potential (RESP) model perform in calculating conformational energies of organic and biological molecules? J Comput Chem 21:1049). The amino acid sequence of human LRRK2 was retrieved from UniProt (The UniProt Consortium (2008) The Universal Protein Resource (UniProt) Nucleic Acids Res 36:D190) and aligned to the template structure sequences specified in the text using MOE’s Protein Align function with default settings followed by manual editing of loop regions, insertions and deletions
Daniels V, Vancraenenbroeck R, Law BMH, Greggio E, Lobbestael E, Gao F, DeMaeyer M, Cookson MR, Harvey K, Baekelandt V, Taymans JM (2011) Insight into the mode of action of the LRRK2 Y1699C pathogenic mutant. J Neurochem 116:304
Fiegen D, Dvorsky R, Ahmadian M (2006) Structural principles of Ras interaction with regulators and effectors. In: Der C (ed) RAS family GTPases, Springer, The Netherlands, p 45
Rojas J, Ras-Gefs E, Santos (2006) Ras Gaps. In: Der C (ed) RAS family GTPases, Springer, the Netherlands, p 15
Albrecht M (2005) LRRK2 mutations and Parkinsonism. Lancet 365:1230
Liu M, Kang S, Ray S, Jackson J, Zaitsev AD, Gerber SA, Cuny GD, Glicksman MA (2011) Kinetic, mechanistic, and structural modeling studies of truncated wild-type leucine-rich repeat kinase 2 and the G2019S mutant. Biochemistry 50:9399
Liu M, Bender SA, Cuny GD, Sherman W, Glicksman M, Ray SS (2013) Type II kinase inhibitors show an unexpected inhibition mode against Parkinson’s disease-linked LRRK2 mutant G2019S. Biochemistry 52:1725
Chen H, Chan BK, Drummond J, Estrada AA, Gunzer-Toste J, Liu X, Liu Y, Moffat J, Shore D, Sweeney ZK, Tran T, Wang S, Zhao G, Zhu H, Burdick DJ (2012) Discovery of selective LRRK2 inhibitors guided by computational analysis and molecular modeling. J Med Chem 55:5536
Estrada AA, Liu X, Baker-Glenn C, Beresford A, Burdick DJ, Chambers M, Chan BK, Chen H, Ding X, DiPasquale AG, Dominguez SL, Dotson J, Drummond J, Flagella M, Flynn S, Fuji R, Gill A, Gunzner-Toste J, Harris SF, Heffron TP, Kleinheinz T, Lee DW, LePichon DE, Lyssikatos JP, Medhurst AD, Moffat JG, Mukund S, Nash K, Scearce-Levie K, Shang Z, Shore DG, Tran T, Tivedi N, Wang S, Zhang S, Zhang X, Zhao G, Zhu H, Sweeney ZK (2012) Discovery of highly potent, selective, and brain-penetrable leucine-rich repeat kinase 2 (LRRK2) small molecule inhibitors. J Med Chem 55:9416
Chan BK, Estrada AA, Chen H, Atherall J, Baker-Glenn C, Beresford A, Burdick DJ, Chambers M, Dominguez SL, Drummond J, Gill A, Keinheinz T, LePichon CE, Medhurst AD, Liu X, Moffat JG, Nash K, Scearce-Levie K, Sheng Z, Shore DG, VandePoel H, Zhang S, Zhu H, Sweeney ZK (2013) Discovery of a highly selective, brain-penetrant aminopyrazole LRRK2 inhibitor. ACS Med Chem Lett 4:85
Estrada AA, Chan BK, Baker-Glenn C, Beresford A, Burdick DJ, Chambers M, Chen H, Dominguez SL, Dotson J, Drummond J, Flagella M, Fuji R, Gill A, Halladay J, Harris SF, Heffron TP, Kleinheinz T, Lee DW, LePichon CR, Liu X, Lyssikatos JP, Medhurst AD, Moffat JG, Nash K, Scearce-Levie K, Sheng Z, Shore DG, Wong S, Zhang S, Zhang X, Zhu H, Sweeney ZK (2014) Discovery of highly potent, selective, and brain-penetrant aminopyrazole leucine-rich repeat kinase 2 (LRRK2) small molecule inhibitors. J Med Chem 57:921
Drolet RE, Sanders JM, Kern JT (2011) Leucine-rich repeat kinase 2 (LRRK2) cellular biology: a review of recent advances in identifying physiological substrates and cellular functions. J Neurogenet 25:140
