Abstract
Seven aerobic bacterial strains were isolated from pulp paper mill waste and screened for pentachlorophenol (PCP) tolerance on PCP containing mineral salt agar medium (MSM). The organism was characterized by 16S rDNA sequencing which showed 99.7% sequence similarity with Serratia marcescens. PCP degradation was routinely monitored with spectrophotometric analysis and further confirmed by HPLC analysis. Among seven strains, ITRC S7 was found to degrade up to 90.33% of 1.127 mM (300 mg/l) of PCP and simultaneous release of chloride ion (2.435 mM) emphasized the bacterial dechlorination in the medium in presence of glucose as an additional carbon and energy source under optimized condition within 168 h incubation. In absence of glucose bacterium was unable to utilize PCP indicating the phenomenon of co-metabolism. Bacterium was identified as S. marcescens (AY927692), was a novel and potential aerobic bacterial strain capable of degrading PCP in axenic condition. Further, this strain may be used for bioremediation of PCP containing pulp paper mill waste in the environment.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Pentachlorophenol (PCP) is a major environmental pollutant discharged from pulp and paper mills, tanneries, distilleries, dye and paint manufacturing and pharmaceutical industries (Crosby 1981). Among chlorophenols (CPs), PCP is a wide spectrum biocide with applications in agriculture, industry and public health. The US Environmental Protection Agency has registered PCP in the list of priority pollutants. They have had now restricted the sale and use of pesticide products containing PCP (EPA 1987).
PCP from the pulp bleaching process is found both in free (hexane extractable) and bound (extractable with strong alkali) forms in dissolved organic matter and particles (Schnell et al. 2000). High and low molecular weight chlorinated compounds are produced by complex reactions between chlorine and lignin in the wood pulp (Yeber et al. 2000). Bleached chemical pulp mill effluents have been identified as toxic according to the Canadian Environmental Protection Act due to the presence of high quantity of CPs (Schnell et al. 2000). In addition, PCP is very harmful to microorganisms because it destroys membrane function (Copley 2000) due to its ability to uncouple oxidative phosphorylation (Ito and Ohnishi 1982). It has been reported as mutagenic even at low level of concentration. It has inherent toxicity to be bioaccumulated in various food chains of biological system. PCP is toxic to living organisms and has a high half-life in environment because of the lack of biodegradative enzymes in the indigenous organisms (Bellinaso et al. 2001).
Different researchers have reported PCP degrading microorganisms isolated from the natural environment. Saber and Crawford (1985) reported bacteria of the genus Flavobacterium utilizing 100 mg/l of PCP as sole source of carbon and energy. Schenk (1989) found the Arthrobacter sp. ATCC 33790 capable of utilizing PCP. Shah and Thakur (2002) have also observed that a Pseudomonas fluorescens could degrade PCP up to 100 mg/l in 120 h. However, the reported concentration of PCP degradation is less than that of PCP discharged by pulp paper industry. Therefore, there is need to isolate more potent bacterial strain which can degrade higher concentrations of PCP. There could still be a number of unidentified bacterial strains having capability of PCP degradation at higher concentrations. Therefore, this study is aimed to isolate and characterize more potent aerobic bacterial strains, which have the capacity to degrade PCP at higher concentrations than reported earlier.
Materials and methods
Chemicals
All reagents used were of analytical grade. Synthetic PCP (FW 266.30) was purchased from Sigma chemicals (USA). All solutions were prepared in Milli-Q water (Elix, Millipore purification system, France).
Sample collection
Pulp paper mill sludge samples contain significant amount of PCP (50.31 mg/l) (Raj et al. 2005) were collected from M/s Century pulp and paper mill, Lalkuan, Nainital, Uttaranchal (India) located at foothills of Himalayas (79° 10′ E and 29° 3′ N). The samples were taken into pre-sterilized test tube and immediately preserved at 4 °C.
