Abstract
Introduction
Amyotrophic lateral sclerosis (ALS) and Parkinson’s disease (PD) are two severe neurodegenerative disorders for which the disease mechanisms are poorly understood and reliable biomarkers are absent.
Objectives
To identify metabolite biomarkers for ALS and PD, and to gain insights into which metabolic pathways are involved in disease.
Methods
Nuclear magnetic resonance (NMR) metabolomics was utilized to characterize the metabolite profiles of cerebrospinal fluid (CSF) and plasma from individuals in three age, gender, and sampling-date matched groups, comprising 22 ALS, 22 PD and 28 control subjects.
Results
Multivariate analysis of NMR data generated robust discriminatory models for separation of ALS from control subjects. ALS patients showed increased concentrations of several metabolites in both CSF and plasma, these are alanine (CSF fold change = 1.22, p = 0.005), creatine (CSF-fc = 1.17, p = 0.001), glucose (CSF-fc = 1.11, p = 0.036), isoleucine (CSF-fc = 1.24, p = 0.002), and valine (CSF-fc = 1.17, p = 0.014). Additional metabolites in CSF (creatinine, dimethylamine and lactic acid) and plasma (acetic acid, glutamic acid, histidine, leucine, pyruvate and tyrosine) were also important for this discrimination. Similarly, panels of CSF-metabolites that discriminate PD from ALS and control subjects were identified.
Conclusions
The results for the ALS patients suggest an affected creatine/creatinine pathway and an altered branched chain amino acid (BCAA) metabolism, and suggest links to glucose and energy metabolism. Putative metabolic markers specific for ALS (e.g. creatinine and lactic acid) and PD (e.g. 3-hydroxyisovaleric acid and mannose) were identified, while several (e.g. creatine and BCAAs) were shared between ALS and PD, suggesting some overlap in metabolic alterations in these disorders.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Amyotrophic lateral sclerosis (ALS) and Parkinson’s disease (PD) are two relentless progressive adult-onset neurodegenerative disorders characterized by loss of specific groups of neurons and linked to protein aggregation. ALS is initially associated with a progressive degeneration of the upper and lower motor neurons, subsequently also affecting other parts of the nervous system (Kiernan et al. 2011) and leading to muscle atrophy and death within 2–5 years after onset (Tandan and Bradley 1985). PD is associated with the loss of dopaminergic neurons within the substantia nigra and is primarily manifested by loss of movement control (Lang and Obeso 2004).
ALS and PD are both protein misfolding diseases, involving either the formation of neuronal cytoplasmic inclusions containing TDP43 or misfolded SOD1 in different subtypes of ALS (Forsberg et al. 2010), or intra-neuronal α-synuclein-rich protein aggregates in PD (Goedert 2001). Although considerable research efforts have not yet revealed the exact details about the underlying disease mechanisms, there are likely mechanistic similarities with regard to protein homeostasis and aggregation, oxidative stress, mitochondrial dysfunction and autophagy (Knowles et al. 2014; Menzies et al. 2015; Lin and Beal 2006; Shaw 2005). Furthermore, emerging evidence suggests that there is a prion-like disease propagation involving SOD1 in ALS and α-synuclein in PD (Polymenidou and Cleveland 2011; Holmqvist et al. 2014; Ekhtiari et al. 2016). There are at present only symptomatic treatment strategies available for the two disorders, underlining a need for disease modifying treatments. Moreover, there is no unique diagnosis-specific biochemical test for the two diseases; instead a comprehensive clinical assessment of the patient is carried out according to the revised EFNS consensus guidelines for managing ALS (Andersen et al. 2012) or the United Kingdom Parkinson’s disease society brain bank (UK PDSBB) criteria (Gibb and Lees 1988). Despite the use of these criteria, it is still difficult to make a diagnosis, especially at an early stage, where incorrect diagnosis is common (Rajput et al. 1991). Robust biomarkers for these diseases would serve several critical functions; obtaining a straightforward and precise early diagnosis; monitoring the disease progression; and assessing therapeutics in clinical trials. In addition, metabolite biomarkers give important clues to the metabolic pathways and pathogenic mechanisms involved in the two diseases, and consequently may identify novel therapeutic targets.
One important field for biomarker discovery is metabolomics/metabonomics, an approach where the quantitative analysis of metabolites produced by a biological system can identify precise effects of a pathophysiological condition (Nicholson and Lindon 2008). Metabolomics usually relies on nuclear magnetic resonance (NMR) spectroscopy or mass spectrometry (MS) methods to identify and quantify metabolites, where NMR has the advantage of being quantitative, highly reproducible and straight-forward to use due to minimal sample-handling, while the higher sensitivity of MS can detect a greater number of metabolites from a given sample (Quinones and Kaddurah-Daouk 2009; Dunn et al. 2011). Solution NMR techniques are typically used for studying various biofluids, and the quantitative nature of NMR is particularly advantageous for result comparisons across different studies and laboratories. NMR measurements have shown excellent consistency between different laboratories and magnetic field strengths (Keun et al. 2002; Lindon et al. 2003). NMR metabolomics/metabonomics plays an important role in biomarker discovery, as previously reviewed (Smolinska et al. 2012a; Zhang et al. 2012), and may also provide details about the crosstalk between the metabolomes of the host and gut microbes (Li et al. 2011).
Extensive characterization of the metabolomes in CSF and blood (Sinclair et al. 2010; Smolinska et al. 2012b; Nicholson et al. 1995; Mandal et al. 2012; Psychogios et al. 2011; Wishart et al. 2008), as well as in urine and saliva (Want et al. 2010; Bouatra et al. 2013; Dame et al. 2015), has facilitated the creation of the human metabolome database (Wishart et al. 2013). Previous NMR- and MS-metabolomic studies of ALS- and PD-patients have described metabolic changes within these groups compared to control groups, both in cerebrospinal fluid (CSF) and blood (serum/plasma) (Jove et al. 2014). Since these two neurodegenerative protein misfolding diseases share common characteristics, it is surprising that, to date, no study has included both ALS and PD patients for direct and detailed comparisons.
In the present study, solution NMR metabolomics was used to characterize the metabolome in CSF and plasma from individuals in three age, gender, and sampling-date matched groups, comprising ALS patients, PD patients, and a control group. This stringent study design provides a unique possibility to identify common and specific metabolic markers for ALS and PD.
2 Materials and methods
2.1 Participants
Participants were selected from the neurobiobank at the Department of Pharmacology and Clinical Neuroscience, Umeå University Hospital, Umeå, Sweden. This biobank provided CSF and plasma samples from 72 age, gender and sampling-date matched participants divided into three groups: the ALS and the PD disease groups and the control group, see Table 1 for group characteristics. Each individual in one disease group has a matched individual in the other disease group, as well as one or two matched control subjects, enabling direct pair-wise comparisons, and reducing known confounds of metabolite analysis, specifically gender, age of patient and age of samples (see Table S1). The 22 ALS patients (male/female: 15/7) were diagnosed according to the revised EFNS Consensus Guidelines for Managing ALS criteria (Andersen et al. 2012), and the 22 PD patients (male/female: 15/7) according to UK PDSBB criteria (Gibb and Lees 1988). Patients were included only if the diagnosis was definite according to the above criteria and if both plasma and CSF were available and had been collected at the same time. Included PD patients had a mean disease duration from symptom onset to collection of samples of 32.5 ± 28.7 months (range 6–97), a mean UPDRSIII score [unified Parkinson’s disease rating scale (motor score); (Fahn et al. 1987)] of 23.2 ± 11.7 (range 5–48), and a median Hoehn-Yahr score of 2 (range 1–3). The 28 subjects (male/female: 19/9) in the control group were, with one exception, patients with disorders other than neurodegenerative diseases (headache and pain 8, disturbed balance 3, gait disturbance 3, Guillain–Barre syndrome 3, Kennedy’s disease 1, another 10 diagnoses were represented by a single individual).
