Abstract
Four amino acid Schiff base complexes—one with a 3D network structure ({[Cd2(PYSA)2(H2O)]·3H2O}n (1)) and the other three with a supramolecular network structure {[Co(PYSA) (H2O)2]·H2O}n (2), {([Zn(PYSA)(H2O)2]·H2O}n (3), {[Mn(PYSA)(H2O)2]·H2O}n (4)), (PYSA = (S-2-(pyridine-4-ylmethylamino) succinic acid)—were successfully obtained under hydrothermal conditions. Among the four complexes, photoluminescence analysis revealed that complex 1 exhibited high selectivity and sensitivity in the detection of iron ions. In particular, it was able to qualitatively and quantitatively detect iron ions at low concentrations without interference from other ions. With a Fe3+ concentration of 0.01 mol/L, the detection limit was 0.81 μM, Ksv = 2.46 × 104 L/mol. In addition, complex 2 exhibited excellent catalytic performance toward 4-nitrophenol (4-PNP).
Graphic abstract
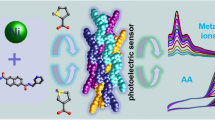
Four amino acid Schiff base complexes 1–4 were synthesized by a hydrothermal method. Among the four, complex 1 exhibited good fluorescence sensing performance in the detection of Fe3+ in water. In addition, the catalytic activity of complex 2 for 4-PNP was preliminarily investigated, which showed an ability to reduce 4-PNP to 4-PAP.

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Coordination complexes have attracted much attention because of their good magnetic [1,2,3,4,5,6,7], luminescence [8,9,10,11,12], adsorption [13,14,15,16,17,18,19], catalytic [20,21,22,23,24,25,26,27,28] and other properties [29, 30]. They have excellent features including large pore size, low density and very large specific surface area. Given the important role of chirality in nature, the synthesis of pure chiral complexes has garnered the interest of researchers. Recent studies have shown that the use of chiral ligands [31] for the construction of chiral complexes is a highly effective method.
Amino acids with both amino and carboxyl groups are an important class of organic compounds, as amino and carboxyl groups have good metal-chelating ability. However, most of the amino acids in proteins are α-amino acids, and they form weak complexes with metals. Therefore, we often use terminal amino acids to react with active aldehyde to form Schiff base carboxylic acid compounds containing carboxyl groups, such as pyridine and pyrazines, owning to their strong coordination ability. With this method, we can design and synthesize amino acid derivative ligands with multiple functional groups, and a series of novel and tunable complexes can be synthesized. A number of amino acid-modified framework structures have been synthesized through judicious assembly of appropriate metal ions (or clusters) and bi- or multifunctional ligands. In such materials, functionality can be introduced from either the inorganic vertexes or the organic links. Furthermore, the molecular-level integration of inorganic and organic links makes it possible to initiate or enhance desirable properties. Therefore, the selection or design of organic ligands containing predefined features such as electronic properties and versatile binding modes is crucial to the construction of metal organic framework complexes. In contrast to the large and growing number of works on the synthesis of complexes with common ligands containing O or N donors, to the best of our knowledge there have been only a few reported studies based on amino acid-modified Schiff base ligands. According to the literature [31, 32], the family of amino acid complexes exhibits good performance in terms of fluorescence, catalysis and adsorption. For instance, in 2003, Qu et al. [33] used ZnCl2 and CdCl2 as metal sources, reacted with (S)-3-cyanophenylalanine (s-tpa) and NaN3 under hydrothermal conditions, to obtain two 3D isophane networks with identical SrAl2 topological structure. In 2008, Ma et al. [34] reported octahedral complexes in water using Mn(OAc)2·4H2O as the source of metal, and H3L (H3L = N-[(3-carbonyl phenyl)-sulfonic group]-glycine) and bipy (bipy = 4, 4′-al pyridine) as the ligands. In 2010, Li et al. [35] used Zn(CH3CO2)2·2H2O as a metal source and N-(2-methyl-pyridine)-l-alanine (l-hpala) and N-(2-methypyridine)-d-alanine (d-hpala) as ligands to react in methanol solvent. They obtained corresponding complexes [Zn(l-pala)2]·3H2O and [Zn(d-pala)2]·3H2O, and carefully studied the structures of the complexes and their photoluminescence properties.
