Abstract
Traditional statistical models allow population based inferences and comparisons. Machine learning (ML) explores datasets to develop algorithms that do not assume linear relationships between variables and outcomes and that may account for higher order interactions to make individualized outcome predictions. To evaluate the performance of machine learning models compared to traditional risk stratification methods for the prediction of major adverse cardiovascular events (MACE) and bleeding in patients with acute coronary syndrome (ACS) that are treated with antithrombotic therapy. Data on 24,178 ACS patients were pooled from four randomized controlled trials. The super learner ensemble algorithm selected weights for 23 machine learning models and was compared to traditional models. The efficacy endpoint was a composite of cardiovascular death, myocardial infarction, or stroke. The safety endpoint was a composite of TIMI major and minor bleeding or bleeding requiring medical attention. For the MACE outcome, the super learner model produced a higher c-statistic (0.734) than logistic regression (0.714), the TIMI risk score (0.489), and a new cardiovascular risk score developed in the dataset (0.644). For the bleeding outcome, the super learner demonstrated a similar c-statistic as the logistic regression model (0.670 vs. 0.671). The machine learning risk estimates were highly calibrated with observed efficacy and bleeding outcomes (Hosmer–Lemeshow p value = 0.692 and 0.970, respectively). The super learner algorithm was highly calibrated on both efficacy and safety outcomes and produced the highest c-statistic for prediction of MACE compared to traditional risk stratification methods. This analysis demonstrates a contemporary application of machine learning to guide patient-level antithrombotic therapy treatment decisions.
Clinical Trial Registration ATLAS ACS-2 TIMI 46: https://clinicaltrials.gov/ct2/show/NCT00402597. Unique Identifier: NCT00402597. ATLAS ACS-2 TIMI 51: https://clinicaltrials.gov/ct2/show/NCT00809965. Unique Identifier: NCT00809965. GEMINI ACS-1: https://clinicaltrials.gov/ct2/show/NCT02293395. Unique Identifier: NCT02293395. PIONEER-AF PCI: https://clinicaltrials.gov/ct2/show/NCT01830543. Unique Identifier: NCT01830543.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Highlights
-
Approximately 15% of acute coronary syndrome patients experience a recurrent event in 1 year.
-
Identification of those who would benefit from intensified antithrombotic strategies is important.
-
This analysis was the first to evaluate the performance of machine learning algorithms for prediction of major adverse cardiovascular and bleeding events in acute coronary syndrome patients.
-
Compared to the TIMI risk score and a surrogate risk score, developed in this dataset, machine learning modestly improved the c-statistic for MACE.
-
Importantly, the machine learning model demonstrated remarkable calibration on both efficacy and safety outcomes.
-
This analysis demonstrates a contemporary application of machine learning models to assist in clinical decision making by concurrently stratifying bleeding and thrombotic risk.
Introduction
Approximately 15% of acute coronary syndrome (ACS) patients experience recurrent cardiovascular (CV) events within 1 year [1,2,3]. Individualized patient-level prediction of major adverse cardiovascular events (MACE) and bleeding events among patients with ACS may help to identify those who would benefit from intensified antithrombotic strategies and finally, tailor a personalized approach [4,5,6]. Traditional statistical methods allow inferences to be made regarding populations, but are less powerful than machine learning in making predictions regarding individual patients. Traditional risk stratification scores are built using parametric and semi-parametric regression scoring systems that have several limitations, including primary reliance on linear models and limited capability in explorations of higher order interactions [7, 8]. This is particularly true among patients with extreme risk profiles, as the underlying parametric assumption extrapolates their risk profiles based on population means. These limitations result in sub-optimal performance of traditional risk prediction scores.
