Abstract
The aerobic cometabolism of chlorobenzoic acids (CBAs) by Rhodococcus sp. R04 was accomplished by augmenting the medium with organic carbon sources. In mineral medium supplemented with glucose (MMG), 0.5 mM 2-CBA was incompletely metabolized after the 5-day incubation, while the near-complete disappearance of 0.5 mM 4-CBA was monitored. Over the 5-day incubation period, the concentration of chloride increased to 0.17 mM in bottles containing 4-CBA, glucose and strain R04; whereas in cultivation with 2-CBA the chloride content was about 0.1 mM. After 5-day incubation, 28.5% 4-CBA was remained in mineral medium supplemented with ethanol (MME), and the relatively low values of chloride were released. To our knowledge, it is first report that the feasibility of using ethanol as an added substrate for cometabolic degradation of CBA by aerobic polychlorinated biphenyl (PCB)-degrading bacteria. The specific activities of (chloro)benzoate 1,2-dioxygenase and (chloro)catechol 1,2-dioxygenase activities were detected in cell-free extracts (CFEs) of strain R04. These results suggest that the initial degradation of CBAs occurred most likely prior to chloride release.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
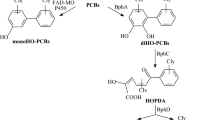
Chlorobenzoic acids (CBAs) are released into environment as partial degradation products of other xenobiotic compounds such as PCBs and benzoate herbicides. Due to their persistence in the environment, considerable problems arise from these compounds with a haloaromatic ring as a structural feature. Bacterial degradation of CBAs has been studies extensively (Pieper 2005). The majority of organisms are able to degrade CBAs via the formation of (chloro)catechol as central intermediate formed by the dioxygenation of CBAs. (Chloro)catechol may further be mineralized through the modified ortho-pathway, which involves the enzyme (chloro)catechol-1,2-dioxygenase, and dechlorination occurs after ring cleavage. Alternatively, mineralization of chloroaromatics via a meta-cleavage pathway with an unusual (chloro)catechol-2,3-dioxygenase have been reported (Mars et al. 1997; Kaschabek et al. 1998; Ajithkumar and Kunhi 2000).
Chlorobenzoic acids (CBAs) generally are transformed slowly by aerobic PCB-degrading bacteria, degradation of which appeared to be the rate limiting step in the overall PCB-degradation process (Adriaens and Focht 1991a), although there was reports that bacteria in principal are able to use chloroaromatics as growth substrates with total elimination of chloride (Kim and Picardal 2001). Due to their toxicity, bacterial growth on halogenated aromatics as sole energy and carbon source is often suboptimal, showing long acclimation lag and slow growth rates. It is quite common that an organic compound is chosen as a growth substrate. Several examples of cometabolism have been reported when the medium contains an alternate carbon source which can serves as sources of carbon and energy to support cell growth and induce the metabolic pathway (Kohler et al. 1988; Adriaens and Focht 1991b; Corbella et al. 2001). Necessary co-substrates NADH or NADPH used in many aerobic cometabolic reactions may be regenerated when glucose is used for cometabolic degradation (Wang and Loh 1999). Especially when the selected growth substrate is a conventional carbon source, the design of cometabolic systems may aid in reducing the toxicity and growth inhibition of xenobiotics, thereby increasing the transformation rate of xenobiotics.
The objective of the work is to study the influence of supplementary carbon sources on improving the biotransformation rates of mono-chloro benzoic acids by a PCB-degrading strain, Rhodococcus sp. R04 (Sun and Qian 2002; Yang et al. 2004). Since glucose is known to facilitate degradation of some chlorinated pollutants (Loh and Wang 1998; Wang and Loh 1999; Corbella et al. 2001) it was tested as the selected carbon source. Ethanol was also tested because of its potential use as PCB solvent in laboratory studies. Meanwhile the expression of the (chloro)benzoate dioxygenase and the induction of (chloro)catechol 1,2-dioxygenase activities were investigated.
Materials and methods
Chemicals
CBAs (98% purity), catechol (98% purity) and 4-chlorocatechol (97% purity) were obtained from Sigma–Aldrich chemical Co. (USA); Biphenyl was purchased from the Academy of Chinese Army Medicine, all other chemicals used in this study were analytical Reagent grade and were procured from standard companies.
Bacterial strain, media
Rhodococcus sp. R04 used in this study was isolated from a soil sample collected from an oil field in Northern China with biphenyl as the sole carbon and energy source, and preserved in our laboratory. The organism was routinely grown at 30°C in a basal MM (mineral medium, pH 7.0) (Yang et al. 2004).