Because of its advantages including rapid, accurate and efficient sensing, fluorescence quenching technology has been developed as an effective method for the detection of pollutants [36,37,38,39]. Therefore, the design and synthesis of a series of complexes for fluorescence sensing is a research hotspot at present. Moreover, ions play an important role in our lives [40]. Iron ions as an essential trace element are an indispensable component in the human body. Iron is an important raw material for producing blood cells from bone marrow, and also plays a vital role in the transport of electrons in the respiratory chain. When the human body is deficient in iron, anemia and immune system impairment are likely to occur. On the other hand, too much iron will affect the function of the pancreas and increase the risk of heart failure, diabetes, liver cirrhosis and other diseases [41]. In recent years, the rapid development of industrial technology has led to severe issues with global water resources. Certain heavy metal ions and organic molecules have been found to be the main pollutants in water. An increase or decrease in iron(III) concentration can cause irreversible damage to the water environment [42, 43]. Iron(III) ions affect the metabolism of planktonic algae, and have shown ecological benefits for the growth of planktonic algae, chlorophyll synthesis, nitrogen and carbon fixation [44]. They are also important for environmental protection and human health. Therefore, reliable methods are needed for the qualitative, quantitative and selective detection of iron ions in water.
In this work, we continue to research and expand the new family of backbone compounds with amino modification and fluorescence detection functions. Amino acid-modified framework complexes 1–4 were synthesized by the reaction of an amino acid-modified Schiff base ligand, which was obtained by reacting aspartic acid and 4-pyridinaldehyde, and Cd(NO3)2·4H2O, Co(NO3)2·6H2O, Zn(NO3)2·6H2O as well as Mn(CH3COO)2·4H2O, respectively. We also studied the fluorescence properties of complex 1 with regard to the qualitative and selective detection of Fe3+.
Experimental
Materials and methods
All chemicals purchased commercially were of reagent grade and were used without further purification. Ligand was synthesized according to a previously described method [31, 45]. Elemental analysis for C, H and N was conducted on a PerkinElmer 240C automatic analyzer. All IR measurements were obtained using a Bruker AXS TENSOR-27 FT-IR spectrometer with pressed KBr pellets in the range of 400–4000 cm−1 at room temperature. UV–visible–near-infrared (NIR) spectra for complexes 1–4 were recorded on a JASCO V-570 UV-visible/NIR microspectrophotometer (200–2500 nm, in the form of solid samples). X-ray powder diffraction patterns were obtained with a Bruker D8 Advance equipped with Cu-Kα radiation in the range of 5° < 2θ < 55°, with a step size of 0.02° (2θ) and a count time of 2 s/step. Thermogravimetric analysis (TG) was performed on a PerkinElmer Diamond Thermogravimetric/Differential Thermal Analyzer (TG/DTA) under a N2 atmosphere over a temperature range of 30–800 °C. The photoluminescence spectra of the coordination polymers were measured on a HORIBA FluoroMax-4 spectrofluorometer with a time-correlated single-photon counting (TCSPC) accessory and equipped with a SpectraLED pulsed LED source at room temperature (200–1000 nm).
Synthesis of (S-2-(pyridine-4-ylmethylamino) succinic acid hydrochloride (PYSA·HCl) [31, 45]
For the synthesis of PYSA·HCl, l-aspartic acid (6.7 g 0.05 mmol) and NaOH (4 g, 0.1 mmol) were stirred in 20 mL deionized water and then poured into a 250-mL three-necked flask and stirred for 30 min. 4-Pyridinecarboxaldehyde (5.4 g, 0.05 mmol) was slowly added dropwise to the mixture under ice-water conditions and stirred for 2 h. NaBH4 (1.9 g, 0.5 mol) was dissolved in 10 mL water, and the solution was added dropwise to the mixture, with stirring, in an ice-water bath. After 3 h of stirring, the pH was adjusted to 3–4 with 6 mol/L HCl. The resulting solid was filtered off, washed with 50 mL hot methanol and then filtered. The solution was collected and concentrated on a rotary evaporator to give a white solid, which was named PYSA·HCl. The specific synthetic route is shown in Scheme 1 [IR (KBr, cm−1): 3445, 3052, 3001, 2983, 2858, 1715, 1615, 1429, 1373, 1323, 1168, 1102, 1060].
Synthesis of the complexes
{[Cd2(PYSA)2(H2O)]·3H2O}n (1)
A mixture of Cd(NO3)2·4H2O (77.1 mg, 0.25 mmol) and PYSA·HCl (13 mg, 0.05 mmol) was added to a mixed solvent of MeOH (2 mL), H2O (2 mL) and NH4HCO3 (1 mol/L) and then heated for 4–6 h. Colorless crystals of 1 were obtained at 100 °C, which were filtered, washed with water and dried at room temperature (yield: 24% based on Cd (II)). Elemental analysis (%) calculated value for 1: C 32.20, N 7.50, H 3.76. Found: C 32.40, N 7.53, H 3.64. IR (KBr, cm−1): 3428, 3229, 3070, 2973, 2864, 1625, 1591, 1427, 1408, 1347, 1327, 1238, 1123, 1089.
