Abstract
Tobacco etch virus (TEV) strains HAT, Mex21, and N have been the focus of numerous studies to dissect a host resistance mechanism in Capsicum spp. Little is known, however, about their general pathogenicity and genomic sequence data are not available on the TEV strains Mex21 and N. Four Nicotiana spp. were evaluated after inoculation with each TEV strain. Nicotiana tabacum ‘Kentucky 14’ and N. clevelandii plants expressed varied systemic symptoms dependent on the TEV strain; however, disease severity increased from HAT (mild mosaic symptoms) to Mex21 (more severe mosaic symptoms with stunting) to N (severe chlorosis and stunting). Nicotiana tabacum ‘Samsun’ plants developed relatively milder symptoms and N. glutinosa plants remained symptomless, although they were systemically infected. The genome of each TEV strain was sequenced and shown to consist of 9,495 nucleotides and a polyprotein of 3,054 amino acids. Comparison of their nucleotide sequences relative to the original HAT sequence (GenBank Accession No. M11458) revealed 95, 92, and 92 % identity for HAT-AU (from Auburn University), Mex21, and N, respectively. HAT-AU had 91 % sequence identity with Mex21 and N, while Mex21 and N were more closely related with 98 % nucleotide sequence identity. Similarly, the amino acid sequence identities for the full-length polyprotein ranged from 95 % for HAT-AU when compared with N to a high of 98 % identity between Mex21 and N.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Tobacco etch virus (TEV) is a member of the genus Potyvirus, in the family Potyviridae, which includes the largest number of virus species among the plant viruses [1]. Potyviruses are flexuous rod-shaped particles that possess a positive sense, single-stranded RNA genome of approximately 10 kb [1]. The potyvirus genome encodes a single polyprotein that was shown to be cleaved into ten individual proteins [2], with an additional open reading frame, PIPO, recently shown to occur within the P3 coding region [3]. Numerous potyviral proteins were shown to form inclusions in the infected cell, some of which develop in the cytoplasm, whereas others develop in the nucleus [4]. The viral RNA has a protein covalently linked to its 5′-terminus, referred to as the VPg, [5] and a 3′ poly (A) tract [6].
TEV is widely distributed in North, Central and South America, and more recently found in Europe and Asia [7]. TEV has a moderately wide host range including 149 plant species in 19 families [8, 9], although most known hosts are in the Solanaceae. Plant virus-induced losses in tobacco have been reported in Georgia and North Carolina (U.S.A.) and China [10–13]. TEV and the Potyvirus, Tobacco vein mottling virus, caused losses estimated to be $2 million in the mid-1980s in North Carolina [10]. Tomato spotted wilt virus (genus Tospovirus) was reported to cause tobacco stand losses as high as 40 % in Georgia and North Carolina [11]. In China, plant viruses are considered the limiting factors in tobacco production, with losses averaging 20–70 % annually [13].
The first report of the full-length sequence for TEV was for the HAT strain [14] and shown to consist of 9,496 nucleotides with a predicted translational product of a 3,054 amino acid polyprotein [14]. Since the first publication of the HAT sequence, four additional TEV strains (NW, 7DA, SD1, and Shannxi) have had their full-length genome sequenced and made available on the Genbank database.
The three TEV strains compared in this study have been the focus in recent studies on genetic resistance in Capsicum spp. to virus infection [15–17]. Despite the importance of these studies and additional studies that included these TEV strains [18, 19], comparative evaluation of their pathogenicity is lacking and full-length genome sequence data are not available for HAT-AU (the HAT strain maintained at Auburn University), Mex21, and N. In this study, the pathogenicity of three TEV strains in different Nicotiana spp. is described along with selected characteristics of their completely sequenced genomes.