Franzini M, Ye ZM, Adler M, Aubele DL, Garofalo AW, Gauby S, Goldbach E, Probst GD, Quinn KP, Santiago P, Sham HL, Tam D, Troung A, Ren Z (2013) Triazolopyridazine LRRK2 kinase inhibitors. Bioorg Med Chem Lett 23:1967
Garofalo AW, Adler M, Aubele DL, Brigham EF, Chian D, Franzini M, Goldbach E, Kwong GT, Motter R, Probst GD, Quinn KP, Ruslim L, Sham HL, Tam D, Tanaka P, Troung AP, Ye XM, Ren Z (2013) Discovery of 4-alkylamino-7-aryl-3-cyanoquinoline LRRK2 kinase inhibitors. Bioorg Med Chem Lett 23:1974
Garofalo AW, Adler M, Aubele DL, Bowers S, Franzini M, Goldbach E, Lorentzen C, Neitz RJ, Probst GD, Quinn KP, Santiago P, Sham HL, Tam D, Troung AP, Ye XM, Ren Z (2013) Novel cinnoline-based inhibitors of LRRK2 kinase activity. Bioorg Med Chem Lett 23:71
Deng X, Elkins JM, Zhang J, Yang Q, Erazo T, Gomez N, Choi HG, Wang J, Dzamko N, Lee JD, Sim T, Kim N, Alessi DR, Lizcano JM, Knapp S, Gray NS (2013) Structural determinants for ERK5 (MAPK7) and leucine rich repeat kinase 2 activities of benzo[e]pyrimido-[5,4-b]diazepine-6(11H)-ones. Eur J Med Chem 70C:758
Anand VS, Braithwaite SP (2009) LRRK2 in Parkinson’s disease: biochemical functions. Febs J 276:6428
Yun H, Heo HY, Kim HH, DooKim N, Seol W (2011) Identification of chemicals to inhibit the kinase activity of leucine-rich repeat kinase 2 (LRRK2), a Parkinson’s disease-associated protein. Bioorg Med Chem Lett 21:2953
Deng X, Dzamko N, Prescott A, Davies P, Liu Q, Yang Q, Lee JD, Patricelli MP, Nomanbhoy TN, Alessi DR, Gray NS (2011) Characterization of a selective inhibitor of the Parkinson’s disease kinase LRRK2. Nat Chem Biol 7:203
Troxler T, Greenidge P, Zimmerman K, Desrayaud S, Drukes P, Schweizer T, Stauffer D, Rovelli G, Shimshek DR (2013) Discovery of novel indolinone-based, potent, selective and brain penetrant inhibitors of LRRK2. Bioorg Med Chem Lett 23:4085
Choi HG, Zhang J, Deng X, Hatcher JM, Patricelli MP, Zhao Z, Aless DR, Gray NS (2012) Brain penetrant LRRK2 inhibitor. ACS Med Chem Lett 3:658
Zhang J, Deng X, Choi HG, Alessi DR, Gray NS (2012) Characterization of TAE684 as a potent LRRK2 kinase inhibitor. Bioorg Med Chem Lett 22:1864
Reith AD, Bamborough P, Jandu K, Andreotti D, Mensah L, Dossang P, Choi HG, Deng X, Zhang J, Alessi DR, Gray NS (2012) GSK2578215A; a potent and highly selective 2-arylmethyloxy-5-substitutent-N-arylbenzamide LRRK2 kinase inhibitor. Bioorg Med Chem Lett 22:5625
Liu M, Poulose S, Schuman E, Zaitsev AD, Dobson B, Auerbach K, Seyb K, Cuny GD, Glicksman MA, Stein RL, Yue Z (2010) Development of a mechanism-based high-throughput screen assay for leucine-rich repeat kinase 2-Discovery of LRRK2 inhibitors. Anal Biochem 404:186
Wager TT, Hou X, Verhoest PR, Villalobos A (2010) Moving beyond rules: the development of a central nervous system multiparameter optimization (CNS MPO) approach to enable alignment of druglike properties. ACS Chem Neurosci 1:435
Labbe C, Ross OA (2014) Associated studies of sporadic Parkinson’s disease in the genomic era. Curr Genomics 15:2
Hughes JD, Blagg J, Price DA, Bailey S, Decrscenzo GA, Devraj RV, Ellsworth E, Fobian YM, Gibbs ME, Gilles RW, Greene N, Huang E, Krieger-Burke T, Loesel J, Wager T, Zhang Y (2008) Physiochemical drug properties associated with in vivo toxicological outcomes. Bioorg Med Chem Lett 18:4872
Edwards AM, Isserlin R, Bader GD, Frye SV, Willson TM, Yu FH (2011) Too many roads not taken. Nature 470:163
