Abstract
The waste generated from industrial processes and operations including domestic wastes when treated partially and disposed in soil–water environment enter to lakes, streams, rivers, oceans and other water bodies. The pollutants get dissolved or lie suspended in water or get deposited on soil sediment beds. This results on aquatic and terrestrial pollution which ultimately impact ecosystems causing toxicity to biota and human beings. Industries such as petrochemical, pharmaceutical, insecticides and fertilizers generates the hazardous waste comprising of inorganic and organic compounds. Organic compounds mainly composed polycyclic aromatic hydrocarbons (PAHs), are one of the toxic environmental pollutant. This paper highlights the physicochemical properties, bioremediation treatment and its mechanism for the waste containing PAH. The process of biological remediation depends upon the metabolic action of microbe toward the contaminant which can be achieved by optimum water and nutrient supply and some other limiting factors. The enzymatic degradation gives the molecular approaches for bioremediation. The study also highlighted the molecular approaches which are helpful in revealing functional, structural and communal information about microbial diversity for exploring the routes of degradation pathway of bioremediation process and future scope to bioremediation of PAHs.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Industrial development and excessive use of synthetic chemicals has led to increase in the concentration of persistent organic compounds in the environment, which causes adverse effect in the environment. Aromatic hydrocarbons are the most common pollutant found in the soil and ground water. Polycyclic aromatic hydrocarbons (PAHs) are one class of toxic environmental pollutants that have accumulated in the environment due to several natural and anthropogenic activities. The largest release PAHs is due to incomplete combustion of organic compounds during the course of industrial processes and other human activities. PAHs are a type of organic compounds that consist of two or more fused benzene rings and/or pentacyclic molecules that are arranged in various structural configurations. They are mostly used as intermediaries in pharmaceuticals, agricultural products, photographic products, thermosetting plastics, lubricating materials, and other chemical industries. The main concern with PAHs compounds is with respect to potential health risk. Some members of these compounds are regulated as carcinogens and mutagens reported by Mastral and Callen (2000). Removal of PAHs from the environment by microbial biodegradation is one of the promising tools and has been studied extensively (Yuan et al. 2000; Zhang et al. 2004a, b). Biodegradation is a natural process that helps to remove PAHs compound from the environment by microorganisms. It is one of the cost effective methods amongst remedial approaches. Bioremediation has been shown to be effective in remediating soils contaminated with low molecular weight PAHs (Mueller et al. 1991; Banerjee et al. 1995; Kastner and Mahro 1996). However, PAHs having high molecular weights are generally recalcitrant to microbial attack (Park et al. 1990; Cerniglia 1992; Erickson et al. 1993). Microbes thriving in contaminated environment are able to use the contaminant as source of energy because of their genetic adaptability which leads to bioremediation. The lack of microbial activity towards high molecular weight PAHs may be attributed to site specific environmental factors, such as bioavailability of the contaminant, nutrients, redox potential, etc., the limiting factor may be the scarcity of micro-organisms capable of degrading the more highly condensed compounds. Process of bioremediation undergoes to some complex stages and accessibility towards the mechanism and metabolic pathway of microbes is quite difficult but with rapid advancement in biotechnological approaches and molecular techniques it become possible i.e. some of the techniques are polymerase chain reaction (PCR), fingerprinting technique based on PCR Presently, 16S rRNA gene liabrary, denaturing gradient gel electrophoresis (DGGE), single strand conformation polymorphism (SSCP), terminal restriction fragment length polymorphism (T-RFLP) rRNA intergenic spacer analysis, DNA hybridization such as fluorescence in situ hybridization (FISH) and DNA microarray, gene reporters and biosensors were also frequently used. However, as for PAHs biodegradation investigation, the techniques extensively employed were PCR, fingerprinting technique (mainly DGGE), DNA hybridization technique and gene reporters. New sequencing technologies i.e. metagenomics, metaproteomics and metatranscriptomics studies were also highlighted. Metaproteomics approaches utilizing two-dimensional electrophoresis (2-DE), mass spectroscopy (MS) have aided in global analysis of catabolic enzymes involved in microbial biodegradation pathways (Kim et al. 2006a, b; Wilmes et al. 2008). One of the upcoming “omics” technologies known as metabolomics refers to the colossal analyses of primary and secondary proteinaceous metabolites produced by microbial cells under defined physiological conditions (Mapelli et al. 2009). The nucleic acid extraction from soil is the first crucial step in the application of most of the molecular techniques, which have mainly been dominated by diverse variations of PCR. These methods can provide new insights into bacterial and fungal community compositions, their associations and their responses to each other with respect to environmental conditions. Technological advancement, sensitivity and specificity PCR-based finger printing techniques have proved enormously useful in assessing the changes in microbial community structure. Fluorescent in situ hybridization (FISH) can be used to evaluate the distribution and function of bacterial population in situ. DNA microarray techniques have also been developed and used for the evaluation of ecological role and phylogenetic affiliations of bacterial population in the soil (Dubey et al. 2006). 16S ribosomal RNA (rRNA) sequencing is a common amplicon sequencing method is helpful in the phylogenetic and taxonomic comparison and identification of bacteria present within a contaminated environment as discussed by Legge (2012). Scope of this review paper is limited to polycyclic aromatic hydrocarbon and role of microorganisms involving characterization of specific enzymatic activity and gene expression during PAHs degradation through molecular approaches.
1.1 Sources and physico-chemical properties
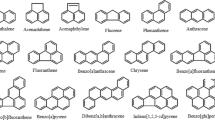
Sources of polycyclic aromatic hydrocarbons (PAHs) can be natural: forest and grass fires, oil seeps, volcanoes, chlorophyllous plants, fungi, and bacteria and anthropogenic: petroleum, electric power generation, refuse incineration, home heating, production of coke, carbon black, coal tar, asphalt and internal combustion engines. In areas remote from urban or industrial activities, the levels of PAHs found in unprocessed foods reflect the background contamination, which originates from long distance airborne transportation of contaminated particles and natural emissions from volcanoes and forest fires. In the neighbourhood of industrial areas or along highways, the contamination of vegetation can be ten-fold higher than in rural areas (Larsson and Sahlberg 1982). Processing of food (drying and smoking) and cooking of foods at high temperatures (grilling, roasting, frying) are major sources generating PAHs (Guillen et al. 1997). The general characteristics of PAHs are high melting and boiling points, low vapor pressure, and very low aqueous solubility (toxipedia). Generally solubility decreases and hydrophobicity increases with an increase in number of fused rings (Wilson and Jones 1993). PAHs are highly lipophilic and therefore very soluble in organic solvents, they also have high octanol water coefficient (Kow), i.e. Naphthalene, Anthracene, Pyrene, Chrysene, Benzo (α) pyrene. Structural representation of PAHs is shown in Fig. 1. PAHs also manifest various functions such as light sensitivity, heat resistance, conductivity, emittability, corrosion resistance, and physiological action. Most PAHs are also fluorescent, emitting characteristic wavelengths of light when they are excited. PAHs have been identified as being of greatest concern with regard to potential exposure and adverse health effects on humans (Table 1).
