Abstract
DNA methylation is one of the most important epigenetic modifications involved in the development and differentiation in plants. Hypocotyl and cotyledon are the two major tissues of cauliflower (Brassica oleracea L. var. botrytis) seedlings. Both tissues show significantly different tissue specificity and regenerative abilities in vitro. However, the characteristics of DNA methylation modification and its roles in regulating the organ development in cauliflower remain largely unknown. In the present study, the DNA methylation status between the hypocotyl and cotyledon of cauliflower seedlings were analyzed. The results indicated that although the hypocotyl and cotyledon of cauliflower seedlings share the same genome, the genomic DNA methylation levels and patterns at CCGG sites were different. Compared with the cotyledon, the hypocotyl showed higher DNA methylation level, and more loci showing methylation pattern adjustments were also discovered. Twelve loci with changes of DNA methylation patterns were further explored. The quantitative expression analysis indicated that eight out of twelve sequenced fragments showed differential expression between the hypocotyl and cotyledon, of which the expression of six sequences was identified to be negative correlation with their DNA methylation status. In addition, three main DNA methyltransferase genes MET1, CMT3 and DRM were first explored in cauliflower. The results indicated that the expression of these three genes was closely associated with the different DNA methylation status in the hypocotyl and cotyledon. These findings provided more information to further explore the roles of DNA methylation modification in tissue differentiation and development of cauliflower.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
DNA methylation is one of the most important epigenetic regulations in plants. It is usually involved in the conversion of cytosine to 5-methylcytosine in CG, CNG and CNN (N: A, T, C or G) sequence contexts, and is mainly controlled by three types of DNA methyltransferases: METHYLTRANSFERASE (MET), CHROMOMETHYLASE (CMT) and DOMAINS REARRANGED METHYLTRANSFERASE (DRM) (Cao et al. 2003; Zubko et al. 2012; Cao and Jacobsen 2002; Finnegan and Kovac 2000; Lindroth et al. 2001). Increasing investigations have identified that DNA methylation modification play important roles in gene expression regulation, genome stability, genomic imprinting and other genomic processes as well as transgene silencing (Bender 2004; Gehring and Henikoff 2007; Henderson and Jacobsen 2007; Grossniklaus et al. 2001; Weinhold et al. 2013). For example, cytosine methylation of the coding parts of SUPERMAN and AGAMOUS genes decreased the transcription of both genes (Sieburth and Meyerowitz 1997; Ito et al. 2003). Methylating the 5′end of FWA transcription factor resulted in the specific expression of FWA only in the Arabidopsis endosperm (Chan et al. 2005; Cao and Jacobsen 2002). Loss of transgene RNA silencing was detected in some methylation-deficient Arabidopsis mutations, and lower level of transgene silencing in roots was found to be associated with reduced DNA methylation levels at non-symmetrical sites (Andika et al. 2006). In addition, in Arabidopsis, endogenous long terminal repeat (LTR)-type retrotransposons have been found to be reactivated by DNA hypomethylation (Tsukahara et al. 2009), and several other transposable elements were also found to be regulated by DNA methylation modification (Kubis et al. 2003).
Characterizing whole-genome or gene-specific DNA methylation status is crucial for understanding the roles of DNA methylation modification. A large body of research has been conducted to understand the significance of DNA methylation status in diverse plants. For example, in Arabidopsis, a general diminishing of DNA methylation was confirmed to be responsible for changes in meristem identity and organ numbers (Ronemus et al. 1996). Loss of genome-wide methylation has been shown to cause aberrant gametophyte development in Physcomitrella patens (Malik et al. 2012). Variation of DNA methylation status has also been reported in oil palm tissue culture-derived explants (Matthes et al. 2001). Similarly, alteration of DNA methylation status in distinct cell lineages of the layers of carnations during vegetative propagation was detected (Yoshida et al. 2004). A higher level of DNA methylation was also detected in seedlings than in flag leaves of rice (Xiong et al. 1999), and significant differences in cytosine methylation status were found to exist between seedlings and adult plants (Sha et al. 2005). Low-paclitaxel yield of the Taxus cell line after long-term culture has been proposed to be associated with higher level of DNA methylation of this cell line (Fu et al. 2012). In addition, during introgressive hybridization of rice and Zizania latifolia Griseb, changes in patterns and the extent of DNA methylation were found to be significant in the introgression lines compared to their parent lines (Dong et al. 2006). Alteration of DNA methylation status was found to be induced by tissue culture in rice pure-lines, F1 hybrids and polyploids (Wang et al. 2013a). The dynamic tissue- and cell type-specific changes that determine the developmental phase of S. sempervirens shoots were also found to involve DNA methylation variations (Huang et al. 2012). Moreover, adjustments in DNA methylation status have frequently been observed in plants responding to diverse environmental stresses (Alvarez et al. 2010; Yu et al. 2013a). For example, cold treatment of maize seedlings could result in global demethylation of root genomic DNA, particularly in nucleosome core regions (Steward et al. 2002). Vitro culture and thermotherapy of grapevine somaclones also caused changes in DNA methylation (Baránek et al. 2010). Pathogenic infection by bacteria Pseudomonas syringae could lead to significant hypomethylation in host genomic loci (Ruiz-Ferrer and Voinnet 2009). Chemically induced demethylation of the rice R gene Xa21G has been shown to abolish silencing of this gene and provide heritable resistance to Xanthomonas oryzae pv. oryzae (Akimoto et al. 2007). Extensive DNA methylation changes in calluses of Arabidopsis have been detected to be induced by kanamycin as a selective agent (Bardini et al. 2003). Salt stress has been shown to cause changes in DNA methylation patterns in rice (Karan et al. 2012), soybean (Song et al. 2012) and maize (Tan 2010). In asexual dandelions, considerable DNA methylation variations were found to be triggered by various stresses, most notably chemical induction of herbivore and pathogen defenses (Verhoeven et al. 2010). Similarly, multiple abiotic stresses such as warming and nitrogen addition were found to cause alternations in cytosine methylation in natural populations of Leymus chinensis (Yu et al. 2013a). All of these findings indicate that different characteristics of DNA methylation status are closely involved in plant growth and development and response to various biotic and abiotic stresses, although how DNA methylation modification functions in these processes remains to be further elucidated.
