Abstract
Flower colour, which is largely determined by anthocyanin, is an important ornamental value of herbaceous peony (Paeonia lactiflora Pall.). Anthocyanin synthesis genes encode enzymes acting at different stages of flavonoid biosynthetic pathway. In this research, we cloned anthocyanin synthesis genes PlANS and PlDFR from red dominated herbaceous peony cultivar ‘Hongyanzhenghui’ and explore their functionality with respect to colour formation by genetic engineering techniques. The over-expression vectors pB1301-PlANS and pB1301-PlDFR were constructed and transformed as heterologous genes into Arabidopsis thaliana (A. thaliana) and tobacco (K326). The quantitative real-time PCR results showed the over expression of PlANS and PlDFR in transgenic plants from 10 to 1600 times. Phenotypic observation and HPLC detection at OD 525 nm also revealed the elevated anthocyanin content in the transgenic Arabidopsis. The a* and c* value of transgenic flower increased 3–4 times compared with control tobacco which are consistent with the observed colour change. Taken together, our findings showed that PlANS, PlDFR is participated in the anthocyanin biosynthesis pathway which could provide a theoretical basis for further research of flower colour in herbaceous peony.
Key message
Our results provided an important research work of two P. lactiflora genes which are related to the red petal colour formation. It provides a theoretical basis for further identification of flower colour-related gene function study in P. lactiflora.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Paeonia lactiflora Pall. (P. lactiflora) is a world well-known flower which is famous for its ornamental, nutritional and medicinal value (Zhao et al. 2016). Especially in China, it is distributed broadly with rich resources and abundant cultivars. Because of its unique ornamental value of bright flower colour and large flowers diameter, it is widely used as cut flower for cultivation and garden landscaping (Hong et al. 2010; Walton et al. 2010). Flower colour is mainly consisted of flavonoids, carotenoids, betalains and other pigments, and flavonoids are discovered to be directly involved in flower pigmentation (Winkel-Shirley 2001; Koes et al. 2005).
Anthocyanins are considered one of the subgroups of water soluble flavonoids. It acts as the main pigments (red to blue) in plants, conferring intense red-to-blue cyanic colours on petals (Mazza 2018). The flavonoid biosynthetic pathway is responsible for synthesis of anthocyanins. High anthocyanins content was found in many fruits such as berries and grapes, coloured leafy vegetables and flowers. In roses, the pigments are composed mainly of anthocyanidin 3,5-O-diglucoside (anthocyanin), which can be further catalysed through glycosylation at different sites of the anthocyanidin molecule (Ogata et al. 2005). In P. lactiflora, most of flower colours were still concentrated on white, pink, red and their derivative, which is determined by flavonoids including anthocyanins and flavones and flavonols (Wu et al. 2016). Our group have reported that there were main anthocyanin composition (peonidin 3,5-di-O-glucoside) (Pn3G5G) and five main anthoxanthin compositions (kaempferol di-hexoside, kaempferol-3-O-malonylglucoside- 7-O-glucoside,quercetin-3-O-galactoside, luteolin-7-O-glucoside and isorhamnetin-3-O-glucoside) were found in double colours herbaceous peony (Wu et al. 2016). Among the 41 P. lactiflora cultivars, only one white P. lactiflora cultivar can not detect anthocyanins while Pn3G5G is identified in the other 3 white P. lactiflora cultivars. In the deep purple or reddish purple P. lactiflora cultivars, they contain four to five anthocyanins, and their contents are much lower than those of purple ones (Jia et al. 2008). However, the molecular mechanism of flower colour formation in P. lactiflora is very few discussed.