1.2 Persistence of PAHs compounds in the environment and its toxicity
There are a variety of mechanisms by which PAHs are degraded in the environment, including chemo-oxidation photooxidation and microbial degradation which is considered as the primary route of degradation of PAHs in soils mentioned by Juhasz and Naidu (2000). Emission into the air of complex mixture of different PAHs including particulate matter as in smoke, PAHs in vapors phase can be absorbed onto airborne particles or to diffusion. Due to low vapor pressure naphthalene and fluorine can exist as vapor phase in the atmosphere, near surface water. Because of their high Kow values they tend to become adsorbed to the organic material of sediments and are taken up by aqueous organisms. The potential for photochemical oxidation of structural isomers benzo[α]pyrene in the atmosphere was studied and a significant difference in photochemical oxidation rate was observed. Benzo[α]pyrene was not photochemically oxidise whereas benzo[α]pyrene was photochemically oxidised (Takata and Sakata 2002). PAHs can be bio-concentrated or bio-accumulated by certain aqueous invertebrate, but such species are less in number which are able to effectively biotransform polycyclic aromatic hydrocarbons. Phytoplankton play an important role in the fate and transport of persistent organic pollutants like PAHs and their consumption as an important initial phase in bioaccumulation was studied by Fan and Reinfelder (2003). Many PAHs are carcinogenic and therefore, of significant concern as environmental contaminants. Numerous studies have indicated that one-, two- and three-ring compounds are acutely toxic as documented by Sims and Overcash (1983), while higher molecular weight PAHs are considered to be genotoxic (Phillips 1983; Lijinsky 1991; Mersch-Sundermann et al. 1992; Nylund et al. 1992). BaP is one of the most potent carcinogenic PAHs, and as such, is the most studied compound of the PAH class (Collins et al. 1991). Metabolism and activation of BaP in mammalian systems was studied by Cooper et al. (1983). Reports states that these intermediates undergo through one of at least four different mechanisms of oxidation and/or hydrolysis before the intermediates combine with and/or attack DNA to form covalent adducts with DNA. DNA adducts can lead to mutations of the DNA, resulting in tumours (Harvey 1996). In mammals when ingested, PAHs are rapidly absorbed into the gastrointestinal tract due to their high lipid solubility (Cerniglia 1984). Vanrooij et al. (1993) were estimated 75 % of the total absorbed amount of PAHs (specifically pyrene) entered the body through the skin, highlighting this as a major exposure route of PAHs. The rapid absorption of PAHs by humans results in a high potential for bio-magnification in the food chain. In general, higher the number of benzene rings, the greater the toxicity of the PAHs (Cerniglia 1992).
1.3 Chemotaxis toward aromatic compounds
Biodegradation research has been focused on catabolic pathway and identification, characterization of catabolic enzyme gene that encode them, identification of pathway intermediates and determination of appropriate condition for pathway induction. The applicability in biodegradation process, there is need to be consider other aspect such as substrate specificity, which determines that whether the organic compounds are accessible to bacteria or fungi for biodegradation process (bioavailability); it can be termed as chemotaxis (bacterium or fungus sensory) which can be conceptualize as how the microorganism find the chemical compound in the environment and how this chemical compound transport into a cell for degradation. It allows the detection of compound and uptake system for these chemical compounds. This is a prerequisite factor for biodegradation. Several factors which affect the bioavailability include hydrophobicity, aqueous solubility, polarity of the compound and characteristic of the medium (soil or water) where the contaminant is present. Recent studies have clearly demonstrated that many bacteria have sensory system that allows them to detect and respond behaviorally to chemical pollutant (Table 2). Fungi also play important role in assessing bioavailability of organic compounds. The strategy of filamentous fungi is to enlarge their external surface and to develop mycelia of high fractal dimension that optimally exploit the three-dimensional space containing the substrate was studied by Nakagaki et al. (2004). Furono et al. (2009) observed the movement of polycyclic aromatic hydrocarbon-degrading Pseudomonas putida PpG7 (NAH7) along mycelium of Pythium ultimum. Some dispersal was also observed in the absence of a chemoattractant. The non-chemotactic derivative strain P. putida G7.C1 (pHG100) was used. The bacterial movement became fourfold more effective when wild type chemotactic was used and salicylate was present as a chemoattractant. No dispersal of bacteria was found in the absence of the fungus. The role of mycelia for the translocation of chemicals was observed. Study suggested that fungi improve the accessibility of contaminants in water.