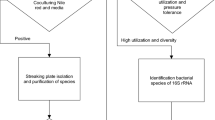
Isolation, purification and screening of PCP degrading bacteria
Screening of PCP tolerant bacteria were done using nutrient enrichment technique by adding one gram pulp paper mill effluent sludge containing (50.31 mg/l of PCP) in mineral salt medium (MSM) of following composition (mg/l): K2HPO4, 85; KH2PO4, 17; MgSO4, 30; FeSO4·7H2O, 30; CaSO4, 30; MnSO4·H2O, 30; (NH4)2 SO4, 17 and purified agar, 18 g/l (pH 7.0 ± 0.2), trace element solution, 1 ml/l amended with different concentrations of PCP (0.376, 0.752, 1.127 and 1.503 mM) along with glucose 1% (w/v) as additional carbon source (Pfennig and Lippert 1966). The flask containing culture was incubated at 30 ± 1 °C and 120 rev/min for 5 days. Thereafter, the bacterial suspension was plated out on to MSM agar with and without glucose. Seven phenotypically different bacterial colonies were picked and purified by repeated streaking on the same medium. These strains were designated as ITRC S1, ITRC S2, ITRC S3, ITRC S4, ITRC S5, ITRC S6 and ITRC S7. The Gram staining of aforesaid strains was done as per standard protocol following Williams et al. (1993). The confirmatory test for the tolerance and accumulation of PCP by these bacteria were performed using bromocresol purple (BCP-16 mg/l) containing MSM (Martins et al. 1997) and control were used without inoculum. The most potent strain capable of degrading PCP was used for further studies and was identified by 16S rDNA sequencing.
Isolation of genomic DNA and amplification of 16S rDNA
Genomic DNA was isolated following the method described by Ausubel et al. (1992) with some minor modifications.
16S rDNA was polymerase chain reaction (PCR) amplified from the genomic DNA template with the help of universal primers 27f (5′-AGAGTTTGATCMTGGCTCAG-3′) and 1492r (5′-TACGGYTACCTTGTTACGACTT-3′) (Escherichia coli 16S rDNA numbering system (Brosius et al. 1978; Weisburg et al. 1991) using a personal thermocycler (Eppendorf, Hamburg, Germany). Reaction mixture contained 70–100 ng of genomic DNA, 1 U of Deep Vent DNA polymerase, 1× Thermopol reaction buffer (10 mM KCl, 20 mM Tris-HCl [pH 8.8]), 10 mM (NH4)2SO4, 2 mM MgSO4 and 0.1% Triton X-100), 200 (μM of each deoxynucleoside triphosphate (New England Biolabs, Beverly, USA) and 20 pmol of each primer (Biobasics, Canada). PCR cycling parameters included an initial denaturation at 95 °C for 5 min, followed by 30 cycles of denaturation at 95 °C for 1 min, annealing at 55 °C for 1 min and extension at 75 °C for 2 min and a final extension for 10 min at 75 °C.
Sequencing of 16S rDNA and phylogenetic analysis
Approximately 1.5 kb amplicon was separated by gel electrophoresis, eluted by Qiaquick gel extraction kit (Qiagen) and was sequenced using the following four forward and one reverse primers, viz. 27f, 357f, 704f, 926f and 1492r (Johnson 1994). The 16S rDNA sequence was determined following the dideoxy chain-termination method using ABI Prism BigDye Terminator Cycle Sequencing Ready Reaction Kit as directed in the manufacturer’s protocol. Sequence reactions were electrophoresed and analysed by ABI 310 Genetic Analyzer (Applied Biosystems, USA). Sequences of closely related taxa of the isolate were retrieved from the GenBank database using BLAST (www.ncbi.nlm.nih.gov/BLAST) and Ribosomal Database Project II (http://rdp.cme.msu.edu/index.jsp) (Altschul et al. 1997; Larsen et al. 1993). For the construction of phylogenetic tree, only sequences from the type strains of species whose names have been validly published were taken into account. All the sequences were aligned using Clustalx program (Thompson et al. 1997) and the alignment was manually corrected. The evolutionary distance dendrogram was constructed using neighbor-joining analysis (Saitou and Nei 1987) and the distances between the sequences were calculated from the models of Jukes and Cantor (1969). Bootstrap analysis was performed to assess the confidence limits of the branching (Felsenstein 1985).