In contrast, machine learning explores large datasets and uses algorithms that can learn from and make predictions on data. Additionally, these models have built-in functionality for variable selection and do not require pre-specification of interaction terms. Over the past decade, machine learning techniques have made substantial advances in many domains, including health care [9, 10]. Thus, recent evidence suggests that machine learning methods may offer a powerful alternative to the conventional methods for risk predictions. However, both the accuracy and the application of machine learning models to predict clinical outcomes in individual ACS patients remain unknown. The aim of this analysis was to evaluate the performance of machine learning models to predict the occurrence of MACE and bleeding events among ACS patients, compared to traditional risk stratification methods and to explore contemporary applications of machine learning models in individualized cardiovascular risk prediction.
Methods
Source of data
Data from 24,178 patients who received at least one dose of study drug were pooled from four large randomized clinical trials. The ATLAS ACS-TIMI 46 and 51 trials enrolled 3491 and 15,526 adult patients with ACS, respectively. Patients were randomized to receive rivaroxaban or placebo in combination with either aspirin alone or dual antiplatelet therapy (DAPT) consisting of aspirin plus a P2Y12 inhibitor. The GEMINI ACS trial enrolled 3037 patients with ACS who were on a P2Y12 inhibitor and randomized them to receive either rivaroxaban or aspirin. Finally, the PIONEER AF-PCI trial enrolled 2124 patients with atrial fibrillation who underwent PCI. Patients were randomized to receive either dual or triple rivaroxaban-based antithrombotic regimens or Vitamin K Antagonist (VKA)-based triple therapy. The study designs and primary results for each of the four randomized clinical trials have been previously published [11,12,13,14].
Outcomes
The efficacy outcome was the composite of cardiovascular (CV) death, myocardial infarction (MI), or stroke within 6 months of randomization. The safety outcome was the composite of TIMI (thrombolysis in myocardial infarction) major and minor bleeding and bleeding requiring medical attention within 6 months of trial randomization.
Predictive models
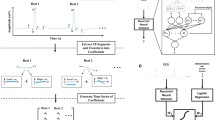
Ensemble learning is a type of machine learning that combines predictions across different candidate models. The Super Learner ensemble method uses cross validation to select weights applied to each candidate model. A total of 23 machine learning algorithms were built using tenfold cross validation. Broadly, the families of candidate models built included Generalized Additive Models (GAMs), Elastic Net (Penalized Logistic Regression), Gradient Boosted Machines (GBMs), Random Forests, a Bayesian logistic regression with default priors, and a naïve Bayes classification model. A total of 48 variables were used for model building (Supplemental Table S1).
For the MACE endpoint, the Super Learner ensemble was compared to three traditional risk stratification tools: (1) TIMI risk score (2) a Surrogate risk score based on the dataset using traditional statistical methods and (3) Stepwise logistic regression. The surrogate model was created by first randomly splitting the data 50/50 into training and test sets, then fitting a logistic regression model to 14 variables thought, a priori, to have the most predictive importance. For the bleeding endpoint, the Super Learner ensemble was compared to a stepwise logistic regression. The detailed statistical methods of these comparator models are described in the supplemental appendix.
Performance measures
Predictive performance was measured in two ways: (1) the ability of the model to discriminate between outcome classes, and (2) the accuracy of the methods probabilistic predictions, called calibration. Discrimination was assessed with a cross-validated concordance statistic (c-statistic). C-statistic comparisons were based on a bootstrapped test of significance. Calibration represents the reliability of models by assessing how closely the predicted risk estimate of a particular patient correlates to the observed event rate for this patient. In this analysis, calibration was assessed via high-resolution non-parametric calibration plots. In the calibration plots, the diagonal line represents perfect calibration with perfect correlation of predicted estimates with observed event rates. Deviations above the diagonal line represent a model that underestimates risk and deviations below the diagonal line represent a model that overestimates risk. The Hosmer–Lemeshow goodness of fit test was used to test for statistical significance between the model and the perfect calibration (diagonal) line. A high p-value on this test is favorable and represents no significant difference from perfect calibration whereby a low p-value represents a significant difference from perfect calibration.
