Abstract
Persian walnut (Juglans regia L) is an economically important species cultivated worldwide for its wood and nuts. Despite the increasing interest in the development of conservation strategies for walnut germplasm, an accurate and full-scale overview of wild genetic resources of J. regia has not been conducted because natural populations are located in regions of Asia historically difficult to access. In this study, we estimated the genetic diversity and spatial genetic structure of 39 autochthonous Persian walnut populations sampled across its Asian range using 14 neutral microsatellite markers. A landscape genetic overlay approach was applied to detect the areas of current reservoirs of walnut genetic diversity in the Asian range and to evaluate the role of landscape in shaping walnut genetic diversity since the Last Glacial Maximum. Although Persian walnut has been highly manipulated by humans over the last 2,000 years, we determined that patches of high genetic diversity still exist in the Caucasus and mountains of Central Asia where J. regia might have survived after Pleistocene glaciations. We detected a clear separation of Persian walnut into four main genetic clusters centered in (1) western Kyrgyzstan, (2) western and south–central Asia, (3) east–central Uzbekistan, and (4) Xinjiang and Shandong provinces (China). Overlay of maps showed a coincidence between groups of walnut populations and potential barriers to gene flow such as the Hindu Kush, Pamir, Tien Shan, and Himalaya mountains and the Karakum, Kyzyl Kum, and Taklamakan deserts. This study claimed the relevance of the preservation of walnut genetic resources in the Asian range.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Landscape genetics is emerging as a powerful approach that combines methods from landscape ecology, spatial analysis, geography, and population genetics to quantify the effects of landscape composition on evolutionary processes such as gene flow and adaptation (Storfer et al. 2010; Holderegger et al. 2010; Manel and Holderegger 2013). In particular, the landscape genetic overlay approach searches for the geographic coincidence of group boundaries, genetic discontinuities, and barriers to gene flow with landscape elements, using Geographic Information Systems (GISs) (Holderegger et al. 2010; Thiel-Egenter et al. 2011; Hepenstrick et al. 2012; Keller et al. 2012). The combination of molecular tools with GIS technologies has provided new opportunities to analyze spatial patterns of neutral or functional genetic diversity in forest tree species (González-Martínez et al. 2006; Eckert et al. 2010; Parchman et al. 2012), in particular in agroforestry species such as Castanea spp. (Mattioni et al. 2013) and Populus spp. (Breen et al. 2012).
Persian or common walnut (Juglans regia L.) is a large, wind-pollinated, monoecious, dichogamous, long-lived, perennial tree cultivated for its high-quality wood and nuts throughout the temperate regions of the world. It is generally accepted that J. regia is native to the mountain ranges of southeastern Europe (Carpathian Mountains) and west–central Asia, extending from Xinjiang Province of Western China, across southwestern Kyrgyzstan and Uzbekistan into the mountains of Nepal, Tibet, northern India, and Pakistan, west through Afghanistan, Turkmenistan, and Iran to the Caucasus (Azerbaijan, Armenia, Georgia, and Eastern Turkey) (McGranahan and Leslie 1991; Hemery et al. 2005; Aradhya et al. 2010). Persian walnut was probably introduced into central and eastern China from Xinjiang or Tibet during the Han Dynasty, about 2,100 years ago (Wang et al. 2008) or may have survived in refugia located in eastern China (Rong-Ting 1990) (a complete list of the earliest traces of radiocarbon-dated Juglans-type fossil pollen in Asia is available in Online Resource 1 and shown in Online Resource 2). It is still debated if Persian walnut was extinguished in Europe during the Pleistocene glaciations or if it survived in refugia located in Italy, Spain, and the Balkans, as suggested by some paleopalynologic studies (Carrion and Sanchez-Gomez 1992; Allen and Huntley 2009). Irrespective of the resolution of that debate, the distribution of J. regia has been strongly modified by human management during the last 2,000 years, especially in the Mediterranean basin (Salas-Salvadó et al. 2011).
Despite its economic importance and the increasing interest in the development of conservation strategies for Persian walnut germplasm, the genetic diversity of J. regia has been investigated primarily on a national or regional scale in the Eurasian continent (Wang et al. 2008; Karimi et al. 2010; Pollegioni et al. 2011). These studies revealed low levels of genetic differentiation among populations/provenances and no robust geographic patterns of genetic diversity. Despite reports of impressive morphological and phenological variation of Persian walnut in Central Asia (Molnar et al. 2011), an accurate, comparative, and full-scale overview of the wild genetic resources of the species in the Asian native range has never been undertaken. The reason is that many autochthonous Asian populations of Persian walnut grow in regions with formidable geographic barriers to collection and a history of geopolitical isolation (Schmidt and Doerre 2011). Nevertheless, in the last decades, the dissolution of the Soviet Union and Chinese economic reforms provided opportunities for establishing scientific collaborations and for assembling new collections of J. regia populations from the Asian range (Woeste and Michler 2011). Surprisingly, no polymorphism was detected in eight natural populations of Persian walnut in Asia using chloroplast PCR-RFLP markers (Fornari et al. 2001). Recently, a repartition of 203 Persian walnut accessions from the native range based on 15 nuclear microsatellites (nSSRs) and neighbor-joining tree analysis showed 5 genetically defined groups that mainly corresponded to major geographic areas of J. regia distribution in Asia (Aradhya et al. 2010). These encouraging results lead us to believe that a study of the large-scale spatial genetic structure of autochthonous Persian walnut populations in the Asian range using nSSR markers, combined with the analysis of geographic data, will provide insight into the following questions: (1) Although J. regia has a long history of domestication and human-mediated dispersal, do ancient reservoirs of genetic diversity (hot spots) still exist in the Asian native range? (2) Have geographic barriers (e.g., mountains, deserts) affected the distribution of current J. regia genetic resources and reduced gene flow among walnut populations? In particular, we wondered if the pattern of genetic variation across the current natural geographic distribution of J. regia bears the imprint of habitat fragmentation and climatic deterioration due to progressive desertification of Central Asia during the Holocene. Insights into these processes may improve the effectiveness of conservation and breeding programs in J. regia. Thus, in the present study, we (1) assessed the genetic diversity and spatial genetic structure of 39 autochthonous Persian walnut populations collected in the Asian range using 14 nSSR markers and (2) applied a landscape genetic overlay approach to evaluate the effects of landscape composition on gene flow and spatial genetic variation of walnut resources in their Asian native range.
Materials and methods
Persian walnut collection
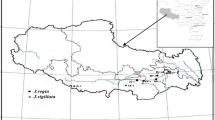
Over the last two decades, the CNR Institute of Agro-environmental and Forest Biology (IBAF, Porano, Italy), CNR Institute of Agricultural Biology and Biotechnology (Milan, Italy), and Earth Trust (Abingdon, Oxfordshire, UK) have monitored and sampled autochthonous Persian walnut populations from the mountainous regions of southwestern, Central, and Eastern Asia. Although we were unable to obtain samples from all areas in which J. regia grows naturally, we assembled a large and unique collection of J. regia, including 39 walnut populations (926 total genotypes) sampled from China (6 populations), Kyrgyzstan (9 populations), Uzbekistan (17 populations), Tajikistan (1 population), Pakistan (2 populations), Iran (1 population), Turkey (1 population), and Georgia (2 populations) (Table 1). These walnut populations represented eight different mountain ranges (Tien Shan, Gissar, Zaamin, Nurata, Pamir, Himalayas, Alborz, and Trans-Caucasus) and fell into seven phytogeographic regions (PRs) (Fig. 1; Table 1). A complete description of the Irano-Turanian (IT2, IT3, and IT4), Central Asiatic (CAR), Sino-Himalayan Forest (SHF), Sino-Japanese Forest (SJF), and Euro-Siberian (ES) PRs were reported by Qiu et al. (2011) and Djamali et al. (2012). The Chinese population Sunbé (Shandong Province) located in the putative Eastern Asian range of J. regia was collected and included in this study. More details of sampled sites were reported by Fornari et al. (2001), Hemery et al. (2005), Mapelli and Vildanova (2009), and Mapelli and Alexandrovsky (2011). Only centuries-old trees grown in ancient forests or in nature reserves were sampled (Table 1). We carefully avoided collecting walnut samples from agricultural and unprotected areas in the Asian range. A complete survey of walnut fossil pollen evidence in the native range has never been compiled and its record is rather sparse. Nevertheless, based on our fossil pollen database (Online Resources 1 and 2) we sampled in macroregions of Asia where J. regia species occurred before the human influence (before Copper Age ~8,000 years BP). Sampled trees in each population were separated by >100 m. Mature leaves were sampled from each of the walnut trees, dehydrated, and then stored at −80 °C at CNR-IBAF (Porano, Italy).