2.2 Sample collection and preparation
CSF was collected through lumbar puncture according to a standard operating procedure, and plasma from non-fasted subjects was obtained from venous blood, using ethylenediaminetetraacetic acid (EDTA) as an anti-coagulant (Otto et al. 2012). Samples were stored at −80 °C until NMR analysis. Samples were thawed at room temperature and thereafter gently shaken to avoid gradient effects. Each CSF NMR-sample was prepared by mixing 100 µl of CSF with 80 µl of NMR buffer. The NMR buffer consists of a sodium phosphate buffer (Scharlau, Germany), pH 7.4, that ensures stable pH, D2O (Cambridge Isotope Laboratories, USA) for providing an NMR lock signal, and sodium-3-trimethylsilylpropionate-2,2,3,3-D4 (TMSP; Cambridge Isotope Laboratories, USA) as an internal reference for alignment and quantification. The resulting sample consisted (apart from CSF) of 50 mM sodium-phosphate, pH 7.4, 10 % D2O and 0.25 mM TMSP. Preparation of the plasma samples were carried out according to the standard operating procedure provided by Bruker Biospin. Briefly, plasma (90 µl) was mixed with equal amount of buffer (90 µl), giving a sample with 37.5 mM sodium phosphate buffer, pH 7.4, 10 % D2O, 0.02 % sodium azide and 2.3 mM TMSP. Finally, each 180 µl sample (CSF and plasma) was briefly centrifuged at 13000g for 30 s (to sediment particulates) before being transferred to a 5 mm heavy-walled NMR-tube (Wilmad-LabGlass, USA).
2.3 NMR spectroscopy and processing
NMR experiments were carried out on a Bruker DRX600 spectrometer, equipped with a 5 mm triple-resonance z-gradient cryo-probe. Additional 2D TOCSY (total correlation spectroscopy, tmix = 60 ms) and NOESY (nuclear Overhauser-effect spectroscopy, tmix = 100 ms) spectra for confirming resonance assignments were carried out on a Bruker 850 MHz Avance III HD spectrometer, equipped with a 5 mm triple-resonance z-gradient cryo-probe. TOPSPIN, version 2.1 or 3.0, was used for spectrometer control and data processing (Bruker Biospin).
For the CSF samples NMR spectra were recorded at 25º C using 1D 1H and 1H Carr-Purcell-Meiboom-Gill (CPMG) experiments, a spectral width of 14 ppm, 16 k data points, excitation sculpting for water suppression, a 200 ms mixing time for the CPMG filter, 192 transients, and 4 s relaxation delay. The CPMG filter was used to suppress broad resonances originating from large molecules, resulting in a significantly improved baseline and avoiding the use of filtration or extraction methods. However, small effects on the signal intensities of the metabolites is unavoidable, as previously described (Jupin et al. 2013, 2014), but these are consistent across samples. The robust excitation sculpting technique was used as this significantly improved the water suppression and baseline, and thus indirectly improved the quantification procedure. However, this technique affects the signal height of resonances in a limited region close to the water resonance (Aranibar et al. 2006; Jupin et al. 2014). For the metabolites that have resonances close to this region (ascorbate, glucose, mannose and pyroglutamate) quantification was carried out using resonances from those metabolites in other spectral regions, thereby no quantification errors due to the excitation sculpting technique were introduced. Spectra were processed with 32 k data points, an exponential window function with 0.5 Hz line broadening, a polynomial base-line correction, and aligned with the TMSP-signal.
NMR data acquisition for the plasma samples were carried out at 37 °C, using the NMR experiments recommended by Bruker. The statistical analysis described below was carried out on data recorded by using a 1D NOESY experiment (Bruker pulse program noesygppr1d), 128 transients, a recycle delay of 4.0 s and a mixing time of 60 ms. Spectra were processed with 32 k data points, an exponential window function with 1 Hz line broadening, a polynomial base-line correction, and aligned using the α-glucose CH2 resonance at 5.25 ppm.
2.4 Spectral analysis
The processed CSF spectra were subjected to a targeted analysis; identification and quantification of metabolites was carried out using the Chenomx NMR Suite software package, version 7.6 (Chenomx Inc., Edmonton, Canada), in which an advanced line-fitting procedure, combined with an extensive database containing NMR reference spectra of 312 metabolites is used, and where the internal TMSP resonance is used as a reference. Each spectrum was analysed at two separate occasions to avoid erroneous identification and quantification. Metabolite identities were confirmed through the use of AMIX (Bruker Biospin), the Human Metabolome Database (Wishart et al. 2013) and literature (Nicholson et al. 1995), as well as through the acquired and analyzed 2D NMR spectra.
A targeted analysis of the plasma spectra was not applicable due to the interference from strong and broad resonances from lipids and proteins. Although the use of a CPMG experiment in part could resolve this problem (Jupin et al. 2013), there were still strong lipid/protein resonances present, interfering with the Chenomx quantification procedure. Instead, an untargeted analysis approach was applied, in which the plasma spectra were reduced to spectral bins, using AMIX (Bruker Biospin), followed by a comparison using multivariate statistics. The spectral region 0.5–9 ppm was included without normalization and divided into bins of 0.004 ppm, excluding the regions where EDTA gives disturbing signals (3.70–3.58, 3.25–3.08, 2.74–2.69 and 2.62–2.55 ppm) and water (5.08–4.37 ppm) (Barton et al. 2010). Furthermore, metabolites are known to bind to albumin, which may potentially interfere with the analyses (Jupin et al. 2013, 2014). Since the plasma is diluted with equal amount of buffer (2 times dilution), and the albumin level within this cohort is homogenous (43 ± 4 mg/ml), it is unlikely that albumin affects the analysis.
2.5 Statistical analysis
Multivariate data analysis (ideally suited for analysis of multiple variables in a complex interaction network) was carried out both on the quantitative metabolite data in CSF, as well as on the binned data from plasma, using SIMCA 13.0 (Umetrics, Umeå, Sweden). Datasets were mean-centered, and scaled (pareto or unit variance), and subsequently subjected to principal component analysis (PCA) to obtain an overview of the data, and orthogonal projection to latent structures discriminant analysis (OPLS-DA) (Trygg and Wold 2002; Bylesjö et al. 2006) to find metabolites that separate the groups. This cohort was additionally designed to allow pair-wise comparisons, and with a new statistical method denoted orthogonal projection to latent structures effect projection (OPLS-EP) analysis, such a pair-wise comparison can now be performed on a multivariate basis (Jonsson et al. 2015). OPLS-DA is excellent for identifying differences among groups and to identify biomarkers, while the matched-pair comparison in the OPLS-EP analysis provides further details on the metabolic effects and mechanisms potentially caused by the disorder (i.e. hidden amongst factors biased by age, gender or sample storage). The final OPLS-DA models were calculated with mean-centering and UV scaling. The OPLS-EP analysis was performed without mean-centering but with UV-scaled datasets; in the cases where dual controls are available, these are averaged before analysis. Derived OPLS-DA models were cross-validated by leaving matched groups out, and for OPLS-EP models the sevenfold cross-validation method was used. The OPLS-EP and OPLS-DA models obtained in CSF were calculated with 1 + 0 + 0 and 1 + 1 + 0 components, respectively, while in plasma 1 + 1 + 0 components were used in both cases. R2X and R2Y are the fractions of the explained variation for the X and Y component, respectively. Q2 indicates the goodness of a prediction and describes the fraction of the total variation predicted by a component, as estimated by cross-validation. Group separations were evaluated through CV-ANOVA (Eriksson et al. 2008) and important metabolites were identified through the first weight vector (w*[1]). Cut-off values of w*[1] were determined empirically by iterative model building to obtain the most significant model. In the binning analysis of the plasma spectra, bins above the w*[1] threshold were translated, via their chemical shifts, into the corresponding metabolite. Quantified metabolites in CSF (from the two disease groups and controls) are further visualized by boxplots, and analyzed by univariate statistics using Wilcoxon rank-sum tests. Pathway analysis was carried out by using the Metaboanalyst (Xia et al. 2012) and HumanCyc web servers (Romero et al. 2005).