Individualized risk predictions
The secondary objective of this study was to explore the ability of the super learner ensemble to produce clinically relevant individual patient risk predictions. This was done by randomly selecting 3000 patients that had both efficacy and safety outcome assessments, and computing risk estimates according to their antithrombotic regimen. The procedure produces four risk estimates for each patient: (1) The predicted probability of a MACE event on rivaroxaban and (2) on the study-specific control regimen, (3) the predicted probability of a bleeding event on rivaroxaban and (4) on the study-specific control regimen. The patient-specific predicted risk of an event on rivaroxaban was plotted against the predicted risk of an event on the study-specific control (Supplemental Fig. S1).
To assess benefit-risk, a two dimensional plot was derived by calculating the difference between the individual patient predicted risk in the control group and predicted risk in the treatment group for both MACE and bleeding for 3000 randomly selected patients. The plot displays the difference in MACE risk estimates on the Y-axis and the difference in bleed risk estimates on the X-axis (Supplemental Fig. S2).
Results
Baseline characteristics
Of the 24,178 pooled patients 22,955 had both ACS and an efficacy outcome assessment and were included in the efficacy dataset. Similarly, 22,936 had both ACS and a safety outcome assessment and were included in the safety dataset.
Baseline characteristics and outcome summary of the patients in the pooled dataset are shown in Table 1. Overall, the mean age was 61.7 years, 49.2% of patients had STEMI, 64.2% of patients were randomized to receive a rivaroxaban-based regimen and 35.8% to a control. Approximately 66% of patients underwent PCI for the index event, 4.2% experienced a MACE event, and 7.5% experienced a TIMI major or minor bleeding event, or bleeding requiring medical attention.
Performance measures
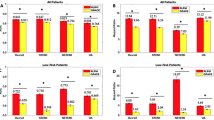
The super learner demonstrated the best discriminative ability for both outcomes achieving a c-statistic of 0.734 for MACE and 0.670 for bleeding (Fig. 1). The best performing candidate model varied according to the outcome. For MACE, the best performing candidate model was the GBM, which achieved a c-statistic of 0.714. For the safety outcome, the best performing candidate model was the Elastic Net, which achieved a c-statistic of 0.669.
Receiver operator characteristics curve. a Shows the receiver operator characteristics curve for the MACE outcome for the super learner ensemble, the best candidate model (GBM), the stepwise logistic regression, the surrogate risk score, and the TIMI risk score. b Shows the receiver operator characteristics Curve for the bleeding outcome for the super learner ensemble, the best candidate model (GBM), and the stepwise logistic regression
The MACE outcome super learner performed significantly better than the TIMI risk score (c-statistic 0.734 vs. 0.489, p < 0.001), the best performing candidate model GBM (c-statistic 0.734 vs. 0.714, p < 0.001), and the surrogate risk score (c-statistic 0.734 vs. 0.644, p < 0.001). The super learner performed similarly to the stepwise logistic regression (0.734 vs. 0.714, p = 0.076). The safety outcome super learner performed similarly to the best candidate model (0.670 vs. 0.669, p = 0.611) and the stepwise regression model (0.670 vs. 0.671, p = 0.946).
Calibration plots suggest that the super learner ensemble demonstrates good calibration for both outcomes (Fig. 2). For the MACE outcome, the super learner calibration plot is close to the perfect calibration line for risk predictions between 0 and 0.3. The Hosmer–Lemeshow test failed to reject good calibration for the super learner (p = 0.612). In contrast, it rejected good calibration for the best performing candidate model (GBM) (p < 0.001), the TIMI risk score (p < 0.001), and stepwise regression model (p < 0.001).