Location of 39 populations of Persian walnut collected across its Asian range: Kyrgyzstan (1–9), Uzbekistan (10–26), China (27–32), Pakistan (33 and 34), Tajikistan (35), Iran (36), Turkey (37), and Georgia (38 and 39). Phytogeographic subdivisions of southwestern and Central Asia proposed by Qiu et al. (2011) (dotted black line) and Djamali et al. (2012) (solid black line) are indicated: Irano-Turanian (IT2, IT3, and IT4), Central Asiatic (CAR), Sino-Himalayan Forest (SHF), Sino-Japanese Forest (SJF), and Euro-Siberian (ES). More details on walnut populations are shown in Table 1
Microsatellite analysis
Dehydrated leaf tissue (60 mg) from each sample was homogenized in a 2-ml microcentrifuge tube containing a 5-mm steel bead cooled with liquid nitrogen using Mixer Mill 300 (Qiagen, Hilden, Germany). Genomic DNA was extracted and purified using a DNeasy96 Plant Kit (Qiagen) and stored at −20 °C.
In this study, all samples were genotyped using 14 unlinked nSSR loci (WGA1, WGA4, WGA9, WGA27, WGA32, WGA69, WGA72, WGA79, WGA89, WGA118, WGA202, WGA276, WGA321, and WGA331) developed in black walnut (Victory et al. 2006) and already selected, sequenced, and used for genetic characterization of Persian walnut and Juglans × intermedia Carr. in Italy (Pollegioni et al. 2009). PCR amplification and visualization of amplified SSR alleles for each sample were conducted as described by Pollegioni et al. (2011). The raw SSR data were collected using Gene Scan Analysis version 3.7 and allele scoring was performed using Genotyper version 3.7 NT (Applied Biosystems, Foster City, CA, USA).
Data analysis
Genetic diversity of SSR loci
Standard measures of genetic diversity, total number of observed alleles (A), effective number of alleles (A e), observed (H O) and expected (H E) heterozygosity, and polymorphic information content (PIC) were computed for each locus and over all loci using POPGENE version 1.32 software (Yeh and Boyle 1997) and CERVUS version 2.0 software (Marshall et al. 1998). The three unbiased estimators of Wright’s F statistics (Weir and Cockerham 1984), within-population inbreeding coefficient f(F IS), total population inbreeding coefficient F(F IT), and among-population genetic differentiation coefficient θ(F ST) were calculated for each locus across all populations and over all loci using hierarchical locus-by-locus analysis of molecular variance (AMOVA) with Arlequin version 3.11 software (Excoffier et al. 2005). We used a nonparametric approach with 1,000 permutations to determine the statistical significance of f(F IS), F(F IT), and θ(F ST). Given that the dependence of F ST values on within-population heterozygosity can lead to an underestimation of the true level of genetic differentiation, we used SMOGD 1.2.5 software (Crawford 2010) to calculate Jost’s unbiased estimator (D est) (Jost 2008) for each locus across all populations and over all loci as an alternative measure of differentiation. Finally, FreeNa software (Chapuis and Estoup 2007) was used to estimate null allele frequencies for each locus and each population following the expectation-maximization algorithm. As suggested by Bagnoli et al. (2009), SSR loci with null allele frequency ranging from 0.1 to 0.3 in more than 30 % of populations were excluded. In addition, F ST estimates were recalculated for each locus [F ST(null)] by applying the ENA method as described by Chapuis and Estoup (2007) with 2,000 bootstrap iterations starting with the adjusted allele frequencies.
Genetic diversity of walnut populations
Genetic diversity was measured for each walnut population across all loci in terms of the mean number of alleles per locus (A), observed heterozygosity (H O), expected heterozygosity (H E), and unbiased expected heterozygosity (UHE) using the GenAlEx software 6.3 (Peakall and Smouse 2006). The observed mean number of alleles per locus is highly dependent on sample size. For this reason, allelic richness (R s) and private allele richness (PAR), which account for variation in sample sizes, were computed by using the rarefaction method with HP-Rare software (Kalinowski 2004). The estimates of R s and PAR were standardized on a minimum sample size of 10 individuals. Subsequently, the Inverse Distance Weighted (IDW) interpolation function implemented in the GIS software ArcGIS 9.3 (ESRI, Redlands, CA, USA) was used to show the geographic patterns of allelic richness (R s) computed for all 39 Persian walnut populations. As described by Murphy et al. (2008), we derived a continuous surface map using a linearly weighted combination of all the sample points such that each input point has a local influence that diminishes with distance to minimize the effect of irregular sampling. The within-population inbreeding coefficient F IS (Weir and Cockerham 1984) was estimated for each population using hierarchical locus-by-locus AMOVA as implemented in Arlequin 3.11 software (Excoffier et al. 2005). The statistical significance of F IS was tested using a nonparametric approach with 1,000 permutations.
Evidence of recent bottleneck events was investigated using the program BOTTLENECK 1.2.02 (Piry et al. 1999). Significance was assessed using Wilcoxon’s signed-rank test, which provides a relatively high power and can be used with as few as four polymorphic loci and any number of individuals. Three models of evolution have been proposed for microsatellite loci in the BOTTLENECK software: stepwise mutation model (SMM), infinite alleles mutation model of loci, and two-phase model (TPM). Pollegioni et al. (2009) determined that TPM most accurately reflected the mutational mechanism of the microsatellite loci used in this study. As recommended by Piry et al. (1999), we used the TPM with 95 % SMM and 5 % multistep mutations. Nevertheless, as reported by Henry et al. (2009), the heterozygosity excess test is sensitive to very recent disturbances. Two additional tests were used to identify bottleneck signatures from larger time scales: the shifted allele distribution analysis (Luikart et al. 1998) and the M ratio test (Garza and Williamson 2001) implemented in BOTTLENECK and Arlequin software, respectively. As proposed by Garza and Williamson (2001), the M ratio estimated for each locus, averaged over loci, was compared with a critical value: any population with a value of M < 0.68 was assumed to have gone through a reduction in size. This test is designed to detect older and longer-lasting bottlenecks than other commonly used methods (Williamson-Natesan 2005).
Spatial genetic structure analysis of walnut populations
Three complementary statistical approaches were used to detect and quantify the spatial genetic structure of walnut. First, a spatial analysis of molecular variance (SAMOVA) was applied to delineate groups of walnut populations using SAMOVA 1.0 software (Dupanloup et al. 2002). This approach iteratively seeks (10,000 iterations) for K groups of geographically adjacent populations to maximize the proportion of total genetic variation due to differences among groups (F CT) and minimize the amount of variation that occurs within groups (F SC). We tested K from 2 to 38 (number of walnut populations −1), selecting 100 random starting conditions. We examined patterns of F CT and F SC for each K to determine the most appropriate number of population groups. The configuration where F SC declined markedly and F CT increased was retained as the best partition of walnut populations.
Second, a Bayesian clustering approach implemented in STRUCTURE 2.3.3 software (Pritchard et al. 2000) was applied. We reconstructed the underlying genetic structure of walnut populations and computed the proportion of membership (Q value) for each predefined population and each individual multilocus genotype in each of the inferred clusters using Markov chain Monte Carlo (MCMC) simulations. STRUCTURE analysis was performed using the admixture model on the whole dataset without a priori population information and with the correlated allele frequencies between populations option. The range of the possible number of clusters (K) tested was from 1 to 39. Based on initial results, a series of 6 independent runs was performed for K between 1 and 12 with a burn-in period of 100,000 steps, followed by 106 MCMC replicates. The ad hoc statistic ∆K (Evanno et al. 2005) was calculated to detect the most likely number of populations. Therefore, the groups inferred by the first STRUCTURE analysis were subsequently reprocessed separately to identify possible substructure (subclusters). The six runs from the most probable number of clusters were averaged by applying the FullSearch algorithm provided by CLUMPP 1.1.2 software (Jakobsson and Rosenberg 2007). The corresponding Q matrices were graphically displayed by using the DISTRUCT software (Rosenberg 2004). After determining the most probable number of clusters, an arbitrary threshold of Q ≥ 0.80 was used to assign populations and/or genotypes to one group. Populations with 0.2 < Q < 0.8 were classified as admixed. Finally, following the approach described by Murphy et al. (2008), the population membership Q values for the K clusters estimated from STRUCTURE and CLUMPP software were spatially interpolated using the IDW function implemented in ArcGIS 9.3. This method provided K clustering surface maps showing the spatial patterns of the inferred genetic clusters.
