Abstract
Detection, molecular characterization, and quantification of Norovirus (NoVs) in a semi-industrial pilot plant were performed in order to assess the efficiency of the secondary biological treatment using two different procedures: natural oxidation ponds and biodisks. A total of 102 wastewater samples were collected from two biological treatment processes in a semi-industrial pilot plant. NoVs GII and NoVs GI were detected and quantified in 65 % (n = 66) and in 1 % (n = 1) of the samples of wastewater from the plant, respectively. The average values of viral content (genome copies/μl) obtained in the effluent of the two lines of treatment showed a substantial reduction in the prevalence and in the viral content of NoVs GII detected from one basin to another of the five watersheds of the oxidation ponds and at the expiration of the biodisk line. The predominant genogroup of NoVs was NoVs GII (65 %), followed by NoVs GI (1 %). The predominant genotype of NoVs GII was GGII.12 (n = 11), followed by GGII.b (n = 1), GGII.1 (n = 1), and GGII.16 (n = 1) and two mixed combinations: GGI.2/GGII.12 (n = 5) and GGI.2/GGII.b (n = 1) were identified. The results obtained in this study represent the first documentation in Tunisia on the effectiveness of biological treatment for the removal of NoVs in the area of the capital of Tunis.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
The prevalence of enteric viruses in wastewater, the efficacy of wastewater treatments in removing such pathogens, and potential health risks from their release into the environment or by recycling the treated wastewater are very important issues in environmental microbiology. The monitoring of wastewater treatment plants (WTPs) may be a suitable approach for the study of circulating viruses in their respective service areas and the persistence of such viruses in treated effluents.
Enteric viruses are considered as the most common pathogens associated with human gastroenteritis. Human Noroviruses (NoVs) are found in all age groups (Atmar 2010; Tian et al. 2014) and are a major cause of foodborne and waterborne outbreaks (Lopman et al. 2012; Gould et al. 2013; Kim et al. 2014; Wilkes et al. 2014). Waterborne NoVs caused by contaminated surface water, drinking waters, mineral waters, recreational waters, and groundwater pose a grave danger to public health (Rutjes et al. 2006; Iwai et al. 2009; Wyn-Jones et al. 2011; Lee et al. 2012). NoVs are considered the second etiologic agent of severe acute gastroenteritis in infants and young children in Tunisia (Sdiri-loulizi et al. 2008; Hassine-Zaafrane et al. 2013) and in different countries of the world (Desai et al. 2012; Van Beek et al. 2013). Once excreted in feces at high concentrations, these viruses are present in wastewater where they are only partially removed by treatment processes. The resistance of these viruses to the treatment process facilitates their diffusion and their transmission by the fecal-oral route that can be directly from person to person or from animal to man or indirectly by ingestion of food or water contaminated (Van den Berg et al. 2005; Stals et al. 2013; Fuentes et al. 2014, Hassine-Zaafrane et al. 2014).
Human NoVs virions are icosahedral particles with a positive sense and single-stranded RNA molecule, belonging to the Caliciviridae family (Martella et al. 2008). Their genomes include three open reading frames (ORFs). ORF1 encodes a polyprotein precursor for several non-structural proteins; ORF2 encodes the capsid protein; and ORF3 encodes a small protein that plays a role in the stability of the viral capsid protein VP1 (Vinjé et al. 2004; Atmar 2010). Agreeing to the nucleotide sequence analysis of the capsid region, NoVs are subdivided into several genogroups (GI to GV) that can be further separated into several clusters or genotypes (Zheng et al. 2006). Genogroups GI, GII, and GIV have been found in humans, though GII seems to be the predominant around the world (Papaventsis et al. 2007; Al-Mashhadani et al. 2008). Genogroup I with 14 clusters and genogroup II with 17 clusters include most of the strains infecting humans (Kageyama et al. 2004; Zheng et al. 2006), but only one NoVs genotype (GGII.4) is the predominant circulating virus associated with global epidemics of gastroenteritis in the world and in Tunisia (Bellilot et al. 2010; Sdiri-Loulizi et al. 2009; Hassine-Zaafrane et al. 2013; Thongprachum et al. 2013; CDC 2013, Van Beek et al. 2013; Moyo et al. 2014; Franck et al. 2014).
Surveillance and prevention of NoVs in the water environment are an internal public health priority in Tunisia. In this study, molecular characterization and gene quantification of NoVs GII and GI from wastewater sampled from an area neighboring two hospitals were performed. The physicochemical analyses and enumeration of two enteric bacteria commonly used as fecal indicators which are the thermo-tolerant fecal coliforms and the fecal Streptococci were performed to assess health water quality. In addition, the effectiveness of secondary biological treatment using two different procedures at a semi-industrial pilot purification plant was studied.
2 Materials and Methods
2.1 The Pilot Plant and Effluent Sampling
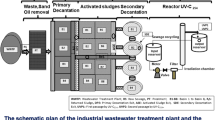
The pilot plant is connected to a sewage network system of the residential city of Tunis (Tunisia). This plant is also drained with contaminated wastewater of two nearby hospitals, and it has four processing lines of wastewater treatment: the line of natural oxidation pond formed by five different basins (B1, B2, B3, B4, and B5), the line of biological rotating disks (biodisks), the line of trickling filter, and finally the line of massif filter (Hassen et al. 1994). From January to December 2011, 102 of wastewater samples were collected. A total of 1 l of wastewater was collected from each basin (B1, B2, B3, B4, and B5) and at the biodisk line (D) at different rate frequency. Bacteriological, physicochemical, and virological analyses of wastewater samples collected were conducted within 3 days after collection.
2.2 Physicochemical and Bacteriological Analyses
The physicochemical analyses were performed for all samples. Samples were analyzed for electrical conductivity (EC), chemical oxygen demand (COD), biological oxygen demand (BOD5), ammonium nitrogen (N-NH4+), suspended solids (SS), nitrate nitrogen (N-N03 −), nitrous nitrogen (N-NO2 −), and ortho-phosphate (PO4-P). Enumeration of fecal indicator bacteria such as thermo-tolerant fecal coliforms and fecal Streptococci was performed according to the standard method of the most probable number MPN (Rodier 1978).