Materials and methods
Virus strains
The TEV strains used in this study included HAT, Mex21, and N. In order to distinguish the original, HAT strain [14], used as a reference in the sequencing sections from the HAT strain maintained at Auburn University, was used throughout this project, the latter will be referred to as HAT-AU. HAT-AU was originally obtained from Dr. T. Pirone, University of Kentucky. Mex21 was obtained from Dr. Molly Jahn, Cornell University, although it was originally provided by Dr. Lowell Black (Louisiana State University) as an isolate from Tampico, Mexico. The N strain was provided by Dr. C. M. Deom, University of Georgia. Each virus was maintained by mechanical passage in Nicotiana tabacum L. ‘Kentucky 14’ with occasional passage through Capsicum annuum L. ‘Calwonder’. The virus strains were propagated in a temperature-controlled greenhouse (mean temperatures of 24 °C for day and 20 °C for night throughout the year) at the Plant Science Research Facility on the campus of Auburn University, AL, U.S.A.
Pathogenicity of the TEV strains in tobacco
The TEV strains were evaluated for their ability to infect different Nicotiana species. Test plants included N. tabacum cv ‘Kentucky 14’, N. tabacum cv ‘Samsun’, N. clevelandii, and N. glutinosa. Seed for each tobacco treatment was sown in Pro-Mix, soilless potting medium (Premier Peat, Riviere-du-Loup, Quebec, Canada) in a 72-well Styrofoam tray (Speedling, Inc., Bushnell, FL). Upon germination, seedlings were individually transplanted to 21.6 cm diameter round plastic pots containing Pro-Mix.
Each TEV strain and a mock-inoculated control were inoculated onto ten plants of each test species when the plants were at the 4–5 leaf stage. Virus was applied by mechanical inoculation of the two oldest leaves (leaves 1 and 2 or sometimes leaves 2 and 3). Inoculum for each TEV strain consisted of systemically infected N. tabacum ‘Kentucky 14’ leaf tissue ground in 50 mM potassium phosphate buffer (pH 7.5). Nicotiana tabacum ‘Kentucky 14’ plants used as inoculum were of similar age and stage of disease development for each virus treatment with individual TEV strains being of similar titers when tested by enzyme-linked immunosorbent assay (ELISA). The mock-inoculation treatment (referred to as the healthy control) consisted of buffer alone.
Plants were evaluated for development of systemic symptoms and tested for virus in young non-inoculated leaves by a commercial TEV-specific ELISA kit according to the manufacturer’s instructions (Agdia, Inc., Elkhart, IN) with modifications [20].
A more detailed evaluation was performed for each TEV strain (comparatively with the healthy control) in N. tabacum ‘Kentucky 14’ plants. Plants were grown and virus inoculations performed as described above. At 24 days post-inoculation (dpi), each leaf on six plants in each treatment was excised from the plant and individually weighed. Stem length and weight were determined after removal of all leaves.
Statistical comparisons were made among treatments (HAT, Mex21, N, and mock-inoculated plants) using t test in SAS 9.3 (SAS Institute Inc, Cary, NC, USA) with differences of means identified at P ≤ 0.05 using LSD mean separation tests.
Sequencing procedures
Each TEV strain was separately propagated in N. tabacum ‘Kentucky 14’ plants, from which systemically infected leaf tissue was harvested and stored at 4 °C until processed. Virus was purified as described previously [5, 21] and viral RNA was isolated using a RNeasy Plant Kit (Qiagen, Germantown, MD) according to manufacturer’s instructions. At the final stage of the virus purification procedure, virus was pelleted by centrifugation at 205,800×g for 1 h at 4 °C. The pelleted virus was resuspended in 450 µl of RLT extraction solution (provided in the RNeasy kit) with the remainder of the isolation as recommended by the manufacturer. The elution of viral RNA from the purification column was carried out using 50 µl of nuclease-free water (provided with the RNeasy kit) and the RNAs were stored at −80 °C until used for sequencing.