PCP degradation assay
The degradation studies were performed by inoculating 1% inoculum (2.5 × 103 cfu/ml) of the most PCP tolerant bacterial strain to 250 ml Erlenmeyer shake flask batch culture containing 99 ml MSM supplemented with 1.127 mM (300 mg/l) PCP in presence of 1% (w/v) glucose as carbon source, incubated at 30 ± 1 °C in a refrigerated incubator shaker (Innova 4230, NJ, USA) at 120 rev/min up to 168 h. The stability of organism in experimental and control (without inoculum) conditions was optimized at different environmental conditions (i.e. temperature, pH and aeration) to ensure that the disappearance of PCP was caused by biodegradation. The culture sample was removed under aseptic conditions. In order to estimate the PCP concentration cells were centrifuged at 5,000 g for 20 min (Remi C-24) and the supernatant was collected. Growth of the bacterial cells was determined in culture medium by measuring absorbance at 620 nm and PCP concentration in the supernatant was quantified by measuring absorbance at 320 nm at every 24 h interval up to 168 h by using UV-Vis spectrophotometer (GBC Cintra-40, Australia). For HPLC analysis the supernatant was initially acidified to pH 2.0 using 1N HCl and subsequently extracted three times using ethyl acetate (99.5%) in 1:1 ratio in a separating funnel by intermittent shaking. The extracted upper organic layer containing residual PCP was filtered through sodium sulphate to absorb excess water. Filtered samples were evaporated to dryness at elevated temperature (50–60 °C) in hot air oven, subsequently resuspended in 5.0 ml acetonitrile (HPLC grade) and analyzed using Waters 515 model equipped with UV-Vis (Waters-2487, Milford, USA) detector operating at 320 nm. Separation was carried out with a reverse phase watersmake 5 μm C-18 column (250 × 4.6) mm and the isocratic mobile phase was acetonitrile and water (70:30, v/v) with a flow rate of 1 ml/min. PCP standard was analyzed under the same conditions and the utilization of PCP was estimated by measuring the peak area of the compound. PCP degradation was also determined by estimation of the chloride ion released in aqueous media at every 24 h interval up to 168 h following the method described by Bergmann and Sanik (1957). Comparing respective standard curves quantitated amount of PCP degraded and chloride ion released.
Statistical analysis
Data were statistically analyzed by an overall one-way analysis of variance (ANOVA) and when differences observed were significant, the mean were compared by Tukey–Kramer Multiple Comparison Test. Linear regression analysis was used to determine the relation between the PCP degradation and chloride ion release were performed using Microsoft Excel 2000. All the experiments in this study were performed in triplicates and values are mean of three (n = 3) ± SD.
Results and discussion
Screening of bacteria for PCP tolerance
Out of the seven strains, ITRC S1, ITRC S2, ITRC S3, ITRC S4, ITRC S5, ITRC S7 were Gram negative and ITRC S6 was found Gram positive. Out of these seven aerobic bacterial isolates, ITRC S7 showed most potent for growth on PCP amended MSM agar in presence of glucose 0.5 and 1% (w/v) during the screening procedure whereas growth was negligible in absence of glucose (Table 1). The strain showed fast and luxuriant growth at various concentrations of PCP such as 0.376, 0.752 and 1.127 mM; however, no growth and change in colour of bromocresol purple were observed at a concentration of 1.503 mM PCP. Strain ITRC S7 was found to be the most potent isolate capable of degrading up to 1.127 mM PCP amended MSM agar in presence of 1% (w/v) glucose (Table 1). Change in colour of bromocresol purple from blue to yellow was due to the acidic pH resulting from release of chloride ion as well as hydrogen from PCP aromatic ring during dechlorination in the medium (Martins et al. 1997).