2.3 Virus Extraction
The recovery of viral particles from the wastewater samples was performed as described in the protocol of the US Environmental Protection Agency by using a method adapted from beef extract and AlCl3 (EPA 1992). Briefly, 10 ml of AlCl3 (0.05 M) were added to 1000 ml of sewage samples and pH value was adjusted to 3.5 using HCl (5 M). The mixture was centrifuged at 4500 rpm for 15 min at 4 °C, and viruses were eluted with 100 ml of extract of beef to 10 % (pH = 9). The mixture was again homogenized and centrifuged at 5000 rpm for 30 min at 4 °C. Viruses were concentrated by precipitation with polyethylene glycol 6000 (PEG 6000) as described by Lewis and Melcaft (1988). After incubation overnight at 4 °C, the extract was centrifuged at 5000 rpm for 30 min at 4 °C and the pellet was suspended in 2 ml of phosphate-buffered saline (PBS, pH = 7). Decontamination of the viral suspension was done by filtration using syringe filters of 0.22 μm. The filtrate was kept until the extraction of viral RNA at −20 °C.
2.4 Viral RNA Extraction
Viral RNA was isolated using 800 μl of sewage extract using an automatic extractor NucliSENS® EasyMag™ platform (bioMérieux, Marcy L’Etoile, France), according to the manufacturer’s instructions, and was stored at −40 °C.
2.5 Virus Detection and Quantification by Real-Time RT-PCR
NoVs GI and GII were detected and quantified by real-time RT-PCR using QNIF2d4, COG2R4 primers and QNIFS4 probe for GII NoVs: JJV1NF1 and JJV1R2/1 primers and JJV1P2/1 and RING-1b1/3 probes which amplify the gene segment coding for the capsid of NoVs GI (Kageyama et al. 2004; Jothikumar et al. 2005; Da Silva et al. 2007; Layman et al. 2009). The reaction was carried out in a 7500 Fast Real-Time PCR system using TaqMan® Fast Virus 1-Step Kit (Applied Biosystems), as recommended by the manufacturer. The reaction condition included a retro transcription step (50 °C for 5 min), activation of the polymerase at 95 °C for 20 s, and 40 cycles of cDNA amplification (95 °C for 20 s and 60 °C for 30 s). The quantification of the viral content was performed utilizing a standard range of ten dilutions (109 to100). The standard amplification was made with the samples.
2.6 Virus Detection by RT-PCR
All positive samples of NoVs GI and NoVs GII revealed by real-time PCR were tested by RT-PCR using several sets of primers in separate reactions. The primer set JV12/JV13 (Vinjé and Koopmans 1996) was used to amplify a fragment of the RNA polymerase gene of NoVs genogroups I and II. The primers sets G1SKF/G1SKR and G2SKF/G2SKR (Kojima et al. 2002) were used to detect a fragment of the capsid gene of NoVs genogroups I and II, respectively. RT-PCRs were performed using a Qiagen One-Step RT-PCR Kit (Qiagen) according to the manufacturer’s instructions and to the cycles of amplification given by the authors of each primer set.
2.7 Sequencing
All the amplified DNA samples were systematically purified from the agarose gel as positive strips cut using a QIAEX II Gel Extraction Kit (Qiagen, reference 20021) or from the PCR products using Amicon column (Ultra 30K. Millipore, reference UFC503096).
Sequencing of the PCR products was performed with the same primers as those for amplification by using the ABI PRISM® BigDye® Terminator cycle sequencing ready reaction kit (Applied Biosystems) and an ABI 3100 automated sequencer (PE Biosystems). The nucleotide sequence of the amplicons were compared to corresponding sequences of NoVs strains available in the GenBank database and in the National Reference Center for Enteric viruses in Dijon (France) using Fasta program.
3 Results
3.1 Physicochemical and Bacteriological Characteristics of Wastewater Samples
The various physicochemical and bacteriological parameters in the different lagoon during the period of sampling ranged on average as follows: 1350–1720 mg/l for electrical conductivity, 58–60 mg/l for SS, 155–240 mg/l for COD, 90–170 mg/l for BOD5, 18–22 mg/l for N-NH4, 18–22 mg/l for P-PO4, 1.6 × 104–1.6 × 106 (CFU/100 ml) for fecal coliforms, and 3.9 × 103–2.5 × 105 (CFU/100 ml) for fecal Streptococci. Similarly, the output values of the biodisk processes fluctuated between 1.4 and 1.7 mg/l for electrical conductivity, 26–40 mg/l for SS, 105–140 mg/l for COD, 30–50 mg/l for BOD5, 8.5–19 mg/l for N-NH4, 0.5–4.2 mg/l for N-NO2, 9–23.5 mg/l for N-NO3, 11–17 mg/l for P-PO4, 3.2 × 105–5.6 × 101 (CFU/100 ml) for fecal coliforms, and 6 × 104–6 × 105 (CFU/100 ml) for fecal Streptococci.
3.2 Prevalence of NoVs
Virological results showed that out of the 102 samples of wastewater analyzed, 66 were found positive for NoVs GII and only one for NoVs GI. Among these samples, 57 (56 %) were isolated from the five different natural pond basins and only 10 (10 %) were collected at the exit of the line of biodisks. The distributions of the overall prevalence of NoVs GII in the different pond basins and at the exit of the biodisk line were, respectively, 21.5 % (n = 22/102), 13 % (n = 13/102), 12 % (n = 12/102), 4 % (n = 4/102), 5 % (n = 5/102), and 10 % (n = 10/102) within the B1, B2, B3, B4, and B5 basins and in the biodisk line (D). The distribution of NoVs GII within the pond basins (B1 to B5) was used to determine the prevalence of NoVs GII in each different basin: 84.6 % (n = 22/26) in B1, 62 % (n = 13/21) in B2, 80 % (n = 12/15) in B3, 33.3 % (n = 4/12) in B4, and 31.2 % (n = 5/16) in B5. At the exit of the biodisk line, 83.3 % (n = 10/12) of samples were contaminated by NoVs GII (Fig. 1, Table 1). NoVs GI was detected in one sample of the basin B5.