3 Results
3.1 Analysis of the CSF metabolome
In the targeted analysis of the CSF data a total of 60 metabolites were unambiguously identified, 38 of which were quantified in every sample and used for subsequent modeling and disease profiling. These results are compiled in the Supplemental Table S2, where average metabolite concentrations for each group, fold-changes and the results from Wilcoxon rank-sum tests have been listed. The spread of concentrations within the ALS, PD, and control groups, are visualized with box-plots in Supplemental Fig. S1. The detected range of metabolite concentrations are in excellent agreement with previous studies of the CSF metabolome (Mandal et al. 2012; Wishart et al. 2008; Stoop et al. 2010). For lactate, a metabolite routinely monitored in all groups by standard clinical chemistry analysis (using the Cobas LACT2 reagent on Cobas C502 analyzers), the agreement is also excellent; NMR gives 1.50 ± 0.22, 1.37 ± 0.22 and 1.29 ± 0.30 mM lactate for the ALS, PD and control groups, respectively, while the corresponding results from the accredited laboratory at Norrlands University Hospital, Umeå, Sweden, were 1.60 ± 0.30, 1.44 ± 0.24 and 1.36 ± 0.23 mM.
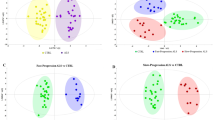
A multivariate overview using PCA revealed no significant outliers, hence all observations were kept for further analysis. The ALS- and the control-groups were discriminated by the OPLS-EP analysis (R2X = 0.29, R2Y = 0.83, Q2 = 0.44, p = 0.027), while the separation with OPLS-DA was not significant. From the significant OPLS-EP model, variables with weight values |w*| > 0.2 were selected for a second round of modelling. Included metabolites were alanine, creatine, creatinine, dimethylamine, glucose, isoleucine, lactate and valine, see Fig. 1a. The resulting OPLS-EP (R2X = 0.65, R2Y = 0.73, Q2 = 0.60, p = 0.002) and OPLS-DA (R2X = 0.55, R2Y = 0.43, Q2 = 0.26, p = 0.008) models provided better group separation and statistically significant models. The cross-validated score plot from the OPLS-EP model, Fig. 1b, showed that most of the matched pairs were driving the model in the same direction. The group separation obtained by using OPLS-DA on the same eight metabolites is shown in Fig. 1c. The score values for both the OPLS-EP and OPLS-DA models as a function of match number are shown in Supplemental Fig. S2A and B. For seven of these eight metabolites (all except creatinine) univariate analysis also identifies considerable concentration changes (p < 0.05).
Multivariate statistical comparison of CSF from ALS patients and control subjects. a Metabolites, with weight values |w*| > 0.2, important for separation between the ALS and control groups. b Cross-validated OPLS-EP score plot of the first component, tcv[1]. c Cross-validated OPLS-DA score plot of the first component, tcv[1] (ALS patients: filled circles; control subjects: open circles)
The PD and the control groups were compared by OPLS-EP and OPLS-DA modelling using all metabolites. No statistically significant models were obtained that could discriminate between the groups. Metabolites with |w*| > 0.15 in the initial OPLS-EP models were selected for a second round of modeling; 15 metabolites fulfilled this criterion, see Fig. 2a. Using this subset of metabolites, a statistically significant OPLS-EP model (R2X = 0.36, R2Y = 0.39, Q2 = 0.27, p = 0.042) was obtained. The cross-validated score plot for this OPLS-EP model is shown in Fig. 2b, and the scores for each matched pair in Supplemental Fig. S3. Smaller subsets of metabolites were tested, but did not improve the discrimination between the PD and control groups (data not shown).
When comparing the ALS group to the PD group using OPLS-EP and OPLS-DA (with all variables), both methods were able to separate between the ALS and the PD group; however, none of the models were significant according to CV-ANOVA. A second round of modeling was carried out using metabolites with weight values |w*| > 0.2. In total, 10 metabolites fulfilled this criterion, see Fig. 3a. The resulting OPLS-EP (R2X = 0.28, R2Y = 0.57, Q2 = 0.42, p = 0.004) and OPLS-DA (R2X = 0.34, R2Y = 0.51, Q2 = 0.34, p = 0.002) models provided discriminatory ability between the ALS and PD groups and both models were statistically significant. Figure 3b shows the cross-validated score plot from the OPLS-EP model, while Fig. 3c shows the scores from the OPLS-DA model. The scores for matched pairs are shown in Supplemental Fig S4.
Multivariate statistical comparison of CSF from ALS and PD patients. a Metabolites, with weight values |w*| > 0.2, important for separation between the ALS and the PD groups. b Cross-validated OPLS-EP score plot of the first component, tcv[1]. c Cross-validated OPLS-DA score plot of the first component, tcv[1] (ALS patients: filled circles; PD patients: open diamonds)
3.2 Analysis of the plasma metabolome
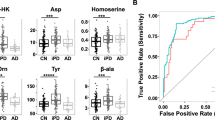
For plasma, the derived spectral bins were analyzed statistically using OPLS-EP and OPLS-DA to identify metabolites capable of discriminating between the ALS, PD and control groups. One sample was excluded from the analysis, since the initial PCA analysis identified it as an outlier caused primarily by highly elevated levels of glucose. Statistically significant models were obtained between the ALS group and the control group using either the OPLS-EP (R2X = 0.37, R2Y = 0.85, Q2 = 0.52, p = 0.010) or the OPLS-DA approach (R2X = 0.32, R2Y = 0.64, Q2 = 0.30, p = 0.004). OPLS-EP variables with weight values |w*| > 0.03 were translated into the corresponding metabolites, shown in Fig. 4a; the OPLS-EP and OPLS-DA scores of these models are shown in Fig. 4b and C. The score values as a function of the matched pair number are shown in the Supplement, Fig. S2C and D. No statistically significant models were obtained for discriminating the PD group from either the control or the ALS groups.
Multivariate statistical comparison of plasma from ALS patients and control subjects. a Metabolites, with weight values |w*| > 0.03, important for the separation between the ALS and the control groups. b Cross-validated OPLS-EP score plot of the first component, tcv[1]. c Cross-validated OPLS-DA score plot of the first component, tcv[1] (ALS patients: filled circles; control subjects: open circles)
4 Discussion
For both ALS and PD we lack a detailed understanding of the underlying disease mechanisms, diagnostic tests and effective disease modifying treatments. It is consequently of great interest to find biomarkers that can support the diagnosis procedure, monitor the disease progression and provide insight about the molecular mechanisms involved. In the present work, CSF and plasma from rigorously matched patients with ALS and PD were analyzed by solution NMR metabolomics, and the obtained metabolite profiles were compared with those of matched control subjects. Multivariate statistical analysis was primarily carried out by using the OPLS-EP and the OPLS-DA approaches. Noteworthy is that the OPLS-EP analysis of this pair-wise matched cohort in general provided stronger models than the OPLS-DA analysis of the aggregate groups. The reason may be that the unpaired group discrimination analysis (i.e. OPLS-DA) may have difficulties in detecting subtle disease-associated variation due to variations from other sources such as gender, age or sample-age within the groups (Jonsson et al. 2015).
