Abstract
D-myo-inositol-3-phosphate synthase (MIPS) catalyzes the reaction from d-glucose 6-phosphate to D-myo-inositol 3-phosphate (MIP), which is the first and rate-limiting step in myo-inositol biosynthesis. In this study, Jatropha curcas MIPS cDNA (JcMIPS) (GenBank accession no. EF 185781) has been isolated using mRNA differential display technology (DDRT) and the rapid amplification of cDNA ends (RACE). The cDNA clone of JcMIPS is comprised of 1,957 bp, encoding 509 amino acids, with a predicted molecular weight of 56.4 kDa. The JcMIPS protein is highly homologous to those from other plant species, ranging from 88.4 to 91.18% homology at the amino acid levels. Real-time quantification polymerase chain reaction (PCR) analysis has revealed that JcMIPS transcripts are highly present in seed and leaf tissues, but are at low levels in stem and flower tissues. Furthermore, the transcription of JcMIPS in leaves is up-regulated by abscisic acid (ABA) (100 μM), drought (30% PEG-6000), NaCl (200 mM), and low-temperature (4°C) treatments. The observed increase of JcMIPS enzyme activity is also detected following treatments with ABA, drought, and NaCl. Interestingly, JcMIPS enzyme activity is only slightly changed following low-temperature treatment.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Abiotic stress, including drought, high salinity, extreme temperatures, and heavy metal toxicity, limits plant development. Plants resist abiotic stress by changing the morphological, biochemical, and metabolic processes (Ingram and Bartels 1996; Lokhande et al. 2010). All of these changes result from the changes of genes expression. Although a large number of genes have been recently identified, such as GmDREB1 (Jin et al. 2010) and PgCDPK1a (Kiselev et al. 2010), their physiological roles in relation to stress are largely unknown in plants (Ingram and Bartels 1996; Seki et al. 2001, 2002).
Jatropha curcas belongs to Euphorbiaceae and it is distributed in many tropical and subtropical areas (He et al. 2009). It can be used as a biological pesticide and in various medicinal purposes. Recently, J. curcas has drawn much attention for its toxicity and high content of seed oil (Singh et al. 1984; Muanza et al. 1995; Misra et al. 2010) and could confer drought and heavy metal tolerance (Zhang et al. 2007; Juwarkar et al. 2008). However, the mechanism of its tolerance to stress remains poorly understood.
Myo-inositol 1-phosphate synthase (MIPS) catalyzes the conversion of d-glucose 6-phosphate to 1-myo-inositol-1-phosphate, the first and rate-limiting step in the biosynthesis of all inositol-containing compounds (Abreu and Aragão 2007). MIPS is a highly conserved enzyme. The binding motif site, which is required for catalytic reaction, is similarly found in amino acid sequences from diverse sources such as bacteria, green algae, fungi, higher plants, and animals (Majumder et al. 2003). MIPS genes have been isolated and characterized from a number of plant species and found to be representing multigene families in some plant species (Ju et al. 2004; Park and Kim 2004). The regulation of MIPS has also been studied in some higher plants. For example, transcripts encoding MIPS have been shown to be up-regulated during osmosis stress in Mesembryanthemum crystallinum (Ishitani et al. 1996). The activity of MIPS enzyme in Oryza sativa L. could be enhanced by salinity (Raychaudhuri and Majumder 1996).
There has been limited molecular information about the MIPS gene in J. curcas so far. In this study, we have cloned and characterized MIPS cDNA from seedlings under drought stress and studied its differential expression in different tissues and under diverse stresses. Furthermore, JcMIPS enzyme activity under abiotic stress and exogenous abscisic acid (ABA) treatment is also discussed.
Materials and methods
Plant materials and stress treatments
The seeds of J. curcas were collected from Panzhihua, Sichuan Province, China. Seeds were sterilized by 0.1% HgCl2 and germinated in seedbeds containing a 1:1:1 (v/v/v) mixture of perlite, peat moss, and sandy loam soil, and irrigated daily with a half-strength Hoagland solution, as described by Saneoka et al. (2001). After the seeds germinated, they were transferred to flower pots and grown at 25°C with a photoperiod of 18 h light/6 h dark. Two weeks later, plants were selected to be used for the stress treatment.