3.3 Reduction of NoVs GII by Wastewater Treatment
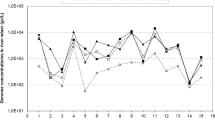
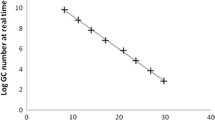
The quantification of NoVs GII content performed for all wastewater revealed some positive samples presenting values from 3.1 × 101 to 54 × 101 genome copies/μl in the two basins (B1 and B2) and from 2.4 × 10−2 to 6 × 101 genome copies/μl (gc/μl) in the three output basins (B3, B4, and B5). At the exit of the biodisk line, the NoVs GII was detected with a high frequency but the values of the viral content are low compared to the two pond basins B1 and B2 (from 6 × 10−1 to 2 gc/μl) (Table 1). The only sample contaminated by NoVs GI was detected with a high viral content around of 2.1 × 102 gc/μl. NoVs GII were present in the two lagoon basins B1 and B2 with important frequencies, 84.6 and 62 % in B1 and B2, respectively, and significant viral loads with average values of 101 and 32.5 gc/μl of NoVs GII in B1 and B2, respectively. Similarly, in the basin B3 and at the exit of the biodisk line (D), NoVs GII was found with high frequencies, 80 and 83.3 % in B3 and biodisk line (D), respectively, but their average values of the viral load are lower than those registered in the two basins B1 and B2 (1.72 gc/μl in B3 and 1.77 gc/μl in biodisk line (D)). However, in the two basins B4 and B5, NoVs GII was encountered with frequencies of 33.3 and 31.2 % in B4 and B5, respectively. Furthermore, the viral loads were lower than those registered in the two basins B1 and B2, 1.68 and 1.77 gc/μl in B4 and in B5, respectively (Table 1). The average values of viral content (gc/μl) recorded in the effluent of the two lines of treatment, natural oxidizing lagoons and biodisks, showed a significant reduction in the prevalence and in the viral content of NoVs GII. This decrease was observed from one basin to another of the five basins of natural lagoon and at the exit of the biodisks (P ≤ 0.05) (Table 1, Figs. 1 and 2).
3.4 Monthly Distribution of NoVs in the Two Lines of Treatments
The monthly distribution of positive NoVs GII during the sampling period (from November to December 2011) showed the presence and the survival of NoVs GII in wastewater having an alkaline pH (7.6 to 8.6) and moderate water temperatures ranging from 10 to 25.7 °C. Its presence was detected not only through the whole sampling period with a peak during the winter and spring season but also an unusual peak during the summer season (July) (Fig. 3, Table 1). Similarly, the variation of the average values of the viral content of NoVs GII over the seasons showed the occurrence of a peak during the cold seasons (37 gc/μl in February, 35 gc/μl in May, and 24 gc/μl in November) and the summer season (51 gc/μl of NoVs GII) (Fig. 4, Table 1). In the pond basin B1, NoVs GII was encountered with important frequencies and viral contents during the four seasons. In the pond basin B2, NoVs GII was detected during the four seasons with a peak during the winter and spring season, with values of 115 gc/μl of NoVs GII in February and 134 gc/μl of NoVs GII in March (Fig. 6, Table 1). In the pond basin B3, NoVs GII are encountered during the four seasons with high frequencies and viral contents lower than those registered in the two ponds basins (B1 and B2), with values of 1.5 to 1.7 gc/μl of NoVs GII (Fig. 6, Table 1). However, in the other basins (B4 and B5), NoVs GII is encountered during the four seasons with frequencies and viral contents lower than those registered in the two ponds basins (B1 and B2), with values of 1.5 to 4 gc/μl of NoVs GII (Fig. 6, Table 1). At the exit of the biodisk line (D), NoVs GII is encountered during the four seasons with high frequencies and viral contents lower than those registered in the two ponds basins (B1 and B2), with values of 1.2 to 1.7 gc/μl of NoVs GII (Fig. 6, Table 1). The only sample contaminated with NoVs GI was detected during December 2011. Results in Table 1 revealed a significant difference between the average values of viral load of NoVs GII (gc/μl) registered over the different seasons in the five basins of natural lagoon and at the exit of the biodisk line (P ≤ 0. 05) (Table 1, Figs. 3, 4, 5, and 6).
3.5 Molecular Characterization of NoVs
All positive samples for NoVs GII by real-time RT-PCR were tested by two conventional RT-PCR using two different pairs of primers. The NoVs GII was detected in 27 samples using the pair of primers G2SKR/GE2KF encoding for a fragment of the capsid gene of NoVs genogroups II. The NoVs GII was detected in 18 samples using the pair of primers JV12/JV13 encoding for a fragment of the RNA polymerase gene of NoVs genogroups I and II. The NoVs GI is not detected in the single positive sample of NoVs GI using the pairs of primers encoding for the capsid gene of NoVs genogroups I and the RNA polymerase gene of NoVs genogroups I and II.
The molecular typing of strains of NoVs GII was performed on all positive samples for NoVs GII by two conventional RT-PCR, but only the genotypes of 20 samples were determined. The genotyping revealed that the GII strains were the most frequent (66 and 1 % for GII and GI, respectively). The predominant genotype of NoVs GII was GII.12 (n = 11), followed by GII.b (n = 1), GII.1 (n = 1) and GII.16 (n = 1) and two mixed combinations: GI.2/GII.12 (n = 5) and GI.2/GII.b (n = 1) were identified (Table 2).
4 Discussion
Among human enteric viruses, NoVs are recognized as being the leading cause of epidemics and sporadic cases of gastroenteritis in all age groups (Atmar 2010; Tian et al. 2014). NoVs are the causative agents of gastroenteritis transmitted by contaminated food and water (Lopman et al. 2012; Sdiri-Loulizi et al. 2010). Waterborne NoVs caused by contaminated surface water, drinking waters, mineral waters, recreational waters, and groundwater pose a grave danger to public health (Rutjes et al. 2006; Iwai et al. 2009; Wyn-Jones et al. 2011; Lee et al. 2012).