Distinct metabolite differences between ALS patients and controls were observed both in CSF and plasma, while robust group discrimination between PD patients and controls, as well as between ALS and PD patients, were only detected in CSF. The metabolites capable of separating these groups are summarized in Table 2, together with affected metabolic pathways and/or physiological functions, and the results are discussed below.
4.1 ALS patients compared to control subjects
In the quantitative and targeted multivariate analysis of CSF metabolites, a clear separation between the ALS group and the control group was observed. Judged by the weight values of the multivariate statistical analysis, eight metabolites were of particular significance, see Fig. 1 and Table 2. The univariate statistical analysis showed that seven of those (all except creatinine) had p values below 0.05. In plasma the multivariate analysis showed that thirteen metabolites are of particular importance (with weight values |w*| > 0.03) for the separation between ALS patients and control subjects, see Fig. 4 and Table 2. Five of the metabolites were found at elevated levels in both CSF and plasma, these were alanine (CSF fold change = 1.22, p = 0.005), creatine (CSF-fc = 1.17, p = 0.001), glucose (CSF-fc = 1.11, p = 0.036), isoleucine (CSF-fc = 1.24, p = 0.002), and valine (CSF-fc = 1.17, p = 0.014), see Table S2.
Among the ALS patients, the CSF and plasma concentrations of creatine were found elevated when compared to the control group, and together with the decreased concentration of creatinine in CSF, this suggest that the creatinine metabolism pathway is disturbed in ALS. These results are in line with several metabolomics studies, in which equivalent changes of the creatine/creatinine levels have been observed in ALS patients (Lawton et al. 2014, 2012; Wuolikainen et al. 2011, 2016). It is however noteworthy that the present NMR study, together with the very recent MS study on the same cohort (Wuolikainen et al. 2016), for the first time have detected increased levels of creatine in CSF. Reduced creatinine level has previously been correlated with ALS both in CSF and blood (Paillisse et al. 2005; Wuolikainen et al. 2011). Since creatinine is primarily formed in the skeletal muscles and the level of CSF creatinine is known to depend on the plasma level (Tachikawa et al. 2008), the reduced CSF level may be linked to the wasting of skeletal muscle mass and not to the neurodegenerative process per se in the central nervous system (CNS). However, since much more creatinine is produced by the non-enzymatic degradation of phosphocreatine than from creatine (Wyss and Kaddurah-Daouk 2000), this altered level in CSF of ALS patients may also be indicative of a compromised creatine to phosphocreatine conversion by creatine kinase within the CNS.
Another group of metabolites, displaying increased concentrations in CSF and plasma from ALS-patients, were the branched-chain amino acids (BCAA) isoleucine and valine (Figs. 1a and 4a; Table 2). The BCAA leucine also has elevated concentration in plasma (Fig. 4a; Table 2). In CSF, leucine is found just below the cutoff value (|w*| = 0.2), with a |w*| = 0.18 (Table S2). These three metabolites are all found in the valine, leucine and isoleucine biosynthesis/biodegradation pathways, and present findings suggest that the ALS disorder may affect these pathways. The BCAAs are also known to affect brain function via an altered neurotransmitter biosynthesis (Fernstrom 2013). Several studies have previously correlated an increased concentration of BCAA, obtained primarily through a diet designed to increase muscle growth, with an accelerated progression of ALS (Beghi 2013; Manuel and Heckman 2011). Interestingly, the BCAA leucine is a key regulator of the mechanistic target of rapamycin complex 1, a key enzyme involved in protein synthesis and degradation (Averous et al. 2014; Jewell et al. 2015). The MS-based metabolomic study on the same cohort observed similar changes in the BCAA levels, thereby supporting these findings (Wuolikainen et al. 2016).
The CSF-levels of dimethylamine are elevated in the ALS patient group when compared to the control group, are therefore important for separation of the two groups statistically (Fig. 1; Table 2). Dimethylamine is produced from asymmetric dimethylarginine (ADMA) via the enzyme dimethylarginine dimethylaminohydrolase 1 (MacAllister et al. 1996). ADMA is an endogenous inhibitor of nitric oxide synthase (Vallance and Leiper 2004), and consequently dimethylamine can be linked to the synthesis of nitric oxide and the regulation of oxidative stress. A previous report in fact observed reduced plasma concentration of ADMA in patients with the ALS disorder (Isobe et al. 2010). A reduced level of ADMA in plasma from ALS patients was also found in the same patient cohort as analyzed here, by MS-based metabolomics (Wuolikainen et al. 2016). Hence, the elevated levels of dimethylamine in CSF combined with reports of reduced levels of ADMA in plasma together indicate perturbations in this pathway in ALS. Since the regulation of oxidative stress is complex and involves other pathways than the one described above, the roles of dimethylamine and ADMA as potential markers for ALS need to be further evaluated in larger patient cohorts.
The increased concentrations of alanine and glucose detected in CSF and plasma from ALS patients, as well as an elevated concentration of lactic acid in CSF, suggest an affected glucose metabolism (Fig. 1 and 4; Table 2) (Tsacopoulos and Magistretti 1996). Elevated levels of glucose and lactic acid in CSF were also detected in a recent longitudinal study (Gray et al. 2015). The observed changes for alanine and glucose are in agreement with the recent MS-based study on the same patient cohort (Wuolikainen et al. 2016), while two other studies did not observe similar changes (Blasco et al. 2010, 2013).
In plasma, the metabolites acetic acid, glutamate, pyruvate, histidine and tyrosine showed increased concentrations (Fig. 4a; Table 2). Elevated CSF- and plasma-levels of glutamate and pyruvate have previously been observed in ALS patients (Blasco et al. 2010, 2013; Lawton et al. 2012; Kumar et al. 2010; Wuolikainen et al. 2016). The increased level of glutamate in plasma/serum has previously been attributed to either the glutamate excitotoxicity in ALS or to an imbalance in the glutamate/glutamine conversion (Kumar et al. 2010). This study (Kumar et al. 2010) did however not observe any significant concentration changes of the aromatic amino acid and neurotransmitter precursor tyrosine, while the antioxidant metabolite histidine in contrast showed a decreased level. Although tyrosine and histidine can be linked to neurotransmission and the reduction of oxidative stress (Wade and Tucker 1998; Milewski et al. 2014), there is to our knowledge no previous direct association with ALS.
Furthermore, we observed increased plasma levels of lipids and triglycerides in ALS patients in comparison to control subjects; changes of the same group of metabolites have been observed in MS-based metabolomic studies of plasma (Lawton et al. 2014, 2012; Wuolikainen et al. 2016). This highlights the need for further studies of the lipidomic metabolic profile of ALS patients.
4.2 PD patients compared to control subjects
In order to distinguish PD patients from control subjects, a set of 15 CSF-metabolites were used together with the OPLS-EP analysis, see Fig. 2 and Table 2. The modelling statistics were poorer than in the comparison between the ALS and control groups; this is most likely due to a great variation among the patients in severity of motor impairment (UPDRSIII motor score 23.2 ± 11.7, range 5–48). Attempts to use subsets of these 15 metabolites did not improve the discrimination. The difficulty in obtaining statistically significant models emphasizes the need for some caution when interpreting the results.