Cauliflower is one of the most important vegetables in the Brassica oleracea species with abundant nutrients, and is eaten widely all over the world. Hypocotyl and cotyledon are the two predominant tissues of cauliflower seedlings, and both originate from the embryo and share the same genomic DNA sequences. However, the fates of these two tissues show significant differences during cauliflower development as well as in the development of other dicotyledons. The cotyledon is the first organ that becomes senescent and ultimately departs from the plant body soon after the true leaves are formed. By contrast, the hypocotyl, which contains the shoot apical meristem (SAM), further develops and eventually gives rise to other organs such as leaf, stem, curd, and floral organ. In addition, tissue culture experiments have indicated that the regenerative abilities of hypocotyl and cotyledon are significantly different in cauliflower. Under the same in vitro conditions, the hypocotyl of cauliflower seedlings can more easily regenerate to form callus and then explant than cotyledon. The hypocotyl and cotyledon of cauliflower seedlings share the same genomic DNA sequences. Hence, it is conceivable that epigenetic regulation, such as DNA methylation, histone modification and micro RNA-mediated gene silencing, none of which affect DNA sequences, is involved in tissue-specific identification and formation of distinct regenerative abilities exhibited by these two tissues. The objective of this work was thus to explore the characteristics of DNA methylation status and the expression patterns of three major DNA methyltransferase genes in the hypocotyl and cotyledon of cauliflower seedlings, and further elucidate their potential roles in cauliflower development. Based on the methylation-sensitive amplified polymorphism (MSAP) method, the genomic DNA methylation status at CCGG sites were analyzed in the hypocotyl and cotyledon of cauliflower seedlings. Several MSAP polymorphic bands showing different DNA methylation patterns were cloned, and their transcript levels were further quantitative analysis. Following that, the partial sequences of three major DNA methyltransferase genes: MET1, CMT3 and DRM were first cloned and identified in cauliflower. Quantitative expression analysis of these three genes was carried out. The relationship between the DNA methylation status and the expression of three DNA methyltransferase genes, and their relationship with the tissue specificity and different regenerative abilities of hypocotyl and cotyledon were also discussed.
Materials and methods
Plant materials
The cauliflower seeds were treated with 75 % ethanol for 5 min and 2 % NaOCl for 10 min, respectively, and then rinsed thrice with sterile distilled water. Subsequently, the seeds were grown in MS solid medium under controlled conditions with a 16/8 h light/dark cycle at 25 and 22 °C, respectively. The hypocotyl and cotyledon of 5-, 10-, and 15-day-old cauliflower seedlings were collected, immediately frozen in liquid nitrogen, and stored at −80 °C for DNA and RNA isolation. To investigate the regenerative abilities of the hypocotyl and cotyledon of cauliflower seedlings in vitro, the hypocotyl and cotyledon of 5-old-seedlings were collected, and cultured in MS solid medium under the controlled conditions (16/8 h light/dark cycle at 25 °C).
DNA and RNA isolation
Genomic DNA was isolated by the CTAB (cetyltrimethylammonium bromide) method with some modifications (Kidwell and Osborn 1992). In brief, 200 mg hypocotyl or cotyledon of 5-, 10- and 15-day-old cauliflower seedlings, respectively, were ground in liquid nitrogen, and the frozen powder was treated with 2 mL lysis buffer containing 100 mM Tris–HCl (pH 8.0), 100 mM EDTA, 1.4 M NaCl, 0.2 % β-mercaptoethanol, 2 % PVP and 1 × CTAB at 56 °C for 20 min. Subsequently, 2 mL phenol:chloroform:isoamyl alcohol (25:24:1, V:V:V) was added. The following steps were as described by our previous study (Wang et al. 2007). To conduct MSAP analysis, equal amount of DNA from the hypocotyl of 5-, 10- and 15-day-old cauliflower seedlings was pooled to construct hypocotyl DNA pool. Similarly, equal amount of DNA from the cotyledon of 5-, 10- and 15-day-old cauliflower seedlings was also pooled to construct cotyledon DNA pool.
Total RNAs were extracted from the hypocotyl and cotyledon of 5-, 10- and 15-day-old cauliflower seedlings, respectively, by using the TRIzol reagent (Invitrogen, USA) according to the manufacturer’s instructions. After the contaminated DNA was digested by RNase-Free DNAse I (Promega, USA) for 30 min at 37 °C, the quality of RNA was tested by 1.2 % denaturing agarose gels and 2 μg total RNA were reverse transcribed to cDNA using the AMV reverse transcriptase (Promega, USA).
MSAP assay
The protocol used for MSAP analysis was conducted mainly as previously described (Xiong et al. 1999), with modifications. Briefly, 300 ng DNA from the hypocotyl and cotyledon pools, respectively, were digested for 48 h at 37 °C with 2 U EcoRI (TaKaRa, Japan) and 2 U HpaII or MspI (Promega, USA) in a final volume of 15 μL. The digested fragments were then ligated to HpaII/MspI adapters and EcoRI adapters using the T4 DNA ligase (TaKaRa, Japan) in a final volume of 20 μL, and incubated at 16 °C for 12 h. Thereafter, the following reactions of pre-amplification and selective amplification using the pre-selective primers and selective primers (Online resource 1), respectively, were conducted (Wang et al. 2009). The denatured PCR products were separated on a 6 % denaturing polyacrylamide gel at 35 W for 1.5 h, and then the gels were stained with silver (Chalhoub et al. 1997). Only clear, reproducible bands that appeared in three independent experiments were scored, and these bands were scored in a binary character matrix, with “1” indicating the presence and “0” indicating the absence at a particular position. Statistical analysis was performed by the U tests (Zhao et al. 2008).
