Abstract
Lung Cancer is the most fast growing cancer around the world and is mostly diagnosed at an advanced stage. Due to enhancement in medical imaging modalities like Computed Tomography (CT) scans there is a need for computer aided detection system to classify the lung nodule into benign and malignant type with maximum accuracy to prevent delay in diagnosis. Many state-of-art methods used so far classify the images by applying machine learning algorithms on manually extracted features from imaging modalities. But in the recent years many deep learning techniques are being used in classification of lung nodule and have shown promising results when compared to other state-of-art methods. In this paper we have surveyed around 108 research papers to focus on the contribution of deep learning methodologies in detection of malignant tumor in Lung CT scan. This paper discusses variation applied on deep learning architecture to improve the accuracy of the classification system and a comprehensive comparison between various deep learning methods used so far for lung nodule classification. After reviewing each paper, this survey also presents challenges and opportunities in classifying lung nodule by using advanced deep learning strategies. The paper concludes with the need to address new issues in nodule classification with an aim to detect the malignant lesion at an early stage.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
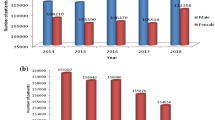
Lung Cancer has been the leading causes of death worldwide, with 1.8 million deaths predicted in 2018 and incidence of 2.1 million new cancer cases. According to GLOBOCAN 2018 survey, lung cancer persists to be most common cancer (11.6% of entire cancer cases) and has the highest mortality rate of 18.4% (of the entire cancer population) [1]. Lung Cancer is mostly found in men than in women with a ratio of 4.5:1 [2]. It is the leading cause of death among men and women in 93 and 23 countries respectively. In 2018, there were 1,184,947 and 5, 76,060 death cases in men and women respectively due to lung cancer [1]. The Global Disease Burden of Lung Cancer is likely to increase in the first half of century due to increasing trends in lung cancer incidence and mortality and very less improvement in survival globally [3]. It is observed that there are substantial delays from development of symptoms to the first initial treatment in most of the lung cancer patients [4]. Majority of patients are mostly diagnosed at a much later stage of lung cancer and the death mostly occurs in the first year after diagnosis [5].
Lung cancer is uncontrolled growth of cells in lung parenchyma which if not treated at early stage can spread to distant location in the body and cause metastatic complications. Cancer that starts from the lung is termed as primary lung cancer. Lung cancers are classified as either small-cell lung carcinoma (SCLC) or non-small-cell lung carcinoma (NSCLC) histopathologically, the SCLCs constituting 15% and NSCLCs constituting 85% of entire lung cancer population, respectively [6]. NSCLC are further sub-divided into three types: Adenocarcinoma, Squamous-cell carcinoma and Large-cell carcinoma [7]. Among all lung cancers, Adenocarcinoma is the most common lung carcinoma; contributing to 40% of entire lung cancer incidents. 25–30% of entire lung cancer population suffer from Squamous cell carcinoma [6, 7], whereas Large cell carcinoma comprises 5–10% of total lung cancer [8]. Categorizing type of malignancy based on histopathology and immunohistochemistry is needed for prognostication and deciding the line of treatment which varies accordingly.
One of the important risk factor of lung cancer is Cigarette Smoking [7, 8]. Cigarette smoke consist of tar made up of 3500 carcinogenic substances [9],which are further converted into DNA adducts by enzymes. DNA adducts causes transition in tumor suppressor gene [10]. Although smoking is considered as a prominent risk factor for developing lung cancer, 10–25% of total lung cancer occurs in never smokers [11]. Some of the other risk factors are air pollution, arsenic substances, radon, asbestos and medical history of chronic bronchitis, pneumonia, tuberculosis, emphysema, asthma [12].
Non-Small Cell Lung Cancer has 4 main stages: (a) Stage IA1, IA2, IA3, IB (b) Stage IIA, IIB (c) Stage IIIA, IIIB, IIIC (d) Stage IVA, IVB which is based on the location and size of primary tumor (T), status of lymph node (N) and distant metastasis (M) [13, 14]. Overall stages of 0, I, II, III, IV can be assigned by combining T, N and M values. Descriptions of each of the stages are mentioned in Table 1.
Staging of cancer, which is based on CT and PET-CT scans, is important to prognosticate and to decide the line of management. Hence there is a need to use deep learning strategies to automatically and accurately detect stages in cancer.
1.1 Need for Diagnostic Tool
Unfortunately the symptoms of lung cancer appear late in the course of the disease where it is non-curable. However, there are two major problems: (1) Lack of recognition of the presenting symptoms from lung cancer and (2) Additional time taken to complete diagnostic investigations [4]. The symptoms of lung cancer being common to many other non-serious lung diseases, the diagnosis of lung cancer are usually missed in the early stages. A study in Southern Norway noted that more than two-third of patients was already in advanced stage of lung tumor at the time of diagnosis (stage IIIB and IV) [15]. Delay in diagnostic imaging results in increase in tumor size and stage which negatively impacts the prognosis of lung cancer [16].
Several imaging modalities can be used to diagnose and determine the stage of lung cancer such as Chest X-ray, Computed Tomography (CT) Scan, Positron Emission Tomography (PET) Scan, MRI, CE-CT (Contrast-Enhanced Computed Tomography) etc. Chest X-ray has also been used for screening but did not help much in early detection of tumor. A false negative chest x-ray contributes to substantial delay in diagnosis [17]. Magnetic resonance imaging can identify hepatic metastases which are greater than 6 mm in diameter but may miss small pulmonary metastases [18]. In recent years, low-dose CT scan (LDCT) has been used in screening people who are at a higher risk of developing lung cancer in future. Screening is the test used to find disease in people who have not developed symptoms. LDCT scans can help find abnormal areas in the lungs that may be labeled as cancer. Research has shown that use of LDCT scans in screening high risk people of lung cancer saved more lives when compared to x-ray. In 2011, it was shown by National Lung Screening Trial (NLST) that participants who were screened using LDCT scans had (15–20%) lower risk of dying from lung carcinoma than those screened using chest X-rays [19]. Integrated PET/CT combined the benefits of metabolic and morphological information from PET and CT scan in the staging of Non-small-cell lung cancer [20, 21].
2 Deep Learning
Deep Neural Network (DNN) is a subfield of machine learning and has been widely used in medical image processing domain with promising results and it is expected that it will hold 300 million dollars in medical-imaging market by 2021 [22]. However the two major challenges of deep learning are usability and scalability [23]. The input to neural network is manually extracted features whereas Deep Neural network can extract features on its own from multiple hidden layers. Deep Learning has several advantages: (a) It can directly extract features from the training set. (b) Feature selection process is simple. (c) The three steps of extracting feature, selecting feature and classification can be integrated in the same deep learning architecture [24]. Deep Learning in chest radiology is active area of research in recent years.
Several deep learning networks are being used today namely: Recurrent Neural Network (RNN) [25], Recursive Neural Network (RvNN) [26], Deep Belief Network (DBN) [27, 28], Deep Boltzmann Machine (DBM) [29, 30], Auto encoder [31, 32], Variation Auto Encoder (VAE) [33], Stacked Auto Encoder [34], Deep Residual Network [35, 36], Convolution Neural Network (CNN) [37], Generative Adversarial Network (GAN) [38]. A few categories of deep learning architecture [39] is shown in Fig. 1.
Recurrent Neural Network [25] connects the nodes like a directed graph and remembers the previous input to predict the output. Recursive Neural Network [26] is a special case of Recurrent neural network which applies same set of weight for a structured input. Both Recurrent neural network and Recursive neural network is used for natural language processing. Deep Belief Network [27, 28] is a deep network of multiple connected hidden layers of restricted boltzmann machine, but the units within the layer are not connected. Deep Boltzmann Machine is a unsupervised learning process similar to DBN, but has undirected connection of hidden layers. DBN can be used for speech and image recognition. They also observed that the accuracy of the unsupervised approach used by Deep Convolution Belief Network increases with addition of new unlabeled data [40]. Autoencoder has a encoder and decoder neural network which learns the data through a iterative process. Variation Auto Encoder is a type of autoencoder that regularises the training process. Sparse Autoencoder is an autoencoder which activates only few hidden layer based on the error that occurs while encoding the input. Stacked Autoencoder is a sequence of sparse encoder where the output of one hidden layer is input to next hidden layer. Autoencoders are mostly used in denoising image and extraction of features from image. Convolution neural network has a sequence of convolution layers to extract features from images,followed by fully connected layer. Generative Adversial Network uses generator and discriminator model which in turn push each other to improve their performance. Deep Residual Network is convolution network that skips of some layers. CRBM are composed of Convolution Restricted Boltzmann machine. CNN have proved to provide best results with respect to image classification.
Two main issues with respect to deep neural network are: over fitting and processing time. Over fitting occurs when the network learns pattern present in training set but cannot learn patterns beyond it. Hence the training set has to be huge and maintain a perfect balance to cover all possible patterns that could occur in real time [41].