Third, the presence of genetic barriers, corresponding to geographic zones of largest genetic differentiation among walnut populations, was investigated using Monmonier’s maximum difference algorithm as implemented in BARRIER 2.2 software (Manni et al. 2004). The geographic coordinates of each sampling site were connected by Delauney triangulation, and the corresponding Voronoï tessellation was derived. Once the network was obtained, each edge of the polygons was associated to its pairwise Nei’s (1972) genetic distance (D A) calculated using GENDIST in the PHYLIP software package (Felsenstein 2005). We tested from one to seven genetic barriers, and their significance was evaluated by means of 100 resampled bootstraps matrices of Nei’s (1972) genetic distances calculated using SEQBOOT and GENDIST in the PHYLIP software package. Genetic barriers with arbitrary bootstrap support of P > 0.50 were considered.
Correlation between genetic and landscape data
To examine the putative loss of walnut genetic diversity along the recolonization routes from refugial areas, the correlations between three geographic variables and Persian walnut genetic diversity parameters were evaluated. We performed a simple linear regression analysis, followed by the sequential Bonferroni correction (XLSTAT2010, http://www.xlstat.com) of allelic richness (R s) and unbiased expected heterozygosity (UHE) against longitude, latitude, and elevation.
The effect of a few selected geographic features on walnut population divergence within the Asian native area was also investigated. We examined pairwise F ST to quantify the effect of geographic distances (longitude and latitude), elevation differences, mountain ridges, and presence of deserts on gene flow among walnut populations using nonparametric pairwise simple and partial Mantel tests (Mantel 1967; Smouse et al. 1986) The P value for the Z score of the Mantel association parameter was inferred using 5,000 permutations as implemented in ZT software (Bonnet and Van der Peer 2002). We first tested for isolation by distance (IBD), i.e., the correlation between genetic and geographic distances. Assuming a nonlinear distribution of sampling sites, we regressed Slatkin’s linearized [F ST/(1 − F ST)] pairwise values against the corresponding natural logarithm of geographic distances. Connecting lines for all pairwise population comparisons were generated using ArcGis 9.3 and overlaid on the GTOPO30 global digital elevation model (DEM) map and global desertification vulnerability map (USDA—Natural Resources Conservation Service, http://soils.usda.gov/use/worldsoils/mapindex/desert.html). A pair of sites was considered to be separated by a geographic barrier if the straight line connecting these two sites intersected one or more ridges (elevation >2,000 m.a.s.l.) and/or deserts. As suggested by Balkenhol et al. (2009), we created a categorical matrix where the presence or absence of boundaries was indicated as 2 (mountain ridges and desert), 1 (mountain ridges or desert), or 0. Given that the geographically distant populations are often also separated by landscape boundaries, leading to spurious correlations, we performed a partial Mantel test to calculate the partial correlation between linearized F ST and mountain ridges/deserts after controlling for straight-line geographic distance. Subsequently, we also observed spatial coincidence between landscape elements and statistically significant genetic discontinuities among the 39 walnut populations. Sampling sites and genetic barriers with P ≥ 0.50 were overlaid on the GTOPO30 global DEM map using ArcGis 9.3. Elevations in GTOPO30 are regularly spaced at 30″, approximately 1 km (http://eros.usgs.gov/#/Find_Data/Products_and_Data_Available/GTOPO30).
Results
Genetic diversity of walnut populations
All 14 SSR loci used in the present study were highly polymorphic in the sampled walnut populations, and none of them showed evidence of null alleles (for a complete description of the SSR loci, see Online Resource 3).
Genetic diversity parameters were congruent and varied substantially among J. regia populations (Table 2). In particular, two geographic areas showed high values of R s: one in south–central Asia including sites in Tibet (32-Dashuicun), Kashmir–western Himalaya (34-Hunza Valley and 33-Gilgit Valley, Pakistan), northern Pamir ridges (35-Shouli, Tajikistan), Zaamir Mountain (15-Djarkurgan, Uzbekistan), Gissar Mountain (18-Bakhmal, Uzbekistan), Western Tien Shan (19-Karankul and 17-Bostanlyk, Uzbekistan), and three Trans-Caucasus sites (37-Anatolia, Turkey; 38-Lagodekhi; and 39-Skra, Georgia) (Fig. 2). Four sites in Eastern Tien Shan (27-Goungliu-1, 28-Goungliu-2, 29-Goungliu-3, and 30-Urumqi) were characterized by a relatively high level of genetic diversity (Fig. 2; Table 2). Of 182 total alleles detected, 34 were unique to a single geographic site and 16 were exclusive to 34-Hunza Valley (8) and 36-Karaj (8) (Table 2). No significant relationships existed between the genetic diversity (in terms of R s or UHE) within walnut populations and geographic (longitude, latitude, and elevation) variables. Overall F IS ranged from −0.039 (35-Shouli) to 0.268 (28-Gongliu-2) and was significantly positive for 19 of the 39 walnut populations, indicating a surplus of homozygotes in those populations (Table 2). Wilcoxon’s signed-rank test revealed a significant (P < 0.05) reduction in effective population size (heterozygosity excess) for only two walnut populations in the 17-Bostanlyk and 36-Karaj sites (Online Resource 4). A significant deviation (P < 0.05) from the mutation–drift equilibrium under the TPM of mutation (heterozygosity deficiency) was also detected for the 4-Shaidan and 5-Kyzyl-Ungur sites. Nevertheless, all these populations showed a typical L-shaped distribution of allele frequencies, except 36-Karaj and 17-Bostanlyk (Online Resource 4). By contrast, using the M ratio test, we observed a genetic signature consistent with a bottleneck in all the populations. The G–W values ranged from 0.28117 (36-Karaj) to 0.36718 (19-Karankul), which is substantially lower than the critical threshold (0.68) (Online Resource 4).
IDW interpolation of the allelic richness values (R s) calculated for 39 Persian walnut populations (black dots) in the species’ Asian range using 14 nSSR markers (Uzb Uzbekistan, Kyr Kyrgyzstan, Taj Tajikistan). The numbers of walnut populations are shown in Table 1
Spatial genetic structure of Persian walnut populations
The genetic structure of the 39 sampled Persian walnut populations was evaluated using SAMOVA and STRUCTURE clustering analysis.