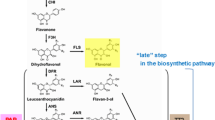
Anthocyanins are structurally diverse but all are based on 17 basic anthocyanidin structures in nature. In the biosynthesis pathway of anthocyanin pigments, the first coloured compound is anthocyanidin (pelargonidin, cyanidin and delphinidin), which is derived from colourless leucoanthocyanidin (Saito et al. 2013). Dihydroflavonol 4-reductase (DFR) provides the first step for anthocyanin production with the formation of leucoanthocyanidins which later are converted to anthocyanidin by anthocyanidin synthase (ANS) (Sakakibara et al. 2019). Recent study indicated CRISPR/Cas9-targeted mutagenesis of OsDFR and two other genes lead to black rice through anthocyanin biosynthesis pathway (Jung et al. 2019). When overexpressing wheat TaDFR in an Arabidopsis dfr mutant, it resulted in significant accumulation of anthocyanins in the respective transgenic plants (Shin et al. 2015). ANS catalyzes the conversion of leucoanthocyanidin to anthocyanidin, a key committed step in biosynthesis of anthocyanins. A 5 bp insertion in the coding region of ANS results in loss of visible and detectable anthocyanin pigments in raspberry (Rafique et al. 2016).
Li et al. introduced the MaANS gene from Morus alba L. into tobacco, which increased flavonoids, anthocyanidin accumulation, and antioxidant potential in transgenic tobacco (Li et al. 2018). The current study has also analyzed the anthocyanin related gene expression patterns of several ornamental plants such as Petunia hybrida and Gentiana scabra, however, there are still many mechanisms for the colour formation of ornamental plants remains further exploration. At present, the research on the colour of peony is mainly focused on the physiological and transcriptome levels (Zhao et al. 2016, 2012). A comprehensive and systematic study of the genes functions which can lay the foundation for the molecular research of colour formation in P. lactiflora still need to be further studied.
In this study, the full-length cDNAs of PlANS and PlDFR were isolated from P. lactiflora. They were both key enzymes in the downstream flavonoids metabolic pathways. The GUS staining results and the transcript levels analysis indicated that PlANS and PlDFR from flower can expressed in Arabidopsis leaves and tobacco flowers. Overexpression of PlANS and PlDFR in Arabidopsis thaliana (A. thaliana) and tobacco (K326) promoted the accumulation of flavonoids, and resulted in the colour changes of transgenic plants. All these results highlighted the importance of PlANS and PlDFR in anthocyanin biosynthesis that may impact both flower industry and molecular breeding technology. The aim of our study on identification and functional analysis of the two anthocyanin synthesis genes are to form novel colour for genetic improvement to provide more materials for new varieties.
Material and methods
Plant materials
The P. lactiflora plants with red petals, called ‘Hongyanzhenghui’ were cultivated in the germplasm repository of Horticulture and Plant Protection College, Yangzhou University, Jiangsu Province, China (32° 30′ N, 119° 25′ E) (Zhao et al. 2016). A. thaliana ecotype Columbia (Col-0) plants and tobacco (K326) were planted with the mixture of 3:1 (v/v) vermiculite and perlite and kept in a growth chamber (SANYO, MLR-359H, Japan).
Plant growth conditions
Three years old P. lactiflora plants were grown on the open-field cultivation conditions without additional fertilizer. The flower petals were picked off in May and used for cloning material. The growth chamber was controlled with a 16 h light (80–100 μmol m−2 s−1 illumination)/8 h darkness cycle and a day/night temperature 23/18 °C, with a relative humidity of 70%. The leaves of A. thaliana at bolting stage were used for qRT-PCR and HPLC detection, and tobacco flowers were used for flower colour identification.
RNA extraction and sequence analysis of two anthocyanin synthesis genes
The fresh petals collected from the flowers of ‘Hongyanzhenghui’ (red colour) at the same location were placed in liquid nitrogen immediately and then preserved at − 80 °C for RNA extraction. Total RNA was extracted from the petals with a Plant RNA kit (TaKaRa, Japan), following the manufacturer’s protocol. Total leaf RNA (1 μg) was reverse-transcribed in a 10 μl reaction based on the superscript first-strand synthesis system (Invitrogen, USA). Then the cDNAs were synthesized and 1 μl cDNA was used as PCR template. Using 5′ and 3′ end primers synthesized commercially (Genery, China) were designed based on the ORF sequence of the GenBank accession ANS (JQ070805.1), DFR (JQ070804.1). PCR reaction was as follows: one cycle of 94 °C (3 min); 35 cycles of 94 °C (30 s), 59 °C (30 s), 72 °C (1 min); and one cycle of 72 °C 10 min; 4 °C end. After testing by 1% (w/v) agarose gel electrophoresis, the PCR products were cloned into the pMD19-T Simple Vector (Takara, Japan) and sequenced. All of gene specific primers (Table 1) were designed through Primer Premier 6.0. Multiple alignment of deduced protein sequences of PlANS and PlDFR with those ones from other species were assembled with DNAMAN 7.0.2. A Neighbor–Joining phylogenetic tree was generated with MEGA-X, using the Poisson correction method and 1000 bootstraps.