Calibration plots. a Shows the calibration plot for the MACE outcome for the super learner ensemble, the best candidate model (GBM), the stepwise logistic regression, the surrogate risk score, and the TIMI risk score. The 45° diagonal line represents perfect calibration between predicted risk estimates and observed risk. b Shows the calibration plot for the bleeding outcome for the super learner ensemble, the best candidate model (GBM), and the stepwise logistic regression. The 45° diagonal line represents perfect calibration between predicted risk estimates and observed risk
Inspection of the calibration plots for the safety outcome leads to similar conclusions. The super learner demonstrated excellent calibrations for predicted risks between 0 and 0.25. Visually, the best performing candidate model and the stepwise regression were less well-calibrated compared to the super learner ensemble. Formally, the Hosmer–Lemeshow test failed to reject good calibration for the super learner (p = 0.970), the best performing candidate model (Elastic Net) (p = 0.993), and the stepwise regression (p = 0.088).
Individualized risk predictions
The super learner-derived MACE risk estimates were plotted on rivaroxaban vs. control (Fig. 3a). The model predicted that approximately 81% of patients fall above the diagonal line and would have reduced MACE risk on rivaroxaban. Approximately 5% (N = 135) of the 3000 randomly selected patients were below the diagonal line and are predicted to have decreased risk of bleeding with rivaroxaban, compared to the control (Fig. 3b). The combined benefit-risk plot conveys the patient level risk prediction results in a single plot. Using this method, individual differences in treatment benefit or harm may be discerned (Fig. 4).
Individualized predicted risk plot. The points on the predicted risk plot represent a patient’s risk profile for a particular outcome. The diagonal line represents equal risk of the outcome with and without rivaroxaban treatment. Patients below the 45° line are predicted to have lower risk on rivaroxaban versus control. Patients above the 45° line are predicted to have higher risk on rivaroxaban versus control
Individualized benefit-risk plot. The points on the plot represent a patient’s individual predicted benefit-risk profile, based on a combination of that patient’s characteristics. A positive value on the Y-axis represents reduced MACE risk with rivaroxaban treatment and a positive value on the X-axis represents reduced risk of bleed on rivaroxaban, compared to the control arm
Discussion
This analysis is the first report of a machine learning model to predict MACE and bleeding outcomes among patients with ACS enrolled in randomized controlled trials. The super learner ensemble method demonstrated improved performance compared to traditional risk stratification methods. The super learner model improved the c-statistic for predicting ischemic risk compared to the TIMI risk score, a surrogate risk score derived from the dataset, but similar discrimination as the stepwise logistic regression. For bleeding risk, it demonstrated a similar c-statistic as the logistic regression model. The machine learning model produced risk estimates that were highly calibrated with observed efficacy and bleeding outcomes. The calibration of the machine learning model exceeded that of logistic regression and risk scores. This analysis additionally explored a new application of the super learner method to assist the treatment decision making process.
On an individual level, patients and physicians are primarily concerned about the accuracy of a prognostic estimate after an ACS event, and not merely about the overall discrimination of outcomes. Thus, beyond the c-statistic calibration is a key component of risk stratification tools. Indeed, the predicted risk in a poorly calibrated model may over or under estimate the observed event rate. In contrast, in a well-calibrated model, a 5% predicted risk of event corresponds to an observed 5% event rate. Therefore, the most significant advance in this analysis was the high calibration of the super learner model as compared to logistic regression and conventional risk score techniques.
Previous studies have demonstrated inconsistent results regarding the performance of machine learning algorithms as compared to regression models [15,16,17,18,19,20,21,22,23]. In the current analysis, there was a dissociation in the performance of the super learner method in the efficacy endpoint versus the safety endpoint. There may be several reasons for this inconsistency. First, there are numerous types of machine learning models that may fit and perform differently in different datasets. Even among the same types of models, there are countless combinations of tuning parameters that may influence model performance. However, the super learner method provides the advantage of combining any number of models to arrive at the best combined estimate that performs at least as well as the best candidate model. Second, one of the components of the bleeding outcome (bleeding requiring medical attention) is a sensitive endpoint but is non-specific and thus, may be more difficult to predict. Moreover, additional variables that are not captured in the dataset may be associated with the occurrence of an outcome (e.g. genetic or environmental factors) and could improve model performance.