Drought and salinity stress treatments were carried out by incubating the seedlings in 100 ml of Murashige–Skoog (MS) liquid medium, containing 30% PEG-6000 and 200 mM NaCl at 25°C for 24, 48, and 72 h, respectively. The degree of water stress was determined by the relative water content (RWC) of the plants. The RWC was measured as described by Ma et al. (2006). The seedlings were incubated at 4°C for 72 h for the low-temperature treatment. For exogenous ABA treatment (ABA dissolved in methanol and 2.5% methanol used as a control), the seedlings were carried out by incubating plants in MS medium containing 100 μM ABA at 25°C for 24 h. These plant tissues were immediately frozen in liquid nitrogen and kept at −80°C until use.
RNA isolation and mRNA differential display
To search for the differentially expressed cDNAs, mRNA differential display analysis was performed as described by Liang and Pardee (1992), with slight modifications. Total RNA was extracted from 2-week-old light-grown J. curcas leaves subjected to 72-h drought stress using plant RNA isolation reagent (QIAGEN RNeasy Plant Mini Kit, QIAGEN, Holland) following the manufacturer’s instructions. Total RNA from untreated leaves was used as a control. To remove contaminating DNA, isolated RNA was treated with RNase-free DNase, and then used for reverse transcription polymerase chain reaction (RT-PCR).
RT-PCR was followed by PCR amplification using three anchored primers (APs) and each of the ten arbitrary primers (ARPs). The primers are shown in Table 1. Twenty-five cycles of PCR were carried out, each consisting of 3 min at 95°C, 30 s at 94°C, 2 min at 42°C, and 1 min at 72°C. Then, ten cycles of PCR were carried out, each consisting of 30 s at 94°C, 1 min at 50°C, and 1 min at 72°C, with an additional 10 min at 72°C in an automatic thermal cycler (Bio-Rad, USA). PCR products were separated on 6% polyacrylamide sequencing gels and visualized by silver staining. Bands of interest were recycled from the gels and reamplified with M13 primer and T7 primer.
Reverse Northern blot, 5′-RACE, and RT-PCR amplification
Reverse Northern blot was carried out to reduce false-positives arising from the differential display. Each reamplification mixture was dotted on Hybond N+ membrane filter (Schleicher & Schuell, Dassel, Germany). Total cDNA probes were synthesized by reverse transcription of 2 μg total RNA for drought-treated and untreated J. curcas leaves. Hybridization was performed following the manufacturer’s instructions (DIG High Prime DNA Labeling and Detection Starter Kit II, Roche, Switzerland). The cDNA fragments were cloned into the pMD18-T Easy vectors and were sequenced (Invitrogen, China). One of these differential display segments shared high similarity to other plant MIPS. The full length of cDNA was cloned by RT-PCR and 5′-rapid amplification of cDNA ends (5′-RACE). The sequences of all primers used are listed in Table 2.
Real-time quantification PCR analysis
SuperScript™ II Reverse Transcriptase (Invitrogen, USA) was used to transcribe 2.5 μg total plant RNA into cDNA. J. curcas 18S ribosomal RNA (GenBank accession no. AY823528) was used as an internal standard in the real-time PCR reaction. All primers are as listed in Table 3.
Real-time PCR was carried out in a BioRad iCycler using SYBR Green qPCR Supermix-UDG (Invitrogen, Carlsbad, USA). The program was as following: 45 cycles of 2 min at 50°C, 5 min at 95°C, 15 s at 55°C, 20 s at 72°C, and the data collection during the extension step (3 min at 72°C). Melting curves were run immediately after the last cycle to exclude any influence of primer–dimer pairs. Cycle numbers at which the fluorescence passed the cycle threshold (Ct) were further analyzed using the DDRT method and the Relative Expression Software Tool (REST) (Pfaffl et al. 2002).