The different samples of wastewater analyzed in the present study and tested at various levels of a semi-industrial pilot plant showed the potential role of treated wastewater as a possible route of transmission of NoVs and highlighted the high risk of infection related to the reuse of this kind of water for different purposes, mainly the agricultural reuse and/or their release in bathing and recreational water. Moreover, this study dealt with the effectiveness of the wastewater treatment concerning the abatement of NoVs since these viruses are considered among the main causes of clinical problems and of a high public health concern; thereby, we tried to assess the viral content circulating in wastewater sampled at different levels of a semi-industrial pilot plant situated in a residential area and drained with the wastewater of two hospitals. The molecular characterization of strains of NoVs was performed to compare virus strains in circulation in the case area of the Grand Tunis.
In the present study, the simultaneous analysis of some various physicochemical and bacteriological parameters performed in the two lines of treatment (pond basins B1 to B5 and biodisks) showed in general an average contrariwise result: poor physicochemical water quality and a good bacteriological quality in the case of the lagoon basin line, and unlike an excellent physicochemical water quality and a poor bacteriological quality in the case of the biodisk line. A similar study done in Tunisia by Hassen et al. (1994) reported comparable results as obtained in the present study showing the low effectiveness of removal of the main physicochemical parameter such DBO5 and the elimination of pathogens such as Streptococci and fecal coliforms by the natural lagoon basins and an inversely result for the biodisk line. On one side, it is well known that the residence time in treatment systems plays a fundamental role in the overall efficiency of removal of the various physicochemical and biological parameters of water quality. On another side and in considering that the residence time in the lagoon system is generally quite high (on average 30 days), it is obvious that the biological parameter removal is very important compared to that usually registered for physicochemical parameters (Hassen et al. 1994). By against, the residence time in the biodisk system is very short, with 4 to 5 h, it would actually be a biofiltration and the physicochemical water quality is greatly improved related to that usually recorded in the case of the lagoon system (Hassen et al. 1994). Indeed, it is well known that the biodisk treatment consists essentially on a sewage clarification (Hassen et al. 1994).
In this study, the high rate (65 %) of detection of NoVs GII in wastewater is not comparable to rates recorded in previous studies in Tunisia. The first clinical study has reported a prevalence of NoVs of 17.4 % between 2003 and 2005 in Tunisia (Sdiri-Loulizi et al. 2008). Similarly, the first environmental study has reported a low prevalence of NoVs (4.4 %) in sewage samples and (1.6 %) in shellfish samples between January 2003 and April 2007 in Tunisia (Sdiri-Loulizi et al. 2010). The second environmental study has showed an increase in the prevalence of NoVs (37.1 %) between 2007 and 2010 (Hassine-Zaafrane et al. 2014). Another study in Tunisia has revealed a prevalence of NoVs around of 35 % in shellfish samples (El-Amri et al. 2006). The high prevalence of NoVs GII (65 %) in wastewater samples confirms that the NoVs are considered the second etiological agent of viral gastroenteritis in children in Tunisia and in the world (Sdiri-Loulizi et al. 2008; Patel et al. 2008; Hassine-Zaafrane et al. 2013). In this study, the rate of detection of NoVs GII (65 %) in wastewater is not similar with rates recorded in previous studies such as Tunisia, Brazil, Venezuela, China, France, Greece, and Spain (Hassine-Zaafrane et al. 2014; Sdiri-Loulizi et al. 2010; Miagostovisch et al. 2008; Rodriguez-Diaz et al. 2009; He et al. 2011; Sima et al. 2011; Kokkinos et al. 2011; Calqua et al. 2013). These dissimilar detection rates could be explained by the difference in the conditions of study such as the sampling season and the sampling methods and the lack of hygiene and sanitation in countries. The stability of NoVs GII in waters has been previously described, and its resistance to the physicochemical processes used for sewage treatment might facilitate its transmission and its dissemination (Van den Berg et al. 2005).
The detection and the quantification of NoVs GII at the exit of the two treatment lines were carried out in a semi-industrial pilot plant in order to evaluate the effectiveness of these two lines of secondary biological treatment concerning the elimination of these viruses. The results showed a significant decrease in the prevalence and concentration of NoVs GII in the treated effluent. These results were comparable to those reported in other studies in Italy, in Brazil, and in China (La Rosa et al. 2010a, b; Prado et al. 2011; He et al. 2011). So, in considering this moderate efficiency of the virus abatement after these two lines of secondary treatment, it appeared that the remaining viral content of this pathogen in wastewater could reach the receiving water and the natural environment. Hence, this study highlighted the lack of sanitary quality of the treated wastewater for various recycling purposes and reuse and the need to perform tertiary treatment of disinfection in such situation, i.e., by UV irradiation. These data are in agreement with previous studies showing that NoVs are present in high levels not only in raw but also in treated wastewaters (Laverick et al. 2004; Van den Berg et al. 2005; Da Silva et al. 2007; La Rosa et al. 2010a, b; Maunula et al. 2012, Hassine Zaafrane et al. 2014).
The seasonal distribution of NoVs GII in this study confirmed the result of two earlier Tunisian studies achieved commonly on patient and wastewater sampling since the results revealed that the NoVs GII were detected in the population and in the wastewater during the winter season and an unusual peak from May to September (Sdiri-Loulizi et al. 2008; 2010). Similarly, in Finland, NoVs were present during spring and winter season in river water and in treated wastewater released into the river water (Maunula et al. 2012). The results obtained in this study are dissimilar to those obtained in two recent studies in Sweden and in Italy showing the abundance of NoVs GII in wastewater during spring and summer season (Nordgren et al. 2009; La Rosa et al. 2010a). However, in China, the peak of NoVs detection in wastewater was registered in cold weather from October to March (He et al. 2011). The change in the temporal seasonal distribution of NoVs GII can be explained by the variability in the average temperature, the humidity, the immune status of the host, and the socioeconomic situation in each country.