Cloning of MSAP fragments
The bands, which represent the adjustment of DNA methylation patterns in certain loci detected by MSAP analysis, were isolated and re-amplified with the appropriate selective primer combinations. The corresponding PCR products were subsequently ligated into a pEASY-T1 vector (Transgene, China) and transformed into DH5α, and sequenced with an ABI3770 sequencer (Applied Biosystems, USA). The advanced homology analysis of the sequences was performed using the Blast tool (http://www.ncbi.nlm.nih.gov).
DNA gel blot analysis
To confirm the reliability of MSAP method in detecting the DNA methylation status, Southern blot analysis was conducted. In brief, a total of 12 μg of genomic DNA from the hypocotyl and cotyledon DNA pools, were digested with 15 U EcoRI/10 U HpaII and 15 U EcoRI/10 U MspI, respectively. The two digested products of each sample were loaded next to each other on a 0.8 % agarose gel, and blotted onto a nylon membrane by capillary transfer with alkali solution (0.4 M NaOH). The probes were selected from the sequenced MSAP bands. The following probes preparation, hybridization, and detection were performed using a DIG DNA labeling and detection kit (Roche, Germany) according to the manufacturer’s instructions.
Cloning of partial sequences of MET1, CMT3 and DRM
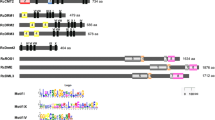
Based on the reported DNA sequences of MET1, CMT3 and DRM deposited in the GenBank database, the specific primers used to amplify the corresponding genes in cauliflower were designed. The PCR product of each gene was cloned and sequenced with an ABI3770 sequencer (Applied Biosystems, USA). The gene sequences obtained from the cauliflower with the corresponding sequences from other plant species (Online resource 2) were further analyzed by Clustal W software (Thompson et al. 1994), and phylogenetic analysis was conducted using the Neighbor-Joining method (Saitou and Nei 1987) by MEGA 5 (Tamura et al. 2007).
Quantitative real-time RT-PCR analysis
The quantitative analysis of possible genes with the adjustment of DNA patterns and the three DNA methyltransferase genes in the hypocotyl and cotyledon of cauliflower seedlings were performed by qRT-PCR, respectively. In brief, 100 ng the first-strand cDNA of each sample was used for the real-time PCR analysis. Real-time qPCR was performed with iQ SYBR Green Supermix (Roche, Germany) using the iCycler iQ5 system (Bio-Rad, USA) and specific primers (Online resource 3). Three independent biology replicates were analyzed per sample. Expression values were normalized using the house-keeping gene actin and relative expression levels of each sample were performed using (Ct) 2−Δ(ΔCt) method.
Results
Distinct regenerative abilities showed by the hypocotyl and cotyledon of cauliflower seedlings
Hypocotyl and cotyledon are two major tissues of cauliflower seedlings. However, significant differences were found in the developmental processes of these two tissues. As a terminal differentiation organ, the cotyledon shows wilt and fall off soon after the generation of true leaves. By contrast, the hypocotyl possessing the SAM can continue to grow and form leaves, stem, and floral organs. In addition, tissue culture experiments indicated that the regenerative abilities of the hypocotyl and cotyledon are significantly different in cauliflower. Under the same condition, the hypocotyl of cauliflower seedlings regenerates to form callus and then explants after about 10 and 15 days, respectively. However, the formation of explants appears to be more difficult from the cotyledon of cauliflower seedlings (Fig. 1).
The regenerative abilities of the hypocotyl and cotyledon of cauliflower seedlings in vitro. a The 5-old-cauliflower seedlings, b the explant culture of hypocotyl for 5 days, c the explant culture of hypocotyl for 15 days, d the explant culture of hypocotyl for 25 days, e the explant culture of cotyledon with petiole for 5 days, f the explant culture of cotyledon without petiole for 5 days, g the explant culture of cotyledon with petiole for 35 days, h the explant culture of cotyledon without petiole for 35 days
DNA methylation levels in the hypocotyl and cotyledon of cauliflower seedlings
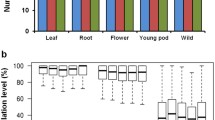
Sixty-four different selective amplification primer pairs were used for MSAP analysis, of which the amplified results of fifty-one primer pairs showed good repeatability and high polymorphism (Online resource 1). Therefore, the MSAP bands amplified by these 51 primer pairs were used for follow-up data analysis. Based on the results of denaturing polyacrylamide gel electrophoresis (PAGE), a total of 1,583 loci were detected in the hypocotyl and cotyledon of cauliflower seedlings, of which more than half did not show methylation. Methylated loci accounted for 43.52 and 41.12 % of the total loci in the hypocotyl and cotyledon, respectively (Table 1). This result showed that compared with the cotyledon, the DNA methylation level at CCGG sites was higher in the hypocotyl of cauliflower seedlings. Further analysis of different types of DNA methylation showed that the semi-methylation ratio of the hypocotyl (12.51 %) was lower than that of the cotyledon (15.22 %). By contrast, the full methylation ratio of the hypocotyl (24.76 %) was higher than that of the cotyledon (20.85 %). Similarly, the ratio of another type of DNA methylation was higher in the hypocotyl than in the cotyledon, although the status of these DNA methylation sites was not clearly distinguished by MSAP analysis (Table 1). Thus, these data indicated that the total DNA methylation level as well as the ratio of different types of DNA methylation at CCGG sites were different in the two major tissues of cauliflower seedlings.