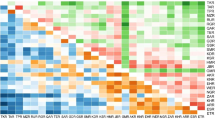
SAMOVA indicated five as the most appropriate number of population groups (F SC = 0.07236, F CT = 0.11351, P < 0.001; see Online Resource 5). The subsequent STRUCTURE analysis corroborated the SAMOVA results and provided additional information on the level of genomic admixture among populations. The ad hoc statistic ∆K of Evanno et al. (2005) strongly supported a hierarchy of clusters, with K = 2 as the most likely primary structure (∆K = 750.68), followed by K = 3 (∆K = 86.17) and K = 4 (∆K = 18.29). All subsequent ∆K values were negligible (Online Resource 6). Nevertheless, the statistical approach of Evanno et al. (2005) may be problematic, as it most often reveals K = 2 as the most likely number of clusters in the vast majority of the cases (uppermost hierarchical level). This outcome is often misleading and in conflict with the decision criteria suggested by Pritchard et al. (2000). Taking into consideration both the hierarchy of clusters detected by the ad hoc statistic ∆K of Evanno et al. (2005) and the distribution of log-likelihood values of data L(K) as a function of K averaged over six replicates (Online Resource 6), K = 4 was the best representation of the underlying hierarchical structure of the Persian walnut populations (Fig. 3a). Subsequently, we determined the populations’ Q membership values (Online Resource 7) for the four clusters inferred by STRUCTURE using the IDW function (Fig. 4). As also indicated by SAMOVA, the first group (cluster 1) was comprised of all nine Kyrgyz populations (1-Ak-Terek, 2-Sharap, 3-Yaradar, 4-Shaidan, 5-Kyzyl-Ungur, 6-Katar-Yangak, 7-Kyok-Sarau, 8-Kyr, and 9-Ters-Kolt) sampled in the walnut forests of the Western Tien Shan located in the IT4 PR (Q 1 ≥ 0.8956) (Figs. 3a and 4). Similarly, the second group (cluster 2) included samples from three Trans-Caucasus sites (37-Anatolia, Turkey, IT2 PR; 38-Lagodekhi, 39-Skra, Georgia, ES PR), Alborz ridges (36-Karaj, Iran, IT2 PR), northern Pamir ridges (35-Shouli, Tajikistan, IT4 PR), Kashmir–western Himalayas (33-Gilgit Valley and 34-Hunza Valley, Pakistan, CAR-ITA4 PR), and Tibetan–eastern Himalaya (32-Dashuicum, China, SHF PR) (Q 2 ≥ 0.8001) (Figs. 3a and 4). In addition, 32 walnut trees (89 %) collected in 19-Karankul (eastern Uzbekistan) and 10 of 67 walnut trees (14.9 %) from 28-Gongliu-2 (Xinjiang Province, China) were unambiguously assigned to cluster 2 (Q 2 ≥ 0.800). The third group (cluster 3) assembled all seven populations from Nurata ridges (20-Farish, 21-Andigen, 22-Katta-Bogdan, 23-Khayat, 24-Yamchi, 25-Karri, and 26-Madjerum) located in east–central Uzbekistan (IT3 PR) (Q 3 ≥ 0.9363) (Figs. 3a and 4). A fourth group of population (cluster 4) was detected and included sites of northern and eastern China, i.e., four sites of Eastern Tien Shan mountains, Xinjiang Province, CAR PR (Gongliu Wild Walnut Nature Reserve 27-Gongliu-1, 28-Gongliu-2, and 29-Gongliu-3 and Urumqi County 30-Urumqi) and one site from Shandong Province, SJF PR (31-Sunbè) (Q 4 ≥ 0.8001) (Figs. 3a and 4). The remaining six walnut populations on the Western Tien Shan mountains (10-Kamchik, 12-Sidjak, 13-Charvak, 14-Nanai, 16-Bogustan, and 17-Bostanlyk) and three populations from Fergana Valley (11-Yakkatut), Zaamin Mountain (18-Bakhmal), and Gissar Mountain (15-Djarkurgan) in eastern Uzbekistan (IT4 and IT3 PR) were admixtures between cluster 1 (0.4401 ≤ Q 1 ≥ 0.7780) and cluster 3 (0.1180 ≤ Q 3 ≥ 0.3525) and constituted a fifth group (Figs. 3a and 4; Online Resource 7).
Population structure inference for 926 Persian walnut samples by Bayesian assignment using STRUCTURE (Pritchard et al. 2000) for K = 4 clusters (a). Substructure inference for 212 walnut samples included in the inferred cluster 2 using K′ = 4 subclusters was also reported (b). Different colors in the same line indicate the individual’s estimated membership percentage in K clusters (Q value)
IDW interpolation of the estimated mean population membership values (Q i ) in the K = 4 clusters inferred by STRUCTURE (Pritchard et al. 2000) for 39 Persian walnut populations (black dots) in the species’ Asian range. The area bounded by brown indicates the strongest genetic similarity between populations belonging to the same cluster, and the gradual change to yellow indicates a gradual decrease in genetic similarity. The numbers of walnut populations are shown in Table 1 (Uzb Uzbekistan, Kyr Kyrgyzstan, Taj Tajikistan)
Subsequent STRUCTURE analysis within each of the previously inferred clusters did not reveal any genetic substructure, except for cluster 2. The analysis of log-likelihood values of data L(K′) as a function of K′ averaged over six replicates and ad hoc statistic ∆K of Evanno et al. (2005) detected a genetic subdivision of cluster 2 into four subclusters (K′ = 4, ∆K = 198.34; see Online Resource 8) dividing walnut trees into 28-Gongliu-2 and 32-Dashuicum (subcluster 1), 38-Lagodekhi and 39-Skra (subcluster 2), 37-Anatolia, 36-Karaj, 35-Shouli, and 19-Karankul (subcluster 3), and 33-Gilgit Valley and 34-Hunza Valley (subcluster 4) (Figs. 3b and 5; Online Resource 7).
IDW interpolation of the estimated mean population membership values (Q i ) in the K′ = 4 subclusters of cluster 2 (212 Persian walnut samples) inferred by STRUCTURE (Pritchard et al. 2000) (19-Karankul, 28-Gongliu-2, 32-Dash, 33-Gilgit, 34-Hunza, 35-Shouli, 36-Karaj, 37-Anatolia, 38-Lagodekhi, and 39-Skra)
IBD and landscape effect on walnut population structure
The pairwise linearized genetic differentiation values [F ST/(1 − F ST)] and the natural logarithm of geographic distances (straight-line distances in km) among sampling sites were significantly correlated (Mantel test r = 0.6248, P = 0.0002). However, space (latitude and longitude) and landscape features may act simultaneously on gene flow, influencing the genetic structure of populations. In this study, simple Mantel tests revealed that the presence of mountain ridges and/or deserts was positively correlated with pairwise genetic divergence (linearized F ST; r = 0.6683, P = 0.0002) and straight-line geographic distances (r = 0.7651, P = 0.0002) between walnut populations. Moreover, the partial correlation of linearized F ST and ridges/desert matrices remained significant but low after controlling for the effect of distances (natural logarithm of a straight line) among populations (partial Mantel test r = 0.3783, P = 0.0002).
Indeed, Monmonier’s maximum difference algorithm implemented in BARRIER identified five statistically significant (0.60 < P > 0.99) genetic barriers (Fig. 6). The main genetic boundary separated populations of western Asia (Georgia, Turkey, and Iran) from populations in Central and East Asia. This boundary was probably strengthened by the previously mentioned sampling gap between Georgia and Central Asia. An additional barrier splits the westernmost populations into subregions, dividing Georgian populations from Turkish and Iranian populations (Fig. 6). Within Central Asia, further genetic boundaries divided the remaining walnut populations into four main groups: (1) east–central Uzbekistan populations, (2) eastern Uzbekistan and western Kyrgyzstan populations, (3) five populations of northeastern China, and (4) five populations sampled in Tajikistan, Pakistan, and Tibet in south–central Asia. The overlay of sampling sites and the five computed genetic barriers on the GTOPO30 global DEM map (Fig. 6) roughly indicated the Hindu Kush, Pamir, Tien Shan, and Himalaya mountains as putative physical barriers to gene flow between Persian walnut populations. The use of a global desertification vulnerability map also indicated that Karakum, Kyzyl Kum, and Taklamakan deserts in west–central Asia probably affected the genetic differentiation of J. regia populations (Fig. 6).