This study describes methods for sample collection and processing of wastewater for molecular detection of enteric viruses, and it also describes both traditional and real-time RT-PCR assays for NoVs diagnosis. Traditional RT-PCR methods are usually used for diagnosis, but they involve the usage of a confirmatory test. More recently, real-time RT-PCR assays have been developed allowing a rapid and accurate identification of NoVs in wastewater samples. The ability to rapidly detect and quantify viruses in environmental samples using real-time PCR represents a considerable advancement, holding great potential for environmental applications. This approach allowed us to overcome most of the disadvantages associated with both conventional cell culture and direct PCR assays, reducing the time needed for the detection of infectious viruses. The real-time PCR has been demonstrated to be a powerful tool for rapid determination of human NoVs in environmental samples and represents a considerable advancement in pathogen quantification in aquatic environments. Other works have demonstrated the effectiveness of real-time RT-PCR for detection of human NoVs in wastewater and in fecal samples (Atmar et al. 2011; La Rosa et al. 2010b).
Genotypes of NoVs GII were identified in 20 positive samples. The genotyping study showed that the genotype GGII-12 (n = 11) was the predominant, followed by GII.b (n = 1), GGII.1 (n = 1), and GGII.16 (n = 1) with two mixed combinations, GGI.2/GGII.12 (n = 5) and GGI.2/GGII.b (n = 1). These results are discordant with those obtained in other Tunisian studies. In Tunisia, clinical studies have demonstrated that the GGII.4 was the most prevalent genotype between 2003 and 2010. Interestingly, the hunter variant, which was the unique GII.4 variant observed from 2003 to April 2007 in the region of Monastir, replaced the 2006b variant between 2007 and 2010 (Sdiri-Loulizi et al. 2009; Hassine-Zaafrane et al. 2013). Similarly, an environmental study has revealed that the GGII.4 was the most prevalent genotype in wastewater samples in the region of Monastir (Sdiri-Loulizi et al. 2010). In previous Tunisian studies, GGI.2 genotype was detected in sewage, in selfish, and fecal samples, but GGII.1 and GGII.12 were only detected in fecal material and in wastewater, respectively (Sdiri-Loulizi et al. 2009, 2010; Hassine-Zaafrane et al. 2013, 2014). This study showed the presence of new genotypes such as GGII.1, GGII.b, and GGII.16 in the residential city of Tunis. GI.2, GI.5, GI.6, and GI.1 genotypes were identified in sewage, in river, in ground, and in marine waters in Thailand, in Italy, in Korea, and in Hong Kong (La Rosa et al. 2010a; Kittigul et al. 2012; Lee et al. 2012; Yang et al. 2012). GII.4 genotype is the most prevalent genotype in sewage in Italy and in Greece, in five rivers of Argentina, in marine waters in Hong Kong and in Korean groundwater (La Rosa et al. 2010a; Kokkinos et al. 2011; Fernández et al. 2012; Yang et al. 2012; Lee et al. 2012). In Italy and in Argentina, GII.b, GII.6, GII.2, GII.7, and GII.17 genotypes were detected in sewage and in river water (La Rosa et al. 2010a; Fernández et al. 2012).
This study revealed a predominance of NoVs GII compared to NoVs GI. These results are similar to those reported in other studies conducted on clinical and wastewater samples in Tunisia (Sdiri-Loulizi et al. 2008, 2009, 2010). On the contrary, a recent Tunisian study revealed a dominance of NoVs GI compared to NoVs GII in sewage samples (Hassine-Zaafrane et al. 2014). However, the significant presence of NoVs GI with high rates and viral content in wastewater samples has been described in Italy, in France, in Sweden, and in Thailand (Da Silva et al. 2007; Nordgren et al. 2009; La Rosa et al. 2010a, b; Kittigul et al. 2012).
Moved over the considerable load of viral gastrointestinal disease and in the absence of costly epidemiological monitoring systems, virological monitoring of sewage flowing into a wastewater treatment plant may represent an inexpensive and effective approach that would earmark for an ongoing follow-up on the occurrence of pathogenic viruses in the plant’s service area. The monitoring of effluent samples, on the other hand, may contribute to the understanding of the viral removed through wastewater treatment and to the prevention disease. The different samples of wastewater analyzed in the present study and tested at various levels of a semi-industrial pilot plant showed the potential role of treated wastewater as a possible route of transmission of NoVs and highlighted the high risk of infection related to the reuse of this kind of water for different purposes, mainly the agricultural reuse and/or their release in bathing and recreational water. Moreover, this study dealt with the effectiveness of the wastewater treatment concerning the abatement of NoVs since these viruses are considered among the main causes of clinical problems and of a high public health concern; thereby, we tried to assess the viral content circulating in wastewater sampled at different levels of a semi-industrial pilot plant situated in a residential area and drained with the wastewater of two hospitals. The molecular characterization of strains of NoVs was performed to compare virus strains in circulation in the case area of the Grand Tunis.
The main goals of this study were to assess the efficiency of two secondary biological treatment lines (natural oxidation ponds and biodisks) concerning the NoVs removal, to indicate the impact of wastewater on the potential spreading and dissemination of these infectious particles in the environment, and to establish important standards for the management policy of wastewater treatment and for public health interest. A moderate improvement of the water bacteriological and virological quality was insured by the natural lagoon basins and the biodisk procedure treatment. Moreover, these two types of treatment have highlighted the inadequacy of the sanitary quality of treated wastewater at the exit of these two lines for recycling and reuse purposes since there were still some infectious NoVs in water. So, tertiary treatment of disinfection is essential and needed for the removal of infectious particles at high rates and at a safe level.
References
Al-Mashhadani, M. N., Nakagomi, O., Dove, W., Ahmed, H., Nakagomi, T., Hart, C. A., & Cunliffe, N. A. (2008). Norovirus gastroenteritis among children in Iraqi Kurdistan. Journal of Medical Virology, 80, 506–509.