{[Co(PYSA)(H2O)2]·H2O}n (2)
Co(NO3)2·6H2O (59.48 mg, 0.25 mmol) and PYSA·HCl (13 mg, 0.05 mmol) were heated at 100 °C in a mixed solvent of MeOH (2 mL), H2O (2 mL) and NH4HCO3 (1 mol/L, 0.2 mL). After 4–6 h, pink crystals of 2 were obtained, which were filtered, washed with H2O and dried at room temperature (yield: 53% based on Co (II)). Elemental analysis (%) calcd for 2: C 35.82, N 7.20, H 4.78. Found: C 35.14, N 7.38, H 4.98. IR (KBr, cm−1): 3426, 3216, 3081, 2895, 2857, 1625, 1562, 1435, 1415, 1389, 1320, 1222, 1112, 1086.
{[Zn(PYSA)(H2O)2]·H2O}n (3)
Heating a mixture of Zn(NO3)2·6H2O (74.25 mg, 0.25 mmol) and PYSA·HCl (13 mg, 0.05 mmol) in a mixed solvent of MeOH (2 mL), H2O (2 mL) and NH4HCO3 (1 mol/L, 0.2 mL) at 100 °C for 4–6 h, we achieved colorless crystals of complex 3, which were filtered, washed with H2O and dried at room temperature (yield: 46% based on Zn (II)). Elemental analysis (%) calcd for 3: C 35.19, N 7.40, H 4.60. Found: C 35.70, N 7.50, H 4.62. IR (KBr, cm−1): 3383, 3285, 3092, 2918, 2859, 1614, 1585, 1429, 1409, 1337, 1307, 1169, 1113, 1070.
{[Mn(PYSA)(H2O)2]·H2O}n (4)
In addition to changing M(NO3)2·6H2O (M = Cd2+, Co2+, Zn2+) to Mn(CH3COO)2·4H2O, the synthesis method for complex 4 was similar to those for complexes 1, 2 and 3. A colorless crystal of complex 4 was obtained (yield: 70% based on Mn (II)). Elemental analysis (%) calcd for 4: C 36.25, N 8.50, H 4.80. Found: C 36.19, N 8.71, H 4.75. IR (KBr, cm−1): 3373, 3271, 3020, 2916, 2893, 1612, 1573, 1423, 1403, 1361, 1327, 1225, 1116, 1028.
X-ray crystal structure determination
Suitable single crystals of the four complexes were mounted on glass fibers for X-ray measurement. Reflection data were collected at room temperature on a Bruker AXS SMART APEX II CCD diffractometer with graphite-monochromatized Mo-Kα radiation (λ = 0.71073 Å). All the measured independent reflections (I > 2σ (I)) were used in the structural analyses, and semi-empirical absorption corrections were applied using the SADABS program [46]. The structures were determined by direct methods using the SHELX-97 program [47, 48]. All non-hydrogen atoms were refined anisotropically. The hydrogen atoms of the organic frameworks were geometrically fixed at calculated positions and refined using a riding model. The crystallographic data and the structure refinement details for complexes 1–4 are given in Table S1. Selected bond lengths are listed in Table S2. The OMIT commands in complex 1 were utilized to remove poor reflections affected by the beam stop. Structures were checked for additional symmetry using PLATON [49].
Results and discussion
Structural analysis
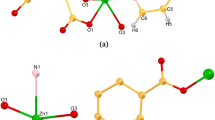
{[Cd2(PYSA)2(H2O)]·3H2O}n (1). Complex 1 crystallized in the orthorhombic space group P212121 with one molecule of 1 in the asymmetric unit. As shown in Fig. 1a, the asymmetric unit contains two cadmium atoms, two PYSA ligands, one water ligand and three lattice H2O molecules. Two cadmium atoms in the structural unit have two coordination modes. The central metal Cd1 is coordinated with two nitrogen atoms (N1, N2) and four oxygen atoms (O1, O2, O3, O9) from the ligand to form a twisted octahedral configuration. The other metal Cd2 is coordinated with two nitrogen atoms (N3, N4) and three oxygen atoms (O4, O6, O7) from the ligand and one oxygen atom (O5) from the coordinated water ligand. The ligand is connected to four Cd atoms, and each carboxyl oxygen atom is in a monodentate and chelating coordination. Nitrogen atoms of amino acid moiety also participate in monodentate coordination, as shown in Fig. 1b. The coordination modes of the entire ligand are described as μ4-ηO1ηO1ηN1ηO2ηN1. The lengths of the Cd1–O coordinate bonds are in the range of 2.2121(19)–2.485(2) Å, and the lengths of the Cd2–O coordinate bonds are 2.2753(19)–2.3878(19) Å. The lengths of the Cd1–N coordinate bonds are 2.346(2)–2.394(2) Å, and the lengths of the Cd2–N bonds are 2.346(2)–2.395(2) Å, which is similar to those reported recently [31].
In Fig. 1c, each Cd atom is connected by an oxygen atom from the ligand to form a 1D chain structure in complex 1. And then, as is shown in Fig. 1d, the 1D chain forms a 2D planar structure through a carboxyl oxygen atom. Finally, a 3D network structure is formed by oxygen atoms and nitrogen atoms which are from the ligand (Fig. 1e).