The primers were designed using Primer 3 version 0.4.0 (http://frodo.wi.mit.edu/primer3/) and analyzed using the oligoanalyzer tool provided by IDT (http://www.idtdna.com/analyzer/Applications/OligoAnalyzer/). The complete list of primers is available upon request. The primers were designed to sequence both DNA strains of each PCR amplified fragment. Purified viral RNA (100 ng) was used as a template to synthesize cDNA using SuperScript III Reverse Transcriptase (Invitrogen, Carlsbad, CA), with addition of 0.1 µM of reverse primer TEV-VPg Rv and 400 µM dNTPs. PCR reactions were made with 2 µl of cDNA added to a cocktail containing 1× HF buffer (as described by Finnzymes, Espoo, Finland), 2 mM MgCl2, 0.2 mM dNTP mix, 0.5 μM TEV-VPg Fw primer, 0.5 μM TEV-VPg Rv primer, and 1 U of Phusion DNA Polymerase (Finnzymes). Reactions were carried out using a Multigene Gradient thermal cycler (Labnet International Inc., Edison, NJ), with initial denaturation at 94 °C for 2 min, followed by 30 cycles of 94 °C for 60 s, 50 °C for 30 s, and 72 °C for 60 s, and a final extension step at 72 °C for 10 min.
Reverse transcription-PCR (RT-PCR) products were sequenced by Lucigen Corporation (Middletown, WI). The identity of the nucleotide sequences was assessed using the basic local alignment search tool (BLASTN) from the NCBI web site [22], then translated to amino acid sequence, and compared using the multiple alignment program ClustalW [23] through BioEdit (v 7.0.9). A further analysis to determine conserved amino acids and regions was made with Jalview [24].
Rapid amplification of 5′ cDNA ends (5′ RACE)
The 5′-terminus sequences for HAT-AU, Mex21, and N RNAs were determined using the 5′ RACE System (Rapid Amplification of cDNA Ends), version 2.0 (Invitrogen), according to the manufacturer’s instructions. To purified virus preparations (250 µl), a 10 % sodium dodecyl sulfate solution was added to achieve 0.5 % and incubated 3 min at 37 °C. Proteinase K was added to a concentration of 0.5 µg/ml and allowed to incubate at 37 °C for 23 min. To this mixture, 200 µl of chloroform and 300 µl of Tris-saturated phenol were added, mixed vigorously by vortex, and centrifuged at 10,000 rpm for 5 min in a Sorvall microfuge. The viral RNA was precipitated with sodium acetate, pH 6.0 (0.1 M final concentration) and 2.5 volumes of absolute ethanol. This procedure was used in order to remove most of the VPg from the 5′-terminus of the viral RNA [25].
The first strand cDNA to each virus genome was synthesized from approximately 5 μg of purified HAT-AU, Mex21, and N RNAs using primers HAT_GSP1 (5′-GAGTGGTGTGTAAGGCAAGTCT-3′), TEV_NGSP1 (5′-AGTGGCGTGTAAGGTAAGTCTG-3′), and TEV_NGSP1, respectively. Purified and dC-tailed HAT-AU, Mex21, and N-specific cDNAs were amplified by PCR using nested primers HAT_GSP2 (5′-CCTCGGTAACGTAGTCTTCTC-3′), TEV_NGSP2 (5′-TCAGTAACGTAGTCTTCTCCGA-3′), and TEV_NGSP2, respectively, and a dC-tail-specific abridged anchor primer. PCR was performed using Taq DNA polymerase (Finnzymes) for 35 cycles using the following thermal cycling conditions: 94 °C for 2 min, then 35 cycles of 94 °C for 30 s, 55 °C for 30 s, 72 °C for 1 min, and a final extension at 72 °C for 5 min. PCR products were column purified (Omega Bio-Tek, Norcross, GA) and sequenced at the Auburn University Sequencing Facility using respective virus-specific nested primers. The 5′-terminus sequences for HAT-AU, Mex21, and N were then assembled to their respective sequences to complete the full-length genome sequence for each TEV strain.