2 Biological treatment of PAH compounds
Biological treatment mainly relies on microbial remediation strategies which are developed to promote the microbial metabolism of contaminants, by providing appropriate water, air and nutrient supply. This is accomplished by the biostimulation (the addition of a bulking agent such as wood chips and/or nutrients such as N/P/K) and bioaugmentation (often an inoculum of microorganisms with known pollutant transformation abilities) of the contaminated environment. The pathways of microbial degradation tend to show broad substrate specificity and occur both aerobically and anaerobically (Harayama 1997). Bioremediation of PAHs contaminated soils, sediments, and water can be accomplished in a variety of ways, e.g. in situ treatment or ex situ methods such as bio-piling and composting. Several bacterial strains, have been isolated were identified on the basis of phospholipid analysis, FAME analysis, G1C content and 16S rRNA gene sequence analysis as species of Pseudomonas, Microbacterium, Paracoccus, Arthrobacter and Mycobacterium from Greek soils contaminated with PAH-containing waste from a wood processing industry, a steel industry and an oil refinery industry (Zhang et al. 2004a, b; Kallimanis et al. 2007). Generally unfavorable site conditions may cause variability in the applicability of bioremediation; therefore a thorough understanding of site conditions will allow optimization of bioremediation and subsequently more effective results. The basis of microbial metabolism of PAHs includes the oxidation of the aromatic ring, followed by the systematic breakdown of the compound to PAHs metabolite and/or carbon dioxide. PAHs degrading microorganisms are ubiquitously distributed in the natural environment, such as in soils (bacteria and non-ligninolytic fungi) and woody materials (ligninolytic fungi) as mentioned by Lee and Lee (2001). A study concluded by Sinha et al. (2009; Khan et al. 2001) during bacterial degradation of pyrene, in M. vanbaalenii PYR-1 cultures, First pathway indicates that pyrene hydroxylation takes place at the 1,2 positions, leading to the formation of 4-hydroxy-perinaphthenone which is the ultimate product so far found. Several key metabolites obtained were pyrene-4,5-dione, cis-4,5-pyrene-dihydrodiol, phenanthrene-4,5-dicarboxylic acid, and 4-phenanthroic acid are obtained during biotransformation of pyrene by gram-negative bacteria and accumulates Pyrene-4,5-dione as a final product and is further utilized as growth substrate (Liang et al. 2006). Fungal variation occurs in wide range of habitats in freshwater and the sea, soil, litter, in dung and in living organisms. It play very important role in mineralization and transformation of complex organic compounds into simpler compounds. According to (Pozdnyakova 2012) ligninolytic fungi, such as Phanerochaete chrysosporium, Bjerkandera adusta, and Pleurotus ostreatus, have the capacity of PAH degradation and the enzymes involved in the degradation of PAHs are ligninolytic and include lignin peroxidase, versatile peroxidase, Mn-peroxidase, and laccase. There are three main pathways followed by bacteria and fungi (Bamforth et al. 2005) shown in Fig. 2.
Algal species including Chlamydomonas sp., Dunaliella sp., and Scenedesmus obliquus, Selenastrum capricornutum have been reported as being able to biotransform and bioaccumulate naphthalene, phenanthrene and pyrene and biotransform benzo[a]pyrene (Semple et al. 1999). Microbial consortia isolated from aged oil-contaminated soil were used to degrade 16 polycyclic aromatic hydrocarbons in soil and slurry phases. The three microbial consortia (bacteria, fungi and bacteria–fungi complex) could degrade polycyclic aromatic hydrocarbons (PAHs), and the highest PAH removals were found in soil and slurry inoculated with fungi. PAHs biodegradation in slurry was lower than in soil for bacteria and bacteria–fungi complex inoculation treatments. Degradation of three- to five-ring PAHs treated by consortia was observed in soil and slurry, and the highest degradation of individual PAHs (anthracene, fluoranthene, and benz(a)anthracene) appeared in soil and slurry. Inoculation of microbial consortia (bacteria, fungi and bacteria–fungi complex) isolated from in situ contaminated soil to degrade PAHs could be considered as a successful method (Li et al. 2008a, b).
2.1 Role of enzyme for biological remediation
Enzymology may be regarded as being at the interface between, on the one hand, biological discovery and protein engineering (Harford-cross et al. 2000) and environmental biotechnology. Enzymatic treatment of PAHs-contamination is an alternative to conventional bioremediation (Gianfreda and Bollag 2002; Gianfreda and Rao 2004; Ruggaber and Talley 2006). Microorganisms can be genetically ‘engineered’ to express specific xenobiotic metabolizing enzymes that would degrade even the most recalcitrant pollutants. The advantages of enzymatic remediation over microbial treatment include high reaction activity towards recalcitrant pollutants, lower sensitivity to the concentrations of contaminants, coverage of a wide range of physicochemical gradients in the environmental matrix and easy control of field application. With a necessity of improvement in biological remediation techniques, enzyme technology has been receiving increased attention. Fungal enzymes, which include lignin and manganese peroxidase and laccase, are responsible for the oxidative biodegradation of PAHs into CO2 and H2O. In the past several years, several oxido-reductases such as laccases and cytochrome P450 monooxygenases (CYPs) have been exploited for the enzymatic degradation of PAHs. Laccases belong to a group of multicopper enzymes that can catalyze the oxidation of a wide variety of phenolic compounds including PAHs (Majcherczyk et al. 1998; Alcade et al. 2002). Like other ligninolytic enzymes, laccase is difficult to express in non-fungal systems and knowledge of structure–function relations underlying the key functional properties of laccase is limited. Hence, directed evolution holds exciting potential for improving the performance of the enzyme. In a study undertaken by Bulter et al. (2003) the laccase gene from Myceliophthora thermophia (MtL), which was previously expressed only in Asperigillus oryzae, was transformed into Saccharomyces cerevisiae and subjected to directed evolution. PAHs can be oxidized by CYP enzymes to form catechols, which are then degraded by other enzymes, including catechol dioxygenases to harmless products and incorporated into the tricarboxylic acid cycle of microorganisms. Wild-type CYP101 (P450cam) from Pseudomonas putida has been shown to have an inherently low activity (<0.01 min−1) towards the PAH substrates phenanthrene, fluoranthene, pyrene and benzo[a]pyrene (Harford-cross et al. 2000). Therefore, CYP enzymes have been subjected to a number of rational design studies to enhance their catalytic performance. Based on the crystal structures of CYP101, selective mutations were performed on the active site residues F87 and Y96 of the enzyme. For all PAH substrates studied, the absolute oxidation rates (approximately 1 min−1) of the mutants, Y96A, Y96F, F87A/Y96A and F87L/Y96F, were increased by two to three orders of magnitude relative to the wild-type enzyme. Carmichael and Wong (2001), introduced two mutations into CYP102, R47L and Y51F, and found that the oxidation activity of the enzyme for phenanthrene and fluoranthene was increased by 40- and ten-fold, respectively. The double mutant was then used as a basis for further engineering of the active site. When the A264G mutation was introduced to the base mutant, NADPH turnover, PAH oxidation and coupling efficiency of the enzyme was greatly improved. The most active mutants showed more than a 200-fold increase in PAH oxidation activity compared the wild type enzyme. Another mutation, F87A, resulted in a larger space in the substrate binding pocket of the enzyme, leading to better accommodation of larger fluoranthene and pyrene molecules in the vicinity of the haem site, and hence a more efficient PAH oxidation.