Decreased concentration of 3-hydroxyisovalerate in the CSF of PD patients has previously been observed (Öhman and Forsgren 2015; Trupp et al. 2014), and can potentially be linked to tyrosine degradation (Trupp et al. 2014). In agreement with a previous report (Öhman and Forsgren 2015), the metabolite mannose, involved in glucose metabolism (Sharma et al. 2014), also has a decreased concentration in the CSF of PD patients. Alanine, dimethylamine and glucose all showed an increased concentration, contrary to several studies (Lewitt et al. 2013; Mally et al. 1997; Molina et al. 1997; Öhman and Forsgren 2015). The BCAA isoleucine, leucine and valine were all found at higher concentrations in the PD patients, similar to the ALS patient group above. Elevated levels of BCAAs in PD patients have previously been observed (Van Sande et al. 1971; Lakke and Teelken 1976; Lakke et al. 1987), although the opposite has also been reported (Molina et al. 1997). As described above, BCAAs are linked in biosynthesis and degradation pathways as well as to neurotransmitter biosynthesis of relevance for the PD pathology. We have also detected altered levels of degradation products from leucine including 3-hydroxyisovaleric acid and acetoacetic acid (Table 2), supporting the model of disrupted leucine regulation in PD. The reduced concentration of acetoacetic acid may also implicate an effect on ketone body synthesis or degradation. Two metabolites with increased concentration in CSF were tyrosine and histidine; they can be linked to neurotransmission and the reduction of oxidative stress (Wade and Tucker 1998; Milewski et al. 2014). The four remaining metabolites, citric acid, creatine, dimethylsulfone and fructose, all display elevated levels in PD patients when compared to controls. Dimethylsulfone is involved in the methanethiol pathway, in which it is produced as a breakdown product of the nitric oxide synthase cofactor methanethiol (Engelke et al. 2005). Fructose is linked to the mannose and fructose pathway, and citric acid to numerous metabolic pathways; future studies of larger patient cohorts will clarify their importance.
4.3 ALS patients compared to PD patients
Multivariate analysis showed that 10 CSF-metabolites could clearly distinguish ALS patients from PD patients, see Fig. 3 and Table 2. As described above creatine, dimethylamine and lactic acid were important to discriminate ALS patients from controls; here it is further shown that these metabolites were also discriminatory between ALS and PD patients. This is likely due to the similarity of the metabolic profiles for the PD and control groups, as indicated by their poor separation in the multivariate models. The two metabolites acetone and acetoacetic acid were shown to be important for the separation between ALS and PD; both metabolites are ketone bodies, and involved in pathways known to play roles in both ALS and PD (Zhao et al. 2006; Paoli et al. 2014). Acetone, acetoacetic acid, 2-hydroxybutyric acid, lactic acid and isopropyl alcohol are all involved in the propanoate/propionic acid metabolism, and therefore implicate a difference in this pathway. 2-hydroxybutyric acid has previously been identified as an early marker for diabetes (Gall et al. 2010), suggesting that this is another indication of altered glucose metabolism in ALS. Lactic acid is together with formic acid also involved in pyruvate metabolism. An elevated level of ascorbic acid has previously been observed in CSF from ALS patients (Blasco et al. 2010, 2013); both ascorbic acid and myo-inositol are part of the ascorbate and aldarate metabolism, which may reflect a difference in this pathway between ALS and PD patients.
5 Concluding remarks
The present NMR metabolomics study revealed significant differences in the metabolic profiles of ALS patients compared to the profiles of age, gender, and sampling-date matched control subjects and PD patients. The discrimination between PD patients and controls is however less significant. Several metabolites have been identified as potential biomarkers for ALS, many of which have been linked to specific metabolic pathways involved in the ALS pathophysiology (Table 2). Interestingly there are both shared and unique metabolic markers for the ALS and PD disorders. Although these findings in part are validated by previous studies and by the very recent MS-based study on the same patient cohort (Wuolikainen et al. 2016), further validation of the identified metabolic markers is necessary using larger patient cohorts.
Abbreviations
- ALS:
-
Amyotrophic lateral sclerosis
- PD:
-
Parkinson’s disease
- NMR:
-
Nuclear magnetic resonance
- MS:
-
Mass spectrometry
- EDTA:
-
Ethylenediaminetetraacetic acid
- CSF:
-
Cerebrospinal fluid
- CPMG:
-
Carr-Purcell-Meiboom-Gill
- PCA:
-
Principal component analysis
- OPLS:
-
Orthogonal projection to latent structures
- DA:
-
Discriminant analysis
- EP:
-
Effect projection
- CV-ANOVA:
-
Analysis of variance testing of cross-validated predictive residuals
- CNS:
-
Central nervous system
- BCAA:
-
Branched chain amino acids
References
Andersen, P. M., Abrahams, S., Borasio, G. D., de Carvalho, M., Chio, A., Hardiman, O., et al. (2012). EFNS guidelines on the clinical management of amyotrophic lateral sclerosis (MALS)–revised report of an EFNS task force. European Journal of Neurology, 19(3), 360–375. doi:10.1111/j.1468-1331.2011.03501.x.
Aranibar, N., Ott, K. H., Roongta, V., & Mueller, L. (2006). Metabolomic analysis using optimized NMR and statistical methods. Analytical Biochemistry, 355(1), 62–70. doi:10.1016/j.ab.2006.04.014.
Averous, J., Lambert-Langlais, S., Carraro, V., Gourbeyre, O., Parry, L., B’Chir, W., et al. (2014). Requirement for lysosomal localization of mTOR for its activation differs between leucine and other amino acids. Cellular Signalling, 26(9), 1918–1927. doi:10.1016/j.cellsig.2014.04.019.
Barton, R. H., Waterman, D., Bonner, F. W., Holmes, E., Clarke, R., Procardis, C., et al. (2010). The influence of EDTA and citrate anticoagulant addition to human plasma on information recovery from NMR-based metabolic profiling studies. Molecular BioSystems, 6(1), 215–224. doi:10.1039/b907021d.
Beghi, E. (2013). Are professional soccer players at higher risk for ALS? Amyotroph Lateral Scler Frontotemporal Degener, 14(7–8), 501–506. doi:10.3109/21678421.2013.809764.
Blasco, H., Corcia, P., Moreau, C., Veau, S., Fournier, C., Vourc’h, P., et al. (2010). 1H-NMR-based metabolomic profiling of CSF in early amyotrophic lateral sclerosis. PLoS ONE, 5(10), e13223. doi:10.1371/journal.pone.0013223.
Blasco, H., Corcia, P., Pradat, P. F., Bocca, C., Gordon, P. H., Veyrat-Durebex, C., et al. (2013). Metabolomics in cerebrospinal fluid of patients with amyotrophic lateral sclerosis: an untargeted approach via high-resolution mass spectrometry. Journal of Proteome Research, 12(8), 3746–3754. doi:10.1021/pr400376e.
Bouatra, S., Aziat, F., Mandal, R., Guo, A. C., Wilson, M. R., Knox, C., et al. (2013). The human urine metabolome. PLoS ONE, 8(9), e73076. doi:10.1371/journal.pone.0073076.
Bylesjö, M., Rantalainen, M., Cloarec, O., Nicholson, J. K., Holmes, E., & Trygg, J. (2006). OPLS discriminant analysis: combining the strengths of PLS-DA and SIMCA classification. Journal of Chemometrics, 20(8–10), 341–351. doi:10.1002/cem.1006.