Atmar, R. L. (2010). Noroviruses-State of the Art. Food Environmental Virology, 2(3), 117–126.
Atmar, R. L., Neil, F. H., & Le Guyader, F. C. (2011). Detection of human Caliciviruses in fecal samples by RT-PCR. Methods in Molecular Biology, 665, 39–50.
Bellilot, G., Kamel, A. H., Estinney, M., Ambert-Balay, K., & Pothier, P. (2010). Evidence of emergence of new GII.4 norovirus variants from gastroenteritis outbreak survey in France during the 2007-to-2008 and 2008-to-2009 winter seasons. Journal of Clinical Microbiology, 48(3), 994–998.
Calqua, B., Fumian, T., Russiñol, M., Rodriguez-Manzano, J., Mbayed, V. A., Bofill-Mas, S., Miagostovich, M., & Girones, R. (2013). Detection and quantification of classic and emerging viruses by skimmed-milk flocculation and PCR in river water from two geographical areas. Water Research, 47(8), 2797–2810.
CDC. (2013). Emergence of new norovirus strain GII.4 Sydney—United States 2012. MMWR. Morbidity and Mortality Weekly Report, 62(3), 55.
Da Silva, A. K., Le Saux, J. C., Parnaudeau, S., Pommepuy, M., Elimelech, M., & Le Guyader, F. S. (2007). Evaluation of removal of noroviruses during wastewater treatment, using real-time reverse transcription-PCR: different behaviors of genogroups I and II. Applied and Environmental Microbiology, 73, 7891–7897.
Desai, R., Henbree, C. D., Handel, A., Mattwess, J. E., Dickey, B. W., McDonald, S., Hall, A. J., Parashar, U. D., Leon, J. S., & Lopman, B. (2012). Severe outcomes are associated with genogroup 2 genotype 4 norovirus outbreaks: a systematic literature review. Clinical Infectious Diseases, 55(2), 189–193.
El-Amri, D. E., Aouni, M., Panaudeau, S., & Le Guyader, F. S. (2006). Detection of human enteric viruses in selfish collected in Tunisia. Letters in Applied Microbiology, 43(4), 399–404.
EPA. (1992). Environmental protection agency: standard for the disposal of sewage sludge. In Federal Register part 503, 9387–9404
Fernández, M. D., Torres, C., Poma, H. R., Riviello-López, G., Martinez, L. C., Cisterna, D. M., Rajal, V. B., Nates, S. V., & Mbayed, V. A. (2012). Environmental surveillance of norovirus in Argentina revealed distinct viral diversity patterns, seasonality and spatio-temporal diffusion processes. Science of the Total Environment, 15(437), 262–269.
Fuentes, C., Guix, S., Pérez-Rodriguez, F. J., Fuster, N., Carol, M., Pintó, R. M., & Bosch, A. (2014). Standardized multiplex one-step qRT-PCR for hepatitis A virus, norovirus GI and GII quantification in Bivalve mollusks and water. Food Microbiology, 40, 55–63.
Franck, K. T., Fonager, J., Ersbøll, A. K., & Böttiger, B. (2014). Norovirus epidemiology in community and health care settings and association with patient age, denmark. Emerging Infectious Diseases, 20(7), 1123–1131.
Gould, L. H., Walsh, K. A., Vieira, A. R., Herman, K., Williams, I. T., Hall, A. J., Cole, D., & Division of Foodborne, Waterborne and Environmental Diseases, National Center for Emerging and Zoonotic Infectious Diseases. (2013). Surveillance for foodborne disease outbreaks—United States, 1998–2008. M M W R Surveillance Summaries, 62(2), 1–34.
Hassen, A., Jedidi, N., Kallali, H., Ferchichi, M., Ghrabi, A., Chebbi, F., Saidi, N., Shayeb, H., & Ennabli, M. (1994). Elimination des indicateurs et des métaux lourds au cours de traitement des eaux usées domestiques dans une station pilote semi-industrielle. Science Technology de l’Eau, 27(4), 34–41.
Hassine-Zaafrane, M., Sdiri-Loulizi, K., Kaplon, J., Ben Salem, I., Pothier, P., Aouni, M., & Ambert-Balay, K. (2014). Molecular detection of human Noroviruses in influent and effluent samples from two biological sewage treatment plants in the region of Monastir, Tunisia. Food Environmental Virology, 6, 125–131.
Hassine-Zaafrane, M., Sdiri-Loulizi, K., Kaplon, J., Salem, I. B., Pothier, P., Aouni, M., & Ambert-Balay, K. (2013). Prevalence and genetic diversity of norovirus infection in Tunisian children (2007–2010). Journal of Medical Virology, 85(6), 1100–1110.
He, X. Q., Cheng, L., Zhang, D. Y., Xie, X. M., Wang, D. H., & Wang, Z. (2011). One-year monthly survey of rotavirus, astrovirus and norovirus in three sewage treatment plants (STPs) in Beijing, China and associated health risk assessment. Water Science and Technology, 64(6), 1202–1210.
Iwai, M., Hasegawa, S., Obara, M., Nakamura, K., Horimoto, E., Takizawa, T., Kurata, T., Sogen, S., & Shiraki, K. (2009). Continuous presence of noroviruses and saporoviruses in raw sewage reflects infections among inhabitants of Toyama, Japan (2006 to 2008). Applied and Environmental Microbiology, 75(5), 1264–1270.
Jothikumar, N., Lowther, J. A., Henshilwood, K., Lees, D. N., Hill, V. R., & Vinjé, J. (2005). Rapid and sensitive detection of noroviruses by using TaqMan based one-step reverse transcription-PCR assays and application to naturally contaminated shellfish samples. App Environmental Microbiology, 71(4), 1870–1875.
Kageyama, T., Shinohara, M., Uchida, K., Fukushi, S., Hoshino, F. B., Kojima, S., Takai, R., Oka, T., Takeda, N., & Katayama, K. (2004). Coexistence of multiple genotypes, including newly identified genotypes, in outbreaks of gastroenteritis due to norovirus in Japan. Journal of Clinical Microbiology, 42, 2988–2995.