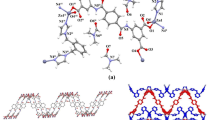
{[Co(PYSA)(H2O)2]·H2O}n (2). Single-crystal X-ray diffraction data show that complex 2 crystallized in a tetragonal P4(3) space group. In the asymmetric unit (Fig. 2a), complex 2 consists of one cobalt atom, one ligand PYSA, two water ligands and one lattice H2O molecule. The cobalt atom center is found in a six-coordinate geometry, bound by two oxygen atoms (O1, O3), two nitrogen atoms (N1#1, N2) from the PYSA ligand and two oxygen atoms (O5, O6) from the coordinated water ligand, which constitutes an octahedral configuration. The lengths of the Co–O bonds are in the range of 2.0736(18)–2.1689(19) Å, and the lengths of the Co–N bonds are 2.1131 (15)–2.1792(16) Å. These are similar to the bond length range of Co–O and Co–N bonds reported previously [31]. Also, as shown in Fig. 2a, the coordination modes of ligand PYSA can be described as μ2-ηO1ηO1ηN1ηN1. The oxygen and nitrogen atoms of each PYSA2− ligand adopt a monodentate coordination mode to connect adjacent Co2+ cations.
In the stacked structure, neighboring Co atoms are connected by one PYSA ligand to form a one-dimensional chain in two different directions along the a-axis and b-axis (Fig. 2b). In addition, the 1D chain structure of complex 2 forms a 2D structure connected by the hydrogen bonds, which are formed by the hydrogen atoms from the coordinated water molecule and carboxyl oxygen in the ligand (Fig. 2c). Figure 2d shows how the 2D planar structure is transformed into a 3D supramolecular network structure through hydrogen bonds. Details of the hydrogen bonds are listed in Table S4.
{[Zn(PYSA)(H2O)2]·H2O}n (3). Single-crystal X-ray diffraction data show that complex 3 is the same as complex 2. It is also found in a tetragonal P4(3) space group. In Fig. 3a, the asymmetric structural unit contains one zinc atom, one PYSA ligand, two water ligands and one lattice H2O molecule. The central metal atom Zn (II) is coordinated by two N atoms (N1#1, N2), two oxygen atoms (O2, O3) from the carboxyl group of the ligand PYSA, and two oxygen atoms (O5, O6) from coordinate water molecules to form a six-coordinated octahedron configuration. The ligand PYSA connection mode is μ2-ηO1ηO1ηN1ηN1. One end of the ligand links a metal zinc atom through a nitrogen atom on the pyridine ring, and the other end links the other zinc atom with the chelated form via two oxygen atoms of two carboxyl groups and a nitrogen atom from the ligand. The lengths of the Zn–O coordinate bonds are 2.095(3)–2.169(3) Å, and the lengths of the Zn–N coordinate bonds are in the range of 2.088(2)–2.189(3) Å. These are similar to the range of bond lengths reported in the literature for Zn–O bonds (2.082(2)–2.163(3) Å) and Zn–N bonds (2.084(2)–2.174(3) Å) [31].
In the stacked structure, the adjacent zinc atom is connected by the carboxyl group and the pyridine nitrogen atom in the ligand to form a 1D chain (Fig. 3b). Interestingly, the 1D chain is linked to form a 2D planar structure by the intermolecular hydrogen bonding O5–H5B···O4#7 and O6–H6A···O4#7 in the complex structure (Fig. 3c). The 2D planar structure of the complex then forms a 3D supramolecular network structure connected by hydrogen bonds (Fig. 3d). A detailed list of the hydrogen bonds is given in Table S4.
{[Mn(PYSA)(H2O)2]·H2O}n (4). Single-crystal X-ray diffraction data show that complex 4 is also found in the tetragonal P4(3) space group. The asymmetric unit contains one manganese atom, one ligand PYSA, two water ligands and one lattice H2O molecule (Fig. 4a). A central metal atom Mn, the two oxygen atoms (O4, O5), two nitrogen atoms (N1, N2) from the ligand PYSA, and two oxygen atoms (O1, O2) from the water ligands form a six-coordinated octahedral configuration. The lengths of the Mn–O coordinate bonds are in the range of 2.1750(19)–2.2139(19) Å, and the lengths of the Mn–N bonds are 2.2082(16)–2.2903(17) Å. Fig. 4a shows the ligand coordination modes, which can be described as μ2-ηO1ηO1ηN1ηN1, which is the same as that observed in complexes 2 and 3. In particular, the nitrogen atom on the pyridine ring adopts a monodentate coordination mode, while the oxygen atom from the carboxyl group and the nitrogen atom from the Schiff base moiety(C=N) adopt a monodentate chelation coordination mode.