Dame, Z. T., Aziat, F., Mandal, R., Krishnamurthy, R., Bouatra, S., Borzouie, S., et al. (2015). The human saliva metabolome. Metabolomics, 11(6), 1864–1883. doi:10.1007/s11306-015-0840-5.
Dunn, W. B., Broadhurst, D. I., Atherton, H. J., Goodacre, R., & Griffin, J. L. (2011). Systems level studies of mammalian metabolomes: the roles of mass spectrometry and nuclear magnetic resonance spectroscopy. Chemical Society Reviews, 40(1), 387–426. doi:10.1039/b906712b.
Engelke, U. F., Tangerman, A., Willemsen, M. A., Moskau, D., Loss, S., Mudd, S. H., et al. (2005). Dimethyl sulfone in human cerebrospinal fluid and blood plasma confirmed by one-dimensional (1)H and two-dimensional (1)H-(13)C NMR. NMR in Biomedicine, 18(5), 331–336. doi:10.1002/nbm.966.
Eriksson, L., Trygg, J., & Wold, S. (2008). CV-ANOVA for significance testing of PLS and OPLS (R) models. Journal of Chemometrics, 22(11–12), 594–600. doi:10.1002/cem.1187.
Ekhtiari Bidhendi, E., Bergh, J., Zetterström, P., Andersen, P. M., Marklund, S. L., & Brännström, T. (2016). Two superoxide dismutase prion strains transmitting amyotrophic lateral sclerosis. The Journal of Investigation (in press).
Fahn, S., Elton, R., & Commitee TUD. (1987). Unified Parkinson’s disease rating scale. In S. Fahn, C. Marsden, D. Calne, & M. Goldstein (Eds.), Recent developments in Parkinson’s disease (pp. 153–163). Florham Park: Macmillan Healthcare Information.
Fernstrom, J. D. (2013). Large neutral amino acids: dietary effects on brain neurochemistry and function. Amino Acids, 45(3), 419–430. doi:10.1007/s00726-012-1330-y.
Forsberg, K., Jonsson, P. A., Andersen, P. M., Bergemalm, D., Graffmo, K. S., Hultdin, M., et al. (2010). Novel antibodies reveal inclusions containing non-native SOD1 in sporadic ALS patients. PLoS ONE, 5(7), e11552. doi:10.1371/journal.pone.0011552.
Gall, W. E., Beebe, K., Lawton, K. A., Adam, K. P., Mitchell, M. W., Nakhle, P. J., et al. (2010). Alpha-hydroxybutyrate is an early biomarker of insulin resistance and glucose intolerance in a nondiabetic population. PLoS ONE, 5(5), e10883. doi:10.1371/journal.pone.0010883.
Gibb, W. R., & Lees, A. J. (1988). A comparison of clinical and pathological features of young- and old-onset Parkinson’s disease. Neurology, 38(9), 1402–1406.
Goedert, M. (2001). Alpha-synuclein and neurodegenerative diseases. Nature Reviews Neuroscience, 2(7), 492–501. doi:10.1038/35081564.
Gray, E., Larkin, J. R., Claridge, T. D., Talbot, K., Sibson, N. R., & Turner, M. R. (2015). The longitudinal cerebrospinal fluid metabolomic profile of amyotrophic lateral sclerosis. Amyotroph Lateral Scler Frontotemporal Degener, 16(1–8), 456–463. doi:10.3109/21678421.2015.1053490.
Holmqvist, S., Chutna, O., Bousset, L., Aldrin-Kirk, P., Li, W., Bjorklund, T., et al. (2014). Direct evidence of Parkinson pathology spread from the gastrointestinal tract to the brain in rats. Acta Neuropathologica, 128(6), 805–820. doi:10.1007/s00401-014-1343-6.
Isobe, C., Abe, T., & Terayama, Y. (2010). Decrease in asymmetrical dimethylarginine, an endogenous nitric oxide synthase inhibitor, in cerebrospinal fluid during elderly aging and in patients with sporadic form of amyotrophic lateral sclerosis. Neurosignals, 18(1), 43–48. doi:10.1159/000312527.
Jewell, J. L., Kim, Y. C., Russell, R. C., Yu, F. X., Park, H. W., Plouffe, S. W., et al. (2015). Metabolism differential regulation of mTORC1 by leucine and glutamine. Science, 347(6218), 194–198. doi:10.1126/science.1259472. (New York).
Jonsson, P., Wuolikainen, A., Thysell, E., Chorell, E., Stattin, P., Wikström, P., et al. (2015). Constrained randomization and multivariate effect projections improve information extraction and biomarker pattern discovery in metabolomics studies involving dependent samples. Metabolomics, 11(6), 1667–1678. doi:10.1007/s11306-015-0818-3.
Jove, M., Portero-Otin, M., Naudi, A., Ferrer, I., & Pamplona, R. (2014). Metabolomics of human brain aging and age-related neurodegenerative diseases. Journal of Neuropathology and Experimental Neurology, 73(7), 640–657. doi:10.1097/NEN.0000000000000091.
Jupin, M., Michiels, P. J., Girard, F. C., Spraul, M., & Wijmenga, S. S. (2013). NMR identification of endogenous metabolites interacting with fatted and non-fatted human serum albumin in blood plasma: fatty acids influence the HSA-metabolite interaction. J Mag Reson, 228, 81–94. (San Diego, Calif: 1997). doi:10.1016/j.jmr.2012.12.010.
Jupin, M., Michiels, P. J., Girard, F. C., Spraul, M., & Wijmenga, S. S. (2014). NMR metabolomics profiling of blood plasma mimics shows that medium- and long-chain fatty acids differently release metabolites from human serum albumin. J Mag Reson, 239, 34–43. (San Diego, Calif: 1997). doi:10.1016/j.jmr.2013.11.019.
Keun, H. C., Ebbels, T. M., Antti, H., Bollard, M. E., Beckonert, O., Schlotterbeck, G., et al. (2002). Analytical reproducibility in (1)H NMR-based metabonomic urinalysis. Chemical Research in Toxicology, 15(11), 1380–1386. doi:10.1021/tx0255774.
Kiernan, M. C., Vucic, S., Cheah, B. C., Turner, M. R., Eisen, A., Hardiman, O., et al. (2011). Amyotrophic lateral sclerosis. Lancet, 377(9769), 942–955. doi:10.1016/S0140-6736(10)61156-7.
Knowles, T. P. J., Vendruscolo, M., & Dobson, C. M. (2014). The amyloid state and its association with protein misfolding diseases. Nature Reviews Molecular Cell Biology, 15(6), 384–396. doi:10.1038/nrm3810.
Kumar, A., Bala, L., Kalita, J., Misra, U. K., Singh, R. L., Khetrapal, C. L., et al. (2010). Metabolomic analysis of serum by (1) H NMR spectroscopy in amyotrophic lateral sclerosis. Clinica Chimica Acta, 411(7–8), 563–567. doi:10.1016/j.cca.2010.01.016.
Lakke, J. P., & Teelken, A. W. (1976). Amino acid abnormalities in cerebrospinal fluid of patients with parkinsonism and extrapyramidal disorders. Neurology, 26(5), 489–493.
Lakke, J. P., Teelken, A. W., vd Voet, H., & Wolthers, B. (1987). Amino acid abnormalities in cerebrospinal fluid and blood serum of patients with Parkinson’s disease, other heredodegenerative disorders and head injuries. Advances in Neurology, 45, 243–247.
Lang, A. E., & Obeso, J. A. (2004). Challenges in Parkinson’s disease: restoration of the nigrostriatal dopamine system is not enough. Lancet Neurology, 3(5), 309–316. doi:10.1016/S1474-4422(04)00740-9.