16S rDNA sequencing of strain ITRC S7 and phylogenetic analysis
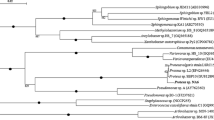
The results of the conventional identification techniques were supported by phylogenetic analysis. Hagstrom et al. (2000) found that 16S rDNA sequence having similarity of ≥97% is a reasonable level of grouping bacteria into species. Using universal primers 1442 bp of 16S rDNA of strain ITRC S7 was amplified, sequenced and submitted to the GenBank and the accession number AY927692 was obtained. Nucleotide–nucleotide BLAST searches of the obtained sequence showed 99% sequence similarity with the first hit of the search result i.e. Serratia marcescens strain AU736. Therefore, the most potent PCP-degrading isolate could be characterized as a strain of S. marcescens and was designated as S. marcescens strain ITRC S7. To understand the taxonomic position of strain ITRC S7 among other type strains of the genus Serratia, a detailed phylogenetic study was performed as shown in (Fig. 1). The evolutionary distance dendrogram revealed that all the sequences obtained from the database related to strain ITRC S7 fell into the class of γ-Proteobacteria, a lineage of domain Bacteria. The phylogenetic analysis showed 99% similarity with S. marcescens at the level of 16S rDNA sequence and also revealed its phylogenetic affiliation with some of the other members of Serratia species.
Phylogenetic neighbour-joining tree based on 16S rDNA sequences showing type strains of Serratia species related to strain ITRC S7 . Escherichia coli ATCC 11775T (X80725) was used as the outgroup. Numbers at nodes indicate levels of bootstrap support ≥50% based on a neighbour-joining analysis of 1000 resampled datasets. GenBank accession numbers are given in parentheses. Bar, 10 nucleotide substitution per 100 nucleotides
Degradation of PCP by strain ITRC S7
Figure 2a representing the growth curve of strain ITRC S7 indicated a prolonged lag phase followed by an exponential phase at approximately 140 h in presence of glucose. Up to 90.33% of PCP was significantly (ANOVA, P < 0.05) degraded within 168 h of incubation at 30 ± 1 °C, pH 7.0 and 120 rev/min (optimized condition) (Table 2). The simultaneous removal of PCP and release of chloride ion (2.435 mM) in the medium also emphasized the significant (ANOVA, P < 0.05) bacterial dechlorination and degradation of PCP (Table 2, Fig. 2b). In addition, no growth and degradation was observed in control sample during experiment (Fig. 2b).
(a) Growth curve (■–■) of Serratia marcescens (AY927692) in 1% glucose containing MSM, in the presence of 1.127 mM (300 mg/l) of PCP at optimized condition (30 ± 1 °C, pH 7.0 and 120 rev/min). All the experiments in this study were performed in triplicate and values are mean of three replicates. (b) PCP degradation and chloride ion release of Serratia marcescens (AY927692) in 1% glucose containing MSM, in the presence of 1.127 mM (300 mg/l) of PCP and its control (without inoculum) at optimized condition (Temperature 30 ± 1 °C, pH 7, 120 rev/min): (◆–◆) indicates PCP degradation (R2 = 0.986), (■–■) indicates chloride ion release (R2 = 0.9863) and (▴–▴) indicates control (R2 = 0.9837). (c) Effect of temperature on PCP degradation and chloride ion release of Serratia marcescens (AY927692). PCP degradation: (■–■) indicates 20 °C (R2 = 0.9871), (•–•) indicates 37 °C (R2 = 0.9873) and chloride ion release: (□–□)indicates 20 °C (R2 = 0.9869), (Ο–Ο) indicates 37 °C (R2 = 0.9871). (d) Effect of pH on PCP degradation and chloride ion release of Serratia marcescens (AY927692). PCP degradation: (■–■) indicates pH 6.0 (R2 = 0.9895), (•–•) indicates pH 9.0 (R2 = 0.9851) and chloride ion release: (□–□)indicates pH 6.0 (R2 = 0.9862), (Ο– Ο) indicates pH 9.0 (R2 = 0.9847). (e) Effect of aeration on PCP degradation and chloride ion release of Serratia marcescens (AY927692). PCP degradation: (■–■) indicates 50 rev/min (R2 = 0.983), (•–•) indicates 200 rev/min (R2 = 0.9862) and chloride ion release: (□–□)indicates 50 rev/min (R2 = 0.9844), (Ο–Ο) indicates 200 rev/min (R2 = 0.9863)
The rate of PCP degradation was not favoured by the variation at different environmental conditions when compared to the optimized condition as shown in Table 2 and Fig. 2 (c, d and e). Further, HPLC analysis confirmed the degradation of PCP (approx 90.33%) by showing disappearance of the peak of PCP as compared to control (Fig. 3).