One promise of precision medicine is to identify patients most likely to benefit from treatment and to withhold treatment from those in whom treatment is more likely to cause harm. Notably, the current analysis demonstrates a novel application of machine learning to assist in clinical decision making. Prior traditional analysis methods estimate mean treatment benefits of antithrombotic regimens across populations enrolled in clinical trials. Even if overall classification rates remain similar, this one-size-fits-all approach does not allow for identification of patients more likely to derive larger relative risk reductions from therapy. The machine learning methods employed here incorporate non-linear mappings from exposures to risk such that a differential benefit of rivaroxaban in ACS patients may be identified. The benefit-risk plots of different antithrombotic regimens may be easily visualized and understood by both physicians and patients to facilitate shared decision making. An application for handheld devices can allow real time calculation and display of these results.
A common problem with machine learning models is difficulty in incorporating prior scientific knowledge. It is also difficult to understand exactly how the super learner makes use of the variables to arrive at predictions due to their “black box” nature [24, 25]. In the absence of causal pathways linking exposure to outcomes, the role of machine learning algorithms should not supersede clinical judgement, but rather serve as a tool to guide clinical decisions. Machine learning algorithms may supplement physician decision making by accounting for interactions among variables that clinicians may not be aware of.
Interest in the potential for machine learning in healthcare has recently increased [26]. There have been suggestions that machine learning will drive changes in patient care within a few years, specifically in clinical settings that rely on the accuracy of prognostic models and those based on pattern recognition [9, 27]. For example, deep learning algorithms demonstrated high accuracy in detecting diabetic retinopathy [28], malignant melanoma [29], and in predicting mortality in patients admitted to the ICU [30]. Personalized benefit-risk estimates are one viable application of machine learning algorithms in cardiovascular medicine. Machine learning models may also be used for the enrichment of clinical trials with high-risk individuals that may benefit from a particular investigational therapy, magnifying the expected effect size, increasing power and thus, reducing the sample size.
Limitations
First, the TIMI score was designed to predict short-term mortality and was only available for a subset of patients. However, it is unlikely that the performance estimate is biased enough to compensate for the approximate 20-point difference in c-statistic values. The Killip class variable, required to calculate the GRACE score (which is validated for long term ischemic outcomes) was not available in this dataset. Second, clinical trials are not representative samples of patient populations, possibly limiting the generalizability of the model. Therefore, the model needs to be evaluated in an external data set. Third, the clinical trials evaluated different dosing regimens of rivaroxaban, different control arms, and enrolled ACS patients with and without atrial fibrillation. The differences in enrollment criteria between the included studies introduces population heterogeneity and is associated with different baseline MACE rates. Though in principle, the inclusion of diagnostic variables like atrial fibrillation could account for this difference, in practice, heterogeneous populations often confound prediction tools. More accuracy could possibly be obtained by fitting the model on a subset of patients, though loss of power from decreasing the size of training data could mitigate the effect of having a more homogeneous population. Furthermore, the studies employed different lengths of follow-up, which could also bias our estimator. Further refinement of the model is needed to provide dose and control-specific prediction estimates. Fourth, the super learner ensemble was not compared to a validated bleeding risk score such as the CRUSADE score as the variables required to calculate it were not available in the dataset. Finally, the super learner model was highly calibrated on MACE risk estimates from 0 to 0.3 but predicted few patients (n = 15) with a risk estimate above 0.3, and consequently over-fits the data in this range. Similarly, for the bleeding outcome, a relatively small proportion of patients had a predicted risk above 0.25 (n = 78). However, for both bleeding and efficacy outcomes, patients with a probability above 0.25 are considered at extremely high risk and would warrant maximal medical therapy (for MACE) and caution (for bleeding). Thus, despite loss of calibration in this extremely high-risk range, the model demonstrates excellent calibration for the most clinically relevant range for which the nuances of individual patient characteristics need to be discerned for appropriate clinical decision making.