Studies with resting cell
Resting cells were prepared as described previously (Yang et al. 2004). Cell suspensions of Rhodococcus sp. R04 were incubated aerobically on MM, MMG (11.2 g l−1), or MME (1/100 v/v). Each CBA was provided at a final concentration of 0.5 mM. Controls were incubated in corresponding mediums without CBAs. Cells were incubated at 30°C at 250 rev/min in a gyratory shaker.
Cell growth and dechlorination assay
Biomass concentrations were determined spectrophotometrically by measuring the optical density of cell cultures at 600 nm and subsequent calculation of the cell dry weight by means of a calibration curve. Inorganic chloride was estimated turbidimetrically by measuring AgCl precipitation at 525 nm (Hickey and Focht 1990). Chloride was quantified by reference to a standard curve that was linear from 0.1 to 1 mM. All the experiments were conducted in triplicate.
Analytical procedures
The culture supernatant with residual chlorobenzoates after removing the cells by centrifugation was acidified with HCl to pH 3.0, then detected by HPLC. The analysis was carried out using C18 reverse-phase column (5 µm, 3.90 mm × 150 mm). Aqueous solvent containing 1 ml of ortho-phosphoric acid and 600 ml of methanol per liter (Rodrigues et al. 2001) was used as mobile phase with a flow rate of 1 ml/min. Mono-chloro benzoic acids were monitored in the eluate at 230 nm and compared to the peaks obtained with standard solutions. Polar metabolites were also analyzed by HPLC with the same C18 reverse-phase column at 254 nm. A mobile phase of 45% acetonitrile and 55% acetic acid (40 mM) was used (Arensdorf and Focht 1994).
Assay of enzymes
Cells grown to mid exponential phase were harvested by centrifugation (10 min, 8000×g, 4°C), and washed twice with 0.05 M sodium phosphate buffer (pH 7.5), then resuspended in the same buffer or in 45 mM Tris–HCl (pH 7.5) containing 1 mM dithiothreitol. For making cell-free extracts (CFEs), the cells were disrupted by ultrasonication at 40 W for 13 min with intermittent bursts of 3 s. The cell debris was removed by centrifugation at 12000×g for 20 min. The clear supernatant solution was used as a source of crude enzymes. All the assays with the crude enzymes were carried out immediately at 25°C.
(Chloro)benzoate 1,2-dioxygenase activity was assayed spectrophotometrically by monitoring the decrease of NADH at 340 nm (Romanov and Hausinger 1994). The reactions were started by addition of the corresponding (chloro)benzoate to a final concentration of 0.7 mM. (Chloro)catechol 1,2-dioxygenase was measured according to the method described by Dorn and Knackmuss (1978). (Chloro)catechol 2,3- dioxygenase was assayed at 375 nm by determining formed reaction product, 2-hydroxymuconic semialdehyde or its chlorinated derivative (Nozaki 1970).
Protein estimation
The protein content of extracts was estimated by the method of Lowry et al. (1951) using bovine serum albumin as the standard.
Results
In this study, the ability of strain Rhodococcus sp. R04 to metabolize mono-chloro benzoic acids grown on MM medium was tested. Neither disappearance of mono-chloro benzoic acids nor release of chloride could be detected in the cultures exposed to CBAs during the 5-day incubation period (data not shown), suggesting that biodegradation of 2-CBA or 4-CBA by resting cells of Rhodococcus sp. R04 was failure in these conditions.
Degradation of chlorobenzoic acids in ethanol-supplemented media
In order to study the effect of ethanol on the degradation of 2-CBA and 4-CBA, resting cells of strain R04 was grown on MM medium supplemented with 1% ethanol and exposed to chlorobenzoic acids. The resting cells transform into growing cells after cultivation in the presence of ethanol (Fig. 1). Growth on ethanol as supplemented carbon source was very efficient: inoculated cultures took, on the average, 5 days to reach 3.7 mg/ml. The growth curves on 4-CBA were different to that of control in ethanol-supplemented media, biomass on the control was sharply increased after 24 h incubation, but biomass on 4-CBA did not nearly increase within the same time, after the lag phase, the cells grew exponentially (Fig. 1). 28.5% 4-CBA were remained in the culture supernatants after 5 days incubation, and the relatively low values of chloride released in 4-CBA grown cultures were monitored (Fig. 2). There was lag period between 4-CBA disappearance and chloride ions release. However, 2-CBA was shown not to be transformed in these conditions and the concentration of chloride was almost same as the control. The cells exposed to 2-CBA and ethanol grew exponentially without lag phase. This indicates that 2-CBA had no apparent effects on the early growth of strain R04.