Five statistically significant genetic barriers among 39 Persian walnut populations identified by BARRIER (Manni et al. 2004). Thin black lines represent the Voronöi tessellation derived from the Delaunay triangulation performed by using ArcGIS 9.3 and solid red line indicates statistically significant genetic boundaries. GTOPO30 global DEM map and global desertification vulnerability map (USDA—Natural Resources Conservation Service, http://soils.usda.gov/use/worldsoils/mapindex/desert.html) were overlaid
Discussion
Genetic diversity of Persian walnut across its Asian range
Although J. regia has a long history of cultivation and human-mediated dispersal (Salas-Salvadó et al. 2011), our investigation revealed two Asian regions containing local foci of high diversity: one in the Caucasus region (Georgia and Turkey) and the other in the mountainous regions of Central Asia that included the Himalayas (Tibet, China and Kashmir, Pakistan), northern Pamir (Tajikistan), Zaamir ridges, northern Gissar ridges, and part of Tien Shan mountains (eastern Uzbekistan and Xinjiang Province, China). These geographic areas may sustain remnants of ancient centers of diversity for the species. Based on the fossil pollen evidence, these peaks of genetic diversity occurred in macroregions where J. regia might have survived after Pleistocene glaciations in Asia. As reported in Online Resources 1 and 2, the earliest traces of radiocarbon-dated Juglans-type fossil pollen in Central Asia indicated that Persian walnut appeared in the Upper Pleistocene (12,000–40,000 years BP) and the beginning of the Holocene (10,000–12,000 years BP) in the Himalayan mountain ranges of Nepal, northwestern India, in the western Kunlun mountains of the Tibetan Plateau, and in the Pamir mountains of southern Tajikistan. Recent studies revealed that walnut pollen was constantly present in the Oligocene–Pliocene stratigraphic sequences detected in two independent Cenozoic sediments of the northern foreland basins of the Eastern Tien Shan mountains (Sun and Zhang 2008; Tang et al. 2011). In addition, although we evaluated only three populations in Georgia and Turkey, the Trans-Caucasian region may be an ancient center of diversity for J. regia and an important glacial refugium for several thermophilous plants, including oak species (Petit et al. 2002), and Castanea sativa (Mattioni et al. 2013). As indicated by the earliest traces of radiocarbon-dated Juglans-type fossil pollen (Online Resources 1 and 2), the most consistent shelter zone for walnut species in the Middle and Near East was the Trans-Caucasus, where walnut was abundant and persistent from the beginning of the Copper Age and/or Bronze Age. A few important geographic sites contained evidence of the presence of Juglans pollen during the Upper Paleolithic in Georgia (Dzudzuana Cave) (Bar-Yosef et al. 2011) and northwestern Iraq (Shanidar Cave, Zagros mountains) (Henry et al. 2011). Paleobotanical data from one of the oldest early Paleolithic sites in Eurasia (Dmanisi, Georgia) indicated that the forest–steppe environment in which hominins (Homo georgicus) settled in the Lesser Caucasus (1.8–1.7 Ma BP) was mostly temperate and dominated by deciduous forest trees such as species of Juglans, Castanea, and Cedrus (Messager et al. 2011).
As reported for several forest tree species of the Northern Hemisphere, glacial refugia are expected to harbor higher levels of genetic diversity compared with those areas that have been recolonized after the last ice age (Krebs et al. 2004). Widmer and Lexer (2001) described glacial refugia as sanctuaries of allelic richness. In this study, we determined that regions of high allelic richness corresponded with areas where ancient deposits of pollen were recorded (Online Resources 1 and 2), which supports the persistence of J. regia in fragmented forests of western and Central Asia during the cold and dry climate of the Pleistocene. Our results were also consistent with the “Phalanx model” of spread formulated to account for genetic patterns observed in temperate forests of Northern and Eastern Asia (Qiu et al. 2011). Based on this model, the high topographic heterogeneity of Asia provided suitable habitats for the survival of forest tree species during both glacial and interglacial cycles. Nevertheless, taking into consideration the long-term economic and cultural significance of walnut cultivation in Asia (Gunn et al. 2010), we cannot totally rule out the possibility that patches of high genetic diversity in the Caucasus and mountains of Central Asia may also be the result of human-mediated admixture.
Finally, the role of the walnut forests of Western Tien Shan in western Kyrgyzstan during the Holocene is of particular interest. They are considered impoverished relicts of mesophyllic forests of the Tertiary (Kolov 1998), but the current consensus is that the Juglans forests of western Kyrgyzstan originated at the end of the Pleistocene, approximately 25,000–30,000 years BP (Hemery et al. 2005). Nevertheless, more recent pollen and charcoal analyses of the sediments collected in the Fergana region revealed a putative anthropogenic origin of walnut forests in the core regions of western Kyrgyzstan (Beer et al. 2008). Juglans fossil pollen appeared during the Copper Age at Ak-Terek site (~7,500 years BP), but surprisingly, it became predominant only after the Iron Age, approximately 2,000 years BP. In this study, the levels of genetic diversity computed for all nine Kyrgyz walnut populations and three out of seven Uzbek populations sampled in the Western Tien Shan mountain were consistently low (R s ≤ 3.97) compared with the remaining walnut collection, possibly indicating a progressive recolonization of these areas from adjacent sites due to founder events and/or human transfers. Our results do not imply that wild walnut trees were entirely absent from the Fergana Range and the southeast-facing slopes of the Chatkal Range of Jalal-Abad Province after the Last Glacial Maximum (LGM). Nevertheless, the present data do not contradict the finding of Beer et al. (2008), which suggested that part of Juglans forests might have been established under the leadership of Arslan-Bop (1,120 years BP), the legendary founder of the Uzbek village of Arslanbop, using local walnut germplasm grown naturally.
Despite the presence of high levels of genetic diversity, significantly positive values of inbreeding (F IS) were observed in 48.7 % of the sampled walnut populations. Each of the sampled J. regia populations was almost completely isolated geographically and represented islands surrounded by arid continental lowland, mountain slopes, and highland steppes. Geographic isolation and the reduction/fragmentation of natural stands typically promote inbreeding over time (Gao et al. 2007), but it is difficult to make strong conclusions with respect to the role of bottlenecks in producing the existing pattern of walnut diversity in Asia. Using Wilcoxon’s signed-rank test combined with shifted allele distribution analysis, clear signals of recent reduction in effective population size were observed in only 2 (36-Karaj and 17-Bostanlyk) of the 39 walnut populations. This result is difficult to interpret because, as suggested by Henry et al. (2009), Wilcoxon’s signed-rank test is sensitive to recent disturbances but often fails to detect bottlenecks that occurred in the distant past. Although Bostanlyk has the highest density of walnut in eastern Uzbekistan, its distribution has been reduced by 50 % over the last 60 years owing to recent anthropogenic activities (Mapelli and Vildanova 2009). Similarly, in the last century, most of the wild walnut populations in Iran have been used extensively by rural communities for wood production (Karimi et al. 2010). By contrast, using the M ratio test of Gaza and Williamson (2001), we detected signatures of bottlenecks in all walnut populations that we sampled. It is not unreasonable that most of the Persian walnut populations sampled in Asia are affected by inbreeding due to geographic isolation and population decline. Nevertheless, these results should be considered with caution because the M ratio test is not completely reliable. As reported by Peery et al. (2012), M ratio tests can be quite sensitive to mistaken assumptions about the proportion of multistep mutations in the TPM, sometimes leading to an inference of the presence of bottlenecks in stable populations.
IBD and landscape genetic effect on J. regia population structure
SAMOVA and Bayesian clustering approaches produced convergent results, dividing the samples into four main clusters: cluster 1, Kyrgyzstan genotypes centered in the Western Tien Shan mountains; cluster 2, walnut trees in western and southern Central Asia, including Georgia and Turkey (Caucasus), Iran (foothills of the Alborz ridges), Karankul (Western Tien Shan mountains), Tajikistan (Pamir), Pakistan, and Tibet (Himalaya); cluster 3, walnut trees in east–central Uzbekistan (Nurata ridges); and cluster 4, Chinese genotypes collected in the Xinjiang (Eastern Tien Shan mountains, Urumqi) and Shandong provinces. Walnut populations from Western Tien Shan in eastern Uzbekistan exhibited an admixture between clusters 1 and 3. Cluster analysis also showed a subtle genetic subdivision of cluster 2 into four subclusters, separating all Tibetan walnut trees and 14.9 % of Gongliu-2 genotypes (subcluster 1) from Georgia (subcluster 2), Turkey, Iran, Tajikistan, and Uzbekistan (Karankul) (subcluster 3), and Pakistan (subcluster 4). Thus, most population groups of J. regia were confined to particular mountain ranges, showing a distinct phylogeographic structure. As already postulated for 9 J. regia populations in Yunnan Province of China (Wang et al. 2008) and 203 Persian walnut accessions from the Eurasian native range (Aradhya et al. 2010), the observed separation of J. regia populations reflects, at least in part, a complex topography of habitats—the presence of high mountains, deep valleys, and deserts—that prevented gene flow among populations during the Holocene. Furthermore, as indicated by the fossil pollen database (Online Resources 1 and 2), the geographic complexity of Asia might have provided multiple isolated refugia, initiating genetic differentiation observed among current Persian walnut populations. Geographic complexity is likely to have shaped the genetic composition of many endemic forest tree species across eastern and southern China, including Juglans mandshurica (Bai et al. 2010), Fagus engleriana (Lei et al. 2012), and Taxus wallichiana Zucc. (Gao et al. 2007).