3 Deep Learning for Lung Nodule Classification
3.1 Preliminaries
Detecting nodule in early stage can improve the chances of survival of the patient. Determining the cancer stages can also help in providing appropriate treatment to the patients. Interpreting the diagnostic images is quite challenging for the radiologist as the dimensionality of CT images are quite huge and nodule could be of smaller size. Some of the features that effect visibility of nodule are density, size, location and contour. Predicting lesion of smaller size (less than 5 mm) is difficult. Even in LDCT scans factors which may lead to lesions being missed are: (1) Location close to structures like the skeletal system, blood vessels, pulmonary hila, diaphragms or airways which can limit visibility. (2) Low Nodule density like ground-glass [42] or sub solid density. (3) Inter-Observer variation [43]. Peri-fissural nodules are solid homogeneous nodule with smooth margin attached to the fissure (typical) or not attached to the fissure (atypical). Both the peri-fissural nodules are non malignant.
And hence computer aided Lung Cancer detection is needed to assist the radiologist to make accurate decisions. CAD output acts as a second opinion to assist radiologist by improving the accuracy of radiological diagnosis and also reduces the image reading time. CAD system are divided into two types: 1) CADe (Computer Aided Detection) System 2) CADx (Computer Aided Diagnostic) [44]. CADe can be used to detect lesion whereas CADx is used to determine malignancy in tumors along with staging [24]. The four main steps in CADe are: segmentation of the lung parenchyma, detection of the candidate nodules, false positive reduction and classification. The most crucial steps in computer aided detection are to reduce the false positive rate in lung nodule detection [45]. The 3 main steps in CADx are: extraction of feature, feature subset selection and classification which have to be integrated together [46].
Machine Learning tools and techniques have been widely used in diagnosis and prognosis of lung cancer. They can detect lung cancer at a very low radiation dose of 0.11 mSv. Neural networks were found to have higher sensitivity of 91.5–95.9% for lung cancer detection at standard dose [47].
The two basic approaches used in classification of lung nodules are: (1) Radiomics approach which is based on extraction of image features from Lung CT scan. (2) Convolutional Neural Networks (CNN). Radiomics approach builds the model by extracting either 2D or 3D image features of lung nodules. It also requires appropriate Lung Segmentation and Feature Extraction algorithm to classify the tumor whereas CNN do not necessarily require segmentation and feature extraction phase. However CNN requires very large dataset.
The five basic steps in Lung Nodule Classification using Radiomics approach are: Image Acquisition, Image Pre-processing, Image Segmentation, Feature Extraction and Lung nodule Classification as shown in Fig. 2. In Image Acquisition, we extract the image from either 64, 256 or 320 slice CT scanner. Image preprocessing stage enhances the image by removing noise. The tumor is then segmented from the region of interest using various segmentation algorithm in the image segmentation stage. Several features such as texture, gradient, shape are then extracted from the segmented image. The classification algorithm then uses these extracted features to classify the tumor into malignant and benign.
The basic steps involved in classification of lung nodule, using deep Convolutional Neural Network are shown in Fig. 3. In CNN model the image is passed through a series of convolution layer with filter, pooling layer, fully connected layer and softmax layer. The first convolution layer extracts low level features from the input image where as the subsequent layers extract the semantic features. In convolution layer, a kernel slides over the input to produce output using dot product, which is concatenated with bias and a non linear activation function like ReLU is applied on it. The output of the convolution layer is sent to the pooling layer to reduce the dimensionality of image at the same time retaining the necessary information of image. A sequence of convolution and pooling layer extract high level features from the image. The feature map is then flattened to a one dimensional vector and fed to a fully connected network. A fully connected neural network contains multiple hidden layer with weights and bias. Neural network uses a non-linear activation function as they allow back propagation whereas linear activation function does not support back propagation as the derivative of function is not related to inputs. The performance of the neural network will not improve with increase in hidden layers until we use a non-linear activation function. Finally a activation function such as sigmoid or softmax is applied to classify the object based on the probability from 0 to 1.
3.2 Data Acquisition
Lung CT scans can be obtained from various sources namely: LIDC/IDRI [48], NLST [19], LUNA16(Lung Nodule Analysis 2016) dataset [49], VIA/I-ELCAP [50], SPIE-AAPM Lung CT Challenge [51], Kaggle Data Science Bowl dataset [52], LISS (Lung CT Imaging Signs) [53], RIDER (Reference Image Database to Evaluate Therapy Response) [54], Lung CT Diagnosis [55]. LIDC/IDRI collects a dataset of 1018 cases with slice thickness varying from 0.45 mm to 5.0 mm and an associated XML file to record two-phase annotations from 4 experienced thoracic radiologists for the nodules > = 3 mm. Malignancy level of each nodule is rated from (1– 5) where 1 means low and 5 means high. Radiologist also rated them on variety of aspects such as internal structure, malignancy, texture, margin, sphericity, subtlety, calcification, lobulation and spiculation. Nodules with average malignancy level less than 3 are considered as non–malignant by few researchers [56]. NLST has collected 75,000 low dose CT screening images which can be utilized for research purposes and mostly consist of baseline scan(T0) followed by 2 successive scans(T1 and T2). LUNA dataset was collected from LIDC/IDRI database with a total of 888 CT scans. VIA/I-ELCAP database consist of 50 low dose CT scans obtained at a single breath hold and 1.25 mm slice thickness, obtained from ELCAP and VIA research groups. SPIE-AAPM-NCI consists of 70 lungs CT scan, most of which are contrast enhanced. Kaggle Data Science Bowl (DSB) dataset consist of 1397 instances of low dose CT scan in DICOM format along with series of axial 2D slices of size 512X512. DSB datasets mostly contains nodules of larger diameter average of 13.68 mm) and mostly located at the main bronchus. LISS database comprises of 271 Lung CT scans which are divided into 9 classes namely: Grand Grass Opacity(GGO), Cavity and Vacuolus(CV),Bronchial Mucus Plugs(BMP), Spiculation(S), Lobulation(L), Obstructive Pnemonia(OP), Calcification(C), Plueral Indentation(PI), Air Brochogram (AB). RIDER Lung CT comprises of 32 lung CT scans of patient having Non small cell lung cancer each of whom underwent two CT scans.
3.3 Image Preprocessing
Image preprocessing is needed to improve the quality of the grayscale CT image by reducing the noise. CT images are mostly effected by Gaussian noise, Poisson noise and artifacts. Several denoising technique like dictionary, filter and transform based approaches can be used to remove noise and artifacts from CT and Low dose CT images. Effective preprocessing technique can facilitate accurate segmentation of lung nodule. F Liao et al. converted the CT lung image into Hounsfield Unit, which is binarized using threshold, selected the connected part of lung, separated the left and right lung with erosion and dilation, computed the convex hull, dilated and combined left and right mask and filled the mask with luminance [57]. Density tendency correction could also be used segmenting lung nodule [58]. CNN architecture like Alexnet mostly needs RGB image of same dimension and hence Haritha et al. and Lei et al. transferred the image from grayscale to RGB and then performed thresholding, segmentation, erosion, dilation to obtain the mask of CT image [59, 60]. However Rahul paul et al. took information from R channel and ignored the B and G channel instead of converting grayscale to RGB image [61]. Allison et al. showed that accuracy of CNN was better on unsmoothed image than on smoothed image. As the lung volumes are too large and are of varying size, feeding the data in deep learning model is challenging and hence a resizing is needed. CT images are preprocessed and reduced to either 128X128 size [62, 63] and 150X150 sizes [60] for 2D images or 20X20X10 for 3 D images [64] based on the maximum size of nodules. Gaussian scale space filter [65, 66] or Gabor filter [67] could also be used to enhance the image. As the nodule size varies, bi-cubic interpolation could also be applied on nodule images [61, 68]. Kazuki et al. pre-processed the CT image by applying isotropic voxel [58] and Gaussian filter to the original images and performed binarization, thresholding, binary inversion, interpolation and thus extracted the lung field [69]. In order to extract 3 dimensional features from CT scan multiple views of CT scans could be combined. Tiantian et al. [56] combined the scans from multiple views using median intensity projection and enhanced using bilinear interpolation. Teramoto et al. [70] extracted the nodule from CT and PET images separately and then combined them using logical OR function where in active contour filter was used to extract the nodule from CT images. Training the model with very few training samples can cause under fitting issues, whereas over learning the training datasets, including the noise can cause over fitting problem. Overfitting problem can be addressed using data augmentation [34, 56, 64, 67, 69, 71,72,73,74,75,76,77], regularization [71, 78], dropout [79], translation [80, 81]. Data augmentation can also be performed by implementing image preprocessing techniques such rotating [77] and flipping the images horizontally and vertically [57, 68] or by using random scaling [82], adding noise [56, 72] so as to increase the number of training samples. Severe class imbalance in the dataset, can cause misclassification of minority class samples, which can also be addressed using data augmentation.