Lawton, K. A., Cudkowicz, M. E., Brown, M. V., Alexander, D., Caffrey, R., Wulff, J. E., et al. (2012). Biochemical alterations associated with ALS. Amyotroph Lateral Scler, 13(1), 110–118. doi:10.3109/17482968.2011.619197.
Lawton, K. A., Brown, M. V., Alexander, D., Li, Z., Wulff, J. E., Lawson, R., et al. (2014). Plasma metabolomic biomarker panel to distinguish patients with amyotrophic lateral sclerosis from disease mimics. Amyotroph Lateral Scler Frontotemporal Degener, 15(5–6), 362–370. doi:10.3109/21678421.2014.908311.
Lewitt, P. A., Li, J., Lu, M., Beach, T. G., Adler, C. H., Guo, L., et al. (2013). 3-hydroxykynurenine and other Parkinson’s disease biomarkers discovered by metabolomic analysis. Movement Disorders, 28(12), 1653–1660. doi:10.1002/mds.25555.
Li, J. V., Ashrafian, H., Bueter, M., Kinross, J., Sands, C., le Roux, C. W., et al. (2011). Metabolic surgery profoundly influences gut microbial-host metabolic cross-talk. Gut, 60(9), 1214–1223. doi:10.1136/gut.2010.234708.
Lin, M. T., & Beal, M. F. (2006). Mitochondrial dysfunction and oxidative stress in neurodegenerative diseases. Nature, 443(7113), 787–795. doi:10.1038/nature05292.
Lindon, J. C., Nicholson, J. K., Holmes, E., Antti, H., Bollard, M. E., Keun, H., et al. (2003). Contemporary issues in toxicology the role of metabonomics in toxicology and its evaluation by the COMET project. Toxicology and Applied Pharmacology, 187(3), 137–146. doi:10.1016/S0041-008X(02)00079-0.
MacAllister, R. J., Parry, H., Kimoto, M., Ogawa, T., Russell, R. J., Hodson, H., et al. (1996). Regulation of nitric oxide synthesis by dimethylarginine dimethylaminohydrolase. British Journal of Pharmacology, 119(8), 1533–1540. doi:10.1111/j.1476-5381.1996.tb16069.x.
Mally, J., Szalai, G., & Stone, T. W. (1997). Changes in the concentration of amino acids in serum and cerebrospinal fluid of patients with Parkinson’s disease. Journal of the Neurological Sciences, 151(2), 159–162. doi:10.1016/S0022-510X(97)00119-6.
Mandal, R., Guo, A. C., Chaudhary, K. K., Liu, P., Yallou, F. S., Dong, E., et al. (2012). Multi-platform characterization of the human cerebrospinal fluid metabolome: a comprehensive and quantitative update. Genome Med, 4(4), 38. doi:10.1186/gm337.
Manuel, M., & Heckman, C. J. (2011). Stronger is not always better: could a bodybuilding dietary supplement lead to ALS? Experimental Neurology, 228(1), 5–8. doi:10.1016/j.expneurol.2010.12.007.
Menzies, F. M., Fleming, A., & Rubinsztein, D. C. (2015). Compromised autophagy and neurodegenerative diseases. Nature Reviews Neuroscience, 16(6), 345–357. doi:10.1038/nrn3961.
Milewski K, Hilgier W, Albrecht J, Zielinska M (2014). The dimethylarginine (ADMA)/nitric oxide pathway in the brain and periphery of rats with thioacetamide-induced acute liver failure: modulation by histidine. http://www.ncbi.nlm.nih.gov/pubmed/25523831.
Molina, J. A., Jimenez-Jimenez, F. J., Gomez, P., Vargas, C., Navarro, J. A., Orti-Pareja, M., et al. (1997). Decreased cerebrospinal fluid levels of neutral and basic amino acids in patients with Parkinson’s disease. Journal of the Neurological Sciences, 150(2), 123–127. doi:10.1016/S0022-510X(97)00069-5.
Nicholson, J. K., & Lindon, J. C. (2008). Systems biology: metabonomics. Nature, 455(7216), 1054–1056. doi:10.1038/4551054a.
Nicholson, J. K., Foxall, P. J. D., Spraul, M., Farrant, R. D., & Lindon, J. C. (1995). 750-Mhz H-1 and H-1-C-13 Nmr-spectroscopy of human blood-plasma. Analytical Chemistry, 67(5), 793–811. doi:10.1021/ac00101a004.
Öhman, A., & Forsgren, L. (2015). NMR metabonomics of cerebrospinal fluid distinguishes between Parkinson’s disease and controls. Neuroscience Letters, 594, 36–39. doi:10.1016/j.neulet.2015.03.051.
Otto, M., Bowser, R., Turner, M., Berry, J., Brettschneider, J., Connor, J., et al. (2012). Roadmap and standard operating procedures for biobanking and discovery of neurochemical markers in ALS. Amyotroph Lateral Scler, 13(1), 1–10. doi:10.3109/17482968.2011.627589.
Paillisse, C., Lacomblez, L., Dib, M., Bensimon, G., Garcia-Acosta, S., & Meininger, V. (2005). Prognostic factors for survival in amyotrophic lateral sclerosis patients treated with riluzole. Amyotroph Lateral Scler Other Motor Neuron Disord, 6(1), 37–44. doi:10.1080/14660820510027035.
Paoli, A., Bianco, A., Damiani, E., & Bosco, G. (2014). Ketogenic diet in neuromuscular and neurodegenerative diseases. Biomed Res Int, 2014, 474296. doi:10.1155/2014/474296.
Polymenidou, M., & Cleveland, D. W. (2011). The seeds of neurodegeneration: prion-like spreading in ALS. Cell, 147(3), 498–508. doi:10.1016/j.cell.2011.10.011.
Psychogios, N., Hau, D. D., Peng, J., Guo, A. C., Mandal, R., Bouatra, S., et al. (2011). The human serum metabolome. PLoS ONE, 6(2), e16957. doi:10.1371/journal.pone.0016957.
Quinones, M. P., & Kaddurah-Daouk, R. (2009). Metabolomics tools for identifying biomarkers for neuropsychiatric diseases. Neurobiology of Diseases, 35(2), 165–176. doi:10.1016/j.nbd.2009.02.019.
Rajput, A. H., Rozdilsky, B., & Rajput, A. (1991). Accuracy of clinical diagnosis in parkinsonism—a prospective study. Canadian Journal of Neurological Sciences, 18(3), 275–278.
Romero, P., Wagg, J., Green, M. L., Kaiser, D., Krummenacker, M., & Karp, P. D. (2005). Computational prediction of human metabolic pathways from the complete human genome. Genome Biology, 6(1), R2. doi:10.1186/gb-2004-6-1-r2.
Sharma, V., Ichikawa, M., & Freeze, H. H. (2014). Mannose metabolism: more than meets the eye. Biochem Biophys Res Commun, 453(2), 220–228. doi:10.1016/j.bbrc.2014.06.021.
Shaw, P. J. (2005). Molecular and cellular pathways of neurodegeneration in motor neurone disease. J Neurol Neurosurg Psychiatr, 76(8), 1046–1057. doi:10.1136/jnnp.2004.048652.
Sinclair, A. J., Viant, M. R., Ball, A. K., Burdon, M. A., Walker, E. A., Stewart, P. M., et al. (2010). NMR-based metabolomic analysis of cerebrospinal fluid and serum in neurological diseases—a diagnostic tool? NMR in Biomedicine, 23(2), 123–132. doi:10.1002/nbm.1428.
