Abstract
The small planorbid snail Gyraulus cf. laevis (Alder) from Lake Mývatn in Iceland was found to emit large-tailed cercariae with 19 collar spines, and three-spined sticklebacks Gasterosteus aculeatus L. were infected with metacercariae of a species of Petasiger Dietz, 1909. Comparative sequence analysis using ND1 mitochondrial DNA sequences revealed that the rediae and cercariae are conspecific with P. islandicus Kostadinova & Skirnisson, 2007, recently described from an isolated population of the horned grebe Podiceps auritus (L.) at the lake. The redia, cercaria and metacercaria are described and compared with related forms.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
As part of a study of a demographically isolated population of the horned grebe Podiceps auritus (L.) at Lake Mývatn, Iceland, we recently described Petasiger islandicus Kostadinova & Skirnisson, 2007 based on abundant material of adult worms (Kostadinova & Skirnisson, 2007). Subsequent investigation of the digenean fauna at the lake revealed that the small planorbid snail Gyraulus cf. laevis (Alder) emitted cercariae of the ‘Magnacauda’ group; we also found that three-spined sticklebacks Gasterosteus aculeatus L. were infected with metacercariae of a species of Petasiger Dietz, 1909.
Comparative analysis revealed that sequences obtained from the intramolluscan larval stages are almost identical with those of the adult worms upon which P. islandicus was described. This paper reports on the life-cycle of P. islandicus elucidated with the aid of molecular evidence and describes the life-history stages in intermediate hosts of this species.
Materials and methods
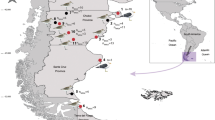
Samples of suspected intermediate hosts of Petasiger islandicus, the freshwater snail Gyraulus cf. laevis and the three-spined stickleback Gasterosteus aculeatus, were collected from Lake Mývatn in Iceland (65°37′N, 16°59′W) and examined for infections during October, 2009 and October, 2011. Rediae and cercariae were fixed in either 100% molecular grade ethanol or hot 4% formalin. The latter were stained with iron-acetocarmine, dehydrated through an alcohol series, cleared in dimethyl phthalate and examined as permanent mounts in Canada balsam. Metacercariae were collected from fishes fixed in hot 4% formalin. Measurements in the text are based on formalin-fixed material, unless otherwise stated, and are in micrometres. Means ± standard deviations are given in parentheses. Two ratios used by Kostadinova & Chipev (1992) in their preliminary key to Palaearctic large-tailed echinostome cercariae were also calculated for both live and fixed cercariae: (i) LT/LB, the length of tail in relation to the length of the body, describing the relative length of the tail; and (ii) LT/WT, the length of the tail in relation to its maximum width, indicating the shape of the tail. Voucher material is deposited in the British Museum (Natural History) Collection at the Natural History Museum, London, UK (BMNH).