In this study, Mantel tests supported an isolation-by-distance model of expansion for J. regia in its Asian range. The patterns of Persian walnut migration during the past 25,000 years might be partially explained by pollen/seed dispersal. At maturity, each catkin of walnut staminate flowers releases two million pollen grains that are subsequently dispersed by wind. As reported by Impiumi and Ramina (1967), when wind speed was low (0.5 m/s), common walnut pollen showed uniform density for 160 m surrounding a source. In addition, seeds of Persian walnut can be dispersed by birds and scatter-hoarded by rodents. Two rodent species, the Japanese squirrel (Sciurus lis) (Tamura et al. 1999) and the field mouse (Apodemus speciosus) (Tamura et al. 2001), transport Juglans ailantifolia seeds at a mean distance of 168 and 7 m, respectively. A recent increase in the population size of the native rook (Corvus frugilegus), combined with large-scale land abandonment, led to the expansion of the non-native Persian walnut in seminatural habitats of Poland (Lenda et al. 2012). Rooks carried walnuts for distances that were usually below 500 m, although sometimes up to or over 1 km. Thus, these dispersers might have a role in the local recolonization of walnut in Asia from multiple glacial refugia after LGM. As reported by Hewitt and Kellman (2002), in patchy systems, where suitable habitat is sparsely distributed and extensively separated, large-seeded and unspecialized species such as Juglans spp. showed limited dispersion that caused fragmentation and increased intrapopulation differentiation. In this study, the partial Mantel tests revealed that the spatial genetic structure of J. regia populations might have been simultaneously influenced by a combination of geographic distance and landscape resistance. Notably, the presence of mountain ridges and/or deserts was positively correlated with pairwise genetic divergence (F ST). We caution that Mantel tests and related methods for genetic studies have been questioned recently for showing inflated type 1 error rate (Guillot and Rousset 2013). Nevertheless, as reported by Balkenhol et al. (2009), Mantel tests may be appropriate for preliminary analyses in the early stages of landscape genetic studies and combined with alternative approaches. Although BARRIER analysis could be influenced by incomplete sampling of the distribution of J. regia in Asia, five obvious genetic barriers (a, b, c, d, and e) with bootstrap support of >50 % were detected, corroborating SAMOVA and STRUCTURE results. As reported by Holderegger et al. (2010), the overlay approach remains purely visual, but in this study, the construction of synthetic maps using GIS technologies allowed us to roughly indicate the Hindu Kush, Pamir, Tien Shan, and Himalaya mountains and the Karakum, Kyzyl Kum, and Taklamakan deserts as putative barriers to gene flow. Analysis of paleoclimatic records of effective moisture demonstrated that the most significant, widespread, and severe desertification occurred in Central Asia during the early Holocene prior to 8,000 years BP (Jin et al. 2012). In addition, a subsequent major transition from relatively humid to more arid conditions was observed between 6,900 and 4,900 years BP in the mountain regions of Central Asia (Ricketts et al. 2001). Relatively stable conditions analogous to the modern situation were observed after 4,000 years BP.
Thus, without overlooking the influence of irregular sampling, we believe that the sharp genetic structure detected for Persian walnut across its Asian range can be attributed to repeated events of population fragmentation and genetic isolation after postglacial expansion. Most important to this process were high mountain ridges, which acted as physical barriers to gene flow, and the climatic deterioration of J. regia habitat associated to the progressive desertification of Central Asia during the Holocene. This scenario matches the vegetation reconstruction of Gallimore et al. (2005), who suggested a long-term fragmentation of temperate deciduous forests in the Central and Eastern Asia after LGM. This model is also supported by the partial coincidence of the genetic structure of Persian walnut and the classification of populations into PRs. We conclude that, in addition to (or despite) any anthropogenic effects on the evolution of J. regia, the complex topography of Central Asia had a major role in shaping the observed PRs of the Asiatic continent (Djamali et al. 2012) and also affected the spatial genetic distribution of Persian walnut resources in their Asian range. In addition to topography, it seems likely that human-mediated dispersal processes might have also influenced the spatial genetic patterns of J. regia over the centuries. In this study, strong genetic relationships were detected among four Persian walnut populations from Turkey, Iran, Tajikistan, and Uzbekistan (Karankul) (subcluster 3). The presence of subcluster 3, grouping four geographically distant and separated populations, cannot be explained in terms of natural pollen/seed dispersal mechanisms. A preliminary genetic analysis of 90 Eurasian walnut populations using nSSR markers suggested that ancient trade routes, passing from China into Uzbekistan, Persia, Turkey, and Greece (e.g., Silk Road), might have a role in transferring walnut seeds throughout the Eurasian continent (data not shown). In particular, these “green corridors” may have facilitated human-mediated gene flow among Persian walnut populations grouped in subcluster 3, reducing the level of genetic structure even in the presence of enormous geographic distances. Although only wild centuries-old walnut trees from ancient forests or in nature reserves were sampled, we cannot exclude an ancient anthropogenic effect on subcluster 3. The role of trading routes, human migration, and historical and anthropogenic barriers in shaping part of the genetic structure of Persian walnut populations in Asia is currently under evaluation.
In conclusion, we believe that our work will provide useful information for improved forest management practices and for developing new conservation strategies for Persian walnut germplasm. Although Persian walnut has been manipulated by humans, we determined that high levels of genetic diversity still exist in the Asian range. A clear separation of Persian walnut in different genetic groups was also detected. The overlay approach indicated that the presence of potential barriers to gene flow, such as the Hindu Kush, Pamir, Tien Shan, and Himalaya mountains, and the progressive desertification of Central Asia during the Holocene may have promoted the fragmentation and isolation of Persian walnut populations. This study claimed the relevance of the preservation of walnut genetic resources in the Asian range.