Kim, A.N., Park, S.Y., Bae, S.C., Oh, M.H., Ha, S.D. (2014). Survival of Norovirus surrogate on various food-contact surfaces. Food Environ Virol PMID:24919545
Kittigul, L., Panjangampatthana, A., Pombubpa, K., Taweekate, Y., Pungchitton, S., Diraphat, P., & Siripanichgon, K. (2012). Detection and genetic characterization of norovirus in environmental water samples in Thailand. Southeast Asian Journal of Tropical Medicine and Public Health, 43(2), 323–332.
Kojima, S., Kageyama, T., Fukushi, S., Hoshino, F. B., Shinohara, M., Uchida, K., Natori, K., Takeda, N., & Katayama, K. (2002). Genogroup-specific PCR primers for detection of Norwalk-like viruses. Journal of Virological Methods, 100(1–2), 107–114.
Kokkinos, P., Ziros, P., Meri, D., Filippidou, S., Kolla, S., Galanis, A., Vantarakis, A., & Environmental surveillance. (2011). An additional/alternative approach for virological surveillance in Greece? International Journal of Environmental Research and Public Health, 8(6), 1914–22.
La Rosa, G., Laconelli, M., Pourshaban, M., & Muscillo, M. (2010a). Detection and molecular characterization of noroviruses from five sewage treatment plants in central Italy. Water Research, 44(6), 1777–84.
La Rosa, G., Pourshaban, M., Iaconelli, M., & Muscillo, M. (2010b). Quantitative real-time of enteric viruses in influent and effluent samples from wastewater treatment plants in Italy. Annals Ist Super Sanita, 46(3), 266–273.
Layman, W. H., Walsh, J. F., Kotch, J. B., Weber, D. J., Gunn, E., & Vinjé, J. (2009). Prospective study of etiologic agents of acute of gastroenteritis outbreaks in child care centers. Journal de Pediatria, 154(2), 253–257.
Lee, G. C., Jheonq, W. H., Junq, G. S., Oh, S. A., Kim, M. J., Rhee, O. J., ParK, S., & Lee, C. H. (2012). Detection and molecular characterization of human noroviruses in Korean groundwater between 2008 and 2010. Food Environmental Virology, 4(3), 115–23.
Lewis, G. D., & Melcaft, T. G. (1988). Polyethylene glycol precipitation for recovery of pathogenic viruses, including hepatitis A virus and Human Rotavirus, from oyster, water and sediment samples. Applied and Environmental Microbiology, 54, 1983–1988.
Lopman, B., Gastañaduy, P., Park, G. W., Hall, A. J., Parashar, U. D., & Vinjé, J. (2012). Environmental transmission of norovirus gastroenteritis. Current Opinion Virology, 2(1), 96–102.
Martella, V., Lorusso, E., Decaro, N., Elia, G., Radogna, A., D’Abramo, M., Desario, C., Cavalli, A., Corrente, M., Camero, M., Germinanio, C. A., Banyai, K., Di Martino, B., Marsilio, F., Carmichael, L. E., & Buonavoglia, C. (2008). Detection and molecular characterization of a canine norovirus. Emerging Infectious Diseases, 14, 1306–1308.
Maunula, L., Söderberg, K., Vahtera Vuorilehto, V. P., von Bonsdorff, C. H., Valtari, M., Laakso, T., & Lahti, K. (2012). Presence of Human noro-and adenoviruses in river and treated wastewater, a longitudinal study and method comparison. Journal of Water and Health, 10(1), 87–99.
Miagostovisch, M. P., Ferreira, F. F., Guimaraes, F. R., Fumian, T. M., Dniz-Mends, L., Luz, S. L., Silvia, L. A., & Leite, J. P. (2008). Molecular detection and characterization of gastroenteritis viruses occurring naturally in the stream waters of Manaus, central Amazonia, Brazil. Applied and Environmental Microbiology, 74(2), 375–82.
Moyo, S., Hanevik, K., Blomberg, B., Kommedal, O., Vainio, K., Maselle, S., & Langeland, N. (2014). Genetic diversity of norovirus in hospitalized diarrhoeic children and asymptomatic controls in Dar Es Salaam, Tanzania. Infection, Genetics and Evolution, 26C, 340–347.
Nordgren, J., Matussek, A., Mattsson, A., Svensson, L., & Lindgren, P. E. (2009). Prevalence of norovirus and factors influencing virus concentrations during 1 year in a full-scale wastewater treatment plant. Water Research, 43(4), 1117–1125.
Papaventsis, D. C., Dove, W., Cunliffe, N. A., Nakagomi, O., Combe, P., Grosjean, P., & Hart, C. A. (2007). Norovirus infection in children with acute gastroenteritis, Madagascar, 2004–2005. Emerging Infectious Diseases, 13, 908–911.
Patel, M. M., Widdowson, M. A., Glass, R. I., Akazawa, K., Vinjé, J., & Parashar, U. D. (2008). Systematic literature review of the role of noroviruses in sporadic gastroenteritis. Emerging Infectious Diseases, 14, 1224–1231.
Prado, T., Silva, D. M., Guilayn, W. C., Rose, T. L., Gaspar, A. M., & Miagostovish, M. P. (2011). Quantification and molecular characterization of enteric viruses detected in effluents from two hospital wastewater treatment plants. Water Research, 45(3), 1287–1297.
Rodier, (1978). L’analyse de l’eau. Eaux naturelles, eaux résiduaires, eaux de mer. 6th Édn, Dunod, Paris, ISBN: 2100496360, 1383.
Rodriguez-Diaz, J., Querales, L., Caraballo, L., Vizzi, E., Liprandi, F., Takiff, H., & Betancourt, W. Q. (2009). Detection and characterization of waterborne gastroenteritis viruses in urban sewage and sewage polluted river waters in Caracas, Venezuela. Application Environmental Microbiology, 75(2), 387–94.