DNA was extracted from a single ethanol-fixed adult individual (3 replicates) and 10–20 pooled cercariae or 5–10 rediae in guanidine lysis buffer, precipitated with isopropanol and dissolved in distilled water, following the protocol of Tkach & Pawlowski (1999). Three replicate sequences were obtained from each life-cycle stage of P. islandicus. Additional comparative sequences for P. neocomense Fuhrmann, 1927 and P. phalacrocoracis (Yamaguti, 1939) were obtained from isolates of individual adult worms. Polymerase chain reaction (PCR) amplifications of partial fragments of the mitochondrial gene nicotinamide adenine dinucleotide dehydrogenase subunit 1 (ND1) was carried out using PCR beads (Amersham Pharmacia Biotech) in a total volume of 25 μl (10 pmol of each primer) with 50 ng gDNA, using the following thermocycling profile: 35 cycles with 30 s DNA denaturation at 94°C, 20 s primer annealing at 48°C, and 45 s at 72°C for primer extension. The following PCR primers were used: forward NDJ11: 5′-AGA TTC GTA AGG GGC CTA ATA-3′ and reverse NDJ2a: 5′-CTT CAG CCT CAG CAT AAT-3′ (Kostadinova et al., 2003). NDJ11 is equivalent to JB11 in Morgan & Blair (1998). Partial (domains D1–D3; c.1400 bps) 28S rDNA sequences were amplified using primers U178F (5′-GCA CCC GCT GAA YTT AAG-3′) and L1642R (5′-CCA GCG CCA TCC ATT TTC A-3′) (Lockyer et al., 2003). The same PCR profile but with an annealing temperature of 56°C was applied for 28S rDNA amplification. PCR products were purified using a QIAquick PCR purification kit (Qiagen). PCR fragments were sequenced directly for both strands using the PCR primers [plus LSU1200R (5′-CAT AGT TCA CCA TCT TTC GG-3′; Lockyer et al., 2003) for 28S] with ABI BigDye chemistry (ABI Perkin-Elmer), alcohol-precipitated, and run on an ABI Prism 3130xl automated sequencer. Sequences were assembled and edited using Bioedit v7.0.5. (©1997–2005, Hall, 1999), and submitted to GenBank under accession numbers JQ425587–JQ425593. Sequences were aligned with Clustal W implemented in MEGA 5 (Tamura et al., 2011) with reference to the amino acid translation, using the echinoderm and flatworm mitochondrial code (Telford et al., 2000). Genetic distances (uncorrected p-distance) were calculated and visualised in a neighbour-joining (NJ) consensus tree with MEGA 5 using the Kimura 2-parameter model of base substitution.
Molecular identification
The newly generated partial ND1 sequences for the isolates of Petasiger islandicus ranged between 467 and 502 bp and those for P. neocomense ranged between 475 and 513 bp. To aid future identification and studies at the supraspecific level, we also obtained sequences of the 28S rRNA gene for P. islandicus (1,411–1,496 bp) and P. phalacrocoracis (1,010 bp). Unfortunately, we failed to obtain ND1 sequences for the latter species and 28S sequences for P. neocomense. Nevertheless, at least one genotype per gene region of each Petasiger species was available for comparative sequence analysis. ND1 sequences for four isolates of P. islandicus and two isolates of P. neocomense were aligned together with the sequence for the ND1 of Echinostoma revolutum (Frölich, 1802) (AY168933) of Kostadinova et al. (2003) as an outgroup taxon. The ND1 alignment from the trimmed sequences (to match the shortest sequence) included two unique haplotypes for P. islandicus and incorporated a total of 471 characters; it contained no insertion or deletions. The consensus tree from neighbour-joining (NJ) analysis in Fig. 1 visualises the relationships of the two groups of unique ND1 haplotypes studied. Divergence in the ND1 sequence between isolates of P. islandicus ranged from 0% to 0.2% and that between P. islandicus and P. neocomense ranged between 28.2 and 28.5% (133–134 nucleotide sites) (Fig. 1). Divergence in the 28S sequence between P. islandicus and P. phalacrocoracis was 3.9% (39 nucleotide sites). These comparisons clearly indicate that the cercarial and redial isolates collected from Gyraulus cf. laevis obtained during 2009 and 2011 are conspecific with P. islandicus and quite divergent from the two Palaearctic species of Petasiger currently available for comparison.
Consensus tree for Petasiger material from a neighbour-joining analysis of sequences of the mitochondrial gene nicotinamide adenine dinucleotide dehydrogenase subunit 1 (ND1). The evolutionary distances were computed using the Kimura 2-parameter method with MEGA5 using Echinostoma revolutum as an outgroup. Isolate code, year of collection and the stage of life-cycle are indicated for Petasiger islandicus. Abbreviations: R, rediae; C, cercariae; A, adult
Petasiger islandicus Kostadinova & Skirnisson, 2007
First intermediate host: Gyraulus cf. laevis (Alder) (Prosobranchia: Planorbidae).