Expression vector construction
Based on the ORF sequences of target genes PlANS and PlDFR which contend the SacI and SmaI restriction cut sites, the PCR products were digested by the restriction enzymes (Takara, Japan), and then purified. The purified products were combined to the expression vector pCAMBIA1301 (pB1301) through T4 DNA Ligase (Takara, Japan). The pB1301 was driven by the double Cauliflower mosaic virus (CaMV) 35S promoters followed by the GUS reporter gene. Primers (PlANS and PlDFR) used to get recombinant were shown in Table 1.
Transformation of two anthocyanin synthesis genes into A. thaliana and tobacco
The empty expression vector pB1301 and the pB1301-ANS, pB1301-DFR plasmids were used for the transformation of competent cells of Agrobacterium tumefaciens strain EHA105. Selected colonies were validated by PCR using primers (ANS and DFR; Table 1); ultrapure water as template was used as negative control. Arabidopsis Col-0 plants were transformed using the floral-dip method (Clough and Bent 1998). The inflorescence of A. thaliana was soaked with 1/2 MS infection liquid with transformed Agrobacterim for 1 min, and then cultivated under dark condition for 12 h. The seeds collected were recorded as T1 (the first generation) seeds. Transgenic seeds were all screened on MS medium containing 25 mg L−1 hygromycin (Hyg). Seeds were screened until the homozygous T3 (the third generation) transgenic lines were obtained.
The tobacco plants were transformed by Agrobacterium mediated leaf disc method to infect tobacco (Grimberg et al. 2015). When the differentiated adventitious buds reached more than 2 cm, they were cut off and inserted into rooting medium. The rooting seedlings of tobacco (T1) were transferred into green chamber. The flower colour was observed and recorded after flowering.
GUS staining
The homozygous T3 generation of A. thaliana and transgenic tobacco callus were detected by histochemical GUS staining method, and the nontransgenic plants of the same period was used as negative control. Both of them were placed in 20 μl X-Gluc staining solution and incubated overnight at 37 °C. After staining, the plant tissues were soaked with 70% ethanol for 15 min to remove chlorophyll and repeated decolourization for at least 3 times. The stained transgenic A. thaliana and tobacco callus were detected.
Quantitative real-time PCR analysis
Quantitative real-time PCR (qRT-PCR) was conducted to investigate the expression levels of PlANS and PlDFR in the leaves of A. thaliana. A Trizol Kit (Aidlab, China) was used to extract total RNA from the leaves of Col-0 and transgenic A. thaliana. A 1 μg sample of total RNA was treated with DNase to eliminate genomic DNA contamination. First strand cDNA was synthesized by using Oligo dT (TaKara, Japan), according to the manufacturer’s instruction. The cDNAs were amplified with qRT-PCR primers (Table 1). The presence of DNA residues was tested using 2 μl of cDNA as the template in a 25 μl standard PCR reaction with A. thaliana Actin2 (AK230311.11). The qRT-PCR reaction was performed by denaturation at 94 °C for 30 s, followed by 35 cycles of 5 s denaturation at 94 °C, 30 s annealing at 52.2 °C, and 30 s extension at 72 °C, with a final extension at 72 °C for 5 min and 4 °C forever. qRT-PCR was performed with SYBR Green qPCR Mix (Takara, Japan) and the Bio-Rad CFX96 Real-Time PCR System (BIO-RAD, USA).