3.4 Image Segmentation
Lung nodule segmentation is a major task in lung nodule classification. We need to segment the lung parenchyma followed by the nodule segmentation in order to inspect the nodule for malignancy. Pulmonary nodule is segmented in training and testing phases. Four different types of pulmonary nodules are solid, semi-solid, non-solid and calcified. The surface of the large solid nodule (> 10 mm) have different intensity range than the smaller lesions which is difficult to capture using solid nodule detection algorithm. The most challenging step would be to separate out the juxta-pleural and juxta-vascular nodules as the contrast of large solid nodule with pleura is low [83]. Candidate nodule can be segmented using local geometric based filter and is found to increase the sensitivity of 3D CNN. Most of the lung nodules are segmented manually based on annotation provided by radiologist [84]. To facilitate the development of lung nodule detection, LUNA dataset also provides a annotation file which provides the x, y, z coordinates of nodule in the CT scan along with its ground truth labels and hence does not require the need for segmentation of lung nodule. Shou wang et al. used both, box and center region-of-interest [85, 86]. Since the shape of the nodule is irregular, we extract a bounding box of the nodule manually [87] or by using deep convolution neural network such as Faster region CNN [7, 44, 82, 87] or fully convolution neural network [57]. Segmentation of the nodule can be automated using different techniques namely border analysis, region based model [67, 85], shape and probabilistic models. Regional method includes regional growth, threshold and clustering method. Threshold method includes iterative, maximal interclass variance method [85], entropy method. Markov gibbs random field was a conventional approach to extract region of interest [88]. Clustering algorithm like k-means clustering [89] and fuzzy c-means clustering algorithm have also shown better accuracy in segmentation of lung nodule [90]. Recently deep learning techniques are also used in lung segmentation. Jeovane H. et al. used fully convolution networks (FCNs) combined with conditional random field for segmenting lung image [80, 91]. Taolin et al. [78] applied binary mask, thresholding, erosion, dilation, closure operation to segment the lung image. Kazuki et al. [42] removed the vessel and bronchial region by using 3D line filter based on Hessian matrix, and extracted the candidate regions using density gradient, volume and sphericity and then segmented the lung using threshold segmentation. They also reduced the false positive shadow of blood vessel using adaptive ring filter. Mitsuaki et al. [58] extracted the nodule from temporal subtraction images using super pixel and graph cut algorithm, retrieved 5 different features of nodule such as standard deviation, sphericity, slenderness, average density, slenderness and max vector degree of concentration and reduced the false positive nodules using SVM. Rahul paul et al. [61] showed that the accuracy of classification is relatively higher when features are extracted from cropped tumor of size 56X56 than the features extracted from warped tumor patches or patches obtained from sliding windows. Combination of features gathered from multiple view of CNN provides better segmentation results even on ground glass opacity nodules as suggested by Wang [92]. Some negative sample appears similar to nodule and hence difficult to classify. This problem can be solved using hard negative mining [57, 69]. Since most of the false positive in nodule detection is also due to airways in lungs, a 3D segmentation algorithm is needed [66].
3.5 Feature Extraction
Two types of features can be extracted: (1) Hand-crafted features (2) Features extracted from convolution neural network. Some of the handcrafted features include texture [61, 64, 67, 68, 90, 93,94,95,96,97,98,99,100,101],wavelet [93, 99], Fourier descriptor [94], shape [32, 64, 67, 70, 95, 98, 100, 101], gradient [95, 97, 101], density [97, 98], Histogram of Oriented Gradients (HoG) features [102], statistical [95, 96, 103], size [32, 64, 68]. Hand-crafted features describe low-level visual features instead of high-level features [85]. A study by Kuruvilla et al. [63] showed that statistical features like kurtosis, mean, standard deviation, central moment increases the accuracy of classification system. Metastasis features such as body temperature, insomnia, bodyweight loss, constipation, breathlessness, heart rate, hypercalcaemia, temperature, fatigue, blood pressure could also be used to predict the stages of lung tumor [67]. Ground glass opacity is quite difficult to detect but is considered to show a spherical shape when viewed in 3 dimensions and shape feature can be extracted to detect a Ground glass opacity tumor. CNN does not require tumor segmentation and feature selection phase.
Nodule detection includes two sub tasks: (1) to detect all positive nodules (2) To reduce the false positive nodule [44, 57, 82, 87]. Several variations in CNN have been utilized. Few of them are usage of Particle Swarm optimization with CNN to optimize the hyper parameters [104], Multi-Crop Convolution Neural Network (MC-CNN) [105], Multi Scale Convolution (MS-CNN) [65, 66, 103, 106,107,108,109], Multi-Level based Deep CNN (ML-CNN) [110,111,112,113], Convolutional Deep Belief network [40], Multi-Pathway CNN architecture (MP-CNN) [114], Multi-Resolution CNN (MR-CNN) [115], Region based CNN (R-CNN) [44, 82, 87, 116], Multi-View CNN (MV-CNN) [72, 79, 80, 103, 117, 118, 119], CNN with dense and shortcut connection [60], Contextual CNN [45, 120], CNN with multi channel region of interest [97], combining output of Multi View CNN using data fusion technique [117]. Region based CNN selects a pre fixed number of region proposals which can further be used for nodule classification [87]. Multi Crop CNN concatenates feature maps obtained by max-pooling the convolution and centre cropped convolution features [121]. Multi scale CNN takes nodule patches from multiple scale or size and combines them in parallel [65]. Multi view CNN concatenates multiple view of lung nodule into single view [72]. Multi level CNN combines multiple CNN with different kernel size together [113]. Several CNN architectures has been proposed for both segmentation and classification. Few of the CNN architecture used are LeNet [100, 122], AlexNet [42, 58, 59, 61, 78, 97, 122,123,124], StochasticNet [75], ZedNet [125], GoogleNet [56, 126], VGGNet [92, 133], ResNet [36, 64, 79, 80, 82, 111, 123, 127,128,129], DenseNet [107, 130,131,132], Tumornet [67, 72], U-Net [44, 57, 69, 80, 114, 120, 126, 133, 134], Dual-path Network [44], Overfeat [135, 136], ReCTnet [137], Xception [138]. Alex-net contains 8 layers, which comprises of 5 convolution layers, 3 intermediate pooling layers and finally 3 fully connected layers [97]. TumorNet comprises 5 convolution layers with max-pooling layer in first, second and fifth convolution layers, 3 fully connected layers and a softmax layer [72]. ResNet consist of multiple small network connected by shortcut connection [128]. U-Net consists of contraction and expansion path mostly used for medical image segmentation [114]. Dense-Net is convolution network where in each layer is connected to every other layer [107]. Dual Path network can gather the benefit of Residual network and dense network by eliminating gradient descent problem and by exploring new features respectively [44]. Few research papers have also integrated hand crafted features with CNN features in order to improve the accuracy [61, 71, 139, 140]. Rahul paul et al. [61] observed that the accuracy of integrated feature classified using naive bayes showed improvement of 82.5% over CNN feature alone classified using random forest (75%). Transfer learning can also be used which uses pre-trained networks to extract features and can reduce the over fitting problem when the dataset are comparatively small [59, 61, 68, 77, 141, 142].
There are 2 types of CNN features: (1) 2D CNN features [61, 73,74,75, 77, 86, 104, 110, 112, 114, 117, 122, 124, 129, 138, 141, 143,144,145,146,147] (2) 3D CNN features [41, 44, 45, 57, 60, 64, 66, 69, 71, 78, 79, 82, 87, 94, 105,106,107,108, 111, 115, 118, 120, 120, 127, 128, 130, 132, 134, 139, 141, 142, 148,149,150,151,153]. 2D CNN mostly considers the CT slice which had the largest area. However 2D CNN could not extract spatial information and correlation between slices. Some researchers also used both 2D and 3D features [107]. Various cross-sectional images can be used in axial, coronal, sagittal views to capture 3D features. 3D data analysis requires higher computation cost and hence 3D data can also be represented in 2D patch using trigonometric sampling approach to lower the computation cost [138]. Some researchers have used CNN for both detection and classification [134, 138]. The discrimination of nodule from tissues such as blood vessels is difficult from axial slices [69]. Lei et al. extracted 3D feature by applying CNN on 20 unified slices.
A detailed summary of Features extracted in few of the research articles are discussed below in Table 2.
3.6 Classification Algorithm
Some of the most commonly used classifiers for classifying nodules are Random Forest [61, 65, 68, 135, 139, 156, 160], Artificial Neural Network [16, 100], support-vector machine (SVM) [67, 70, 77, 98, 103, 136, 154], Linear Discriminant Analysis (LDA) [93], Decision Tree [74, 84, 93], Boosting [44, 94, 143], Linear Regression [102], Logistic Regression [64, 140], Random Forest Regression [156], Gaussian Process Regression [72], Naïve Bayes [61, 160], DBScan [106], Multi Kernel Learning [95, 101].
Several other deep architectures also seem to show promising result like Deep Belief Network (DBN) with RBM [40, 93, 99, 155], Deep Residual Network (DRN) [36, 82, 111], Deep Reinforcement Learning (DRL) [105, 162], Multi-layer perceptron model [96], Deep Denoising autoencoder [32, 98, 99, 102, 140], Deep Sparse Auto-encoder [62, 140], Generative Adversarial Network [120], Deep Stacked Autoencoder [34, 88], Convolutional Neural Network [34, 41, 42, 45, 56,57,58,59,60, 63, 64, 66,67,68,69, 71,72,73, 76, 78,79,80,81, 86, 87, 97, 99, 101, 104, 107, 108, 110,111,112, 114, 115, 117, 118, 120,121,122,123,124,126,127,128,129,130,131,132,132,133,134,135,136, 138, 141,142,143,144,145,146,147,148,149,150,151,153, 155, 161]. Ensemble model can also be used which combines several deep learning model to form optimal model. Final output in an ensemble model can be obtained by combining the prediction from multiple ensembles by using voting, averaging probabilities, max, median etc. Allison et al. [63] found that voting system did not improve the accuracy of system while combining the result of 2D CNN model applied on both smoothed and unsmoothed images. Some study also found that scaled logistic output function outperforms a softmax function [64]. A DBN is a feed-forward neural network with multiple hidden layers which is composed of Restricted Boltzmann Machine (RBM) [93]. An auto encoder is a 2 layered network that takes binary input and encodes/decodes the data using linear and non-linear transformation [84]. A Reinforcement Learning models data with incremental feedback rather than labeled data [105]. A Sparse Auto Encoder (SAE) can learn features from unlabeled data and a Deep Sparse Auto Encoder is made up of multiple layers of SAE which is trained layer by layer [62]. A Denoising Auto Encoder (DAE) contaminates the original image by adding noises but trains the model to output image similar to original image without contamination. Stacked Denoising Auto Encoder (SDAE) uses a stack of such denoising autoencoder to form a deep architecture [98]. Convolutional Neural Network (CNN) consists of input, hidden and output layer, where the hidden layer consists of convolution, ReLU, pooling, fully-connected layers.