2.2 Aerobic degradation of PAHS
The basic mechanism behind the aerobic microorganisms to overcome the problem of degradation is linked with oxygenases that primarily reduce elemental oxygen to activate it, and allow it to insert into substrate. Aerobic condition favors the electron acceptor has lower standard reduction potential values and convert into less standard Gibbs free energy change when coupled with any given substrate.
The initial catabolic step in aerobic oxidation of a PAH molecule occurs via formation of dihydrodiol by a multicomponent dioxygenase enzyme system which is further metabolized by either an ortho-or a meta pathway, leading to intermediates such as protocatechuates and catechol. These intermediates compounds are processed through either ortho-meta cleavage leading to central intermediate compounds such as protocatechuates, catechols, gentisates, homoprotoocatechuates, homogentistases, hydroquinones and hydroxyquinols which are further transformed to tricarboxylic acid cycle intermediates (Harayama and Timmis 1992) finally channeled into intermediates of kreb’s cycle. The principal mechanism for the aerobic bacterial metabolism of PAHs is the initial oxidation of the benzene ring by the action of dioxygenase enzymes to form cis-dihydrodiols. These dihydrodiols are dehydrogenated to form dihydroxylated intermediates, which can further metabolised via catechols to carbon dioxide and water. The ability of the microorganisms to degrade PAHs is reported the involvement of Beijerinkia sp. in oxidation of benzo(a)pyrene and benzo(a)Anthracene to dihydrodiols (Gibson and Subranian 1984; Jain et al. 2005).
2.2.1 Advantages and disadvantages
The aerobic pathway releases a substantial amount of energy. A portion is used by the microorganisms for synthesis and growth of new microorganisms. Aerobic treatment yields better effluent quality. Aerobic co-metabolism has unique benefits derived from the advantages of aerobic respiration. These advantages and benefits can reduce costs and risks associated with soil and ground water remediation. There is a drawback of aerobic degradation process that it requires high power Supply and energy consumption.
2.3 Anaerobic degradation of PAHS
Anaerobic degradation occurs in the absence of oxygen, under anaerobic conditions, aromatic compounds are able to serve as electron donating substrates of primary metabolism, several compound acts as terminal electron acceptor (TEA) and oxidation of aromatic compounds support the growth of microorganisms. A basic pathway for biotransformation of PAHs under anaerobic condition is shown in Fig. 3. (Widdle and Rabbus 2001). Phthalate compound degradation is mainly carried out by anaerobic methanogens (Methanospirillum hungatei, Methanosaeta concilii, Syntrophobacter fumaroxidens), producing acetate and methane as end products by decarboxylation initially, then reduction followed by ring cleave and ultimately pave to the β-oxidation pathway (Qiu et al. 2004; Zhang and Bennet 2005). The biotransformation of pyrene and benzo[a]pyrene (BaP) is well studied in different bacterial species such as Mycobacterium vanbaalenii PYR-1, M. flavescens PYRGCK, Mycobacterium RJGII-135, Mycobacterium KR2 and MycobacteriumAP1. Different pathways have been anticipated. Mycobacterium KMS has been used to study the metabolites produced during pyrene transformation. First pathway indicates that pyrene hydroxylation takes place at the 1,2 positions, leading to the formation of 4-hydroxy-perinaphthenone which is the ultimate product so far found only in M. vanbaalenii PYR-1 cultures (Khan et al. 2001; Sinha et al. 2009).
2.3.1 Advantages and disadvantages
Anaerobic bioreactors have several potential advantages over aerobic processes. Due to omission of aeration and the conversion of organic matter to methane, which is an energy source by itself and can be used for temperature control so, less energy is required in this process. High energy recovery, increase in hydrolysis rate at increasing biodegradability suggests that the rate of hydrolysis of particulate organic matter is determined by the adsorption of hydrolytic enzymes to the biodegradable surface sites and the gas generated after through the various stages of breakdown of complex organic compounds can be used as biofuel (Veeken and Hamelers 1999). Apart from the benefits of anaerobic process there is one shortcoming that it need for a longer start-up Period time to develop necessary biomass and the fact that they are much more sensitive to the temperature value.
2.4 Co-metabolism
Co-metabolism is especially important for the degradation of mixtures of polycyclic aromatic hydrocarbons (Chauhan et al. 2008). Co-metabolism entails the parallel oxidation of a non-growth substrate during growth of bacteria on a compatible carbon energy source. It also describes oxidation of non-utilizable substrates by resting cell suspensions grown at the disbursement of substances that are capable of supporting microbial growth. Co-metabolism refers to oxidation of substrate without utilization of energy derived from the oxidation to support microbial growth. Cometabolism is often observed for PAHs compounds which do not enable growth as a single carbon- and energy source (Janke and Fritsche 1985). This is due to long degradation pathways and unnatural structures. Cometabolism of benzo-α-pyrene is significantly increased with enrichment of the soil with phenanthrene as growth substrate for the non growth substrate of pyrene, 3,4 bezopyrene, 1,2 benzoanthracene, 1,3,5,6 dibenzanthracene (McKenna and Heath 1976).