For the amplification of rare mRNA species, 1 μg of total RNA was reverse transcribed using the iScript Select cDNA Synthesis Kit (Bio-Rad) with an oligo dT primer and enhancer according to the manufacturer’s protocol. PCR products from templates of leaves of unstressed and stressed plants were compared by standard agarose gel electrophoresis.
Preparation of crude enzyme and enzyme assay
Root, stem, leaf, flower, and seed (5 g of each) tissues were homogenized in 50 ml of ice-cold buffer A (20 mM Tris–HCl pH 7.5, 10 mM NH4Cl, 10 mM β-mercaptoethanol, 2 mM PMSF). The homogenate was centrifuged at 8,000g at 4°C for 45 min. The supernatant was stored at 4°C for protein concentration and the JcMIPS enzyme activity assay. The enzyme was assayed colorimetrically by the periodate oxidation method (Barnett et al. 1970). One unit of enzyme activity is defined as the μmol amount of 1 mg protein converting the substance d-glucose-6-phosphate to inositol-1-phosphate during 1 h (μmol h−1 mg−1). Protein was determined according to the method of Bradford (1976) with BSA as a standard.
Results
Screening the differential expression genes under drought stress by DDRT-PCR
Some partial cDNA fragments were identified by mRNA differential display technology (DDRT-PCR) induced by water deficit from leaves of 2-week-old light-grown J. curcas plants (Fig. 1). A 527-bp fragment that showed significant positive signal in reverse Northern blot was selected for further investigation. Nucleotide BLAST search suggested that the isolated cDNA fragment shared high similarity with other known MIPS genes, indicating that the partial MIPS gene of J. curcas (JcMIPS) was obtained (data not shown).
The partial results of mRNA differential display PCR. a Electrophoresis of mRNA differential display PCR between control and treatment (1, 3, 5, 7: control; 2, 4, 6, 8: treatment). b Electrophoresis of denaturing polyacrylamide gel of mRNA differential display PCR between control and treatment. The arrows indicate different bands of PCR-amplified cDNA fragments between control and treatment. c Reverse Northern blot analysis between control and treatment. The arrows indicate the positive dots
Cloning and characterization of the full-length cDNA encoding JcMIPS
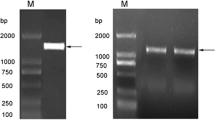
A J. curcas MIPS cDNA (JcMIPS) was isolated from J. curcas seedlings using the DDRT-PCR and RACE systems. The 1,957-bp cDNA sequence contained a 1,530-bp coding sequence that could encode a protein of 509 amino acids, which correspond to a polypeptide with a predicted molecular weight (MW) of 56.4 kD (Fig. 2). The nucleotide sequence of the JcMIPS gene was deposited in the GenBank database under accession number EF 185781.
Sequence alignment showed that JcMIPS shared a high similarity to the other plant MIPS from Glycine max (89.22%, AY038802), Citrus paradisi (88.63%, Z32632), Phaseolus vulgar (88.63%, AJ853494), Brassica napus (88.43%, FJ429169), and Arabidopsis thaliana (87.08%, U04876). The JcMIPS protein had the highest identity with Nicotiana paniculata MIPS (91.18%, AB032073) (Fig. 3).
Alignment of the amino acid sequences of myo-inositol-1L-phosphate synthase from Jatropha curcas (EF185781), Glycine max (AY038802), Citrus paradisi (Z32632), Phaseolus vulgar (AJ853494), Brassica napus (FJ429169), Arabidopsis thaliana (U04876), and Nicotiana paniculata (AB032073). The four highly conserved motifs (I, II, III, and IV) are indicated by the gray box outline
In addition, four highly conserved motifs similar to those of other plant MIPS were also found in the JcMIPS polypeptide. The four conserved domains are GWGGNNG (domain I), LWTANTERY (domain II), NGSPQNTFVPGL (domain III), and SYNHLGNNDG (domain IV) (Fig. 3), which are essential for MIPS functions such as the binding of cofactor NAD+ and catalysis of the reaction (Majumder et al. 1997; Bachhawat and Mande 1999).