HPLC detection
The anthocyanins were extracted from the leaves of transgenic A. thaliana using extraction buffer and examined by high-performance liquid chromatography (HPLC) (Wu and Prior 2005; Ni et al. 2008). Each sample (1.0 g fresh weight) was ground in a mortar with liquid nitrogen. Then 6 ml of a mixture of extraction buffer (70:0.1:29.9; CH3OH:HCl:H2O) was added and vibrated at 4 °C for 24 h. The mixture was then centrifuged at 12,000 rpm for 10 min at 4 °C and each supernatant was filtered through a 0.22 μm Millipore ™ filter (Millipore Corp., Billerica, MA, USA).
Chromatographic analysis were carried out on a high-performance liquid chromatograph (Dionex, USA) equipped with a P680 HPLC Pump, Thermostatted Column Compartment TCC-100 and a Dionex Photodiode Array DAD-100 Detector (DAD). Anthocyanins were detected under the wavelength of 525 nm through HPLC column: TSK gel ODS-80Ts QA (4.6 mm × 250 mm2) (Tosoh, Japan), the column temperature was 35 °C. Extracts were firstly passed through a 0.45 µm pore syringe filter (Anpel, China) and then a volume of 10 µL solution was injected for HPLC analysis. Flow rate was 0.8 mL min−1 and the mobile phase consisted of 10% formic acid water solution as solvent A (10:90; v/v, HCOOH: H2O) and formic acid/acetonitrile/water (10:40:50; v/v/v, HCOOH: CH3CN: H2O) as solvent B. The linear gradient profile was 10% B at 0 min, 20% B at 30 min, 30% B at 50 min, 40% B at 60 min, 50% B at 80 min, and then returned to10% B in 90 min. Data were analyzed with the Chromeleon software (Dionex, USA).
Results
Isolation and sequence analysis of two anthocyanin synthesis genes
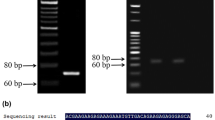
PlANS (Genbank JQ070805.1) and PlDFR (Genbank JQ070804.1) were used for primers design during homologous-based cloning. PCR specific primers were designed according to both sequences above (ANS-cloning and DFR-cloning; Table 1). The cDNA from red flower peony ‘Hongyanzhenghui’ was used for PCR amplification. The full-length cDNA of cloned ANS and DFR were 1111 bp, 1221 bp containing 1065 bp, 1095 bp ORF with a stop codon and encoded a protein of 354, 364 amino acids, respectively. BLASTn analysis on NCBI showed that the cloned PlANS and PlDFR have 100% similarity with the published genes sequence.
Both PlANS and PlDFR amino acid sequences were aligned with sequences from different species which were downloaded from NCBI (Fig. 1). To further investigate the evolutionary relationships of PlANS and PlDFR with other plants, we constructed a phylogenetic tree using MEGA-X (Kumar et al. 2018) (Fig. 2). The amino acids sequence of two genes shared similarity at different extent with other species which have been shown to be involved in anthocyanin biosynthesis. From this, we inferred that PlANS and PlDFR may have a related function to other homologous genes.
Multiple alignment of deduced protein sequences of PlANS and PlDFR. a Multiple alignment of deduced protein sequence of PlANS with ANSs from other species. The blue bars highlight the conserved domains. Gerbera hybrid (AAY15743.2), Malus domestica (AAD26205.1), Paeonia lactiflora (AFI71900.1), Paeonia suffruticosa (AEN71543.1), Rosa hybrid (BAF96590.1). b Multiple alignment of deduced protein sequence of PlDFR with DFRs from other species. The blue bars highlight the NADP-binding site and substrate specificity domain, respectively; the filled red circles indicate the active sites of the proteins. Gerbera hybrid (CAA78930.1), Malus domestica (AAD26204.1), Oryza sativa (CAA69253.1), Paeonia lactiflora (AFI71899.1), Paeonia suffruticosa (AMW36065.1). (Color figure online)
The generation of over-expressors of PlDFR and PlANS
The full-length ORFs of PlANS and PlDFR were subcloned into the expression vector pB1301 and sequenced to be right (Fig. 3). The reconstructed expression vectors pB1301-PlANS and pB1301-PlDFR were also assessed by SacI and SmaI digestion, agarose gel electrophoresis proved that both strips size are correct. Finally, the correct expression vectors were transformed into Agrobacterium tumefaciens strain EHA105 which was used for plant transgenic study.