Although CNN has shown promising results, several variation of CNN is being proposed. Multi path CNN proposed by Sori et al. have shown better results than CNN by concatenating the features from multiple paths of CNN [114]. Tajbakhsh et al. found that massive training artificial neural network shows better accuracy when compared to CNN with less training set [153]. Multi scale CNN as proposed by Wei Shen et al. showed better results with noisy data thus avoiding the data pre-processing stage [58]. Researchers have also used ensemble classifiers which combines several models into one predictive model to improve the performance of classification system [68]. Xie also showed that the concatenation of CNN features with texture and shape provides accurate results to represent heterogeneity of nodule but increases the computation time [83]. Wei Shen also proposed multi crop CNN to predict the nodule semantic attribute by using max-pooling on multiple cropped region of the convolution [93]. Yuan fused the features from multi view, multi scale CNN along with geometrical features to classify different types of nodule based on appearance and showed that the accuracy was better than using multi scale or multi view CNN alone [103]. The prediction of multiple CNN on different preprocessed images were ensemble using voting system by Allison to reduce the false positive rate in classification [63]. Devinder used autoencoder to extract the features which maps the semantic features [84]. Giovanni showed that CNN parameters can also be optimized using optimization algorithm like particle swarm optimization algorithm which increases the sensitivity of classification system [104]. Masood et al. [67] found that deep fully convolution network shows better performance than tumornet. It was also observed that accuracy of Alexnet was better than Resnet [124]. Albert et al. [126] found that the number of false positive increased with the use of U-Net and hence the output of U-Net was combined with Vanilla and GoogleNet to reduce the false positive. Chen et al. found that the sensitivity of the classification system increases with increase in convolution layers whereas sensitivity decreases with increase in kernel size [141]. Xinyu et al. observed that the increase in number of CRBM layers in DCBN (Deep convolution belief network) increases the accuracy of classification [40]. Kim et al. also found that sensitivity can be increased by combining morphological features with features extracted from Stacked Denoised Autoencoder [98]. A comparative study of various deep learning algorithm used for lung nodule classification and its performance is shown in Table 3.
4 Conclusion
Currently Deep Learning is used for lung segmentation, feature extraction, nodule detection, false positive reduction and classification. Most of Machine Learning algorithm needs to understand the data to extract relevant features for classification. Figure 4 gives a brief summary of contribution of different deep learning architecture used in the classification of lung nodule.
Feature selection hugely impacts the performance of the system. However feature selection process is automated in deep learning algorithm like CNN. Due to the success of CNN in most of the image recognition application, CNN is mostly being used in medical image processing for detection of nodules. The advantages of using CNN architecture in lung nodule classification are:1) CNN is most commonly used approach in detection of lung nodule and also outperforms the conventional models [155]. 2)Both feature extraction and classification are combined together by convolution, pooling and fully connected. 3)CNN does not require a segmentation layer. 4) Feature selection is also not needed in CNN as the features are selected by the convolution layer. 5) Convolutional Neural Network has proved to provide better results with respect to lung nodule classification and hence most of the research is focused on improvising CNN. 6)CNN also provides better performance when compared to DNN and Stacked AutoEncoders [34]. 7) A CNN feature also proves to be better than other hand crafted features like shape, size and texture [136]. Some of the challenges faced in using CNN are: 1)CNN works well on huge datasets but gathering real life dataset is quite challenging because of the time it takes. 2)Overfitting problem can also arise if there is imbalance in the dataset. 3)CNN requires several hyper parameters to be set which requires lot of trial and expertise. 4)CNN also requires very large computation and huge processing time and hence require high end GPU’s.
Although size is an important factor for determining malignancy [64], size alone cannot help in determining the malignancy levels [154]. An ensemble model which combines the quantitative features of CT image with CNN seems to improve the accuracy of model [68]. Combination of high level features such as lobulation, margin, calcification, spiculation, sphericity and CNN features improves the classification accuracy [72, 136, 160]. The relationship between adjacent consecutive frames cannot be analyzed with 2D CNN. However 3D CNN can extract spatio-temporal data, but the computation cost would be higher. 3D multi-view CNN was found to provide better accuracy than 2D Multiview CNN and also does not require a very deep network [79, 132]. Most of the experiments also show that 3D CNN shows better sensitivity than 2D CNN [60]. As 3D-CNN is expensive, several variations in 3D-CNN have been proposed to reduce the number of computation. Figure 5 gives a brief summary of number of researcher using 3D CNN and 2D CNN.
Multi-view CNN has lower error rate than single-view CNN [79]. Multi scale CNN can help detect pleural (between chest wall and lung) nodule [76]. Multi scale CNN works better even on noisy input [79]. Figure 6 gives a brief summary of number of researcher using different variation in CNN.
Several CNN Architectures are being used for classification, but Res-Net, Alex-Net was most commonly used. Different CNN Architecture could also be ensemble in order to improve the sensitivity the classification system. Combination of CNN with RNN or LSTM have shown better results than using each architecture individually. CT prediction can be combined with clinical probabilities to improve the prediction. However there is no previous studies combining clinical data with CT data. Figure 7 gives a brief summary of contribution of different deep CNN architecture with respect to lung nodule classification.
CNN works better with large dataset. Transfer learning is also being used by several researchers [56, 59, 61, 68, 160] which reduces over fitting problem. Use of 2D CNN instead of 3D CNN can increase the volume of lung CT scan for training. Integrating medical images from multiple sources also tend to improve the accuracy of the system [108]. CNN parameters such as kernel size, batch size, learning rate, weight initialization play an important role in effecting the performance of the CNN network [122]. Convolution layer and kernel size affect the sensitivity of the CNN model [141]. Setting up the hyper-parameters for deep neural network is quite a challenging task. Some study also show that performance can be improved by increasing the number of epoch [59, 63]. Classification error can also be reduced by augmenting the data if the dataset is small [73]. Combining convolution network of different configuration gives better sensitivity [125].
Although there are great progress made in the field of lung cancer classification, some of the areas still needs to be explored: 1) Most of the research papers have focused on binary classification of lung tumors as benign and malignant. Malignancy can also be determined using type of nodules which is based on the position and its intensity. Some of the nodule types are well circumscribed (W), non-nodule (N), juxta pleural (J), ground glass opacity (G), pleural tail (P), vascularized (V). However very few of the research papers have focused on ternary classification where in the tumors are classified into benign, primary malignant and metastasis malignant as shown by Kang [75], solid, semi-solid, ground glass opacity as shown by Wei [86, 156], nodule type like W, N, J, G, P, V as shown by Liu [76]. 2) Very few papers have focused on detection of nodules below 3 mm diameter. However Patrice [148] classified micro-nodules from non-nodules. Most of the early stage malignant tumors are smaller in size and if these tumors are detected at early stage,it might increase the life expectancy of the individual. 3) Detection of ground glass optical and non-nodules are difficult and are explored by very few researchers like [76]. 4) Segmenting out large solid nodules attached to the pleural wall is quite challenging and needs to be explored. 5)Some researchers have used private dataset which becomes very difficult to compare the results with different algorithm and hence there is a need of global dataset which can be used by all researchers. 6)One of the major challenges in lung cancer detection is to differentiate between early stage small cancerous lesion and benign nodule which can be confirmed by both invasive and non-invasive method. The invasive method includes use of biopsy whereas non-invasive method requires follow-up of CT scans periodically. The invasive method can cause complications like bleeding and infection of wounds whereas non-invasive method can increase the risk of radiation. Hence there is a need for advanced deep learning technique to identify the morphological difference between early stage cancerous nodule and benign noduleto overcome the drawback of invasive and non-invasive cancer detection technique.7) Additional information of patients such as the medical history, genetic report can also be analyzed and fused with the deep features extracted from lung scan images to improve the efficiency of automated tumor detection. And hence a strong collaboration between physician and researchers needed to design a full proof model. 8)Due to privacy issues and to prevent the use of sensitive information, CT datasets cannot be easily shared for research purpose. 9)Majority of the researchers have limited their work to lung nodule detection; there is scope for utilization of deep learning techniques for automatic staging of cancer. Although many researchers have used various deep learning models with the aim to improve the accuracy of lung nodule classification, there is still scope of improvement in addressing the above challenges with the aim to detect the malignant tumors at an early stage.
References
Bray, F., Ferlay, J., Soerjomataram, I., Siegel, R. L., Torre, L. A., & Jemal, A. (2018). Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA: Cancer Journal for Clinicians, 68, 394–424.