In the stacked structure, the central metal atom Mn (II) connects the ligand to form a 1D chain structure (Fig. 4b). The 1D chain structure is formed by the hydrogen bond O1–H1C···O6#4 and O2–H2C···O6#4 along the ac plane direction to form a 2D layered structure (Fig. 4c). The 2D structure is transformed into a 3D supramolecular network structure through other hydrogen bonds. as shown in Fig. 4d. The hydrogen bonds are listed in detail in Table S4.
Comparison of the structures
Comparing the structures of complexes 1–4, it is found that although the ligand is identical and the metals in complexes 2, 3 and 4 show the same coordination mode and coordination number, complex 1 is obviously different from the other three coordination modes and structures. Specific structural comparisons are listed in Table S3. The connection mode of the ligand PYSA is μ2-ηO1ηO1ηN1ηN1 in complexes 2, 3 and 4, while for PYSA in complex 1 it is μ4-ηO1ηO1ηN1ηO2ηN1. On the other hand, the lengths of the M–N/O bonds of complexes 2, 3 and 4 are basically the same, possibly due to the same coordination number and the coordination pattern around the metal atoms. In addition, complexes 2, 3 and 4 connect one-dimensional chains through hydrogen bonds to form a two-dimensional planar structure, which is then transformed into a three-dimensional supramolecular network structure through the hydrogen bonds. Complex 1 is gradually transformed from 1D to 2D through the atoms in the ligand, eventually forming a 3D network structure. The structure of complexes 1, 2 and 3 is similar to that reported in the literature [31] in terms of the crystallographic parameters including crystallographic data, bond length, bond angle and coordination number.
Spectral properties
IR spectra
In the IR spectrum of complex 1, the peak at 3428 cm−1 is attributed to the characteristic peak of \(\nu_{{{\text{OH}}^{ - } }}\) in water. The characteristic peaks of νN–H and νAr–H appear at 3229 cm−1 and 3070 cm−1. Also, the peaks at 1625 cm−1 and 1408 cm−1 are characteristic peaks of the stretching vibration of \(\nu_{{{\text{as}}({\text{COO}}^{{^{ - } }} )}}\) and \(\nu_{{{\text{s}}({\text{COO}}^{{^{ - } }} )}}\), respectively. A redshift is observed with respect to the stretching vibration peaks of \(\nu_{{{\text{as}}({\text{COO}}^{{^{ - } }} )}}\) of the ligand, indicating that the ligand PYSA (\(\nu_{{{\text{as}}({\text{COO}}^{{^{ - } }} )}}\): 1715 cm−1) is coordinated with the metal. The peaks at 1591 cm−1, 1427 cm−1, 1347 cm−1 and 1327 cm−1 are stretching vibrations of νC=C and νC=N. Because of the same ligand and the similarity of the crystal structure, only complex 1 is discussed in detail. Detailed analyses of the IR spectral data for complexes 1–4 and the ligand are shown in Table S5 and Fig. S1.
UV–Vis spectra
All UV–Vis absorption spectra of ligand (PYSA) and complexes 1–4 were recorded in the form of solid samples. Two high-energy absorption bands of ligand (PYSA) appeared at 213 nm and 258 nm, which were assigned to the π–π* and n–π* transition of the ligand (PYSA). Similar absorption bands in complexes 1–4 were observed at 219 nm and 262 nm for 1, 216 nm and 293 nm for 2, 219 nm and 264 nm for 3, and 260 nm and 324 nm for 4. The peaks observed at 400 nm for 1, 341 nm for 2, 343 nm for 3 and 381 nm for 4 were attributed to the charge transition from the ligand (PYSA) to Cd (II), Co (II), Zn (II) and Mn (II) atoms (LMCT). The bands appearing at 521 nm, 1182 nm and 1440 nm were attributed to the d–d*transition of the Co2+ center for complex 2. Detailed analyses of the UV–Vis absorption bands of complexes 1–4 and PYSA are given in Table S6 and Fig. S2.
Powder X-ray diffraction analysis
The phase purity of complexes 1–4 was investigated by the powder X-ray diffraction (PXRD) patterns in the 2θ range of 5°–55°. As shown in Figure S3, the diffraction peaks of the as-synthesized samples are almost coincident in position with the simulated patterns calculated from the single-crystal diffraction data, confirming that the as-synthesized samples have high phase purity.
Thermal analysis
To ascertain the thermal stability of complexes 1–4, TG analysis was carried out under a flowing nitrogen atmosphere. In the range of 30–230 °C, the weight loss was 8.06% (calcd: 9.70%) for 1, 15.9% (calcd: 16.12%) for 2, 14.67% (calcd: 15.84%) for 3 and 17.14% (calcd: 15.87%) for 4, corresponding to the removal of the coordinated and lattice water molecules, and the major structures of 1–4 then began to decompose after 320 °C, 360 °C, 330 °C and 320 °C, respectively. The TG curves are shown in detail in Fig. S5.