DNA methylation patterns between the hypocotyl and cotyledon of cauliflower seedlings
The characteristics of DNA methylation patterns at the detected 1,583 loci were further analyzed between the hypocotyl and cotyledon of cauliflower seedlings. In total, four types containing 15 subtypes of different DNA methylation patterns were detected (Fig. 2; Table 2). Data analysis indicated that more than 60 % of the 1,583 loci did not show the changes in amplification patterns, of which most of the loci were not methylated (43.15 %), and proportions of semi-methylated and fully methylated loci were 4.23 and 13.27 %, respectively (Table 2). The remaining about 40 % of loci were found to show adjustments in DNA methylation patterns between the hypocotyl and the cotyledon (Table 2). Compared with the cotyledon, 20.66 % of the loci underwent methylation pattern adjustments (type B), and 17.37 % of the loci in the hypocotyl underwent changes in demethylation patterns (type C). These data indicate that the trend of methylation pattern adjustments appears to be more predominant in the hypocotyl. In addition, another type of DNA methylation pattern adjustment, which was undistinguished by the MSAP method (type D), were also detected, and these loci accounted for only 1.33 % of the total detected loci (Table 2).
Representative DNA methylation patterns that were detected in the hypocotyl and cotyledon of cauliflower seedlings by MSAP method. The red boxes indicate the different types of DNA methylation patterns in hypocotyl and cotyledon; H indicates the selective amplification results used the genomic DNA digested with HpaII and EcoRI as templates; M indicates the selective amplification results used the genomic DNA digested with MspI and EcoRI as templates; Hy and Co are the abbreviation of hypocotyl and cotyledon, respectively; L indicates the 50 bp molecular marker; E 32 /H 47 , E 33 /H 25 , E 33 /H 47 , E 37 /H 48 and E 39 /H 83 indicate the different MSAP primer combinations, respectively. (Color figure online)
Cloning and sequence analysis of polymorphic MSAP fragments
Thirteen loci with adjustments of DNA methylation patterns between the hypocotyl and cotyledon were randomly selected. The corresponding polymorphic fragments were isolated, and among them, twelve fragments were successfully cloned (Table 3). Sequence analysis indicated that 4 out of the 12 sequenced fragments showed high sequence homology with reported encoding sequences in other plants (Table 3). For example, E33H25-9-3 and E37H48-3-1 were found to be homologous with the sequences encoding the seed storage and ARM repeat superfamily proteins in Arabidopsis, respectively. To further validate the DNA methylation patterns that were revealed by the MSAP analysis, three sequenced fragments (E33H50-16-3, E33H48-13-9 and E33H61-17-1) were randomly selected as probes for a Southern blot analysis. The results identified that the Southern hybridization patterns of E33H50-16-3 and E33H48-13-9 were fully consistent with those detected in MSAP assay. The hybridization pattern of E33H61-17-1 was also coincident with the MSAP assay, except other redundant hybrid signal was showed (Fig. 3). Nevertheless, these results validated that the MSAP method were reliable to explore the DNA methylation status.
DNA gel blot analysis using the isolated fragments with changes of DNA methylation patterns as probes. H indicates that the DNA samples were digested with EcoRI/HpaII; M indicates that the DNA samples were digested with EcoRI/MspI; E33H48-13-9, E33H61-17-1 and E33H50-16-3 indicate the different probes, which were detected to show changes of DNA methylation patterns by MSAP analysis; Hy and Co are the abbreviation of hypocotyl and cotyledon, respectively
Transcript expression of sequenced possible genes with DNA methylation pattern adjustment
The changes of DNA methylation level or pattern were closely related to the regulation of gene expression. Consequently, qRT-PCR analysis was conducted to explore the transcript expression of twelve sequenced fragments with adjustments of DNA methylation patterns in hypocotyl and cotyledon. The results indicated that except four sequenced fragments (E33H25-8-1, E33H48-13-9, E37H49-4-1 and E39H83-1-2) originating from four different loci, the other eight sequenced fragments all showed differential expression between the hypocotyl and cotyledon. Among them, the expression of most of the fragments (6/8) was showed to be negative correlation with their corresponding DNA methylation status (Fig. 4a–h). These results further confirmed that the characteristics of DNA methylation status could significantly affect the gene transcription expression in early development of cauliflower.
Identification and expression analysis of MET1, CMT3 and DRM in cauliflower
Three main types of DNA methyltransferases (CMT, DRM and MET) have been identified in diverse plants. To explore the DNA methyltransferases in cauliflower and elucidate their roles on the DNA methylation status of hypocotyl and cotyledon, partial sequences of three DNA methyltransferase genes, namely, MET1, CMT3 and DRM were cloned from cauliflower (accession numbers: KF651133-KF651135). Sequence analysis showed that MET1, CMT3 and DRM from cauliflower were all highly homologous with those of the corresponding genes in Arabidopsis. Further phylogenetic analysis indicated that MET1 and CMT3 were also highly conserved in other plants (Online resource 4). However, homology of cauliflower DRM with that of other plants, except Arabidopsis, was very low (data not shown). Quantitative analysis of these three DNA methyltransferase genes was conducted. The results indicated that all three of these genes showed differential expression, and the expression levels of MET1 and CMT3 in the hypocotyl were significantly higher than those in the cotyledon. By contrast, expression of DRM was significantly higher in the cotyledon than that in the hypocotyl (Fig. 4i).