There was no change in colour of bromocresol purple on PCP amended MSM agar medium in absence of glucose (Table 1). This phenomenon indicated that PCP degradation was a result of co-metabolism process, where glucose was utilized initially followed by PCP degradation. This finding was similar as reported earlier by Premalatha and Rajkumar (1994) for PCP degradation by Pseudomonas aeruginosa. The simultaneous release of chloride ion also corroborated the previous findings of dechlorination during PCP degradation as described by Mohn and Kennedy (1992). Variation in pH, temperature, aeration and increased PCP concentration did not favour the PCP degradation phenomenon. Alteration of optimized pH might be inhibitory to the activity of the enzymes responsible for PCP degradation in bacteria (Miller et al. 2004).
Our results suggest that the degradation of PCP by strain ITRC S7 does not go to completion because of the slow degradation rate of the metabolic intermediate. Results obtained for degradation of PCP through HPLC analysis of the extractable products show formation of tetrachloro-p-hydroquinone and 6-chlorohydroquinol in the extracts. The metabolites released during degradation of PCP was extracted and analyzed by HPLC. The culture medium was extracted twice with ethyl acetate. The extracted solution was acidified to pH 2.0 and extracted twice with ethyl acetate (acidic extract). PCP (RT 1.014) was degraded at 168 h and two new peaks appeared which were detected as tetrachloro-p-hydroquinone (TeCH) (RT 0.787) and 6-chlorohydroquinol (RT 0.916). The result of the study indicated conversion of PCP into TeCH, which further converted to 6-chlorohydroquinol. The identification of intermediary metabolites is a significant step in complete mineralization of PCP. Extraction methods are critical steps and selection of time points for sampling is another step, which must be optimized for degradation of PCP. Two methods are adopted for extraction of PCP and its intermediary metabolites for the complete mineralization of PCP. The main reason for this was the lack of sufficient information on the inhibitory effect of metabolites on microbial growth (data not shown).
The bacterial enzyme PCP-4-monooxygenase from Flavobacterium sp. strain ATCC 39723 catalyzes the oxygenolytic removal of the first chlorine from PCP. PCP-4-monooxygenase is a FAD-binding, NADPH-requiring oxygenase with similar functional domains as other bacterial flavoprotein monooxygenase specific for phenolic substrate (Lange et al. 1996). PCP-4-monooxygenase converts PCP to 2, 3, 5, 6-tetrachloro-p-hydroquinone (TeCH) in the presence of oxygen and NADPH, which is further converted to trichlorohydroquinone by reductive dechlorination and subsequently 2, 6-dichlorohydroquinone (DiCH) was formed by reductive dehalogenase enzyme (Xun and Orser 1991a; Xun et al. 1992a;). They showed that pcp A has novel ring-cleavage dioxygenase activity that cleaves aromatic rings with two hydroxyl group para to each other and that pcp A converts 2, 6-DCHQ to 2-chloromaleylacetate. This enzyme not only catalyzes dehalogenation but also removes hydrogen, nitro, amino and cyno groups from the benzene ring at the para position in relation to the hydroxyl of phenol (Xun et al. 1992b). Dechlorination of PCP by Flavobacterium sp. and PCP induces the presence of several proteins in the bacterium (Xun and Orser 1991b). Identification, purification, and characterization of PCP-4-monooxygenase that converts PCP to TeCH from P. fluorescens are also reported by Shah and Thakur (2003).