Photoluminescence properties and organic solvent sensing of complex 1
The solid-state emission spectra of complex 1 were recorded at room temperature. Upon excitation at 306 nm for 1, the maximum emission peak of the complex was observed at 440 nm (Fig. S6). For further study, the luminescence emission of complex 1 dispersed in different small molecule solvents including N,N-dimethylformamide (DMF), ethanol, methanol, water, acetonitrile, hexane and acetone was investigated at room temperature (Fig. 5). As shown in Fig. 5, the emission peaks of complex 1 exhibit different phenomena in different solvents because of the different polarities of the solvents (hexane < acetone < ethanol < methanol < DMF < acetonitrile < water). The emission peak is blueshifted in a solvent with low polarity (such as hexane) and redshifted in a solvent with high polarity (such as water). Hexane and DMF are toxic, so we chose water as the solvent to investigate the luminescence properties of the complex in water suspension. After immersion of complex 1 in water for 24 h and drying in vacuum, PXRD measurement showed that the complex was stable in water (Figure S4). Therefore, the strong emission of complex 1 in solid state and organic suspensions confirms that it has potential application in liquid-phase fluorescence detection.
Detection of Fe3+ metal cations
The fluorescence properties of complex 1 in various solvents, especially in aqueous solution, encouraged us to explore its use for the detection of various kinds of metal ions closely associated with water environments and human health. In order to further explore the fluorescence properties of complex 1, aqueous solutions of 200 μL, 0.01 mol/L M(NO3)x (M = Na+, Ca2+, Cr3+, Mg2+, Cd2+, Ba2+, Cu2+, Al3+, Pb+, Ni2+ and Fe3+) were respectively added to a suspension of complex 1 (1 mg/mL, 2 mL), and the fluorescence data for complex 1 were then collected under the same conditions. Figure 6 shows the emission intensity of complex 1 in the presence of various metal cations. The fluorescence is quenched by Na+, Ca2+, Cr3+, Mg2+, Cd2+, Ba2+, Cu2+, Al3+, Pb2+, Ni2+ and Fe3+. It is worth noting that Fe3+ produces a more significant quenching effect (when 200 μL is added up to 99.3%), indicating that 1 is a promising fluorescence sensor for distinguishing Fe3+ from numerous other metal ions.
In addition, in order to study the effect of Fe3+ concentration on fluorescence quenching, we added 20–200 μL of Fe3+ to the suspension of complex 1. We found that at an excitation wavelength of 305 nm, the emission peak intensity at 380 nm gradually decreased with the increase in Fe3+ ion concentration. When the concentration of Fe3+ increased from 10−4 to 10−3 mol/L, the quenching rate reached 99.1%. These results show that complex 1 has good fluorescence sensing ability for the detection Fe3+(Fig. 7).
As can be seen from Fig. 8, with the increasing concentration of Fe3+, the S–V curve showed a good linear relationship in a low linear range of 0.1–0.5 mM (R2 = 0.9891). We used the Stern–Volmer (SV) equation to calculate the Ksv, and obtained a value of 2.46*104 L/mol, which may be due to its static quenching. Moreover, irrespective of the presence or absence of Fe3+, the fluorescence lifetime of complex 1 did not change, further supporting this view (Fig. 9, Table 1). The corresponding detection limit (\(3\sigma /K_{\text{SV}}\)) reached 0.81 μM, indicating that complex 1 may be a promising fluorescent probe for the detection of Fe3+. Compared with the results of other studies, the detection ability of complex 1 for Fe3+ ions is superior to that reported in Cd-MOFs [50,51,52,53].
In addition, it can be seen from Fig. 10 that the other ions had no effect on the quenching efficiency of Fe3+, proving that complex 1 is capable of specific detection of Fe3+.
Catalytic reduction of 4-PNP of complex 2
It was found that although complex 2 did not have good fluorescence sensing performance, it showed good performance in terms of catalysis. Therefore, we used complex 2 to conduct a preliminary study on the catalytic reduction of 4-PNP. Twenty milliliters of 4-PNP of a 0.01 mmol/L solution was mixed with 2 mL of 0.2 mol/L NaBH4 solution at room temperature (note: an ice bath was required when the NaBH4 solution was added). Six milligrams of complex 2 was then added to the mixed solution. The UV–visible absorption spectra were recorded until the absorption peak completely disappeared. We found that as the catalytic reaction proceeded, the 4-PNP was reduced to 4-PAP under the catalytic action of complex 2.