Phylogenetic comparison of TEV strains
The complete genome sequences of TEV strains SD1, Shannxi, TEV7DA, HAT, TEVGEN, and TEVNWP, as well as Potato virus Y (PVY) strain N:O (PVY N:O) (accession numbers EF470242, JN711120, DQ986288, M15235, M15239, L3871, and EF026076, respectively) were collected from the Genbank database. The genome sequences were aligned using ClustalW algorithm in MEGA5.05 [26]. The PVY N:O sequence was used as an outlier. The evolutionary history of those strains was inferred using the Maximum Likelihood method based on the Tamura-Nei model [27].
Results
Infection of Nicotiana spp.
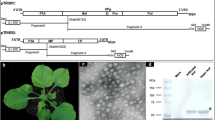
The disease symptoms varied among Nicotiana species and TEV strain. In N. tabacum ‘Kentucky 14’, all plants in each virus treatment developed systemic vein-clearing followed by different degrees of mosaic. HAT-AU-infected plants developed clearly apparent mosaic symptoms (Fig. 1a) and, although all newly emerging leaves continued to express mosaic symptoms, the plants were not overtly stunted compared with healthy control plants. Mex21-infected plants developed a more severe mosaic symptom (Fig. 1a) that was blotchy with dark green, puckered islands. Older infected leaves had extensive etching symptoms. These plants were smaller than healthy control plants. Nicotiana tabacum ‘Kentucky 14’ plants infected with N had a severe chlorotic/mosaic symptom with increased severity in newly emerged leaves (Fig. 1a). These plants were severely stunted with small, deformed leaves in the upper portion of the plant.
Each of the TEV strains induced a mild systemic vein-clearing in N. clevelandii plants. The vegetative growth of HAT-AU-infected plants was similar in size (leaf number and leaf size) to that of healthy control plants; however, floral stems tended to be slightly shorter but more numerous than those of healthy control plants (Fig. 1b). Nicotiana clevelandii plants infected with Mex21 had reduced vegetative growth (fewer and smaller leaves) and smaller floral stems that were also fewer in number than occurred for healthy control plants (Fig. 1b). N-infected plants had extensive chlorosis on vegetative leaves that eventually became necrotic. Vegetative growth for such plants was much less than for the other treatments and, of particular importance, floral stems did not develop, instead new, small leaves were bunched together at the central stem with (for some plants) a single flower (Fig. 1b). ELISA analysis of non-inoculated leaves indicated the presence of each virus in all inoculated plants.
Nicotiana tabacum ‘Samsun’ plants infected with each of the TEV strains developed systemic vein-clearing symptoms followed by a chlorotic, mild mosaic symptom. Plants infected with Mex21 or N were more chlorotic than those infected with HAT-AU and were also smaller in size. Young emerging leaves of N-infected N. tabacum ‘Samsun’ plants were chlorotic and deformed; these plants were more stunted than plants infected with either of the other two viruses (data not shown). TEV strain infection in systemically infected leaves from all plants within each virus treatment was confirmed by ELISA.
None of the N. glutinosa plants inoculated with each of the three different TEV strains developed visible symptoms. ELISA analysis of non-inoculated leaves for virus infection, however, indicated that all plants within each virus treatment were infected.
Effects on N. tabacum ‘Kentucky 14’
The effects on N. tabacum ‘Kentucky 14’ growth and development were evaluated relative to infection by each TEV strain in comparison with mock-inoculated control plants. The length of the stem, from soil-line to apical tip taken at 24 dpi, did not differ significantly for healthy control and HAT-AU treatments (mean values of 24 and 21 cm, respectively); however, plants in the Mex21 treatment had significantly shorter stems (15 cm) (Fig. 2). The stem length of plants infected with N was significantly shorter than all other treatments (11 cm).