Liou GY, Gallo KA (2009) New biochemical approaches towards understanding Parkinson’s disease-associated kinase, LRRK2 Biochem. J 424:e1
Covy JP, Giasson BI (2009) Identification of compounds that inhibit the kinase activity of leucine-rich repeat kinase 2. Biochem Biophys Res Commun 378:473
Davis MI, Hunt JP, Herrgard S, Ciceri P, Wodicka LM, Pallares G, Hocker M, Treiber DK, Zarrinkar PP (2011) Comprehensive analysis of kinase inhibitor selectivity. Nat Biotech 29:1046
Ramsden N, Perrin J, Ren Z, Lee BD, Zinn N, Dawson VL, Tam D, Bova M, Lang M, Drewes G, Bantscheff M, Bard F, Dawson TM, Hopf C (2011) Chemoproteomics-based design of potent LRRK2-selective lead compounds that attenuate Parkinson’s disease-related toxicity in human neurons. ACS Chem Biol 6:1021
Lee J, Song HJ, Koh JS, Lee HK, Kim Y, Chang S, Kim HW, Lim SH, Choi JS, Lim SH, Kim SW (2011) Patent Application WO2011060295
Baker-Glenn C, Burdick DJ, Chambers M, Chan BK, Chen H, Estrada, A, Guzner JL, Shore D, Sweeney ZK, Wang S, Zhao G (2011) Patent Application WO2011151360
Nicols PL, Eatherton AJ, Bamborough P, Jandu KJ, Phillips OJ, Andreotti D (2011) Patent Application WO2011038572
Galatsis P, Hayward MM, Kormos BL, Wager TT, Zhang L, Stepan AF, Henderson JL, Kurumbail RG, Verhoest PR (2014) Patent Application WO2014001973
Di L, Whitney-Pickett C, Umland JP, Zhang H, Zhang X, Gebhard DF, Lai Y, Federico JJ, Davidson RE, Smith R, Reyner EL, Lee C, Feng B, Rotter C, Varma MV, Kempshall S, Fenner K, El-kattan AF, Liston TE, Troutman MD (2011) Development of a new permeability assay using low-efflux MDCKII cells. J Pharm Sci 100:4974
Feng B, Mills JB, Davidson RE, Mireles RJ, Janiszewski JS, Troutman MD, de Morais SM (2008) In vitro P-glycoprotein assays to predict the in vivo interactions of P-glycoprotein with drugs in the central nervous system. Drug Metab Disposit 36:268
Finlayson K, Sharkey J (2004) A High-throughput binding assay for HERG. In: Yan Z, Caldwell GW (eds) Methods in pharmacology and toxicology. Humana, Totowa, pp 353–368
Greene N, Aleo MD, Louise-May S, Price DA, Will Y (2010) Using an in vitro cytotoxicity assay to aid in compound selection for in vivo safety studies. Bioorg Med Chem Lett 20:5308
Marroquin LD, Hynes J, Dykens JA, Jamieson JD, Will Y (2007) Circumventing the Crabtree effect: replacing media glucose with galactose increases susceptibility of HepG2 cells to mitochondrial toxicants. Toxicol Sci 97:539
Dundee Panel Data (2014) http://www.kinase-screen.mrc.ac.uk/kinase-inhibitors. Accessed 2 June 2014
Okerberg ES, Wu J, Zhang B, Samii B, Blackford K, Winn DT, Shreder KR, Burbaum JJ, Patricelli MP (2005) High-resolution functional proteomics by active-site peptide profiling. Proc Natl Acad Sci 102:4996
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer-Verlag Berlin Heidelberg
About this chapter
Cite this chapter
Galatsis, P., Henderson, J.L., Kormos, B.L., Hirst, W.D. (2014). Leucine-Rich Repeat Kinase 2 (LRRK2) Inhibitors. In: Hopkins, C. (eds) Novel Therapeutic Approaches to the Treatment of Parkinson’s Disease. Topics in Medicinal Chemistry, vol 18. Springer, Cham. https://doi.org/10.1007/7355_2014_69
Download citation
DOI: https://doi.org/10.1007/7355_2014_69
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-39924-9
Online ISBN: 978-3-319-39926-3
eBook Packages: Chemistry and Materials ScienceChemistry and Material Science (R0)