Second intermediate host: Gasterosteus aculeatus L. (Teleostei: Gasterosteidae).
Locality: Lake Mývatn (65°37′N, 16°59′W), Iceland.
Prevalence: In Gyraulus cf. laevis: 3.6% (October, 2009; n = 195); 7.0% (October, 2011; n = 100). Exclusively, snails with a diameter exceeding 3.8 mm (up to 4.2 mm) shed cercariae. In Gasterosteus aculeatus: 100% (October, 2011; n = 10).
Voucher material: BMNH 2012.1.9.1-4 (cercariae); BMNH 2012.1.9.5 (metacercariae).
Representative DNA sequences: Partial (D1-D3) 28S rRNA gene: GenBank accession no. JQ425592. Partial ND1 mitochondrial gene: GenBank accession nos JQ425587–JQ425590.
Comparative DNA sequences: Petasiger phalacrocoracis (adult ex Phalacrocorax carbo; Czech Republic): partial (D1-D3) 28S rRNA gene: GenBank accession no. JQ425593. Petasiger neocomense (adult ex Podiceps cristatus; Czech Republic): partial ND1 mitochondrial gene: GenBank accession no. JQ425591.
Morphology of the life-cycle stages
Redia (Fig. 2)
[Based on 20 specimens from snail hepatopancreas and gonad, fixed in hot 4% formalin.] Body slightly brownish, elongate, 650–1,838 × 114–279 (1,198 ± 355 × 197 ± 48). Collar 76–288 (192 ± 61) wide, divided into 4 lobes: 2 lateral, one dorsal and one ventral. Birthpore prominent, immediately posterior to collar. Pharynx 40–85 × 28–70 (55 ± 12 × 44 ± 9). Gut saccate, reaches close to posterior locomotory processes located in posterior fifth of body.
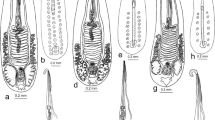
[Measurements from 15 specimens fixed in hot 4% formalin.] Body whitish, elongate-oval, small, 155–223 (191 ± 25) × 70–107 (85 ± 11). Tegument thick; entire body surface covered with minute scale-like spines. Collar 33–48 (43 ± 4) × 44–69 (55 ± 8), with 19 spines (4 angle spines on each side and 11 dorsal spines in single row). Tail leaf-like (LT/WT ratio 3.8–6.7 and 4.5–5.0 for fixed and live cercariae, respectively), massive, brownish, with festooned margins, 740–972 (847 ± 80) long, much longer than body (LT/LB ratio 3.5–5.8 and 3.8–6.7 for fixed and live cercariae, respectively), with maximum width 145–224 (185 ± 24); accessory excretory vesicle in narrow anterior region of tail. Tail musculature consists of thickly set, fine circular muscle fibres; axial longitudinal muscles extend medially throughout length of tail and numerous fine oblique muscle fibres originate from lateral margins and terminate medially along tail. String of clearly separated large, dark brown cells present along median axis of tail in immature cercariae (Fig. 4).
Oral sucker ventro-subterminal, 30–42 (36 ± 3) × 29–43 (35 ± 4). Ventral sucker muscular, postequatorial, 34–45 (39 ± 4) × 34–56 (41 ± 6). Prepharynx distinct, 11–23 (17 ± 4); pharynx elongate-oval, 15–23 (18 ± 2) × 10–14 (11 ± 1); oesophagus long; intestinal bifurcation just anterior to ventral sucker; caeca reach anterior margin of excretory vesicle. Cystogenous glands numerous, 4–9 (7 ± 1) × 3–6 (4 ± 1), with rhabditiform contents, occupy most of body posterior to pharynx. Penetration glands few, small, indistinct, present on both sides of oesophagus. Genital primordia in 2 cellular masses just anterior and posterior to ventral sucker, connected by chain of cells.
Main excretory ducts filled with large refractive granules mostly formed by fusion of 2–4 smaller ones, situated between posterior margin of pharynx and posterior half of ventral sucker, connected to anterior corners of excretory vesicle. Excretory vesicle bipartite. Flame-cell formula not determined.