Bar graphs representing growth parameters for Nicotiana tabacum cv ‘Kentucky 14’ plants inoculated with Tobacco etch virus (TEV) strains HAT-AU, Mex21, and N as well as a healthy control treatment. Growth parameters included stem length, a measure taken from soil-line to apical tip (left graph) and stem weight, a measure of the aboveground stem taken after removal of all leaves (right graph). Statistical comparisons were made using SAS among treatments within each respective growth parameter. Different letters represent a significant difference of the means at P ≤ 0.05 using LSD mean separation tests
The weight of stems, with all leaves excised, did not differ significantly for healthy control and HAT-AU treatments (33 and 28 g, respectively) but was significantly less for plants in the Mex21 and N treatments (14 and 9 g, respectively) (Fig. 2).
At 24 dpi, each leaf for plants within a treatment was excised from the stem and its fresh weight measured (Fig. 3). Measurements were initiated at leaf 4 (leaf 1 being the oldest leaf along the stem) because leaves 1 through 3 were often dead and/or missing for plants in the virus treatments. Each TEV strain was inoculated onto leaves 1 and 2 (sometimes 2 and 3) when plants were at the 5 leaf stage of development. The first leaf to express systemic symptoms was leaf 6; leaves 6, 7, and sometimes 8 expressed systemic vein-clearing symptoms. All subsequent leaves expressed varied degrees of chlorosis, mosaic or combinations of both and, if present, leaf deformation. The weight of leaf 4 did not differ significantly among treatments but leaf 5 (which remained symptomless) weighed significantly more for healthy control and HAT-AU treatments than for Mex21 and N treatments (Fig. 3). The first symptomatic leaf (leaf 6) weighed significantly less for all virus treatments than for the healthy control plants and leaf 6 for the HAT-AU treatment weighed significantly more than that for Mex21 and N treatments. Leaves 7, 8, and 9 had the greatest differences among treatments with leaves from the healthy control having significantly greater weight, followed by plants in the HAT-AU treatment which was significantly greater than those for Mex21 and N treatments. The weight of leaves 7, 8, and 9 for plants in the Mex21treatment was significantly greater than those for N-infected plants. The weight of leaf 10 was significantly greater for healthy control and HAT-AU treatments than for Mex21 and N treatments and, leaf 10 for Mex21-infected plants had a significantly greater weight than for N-infected plants. Differences among treatments lessened or did not occur for younger leaves, although leaves from healthy control plants tended to have significantly greater weights than those in the N treatment. In summary, at 24 dpi, all systemically infected leaves that experienced a degree of expansion, e.g., leaves 6 through 9, revealed significant differences in weight among TEV strain treatments. Lesser differences occurred in younger leaves; however, this likely reflected leaf age at 24 dpi and, if allowed more time to expand, continued differences in leaf weight among virus treatments would have occurred.
Line graph presenting weight (g) of individual Nicotiana tabacum cv ‘Kentucky 14’ leaves along the stem (denoted along the x-axis) of plants inoculated with Tobacco etch virus (TEV) strains HAT-AU, Mex21 or N, as well as a healthy control treatment. The rectangular box enclosing data for leaf 6 identifies the first leaf to express symptoms. Statistical comparisons were made using SAS among treatments within each respective leaf number along the stem. Different letters represent a significant difference of the means at P ≤ 0.05 using LSD mean separation tests
TEV strain sequence analysis
Nucleotide and amino acid sequence comparisons included the original HAT sequence [14; GenBank Accession no. M11458], HAT-AU, Mex21, and N. The complete genome sequences of HAT-AU, Mex21, and N were deposited into the GenBank database under the accession numbers KM282187, KM282188, and KM282189, respectively. The complete genome of HAT-AU, Mex21, and N were sequenced and shown to consist of 9,495 nucleotides, not including the poly (A) tail, and a predicted polyprotein of 3,054 amino acids. Comparison of their nucleotide sequences relative to the original HAT sequence [14] revealed 95, 92, and 92 % identity for HAT-AU, Mex21, and N, respectively (Table 1). HAT-AU had 91 % sequence identity with Mex21 and N, while Mex21 and N were more closely related with 98 % nucleotide sequence identity. Similarly, the amino acid sequence identities for the full-length polyprotein ranged from 95 % for HAT-AU when compared with N to a high of 98 % identity between Mex21 and N (Table 2). These data indicate that Mex21 and N are more closely related than relationships among the other viruses used in this study, this includes the relationship between the original HAT and HAT-AU.