Tissue-specific expression of the JcMIPS mRNA in J. curcas
The expression pattern of JcMIPS in various tissues of J. curcas, the JcMIPS gene, was transcriptionally active in all of the five detected tissues at different levels (Fig. 4). The mRNA levels of JcMIPS were high in leaf and seed, but lower in root and flower and the lowest in the stem. The mRNA level of JcMIPS in leaf was nearly eight times higher than that in stem. The expression pattern of JcMIPS demonstrated that this gene was differentially expressed in various tissues.
Expression of the JcMIPS gene in different tissues of J. curcas (given as relative % yield of mRNA level normalized to 18S rRNA). Total RNAs were isolated from different tissues of J. curcas and 1 μg of total RNA was used for the real-time PCR assay. The cDNA fragments were amplified by PCR reaction with primers for the JcMIPS gene and 18S rRNA
Expression analysis of the JcMIPS gene and JcMIPS enzyme activity under abiotic stress and exogenous ABA treatment
Expression patterns of the JcMIPS gene were studied in leaves under abiotic stress and exogenous ABA treatment. When exposed to 30% PEG 6000, 200 mM NaCl, or 4°C cold stress, JcMIPS transcripts in J. curcas seedlings were all up-regulated clearly (Fig. 5a). The mRNA levels of JcMIPS under drought and salt treatments were higher (24 and 36%, respectively) than that of the control. However, the expression of JcMIPS in response to cold treatment was lower and increased only 15% after the seedlings were exposed to a 4°C environment for 72 h. In order to study the transcription level of JcMIPS after exogenous hormone treatment, J. curcas seedlings were treated by 100 μM exogenous ABA at 25°C for 24 h. Figure 5a shows that the transcription level reached 1.56 times that of the control. The data indicates that the expression of JcMIPS was related to drought, salt and cold stress, and exogenous ABA treatment.
Expression analysis of JcMIPS mRNA and changes of enzyme activity under abiotic stress and exogenous ABA treatment. a Expression of the JcMIPS gene in leaves of J. curcas treated by drought (30% PEG-6000, 24 h), salt (200 mM NaCl, 24 h), cold (4°C, 72 h), and ABA (100 μM ABA, 24 h). Total RNAs were isolated from different leaves of J. curcas exposed to different stress treatment, and 1 μg of total RNA was used for real-time PCR. The cDNA fragments were amplified by PCR reaction with primers for the JcMIPS gene and 18S rRNA. The transcript levels of the JcMIPS at each treatment were plotted as percentages of the control. b The changes of activity of JcMIPS enzyme in leaves of J. curcas in control and those treated by different stress treatments (drought, salt, cold, and exogenous ABA). All of the presented data are averages and standard errors (bars) of three independent experiments. The double asterisks indicate that the treatment was significantly different from the control at P < 0.01. The single asterisk indicates that the treatment was different from the control at P < 0.05
The JcMIPS enzyme activity in J. curcas leaf under different stress treatments and ABA induction had been detected. The results showed that JcMIPS enzyme activity was also induced by drought, salt, and exogenous ABA treatment (Fig. 5b). The activity of JcMIPS increased 1.15 and 1.52 times when exposed to 30% PEG and 200 mM NaCl, respectively, for 24 h. However, it seemed that the JcMIPS enzyme activity was not sensitive to cold treatment. There was only a 10% increase when exposed to 4°C for 72 h. Interestingly, the activity of the JcMIPS enzyme suffered sharply from the induction of ABA and increased 2.16 times more than control plants after 100 μM ABA treatment on J. curcas seedlings for 24 h.