Ferlay, J., Shin, H. R., Bray, F., Forman, D., Mathers, C., & Parkin, D. M. (2010). Estimates of worldwide burden of cancer in 2008: GLOBOCAN 2008. International Journal of Cancer, 127(12), 2893–2917.
Cheng, T.-Y. D., Cramb, S. M., Baade, P. D., Youlden, D. R., Nwogu, C., & Reid, M. E. (2016). The international epidemiology of lung cancer: Latest trends, disparities, and tumor characteristics. Journal of Thoracic Oncology, 11(10), 1653–1671.
Ellis, P. M., & Vandermeer, R. (2011). Delays in the diagnosis of lung cancer. Journal of Thoraic Disease, 3(3), 183–188.
Kenfield, S. A., Wei, E. K., Stampfer, M. J., et al. (2008). Comparison of aspects of smoking among the four histological types of lung cancer. Tobacco Control, 17, 198–204.
Denisenko, T. V., Budkevich, I. N., & Zhivotovsky, B. (2018). Cell death-based treatment of lungadenocarcinoma. Cell Death and Disease, 9, 117.
Doll, R., & Hill, A. B. (1999). Smoking and carcinoma of the lung. Preliminary report. 1950. Bulletin of the World Health Organization, 77(1), 84–93.
Gandini, S., EdoardoBotteri, S., Boniol, M., Lowenfels, A. B., Maisonneuve, P., & Boyle, P. (2008). Tobacco smoking and cancer: A meta-analysis. International Journal of Cancer, 122, 155–164.
Smith, C. J., Perfetti, T. A., Rumple, M. A., Rodgman, A., & Doolittle, D. J. (2001). “IARC Group 2B carcinogens” reported in cigarette mainstream smoke. Food and Chemical Toxicolog, 39(2), 183–205.
Olshan, A. F., Weissler, M. C., Pei, H., & Conway, K. (1997). p53 mutations in head and neck cancer: New data and evaluation of mutational spectra. Cancer Epidemiology, Biomarkers & Prevention, 6, 499–504.
Couraud, S., Zalcman, G., Milleron, B., Morin, F., & Souquet, P.-J. (2012). Lung cancer in never smokers—A review. European Journal of Cancer, 48, 1299–1311.
Akhtar, N., & Bansal, J. G. (2017). Risk factors of Lung Cancer in nonsmoker. Current Problems in Cancer, 41(5), 328–339.
Detterbeck, F. C., Postmus, P. E., & Tanoue, L. T. (2013). The stage classification of lung cancer diagnosis and management of lung cancer, 3rd ed: American College of chest physicians evidence-based clinical practice guidelines. Chest, 143(5), e191S–e210S.
Detterbeck, F. C. (2018). The eighth edition TNM stage classification for lung cancer: What does it mean on main street? The Journal of Thoracic and Cardiovascular Surgery, 155(1), 356–359.
Rolke, H. B., Bakke, P. S., & Gallefoss, F. (2007). Delays in the diagnostic pathways for primary pulmonary carcinoma in Southern Norway. Respiratory Medicine, 101(6), 1251–1257.
Byrne, S. C., Barrett, B., & Bhatia, R. (2015). The impact of diagnostic imaging wait times on the prognosis of lung cancer. Canadian Association of Radiologists Journal, 66(1), 53–57.
Bjerager, M., Palshof, T., Dahl, R., Vedsted, P., & Olese, F. (2006). Delay in diagnosis of lung cancer in general practice. British Journal of General Practice, 56(532), 863–868.
Lauenstein, T. C., Goehde, S. C., Herborn, C. U., Goyen, M., Oberhoff, C., Debatin, J. F., et al. (2004). Whole-body MR imaging: Evaluation of patients for metastases. Radiology, 233(1), 139–148.
The National Lung Screening Trial Research Team. (2011). Reduced lung-cancer mortality with low-dose computed tomographic screening. New England Journal of Medicine, 365(5), 395–409.
Chao, F., & Zhang, H. (2012). PET/CT in the staging of the non-small-cell lung cancer. Journal of Biomedicine and Biotechnology, 2012, 783739.
Ricard, F., & Giammarile, F. (2014). PET-CT and diagnostic CT: The synergy of metabolic and morphological data in onco-haematology. Diagnostic and Interventional Imaging, 95(1), 55–62.
Razzak, M. I., Naz, S., & Zaib, A. (2018). Deep learning for medical image processing: Overview, challenges and the future. In N. Dey, A. Ashour, & S. Borra (Eds.), Classification in BioApps. Lecture notes in computational vision and biomechanics (Vol. 26). Berlin: Springer.
Wang, W., Chen, G., Chen, H., AnhDinh, T. T., JinyangGao, B. C., Ooi, K.-L. T., et al. (2016). Deep learning at scale and at ease. ACM Transactions on Multimedia Computing, Communications, and Applications (TOMM), 12(4), 1–25.
Cheng, J.-Z., et al. (2016). Computer-aided diagnosis with deep learning architecture: Applications to breast lesions in US images and pulmonary nodules in CT scans. Scientific Reports, 6, 24454. https://doi.org/10.1038/srep24454.
Li, X., & Wu, X. (2015). Constructing long short-term memory based deep recurrent neural networks for large vocabulary speech recognition. In IEEE international conference on acoustics, speech and signal processing (pp. 4520–4524). IEEE.
Socher, R., Lin, C.C., Manning, C., & Ng, A.Y. (2011). Parsing natural scenes and natural language with recursive neural networks. In International conference on machine learning (pp. 129–136). Omnipress.
Hinton, G. E. (2009). Deep belief networks. Scholarpedia, 4(5), 5947.
Vinod, N., & Hinton, G.E. (2010). Rectified linear units improve restricted boltzmann machines. In Proceedings of the 27th international conference on machine learning (ICML-10) (pp. 807–814).
Salakhutdinov, R., & Hinton, G. (2009). Deep boltzmann machines. In Proceedings of AISTATS 2009 (pp. 448–455). PMLR.
Salakhutdinov, R., & Hinton, G. (2012). An efficient learning procedure for deep Boltzmann machines. Neural Computation, 24(8), 1967–2006.
Hinton, G. E., & Salakhutdinov, R. R. (2006). Reducing the dimensionality of data with neural networks. Science, 313(5786), 504.
Shaffie, A. et al. (2017). A new framework for incorporating appearance and shape features of lung nodules for precise diagnosis of lung cancer. In 2017 IEEE international conference on image processing (ICIP), Beijing (pp. 1372–1376).
Kingma, D.P., & Welling, M. (2013). Auto-encoding variationalbayes. arXiv:1312.6114.
Hamidian, S., Sahiner, B., Petrick, N., & Pezeshk, A. (2017). 3D convolutional neural network for automatic detection of lung nodules in chest CT. Proceedings of SPIE The International Society for Optical Engineering, 10134, 1013409. https://doi.org/10.1117/12.2255795.
Kaiming, H., Zhang, X., Ren, S., & Sun, J. (2016). Deep residual learning for image recognition. In Proceedings of the IEEE CVPR (pp. 770–778).
Schwyzer, M., Ferraro, D. A., Muehlematter, U. J., CurioniFontecedro, A., Huellner, M. W., von Schulthess, G. K., et al. (2018). Automated detection of lung cancer at ultralow dose PET/CT by deep neural networks—Initial results. Lung Cancer. https://doi.org/10.1016/j.lungcan.2018.11.001.
Alex, K., Sutskever, I., & Hinton, G.E. (2012). Imagenet classification with deep convolutional neural networks. In Proceedings of the 25th International Conference on Neural Information Processing Systems - Volume 1 (NIPS’12). Curran Associates Inc., Red Hook, NY, USA (pp. 1097–1105).
Goodfellow, I., Pouget-Abadie, J., Mirza, M., Xu, B., Warde-Farley, D., Ozair, S., Courville, A., & Bengio, Y. (2014). Generative adversarial nets. In Advances in neural information processing systems (pp. 2672–2680). Curran Associates.
Pouyanfar, S. (2018). A survey on deep learning: Algorithms, techniques, and applications. ACM Computing Surveys, 51(5), 92.
Jin, X., Ma, C., Zhang, Y., & Li, L. (2017). Classification of lung nodules based on convolutional deep belief network. In 2017 10th International symposium on computational intelligence and design (ISCID), Hangzhou (pp. 139–142).
Golan, R., Christian, J., & Denzinger, J. (2016). Lung nodule detection in CT images using deep convolutional neural networks. In 2016 International joint conference on neural networks (IJCNN). IEEE.
Hirayama, K., Miyake, N., Lu, H., Tan, J.K., Kim, H., Tachibana, R., Hirano, Y., & Kido, S. (2017). Extraction of GGO regions from chest CT images using deep learning. In 2017 17th International conference on control, automation and systems (ICCAS).
Naidich, D. P. (2004). Lung cancer detection and characterization: Challenges and solutions. In C. D. Claussen, E. K. Fishman, B. Marincek, & M. Reiser (Eds.), Multislice CT. Berlin: Springer.
Zhu, W., Liu, C., Fan, W., & Xie, X. (2018). DeepLung: Deep 3D dual path nets for automated pulmonary nodule detection and classification. In 2018 IEEE winter conference on applications of computer vision (WACV).
Dou, Q. (2017). Multilevel contextual 3-D CNNs for false positive reduction in pulmonary nodule detection. IEEE Transactions on Biomedical Engineering, 64(7), 1558–1567.