The mature polyprotein is processed into individual proteins by proteinases that reside within the polyprotein. The cleavage sequences recognized by these proteinases and the specific cleavage sites have been identified for TEV [28]. Six of the nine recognized cleavage sequences within the polyprotein amino acid sequence were identical among the four TEV strains (Fig. 4). Three of the cleavage sequences varied, all of which were associated with either 6K1 or 6K2 proteins, and the variation occurred in the first or second amino acid of the six amino acid cleavage sequence (Fig. 4). The exact cleavage site within the amino acid cleavage sequence was identical for all cleavage sites among the TEV strains.
The Tobacco etch virus (TEV) genome organization for strains TEV-HAT (original HAT strain sequenced) [14], HAT-AU (HAT strain maintained at Auburn University), Mex21, and N. Individual protein coding regions are above each respective box with the size of each protein indicated within each box (with the exceptions of the 6 K proteins which are below each box). Individual proteins are autocatalytically cleaved from the polyprotein by proteinases within the polyprotein. The six amino acid cleavage sequences are presented for each protein with the specific cleavage site identified with bold letters separated by a slash
Among the seven primary proteins (i.e., P1, HC-Pro, P3, CI, VPg, NIa-proteinase, NIb, and CP), amino acid sequence identities varied from a low of 86.8 % for the P1 proteins of HAT-AU and Mex21 to a high of 99.6 % for the CI proteins of Mex21 and N (Table 2). Overall, the lower amount of amino acid sequence identity occurred among TEV strains for the P1 protein with the highest percentage identity for the CI protein (Table 2).
Phylogenetic comparison of TEV strains
To determine the evolutionary relationships among the HAT-AU, Mex21, and N strains, a phylogenetic tree was constructed based on the complete nucleotide sequences of nine different TEV strains and an outlier virus in the genus Potyvirus, Potato virus Y (PVY) (Fig. 5). The nine TEV strains are grouped into three different clusters each consisting of three strains. The newly sequenced strains, Mex21 and N, are distantly related to HAT-AU. Mex21 formed a monophyletic group with TEVNWP; these two strains are more closely related to each other than to N and HAT. N is more closely related to Mex21 than HAT, which is highly divergent and part of a more distant cluster. HAT and TEVGEN are clonal and shared a more recent common ancestor with strain TEV7DA, which are all clustered together in a single clade.