Discussion
DNA methylation is one of the most important epigenetic modifications, and has been identified to be involved in the growth, development and tissue-specific maintenance of plants (Jin et al. 2013; Braszewska-Zalewska et al. 2013; Vining et al. 2013; Iwasaki et al. 2013; Jullien et al. 2012; Viejo et al. 2012). In these processes, one of the main functions of DNA methylation is to regulate gene expression (Weinhold et al. 2013; Gohlke et al. 2013; Wang et al. 2013b; Kiselev et al. 2013; Li et al. 2012). Generally, if the cytosines of a gene are methylated, especially in the promoter region, expression of the gene is inhibited. By contrast, DNA demethylation is closely associated with the activation of gene expression (Brabbs et al. 2013; Yu et al. 2013b). Therefore, to elucidate the roles of DNA methylation in regulation of gene expression, it is necessary to uncover the whole-genome DNA methylation status or the DNA methylation characteristics of specific genes of interest. Recently, based on analysis of DNA methylomes, a large number of differentially methylated regions have been detected in ripening tomato fruits, and the evidence showed that adjustments in DNA methylation status may ensure the fidelity of this developmental process (Zhong et al. 2013). In fact, variation in DNA methylation status have been discovered in different tissues or different development stages of almost all detected plants, e.g., in the oil palm tissue culture-derived explants (Matthes et al. 2001), the distinct cell lineages of carnations (Yoshida et al. 2004), and the seedlings and flag leaves of rice (Xiong et al. 1999). All of these findings provided crucial clues to further explore the relationship between DNA methylation modification and the growth and development processes in plants. Consistent with the previous reports, different DNA methylation status in the hypocotyl and cotyledon of cauliflower seedlings were also detected in the present study. Further analysis indicated that compared with the cotyledon, the full methylation ratio as well as the ratio of another DNA methylation type which was not distinguished by the MSAP method were obviously higher in the hypocotyl, resulting in higher total DNA methylation level of the hypocotyl. However, the semi-methylation ratio was higher in the cotyledon than in the hypocotyl. Moreover, analysis of the DNA methylation patterns indicated that among the nearly 40 % of detected loci which showed adjustments in their DNA methylation patterns in the hypocotyl and cotyledon, methylation pattern adjustments were identified to be more predominant in the hypocotyl. These results suggested that despite sharing the same genome composition, the hypocotyl of cauliflower seedlings showed higher DNA methylation level, with more loci showing methylation pattern adjustments, compared with the cotyledon. Furthermore, these results implied that expression of more genes in the hypocotyl of cauliflower seedlings may be inhibited. Hypocotyl and cotyledon are two major tissues of cauliflower seedlings, and show significantly different tissue specificity. As a terminal differentiation organ, the main function of the cotyledon is to provide nutrients for seed germination and early development of cauliflower. Therefore, it is conceivable that extensive gene expression and corresponding protein synthesis in the cotyledon are necessary to support the early development of cauliflower. Interestingly, more total RNAs were obtained from the cotyledon despite using the same weights of cotyledon and hypocotyl of cauliflower seedlings (data not shown), which may further imply that gene expression was more active in the cotyledon, even though the hypocotyl shows stronger differentiation and regeneration abilities than the cotyledon in vitro. Accordingly, it can be speculated that the different characteristics of DNA methylation status showed by the hypocotyl and cotyledon of cauliflower seedlings were closely involved in their tissue specificity. Compared with the hypocotyl, lower DNA methylation level and more loci showing demethylation pattern adjustments may be necessary for the cotyledon to activate more genes to support the early development of cauliflower. Nevertheless, further identification of genes that undergo adjustments in DNA methylation patterns and uncovering of their DNA methylation status are required to elucidate the roles of DNA methylation modification in hypocotyl and cotyledon development. In the present study, several loci showing adjustments of DNA methylation patterns were analyzed, and most of them were found to be differentially expressed in the hypocotyl and the cotyledon, implying that the corresponding genes or transcription factors were regulated by DNA methylation modification. Thus, these findings provide new insights into the roles of possible genes regulated by DNA methylation in the early development of cauliflower.
Uncovering how genes are regulated by DNA methylation modification is important to elucidate the roles of DNA methylation in plant growth and development or other biological processes. Moreover, understanding how DNA methylation modification is regulated in plants is also crucial to elucidate the functions of DNA methylation. Investigations have indicated that DNA methyltransferases play crucial roles in the cytosine methylation of DNA sequences in diverse plants, and three main DNA methyltransferases genes MET, CMT and DRM, which play different roles in DNA methylation modification, have been identified (Finnegan and Kovac 2000; Steward et al. 2000; Xiao et al. 2006; Cao and Jacobsen 2002). However, the DNA methyltransferases genes in cauliflower and their roles in DNA methylation modification remain to be studied. Consequently, MET1, CMT3 and DRM were analyzed for the first time in the present study. Sequence alignment analysis indicated that MET1 and CMT3 in cauliflower were highly homologous with the corresponding genes in diverse plants, while DRM only showed high homology with the Arabidopsis DRM gene. These results suggested that the functions of MET1 and CMT3 are highly conserved in plants. MET1 is a homologue of the mammalian maintenance methyltransferase Dnmt1 (Finnegan et al. 1996; Kankel et al. 2003), and has been confirmed to play a role in faithful maintenance of the CG methylation patterns. CMT3 is specific to the plant kingdom, and has been reported to predominantly control CNG methylation (Lindroth et al. 2001). Transcript expression analysis of MET1 and CMT3 indicated that the expression level of both genes were significantly higher in the hypocotyl than in the cotyledon of cauliflower seedlings, which is consistent with the higher full methylation ratio of the hypocotyl. Therefore, this suggests that MET1 and CMT3 may be mainly involved in controlling the fully methylated status of CCGG sites during development of cauliflower seedlings. DRM is another major class of DNA methyltransferase gene, and is targeted by siRNAs generated by the RNA-directed DNA methylation (RdDM) pathway. It is a homologue of mammalian Dnmt3 de novo methyltransferases, and is responsible for de novo methylation of cytosines in all sequence contexts (Xie et al. 2004; Cao and Jacobsen 2002). Different from the MET1 and CMT3 genes, DRM was found to show lower expression level in the hypocotyl than in the cotyledon of cauliflower seedlings, which is consistent with the lower semi-methylated ratio of the hypocotyl. In fact, based on the MSAP analysis, all the detected semi-methylated sites were belonged to asymmetric cytosine methylation. These results suggested that DRM should play important roles in controlling the semi-methylated status of CCGG sites during development of cauliflower seedlings. In addition, previous reports have showed that loss-of-function or inhibiting expression of these DNA methyltransferase genes could result in reactivating the expression of some silenced genes in plants (Chen and Pikaard 1997; Chang and Pikaard 2005; Hudson et al. 2011; Pillot et al. 2010). Moreover, the recent report indicated that enhancing expression of these genes also could lead to random hypermethylation in normally unmethylated loci and result in its silencing (Tyunin et al. 2012). These results suggested that the changes of DNA methyltransferase gene expression can significantly affect the expression of other genes, which then affect the plant growth and development. Accordingly, although the roles of DNA methyltransferase genes in cauliflower are still to be elucidated, the detected different expression profiles of these three genes in the present study implied that they should play crucial roles in regulating gene expression in early development of cauliflower.