Time courses of mRNA expression and enzymatic activity changes of JcMIPS in response to drought stress
Figure 6a shows that the average value of RWC of the control group leaves is 77.6%. This index decreased to 64, 57, and 52% after 24, 48, and 72 h of drought stress treatment, but recovered to 62% quickly after one day of re-watering. Figure 6b shows that the mRNA level of JcMIPS in J. curcas leaf increased gradually with the intensification of drought stress degree and reached 1.24 and 1.45 times that of the control in the first two days, but it decreased on the third day, and nearly decreased to the control level after re-watering treatment. The changes of JcMIPS enzyme activity in J. curcas leaves along with the stress degree were detected (Fig. 6c). The results suggested that JcMIPS activities in J. curcas leaves were further enhanced as plants were exposed to more severe water loss. JcMIPS enzyme activity began to rise slowly (114.7%) at 24 h and continued to rise after 48 h of treatment (124.2%), reaching the peak at 72 h, which was nearly one times higher than the control. After one day of re-watering, it decreased quickly and was similar to the level after 24 h of treatment.
Time courses of JcMIPS mRNA expression and activity changes of JcMIPS enzyme in response to drought stress. D24, D48, and D72 represent drought treatment for 24, 48, and 72 h, respectively; R represents re-watering after drought treatment for 72 h. a Changes of the relative water content (RWC) of J. curcas leaves at different stress treatments. The presented data are averages and standard errors (bars) of three independent experiments. b Expression of the JcMIPS gene upon different drought level treatments. The transcript levels of the JcMIPS at each treatment were plotted as percentages of the control. c The enzymatic activities of JcMIPS in the leaves of J. curcas as control and treated by drought. All of the presented data are averages and standard errors (bars) of three independent experiments. The double asterisks indicate that the treatment was significantly different from the control at P < 0.01. The single asterisk indicates that the treatment was different from the control at P < 0.05
Discussion
In this study, the cloning of JcMIPS and its expression in response to stress tolerance in J. curcas provided opportunities for developing J. curcas lines capable of growing on poor and marginal soils and yielding acceptable levels of oil. Furthermore, JcMIPS can also be used for the genetic improvement of other agriculturally important crops for tolerance to salt and drought stresses (Eswaran et al. 2010).
The expression pattern was shown to be organ-specific in different plant species, such as sesame (Chun et al. 2003). In soybean, MIPS1 was highly expressed in developing seeds, but it was not expressed or minimally expressed in flowers, leaves, and roots (Chappell et al. 2006). Our data suggested that JcMIPS transcription was the highest in seed and the lowest in stem. The MIPS gene in different organs belonged to the multigene family. They may play different functions in plants but would coordinate inositol metabolism with cellular growth (Ishitani et al. 1996; Majumder et al. 1997).
The studies revealed that the transcription of MIPS could be induced by different abiotic stresses (Ishitani et al. 1996; Abreu and Aragão 2007). Our results showed that JcMIPS transcription would be up-regulated by drought, salt, and low-temperature stress, and it is the most sensitive to salt stress. However, JcMIPS transcription, but not the enzyme activity, is slightly induced by low-temperature treatment. Luo et al. (2005) showed that the seedlings of J. curcas had a lower tolerance to low temperature. The results indicated that JcMIPS may participate in the response of drought stress but not cold stress.
JcMIPS transcripts would increase after drought stress, but decreased dramatically after 24 h of re-watering. The changes of gene expression at the enzymatic level also are consistent with the transcriptional level. The RWC in J. curcas leaf continued decreasing with the prolongation of stress treatment. When the drought stress degree was aggravated, the JcMIPS gene transcription and protein translation level would gradually increase. The RWC increased in leaves, indicating that the stress degree was mitigating. At the same time, the JcMIPS transcription level and enzymatic activity would decrease. So, JcMIPS was closely related to J. curcas in response to water stress.
Most of the stress-responsive genes, including the MIPS gene, have been found to be induced by exogenous application of ABA, which plays an important role in signal transduction pathways responding to abiotic stress. The endogenous ABA content would increase when plants perceived drought and salt stress, which mediate rapid physiological response through gene transcription (Bray 1997; Leung and Giraudat 1998). Our research is similar to the previous results. JcMIPS is induced fiercely by exogenous ABA. The up-regulation degree of JcMIPS induced by ABA treatment is higher than the other stresses.