The cercariae were positively phototactic, described a figure of 8 while swimming and took up a vertical resting position.
Metacercaria (Fig. 7)
[Based on 15 specimens fixed in hot 4% formalin.] Encysted on inner surface of pharynx and anterior part of oesophagus of host. Metacercarial cyst invariably ovoid, 100–109 (103 ± 3) × 58–74 (67 ± 6), covered with thin [2–4 (3 ± 1)] layer of parasite origin enveloped by thick [3–10 (8 ± 4)] layer of connective tissue of host origin. Body covered with blunt spines. Collar bears 19 spines. Main excretory ducts filled with large, composite excretory granules.
Discussion
Although records of Petasiger spp. have been relatively abundant, especially in grebes from the Palaearctic (Faltýnková et al., 2008), data on their life-cycles and life-history stages in the intermediate hosts are still scarce. Thus far, life-cycles of two species, P. grandivesicularis Ishii, 1935 and P. neocomense Fuhrmann, 1927, have been completed experimentally within the Palaearctic (Karmanova, 1971; Kostadinova & Chipev, 1992) and two further unidentified cercariae of the ‘Magnacauda’ group, possessing 19 collar spines, have been described from Planorbis carinatus Müller in pools near Birmingham, UK, i.e. Cercaria rashidi Nasir, 1962 and C. titfordensis Nasir, 1962 (see Nasir, 1962). Comparative analysis of molecular data is a powerful tool that can help elucidate trematode life-cycles by linking life-cycle stages; this is especially important for parasites of wildlife, since laboratory experiments with the use of natural definitive hosts, such as grebes, is deemed impossible.
Comparative ND1 sequence analysis carried out by us confidently confirmed the link between the redial and cercarial isolates ex Gyraulus cf. laevis and the adult stages of P. islandicus parasitising Podiceps auritus in Lake Mývatn. Sequence divergence in the ND1 between these isolates was notably low, much lower than interspecific divergence (0–0.2 vs 28.2–28.5%), thus strongly supporting the conspecificity of the isolates from Lake Mývatn. The fact that isolates of the redial and cercarial stages collected during different years represented P. islandicus indicates that this is the only species of Petasiger completing its life-cycle in the lake. Although we could not obtain ND1 sequences from the metacercariae ex Gasterosteus aculeatus, their morphology and the specific site in the host indicate conspecificity with P. islandicus. We suggest, therefore, that the life-cycle of P. islandicus is completed using Gyraulus cf. laevis and Gasterosteus aculeatus as intermediate hosts. The latter, three-spined stickleback, is the most abundant fish species, with a density reaching up to 100–200 fish m−2, in the north basin of Lake Mývatn (Einarsson et al., 2004) and represents the main food-source of Podiceps auritus in its north-European range (Fjeldså, 1973), thus supporting our suggestion.
The cercaria of P. islandicus keys down to Cercaria titfordensis using the simple key to the known large-tailed echinostome cercariae from the Palaearctic presented by Kostadinova & Chipev (1992) but differs from the latter in both the relative length of the tail (LT/LB 3.9–7.7 vs 3.5–3.8) and its shape, i.e. more elongate in P. islandicus (LT/WT 4.5–5.0 vs 2.7). Kostadinova & Chipev (1992) suggested that C. titfordensis and the cercaria of P. neocomense described by Karmanova (1971) are conspecific. Although currently no additional data exist to support this morphologically, we have observed substantial ND1 sequence divergence between P. islandicus and P. neocomense, thus supporting the distinct status of the two species.
References
Einarsson, Á., Stefánsdóttir, G., Jóhannesson, H., Ólafsson, J. S., Gíslason, G. M., Wakana, I., Gudbergsson, G., & Gardarsson, A. (2004). The ecology of Lake Mývatn and the River Laxá: variation in space and time. Aquatic Ecology, 38, 317–348.