Chloride release kinetics and substrate disappearance in MME cultures without CBAs (control) or exposed to 2-CBA and 4-CBA (0.5 mM). Symbols: ∆, remaining 2-CBA; ■, remaining 4-CBA; □, chloride ion production in bottles with 4-CBA and strain R04; ▲, chloride ion production in control bottles and strain R04
Degradation of chlorobenzoic acids in glucose-supplemented media
In order to study the feasibility of using glucose as a growth substrate to cometabolize chlorobenzoic acids, resting cells of strain R04 were grown on glucose and exposed to chlorobenzoic acids. Interestingly, the resting cells also transform into growing cells after cultivation in the presence of glucose (Fig. 3). The growth curves of strain R04 on chlorobenzoic acids were similar to that of the control, biomass achieved by 2-CBA or 4-CBA supplied medium was slightly lower than that of the control during the 5-day incubation period (Fig. 3). Figure 4 shows that after the 5-day incubation, 2-CBA was incompletely metabolized; the near-complete disappearance of 0.5 mM 4-CBA was monitored by HPLC. As biomass increased over the 5-day incubation period, the concentration of chloride increased to 0.17 mM in MMG containing 4-CBA, whereas in cultures with 2-CBA the chloride content was about 0.1 mM (Fig. 4). There was also lag period between CBAs disappearance and chloride ions release.
Chloride release kinetics and substrate disappearance in MMG cultures without CBAs (control) or exposed to 2-CBA and 4-CBA (0.5 mM). Symbols: ■, remaining 4-CBA; ◊, remaining 2-CBA; □, chloride ion production in bottles with 4-CBA and strain R04; ▲, chloride ion production in bottles with 2-CBA and R04; ×, chloride ion production in control bottles and strain R04
Enzyme activities
Table 1 shows the levels of (chloro)benzoate 1,2-dioxygenase activity and (chloro)catechol 1,2-dioxygenase activity as monitored by a spectrophotometric assay in CFEs prepared from 2-CBA, 4-CBA, benzoate-grown cells or control. Compared to cultures grown on MME, higher activities of (chloro)catechol 1,2-dioxygenase were recorded in glucose-fed cells. The high activities of (chloro)catechol 1,2-dioxygenase found in MMG agree with the observed ability for 4-CBA degradation. It is important to note that the ratio of (chloro)catechol 1,2-dioxygenase activity for catechol versus 4-chlorocatechol varies for extracts from cells grown with different carbon sources (e.g., the ratios are from 3.2 to 5.3 for extracts from cells grown on ethanol or glucose exposed to CBAs, but on benzoate the ratios are 16.7). These results are consistent with the presence of at least two distinct enzymes (possibly with overlapping specificities) that appear to be regulated differentially (Romanov and Hausinger 1994). The strain R04, like some chlorocatechol-degrading bacteria described in the literature (Qi et al. 2007) seems to have two distinct pyrocatechases (a catechol 1,2-dioxygenase, capable of catabolizing catechol, and a chlorocatechol 1,2-dioxygenase, which has relaxed substrate specificity and is thus active with catechol and substituted catechols (Dorn and Knackmuss 1978). (Chloro)catechol 2,3-dioxygenase activity was not detected in CFEs obtained from R04 cells.
Discussion
CBAs could be produced by the biphenyl pathway of Rhodococcus sp. R04 during degradation of diverse chlorobiphenyls (Yang et al. 2004, 2007). 3-CBA and 4-CBA were, respectively monitored by GC-MS in cultures from strain R04 grown on 3-chlorobiphenyl and 4-chlorobiphenyl (unpublished data). In this study, biodegradation of 2-CBA or 4-CBA by strain R04 grown on MM medium was failure, although 3-CBA was shown to be transformed by resting cells (unpublished data). The inability to mineralize these compounds may be related to the lack of induction of key metabolic reactions, the appearance of non-metabolizable intermediates, or the lack of suitable systems for uptake (Corbella et al. 2001). Our research focus on the degradation and dechlorination of 2-CBA and 4-CBA by aerobic PCB-degrading bacteria R04 grown on MM medium supplemented with organic carbon sources. Nies and Vogel observed that several organic substrates including methanol, glucose, acetone and acetate appear to accelerate dechlorination of Arclor 1242 in anaerobic sediments (Nies and Vogel 1990). To our knowledge, it is first report that the feasibility of using ethanol as an added substrate for cometabolic degradation of CBA by aerobic PCB-degrading bacteria (Fig. 2), albeit degradation of 2-CBA was not been detected in same conditions.