Screening and assay of PlDFR and PlANS genes expression in A. thaliana
The Agrobacterium tumefaciens strain EHA105 carrying the pB1301-PlANS and pB1301-PlDFR and empty pB1301 vectors were used for A. thaliana transformation. Transgenic lines of A. thaliana were selected on MS medium supplemented with Hygromycin (Hyg) 25 mg L−1 (Fig. 4a). The phenotype of the T1 and T2 lines was recorded. The resistance phenotype of T2 seeds were stained by GUS. Histochemical staining of GUS activity was carried out to screen transgenic positive plants which were turned into blue (Fig. 4b). After sowing T2 seedlings, T3 seeds were collected as homozygous lines. Leaves of T3 seedlings were used to extract genomic DNA for PCR detection with specific primers (ANS and DFR; Table 1). Both NPTII region and 35S promoter sequences were used to detect transgenic lines. Correct stripe lines can be recognized as positive one.
Screening and identification of homozygous OX-PlANS and OX-PlDFR Arabidopsis. a The red arrow indicates the green T1 seedings of successful transformation. The red outline indicates homozygous T2 seedings. b Histochemical staining of GUS in transgenic Arabidopsis compared with it in Col-0. c Phenotypes of Col-0, the empty vector pB1301 and transgenic Arabidopsis overexpressing PlANS and PlDFR. d–g Indentification of overexpression lines by real-time quantitative PCR (qRT-PCR) of PlANS and PlDFR genes. Actin gene expression level in different plant of Col-0, pB1301, PlANS (d) and PlDFR (e) were amplified, respectively. f, g The values on the y-axis were relative expression level. Different letters above bars indicate statistical significance (P < 0.05). (Color figure online)
The transfected homozygous A. thaliana (pB1301) and Col-0 were used as control groups and the results were shown in Fig. 4c. With SD treatment for 2 weeks, the leaves of both OX-PlANS and OX-PlDFR plants turned red at different degrees, OX-PlANS plants are redder than OX-PlDFR ones (Fig. 4c). In contrast, the colours of Col-0 and pB1301 plants leaves were keeping green and not changed. qRT-PCR results (Fig. 4d–g) from T3 homozygous were consistent with the phenotypes. The relative expression level of ANS in two ANS-transgenic homozygous lines was about 800–1600 times higher than that in the control groups, and the expression level of OX-ANS 8# was higher than OX-ANS 6#. The relative expression level of six DFR-transgenic homozygous lines was 10–250 times higher than control group, with the highest level in OX-DFR 2#.
To examine whether anthocyanins accumulated in A. thaliana leaves were consistent with the PlANS and PlDFR expression level, the anthocyanin content in leaves of transgenic plants were detected by HPLC (Fig. 5). The results revealed that their contents in the control and transgenic plants were quite different. At 525 nm, there were obvious peaks of anthocyanins in OX-ANS 8# (pink line) and OX-DFR 2# plants (blue line) compared with Col-0 (black line).
Screening and assay of two anthocyanin synthesis genes expression in tobacco
The process of genetic transformation and regeneration of transgenic tobacco is shown in Fig. 6a–i. PlANS and PlDFR transgenic tobacco callus were regenerated through Hyg selection. GUS staining of transgenic tobacco callus turned into blue (Fig. 6d). During flowering stage, the flower colour of OX-ANS and OX-DFR plants became darker than control tobaccos, as shown in Fig. 6j. Royal Horticultural Society Colour Chart (RHSCC) measurement showed that the flowers colour of control, pB1301, OX-PlANS 8# and OX-PlDFR 2# were 65A, 73C, 68A, 64B, respectively.