In summary, we have identified the MIPS gene in J. curcas and studied its expression pattern. The data showed that the JcMIPS gene was highly conserved at the amino acid levels compared to other plants, and the differential expressions of JcMIPS and enzyme activity were induced by abiotic stress and ABA treatment. Furthermore, the copy number of the JcMIPS gene would be investigated, which may be of fundamental importance in order to understand this gene across phylogenetic groups.
References
Abreu EFM, Aragão FJ (2007) Isolation and characterization of a myo-inositol-1-phosphate synthase gene from yellow passion fruit (Passiflora edulis f. flavicarpa) expressed during seed development and environmental stress. Ann Bot 99:285–292
Barnett JEG, Brice RE, Corina DL (1970) A colorimetric determination of inositol monophosphates as an assay for d-glucose 6-phosphate–1L-myoInositol 1-phosphate cyclase. Biochem J 119:183–186
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Bray EA (1997) Plant responses to water deficit. Trends Plant Sci 2:48–54
Chappell AS, Scaboo AM, Wu X, Nguyen H, Pantalone VR, Bilyeu KD (2006) Characterization of the MIPS gene family in Glycine max. Plant Breed 125:493–500
Chun JA, Jin UH, Lee JW, Yi YB, Hyung NI, Kang MH et al (2003) Isolation and characterization of a myo-inositol 1-phosphate synthase cDNA from developing sesame (Sesamum indicum L.) seeds: functional and differential expression, and salt-induced transcription during germination. Planta 216:874–880
Eswaran N, Parameswaran S, Sathram B, Anantharaman B, Kumar GRK, Tangirala SJ (2010) Yeast functional screen to identify genetic determinants capable of conferring abiotic stress tolerance in Jatropha curcas. BMC Biotechnol 10:23
He Y, Guo XL, Lu R, Niu B, Pasapula V, Hou P, Cai F, Xu Y, Chen F (2009) Changes in morphology and biochemical indices in browning callus derived from Jatropha curcas hypocotyls. Plant Cell Tiss Organ Cult 98:11–17
Ingram J, Bartels D (1996) The molecular basis of dehydration tolerance in plants. Ann Rev Plant Physiol Plant Mol Biol 47:377–403
Ishitani M, Majumder AL, Bornhouser A, Michalowski CB, Jensen RG, Bohnert HJ (1996) Coordinate transcriptional induction of myo-inositol metabolism during environmental stress. Plant J 9:537–548
Jin TC, Chang Q, Li WF, Yin DX, Li ZJ, Wang DL, Liu B, Liu LX (2010) Stress-inducible expression of GmDREB1 conferred salt tolerance in transgenic alfalfa. Plant Cell Tiss Organ Cult 100:219–227
Ju S, Shaltiel G, Shamir A, Agam G, Greenberg ML (2004) Human 1-D-myo-inositol-3-phosphate synthase is functional in yeast. J Biol Chem 279:21759–21765
Juwarkar AA, Yadav SK, Kumar P, Singh SK (2008) Effect of biosludge and biofertilizer amendment on growth of Jatropha curcas in heavy metal contaminated soils. Environ Monit Assess 145:7–15
Kiselev KV, Turlenko AV, Zhuravlev YN (2010) Structure and expression profiling of a novel calcium-dependent protein kinase gene PgCDPK1a in roots, leaves, and cell cultures of Panax ginseng. Plant Cell Tiss Organ Cult 103:197–204
Leung J, Giraudat J (1998) Abscisic acid signal transduction. Ann Rev Plant Physiol Plant Mol Biol 49:199–222
Liang P, Pardee AB (1992) Differential display of eukaryotic messenger RNA by means of the polymerase chain reaction. Science 257:967–971
Lokhande VH, Nikam TD, Penna S (2010) Biochemical, physiological and growth changes in response to salinity in callus cultures of Sesuvium portulacastrum L. Plant Cell Tiss Organ Cult 102:17–25