Rutjes, S. A., Van Der Berg, H. H., Lodder, W. J., & de Roda Husman, A. M. (2006). Real-time detection of noroviruses in surface water by use of a broadly reactive nucleic acid-sequence-based amplification assay. Applied and Environmental Microbiology, 72(8), 5349–58.
Sdiri-Loulizi, K., Hassine, M., Ouni, Z., Gharbi-Khelifi, H., Chouchane, S., Sakly, N., Neji-Guédiche, M., Pothier, P., Ouni, M., & Ambert-Balay, K. (2010). Detection and molecular characterization of enteric viruses in environmental samples in Monastir, Tunisia, between January 2003 and April 2007. Journal of Applied Microbiology, 109, 1093–1104.
Sdiri-Loulizi, K., Ambert-Balay, K., Gharbi-Khelifi, H., Sakly, N., Hassine, M., Chouchane, S., Guediche, M. N., Pothier, P., & Aouni, M. (2009). Molecular epidemiology of Norovirus gastroenteritis investigated using samples collected from children in Tunisia during a 4-year period: detection of the Norovirus variant GGII.4 as early as January 2003. Journal of Clinical Microbiology, 47(2), 421––429.
Sdiri-Loulizi, K., Gharbi-Khelifi, H., De Rougemont, A., Ouni, Z., Chouchane, S., Sakly, N., Ambert-Balay, K., Hassine, M., Neji-Guédiche, M., Ouni, M., & Pothier, P. (2008). Acute infantile gastroenteritis associated with human enteric viruses in Tunisia. Journal of Clinical Microbiology, 46(4), 1349–1355.
Sima, L. C., Schaefffer, J., Le Saux, J. C., Parnaudeau, S., Elimelech, M., & Le Guyader, F. S. (2011). Calicivirus removal in a membrane bioreactor wastewater treatment plant. Applied and Environmental Microbiology, 77(15), 5170–5177.
Stals, A., Uyttendaele, M., Baert, L., & Van Coilie, E. (2013). Norovirus transfer between foods and food contact materials. Journal Food Production, 76(7), 1202–1209.
Thongprachum, A., Khamrin, P., Chan-It, W., Malasao, R., Chaimangkol, N., Okitsu, S., Mizuguchi, M., Maneekarn, N., Hayakawa, S., & Ushijima, H. (2013). Emergence of norovirus GII/4 2006a and 2006b variants in hospitalized children with acute gastroenteritis in Thailand. Clinical Laboratory, 59(3–4), 271–276.
Tian, G., Jin, M., Li, H., Li, Q., Wang, J., & Duan, Z. J. (2014). Clinical characteristics and genetic diversity of noroviruses in adults with acute gastroenteritis in Beijing, China in 2008–2009. Journal of Medical Virology, 86(7), 1235–1242.
Van Beek, J., Ambert-Balay, K., Botteldoorn, N., Eden, J., Fonager, J., Hewitt, J., Iritani, N., Kroneman, A., Vennema, H., Vinjé, J., White, P. A., Koopmans, M., & NoroNet. (2013). Indications for worldwide increased norovirus activity associated with the emergence of a new variant of genotype II.4 late 2012. Euro Surveillance, 18(1), 8–9.
Van den Berg, H., Lodder, W., van der Poel, W., Vennema, H., & de Roda Husman, A. M. (2005). Genetic diversity of noroviruses in raw and treated sewage water. Research in Microbiology, 156(4), 532–540.
Vinjé, J., Hamidjaja, R. A., & Sobsey, M. D. (2004). Development and application of a capsid VP1 (region D) based reverse transcription PCR assay for genotyping of genogroupe I and II noroviruses. Journal of Medical Virology, 116, 109–117.
Vinjé, J., & Koopmans, M. P. (1996). Molecular detection and epidemiology of small round-structured viruses in outbreaks of gastroenteritis in the Netherlands. Journal of Infectious Diseases, 174, 610–615.
Wilkes, G., Brassard, J., Edge, T. A., Gannon, V., Gottschall, N., Jokinen, C. C., Jones, T. H., Khan, I. U., Marti, R., Sunohara, M. D., Topp, E., & Lapen, D. R. (2014). Long-Term monitoring of waterborne pathogens and microbial source tracking markers in paired agricultural watersheds under controlled and conventional tile drainage management. Applied and Environmental Microbiology, 80(12), 3708–3720.
Wyn-Jones, A. P., Carducci, A., Cook, N., D’Agostino, M., Divizia, M., Fleischer, J., Gantzer, C., Gawler, A., Girones, R., Höller, C., de Roda Husman, A. M., Kay, D., Kozyra, I., López-Pila, J., Muscillo, M., Nascimento, M. S., Papageorgiou, G., Rutjets, S., Sellwood, J., Szewzyk, R., & Wyer, M. (2011). Surveillance of adenoviruses and noroviruses in European recreational waters. Water Research, 45(3), 1025–1038.
Yang, N., Qi, H., Wong, M. M., Wu, R. S., & Kong, R. Y. (2012). Prevalence and diversity of norovirus genogroups I and II in Hong Kong marine waters and detection by real-time PCR. Marine Pollution Bulletin, 64(1), 164–168.
Zheng, D. P., Ando, T., Fankhauser, R. L., Beard, R. S., Glass, R. I., & Monroe, S. S. (2006). Norovirus classification and proposed strain nomenclature. Virology, 346, 312–323.
Acknowledgments
This study is jointly supported by the Laboratory of National Center of Enteric Viruses (Dijon, France) and by the Tunisian Ministry of Higher Education and Scientific Research within the contract program (2010–2013). We are also grateful to our friends and colleagues, French and Tunisian, for their encouragement and support during the progress of this study.
Conflict of Interest
The authors declare that they have no conflicts of interests.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ibrahim, C., Cherif, N., Hammami, S. et al. Quantification and Molecular Characterization of Norovirus After Two Wastewater Treatment Procedures. Water Air Soil Pollut 226, 187 (2015). https://doi.org/10.1007/s11270-015-2402-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11270-015-2402-x