Conclusion
This analysis is the first to evaluate the performance of machine learning algorithms, built on pooled randomized clinical trial data, for the prediction of MACE and bleeding outcomes among patients with ACS. The super learner produced the highest c-statistic for prediction of MACE compared to traditional risk stratification methods including the TIMI risk score, a surrogate risk score derived from the dataset and a stepwise logistic regression. Importantly, the super learner ensemble method demonstrated remarkable calibration on both efficacy and safety outcomes which greatly exceeded that of traditional logistic regression. This analysis also displayed a contemporary application of machine learning models to assist in clinical decision making based on easily interpreted plots of robust individualized predicted risks of efficacy and safety events in coronary artery disease patients on antithrombotic therapy.
Abbreviations
- ACS:
-
Acute coronary syndrome
- ASA:
-
Aspirin
- CV:
-
Cardiovascular
- DAPT:
-
Dual antiplatelet therapy
- MACE:
-
Major adverse cardiovascular events
- MI:
-
Myocardial infarction
- PCI:
-
Percutaneous coronary intervention
- TIMI:
-
Thrombolysis in myocardial infarction
- VKA:
-
Vitamin K Antagonist
References
Kumar A, Cannon CP (2009) Acute coronary syndromes: diagnosis and management, part I. Mayo Clin Proc 84:917–938
Wachira JK, Stys TP (2013) Cardiovascular disease and bridging the diagnostic gap. SD Med. 66:366–369
Writing Group M, Mozaffarian D, Benjamin EJ, Go AS, Arnett DK, Blaha MJ, Cushman M, Das SR, de Ferranti S, Despres JP, Fullerton HJ, Howard VJ, Huffman MD, Isasi CR, Jimenez MC, Judd SE, Kissela BM, Lichtman JH, Lisabeth LD, Liu S, Mackey RH, Magid DJ, McGuire DK, Mohler ER, Moy CS, Muntner P, Mussolino ME, Nasir K, Neumar RW, Nichol G, Palaniappan L, Pandey DK, Reeves MJ, Rodriguez CJ, Rosamond W, Sorlie PD, Stein J, Towfighi A, Turan TN, Virani SS, Woo D, Yeh RW, Turner MB (2016) American Heart Association Statistics C and Stroke Statistics S. Heart Disease and Stroke Statistics-2016 update: a report from the American Heart Association. Circulation 133:e38–e360
Levine GN, Bates ER, Bittl JA, Brindis RG, Fihn SD, Fleisher LA, Granger CB, Lange RA, Mack MJ, Mauri L, Mehran R, Mukherjee D, Newby LK, O’Gara PT, Sabatine MS, Smith PK, Smith SC Jr, Halperin JL, Levine GN, Al-Khatib SM, Birtcher KK, Bozkurt B, Brindis RG, Cigarroa JE, Curtis LH, Fleisher LA, Gentile F, Gidding S, Hlatky MA, Ikonomidis JS, Joglar JA, Pressler SJ, Wijeysundera DN (2016) 2016 ACC/AHA guideline focused update on duration of dual antiplatelet therapy in patients with coronary artery disease: a report of the American College of Cardiology/American Heart Association Task Force on Clinical Practice Guidelines. J Thorac Cardiovasc Surg 152:1243–1275
Amsterdam EA, Wenger NK, Brindis RG, Casey DE Jr, Ganiats TG, Holmes DR Jr, Jaffe AS, Jneid H, Kelly RF, Kontos MC, Levine GN, Liebson PR, Mukherjee D, Peterson ED, Sabatine MS, Smalling RW, Zieman SJ (2014) 2014 AHA/ACC guideline for the management of patients with non-ST-elevation acute coronary syndromes: a report of the American College of Cardiology/American Heart Association Task Force on Practice Guidelines. J Am Coll Cardiol 64:e139–e228