On the other hand, microorganisms with a restricted metabolism are dependent on cometabolism of essential natural compounds. Another distinguishing feature is its high cellular demand, is additionally required for glycine, cysteine, tryptophan, and phospholipid synthesis. The short-chain fatty acid is more suitable than the carbohydrates as the carbon sources for cometabolism (Xie et al. 2009). Dioxygenase and monooxygenase enzymes are responsible for mineralization of various PAHs, including pyrene and BaP. Although the degradation of pyrene increases in organic nutrients to the microcosm inhibited pyrene degradation (Heitkamp and Cerniglia 1989). Study showed that in gram positive bacteria especially mycobacteria, genetical and biochemical data of high molecular weight, polycyclic aromatic hydrocarbon degradation is relatively lower since it possesses extremely resistant cell wall, significantly low growth rate and triggers the activity of cell clumping. In gram positive bacteria, nid, pdo genes encode high molecular weight polycyclic aromatic hydrocarbons (HMWPAH) dioxygenases, whereas in gram negative bacteria nah, pah and phn genes encode low molecular weight polycyclic aromatic hydrocarbons (LMWPAH) dioxygenases. The initial reactions in both the aerobic metabolism and co-metabolism of PAHs are catalyzed by a variety of oxygenase-type enzymes which generate mono- and di-hydroxylated products. It is important to look at the possibilities and extent of both metabolism and cometabolism of those high molecular weight PAHs. Co-metabolic pathway of Napthalene (Denome et al. 1993; Goyal and Zylstra 1997; Kiyohara et al. 1994), Phenanthrene (Moody et al. 2001; Seo et al. 2006, 2007; Prabhu and Phale 2003; Mallick et al. 2007; Krivobok et al. 2003; Pinyakong et al. 2000; Kanaly and Harayama 2000), Fluoranthene (Cerniglia 1992; Kelley et al. 1993; Weissenfels et al. 1991; Kanaly and Harayama 2000) and Benzo-α-pyrene (Schneider et al. 1996; Cerniglia 1992; Heitkamp and Cerniglia 1988; Seo et al. 2009) is shown in Figs. 4, 5, 6, and 7. Gene clusters that code for the catabolism of aromatic compounds are frequently found in mobile genetic elements, as transposons and plasmids, which assist their horizontal gene transfer, and consequently enhance adaptation of specific bacterial genera to novel pollutants. This technique generally employed for the biochemical study of microbial aromatic metabolism (Horvath 1972). Strains that exhibit the phenomenon of co-metabolism in a well-organized manner include Nocardia, Pseudomonas, Xanthomonas, Bacillus, Brevibacterium, Flavobacterium, Aspergillus, Azotobacter, Trichoderma, Vibrio, Achromobacter, Arthrobacter, Hydrogenomonas, Microbacterium, Micrococcus and Streptomyces (Beam and Perry 1973). The manipulation of the catabolic genes from degradative enzymes could solve the problem and boost up the process. The degradation mainly depends upon the adapting response of the microbial communities which include both selective enrichment (resulting in amplification of genes) and genetic changes (mainly includes gene transfer or mutation). Recent approaches have been made to characterize and compare the gene involved in degradation of identical or similar aromatic compounds in nearly diverse or more isolated bacterial genera from diverse topologies (Liang et al. 2006; Kim et al. 2006a, b). The influence of growth medium on cometabolic degradation of PAHs is more effective in nutrient rich medium for enhancement of cometabolic degradation of mixed PAHs concomitant with a rapid removal of metabolites, which could be useful for the bioremediation of mixed PAHs contaminated sites (Zhong et al. 2007).
3 Bacterial degradation of PAHS
Intense research pertaining to bacterial degradation of PAHs which composed of three rings has been well documented. The most commonly reported bacterial species include Acinetobacter calcoaceticus, Alcaligens denitrificans, Mycobacterium sp., Pseudomonas putida, Pseudomonas fluorescens, Pseudomonas vesicularis, Pseudomonas cepacia, Rhodococcus sp., Corynebacterium renale, Moraxella sp., Bacillus cereus, Beijerinckia sp., Micrococcus sp., Pseudomonas paucimobilis and Sphingomonas sp. (Kanaly et al. 2000; Arun et al. 2008). Although most of these bacterial species are reported to degrade low molecular weight PAHs, there are limited reports on degradation of high molecular weight PAHs with more than four benzene rings. Bacterial degradation predominantly occurs via aerobic oxygenase mediated pathways; although an alternative pathway of degradation involving utilization of nitrate as the alternative electron acceptor has been reported. Degradation of PAHs compounds and formation of intermediates by bacterial action (Cerniglia 1992; Juhasz and Naidu 2000; UMBBD 2004) are listed in Table 3. As for mangrove environment, Zhou et al. (2009) utilized 16S rRNA PCR and DGGE to assess the effects of PAHs (before and after PAHs exposure) on the bacterial community. Sequencing of DGGE bands showed that marine bacteria from the genera of Vibrio, Roseobacter, and Ferrimonas were most abundant after PAHs exposure, which suggests that both marine and terrestrial bacteria coexisted in the mangrove sediment (Zhou et al. 2009; Shi et al. 2010). Molina et al. (2009) obtained PAHs degrading microbial consortium C2PL05 from a petrochemical complex in Puertollano (Ciudad Real, Spain) which possessed highly efficient degrading capacity and DGGE analysis revealed uncultured Stenotrophomonas ribotypes as a possible PAHs degrader in the microbial consortium (Molina et al. 2009; Shi et al. 2010). Although several PAHs degrading bacterial species have been isolated, it is not expected that a single isolate would exhibit the ability to degrade completely all PAHs. PCR–DGGE of 16S rRNA gene sequences was used to monitor the bacterial population changes during PAHs degradation of the consortium when pyrene, chrysene, and benzo[a]pyrene were provided together or separately in the TLP cultures (Lafortune et al. 2009; Shi et al. 2010).
3.1 Application of immobilized bacterial cell
Immobilized cells have been used and studied for the bioremediation of numerous toxic chemicals. Immobilization not only simplifies separation and recovery of immobilized cells but also makes the application reusable which reduces the overall cost. Wilson and Bradley (1996) used free suspension and immobilized Pseudomonas sp. to degrade petrol in an aqueous system. Immobilization increase contact between cell and hydrocarbon droplets and enhanced level of rhamnolipids production was studied by Wilson and Bradley (1996; Das and Chandran 2011). Díaz et al. (2002) reported that immobilization of bacterial cells enhanced the biodegradation rate of crude oil compared to free living cells in a wide range of culture salinity. Immobilization can be done in batch mode as well as continuous mode. Packed bed reactors are commonly used in continuous mode to degrade hydrocarbons. Polyvinyl alcohol (PVA) cryogelation was used as an entrapment matrix by Cunningham et al. (2004). Laboratory biopiles was constructed to compare immobilised bioaugmentation with liquid culture bioaugmentation and biostimulation. Immobilised systems were found to be the most successful in terms of percentage removal of diesel after 32 days and immobilized bacteria in alginate beads were able to degrade hydrocarbons, there was no decline in the biodegradation activity was observed in microbial consortium on the repeated use study conducted by Rahman et al. (2006). It can be concluded that immobilization of cells are a promising application in the bioremediation of hydrocarbon contaminated site.