In summary, findings of the present study suggest that the tissue-specific maintenance and regenerative potential of the hypocotyl and the cotyledon of cauliflower seedlings in vitro are closely associated with their corresponding DNA methylation status. Lower DNA methylation level and more loci showing demethylation changes may be necessary in the cotyledons to activate more genes and thus facilitate synthesis of more proteins to support early development of cauliflower. However, compared with the cotyledon, higher DNA methylation level and more loci showing methylation pattern adjustments did not affect the hypocotyl showed higher differentiation and regeneration abilities. In addition, three main DNA methyltransferases genes were first explored in cauliflower, and the findings indicated that differential expression of these genes may contribute to the different DNA methylation status of the hypocotyl and cotyledon of cauliflower seedlings. Loci showing changes in DNA methylation patterns were also detected, and importantly, differential expression of several such loci in hypocotyl and cotyledon were discovered. Taken together, these findings provided more new insights into the roles of DNA methylation in early development of cauliflower.
References
Akimoto K, Katakami H, Kim HJ, Ogawa E, Sano CM, Wada Y, Sano H (2007) Epigenetic inheritance in rice plant. Ann Bot 100:205–217
Alvarez ME, Nota F, Cambiagno DA (2010) Epigenetic control of plant immunity. Mol Plant Pathol 11:563–576
Andika IB, Kondo H, Rahim MD, Tamada T (2006) Lower levels of transgene silencing in roots is associated with reduced DNA methylation levels at non-symmetrical sites but not at symmetrical sites. Plant Mol Biol 60:423–435
Baránek M, Křižan B, Ondrušíková E, Pidra M (2010) DNA-methylation changes in grapevine somaclones following in vitro culture and thermotherapy. Plant Cell Tissue Organ Cult 101:11–22
Bardini M, Labra M, Winfield M, Sala F (2003) Antibiotic-induced DNA methylation changes in calluses of Arabidopsis thaliana. Plant Cell Tissue Organ Cult 72:157–162
Bender J (2004) DNA methylation and epigenetics. Annu Rev Plant Biol 55:41–68
Brabbs TR, He Z, Hogg K, Kamenski A, Li Y, Paszkiewicz KH, Moore KA, O’Toole P, Graham IA, Jones L (2013) The stochastic silencing phenotype of Arabidopsis morc6 mutants reveals a role in efficient RNA-directed DNA methylation. Plant J 75:835–846
Braszewska-Zalewska AJ, Wolny EA, Smialek L, Hasterok R (2013) Tissue-specific epigenetic modifications in root apical meristem cells of Hordeum vulgare. PLoS One 8:e69204
Cao X, Jacobsen SE (2002) Role of the arabidopsis DRM methyltransferases in de novo DNA methylation and gene silencing. Curr Biol 12:1138–1144
Cao X, Aufsatz W, Zilberman D, Mette MF, Huang MS, Matzke M, Jacobsen SE (2003) Role of the DRM and CMT3 methyltransferases in RNA-directed DNA methylation. Curr Biol 13:2212–2217
Chalhoub BA, Thibault S, Laucou V, Rameau C, Höfte H, Cousin R (1997) Silver staining and recovery of AFLP amplification products on large denaturing polyacrylamide gels. Biotechniques 22:216–220
Chan SW, Henderson IR, Jacobsen SE (2005) Gardening the genome: DNA methylation in Arabidopsis thaliana. Nat Rev Genet 6:351–360
Chang S, Pikaard CS (2005) Transcript profiling in Arabidopsis reveals complex responses to global inhibition of DNA methylation and histone deacetylation. J Biol Chem 280:796–804
Chen ZJ, Pikaard CS (1997) Epigenetic silencing of RNA polymerase I transcription: a role for DNA methylation and histone modification in nucleolar dominance. Genes Dev 11:2124–2136
Dong ZY, Wang YM, Zhang ZJ, Shen Y, Lin XY, Ou XF, Han FP, Liu B (2006) Extent and pattern of DNA methylation alteration in rice lines derived from introgressive hybridization of rice and Zizania latifolia Griseb. Theor Appl Genet 113:196–205
Finnegan EJ, Kovac KA (2000) Plant DNA methyltransferases. Plant Mol Biol 43:189–201
Finnegan EJ, Peacock WJ, Dennis ES (1996) Reduced DNA methylation in Arabidopsis thaliana results in abnormal plant development. Proc Natl Acad Sci USA 93:8449–8454
Fu C, Li L, Wu W, Li M, Yu X, Yu L (2012) Assessment of genetic and epigenetic variation during long-term Taxus cell culture. Plant Cell Rep 31:1321–1331
Gehring M, Henikoff S (2007) DNA methylation dynamics in plant genomes. Biochim Biophys Acta 1769:276–286
Gohlke J, Scholz CJ, Kneitz S, Weber D, Fuchs J, Hedrich R, Deeken R (2013) DNA methylation mediated control of gene expression is critical for development of crown gall tumors. PLoS Genet 9:e1003267
Grossniklaus U, Spillane C, Page DR, Köhler C (2001) Genomic imprinting and seed development: endosperm formation with and without sex. Curr Opin Plant Biol 4:21–27
Henderson IR, Jacobsen SE (2007) Epigenetic inheritance in plants. Nature 447:418–424