The process of transformation and regeneration of the transgenic tobacco plants. a Co-culture, b Screening culture for 1–2 weeks, c Differentiation culture for 3–4 weeks, d GUS staining results of hygromycin-resistant callus tissue, e Re-culture of hygromycin-resistant buds, f, g Rooting culture, h, i Domestication and transplantation, j Phenotypes of flowers on the WT, the empty vector pB1301 and overexpressing PlANS and PlDFR tobacco
Furthermore, Portable Imaging Spectrophotometer RM 200QC (X-Rite, USA) was used to detect the degree of flower colour change which was expressed through L*, a*, b* and C* values. L* represents lightness, a* represents the ratio of red/magenta and green, b* represents the ratio of yellow and blue and C* indicates chroma (saturation or purity of colour) (Zhao et al. 2011). The collected values from wild type (seven plants), pB1301 (seven plants), OX-PlANS (seven plants) and OX-PlDFR (seven plants) were analyzed through L*, a*, b* and C* comparison, respectively (Fig. 7). Statistic analysis showed that the L* and b* values of OX-ANS and OX-DFR were dropped obviously, however, their a* and C* values were 3–4 times higher than the wild types (Fig. 7b, d). This result revealed a redder and different degrees of saturation enhanced colour in ANS and DFR transgenic plants.
Discussion
Anthocyanin and anthoxanthin, especially the anthocyanin content was the main reason for giving P. lactiflora petal red, pink and white colours (Zhao et al. 2016). The peony petals showed white colour when the anthocyanin content was low; it will appear red when the anthocyanin content was high. Pn3G5G was reported as the major composition of anthocyanins in peony petals (Jia et al. 2008). Besides, there is a close correlation between PlDFR and PlANS transcript levels and anthocyanin contents in white and red peony cultivars, the transcription level of both genes were higher in red and pink cultivar than the white one (Zhao et al. 2016). In our previous study, we found the expression of PlANS and PlDFR genes were quite different between red and non-red cultivars which were speculated as a main reason for the formation of red colour (Zhao et al. 2012). However, it was unknown whether PlDFR or PlANS has independent function during the colour formation of peony petals.
So far, it was found that the mutation or RNAi of DFR gene could block the pathway of anthocyanin biosynthesis in tobacco and Dendrobium, respectively (Kazama et al. 2013; Ratanasut et al. 2015; Lim et al. 2016). This indicated that DFR gene is necessary for the flower colour formation in several species. The overexpression of the PtrDFR gene can obviously change tobacco petal colour and attend higher accumulation of anthocyanins (Huang et al. 2012). Furthermore, when introduced DFR gene of cranberry into tobacco, the corolla colour became dark pink (Polashock et al. 2002). In our research, after over-expression of PlDFR, the same significant phenotype was found in the floral organs of transgenetic tobacco. It indicated that PlDFR was positively correlated with the accumulation of anthocyanins, and played an important role in the flavonoid biosynthetic pathway. As for homologous genetic transformation, it was also found that the deletion or silencing of ANS gene can cause flower organ of Calla Lily and Torenia fournieri turned to white colour (Seitz et al. 2003; Aida et al. 2000). We cloned PlANS from red flower peony and used heterologous transformation system to clarify their potential functions. Compared to Qi’s research, our transgenic leaves of PlANS transgenic Arobidopsis turned to be red instead of that stem of the transgenic A. thaliana with MnANS turned reddish (Qi et al. 2014). The PlANS gene also caused the deepening of tobacco petals which was similar to the transgenic tobacco of cocoa TcANS (Liu et al. 2013). These results revealed that both PlDFR and PlANS genes can affect biosynthesis of anthocyanins. However, which form of aglycones of anthocyanins was changed in transgenic plants still need further study.
As an ornamental plant, the genes related to pigments synthesis in P. lactiflora have different functions in the formation of flower colour (Tang et al. 2018; Wu et al. 2018a, b). Due to the lack of homologous transformation system in peony, it still stays at the level of chemical composition analysis and transcriptome sequencing and how to regulate the colour of peony is still unclear. In our study, it was demonstrated that separate transferring PlDFR and PlANS could also affect plant colour, and the phenotype of the transgenic PlANS lines are more pronounced than PlDFR transgenic lines. We speculated that it may be caused by the direct influence of ANS on the synthesis of anthocyanins which is located at downstream of the synthesis pathway. Although we did not transfer both PlDFR and PlANS genes together, Rosati’s research work showed that forsythia displayed a novel bronze-orange petal colour when transferred AmANS and MiDFR together, which implies another application approach in the future (Rosati et al. 2003). Furthermore, our results provide a theoretical basis for further identification of flower colour-related gene function study in P. lactiflora.