Way, T. W., et al. (2009). Computer-aided diagnosis of pulmonary nodules on CT scans: Improvement of classification performance with nodule surface features. Medical Physics, 36, 3086–3098.
Schwyzer, M. (2018). Automated detection of lung cancer at ultralow dose PET/CT by deep neural networks—Initial results. Lung Cancer, 126, 170–173.
Armato, S. G., III, McLennan, G., Bidaut, L., et al. (2011). The lung image database consortium (LIDC) and image database resource initiative (IDRI): A completed reference database of lung nodules on CT scans. Medical Physics, 38(2), 915–931.
Lung Nodule Analysis Challenge.<http://luna.grand-challenge.org>
Armato, S. G., III, Hadjiiski, L., Tourassi, G. D., Drukker, K., Giger, M. L., Li, F., et al. (2015). SPIE-AAPM-NCI lung nodule classification challenge dataset. Cancer Imaging Arch. https://doi.org/10.7937/K9/TCIA.2015.UZLSU3FL.
Kaggle Data Science Bowl (KDSB). (2017). https://www.kaggle.com/c/data-science-bowl-2017/data.
Han, G., Liu, X., Han, F., Santika, I. N. T., Zhao, Y., Zhao, X., et al. (2015). The LISS—a public database of common imaging signs of lung diseases for computer-aided detection and diagnosis research and medical education. IEEE Transactions on Biomedical Engineering, 62(2), 648–656.
Armato, S. G., McLennan, G., Meyer, C. R., Reeves, A. P., McNitt-Gray, M. F., Croft, B. Y., et al. (2016). The reference image database to evaluate response to therapy in lung cancer (RIDER) project: A resource for the development of change analysis software. Clinical Pharmacology and Therapeutics, 84(4), 448–456.
Grove, O., Berglund, A. E., Schabath, M. B., et al. (2015). Data from: Quantitative computed tomographic descriptors associate tumor shape complexity and intratumor heterogeneity with prognosis in lung adenocarcinoma, Lung CT-Diagnosis Data. Cancer Imag. Arch.. https://doi.org/10.7937/K9/TCIA.2015.A6V7JIWX.
Fang, T. (2018). A novel computer-aided lung cancer detection method based on transfer learning from GoogLeNet and median intensity projections. In 2018 IEEE international conference on computer and communication engineering technology (CCET).
Liao, F., Liang, M., Li, Z., Hu, X., & Song, S. (2017) Evaluate the malignancy of pulmonary nodules using the 3d deep leaky noisy-or network. arXiv preprint arXiv:1711.08324.
Nagao, M. et al. (2017). Detection of abnormal candidate regions on temporal subtraction images based on DCNN. In 2017 17th International conference on control, automation and systems (ICCAS), Jeju (pp. 1444–1448).
Sathyan, H., & Panicker, J.V. (2018). Lung nodule classification using deep ConvNets on CT image. In 2018 9th International conference on computing, communication and networking technologies (ICCCNT).
Fan, L., Xia, Z., Zhang, X., & Feng, X. (2017). Lung nodule detection based on 3D convolutional neural networks. In: 2017 International conference on the frontiers and advances in data science (FADS).
Paul, R., Hawkins, S.H., Hall, L.O., Goldgof, D.B., & Gillies, R.J. (2016). Combining deep neural network and traditional image features to improve survival prediction accuracy for lung cancer patients from diagnostic CT. In 2016 IEEE international conference on systems, man, and cybernetics.
Sun, B., Ma, C.-H., Jin, X.-Y., & Luo, Y. (2016). Deep sparse auto-encoder for computer aided pulmonary nodules CT diagnosis. In 2016 13th international computer conference on wavelet active media technology and information processing (ICCWAMTIP).
Rossetto, A.M., & Zhou, W. (2017). Deep learning for categorization of lung cancer CT images. In 2017 IEEE/ACM international conference on connected health: Applications, systems and engineering technologies (CHASE), Philadelphia, PA (pp. 272–273).
Luo, Z., Brubaker, M.A., & Brudno, M. (2017). Size and texture-based classification of lung tumors with 3D CNNs. In 2017 IEEE winter conference on applications of computer vision (WACV).
Shen, W., Zhou, M., Yang, F., Yang, C., & Tian, J. (2015). Multi-scale convolutional neural networks for lung nodule classification. In International conference on information processing in medical imaging (pp. 588–599). Springer.
Anirudh, R., Thiagarajan, J.J., Bremer, T., Kim, H. (2016) Lung nodule detection using 3d convolutional neural networks trained on weakly labeled data. In SPIE medical imaging (pp. 978532–978532).
Masood, A., Shenga, B., Li, P., Hou, X., Wei, X., Qin, J., et al. (2018). Computer-assisted decision support system in pulmonary cancer detection and stage classification on CT images. Journal of Biomedical Informatics, 79, 117–128.
Paul, R., Hall, L., Goldgof, D., Schabath, M., & Gillies, R. (2018). Predicting nodule malignancy using a CNN ensemble approach. In 2018 International joint conference on neural networks (IJCNN), Rio de Janeiro (pp. 1–8).
Tang, H., Kim, D.R., & Xie, X. (2018). Automated pulmonary nodule detection using 3D deep convolutional neural networks, In 2018 IEEE 15th international convolutional neural networks symposium on biomedical imaging (ISBI 2018).
Teramoto, A., Fujita, H., Yamamuro, O., & Tamaki, T. (2016). Automated detection of pulmonary nodules in PET/CT images: Ensemble false positive reduction using a convolutional neural network technique. Medical Physics, 43, 2821. https://doi.org/10.1118/1.4948498.
Huang, X., Shan, J., & Vaidya, V. (2017). Lung nodule detection in CT using 3D convolutional neural networks. In 2017 IEEE 14th international symposium on biomedical imaging (ISBI 2017).
Hussein, S., Gillies, R., Cao, K., Song, Q., & Bagci, U. (2017) Tumornet: Lung nodule characterization using multi-viewconvolutional neural network with gaussian process. In IEEE international symposium on biomedical imaging (ISBI) 2017.
Yang, H., & Yu, H. (2016). Deep learning for the classification of lung nodules. arXiv:1611.06651.
Devinder, K. et al. (2017). Discovery radiomics for pathologically-proven computed tomography lung cancer prediction. In ICIAR.
Shafiee, M.J., Chung, A.G., Kumar, D., Khalvati, F., Haider, M., & Wong, A. (2015). Discovery radiomics via stochasticnet sequencers for cancer detection. arxiv preprint arXiv:151103361.
Liu, X., Hou, F., Qin, H., & Hao, A. (2018). Multi-view multi-scale CNNs for lung nodule type classification from CT images. Pattern Recognition. https://doi.org/10.1016/j.patcog.2017.12.022.
Gupta, A., Das, S., Khurana, T., & Suri, K. (2018). Prediction of lung cancer from low-resolution nodules in CT-scan images by using deep features. In 2018 International conference on advances in computing, communications and informatics (ICACCI), Bangalore (pp. 531–537).
Jin, T., Cui, H., Zeng, S., & Wang, X. (2017). Learning deep spatial lung features by 3D convolutional neural network for early cancer detection. In 2017 International conference on digital image computing: Techniques and applications (DICTA).
Kang, G., Liu, K., Hou, B., & Zhang, N. (2017). 3D multi-view convolutional neural networks for lung nodule classification. PLoS ONE, 12(11), e0188290. https://doi.org/10.1371/journal.pone.0188290.
Xie, Y., Xia, Y., Zhang, J., Song, Y., Feng, D., Fulham, M., et al. (2018). Knowledge-based collaborative deep learning for benign-malignant lung nodule classification on chest CT. IEEE Transactions on Medical Imaging. https://doi.org/10.1109/tmi.2018.2876510.
Roth, H.R., Lu, L., Seff, A., Cherry, K.M., Hoffman, J., Wang, S., Liu, J., Turkbey, E., & Summers, R.M. (2014) A new 2.5 d representation for lymph node detection using random sets of deep convolutional neural network observations. In Medical image computing and computer-assisted intervention–MICCAI 2014 (pp. 520–527). Springer.
Dou, Q., Chen, H., Jin, Y., Lin, H., Qin, J., & Heng, P.-A. (2017). Automated pulmonary nodule detection via 3d convnets with online sample filtering and hybrid-loss residual learning. In MICCAI.
Setio, A. A. A., Jacobs, C., Gelderblom, J., & van Ginneken, B. (2015). Automatic detection of large pulmonary solid nodules in thoracic CT images. Medical Physics, 42(10), 5642–5653.
Kumar, D., Wong, A., & Clausi, D.A. (2015). Lung nodule classification using deep features in CT images. In 2015 12th conference on computer and robot vision (June 2015) (pp 133–138).
Wang, S., Liu, Z., Chen, X., Zhu, Y., Zhou, H., Tang, Z., Wei, W., Dong, D., Wang, M., & Tian, J. (2018). Unsupervised deep learning features for lung cancer overall survival analysis. In 2018 40th Annual international conference of the IEEE engineering in medicine and biology society (EMBC).
Wei, L., Cao, P., Zhao, D., & Wang, J. (2016). Pulmonary Nodule Classification with Deep Convolutional Neural Networks on ComputedTomography Images. Computational and Mathematical Methods in Medicine., 2016, 1–7. https://doi.org/10.1155/2016/6215085.