Luo T, Ma DW, Deng WY, Chen F (2005) Effect of low temperature on physiological indexes of Jatropha curcas. Chin J Oil Crop Sci 27(4):50–54
Ma QQ, Wang W, Li YH, Li DQ, Zou Q (2006) Alleviation of photoinhibition in drought-stressed wheat (Triticum aestivum) by foliar-applied glycinebetaine. J Plant Physiol 163:165–175
Majumder AL, Johnson MD, Henry SA (1997) 1L-myo-inositol-1-phosphate synthase. Biochim Biophys Acta 1348:245–256
Majumder AL, Chatterjee A, Ghosh Dastidar K, Majee M (2003) Diversification and evolution of L-myo-inositol 1-phosphate synthase. FEBS Lett 553:3–10
Misra P, Gupta N, Toppo DD, Pandey V, Mishra MK, Tuli R (2010) Establishment of long-term proliferating shoot cultures of elite Jatropha curcas L. by controlling endophytic bacterial contamination. Plant Cell Tiss Organ Cult 100:189–197
Muanza DN, Euler KL, Williams L, Newman DJ (1995) Screening for antitumor and anti-HIV activities of nine medicinal plants from Zaire. Int J Pharmacogn 33:98–106
Park SH, Kim JI (2004) Characterization of recombinant Drosophila melanogaster myo-inositol-1-phosphate synthase expressed in Escherichia coli. J Microbiol 42:20–24
Pfaffl MW, Horgan GW, Dempfle L (2002) Relative expression software tool (REST) for group-wise comparison and statistical analysis of relative expression results in real-time PCR. Nucleic Acids Res 30:e36
Raychaudhuri A, Majumder AL (1996) Salinity-induced enhancement of L-myo-inositol 1-phosphate synthase in rice (Oryza sativa L). Plant Cell Environ 19:1437–1442
Saneoka H, Ishiguro S, Moghaieb REA (2001) Effect of salinity and abscisic acid on accumulation of glycinebetaine and betaine aldehyde dehydrogenase mRNA in Sorghum leaves (Sorghum bicolor). J Plant Physiol 158:853–859
Seki M, Narusaka M, Abe H, Kasuga M, Yamaguchi-Shinozaki K, Carninci P, Hayashizaki Y, Shinozaki K (2001) Monitoring the expression pattern of 1300 Arabidopsis genes under drought and cold stresses by using a full-length cDNA microarray. Plant Cell 13:61–72
Seki M, Narusaka M, Ishida J, Nanjo T, Fujita M, Oono Y, Kamiya A, Nakajima M, Enju A, Sakurai T, Satou M, Akiyama K, Taji T, Yamaguchi-Shinozaki K, Carninci P, Kawai J, Hayashizaki Y, Shinozaki K (2002) Monitoring the expression profiles of 7000 Arabidopsis genes under drought, cold and high-salinity stresses using a full-length cDNA microarray. Plant J 31:279–292
Singh YN, Ikahihifo T, Panuve M, Slatter C (1984) Folk medicine in Tonga. A study of the use of herbal medicines for obstetric and gynaecological conditions and disorders. J Ethnopharmacol 12:305–329
Zhang Y, Wang YX, Jiang LD, Xu Y, Wang YC, Lu DH, Chen F (2007) Aquaporin JcPIP2 is involved in drought responses in Jatropha curcas. Acta Biochim Biophys Sin 39(10):787–794
Acknowledgments
This study was supported by grants from the National Science & Technology Pillar Program (no. 2006BAD07A04), the National Natural Science Foundation of China (no. 31071448), and the International Cooperation Program (no. 2006 DFB63400).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wang, Y., Huang, J., Gou, C.B. et al. Cloning and characterization of a differentially expressed cDNA encoding myo-inositol-1-phosphate synthase involved in response to abiotic stress in Jatropha curcas . Plant Cell Tiss Organ Cult 106, 269–277 (2011). https://doi.org/10.1007/s11240-011-9917-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11240-011-9917-7