References
Aida R, Kishimoto S et al (2000) Modification of flower colour in torenia (Torenia fournieri Lind) by genetic transformation. Plant Sci 153(1):33–42
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16(6):735–743
Grimberg A, Carlsson AS, Marttila S et al (2015) Transcriptional transitions in Nicotiana benthamiana leaves upon induction of oil synthesis by WRINKLED1 homologs from diverse species and tissues. BMC Plant Biol 15:192–209
Hong MH, Kim JH et al (2010) Inhibitory effects of Paeonia suffruticosa on allergic reactions by inhibiting the NF-kappaB/IkappaB-alpha signaling pathway and phosphorylation of ERK in an animal model and human mast cells. Biosci Biotechnol Biochem 74(6):1152–1156
Huang Y, Gou J, Jia Z et al (2012) Molecular cloning and characterization of two genes encoding Dihydroflavonol-4-reductase from Populus trichocarpa. PLoS ONE 7(2):e30364
Jia N, Shu QY, Wang LS (2008) Analysis of petal anthocyanins to investigate colouration mechanism in herbaceous peony cultivars. Sci Hortic 117:167–173
Jung YJ, Lee HJ, Kim JH et al (2019) CRISPR/Cas9-targeted mutagenesis of F3’H, DFR and LDOX, genes related to anthocyanin biosynthesis in black rice (Oryza sativa L.). Plant Biotechnol Rep 13:521–531
Kazama Y, Fujiwara MT, Takehisa H et al (2013) Characterization of a heavy-ion induced white flower mutant of allotetraploid Nicotiana tabacum. Plant Cell Rep 32(1):11–19
Koes R, Verweij W, Quattrocchio F (2005) Flavonoids: a colourful model for the regulation and evolution of biochemical pathways. Trends Plant Sci 10(5):236–242
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35:1547–1549
Li J, Zhao A, Yu M et al (2018) Function analysis of anthocyanidin synthase from Morus alba L. by expression in bacteria and tobacco. Electron J Biotechnol 36:9–14
Lim SH, You MK, Kim DH et al (2016) RNAi-mediated suppression of dihydroflavonol 4-reductase in tobacco allows fine-tuning of flower colour and flux through the flavonoid biosynthetic pathway. Plant Physiol Biochem 109:482–490
Liu Y, Shi Z, Maximova S et al (2013) Proanthocyanidin synthesis in Theobroma cacao: genes encoding anthocyanidin synthase, anthocyanidin reductase, and leucoanthocyanidin reductase. BMC Plant Biol 13:202–221
Mazza G (2018) Anthocyanins in fruits, vegetables, and grains. CRC Press, Florida
Ni J, Qingyan S, Danhua W, Liangsheng W, Zhengan L, Hongxu R et al (2008) Identification and characterization of anthocyanins by high-performance liquid chromatography-electrospray ionization-mass spectrometry in herbaceous peony species. J Am Soc Hortic Sci 133(3):418–426
Ogata J, Kanno Y, Itoh Y et al (2005) Anthocyanin biosynthesis in roses, 2005. Nature 435:757–758
Polashock JJ, Griesbach RJ, Sullivan RF et al (2002) Cloning of a cDNA encoding the cranberry dihydroflavonol-4-reductase (DFR) and expression in transgenic tobacco. Plant Sci 163(2):241–251
Qi XW, Shuai Q, Chen H et al (2014) Cloning and expression analyses of the anthocyanin biosynthetic genes in mulberry plants. Mol Genet Genomics 289(5):783–793
Rafique MZ, Carvalho E, Stracke R et al (2016) Nonsense mutation inside anthocyanidin synthase gene controls pigmentation in yellow raspberry (Rubus idaeus L.). Front Plant Sci 7:1892–1904
Ratanasut K, Monmai C, Piluk P (2015) Transient hairpin RNAi-induced silencing in floral tissues of Dendrobium Sonia ‘Earsakul’ by agroinfiltration for rapid assay of flower colour modification. Plant Cell Tissue Organ 120(2):643–654
Rosati C, Simoneau P, Treutter D et al (2003) Engineering of flower colour in forsythia by expression of two independently-transformed dihydroflavonol 4-reductase and anthocyanidin synthase genes of flavonoid pathway. Mol Breed 12(3):197–208