Ding, J., Li, A., Hu, Z., & Wang, L. (2017) Accurate pulmonary nodule detection in computed tomography images using deep convolutional neural networks. In MICCAI.
Shaffie, A., Soliman, A., Ghazal, M., Taher, F., Dunlap, N., Wang, B., van Berkel, V., Gimel’farb, G., Elmaghraby, A., & El-Baz, A. (2018). A novel autoencoder-based diagnostic system for early assessment of lung cancer. In 2018 25th IEEE international conference on image processing (ICIP).
Kockelkorn, T. J. P., Rikxoort, M., Grutters, C., et al. (2010). Interactive lung segmentation in CT scans with severe abnormalities. IEEE International Symposium on Biomedical Imaging: From Nano to Macro, 14, 564–567.
Meng, Y., Yi, P., Guo, X., Gu, W., Liu, X., Wang, W., & Zhu, T. (2018). Detection for pulmonary nodules using RGB channel superposition method in deep learning framework. In 2018 Third international conference on security of smart cities, industrial control system and communications (SSIC).
Alves, J.H., Moreira Neto, P.M., & Oliveira, L.F. (2018) Extracting lungs from CT images using fully convolutional networks. In 2018 International joint conference on neural networks (IJCNN).
Wang, S. et al. (2017). A multi-view deep convolutional neural networks for lung nodule segmentation. In 2017 39th Annual international conference of the IEEE engineering in medicine and biology society (EMBC), Seogwipo (pp. 1752–1755). https://doi.org/10.1109/embc.2017.8037182.
Lakshmanaprabu, S. K., Mohanty, S. N., Shankar, K., Arunkumar, N., & Ramirez, G. (2018). Optimal deep learning model for classification of lung cancer on CT images. Future Generation Computer Systems, 92, 374–382. (ISSN: 0167-739X).
Xie, Y., Zhang, J., Xia, Y., Fulham, M., & Zhang, Y. (2018). Fusing texture, shape and deep model-learned information at decision level for automated classification of lung nodules on chest CT. Data Information Fusion, 42, 102–110. (ISSN: 1566-2535).
Cao, P., Liu, X., Zhang, J., Li, W., Zhao, D., Huang, M., et al. (2017). A _ 2, 1 norm regularized multi-kernel learning for false positive reduction in Lung nodule CAD. Computer Methods and Programs in Biomedicine, 140, 211–231.
Singh, G. A. P., & Gupta, P. K. (2018). Performance analysis of various machine learning-based approaches for detection and classification of lung cancer in humans. Neural Computing and Applications, 31(10), 6863–6877.
Sun, W., Zheng, B., & Qian, W. (2017). Automatic feature learning using multichannel ROI based on deep structured algorithms for computerized lung cancer diagnosis. Computers in Biology and Medicine, 89(1), 530–539.
Kim, B., Sung, Y.S., & Suk, H. (2016). Deep feature learning for pulmonary nodule classification in a lung CT. In 2016 4th International winter conference on brain-computer interface (BCI), Yongpyong (pp. 1–3).
Sun, W., Zheng, B., & Qian, W. (2016). Computer aided lung cancer diagnosis with deep learning algorithms. In Proceedings of SPIE 9785, Medical Imaging 2016: Computer-Aided Diagnosis, 97850Z. https://doi.org/10.1117/12.2216307.
Xie, Y., Zhang, J., Liu, S., Cai, W., & Xia, Y. (2017). Lung nodule classification by jointly using visual descriptors and deep features. Lecture Notes in Computer Science, 10081, 116–125.
Cao, P., Liu, X., Yang, J., Zhao, D., Li, W., Huang, M., et al. (2017). A multi-kernel based framework for heterogeneous feature selection and over-sampling for com- puter-aided detection of pulmonary nodules. Pattern Recognition, 64(C), 327–346.
Chen, S., Qin, J., Ji, X., Lei, B., Wang, T., Ni, D., et al. (2017). Automatic scoring of multiple semantic attributes with multi-task feature leverage: A study on pulmonary nodules in CT images. IEEE Transactions on Medical Imaging, 36(3), 802–814. https://doi.org/10.1109/TMI.2016.2629462
Yuan, J., Liu, X., Hou, F., Qin, H., & Hao, A. (2018). Hybrid-feature-guided lung nodule type classification on CT images. Computers & Graphics, 70, 288–299.
Silva, G. L. F., Valente, T. L. A., Silva, A. C., Paiva, A. C., & Gattassa, M. (2018). Convolutional neural network-based PSO for lung nodule false positive reduction on CT images. Computer Methods and Programs in Biomedicine, 162, 109–118.
Ali, I., Hart, G. R., Gunabushanam, G., Liang, Y., Muhammad, W., Nartowt, B., et al. (2018). Lung nodule detection via deep reinforcement learning. Frontiers in Oncology, 8, 108. https://doi.org/10.3389/fonc.2018.00108
Yu, G., Xiaoqi, L., Yang, L., Zhang, B., Dahua, Yu., Zhao, Y., et al. (2018). Automatic lung nodule detection using a 3D deep convolutional neural network combined with a multi-scale prediction strategy in chest CTs. Computers in Biology and Medicine, 103, 220–231.
Zhan, J., Xia, Y., Zeng, H., & Zhang, Y. (2018). NODULe: Combining constrained multi-scale LoG filters with densely dilated 3D deep convolutional neural network for pulmonary nodule detection. Neurocomputing, 317(23), 159–167.
Tafti, A.P., Bashiri, F.S., LaRose, E., & Peissig, P. (2018) Diagnostic classification of lung CT images using deep 3D multi-scale convolutional neural network. In 2018 IEEE international conference on healthcare informatics.
Kim, B.-C., Yoon, J. S., Choi, J.-S., & Suk, H.-I. (2019). Multi-scale gradual integration CNN for false positive reduction in pulmonary nodule detection. Neural Networks, 115, 1–10.
Jiang, H., Ma, H., Qian, W., Gao, M., & Li, Y. (2018). An automatic detection system of lung nodule based on multigroup patch-based deep learning network. IEEE Journal of Biomedical and Health Informatics, 22(4), 1227–1237. https://doi.org/10.1109/JBHI.2017.2725903
Jin, H., Li, Z., Tong, R., & Lin, L. (2018). A deep 3D residual CNN for false-positive reduction in pulmonary nodule detection. Medical Physics, 45(5), 2097–2107.
Lyu, J., & Ling, S.H. (2018). Using multi-level convolutional neural network for classification of lung nodules on CT images. In 2018 40th Annual international conference of the IEEE engineering in medicine and biology society.
Kim, B.-C., Yoon, J., Choi, J.-S., & Suk, H.-I. (2019). Multi-scale gradual integration CNN for false positive reduction in pulmonarynodule detection. Neural Networks. https://doi.org/10.1016/j.neunet.2019.03.003.
Sori, W. J., Jiang, F., & Liu, S. (2019). Multi-path convolutional neural network for lung cancer detection. Multidimensional Systems and Signal Processing. https://doi.org/10.1007/s11045-018-0626-9.
Zuo, W., Zhou, F., Li, Z., & Wang, L. (2019). Multi-resolution CNN and knowledge transfer for candidate classification in lung nodule detection. IEEE Access, 7, 32510–32521.
Kido, S., Hirano, Y., & Hashimoto, N. (2018). Detection and classification of lung abnormalities by use of convolutional neural network (CNN) and regions with CNN features (R-CNN). In 2018 International workshop on advanced image technology (IWAIT).
Setio, A. A. A., Ciompi, F., Litjens, G., Gerke, P., Jacobs, C., van Riel, S. J., et al. (2016). Pulmonary nodule detection in CT images: False positive reduction using multi-view convolutional networks. IEEE Transactions on Medical Imaging, 35(5), 1160–1169. https://doi.org/10.1109/TMI.2016.2536809
Dobrenkii, A., Kuleev, R., Khan, A., Ramirez Rivera, A., & Khattak, A. M. (2017). Large residual multiple view 3D CNN for false positive reduction in pulmonary nodule detection. In 2017 IEEE conference on computational intelligence in bioinformatics and computational biology (CIBCB).
Xie, Y., Xia, Y., Zhang, J., Song, Y., Feng, D., Fulham, M., et al. (2018). Knowledge-based collaborative deep learning for benign-malignant lung nodule classification on chest CT. IEEE Transactions on Medical Imaging, 38(4), 991–1004.
Zhao, C., Han, J., Jia, Y., & Gou, F. (2018) Lung nodule detection via 3D U-Net and contextual convolutional neural network. In 2018 International conference on networking and network applications (NaNA) (pp. 356–361).
Shen, W., Zhou, M., Yang, F., Yu, D., Dong, D., Yang, C., et al. (2017). Multi-crop convolutional neural networks for lung nodule malignancy suspiciousness classification. Pattern Recognition, 61, 663–673.
Zhao, X., Liu, L., Qi, S., Teng, Y., Li, J., & Qian, W. (2018). Agile convolutional neural network for pulmonary nodule classification using CT images. International Journal of Computer Assisted Radiology and Surgery, 13(4), 585–595.
Wang, H., Zhou, Z., Li, Y., Chen, Z., Lu, P., Wang, W., et al. (2017). Comparison of machine learning methods for classifying mediastinal lymph node metastasis of non-small cell lung cancer from 18F-FDG PET/CT images. EJNMMI Research, 7, 11.