The effects of CBAs on growth were analyzed. To some extent, the presence of 4-CBA and 2-CBA inhibited the growth of strain R04 using glucose as a growth substrate. Martinez et al. (2007) also observed chlorobenzoate inhibited growth in the PCB-degrading bacterium Burkholderia xenovorans LB400. 4-CBA exhibited a stronger negative effect on ethanol supplemented medium than that on glucose supplemented medium during early 24 h incubation. This indicates the time for physiological changes in response to exposure to a new environment, in which metabolic system of cells can be affected. It may be suitable to use glucose as a carbon source to mineralize CBAs by Rhodococcus sp. R04.
Because strain R04 can degrade chlorobenzoic acids grown on MM supplemented with ethanol or glucose, we want to understand the pathways for chlorobenzoate biodegradation. Table 1 shows specific enzymes activities in cell-free extracts of Rhodococcus sp. R04 grown on different substrates. (Chloro)catechol 2,3-dioxygenase activity, which was responded for extradiol cleavage of the meta-pathway in microorganisms, was not detected in CFEs obtained from R04 cells. However, the specific activities of (chloro)benzoate 1,2-dioxygenase and (chloro)catechol 1,2-dioxygenase activities were present in CFEs of Rhodococcus sp. R04, and Figs. 2 and 4 shows the lag period between chlorobenzoic acids disappearance and chloride ions release, suggesting that the initial degradation occurred most likely prior to chloride release. Although looking for other metabolites such as catechol or chlorocatechol by HPLC in the full grown cultures supernatants was unsuccessful (data not shown), these results obtained give insight in the induction of the pathways for the metabolism of chorobenzoates. Strain R04 may degrade the chlorobenzoic acids through the formation of chlorobenzoate dihydrodiol catalysed by a benzoate dioxygenase by inserting dioxygen to the aromatic ring. Chlorobenzoate dihydrodiol was then converted to chlorocatechol. chlorocatechol as intermediate, may be degraded via the modified ortho-pathway, and dechlorination occurs after ring cleavage.
Strain R04 was able to transform and dehalogenate 2-CBA in MM culture supplemented with glucose, but not to transform and dehalogenate 2-CBA in MM culture supplemented with 1% ethanol. Moreover degradation of 4-CBA seemed to be more efficient on glucose supplemented media than that on ethanol supplemented media. The results obtained infer that glucose could be a good carbon source to cometabolize CBAs. Moreover, the use of glucose would not result in additional environmental pollution. Therefore, although these results hold promise for inducing CBAs degradation and dechlorination, understanding of other useful factors is required.
Conclusions
We investigated the influence of supplementary carbon sources (ethanol and glucose) on improving the biotransformation rates of CBAs by strain R04. To our knowledge, it is first report that the feasibility of using ethanol as an added substrate for cometabolic degradation of CBA by aerobic PCB-degrading bacteria.
Based on the chloride released rate and enzymes activities, a pathway for degradation of chlorobenzoic acids by Strain R04 was proposed: degradation of chlorobenzoates occurs via the formation of chlorocatechols as central intermediates which are further mineralized through a modified ortho-pathway.
References
Adriaens P, Focht DD (1991a) Cometabolism of 3,4-dichlorobenzoate by Acinetobacter sp. strain 4-CB1. Appl Environ Microbiol 57:173–179
Adriaens P, Focht DD (1991b) Continuous co-culture degradation of selected polychlorinated biphenyl congeners by Acinetobacter sp. in an aerobic reactor system. Environ Sci Technol 24:1042–1049. doi:10.1021/es00077a015
Ajithkumar PV, Kunhi AA (2000) Pathways for 3-chloro- and 4-chlorobenzoate degradation in Pseudomonas aeruginosa 3mT. Biodegradation 11:247–261. doi:10.1023/A:1011124220003
Arensdorf JJ, Focht DD (1994) Formation of chlorocatechol meta-cleavage products by a pseudomonad during metabolism of monochlorobiphenyls. Appl Environ Microbiol 60:2884–2889
Corbella ME, Garrido-Pertierra A, Puyet A (2001) Induction of the halobenzoate catabolic pathway and cometabolism of ortho-chlorobenzoates in Pseudomonas aeruginosa 142 grown on glucose-supplemented media. Biodegradation 12:149–157. doi:10.1023/A:1013117805732