Phylogenetic analysis based on the full-length genome sequences of nine Tobacco etch virus (TEV) strains. The sequence of Potato virus Y (PVY) was used as an outlier. The evolutionary history of these viruses was inferred using the Maximum Likelihood method based on the Tamura-Nei model [27]. The branch lengths represent a measure of the number of base substitutions per site (the tree is dawn to scale). All positions in the sequence alignment containing gaps and missing data were eliminated. There were a total of 7,953 positions in the final data set. Evolutionary analyses were conducted in MEGA5.05 with 1,000 repetitions to generate bootstrap support [26]. The nodes of the phylogenetic tree were supported with strong bootstrap values. The sequence accession number of each virus is provided in brackets to the right of the respective virus
Discussion
This report clearly illustrates the differences in symptom type and severity between HAT-AU, Mex21, and N in the different Nicotiana spp. Nicotiana tabacum ‘Kentucky 14’ (a Burley type tobacco variety) and N. clevelandii expressed obvious disease symptoms in response to infection by the TEV strains with increasing severity from HAT-AU (mild) to Mex21 (moderately severe) to N (severe). The initial expression of systemic infection in N. tabacum ‘Kentucky 14’ plants was similar among the TEV strains with regard to the time required for symptoms to develop and type of symptom which consisted of systemic vein-clearing in young leaves. Upon establishment of the systemic infection, however, subsequent symptom type, severity, and effects on plant growth and development differed. Newly emerged leaves of HAT-AU-infected plants expressed mosaic symptoms but with little, if any, apparent effects on plant size. The systemic mosaic symptom induced by Mex21 was more pronounced than that of HAT-AU with reduced leaf size and internode extension resulting in stunted growth. The systemically infected newly emerging leaves of N-infected N. tabacum ‘Kentucky 14’ plants were extremely chlorotic, deformed, and small. Internode extension in the upper portions of the stem of N-infected plants was significantly reduced resulting in development of small, chlorotic leaves bunched together at the top of the plant. Despite these differences in disease severity, relative amounts of virus accumulation in systemically infected leaves was similar when tested using a quantitative ELISA test (data not shown). The lack of correlation between symptom severity and virus titer is not uncommon, as shown for Barley stripe mosaic virus [29], Cucumber mosaic virus [CMV; 30], and in a study evaluating the fitness and virulence effects on TEV variants [31].
The genetic relationship among the TEV strains is quite close, as might be expected since they are strains of the same species. It is not surprising, perhaps, that Mex21 and N are in different clades than HAT-AU based on differences in their pathogenicity in tobacco plants. It was surprising, however, that HAT-AU and the original HAT fell into different clades. The HAT isolate originally sequenced by Allison et al. [14] and HAT-AU was both obtained from the same source (i.e., Dr. Thomas Pirone, University of Kentucky). The differences observed in the HAT and HAT-AU genomic sequences are likely the result of years of passage of HAT-AU through pepper and tobacco hosts [Murphy, unpublished].
This study illustrates that the degree of subtle differences in genetic code can lead to significant differences in disease phenotype. The rate of systemic movement into young developing tissues was relatively similar among the TEV strains, as were the levels of accumulation in systemically infected leaves, suggesting that these factors may not account for the differing effects on host plant physiological processes leading to the different disease phenotypes. The induced disease symptoms for HAT-AU, Mex21, and N vary in type and severity which might suggest a more complicated virus-host interaction to dissect from one strain to another. The availability of the complete genome sequence for each of the TEV strains, however, should provide the needed foundational data to determine the viral determinant(s) responsible for the differences in TEV strain pathogenicity.