Wang, Z., Xu, H., & Sun, M. (2017). Deep learning based nodule detection from pulmonary CT images. In 2017 10th International symposium on computational intelligence and design (ISCID), Hangzhou (pp. 370–373).
Setio, A.A.A. et al. (2016). Validation, comparison, and combination of algorithms for automatic detection of pulmonary nodules in computed tomography images: The LUNA16 challenge. arXiv.org cs.CV.
Chon, A., & Balachandar, N. (2017). Deep convolutional neural networks for lung cancer detection, Stanford University.
Kuan, K., Ravaut, M., Manek, G., & Chen, H. (2017). Deep learning for lung cancer detection: Tackling the Kaggle data science bowl 2017 challenge. Retrieved from arXiv:1705.09435.
Shen, W., Zhou, M., Yang, F., Dong, D., Yang, C., Zang, Y., & Tian, J. (2016). Learning from experts: Developing transferable deep features for patient-level lung cancer prediction. In International conference on medical image computing and computer-assisted intervention. (pp. 124–131). Springer.
Bush, I.F. (2016). Lung nodule detection and classification. Technical report, Stanford Computer Science.
Jung, H., Kim, B., Lee, I., Lee, J., & Kang, J. (2018). Classification of lung nodules in CT scans using three-dimensional deep convolutionalneural networks with a checkpoint ensemble method. BMC Medical Imaging, 18, 48.
Dey, R., Lu, Z., & Hong, Y. (2018). Diagnostic classification of lung nodules using 3D neural networks. In 2018 IEEE 15th international symposium on biomedical imaging (ISBI 2018).
Khosravan, N., & Bagci, U. (2018). S4ND: Single-shot single-scale lung nodule detection. arxiv preprint arXiv:1805.02279.
Shaziya, H., Shyamala, K., & Zaheer, R. (2018). Automatic lung segmentation on thoracic CT scans using U-net convolutional network. In 2018 International conference on communication and signal processing (ICCSP). IEEE.
Dai, C., Xiao, B., Chen, Y., Du, Y., Liang, Y., Zhao, K., & Yan, L. (2018). Automated detection of lung nodules in CT images with 3D convolutional neural networks. In 2018 International conference on network infrastructure and digital content (IC-NIDC). Biology Society (EMBC).
Ciompi, F., de Hoop, B., van Riel, S. J., Chung, K., Scholten, E., Oudkerk, M., et al. (2015). Automatic classification of pulmonary peri-fissural nodules in computed tomography using an ensemble of 2D views and a convolutional neural network out-of-the-box. Medical Image Analysis, 26(1), 195–202.
Van, G., Bram, S., Arnaud, A.A., Jacobs, C., & Ciompi, F. (2015). Off-the-shelf convolutional neural network features for pulmonary nodule detection in computed tomography scans. In SPIE medical imaging (pp. 286–289).
Ypsilantis, P.P., & Montana, G. (2016). Recurrent convolutional networks for pulmonary nodule detection in CT imaging. arXiv.org stat.ML.
Oliveira, D.A.B., & Viana, M.P. (2018). An efficient multi-scale data representation method for lung nodule false positive reduction using convolutional neural networks. In 2018 IEEE 15th international symposium on biomedical imaging (ISBI 2018).
Causey, J. L., Zhang, J., Ma, S., Jiang, B., Qualls, J. A., Politte, D. G., et al. (2018). Highly accurate model for prediction of lung nodule malignancy with CT scans. Scientific Reports, 8, 9286.
Pham, T.D. (2017). Complementary features for radiomic analysis of malignant and benign mediastinal lymph nodes. In 2017 IEEE international conference on image processing (ICIP).
Chen, J., & Shen, Y. (2017). The effect of kernel size of CNNs for lung nodule classification. In 2017 9th International conference on advanced Infocomm technology (ICAIT), Chengdu (pp 340–344).
Yan, X., Pang, J., Qi, H., Zhu, Y., Bai, C., Geng, X., Liu, M., Terzopoulos, D., & Ding, X. (2016). Classification of lung nodule malignancy risk on computed tomography images using convolutional neural network: A comparison between 2D and 3D strategies. In Chen, C.-S., Lu, J., & Ma, K.-K. (Eds.), LNCS 10118 (pp. 91–101).
Xie, H., Yang, D., Sun, N., Chen, Z., & Zhang, Y. (2019). Automated pulmonary nodule detection in CT images using deep convolutional neural networks. Pattern Recognition, 85, 109–119.
Li, W., Cao, P., Zhao, D., & Wang, J. (2016). Pulmonary nodule classification with deep convolutional neural networks on computed tomography images. Computational and Mathematical Methods in Medicine, 2016, 1–7.
Nibali, A., He, Z., & Wollersheim, D. (2017). Pulmonary nodule classification with deep residual networks. International Journal of Computer Assisted Radiology and Surgery, 12, 1799–1808. https://doi.org/10.1007/s11548-017-1605-6
Song, Q., Zhao, L., XingKe, L., & XueChen, D. (2017). Using deep learning for classification of lung nodules on computed tomography images. Journal of Healthcare Engineering, 1, 1–7.
da Silva, G.L.F., Silva, A.C., de Paiva, A.C., & Gattass, M. (2016). Classification of malignancy of lung nodules in CT images using convolutional neural network. In XVI workshop on medical informatics—2016—Porto Alegre, RS (pp. 2481–2489).
Patrice, M., Qi, S., Xu, M., Li, H., Han, F., Teng, Y., et al. (2018). Ensemble learning of multiple-view 3D-CNNs model for micro-nodules identification in CT images. IEEE Access., PP, 1. https://doi.org/10.1109/access.2018.2889350.
Shen, S., Han, S. X., Aberle, D. R., Bui, A. A., & Hsu, W. (2019). An interpretable deep hierarchical semantic convolutional neural network for lung nodule malignancy classification. Expert Systems with Applications. https://doi.org/10.1016/j.eswa.2019.01.048.
Murphy, K., van Ginneken, B., Schilham, A. M. R., de Hoop, B. J., Gietema, H. A., & Prokop, M. (2009). A large scale evaluation of automatic pulmonary nodule detection in chest CT using local image features and k-nearest-neighbour classification. Medical Image Analysis, 13, 757–770. https://doi.org/10.1016/j.media.2009.07.001.a.
Li, X., Kao, Y., Shen, W., Li, X., Xie, G., Lung nodule malignancy prediction using multi-task convolutional neural network. In Proceedings of SPIE 10134, medical imaging 2017: Computer-aided diagnosis (pp. 1013424). https://doi.org/10.1117/12.2253836.
Khosravan, N., & Bagci, U. (2018). Semi-supervised multi-task learning for lung cancer diagnosis. In 2018 40th Annual international conference of the IEEE engineering in medicine and biology society (EMBC).
Winkels, M., & Cohen, T. S. (2019). Pulmonary nodule detection in CT scans with equivariant CNNs. Medical Image Analysis, Medical Image Analysis, 55, 15–26.
da Nóbrega, R.V.M., Peixoto, S.A., da Silva, S.P.P., & Rebouças Filho, P.P. (2018). Lung nodule classification via deep transfer learning in CT lung images. In 2018 IEEE 31st international symposium on computer-based medical systems (CBMS).
Hua, K. L., Hsu, C. H., Hidayati, S. C., Cheng, W. H., & Chen, Y. J. (2015). Computer-aided classification of lung nodules on computed tomography images via deep learning technique. OncoTargets and Therapy, 8, 2015–2022. https://doi.org/10.2147/ott.s80733.
Tu, X., et al. (2017). Automatic categorization and scoring of solid, part-solid and non-solid pulmonary nodules in CT images with convolutional neural network. Scientific Reports, 7, 8533.
Wen, Y., Zhang, K., Li, Z., & Qiao, Y. (2016) A discriminative feature learning approach for deep face recognition. In European conference on computer vision (pp. 499–515). Springer.
He, K., Zhang, X., Ren, S., & Sun, J. (2015). Delving deep into rectifiers: Surpassing human-level performance on imagenet classification. In Proceedings of the IEEE international conference on computer vision (pp. 1026–1034). doi: 10.1.1.725.4861.
Zagoruyko, S., & Komodakis, N. (2016). Wide residual networks. arxiv preprint arXiv:1605.07146.
Paul, R., et al. (2016). Deep feature transfer learning in combination with traditional features predicts survival among patients with lung adenocarcinoma. Tomography, 2(4), 388–395.
Hussein, S., Cao, K., Song, Q., & Bagci, U. (2017). Risk stratification of lung nodules using 3d cnn-based multi-task learning. In IPMI.
Liu, Z., Yao, C., Yu, H. et al. (2019). Deep reinforcement learning with its application for lung cancer detection in medical internet of things. Future Generation Computer Systems.
Zhang, M., Li, H., Lyu, J., Ling, S.H., Su, S. (2019). Multi-level CNN for lung nodule classification with Gaussian Process assisted hyperparameter optimization. arXiv:1901.00276.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Naik, A., Edla, D.R. Lung Nodule Classification on Computed Tomography Images Using Deep Learning. Wireless Pers Commun 116, 655–690 (2021). https://doi.org/10.1007/s11277-020-07732-1
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11277-020-07732-1