According to the reports (Xun and Orser 1991a; Cai and Xun 2002) of PCP degradation pathway, adding glucose facilitated the PCP metabolism by Sphingomonas chlorophenolica. The first possible reason was that some part of the metabolic pathway of PCP might overlap the metabolic pathway of glucose. PCP could be biotransformed into 2, 6 dichlorohydroquinone (2, 6-DCHQ). Subsequently, 2, 6-DCHQ entered the ring cleavage pathway to form maleylacetate and then maleylacetate entered the TCA cycle, which was an important metabolic pathway of glucose. Thus S. chlorophenolica might use some enzymes produced by metabolizing glucose to metabolize PCP. The second possible reason was that the glucose metabolism could provide more NADH to facilitate the first step. Therefore, S. chlorophenolica might use NADH produced by metabolizing glucose to metabolize PCP. The final conjecture could be that S. chlorophenolica used glucose, which were easily biodegraded to increase the biomass and thus increasing the total activity for metabolizing PCP. Further tests should be performed to determine the reason for facilitating PCP degradation using glucose.
Conclusions
S. marcescens strain ITRC S7 is a novel and most potent aerobic bacterial isolate capable of degrading PCP in axenic condition. Therefore, this strain may be used for bioremediation of PCP containing industrial wastes.
References
Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Ausubel FM, Brent R, Kingston RE, Moore DD, Seidman JG, Smith JA, Struhl K (1992) Current Protocols in Molecular Biology. John Wiley and Sons, New York
Bellinaso ML, Henriques JAP, Gaylarde CC (2001) Biodegradation as a biotechnological model for the teaching of biochemistry. World J Microbiol Biotechnol 17:1–6
Bergmann JG, Sanik J (1957) Determination of trace amounts of chlorine in naphtha. Anal Chem 29:241–243
Brosius J, Palmer ML, Kennedy PJ, Noller HF (1978) Complete nucleotide sequence of a 16S ribosomal RNA gene from Escherichia coli. Proc Nat Acad Sci USA 75:4801–4805
Cai M, Xun L (2002) Organization and regulation of pentachlorophenol-degrading genes in Sphingobium chlorophenolicum ATCC 39723. J Bacteriol 184:4672–4680
Copley SD (2000) Evolution of a metabolic pathway for degradation of toxic xenobiotics: the patch work approach. Trends Biochem Sci 25:261–265
Crosby DG (1981) Environmental chemistry of pentachlorophenol. Pure Appl Chem 53:1051–1080
EPA 1987 Final determination and indent to cancel and deny applications for registrations of pesticide products containing pentachlorophenol (including but not limited to its salts and esters) for non-wood uses. US Environmental Protection Agency. Fed Regist 52:2282–2293
Felsenstein J (1985) Confidence limits on phylogenetics: an approach using the bootstrap. Evolution 39:783–791
Hagstrom A, Pinhassi J, Zweifel UL (2000) Biogeographical diversity among marine bacterioplankton. Aquat Microbiol Ecol 21:231–244
Ito H, Ohnishi Y (1982) Escherichia coli mutants resistant to uncouplers of oxidative phosphorylation. Microbiol Immunol 226:1079–1084
Johnson JL (1994) Similarity analysis of rRNAs. In: Gerhardt P, Murray RGE, Wood WA, Krieg NR (eds) Methods for General and Molecular Bacteriology. American Society for Microbiology, Washington DC, pp 683–700
Jukes TH, Cantor CR (1969) Evolution of protein molecules. In: Munro HN (ed) Mammalian Protein Metabolism, vol. 3, Academic Press, New York, pp 21–132
Lange CC, Schnieder BJ, Orser CS (1996) Verification of the role of PCP-4-monooxygenase in chlorine elimination from pentachlorophenol by Flavobacterium sp. strain ATCC 39723. Biochem Biophys Res Commun 219:146–149
Larsen N, Olsen GJ, Maidak BL, McCaughey MJ, Overbeck R, Macke TJ, March TL, Woese CR (1993) The ribosomal database project. Nucleic Acids Res 21:3021–3023