Saito K, Yonekura-Sakakibara K, Nakabayashi R et al (2013) The flavonoid biosynthetic pathway in Arabidopsis: structural and genetic diversity. Plant Physiol Biochem 72:21–34
Sakakibara KY, Higashi Y, Nakabayashi R (2019) The origin and evolution of plant flavonoid metabolism. Front Plant Sci 10:943–959
Seitz C, Oswald N, Börstling D et al (2003) Being acyanic: an unavoidable fate for many white flowers? ISHS Acta Hortic 612(612):83–88
Shin DH, Choi MG, Kang CS et al (2015) Overexpressing the wheat dihydroflavonol 4-reductase gene TaDFR increases anthocyanin accumulation in an Arabidopsis dfr mutant. Genes Genom 38:333–340
Tang Y, Zhao D et al (2018) Daminozide reduces red colour intensity in herbaceous peony (Paeonia lactiflora Pall.) flowers by inhibiting the expression of flavonoid biosynthetic genes. 3 Biotech 8(2):102
Walton EF, Boldingh HL et al (2010) The dynamics of starch and sugar utilisation in cut peony (Paeonia lactiflora Pall.) stems during storage and vase life. Postharvest Biol Technol 58(2):142–146
Winkel-Shirley B (2001) Flavonoid biosynthesis. A colourful model for genetics, biochemistry, cell biology, and biotechnology. Plant Physiol 126(2):485–493
Wu X, Prior RL (2005) Identification and characterization of anthocyanins by high-performance liquid chromatography-electrospray ionization-tandem mass spectrometry in common foods in the United States: vegetables, nuts, and grains. J Agric Food Chem 53(8):3101–3113
Wu Y, Wei M, Zhao D et al (2016) Flavonoid content and expression analysis of flavonoid biosynthetic genes in herbaceous peony (Paeonia lactiflora Pall.) with double colours. J Integr Agric 15(9):2023–2031
Wu Q, Li PC, Zhang HJ et al (2018a) Relationship between the flavonoid composition and flower colour variation in Victoria. Plant Biol 20:674–681
Wu Y, Zhu M, Jiang Y et al (2018b) Molecular characterization of chalcone isomerase (CHI) regulating flower colour in herbaceous peony (Paeonia lactiflora Pall.). J Integr Agric 17(1):122–129
Zhao D, Tao J, Han C et al (2012) Flower colour diversity revealed by differential expression of flavonoid biosynthetic genes and flavonoid accumulation in herbaceous peony (Paeonia lactiflora pall.). Mol Biol Rep 39(12):11263–11275
Zhao D, Zhou C, Tao J (2011) Carotenoid accumulation and carotenogenic genes expression during two types of persimmon fruit (Diospyros kaki L.) development. Plant Mol Biol Rep 29(3):646–654
Zhao D, Wei M et al (2016) Anatomical and biochemical analysis reveal the role of anthocyanins in flower colouration of herbaceous peony. Plant Physiol Biochem 102:97–106
Acknowledgements
This work was supported by funding from the Subproject of National Key R&D Program (2018YFD1000405), Three New Project for Agriculture of Jiangsu Province (LYSX[2016]45), Three New Agricultural Program of Jiangsu Province (SXGC[2017]297).
Author information
Authors and Affiliations
Contributions
JS and MS were responsible for conception and design of experiments. YW was responsible for data analysis, and drafting of the manuscript. JS, DQZ and JT revised the manuscript. All authors read and approved the final manuscript.
Corresponding author
Additional information
Communicated by Klaus Eimert.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Sun, J., Wu, Y., Shi, M. et al. Isolation of PlANS and PlDFR genes from herbaceous peony (Paeonia lactiflora Pall.) and its functional characterization in Arabidopsis and tobacco. Plant Cell Tiss Organ Cult 141, 435–445 (2020). https://doi.org/10.1007/s11240-020-01802-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11240-020-01802-9