Huang LC, Hsiao LJ, Pu SY, Kuo CI, Huang BL, Tseng TC, Huang HJ, Chen YT (2012) DNA methylation and genome rearrangement characteristics of phase change in cultured shoots of Sequoia sempervirens. Physiol Plant 145:360–368
Hudson K, Luo S, Hagemann N, Preuss D (2011) Changes in global gene expression in response to chemical and genetic perturbation of chromatin structure. PLoS One 6:e20587
Ito T, Sakai H, Meyerowitz EM (2003) Whorl-specific expression of the SUPERMAN gene of Arabidopsis is mediated by cis elements in the transcribed region. Curr Biol 13:1524–1530
Iwasaki M, Takahashi H, Iwakawa H, Nakagawa A, Ishikawa T, Tanaka H, Matsumura Y, Pekker I, Eshed Y, Vial-Pradel S, Ito T, Watanabe Y, Ueno Y, Fukazawa H, Kojima S, Machida Y, Machida C (2013) Dual regulation of ETTIN (ARF3) gene expression by AS1-AS2, which maintains the DNA methylation level, is involved in stabilization of leaf adaxial–abaxial partitioning in Arabidopsis. Development 140:1958–1969
Jin X, Pang Y, Jia F, Xiao G, Li Q, Zhu Y (2013) A potential role for CHH DNA methylation in cotton fiber growth patterns. PLoS One 8:e60547
Jullien PE, Susaki D, Yelagandula R, Higashiyama T, Berger F (2012) DNA methylation dynamics during sexual reproduction in Arabidopsis thaliana. Curr Biol 22:1825–1830
Kankel MW, Ramsey DE, Stokes TL, Flowers SK, Haag JR, Jeddeloh JA, Riddle NC, Verbsky ML, Richards EJ (2003) Arabidopsis MET1 cytosine methyltransferase mutants. Genetics 163:1109–1122
Karan R, DeLeon T, Biradar H, Subudhi PK (2012) Salt stress induced variation in DNA methylation pattern and its influence on gene expression in contrasting rice genotypes. PLoS One 7:e40203
Kidwell KK, Osborn TC (1992) Simple plant DNA isolation procedures. In: Beckmann JS, Osborn TC (eds) Plant genomes: methods for genetic and physical mapping. Kluwer Academic Publishers, Dordrecht, pp 1–13
Kiselev KV, Tyunin AP, Zhuravlev YN (2013) Involvement of DNA methylation in the regulation of STS10 gene expression in Vitis amurensis. Planta 237:933–941
Kubis SE, Castilho AM, Vershinin AV, Heslop-Harrison JS (2003) Retroelements, transposons and methylation status in the genome of oil palm (Elaeis guineensis) and the relationship to somaclonal variation. Plant Mol Biol 52:69–79
Li X, Zhu J, Hu F, Ge S, Ye M, Xiang H, Zhang G, Zheng X, Zhang H, Zhang S, Li Q, Luo R, Yu C, Yu J, Sun J, Zou X, Cao X, Xie X, Wang J, Wang W (2012) Single-base resolution maps of cultivated and wild rice methylomes and regulatory roles of DNA methylation in plant gene expression. BMC Genom 13:300
Lindroth AM, Cao X, Jackson JP, Zilberman D, McCallum CM, Henikoff S, Jacobsen SE (2001) Requirement of CHROMOMETHYLASE3 for maintenance of CpXpG methylation. Science 292:2077–2080
Malik G, Dangwal M, Kapoor S, Kapoor M (2012) Role of DNA methylation in growth and differentiation in Physcomitrella patens and characterization of cytosine DNA methyltransferases. FEBS J 279:4081–4094
Matthes M, Singh R, Cheah S-C, Karp A (2001) Variation in oil palm (Eleais guineensis Jacq.) tissue culture-derived regenerants revealed by AFLPs with methylation-sensitive enzymes. Theor Appl Genet 102:971–979
Pillot M, Autran D, Leblanc O, Grimanelli D (2010) A role for CHROMOMETHYLASE3 in mediating transposon and euchromatin silencing during egg cell reprogramming in Arabidopsis. Plant Signal Behav 5:1167–1170
Ronemus MJ, Galbiati M, Ticknor C, Chen J, Dellaporta SL (1996) Demethylation-induced developmental pleiotropy in Arabidopsis. Science 273:654–657
Ruiz-Ferrer V, Voinnet O (2009) Roles of plant small RNAs in biotic stress responses. Annu Rev Plant Biol 60:485–510
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sha AH, Lin XH, Huang JB, Zhang DP (2005) Analysis of DNA methylation related to rice adult plant resistance to bacterial blight based on methylation-sensitive AFLP (MSAP) analysis. Mol Genet Genomics 273:484–490
Sieburth LE, Meyerowitz EM (1997) Molecular dissection of the AGAMOUS control region shows that cis elements for spatial regulation are located intragenically. Plant Cell 9:355–365
Song Y, Ji D, Li S, Wang P, Li Q, Xiang F (2012) The dynamic changes of DNA methylation and histone modifications of salt responsive transcription factor genes in soybean. PLoS One 7:e41274
Steward N, Kusano T, Sano H (2000) Expression of ZmMET1, a gene encoding a DNA methyltransferase from maize, is associated not only with DNA replication in actively proliferating cells, but also with altered DNA methylation status in cold-stressed quiescent cells. Nucleic Acids Res 28:3250–3259
Steward N, Ito M, Yamaguchi Y, Koizumi N, Sano H (2002) Periodic DNA methylation in maize nucleosomes and demethylation by environmental stress. J Biol Chem 277:37741–37746
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Tan MP (2010) Analysis of DNA methylation of maize in response to osmotic and salt stress based on methylation-sensitive amplified polymorphism. Plant Physiol Biochem 48:21–26