Faltýnková, A., Gibson, D. I., & Kostadinova, A. (2008). A revision of Petasiger Dietz, 1909 (Digenea: Echinostomatidae) and a key to its species. Systematic Parasitology, 71, 1–40.
Fjeldså, J. (1973). Territory and the regulation of population density and recruitment in the horned grebe Podiceps auritus arcticus Boje, 1822. Videnskabelige Meddelelser frå Dansk Naturhistorisk Forening, 136, 117–189.
Hall, T. A. (1999). Bioedit: a user-friendly biological sequence editor and analysis program for Windows 95/98/NT. Nucleic Acids Symposium, Ser. 41, 95–98.
Karmanova, E. M. (1971). [The life cycle of Petasiger neocomense (Trematoda, Echinostomatidae).] Trudy Gel’minthologicheskoi Laboratorii, 21, 31–35. (In Russian).
Kostadinova, A., & Chipev, N. (1992). Experimental data on the life-cycle of Petasiger grandivesicularis Ishii, 1935 (Trematoda: Echinostomatidae). Systematic Parasitology, 23, 55–65.
Kostadinova, A., & Skirnisson, K. (2007). Petasiger islandicus n. sp. (Digenea: Echinostomatidae) in the horned grebe Podiceps auritus (L.) (Aves: Podicipedidae) from Iceland. Systematic Parasitology, 68, 217–223.
Kostadinova, A., Herniou, E. A., Barrett, J., & Littlewood, D. T. J. (2003). Phylogenetic relationships of Echinostoma Rudolphi, 1809 (Digenea: Echinostomatidae) and related genera re-assessed via DNA and morphological analyses. Systematic Parasitology, 54, 159–176.
Lockyer, A. E., Olson, P. D., & Littlewood, D. T. J. (2003). Utility of complete large and small subunit rRNA genes in resolving the phylogeny of the Neodermata (Platyhelminthes): implications and a review of the cercomer theory. Biological Journal of the Linnean Society (London), 78, 155–171.
Morgan, J. A. T., & Blair, D. (1998). Relative merits of nuclear ribosomal internal transcribed spacers and mitochondrial CO1 and ND1 genes for distinguishing among Echinostoma species (Trematoda). Parasitology, 116, 289–297.
Nasir, P. (1962). Two new species of giant-tailed echinostome cercariae from Planorbis carinatus (Mtill) [sic]. Transactions of the American Microscopical Society, 81, 132–137.
Tamura, K., Peterson, D., Peterson, N., Stecher, G., Nei, M., & Kumar, S. (2011). MEGA5: Molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Molecular Biology and Evolution, 28, 2731–2739.
Telford, M. J., Herniou, E. A., Russell, R. B., & Littlewood, D. T. J. (2000). Changes in mitochondrial genetic codes as phylogenetic characters: two examples from the flatworms. Proceedings of the National Academy of Sciences, 97, 11359–11364.
Tkach, V., & Pawlowski, J. (1999). Method for DNA isolation from ethanol-fixed specimens using guanidine thiocyanate. Acta Parasitologica, 44, 147–148.
Acknowledgements
We thank Drs David Gibson (Natural History Museum, London, UK) and Tomáš Scholz (Institute of Parasitology, ASCR, České Budějovice, Czech Republic) for their comments. This research was supported by the Czech Science Foundation (projects P505/10/1562 to AK and SG and 206/09/H026 to SG), the Grant Agency of the University of South Bohemia (GAJU) (project 04-135/2010/P to SG) and the Research Fund of the University of Iceland (to KS).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Georgieva, S., Kostadinova, A. & Skirnisson, K. The life-cycle of Petasiger islandicus Kostadinova & Skirnisson, 2007 (Digenea: Echinostomatidae) elucidated with the aid of molecular data. Syst Parasitol 82, 177–183 (2012). https://doi.org/10.1007/s11230-012-9354-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11230-012-9354-y