Dorn E, Knackmuss HJ (1978) Chemical structure and biodegradability of halogenated aromatic compounds. Two catechol 1,2-dioxygenases from a 3-chlorobenzoate-grown pseudomonad. Biochem J 174:73–84
Hickey WJ, Focht DD (1990) Degradation of mono-, di-, and trihalogenated benzoic acids by Pseudomonas aeruginosa JB2. Appl Environ Microbiol 56:3842–3850
Kaschabek SR, Kasberg T, Muller D, Mars AE, Janssen DB, Reineke W (1998) Degradation of chloroaromatics: purification and characterization of a novel type of chlorocatechol 2, 3-dioxygenase of Pseudomonas putida GJ31. J Bacteriol 180:296–302
Kim S, Picardal FW (2001) Microbial growth on dichlorobiphenyls chlorinated on both rings as a sole carbon and energy source. Appl Environ Microbiol 67:1953–1955. doi:10.1128/AEM.67.4.1953-1955.2001
Kohler HP, Kohler-Staub D, Focht DD (1988) Cometabolism of polychlorinated biphenyls: enhanced transformation of Aroclor 1254 by growing bacterial cells. Appl Environ Microbiol 54:1940–1945
Loh KC, Wang SJ (1998) Enhancement of biodegradation of phenol and a nongrowth substrate 4-chlorophenol by medium augmentation with conventional carbon sources. Biodegradation 8:329–338. doi:10.1023/A:1008267607634
Lowry OH, Rosebrough NJ, Farr AL, Randall RJ (1951) Protein measurement with the Folin phenol reagent. J Biol Chem 193:265–275
Mars AE, Kasberg T, Kaschabek SR, van Agteren MH, Janssen DB, Reineke W (1997) Microbial degradation of chloroaromatics: use of the meta-cleavage pathway for mineralization of chlorobenzene. J Bacteriol 179:4530–4537
Martinez P, Agullo L, Hernandez M, Seeger M (2007) Chlorobenzoate inhibits growth and induces stress proteins in the PCB-degrading bacterium Burkholderia xenovorans LB400. Arch Microbiol 188:289–297. doi:10.1007/s00203-007-0247-4
Nies L, Vogel TM (1990) Effects of organic substrates on dechlorination of Aroclor 1242 in anaerobic sediments. Appl Environ Microbiol 56:2612–2617
Nozaki M (1970) Metapyrocatechase (Pseudomonad). Methods Enzymol 17A:522–525. doi:10.1016/0076-6879(71)17235-7
Pieper DH (2005) Aerobic degradation of polychlorinated biphenyls. Appl Microbiol Biotechnol 67:170–191. doi:10.1007/s00253-004-1810-4
Qi Y, Zhao L, Ojekunle ZO, Tan X (2007) Isolation and preliminary characterization of a 3-chlorobenzoate degrading bacteria. J Environ Sci (China) 19:332–337. doi:10.1016/S1001-0742(07)60054-0
Rodrigues JL, Maltseva OV, Tsoi TV, Helton RR, Quensen JF 3rd, Fukuda M, Tiedje JM (2001) Development of a Rhodococcus recombinant strain for degradation of products from anaerobic dechlorination of PCBs. Environ Sci Technol 35:663–668. doi:10.1021/es001308t
Romanov V, Hausinger RP (1994) Pseudomonas aeruginosa 142 uses a three-component ortho-halobenzoate 1,2-dioxygenase for metabolism of 2, 4-dichloro- and 2-chlorobenzoate. J Bacteriol 176:3368–3374
Sun Y, Qian S (2002) Flux control analysis for biphenyl metabolism by Rhodococcus pyridinovorans R04. Biotechnol Lett 24:1525–1529. doi:10.1023/A:1019843620061
Wang SJ, Loh KC (1999) Facilitation of cometabolic degradation of 4-chlorophenol using glucose as an added growth substrate. Biodegradation 10:261–269. doi:10.1023/A:1008347630546
Yang X, Sun Y, Qian S (2004) Biodegradation of seven polychlorinated biphenyls by a newly isolated aerobic bacterium (Rhodococcus sp. R04). J Ind Microbiol Biotechnol 31:415–420. doi:10.1007/s10295-004-0162-5
Yang X, Liu X, Song L, Xie F, Zhang G, Qian S (2007) Characterization and functional analysis of a novel gene cluster involved in biphenyl degradation in Rhodococcus sp. strain R04. J Appl Microbiol 103:2214–2224. doi:10.1111/j.1365-2672.2007.03461.x
Acknowledgments
This work was supported by the National Natural Science Foundation of China (30570017). We thank J.J. Shi for advice with HPLC separations, Dr. Y.F. Ma for dechlorination assay.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zhang, G., Yang, X., Xie, F. et al. Cometabolic degradation of mono-chloro benzoic acids by Rhodococcus sp. R04 grown on organic carbon sources. World J Microbiol Biotechnol 25, 1169–1174 (2009). https://doi.org/10.1007/s11274-009-9996-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11274-009-9996-3