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Tsukahara S, Kobayashi A, Kawabe A, Mathieu O, Miura A, Kakutani T (2009) Bursts of retrotransposition reproduced in Arabidopsis. Nature 461:423–426
Tyunin AP, Kiselev KV, Zhuravlev YN (2012) Effects of 5-azacytidine induced DNA demethylation on methyltransferase gene expression and resveratrol production in cell cultures of Vitis amurensis. Plant Cell Tissue Organ Cult 111:91–100
Verhoeven KJ, Jansen JJ, van Dijk PJ, Biere A (2010) Stress-induced DNA methylation changes and their heritability in asexual dandelions. New Phytol 185:1108–1118
Viejo M, Santamaría ME, Rodríguez JL, Valledor L, Meijón M, Pérez M, Pascual J, Hasbún R, Fernández Fraga M, Berdasco M, Toorop PE, Cañal MJ, Rodríguez Fernández R (2012) Epigenetics, the role of DNA methylation in tree development. Methods Mol Biol 877:277–301
Vining K, Pomraning KR, Wilhelm LJ, Ma C, Pellegrini M, Di Y, Mockler TC, Freitag M, Strauss SH (2013) Methylome reorganization during in vitro dedifferentiation and regeneration of Populus trichocarpa. BMC Plant Biol 13:92
Wang CG, Chen XQ, Li H, Song WS (2007) RNA editing analysis of mitochondrial nad3/rps12 genes in cytoplasmic male sterility and fertile cauliflower (Brassica oleracea var. botrytis) by cDNA-SSCP. Bot Stud 48:13–23
Wang CG, Li H, Xue ZY, Chen CB, Gu Y, Sun DL, Song WQ (2009) Marker-based analysis of genomic structure and DNA methylation in a watermelon (Citrullus lanatus) ploidy series. Bot Stud 50:389–402
Wang X, Wu R, Lin X, Bai Y, Song C, Yu X, Xu C, Zhao N, Dong Y, Liu B (2013a) Tissue culture-induced genetic and epigenetic alterations in rice pure-lines, F1 hybrids and polyploids. BMC Plant Biol 13:77
Wang Y, Wang X, Lee TH, Mansoor S, Paterson AH (2013b) Gene body methylation shows distinct patterns associated with different gene origins and duplication modes and has a heterogeneous relationship with gene expression in Oryza sativa (rice). New Phytol 198:274–283
Weinhold A, Kallenbach M, Baldwin IT (2013) Progressive 35S promoter methylation increases rapidly during vegetative development in transgenic Nicotiana attenuata plants. BMC Plant Biol 13:99
Xiao W, Custard KD, Brown RC, Lemmon BE, Harada JJ, Goldberg RB, Fischer RL (2006) DNA methylation is critical for Arabidopsis embryogenesis and seed viability. Plant Cell 18:805–814
Xie Z, Johansen LK, Gustafson AM, Kasschau KD, Lellis AD, Zilberman D, Jacobsen SE, Carrington JC (2004) Genetic and functional diversification of small RNA pathways in plants. PLoS Biol 2:e104
Xiong LZ, Xu CG, Saghai Maroof MA, Zhang Q (1999) Patterns of cytosine methylation in an elite rice hybrid and its parental lines, detected by a methylation-sensitive amplification polymorphism technique. Mol Gen Genet 261:439–446
Yoshida H, Akimoto H, Yamaguchi M, Shibata M, Habu Y, Iida S, Ozeki Y (2004) Alteration of methylation profiles in distinct cell lineages of the layers during vegetative propagation in carnations (Dianthus caryophyllus). Euphytica 135:247–253
Yu A, Lepère G, Jay F, Wang J, Bapaume L, Wang Y, Abraham AL, Penterman J, Fischer RL, Voinnet O, Navarro L (2013a) Dynamics and biological relevance of DNA demethylation in Arabidopsis antibacterial defense. Proc Natl Acad Sci USA 110:2389–2394
Yu Y, Yang X, Wang H, Shi F, Liu Y, Liu J, Li L, Wang D, Liu B (2013b) Cytosine methylation alteration in natural populations of Leymus chinensis induced by multiple abiotic stresses. PLoS One 8:e55772
Zhao Y, Yu S, Xing C, Fan S, Song M (2008) Analysis of DNA methylation in cotton hybrids and their parents. Mol Biol 42:195–205
Zhong S, Fei Z, Chen YR, Zheng Y, Huang M, Vrebalov J, McQuinn R, Gapper N, Liu B, Xiang J, Shao Y, Giovannoni JJ (2013) Single-base resolution methylomes of tomato fruit development reveal epigenome modifications associated with ripening. Nat Biotechnol 31:154–159
Zubko E, Gentry M, Kunova A, Meyer P (2012) De novo DNA methylation activity of methyltransferase 1 (MET1) partially restores body methylation in Arabidopsis thaliana. Plant J 71:1029–1037
Acknowledgments
We greatly thank Dr. Deling Sun at Tianjin Kernel Vegetable Research Institute, China for kindly providing the cauliflower seeds. This work was performed with financial support from The Natural Science Foundation from The National Natural Science Foundation of China (No. 31100234) and Tianjin (No. 10JCZDJC17900).
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Li, H., Geng, M., Liu, Q. et al. Characteristics of cytosine methylation status and methyltransferase genes in the early development stage of cauliflower (Brassica oleracea L. var. botrytis). Plant Cell Tiss Organ Cult 117, 187–199 (2014). https://doi.org/10.1007/s11240-014-0432-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11240-014-0432-5