The experimental results showed that the UV–visible absorption peak of 4-PNP was 318 nm and the solution color was light yellow. In addition, the absorption peak did not change after mixing for 1 h. This phenomenon indicates that the presence of complex 2 alone did not cause a catalytic reaction. However, after adding 2 mL of NaBH4 solution to the 4-PNP solution, the absorption peak of the 4-PNP solution was redshifted from 318 to 401 nm and the color of the solution turned bright yellow (Fig. 11). The characteristic absorption peak of 4-PAP was 300 nm. When complex 2 was added to the 4-PNP solution of NaBH4, the intensity of the characteristic absorption peak of 4-PNP at 400 nm decreased gradually, but the UV–visible absorption peak of 4-PAP at 300 nm was enhanced. This phenomenon indicates that complex 2 is able to catalytically reduce 4-PNP to 4-PAP (Fig. 12).
Conclusion
In summary, four amino acid complexes 1–4 were synthesized under hydrothermal conditions. Among these, complex 1 showed good detection ability for Fe3+ in terms of fluorescence quenching. The experimental results show that complex 1 can detect Fe3+ qualitatively. Fluorescence quenching titration experiments demonstrated that complex 1 can quantitatively detect Fe3+ at a low concentration of 10−2 mol/L. In addition, the actual water environment was simulated to verify that other metal cations had no effect on the detection of Fe3+ by complex 1. Complex 2, on the other hand, exhibited attractive catalytic performance for the reduction of 4-PNP to 4-PAP.
Supplementary material
The crystallographic data for complexes 1–4 are given in Table S1. The selected bond lengths of complexes 1–4 are listed in Table S2. The details of the structural comparison of complexes 1–4 are provided in Table S3. The detailed hydrogen bonds of complexes 2, 3, 4 are given in Table S4. The attribution of IR for ligand PYSA and complexes 1–4 is shown in Table S5. The detailed characteristics of the UV–Vis spectra of complexes 1–4 are shown in Table S6. The IR spectra of the ligand PYSA and complexes 1–4 are shown in Figure S1. The UV–Vis spectra of the ligand PYSA and complexes 1–4 are shown in Figure S2. The simulated PXRD pattern and measured PXRD spectrum of complexes 1–4 are shown in Figure S3. The PXRD spectra of complex 1 after 24 h soaking in water are shown in Figure S4. Thermal analysis of complexes 1–4 is shown in Figure S5. The solid fluorescence spectra of complex 1 are given in Figure S6.
CIF files have also been deposited at the Cambridge Crystallographic Database Centre (CCDC) and may be obtained from http://www.ccdc.cam.ac.uk by citing CCDC fo complex 1: 1955862; 2: 1955863, 3: 1955513 4:199921.
References
Kurmoo M (2009) Chem Soc Rev 38:1353–1379
Jain P, Ramachandran V, Clark RJ, Zhou HD, Toby BH, Dalal NS, Kroto HW, Cheetham AK (2009) J Am Chem Soc 131:13625–13627
Xiang SC, Wu XT, Zhang JJ, Fu RB, Hu SM, Zhang XZ (2005) J Am Chem Soc 127:16352–16353
Kirillov AM, Karabach YY, Haukka M, Silva MF, Sanchiz J, Kopylovich MN, Pombeiro AJL (2008) Inorg Chem 47:162–175
Gu ZG, Zhou XH, Jin YB, Xiong RG, Zuo JL, You XZ (2007) Inorg Chem 46:5462–5464
Huang YG, Yuan DQ, Pan L, Jiang FL, Wu MY, Zhang XD, Wei W, Gao G, Lee JY, Li J, Hong MC (2007) Inorg Chem 46:9609–9615
Zeng MH, Wang B, Wang XY, Zhang WX, Chen XM, Gao S (2006) Inorg Chem 45:7069–7076
Allendorf MD, Bauer CA, Bhakta RK, Houk RJT (2009) Chem Soc Rev 38:1330–1352
Lee EY, Jang SY, Suh MP (2005) J Am Chem Soc 127:6374–6381
Bauer CA, Timofeeva TV, Settersten TB, Patterson BD, Liu V, Simmons BA, Allendorf MD (2007) J Am Chem Soc 129:7136–7144
Mcmanus GJ, Perry JJ, Perry M, Wagner BD, Zaworotko MJ (2007) J Am Chem Soc 129:9094–9101
Fu RB, Hu SM, Wu XT (2007) Inorg Chem 46:9630–9640
Murray LJ, Dincă M, Long JR (2009) Chem Soc Rev 38:1294–1314