4 Fungal degradation of PAHS
Fungal remediation or mycoremediation is a promising technique for cleanup of contaminated soil. Two main types of fungal metabolism of PAHs degradation; these are mediated by the ligninolytic (also known as the white-rot fungi) and non-ligninolytic fungi. The majority of fungi are non-ligninolytic, as they do not grow on wood, and therefore have no need for the lignin peroxidase enzymes that produced by the ligninolytic fungi. However, many ligninolytic fungi such as Phanerochaete chrysosporium and Pleurotus ostreatus can produce both non-ligninolytic and ligninolytic type enzymes (Hammel et al. 1992; Bezael et al. 1997). The main mechanism involved in the fungal degradation of PAHs is enzymatic transformation by intracellular cytochrome P450 enzymes and extracellular ligninolytic enzymes (Cerniglia 1997). Molecular fingerprinting profiles and selective enumeration showed biostimulation with ground corn cob increased both number and abundance of indigenous aromatic hydrocarbons degraders and changed microbial community composition of soil, which is beneficial to natural attenuation of PAHs. At the same time, bioaugmentation of PAHs with Monilinia strain W5-2 imposed negligible effect on indeginous microbial community. Wu et al. (2008a, b) suggested that fungal remediation is promising in eliminating PAHs especially the part of recalcitrant and highly toxic benzo[α]pyrene, in contaminated soil. It is also the first description of soil bioremediation with Monilinia sp.
4.1 Lygnolytic fungal degradation
There are two types of ligninolytic enzymes; these being peroxidases and laccases. These enzymes are secreted extracellularly, and oxidise organic matter via a non-specific radical based reaction. There are two main types of peroxidase enzyme depending on their reducing substrate type, lignin peroxidase (LP) and manganese peroxidase (MnP), both of which are capable of oxidising PAHs. Laccases, which are phenol oxidase enzymes, are also capable of oxidizing PAHs (Mester and Tien 2000).
Two possible roles of ligninolytic system have been discussed by Collins and Dobson (1996; Steffen et al. 2003): (a) LiP, MnP, and LAC were found to have a pivotal role in the degradation of PAHs, catalyzing the first attack of molecule. (b) Cytochrome P-450 monooxygenase could be responsible for this initial step (Bezalel et al. 1996) White-rot fungi, such as Phanerochaete chrysosporium and Trametes versicolor, were able to mineralize PAHs, and indicated the complete breakdown of PAHs observed by Renner (1999; Pozdnyakova 2012). Enzymes involved in the degradation of PAHs are oxygenase, dehydrogenase and lignolytic enzymes. Fungal lignolytic enzymes are lignin peroxidase, laccase, and manganese peroxidase. They are extracellular and catalyze radical formation by oxidation to destabilize bonds in a molecule. White-rot fungi are a group of fungi that produce ligninolytic enzymes involved in the oxidation of lignin present in wood and other organic matter as mentioned by Haritash and Kaushik (2009). Lignolytic fungal strains responsible for the PAHs degradation and their metabolites listed in Table 4.
4.2 Non lignolytic fungal degradation
The first step in the metabolism of PAHs by nonligninolytic fungi is to oxidise the aromatic ring in a cytochrome P450 monoxygenase enzyme catalyzed reaction to produce an arene oxide (Bezael et al. 1997). In comparison to the oxidation of the aromatic ring by dioxygenase enzymes to form cis dihydrodiols, the monoxygenase enzyme incorporates only one oxygen atom into the ring to form an arene oxide. This is subsequently hydrated via an epoxide-hydrolase catalysed reaction to form a trans dihydrodiol (Baldrian et al. 2000). In addition, phenol derivatives may be produced from arene oxides by the non-enzymatic rearrangement of the compound, which can act as substrates for subsequent sulfation/methylation, or conjugation with glucose, xylose, or glucuronic acid. Most non-ligninolytic fungi are not capable to complete mineralisation of PAHs. These PAHs conjugate are generally less toxic and more soluble than their respective parent compounds. Some of non lignolytic fungal strains are listed in Table 5.
4.3 Application of immobobilized fungal strains
Immobilized form of microorganisms will promote degradation as immobilization is known to offer protection from extremes of pH and toxic compounds in the contaminated soil (Su et al. 2006; Wang et al. 2012). Among all the immobilization methods, physical adsorption on farm byproducts was found to be the optimum method owing to its properties of being highly granular, absorbent, biodegradable and inexpensive (Xu and Lu 2010; Wang et al. 2012). In conclusion, the use of immobilized microorganisms with farm byproducts and nutrients as carrier materials and bulking agents for the remediation of PAH-contaminated site in situ is a promising technology. Wang et al. (2012) studied the degradation of pyrene by the immobilized microorganisms Mucor sp. F2, fungal consortium MF and co-cultures of MB + MF was increased by 161.7 % (P < 0.05), 60.1 % (P < 0.05) and 59.6 % (P < 0.05) after 30 days, respectively. When compared with free F2, MF and MB + MF. Results indicated that immobilization improved stability of laccase to temperature, pH, inhibitors and storage time compared with the free enzyme. Tao et al. (2009) were done research work on strain Sphingomonas sp. GY2B is a high efficient phenanthrene-degrading strain isolated from crude oil contaminated soils that displays broad-spectrum degradation ability towards PAHs and related aromatic compounds. The immobilization of strain GY2B possesses a good potential for application in the treatment of industrial wastewater containing phenanthrene and other related aromatic compounds. A Comparative study of different immobilize fungal strains is shown in Table 6.
5 Factors affecting PAHs degradation
A number of limiting factors have been recognized to affect the biodegradation of PAHs. It is apparent that environmental factors that vary from site to site (such as soil pH, nutrient availability and the bioavailability of the contaminant) can influence the process of bioremediation by inhibiting growth of the pollutant-degrading microorganisms. The main environmental factors that could affect the feasibility of bioremediation are summarized in the following sections.