References
Allen JRM, Huntley B (2009) Last interglacial palaeovegetation, palaeoenvironments and chronology: a new record from Lago Grande di Monticchio, southern Italy. Quat Sci Rev 28:1521–1538
Aradhya M, Woeste K, Velasco D (2010) Genetic diversity, structure and differentiation in cultivated walnut (Juglans regia L). Acta Horticult 861:127–132
Bagnoli F, Vendramin GG, Buonamici A, Doulis G, Gonzàlez-Martìnez C, La Porta N, Magri D, Raddi P, Sebastiani F, Fineschi S (2009) Is Cupressus sempervirens native in Italy? An answer from genetic and paleobotanical data. Mol Ecol 18:2276–2286
Bai WN, Liao WJ, Zhang DY (2010) Nuclear and chloroplast DNA phylogeography reveal two refuge areas with asymmetrical gene flow in a temperate walnut tree from East Asia. New Phytol 188:892–901
Balkenhol N, Waits LP, Dezzani RJ (2009) Statistical approaches in landscape genetics: an evaluation of methods for linking landscape and genetic data. Ecography 32:818–830
Bar-Yosef O, Belfer-Cohen A, Mesheviliani T, Jakeli N, Bar-Oz G, Boaretto E, Goldberg P, Kvavadze E, Matskevich Z (2011) Dzudzuana: an Upper Palaeolithic cave site in the Caucasus foothills (Georgia). Antiquity 85:331–349
Beer R, Kaiser F, Schmidt K, Ammann B, Carraro G, Grisa E, Tinner W (2008) Vegetation history of the walnut forests in Kyrgyzstan (Central Asia): natural or anthropogenic origin? Quat Sci Rev 27:621–632
Bonnet E, Van der Peer Y (2002) ZT: a software tool for simple and partial Mantel tests. Ghent University. J Stat Softw 7:1–12. Available at http://bioinformatics.psb.ugent.be/software/details/ZT
Breen AL, Murray DF, Olson MS (2012) Genetic consequences of glacial survival: the late Quaternary history of balsam poplar (Populus balsamifera L.) in North America. J Biogeogr 39:918–928
Carrion JS, Sanchez-Gomez P (1992) Palynological data in support of the survival of walnut (Juglans regia L.) in the western Mediterranean area during last glacial times. J Biogeogr 19:623–630
Chapuis MP, Estoup A (2007) Microsatellite null alleles and estimation of population differentiation. Mol Biol Evol 24:621–631
Crawford NG (2010) SMOGD: software for the measurement of genetic diversity. Mol Ecol Resour 10:556–557
Djamali M, Brewer S, Breckle SW, Jackson ST (2012) Climatic determinism in phytogeographic regionalization: a test from the Irano-Turanian region, SW and Central Asia. Flora 207:237–249
Dupanloup I, Schneider S, Excoffier L (2002) A simulated annealing approach to define the genetic structure of populations. Mol Ecol 11:2571–2581
Eckert AJ, Bower AD, Gonzàlez-Martinez SC, Wegrzyn JL, Coop G, Neale DB (2010) Back to nature: genetic associations in a landscape context. Mol Ecol 19:3789–3805
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
Excoffier L, Laval G, Schneider S (2005) Arlequin ver. 3.0: an integrated software package for population genetics data analysis. Evol Bioinformatics Online 1:47–50
Felsenstein J (2005) PHYLIP (phylogeny inference package). Department of Genome Sciences, University of Washington, Seattle
Fornari B, Malvolti ME, Taurchini D, Fineschi S, Beritognolo I, McCaglia E, Cannata F (2001) Isozyme and organellar DNA analysis of genetic diversity in natural/naturalized European and Asiatic walnut (Juglans regia) populations. Acta Horticult 544:167–178
Gallimore R, Jacob R, Kutzbach J (2005) Coupled atmosphere–ocean–vegetation simulations for modern and mid-Holocene climates: role of extratropical vegetation cover feedbacks. Clim Dyn 25:755–776
Gao LM, Möller M, Zhang XM, Hollingsworth ML, Liu J, Mill RR, Gibby M, Li DZ (2007) High variation and strong phylogeographic pattern among cpDNA haplotypes in Taxus wallichiana (Taxaceae) in China and North Vietnam. Mol Ecol 16:4684–4698
Garza JC, Williamson EG (2001) Detection of reduction in population size using data from microsatellite loci. Mol Ecol 10:305–318
González-Martínez SC, Krutovsky KV, Neale DB (2006) Forest-tree population genomics and adaptive evolution. New Phytol 170:227–238
Guillot G, Rousset F (2013) Dismantling the Mantel tests. Methods Ecol Evol 4:336–344
Gunn BF, Aradhya M, Salick JM, Miller AJ, Yongping Y, Lin L, Xian H (2010) Genetic variation in walnuts (Juglans regia and J. sigillata; Juglandaceae): species distinctions, human impacts, and the conservation of agrobiodiversity in Yunnan, China. Am J Bot 97:660–671
Hemery G, Savill PS, Thakur A (2005) Height growth and flushing in common walnut (Juglans regia L.): 5-year results from provenance trials in Great Britain. Forestry 78:121–133
Henry P, Miquelle D, Sugimoto T, McCullough DR, Caccone A, Russello MA (2009) In situ population structure and ex situ representation of the endangered Amur tiger. Mol Ecol 18:3173–3184
Henry AG, Brooks AS, Piperno DR (2011) Microfossils in calculus demonstrate consumption of plants and cooked foods in Neanderthal diets (Shanidar III, Iraq; Spy I and II, Belgium). Proc Natl Acad Sci U S A 108:486–491
Hepenstrick D, Holderegger R, Thiel D, Gugerli F (2012) Genetic discontinuities in roe deer (Capreolus capreolus) coincide with fenced transportation infrastructure. Basic Appl Ecol 13:631–638
Hewitt N, Kellman M (2002) Tree seed dispersal among forest fragments: II. Dispersal abilities and biogeographical controls. J Biogeogr 29:351–363
Holderegger R, Buehler D, Gugerli F, Manel S (2010) Landscape genetics of plants. Trends Plant Sci 15:675–683
Impiumi G, Ramina A (1967) Ricerche sulla biologia fiorale e di fruttificazione del noce (J regia) I Osservazioni sulla morfologia fiorale e sul trasporto del polline. Riv Ortofrutticoltura Ital 51:538–543
Jakobsson M, Rosenberg NA (2007) CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodaly in analysis of population structure. Bioinformatics 14:1801–1806
Jin L, Chen F, Morrill C, Otto-Bliesner BL, Rosenbloom N (2012) Causes of early Holocene desertification in arid central Asia. Clim Dyn 38:1577–1591
Jost L (2008) G ST and its relatives do not measure differentiation. Mol Ecol 17:4015–4026
Kalinowski ST (2004) Counting alleles with rarefaction: private alleles and hierarchical sampling designs. Conserv Genet 5:539–543
Karimi R, Ershadi A, Vahdati K, Woeste K (2010) Molecular characterization of Persian walnut populations in Iran with microsatellites markers. J Am Soc Hortic Sci 45:1403–1406
Keller D, Van Strien MJ, Holderegger R (2012) Do landscape barriers affect functional connectivity of populations of an endangered damselfly? Freshwater Biol 57:1373–1384
Kolov O (1998) Ecological characteristics of the walnut–fruit forests of southern Kyrgyzstan. In: Blaser J, Carter J, Gilmour D (eds) Biodiversity and sustainable use of Kyrgyzstan’s walnut–fruit forests. English version. IUCN, Intercooperation, Gland, pp 59–61
Krebs P, Conedera M, Pradella M, Torriani D, Felber M, Tinner W (2004) Quaternary refugia of the sweet chestnut (Castanea sativa Mill.): an extended palynological approach. Veg Hist Archaeobot 13:145–160
Lei M, Wang Q, Wu Z-J, López-Pujol J, Li D-Z, Zhang Z-Y (2012) Molecular phylogeography of Fagus engleriana (Fagaceace) in subtropical China: limited admixture among multiple refugia. Tree Genet Genomes 8:1203–1212
Lenda M, Skórka P, Knops JMH, Moron D, Tworek S, Woyciechowski M (2012) Plant establishment and invasions: an increase in a seed disperser combined with land abandonment causes an invasion of the non-native walnut in Europe. Proc R Soc B 279:1491–1497
Luikart G, Allendorf FW, Cornuet JM, Sherwin WB (1998) Distortion of allele frequency distributions provides a test for recent population bottlenecks. J Hered 89:238–247
Manel S, Holderegger R (2013) Ten years of landscape genetics. Trends Ecol Evol 28:614–621
Manni F, Guerard E, Heyer E (2004) Geographic patterns of (genetic, morphologic, linguistic) variation: how barriers can be detected by using Monmonier’s algorithm. Hum Biol 76:173–190
Mantel N (1967) The detection of disease clustering and a generalized regression approach. Cancer Res 27:209–220
Mapelli S, Alexandrovsky ES (2011) Study of biodiversity and genetic resources of walnut in Uzbekistan. IV: Eco-geographical of walnut stands and form diversity in Nurata ridge. National Research Council, Institute of Agricultural Biology and Biotechnology, Milan, pp 1–56
Mapelli S, Vildanova GV (2009) Study of biodiversity and genetic resources of walnut in Uzbekistan. I: Western Tien-Shan forests. National Research Council, Institute of Agricultural Biology and Biotechnology, Milan, pp 1–36