O’Gara PT, Kushner FG, Ascheim DD, Casey DE Jr, Chung MK, de Lemos JA, Ettinger SM, Fang JC, Fesmire FM, Franklin BA, Granger CB, Krumholz HM, Linderbaum JA, Morrow DA, Newby LK, Ornato JP, Ou N, Radford MJ, Tamis-Holland JE, Tommaso CL, Tracy CM, Woo YJ, Zhao DX, Anderson JL, Jacobs AK, Halperin JL, Albert NM, Brindis RG, Creager MA, DeMets D, Guyton RA, Hochman JS, Kovacs RJ, Kushner FG, Ohman EM, Stevenson WG, Yancy CW, American College of Cardiology Foundation/American Heart Association Task Force on Practice G (2013) ACCF/AHA guideline for the management of ST-elevation myocardial infarction: a report of the American College of Cardiology Foundation/American Heart Association Task Force on Practice Guidelines. Circulation 2013(127):e362–e425
Antman EM, Cohen M, Bernink PJ, McCabe CH, Horacek T, Papuchis G, Mautner B, Corbalan R, Radley D, Braunwald E (2000) The TIMI risk score for unstable angina/non-ST elevation MI: a method for prognostication and therapeutic decision making. JAMA 284:835–842
Eagle KA, Lim MJ, Dabbous OH, Pieper KS, Goldberg RJ, Van de Werf F, Goodman SG, Granger CB, Steg PG, Gore JM, Budaj A, Avezum A, Flather MD, Fox KA, Investigators G (2004) A validated prediction model for all forms of acute coronary syndrome: estimating the risk of 6-month post discharge death in an international registry. JAMA 291:2727–2733
Deo RC, Nallamothu BK (2016) Learning about machine learning: the promise and pitfalls of big data and the electronic health record. Circ Cardiovasc Qual Outcomes 9:618–620
Krittanawong C, Zhang H, Wang Z, Aydar M, Kitai T (2017) Artificial intelligence in precision cardiovascular medicine. J Am Coll Cardiol 69:2657–2664
Mega JL, Braunwald E, Mohanavelu S, Burton P, Poulter R, Misselwitz F, Hricak V, Barnathan ES, Bordes P, Witkowski A, Markov V, Oppenheimer L, Gibson CM, Group AA-Ts (2009) Rivaroxaban versus placebo in patients with acute coronary syndromes (ATLAS ACS-TIMI 46): a randomised, double-blind, phase II trial. Lancet 374:29–38
Mega JL, Braunwald E, Wiviott SD, Bassand JP, Bhatt DL, Bode C, Burton P, Cohen M, Cook-Bruns N, Fox KA, Goto S, Murphy SA, Plotnikov AN, Schneider D, Sun X, Verheugt FW, Gibson CM, Investigators AAT (2012) Rivaroxaban in patients with a recent acute coronary syndrome. N Engl J Med 366:9–19
Gibson CM, Mehran R, Bode C, Halperin J, Verheugt F, Wildgoose P, van Eickels M, Lip GY, Cohen M, Husted S, Peterson E, Fox K (2015) An open-label, randomized, controlled, multicenter study exploring two treatment strategies of rivaroxaban and a dose-adjusted oral vitamin K antagonist treatment strategy in subjects with atrial fibrillation who undergo percutaneous coronary intervention (PIONEER AF-PCI). Am Heart J 169(472–8):e5
Ohman EM, Roe MT, Steg PG, James SK, Povsic TJ, White J, Rockhold F, Plotnikov A, Mundl H, Strony J, Sun X, Husted S, Tendera M, Montalescot G, Bahit MC, Ardissino D, Bueno H, Claeys MJ, Nicolau JC, Cornel JH, Goto S, Kiss RG, Guray U, Park DW, Bode C, Welsh RC, Gibson CM (2017) Clinically significant bleeding with low-dose rivaroxaban versus aspirin, in addition to P2Y12 inhibition, in acute coronary syndromes (GEMINI-ACS-1): a double-blind, multicentre, randomised trial. Lancet 389:1799–1808
Blecker S, Katz SD, Horwitz LI, Kuperman G, Park H, Gold A, Sontag D (2016) Comparison of approaches for heart failure case identification from electronic health record data. JAMA Cardiol 1:1014–1020