Martins JM, Jocteur ML, Chalamet A, Bardin R (1997) Microbial response to repeated applications of low concentrations of pasture. Chemosphere 35:1637–1650
Miller MN, Stratton GW, Murray G (2004) Effects of soil moisture and aeration on the biodegradatic of pentachlorophenol contaminated soil. Bull Environ Contam Toxicol 72:101–108
Mohn WW, Kennedy KJ (1992) Reductive dehalogenation of chlorophenol by Desulfomonite tiedjei DCB-1. Appl Environ Microbiol 58:1367–1370
Pfennig N, Lippert KD (1966) Uber das vitamin B12-Bedurfnis phototropher Schwefelbakterien. Arch Mikrobiol 55:245–256
Premalatha A, Rajkumar GS (1994) Pentachlorophenol degradation by Pseudomonas aeruginosa. World J Microbiol Biotechnol 10:334–337
Raj A, Chandra R, Patel DK (2005) Physico-chemical characterisation of pulp and paper mill effluent and toxicity assessment by a tubificid worm, Tubifex tubifex. Toxicol Int 12:109–118
Saber DL, Crawford RL (1985) Isolation and characterization of Flavobacterium strains that degrade pentachlorophenol. Appl Environ Microbiol 50:1512–1518
Saitou N, Nei M (1987) The neighbour-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Schenk T, Muller R, Morsberger F, Otto MK, Lingens F (1989) Enzymatic dehalogenation of pentachlorophenol by extracts from Arthrobacter sp. strain ATCC 33790. J Bacteriol 171:5487–5491
Schnell A, Stell P, Melcer H, Hudson PV, Carey JH (2000) Enhanced biological treatment of bleached kraft mill effluents removal of chlorinated organic compounds and toxicity. Water Res 34:493–500
Shah S, Thakur IS (2002) Enrichment and characterization of pentachlorophenol degrading microbial community for the treatment of tannery effluent. Pollut Res 20:353–363
Shah S, Thakur IS (2003) Enzymatic dehalogenation of pentachlorophenol by Pseudomonas fluorescens of the microbial community from tannery effluent. Curr Microbiol 47:65–70
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The clustalx windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 24:4876–4882
Weisburg WG, Barns SM, Pelletier DA, Lane DJ (1991) 16S ribosomal DNA amplification for phylogenetic study. J Bacteriol 173:697–703
Williams ST, Sharpe ME, Holt JG (1993) Bergey’s Manual of Systemic Bacteriology. vol. 4, Williams and Wilkins, USA, pp 428
Yeber MC, Freer J, Martinez M, Mansilla HD (2000) Bacterial response to photolytic degradation of 6-chlorovanillin. Chemosphere 41:12257–12261
Xun L, Orser CS (1991a) Purification of a Flavobacterium pentachlorophenol-induced periplasmic protein (pcpA) and nucleotide sequence of the corresponding gene (pcpA). J Bacteriol 173:2920–2926
Xun L, Orser CS (1991b) Purification and properties of pentachlorophenol hydroxylase, a flavoprotein from Flavobacterium sp. strain ATCC39723. J Bacteriol 173:4447–4453
Xun L, Topp E, Orser CS (1992a) Confirmation of oxidation of oxidative dehalogenation of a pentachlorophenol by a Flavobacterium pentachlorophenol hydroxylase. J Bacteriol 174:5745–5747
Xun L, Topp E, Orser CS (1992b) Diverse substrate range of Flavobacterium pentachlorophenol hydroxylase. J Bacteriol 174:5745–5747
Acknowledgements
Authors are thankful to Director, ITRC, Lucknow, for his encouragement. The financial support from Council of Science and Technology, UP is also highly acknowledged.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Singh, S., Chandra, R., Patel, D.K. et al. Isolation and characterization of novel Serratia marcescens (AY927692) for pentachlorophenol degradation from pulp and paper mill waste. World J Microbiol Biotechnol 23, 1747–1754 (2007). https://doi.org/10.1007/s11274-007-9424-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11274-007-9424-5