Marshall TC, Slate J, Kruuk LEB, Pemberton M (1998) Statistical confidence for likelihood-based paternity inference in natural populations. Mol Ecol 7:639–655
Mattioni C, Martin A, Pollegioni P, Cherubini M, Villani F (2013) Microsatellite markers reveal a strong geographic structure in European populations of Castanea sativa (Fagaceae): evidence for multiple glacial refugia. Am J Bot 100:1–11
McGranahan G, Leslie CA (1991) Walnuts (Juglans). In: Moore JN and Ballington JR (eds) Genetic resources of temperate fruit and nut crops, vol. 2. Acta Hort 290:907–951
Messager E, Lebreton V, Marquer L, Russo-Ermolli E, Orain R, Renault-Miskovsky J, Lordkipanidze D, Despriée J, Peretto C, Arzarello M (2011) Palaeoenvironments of early hominins in temperate and Mediterranean Eurasia: new palaeobotanical data from Palaeolithic key-sites and synchronous natural sequences. Quat Sci Rev 30:1439–1447
Molnar TJ, Zaurov DE, Capik JM, Eisenman SW, Ford T, Nikolyi LV, Funk CR (2011) Persian walnuts (Juglans regia L.) in Central Asia. Annu Rep N Nutr Grow Assoc 101:56–69
Murphy MA, Evans JS, Cushman SA, Storfer A (2008) Representing genetic variation as continuous surfaces: an approach for identifying spatial dependency in landscape genetic studies. Ecography 31:685–697
Nei M (1972) Genetic distance between populations. Am Nat 106:283–292
Parchman TL, Gompert Z, Mudge J, Schilkey FD, Benkman CW, Buerkle CA (2012) Genome-wide association genetics of an adaptive trait in lodgepole pine. Mol Ecol 21:2991–3005
Peakall R, Smouse PE (2006) GenAlEx V6: genetic analysis in Excel. Population genetic software for teaching and research. The Australian National University, Canberra, Australia. Molec Ecol Notes 6:288–295. Available at http://www.anu.edu.au/BoZo/genAlEx/
Peery MZ, Kirby R, Reid BN, Stoelting R, Doucet-Bëer E, Robinson S, Vásquez-Carrillo C, Pauli JN, Palsbøll PJ (2012) Reliability of genetic bottleneck tests for detecting recent population declines. Mol Ecol 21:3403–3418
Petit RJ, Brewer S, Bordac S, Burg K, Cheddadi R, Coart E, Cottrell J, Csaikl UM, van Dam BC, Deans JD, Espinel S, Fineschi S, Finkeldey R, Glaz I, Goicoechea PG, Jensen JS, König AO, Lowe AJ, Madsen SF, Mátyás G, Munro RC, Popescu F, Slade D, Tabbener H, de Vries SMG, Ziegenhagen B, de Beaulieu J-L, Kremer A (2002) Identification of refugia and post-glacial colonisation routes of European white oaks based on chloroplast DNA and fossil pollen evidence. Forest Ecol Manag 156:49–74
Piry S, Luikart G, Cornuet JM (1999) BOTTLENECK: a computer program for detecting recent reductions in effective population size from allele frequency data. J Hered 90:502–503
Pollegioni P, Woeste K, Major A, Scarascia Mugnozza G, Malvolti ME (2009) Characterization of Juglans nigra (L.), Juglans regia (L.) and Juglans × intermedia (Carr.) by SSR markers: a case study in Italy. Silvae Genet 58:68–78
Pollegioni P, Woeste K, Olimpieri I, Marandola D, Cannata F, Malvolti ME (2011) Long term human impacts on genetic structure of Italian walnut inferred by SSR markers. Tree Genetics Genomes 7:707–723
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Qiu YX, Fu CX, Comes HP (2011) Plant molecular phytogeography in China and adjacent regions: tracing the genetic imprints of Quaternary climate and environmental change in the world’s most diverse temperate flora. Mol Phylogenet Evol 59:225–244
Ricketts RD, Johson TC, Brown ET, Rasmussen KA, Romanovsky VV (2001) The Holocene paleolimnology of Lake Issyk-Kul, Kyrgyzstan: trace element and stable isotope composition of ostracodes. Palaeogeogr Palaeoclimatol Palaeoecol 176:207–227
Rong-Ting X (1990) Discussion on the origin of walnut in China. Acta Horticult 284:353–361
Rosenberg NA (2004) DISTRUCT: a program for the graphical display of population structure. Mol Ecol Notes 4:137–138
Salas-Salvadó J, Casa-Agustench P, Sala-Huetos A (2011) Cultural and historical aspects of Mediterranean nuts with emphasis on their attributed healthy and nutritional properties. Nutr Metab Cardiovasc 21:S1–S6
Schmidt M, Doerre A (2011) Changing meanings of Kyrgyzstan’s nut forests from colonial to post-Soviet times. Area 43(3):288–296
Smouse PE, Long JC, Sokal RR (1986) Multiple regression and correlation extensions of the Mantel test of matrix correspondence. Syst Zool 35:627–632
Storfer A, Murphy MA, Spear SF, Holderegger R, Waits LP (2010) Landscape genetics: where are we now? Mol Ecol 19:3496–3514
Sun J, Zhang Z (2008) Palynological evidence for the Mid-Miocene climatic optimum recorded in Cenozoic sediments of the Tian Shan Range, northwestern China. Glob Planet Chang 64:53–68
Tamura N, Hashimoto Y, Hayashi F (1999) Optimal distances for squirrels to transport and hoard walnuts. Anim Behav 58:635–642
Tamura N, Hashimoto Y, Hayashi F (2001) Walnut hoarding by the Japanese wood mouse, Apodemus speciosus Temminck. J For Res 6:187–190
Tang Z, Ding Z, White PD, Dong XX, Ji JL, Jiang HC, Luo P, Wang X (2011) Late Cenozoic central Asian drying inferred from a palynological record from the northern Tian Shan. Earth Planet Sci Lett 302:439–447
Thiel-Egenter C, Alvarez N, Holderegger R, Tribsch A, Englisch T, Wohlgemuth T, Ahlmer WR, Colli L, Dalmas JP, Gaudeul M, Gielly L, Jogan N, Lequette B, Linder HP, Martini F, Negrini R, Niklfeld H, Pellecchia M, Prosser F, Rioux D, Rossi P, Scheuerer M, Schönswetter P, Selvaggi A, vanLoo M, Winkler M, IntraBioDiv Consortium, Gugerli F (2011) Break zones in the distributions of alleles and species in alpine plants. J Biogeogr 38:772–782
Victory E, Glaubitz JC, Rhodes OE, Woeste KE (2006) Genetic homogeneity in Juglans nigra (Juglandaceae) at nuclear microsatellites. Am J Bot 93:118–126
Wang H, Pei D, Gu R, Wang B (2008) Genetic diversity and structure of walnut populations in central and southwestern China revealed by microsatellite markers. J Hortic Sci 133:197–203
Weir BS, Cockerham CC (1984) Estimating F-statistics for the analysis of population structure. Evolution 38:1358–1370
Widmer A, Lexer C (2001) Glacial refugia: sanctuaries for allelic richness, but not for gene diversity. Trends Ecol Evol 16:267–269
Williamson-Natesan EG (2005) Comparison of methods for detecting bottlenecks from microsatellite loci. Conserv Genet 6:551–562
Woeste K, Michler C (2011) Juglans. In: Kole C (ed) Wild crop relatives: Genomic and breeding resources, forest trees. Springer, Berlin, pp 77–87
Yeh FC, Boyle TJB (1997) Population genetic analysis of co-dominant and dominant markers and quantitative traits. Belg J Bot 129:157
Acknowledgments
This study was supported by the European Community in the framework of the Seventh Framework Programme under the Marie Curie Actions COFUND project “I-MOVE” (No. 267232) and by the CNR-IBAF (Italian National Research Council–Institute of Agro-environmental and Forest Biology, Porano (Terni, Italy). Samples were obtained in the framework of the Bilateral Projects of the Italian Ministry of Foreign Affairs (L. 401.1990) during the years 1998–2010. The authors thank Marcello Cherubini and Daniela Taurchini for their support in the laboratory and Dr. Claudia Mattioni, Andreas Hamann, and Kevin McKelvey for their critical review of the manuscript. Special thanks goes to the memory of Prof. Erìc Germain (INRA-Bordeaux, France), who provided the Caucasian walnut populations under the EU-AIR 3 CT92 0142 and FAIR III PL96 1887 “W-BRAINS” projects. The use of trade names is for the information and convenience of the reader and does not imply official endorsement or approval by the United States Department of Agriculture or the Forest Service of any product to the exclusion of others that may be suitable.
Data Archiving Statement
Sample and population names of Persian walnut collection, geographic location of Persian walnut populations, and microsatellite raw data for 926 Persian walnut genotypes were submitted to the TreeGene Database (http://dendrome.ucdavis.edu/treegenes/; accession number TGDR017).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by S. C. González-Martínez
Rights and permissions
About this article
Cite this article
Pollegioni, P., Woeste, K.E., Chiocchini, F. et al. Landscape genetics of Persian walnut (Juglans regia L.) across its Asian range. Tree Genetics & Genomes 10, 1027–1043 (2014). https://doi.org/10.1007/s11295-014-0740-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11295-014-0740-2