Frizzell JD, Liang L, Schulte PJ, Yancy CW, Heidenreich PA, Hernandez AF, Bhatt DL, Fonarow GC, Laskey WK (2017) Prediction of 30-day all-cause readmissions in patients hospitalized for heart failure: comparison of machine learning and other statistical approaches. JAMA Cardiol 2:204–209
Weng SF, Reps J, Kai J, Garibaldi JM, Qureshi N (2017) Can machine-learning improve cardiovascular risk prediction using routine clinical data? PLoS ONE 12:e0174944
Mortazavi BJ, Downing NS, Bucholz EM, Dharmarajan K, Manhapra A, Li SX, Negahban SN, Krumholz HM (2016) Analysis of machine learning techniques for heart failure readmissions. Circ Cardiovasc Qual Outcomes 9:629–640
Mansoor H, Elgendy IY, Segal R, Bavry AA, Bian J (2017) Risk prediction model for in-hospital mortality in women with ST-elevation myocardial infarction: a machine learning approach. Heart Lung 46:405–411
VanHouten JP, Starmer JM, Lorenzi NM, Maron DJ, Lasko TA (2014) Machine learning for risk prediction of acute coronary syndrome. AMIA Annu Symp Proc 2014:1940–1949
Hu D, Huang Z, Chan TM, Dong W, Lu X, Duan H (2016) Utilizing Chinese admission records for MACE prediction of acute coronary syndrome. Int J Environ Res Public Health 13:912
Austin PC, Tu JV, Ho JE, Levy D, Lee DS (2013) Using methods from the data-mining and machine-learning literature for disease classification and prediction: a case study examining classification of heart failure subtypes. J Clin Epidemiol 66:398–407
Ambale-Venkatesh B, Yang X, Wu CO, Liu K, Hundley WG, McClelland R, Gomes AS, Folsom AR, Shea S, Guallar E, Bluemke DA, Lima JAC (2017) Cardiovascular event prediction by machine learning: the multi-ethnic study of atherosclerosis. Circ Res 121:1092–1101
Cabitza F, Rasoini R, Gensini GF (2017) Unintended consequences of machine learning in medicine. JAMA 318:517–518
Ng K, Steinhubl SR, deFilippi C, Dey S, Stewart WF (2016) Early detection of heart failure using electronic health records: practical implications for time before diagnosis, data diversity, data quantity, and data density. Circ Cardiovasc Qual Outcomes 9:649–658
Mayer-Schonberger V, Ingelsson E (2017) Big data and medicine—a big deal? J Intern Med 283:418–429
Deo RC (2015) Machine learning in medicine. Circulation 132:1920–1930
Gulshan V, Peng L, Coram M, Stumpe MC, Wu D, Narayanaswamy A, Venugopalan S, Widner K, Madams T, Cuadros J, Kim R, Raman R, Nelson PC, Mega JL, Webster DR (2016) Development and validation of a deep learning algorithm for detection of diabetic retinopathy in retinal fundus photographs. JAMA 316:2402–2410
Esteva A, Kuprel B, Novoa RA, Ko J, Swetter SM, Blau HM, Thrun S (2017) Dermatologist-level classification of skin cancer with deep neural networks. Nature 542:115–118
Pirracchio R, Petersen ML, Carone M, Rigon MR, Chevret S, van der Laan MJ (2015) Mortality prediction in intensive care units with the Super ICU Learner Algorithm (SICULA): a population-based study. Lancet Respir Med 3:42–52
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
All authors report research grant support from Janssen Pharmaceuticals Inc.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Gibson, W.J., Nafee, T., Travis, R. et al. Machine learning versus traditional risk stratification methods in acute coronary syndrome: a pooled randomized clinical trial analysis. J Thromb Thrombolysis 49, 1–9 (2020). https://doi.org/10.1007/s11239-019-01940-8
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11239-019-01940-8