Smolinska, A., Blanchet, L., Buydens, L. M., & Wijmenga, S. S. (2012a). NMR and pattern recognition methods in metabolomics: from data acquisition to biomarker discovery: a review. Analytica Chimica Acta, 750, 82–97. doi:10.1016/j.aca.2012.05.049.
Smolinska, A., Posma, J. M., Blanchet, L., Ampt, K. A., Attali, A., Tuinstra, T., et al. (2012b). Simultaneous analysis of plasma and CSF by NMR and hierarchical models fusion. Analytical and Bioanalytical Chemistry, 403(4), 947–959. doi:10.1007/s00216-012-5871-4.
Stoop, M. P., Coulier, L., Rosenling, T., Shi, S., Smolinska, A. M., Buydens, L., et al. (2010). Quantitative proteomics and metabolomics analysis of normal human cerebrospinal fluid samples. Mol Cell Proteom, 9(9), 2063–2075. doi:10.1074/mcp.M900877-MCP200.
Tachikawa, M., Kasai, Y., Takahashi, M., Fujinawa, J., Kitaichi, K., Terasaki, T., et al. (2008). The blood-cerebrospinal fluid barrier is a major pathway of cerebral creatinine clearance: involvement of transporter-mediated process. Journal of Neurochemistry, 107(2), 432–442. doi:10.1111/j.1471-4159.2008.05641.x.
Tandan, R., & Bradley, W. G. (1985). Amyotrophic lateral sclerosis: part 1. Clinical features, pathology, and ethical issues in management. Ann Neurol, 18(3), 271–280. doi:10.1002/ana.410180302.
Trupp, M., Jonsson, P., Ohrfelt, A., Zetterberg, H., Obudulu, O., Malm, L., et al. (2014). Metabolite and peptide levels in plasma and CSF differentiating healthy controls from patients with newly diagnosed Parkinson’s disease. J Parkinsons Dis, 4(3), 549–560. doi:10.3233/JPD-140389.
Trygg, J., & Wold, S. (2002). Orthogonal projections to latent structures (O-PLS). Journal of Chemometrics, 16(3), 119–128. doi:10.1002/cem.695.
Tsacopoulos, M., & Magistretti, P. J. (1996). Metabolic coupling between glia and neurons. Journal of Neuroscience, 16(3), 877–885.
Vallance, P., & Leiper, J. (2004). Cardiovascular biology of the asymmetric dimethylarginine:dimethylarginine dimethylaminohydrolase pathway. Arteriosclerosis, Thrombosis, and Vascular Biology, 24(6), 1023–1030. doi:10.1161/01.ATV.0000128897.54893.26.
Van Sande, M., Caers, J., & Lowenthal, A. (1971). Cerebrospinal fluid amino acids in extrapyramidal disorders before and after L-DOPA treatment. Z Neurol, 199(1), 24–29.
Wade, A. M., & Tucker, H. N. (1998). Antioxidant characteristics of l-histidine 11The work described in this manuscript was partially sponsored and funded by cytos pharmaceuticals, LLC. Journal of Nutritional Biochemistry, 9(6), 308–315. doi:10.1016/s0955-2863(98)00022-9.
Want, E. J., Wilson, I. D., Gika, H., Theodoridis, G., Plumb, R. S., Shockcor, J., et al. (2010). Global metabolic profiling procedures for urine using UPLC-MS. Nature Protocols, 5(6), 1005–1018. doi:10.1038/nprot.2010.50.
Wishart, D. S., Lewis, M. J., Morrissey, J. A., Flegel, M. D., Jeroncic, K., Xiong, Y., et al. (2008). The human cerebrospinal fluid metabolome. J Chromatogr B Analyt Technol Biomed Life Sci, 871(2), 164–173. doi:10.1016/j.jchromb.2008.05.001.
Wishart, D. S., Jewison, T., Guo, A. C., Wilson, M., Knox, C., Liu, Y., et al. (2013). HMDB 3.0—the human metabolome database in 2013. Nucleic Acids Research, 41, 801–807. doi:10.1093/nar/gks1065. (Database issue).
Wuolikainen, A., Moritz, T., Marklund, S. L., Antti, H., & Andersen, P. M. (2011). Disease-related changes in the cerebrospinal fluid metabolome in amyotrophic lateral sclerosis detected by GC/TOFMS. PLoS ONE, 6(4), e17947. doi:10.1371/journal.pone.0017947.
Wuolikainen, A., Jonsson, P., Ahnlund, M., Antti, H., Marklund, S. L., Moritz, T., et al. (2016). Multi-platform mass spectrometry analysis of the CSF and plasma metabolomes of rigorously matched amyotrophic lateral sclerosis, Parkinson’s disease and control subjects. Molecular BioSystems, 12(4), 1287–1298. doi:10.1039/c5mb00711a.
Wyss, M., & Kaddurah-Daouk, R. (2000). Creatine and creatinine metabolism. Physiological Reviews, 80(3), 1107–1213.
Xia, J., Mandal, R., Sinelnikov, I. V., Broadhurst, D., & Wishart, D. S. (2012). MetaboAnalyst 2.0—a comprehensive server for metabolomic data analysis. Nucleic Acids Research, 40(Web Server issue), W127–W133. doi:10.1093/nar/gks374.
Zhang, A., Sun, H., Wang, P., Han, Y., & Wang, X. (2012). Recent and potential developments of biofluid analyses in metabolomics. J Proteomics, 75(4), 1079–1088. doi:10.1016/j.jprot.2011.10.027.
Zhao, Z., Lange, D. J., Voustianiouk, A., MacGrogan, D., Ho, L., Suh, J., et al. (2006). A ketogenic diet as a potential novel therapeutic intervention in amyotrophic lateral sclerosis. BMC Neurosci, 7, 29. doi:10.1186/1471-2202-7-29.
Acknowledgments
We are indebted to the patients who contributed to this study. We are grateful to the Erling-Persson Family Foundation, the Kempe Foundation and Umeå University for support. Dr. Junfang Wu is grateful for a post-doctoral fellowship from the Kempe foundation. Dr. Anna Wuolikainen, Dr. Miles Trupp, Dr. Pär Jonsson and Dr. Anders Öhman are supported by grants from the Erling-Persson Family Foundation. Dr. Peter Andersen has received grants from the Swedish Research Council, the Knut and Alice Wallenberg Foundation, The Bertil Hållsten Brain Research Foundation, The Brain Research Foundation, Swedish Brain Power Consortium, The Ulla-Carin Lindquist Foundation and Neuroförbundet. Dr. Lars Forsgren has received grants from The Swedish Medical Research Council, The Swedish Parkinson Foundation, The Swedish Parkinson’s disease Association, Västerbotten County Council (ALF), and the King Gustaf V’s and Queen Victoria’s Freemason Foundation. Dr. Anders Öhman has received grants from the Parkinson Foundation in Sweden and Insamlingsstiftelsen at Umeå University.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
The authors report no conflicts of interest related to this study.
Ethical approval
This study was carried out in accordance with the Declaration of Helsinki (WMA 1964), with ethical approval from the Medical Ethical Review Board for northern Sweden, Umeå, Sweden.
Informed consent
Written informed consent was obtained from all subjects.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wu, J., Wuolikainen, A., Trupp, M. et al. NMR analysis of the CSF and plasma metabolome of rigorously matched amyotrophic lateral sclerosis, Parkinson’s disease and control subjects. Metabolomics 12, 101 (2016). https://doi.org/10.1007/s11306-016-1041-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11306-016-1041-6