Chen BL, Zhao XB, Putkham A, Hong KL, Lobkovsky EB, Hurtado EJ, Fletcher AJ, Thomas KM (2008) J Am Chem Soc 130:6411–6423
Pan L, Parker B, Huang XY, Olson DH, Lee JY, Li J (2006) J Am Chem Soc 128:4180–4181
Kesanli B, Cui Y, Smith MR, Bittner EW, Bockrath BC, Lin WB (2005) Angew Chem Int Ed 44:72–75
Sudik AC, Millward AR, Ockwig NW, Côté AP, Kim J, Yaghi OM (2005) J Am Chem Soc 127:7110–7118
Wong-Foy AG, Matzger AJ, Yaghi OM (2006) J Am Chem Soc 128:3494–3495
Humphrey SM, Chang JS, Jhung SH, Yoon JW, Wood PT (2007) Angew Chem Int Ed 46:272–275
Ma L, Abney C, Lin W (2009) Chem Soc Rev 38:1248–1256
Lee JY, Farha O, Roberts J, Scheidt KA, Nguyen S, Hupp J (2009) Chem Soc Rev 38:1450–1459
Wu CD, Lin W (2007) Angew Chem Int Ed 46:1075–1078
Seo JS, Whang D, Lee H, Jun SI, Oh J, Jeon Y, Kim K (2000) Nature 404:982–986
Wang M, Xie MH, Wu CD, Wang YG (2009) Chem Commun 40:2396–2398
Wu CD, Li L, Shi LX (2009) Dalton Trans 34:6790–6794
Horike S, Dinca M, Tamaki K, Long JR (2008) Chem Soc 130:5854–5855
Kazmaierand U, Lindner T (2005) Angew Chem 117:3368–3371
Lipner JP, Stodulski M, Mlynarski J (2010) Angew Chem Int Ed 48:4288–4297
Du JJ, Yuan YP, Sun JX, Peng FM, Jiang X, Qiu LG, Xie AJ, Shen YH, Zhu JF (2011) J Hazard Mater 190:945–951
Campo J, Falvello L, Mayoral I, Palacio F, Soler T, Tomás M (2008) J Am Chem Soc 130:2932–2933
Yang XL, Xie MH, Zou C, Sun FF, Wu CD (2011) CrystEngComm 13:1570–1579
Feng Q, Yan LL, Wu HH, Song HT (2014) Inorg Chem 43:1–4
Qu ZR, Zhao H, Wang S, Li YH, Song YM, Liu YJ, Ye Q, Xiong RG, Abrahams BF, Xue ZL, You XZ (2003) Inorg Chem 42:7710–7712
Ma LF, Wang LY, Huo XK, Wang YY, Fan YT, Wang JG, Chen SH (2008) Cryst Growth Des 8:620–628
Li XF, Liu TF, Hu B, Li GL, Zhang H, Cao J (2010) Cryst Growth Des 10:3051–3059
Ghosh S, Mukherjee PS (2008) Organometallics 27:316–319
Zyryanov G, Palacious M, Anzenbacher P (2008) Org Lett 10:3681–3684
Gole B, Shanmugaraju S, Bar AK, Mukherjee PS (2011) Chem Commun 47:10046–10048
Jackson SL, Rananaware A, Rix C, Bhosale SV, Latham K (2016) Cryst Growth Des 16:3067–3071
He J, Xu JL, Yin JC, Li N, Bu XH (2019) Sci China Mater 62:1655–1678
Carter KP, Young AM, Palmer AE (2014) Chem Rev 114:4564–4601
Aron AT, Loehr MO, Bogena J, Chang CJ (2016) J Am Chem Soc 138:14338–14346
Sahoo S, Sharma D, Bera RK, Crisponi G, Callan JF (2012) Chem Soc Rev 41:7195–7227
Lee MH, Giap TV, Sangjin K, Lee YH, Kang C, Kim JS (2010) Chem Commun 46:1407–1409
Koh LL, Ranford JD, Robinson WT, Svensson JO, Tan ALC, Wu DQ (1996) Inorg Chem 35:6466–6472
Sheldrick GM (2008) Acta Crystallogr A 64:112–122
Sheldrick GM (1996) SADABS, Program for Empirical Absorption Correction for Area Detector Data. University of Göttingen, Göttingen
Sheldrick GM (1997) SHELXS 97, Program for Crystal Structure Refinement. University of Göttingen, Göttingen
Spek AL (2003) J Appl Crystallogr 36:7–13
Zhang J, Zhao LL, Liu YX, Li MY, Li G, Meng XR (2018) New J Chem 42:6839–6847
Tong WQ, Liu WN, Cheng JG, Zhang PF, Li GP, Hou L, Wang YY (2018) Dalton Trans 47:9466–9473
Qiu XT, Shi Q, Zhang DQ, Lin QF, Sun YQ (2018) ChemistrySelect. 23:6611–6616
Chand S, Mondal M, Pal SC, Pal A, Maji S, Mandal D, Das MC (2018) New J Chem 42:12865–12871
Acknowledgements
Financial support for this work was provided by grants from the National Natural Science Foundation of China (No. 21571091) and the Commonweal Research Foundation of Liaoning province in China (No. 20170055).
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wang, C., Zhang, N., Hou, CY. et al. Transition metal complexes constructed by pyridine–amino acid: fluorescence sensing and catalytic properties. Transit Met Chem 45, 423–433 (2020). https://doi.org/10.1007/s11243-020-00394-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11243-020-00394-9