References
F. Rabenstein, M.J. Adams, R. French, J.F. Kreuze, J.J. Lopez-Moya, K. Ohshima, D.C. Stenger, A. Wang, S. Wylie, F.M. Zerbini, Virus Taxonomy. 9th Report of the International Committee on Taxonomy of Viruses, ed. by A.M.Q. King, M.J. Adams, E.B. Carstens, and E.J. Lefkowitz (Elsevier, Amsterdam, 2011), p. 1069
S. Urcuqui-Inchima, A.-L. Haenni, F. Bernardi, Virus Res. 74, 157–175 (2001)
B.Y.-W. Chung, W.A. Miller, J.F. Atkins, A.E. Firth, Proc. Natl. Acad. Sci. USA 105, 5897–5902 (2008)
R.G. Christie, J.R. Edwardson, Fla. Agric. Exp. Stn. Monogr. Ser. no. 9 (1977)
J.F. Murphy, R.E. Rhoads, A.G. Hunt, J.G. Shaw, Virology 178, 285–288 (1990)
V. Hari, A. Siegel, C. Rozek, W.E. Timberlake, Virology 92, 568–571 (1979)
CABI, 2010. Distribution maps of plant diseases. http://www.cabi.org/dmpd/default.aspx?site=165&page=4050&LoadModule=Review&ReviewID=151422
D.D. Shukla, C.W. Ward, A.A. Brunt, Potyviruses: Biology, Molecular Structure and Taxonomy, CAB Int., Wallingford, England (1994)
J.R. Edwardson, R.G. Christie, Viruses Infecting Peppers and Other Solanaceous Crops, vols. I and II. (University of Florida. Gainesville, FL, 1997)
R.C. Rufty, R.O. Miller, G.V. Gooding Jr., Plant Dis. 73, 45–48 (1989)
P. Bertrand, Georgia Tobacco Research-Extension Report, University of Georgia Cooperative Research-Extension Publication number 1-2003, pp 105-116 (2004)
W.C. Wilkinson, L.R. Fisher, W.D. Smith, D.L. Jordan, Tobacco Sci. 47, 44–52 (2007/2008)
L. Zhao, J. Cheng, X. Hao, X. Tian, Y. Wu, Arch. Virol. 157, 2291–2298 (2012)
R. Allison, R.E. Johnston, W.G. Dougherty, Virology 154, 9–20 (1986)
B.-C. Kang, I. Yeam, J.D. Frantz, J.F. Murphy, M.M. Jahn, Plant J. 42, 392–405 (2005)
I. Yeam, J.R. Cavatorta, D.R. Ripoll, B.-C. Kang, M.M. Jahn, Plant Cell 19, 2913–2928 (2007)
K. Perez, I. Yeam, B.-C. Kang, D.R. Ripoll, J. Kim, J.F. Murphy, M.M. Jahn, Mol. Plant–Microbe Interact. 25, 1562–1573 (2012)
C.M. Deom, J.F. Murphy, O. Pagiuo, Mol. Plant–Microbe Interact. 10, 917–921 (1997)
J.F. Murphy, J.R. Blauth, K.D. Livingstone, V. Lackney, M.K. Jahn, Mol. Plant–Microbe Interact. 11, 943–951 (1998)
N. Valesquez, J.F. Murphy, Plant. Pathol. 63, 675–683 (2014)
M.N. Guerini, J.F. Murphy, J. Gen. Virol. 80, 2785–2793 (1999)
S.F. Altschul, W. Gish, W. Miller, E.W. Myers, D.J. Lipman, J. Mol. Biol. 215, 403–410 (1990)
J.D. Thompson, D.G. Higgins, T.J. Gibson, Nucleic Acids Res. 22, 4673–4680 (1994)
A.M. Waterhouse, J.B. Procter, D.M.A. Martin, M. Clamp, G.J. Barton, Bioinformatics 25, 1189–1191 (2009)
J.F. Murphy, W. Rychlik, R.E. Rhoads, A.G. Hunt, J.G. Shaw, J. Virol. 65, 511–513 (1991)
K. Tamura, D. Peterson, N. Peterson, G. Stecher, M. Nei, S. Kumar, Mol. Biol. Evol. 28, 2731–2739 (2011). doi:10.1093/molbev/msr121
K. Tamura, M. Nei, Mol. Biol. Evol. 10, 512–526 (1993)
M.J. Adams, J.F. Antoniw, F. Beaudoin, Mol. Plant Pathol. 6, 471–487 (2005)
A.D. Stewart, J.M. Logsdon Jr., S.E. Kelly, Evolution 59, 730–739 (2005)
F. Escriu, A. Fraile, F. Garcia-Arenal, Phytopathology 90, 480–485 (2000)
P. Carrasco, F. de la Iglesia, S.F. Elena, J. Virol. 81, 12979–12984 (2007)
Acknowledgments
The authors wish to express their gratitude to the Auburn University Plant Science Greenhouse Facility personnel and funding provided by the Alabama Agriculture Experiment Station and the Department of Entomology and Plant Pathology, Auburn University, AL, U.S.A.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Velasquez, N., Hossain, M.J. & Murphy, J.F. Differential disease symptoms and full-length genome sequence analysis for three strains of Tobacco etch virus . Virus Genes 50, 442–449 (2015). https://doi.org/10.1007/s11262-014-1146-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11262-014-1146-9