5.1 Temperature
Temperature has a considerable effect on the ability of the in situ microorganisms to degrade PAHs and, in general, most contaminated sites will not be at the optimum temperature for bioremediation during every season of the year. According to (Margesin and Schinner 2001) the solubility of PAHs increases with an increase in temperature, which increases the bioavailability of the PAH molecules. In addition, oxygen solubility decreases with increasing temperature, which will reduce the metabolic activity of aerobic microorganisms. Biodegradation of PAHs can occur over a wide temperature range however, most studies tend to focus on mesophilic temperatures rather than the efficiency of transformations at very low or high temperatures. Lau et al. (2003) were reported that the laccase and manganese peroxidase enzymes of ligninolytic fungi have a temperature optimum of ~50 and >75 °C respectively in spent-mushroom compost during the degradation of PAHs, with over 90 % degradation of the contaminating PAHs occurring at these temperatures.
5.2 pH
Many sites contaminated with PAHs are not at the optimal pH for bioremediation. For example, Phenanthrene degradation in liquid culture has been investigated by Wong et al. (2002) at a range of pH values (pH 5.5–7.5) with Burkholderia cocovenenas, an organism isolated from a petroleum-contaminated soil, bacterial growth was not significantly affected by the pH. Phenanthrene removal was only 40 % at pH 5.5 after 16 days, whereas at circum-neutral pH values, phenanthrene removal was 80 %. Sphingomonas paucimobilis (strain BA 2) was however more sensitive to the pH of growth media, with the degradation of the PAHs phenanthrene and anthracene significantly inhibited at pH 5.2 relative to pH 7 reported by Kastner et al. (1998).
5.3 Oxygen
For the reduction of complex hydrocarbons into substrates, require an electron acceptor, molecular oxygen being most common. In the absence of molecular oxygen, nitrate, iron, bicarbonate, nitrous oxide and sulfate, have been shown to act as an alternate electron acceptor during hydrocarbon degradation. Bioremediation of organic contaminants (PAHs) can proceed under both aerobic and anaerobic conditions; most work has tended to concentrate upon the dynamics of aerobic metabolism of PAHs. This is in part due to the ease of study and culture of aerobic microorganisms relative to anaerobic microorganisms. To maintain adequate oxygen levels for aerobic metabolism for in situ treatments include hydrogen peroxide for sub-surface contamination. The aerobic biodegradation of hydrocarbons has been reported to be highly relative to anaerobic biodegradation (Rockne and Strand 1998).
5.4 Nutrient availability
Basically, microorganisms require mineral nutrients such as nitrogen, phosphate and potassium (N, P and K) for cellular metabolism and successful growth. In contaminated sites, where organic carbon levels are often high due to the nature of the pollutant, available nutrients can become rapidly depleted during microbial metabolism. Therefore it is common practice to supplement contaminated land with nutrients, generally nitrogen and phosphates to stimulate the in situ microbial community to enhance bioremediation. The amounts of N and P required for optimal microbial growth and hence bioremediation have been previously estimated from the ratio of C:N:P in microbial biomass (100:15:364 and 120:10:165). However, a recent study has shown that optimal microbial growth and creosote biodegradation occurred in soil with a much higher C:N ratio (25:1) than those predicted from the ratio in microbial biomass, with lower C:N ratios (5:1) causing no enhancement in microbial growth (Alexander 1977; Atagana et al. 2003).
5.5 Bioavailability
The effect of physico-chemical and microbiological factors on the rate and extent of biodegradation and is believed to be one of the most important factors in bioremediation. PAH compounds have a low bioavailability, and are classed as hydrophobic organic contaminants. The larger the molecular weight of the PAH, the lower its solubility, which in turn reduces the accessibility of the PAH for metabolism by the microbial cell. In addition, PAHs can undergo rapid sorption to mineral surfaces (i.e. clays) and organic matter (i.e. humic and fulvic acids) in the soil matrix. The longer that the PAH is in contact with soil, the more irreversible the sorption, and the lower is the chemical and biological extractability of the contaminant. This phenomenon is known as ‘ageing’ of the contaminant. Therefore the bioavailability of a pollutant is linked to its persistence in a given environment (Semple et al. 2003; Peltola 2010).
5.6 Surfactant: bioavailability enhancer
Reduction of PAHs from the surface of minerals and organic matter can be achieved by the use of surface-active agents (also known as surfactants or detergents). The common chemical surfactant such as triton x-100, tween 80 and sodium dodecyl sulfate are petroleum derived products. The salient mechanism which are involved in the surfactant—amended remediation are: lowering of interfacial tension, surfactant solubilization of hydrophobic organic compounds and, the phase transfer of organic compounds from soil sorbed to pseudo aqueous phase (Laha et al. 2009).
5.7 Salinity
Few reports have been published documenting the effect of salt on hydrocarbon biodegradation in soil. High salt concentrations can inhibit the activity of microbes that are not adapted to salt. Possible reasons for this effect include direct inhibition of metabolic activity because of unfavorable high osmotic potential of the microbe’s environment (Amatya et al. 2002), and altered solubility or sorption of toxic or essential ions. The inhibitory salt effects observed included longer lag times and decreased rates and extents of mineralization. Børresen and Rike (2007) found that low levels of NaCl (0.3 % w/w NaCl) slightly stimulated rates of hexadecane mineralization in an Arctic soil. This stimulation may be explained if the salt provides a more ironically balanced medium for the microbes, or a medium which disperses clays and thus provides a larger surface area for attachment of cells, or for access to trace nutrients.
6 Molecular techniques for characterization of PAH’s degrading microorganisms
A variety of molecular methods based on direct isolation and analysis of nucleic acids, proteins, and lipids from environmental sample and revealed structural and functional information about microbial community. PAHs microbial degradation were carried out by the following pathway (Source-Zhang et al. 2002).
6.1 Polymerase chain reaction (PCR) amplification technique
With the rapid development of molecular-biology, modern taxonomy prefer sequencing technologies of molecular markers such as 16S rRNA (prokaryots) or 18S rRNA (eukaryots). These technologies allow the identification of colonies isolated from microbial consortia and the establishment of phylogenetic relationships between them (Molina et al. 2009). In addition to taxonomy, PCR combined with other approaches could also be used to estimate in situ bacterial community how pollution affects structure of bacterial community and composition of sediments (Ding et al. 2013). Function and application of different PCR techniques is presented in Table 7. The recent development of real-time PCR devices has made quantitative PCR much easier. Besides single-cell level detection, the quantitative PCR approach utilizing bulk DNA from natural bacterial communities may be an effective approach to monitor target bacteria. Distinguishing closely related strains is difficult because of high conservation of 16s rRNA in prokaryotes, 16s rRNA PCR amplification fragment could not be relied on completely sometimes (Tomotada and NaSu 2001).
6.2 Fingerprinting techniques based on 16s rRNA
Microbial fingerprinting methods are a category of techniques that differentiate microorganisms or groups of microorganisms based on unique characteristics of a universal component or section of a biomolecule (e.g. phospholipids, DNA, or RNA; ITRC EMD team 2011). Fingerprinting techniques based on 16S rRNA such as DGGE, SSCP, T-RFLP and RISA could analysis the diversity and dynamics of the whole community at molecular level. Fingerprinting methods are used to provide an overall view of the microbial community, indications of microbial diversity, and insight into the types of metabolic processes occurring in the microorganisms present in the sample. Microbial fingerprinting methods can identify when adverse conditions (e.g. low pH), either natural or following a remedy (e.g. chemical oxidation). Microbial fingerprinting methods can be used to determine whether the overall microbial community has recovered or responded to remedial actions. Several microbial fingerprinting techniques can be used to identify the predominant microorganisms present in the sample and to describe the microbial community. Information about complex dynamic in microbial communities can undergo by PAHs changes and seasonal fluctuation fingerprinting technique based on 16s rRNA-DGGE, SSCP, T-RFLP and RISA can analyze the diversity and dynamics of the whole community at molecular level. Function and application are discussed in Table 8.
6.3 Fluorescence in situ hybridization
Fluorescence in situ hybridization (FISH) enables in situ phylogenetic identification and enumeration of individual microbial cells by whole cell hybridization with oligonucleotide probes (Amann et al. 1995). The FISH probes are generally 18–30 nucleotides long and contain a fluorescent dye that allows detection of probe bound to cellular rRNA by epifluorescence microscopy. FISH can be combined with flow cytometry for a high resolution automated analysis of mixed microbial populations. The FISH method was used to follow the dynamics of bacterial populations in agricultural soils treated with s-triazine herbicides (Caracciolo et al. 2010).
FISH can detect microorganisms across all phylogenetic levels, whereas immunofluorescence techniques are limited to the species and sub-species levels. FISH is more sensitive than immunofluorescence because non-specific binding to soil particles does not typically occur (Amann et al. 1995). FISH probes can be generated without prior isolation of the microorganism, whereas pure cultures are needed in immunofluorescence studies for generating specific antibodies (Hahn et al. 1992; Hill et al. 2000). FISH is a powerful tool that can be used not only for studying individuals within a population, but also has potential uses for studying population dynamics, tracking microorganisms released into the environment (e.g. for biological control or bioremediation), epidemiology, and microbial ecology of economically important plant pathogens in agricultural soils (Hahn et al. 1992; Kirchhof et al. 1997; Wullings et al. 1998; Hill et al. 2000).
6.4 DGGE
Is commonly used in PAHs biodegradation this technique is based on the electrophoretic separation of PCR generated double stranded DNA acrylamide gel. The use of denaturing gradient and temperature gradient gel electrophoresis (DGGE/TGGE) for separating individual amplicons has been described (Muyzer et al. 1993; Ferris and Ward 1997; Heuer et al. 1997; Muyzer and Smalla 1998). This technique allows to separate mixtures of PCR products that are of the same length but differ in sequence. The separation power of this technique rests with the melting behavior of the double stranded DNA molecule.16s rRNA and DGGE profiling use to assess the effects of PAHs (before and after PAHs exposure) on the bacterial community.
Microbial diversity is underestimated because it is difficult to extract and amplify all microorganism-genome and DNA. Sometime microbial diversity is overestimated when a single strain might presence multiple bands in DGGE gel. 16s rRNA presents multiple copies of chromosome.
6.5 DNA hybridization technique (DNA microarray, DNA probe technique)
DNA microarrays have been used primarily to provide a high-throughput and comprehensive view of microbial communities in environmental samples. The PCR products amplified from total environmental DNA is directly hybridized to known molecular probes, which are attached on the microarrays (Gentry et al. 2006). After the fluorescently labeled PCR amplicons are hybridized to the probes, positive signals are scored by the use of confocal laser scanning microscopy. DNA microarrays has the advantage that the relative amount of transcripts from the whole genome may be easily determined compared to proteomics. Membrane-bound proteins are problematic and roughly 2/3 of E. coli proteins have not been identified by non-gel proteomic techniques (Han and Lee 2006; Wood 2008). However, transcriptome profiling often assumes that changes in transcription may be used to predict changes in protein formation that may not always be correct but is often true for procaryotes since regulation occurs primarily at the level of transcription. DNA microarrays used in microbial ecology could be classified into different types of microarrays for environmental studies (DeSantis et al. 2007) Detailed applications is presented in Table 9.
6.6 DNA shuffling
Directed evolution or DNA shuffling (Crameri and Stemmer 1995; Crameri et al. 1996; Stemmer 1994a, b; Wood 2008) is a powerful mutagenesis technique that mimics the natural molecular evolution of genes. It can introduce multiple mutations into a gene in order to create new enzymatic activity. It is still difficult to rationally predict the amino acid changes that occur during DNA shuffling and that are necessary to create the new activity. DNA shuffling has been used successfully to create a biocatalyst with higher degradation rates for polyaromatic hydrocarbons (naphthalene, phenanthrene, fluorene, and anthracene).
6.7 Applications of new sequencing technologies: meta-approaches
Particularly meta-transcriptomics, and meta-proteomics are new approaches which in the coming years will be more and more routinely used in environmental studies. These refer to the collection and analysis of transcription (mRNA) and protein profile information from microbial communities. Metagenomic libraries are a powerful molecular technique which has been flourished for the identification of the desired catabolic genes. Basically, metagenomic is a culture dependent genomic analysis; it is either function driven approach or sequence driven approach, of total microbial communities, which provides access to retrieve unknown sequences (Schloss and Handelsman 2003). Metagenomics provides a view not only of the community structure (species phylogeny, richness, and distribution) but also of the functional (metabolic) potential of a community because virtually about all genes are captured and sequenced. In principle, metagenomic techniques are based on the concept that the entire genetic composition of environmental microbial communities could be sequenced and analyzed in the same way as sequencing a whole genome of a pure bacterial culture.
Metaproteomics commonly known as environmental proteomics, deals with the large-scale study of proteins expressed by environmental microbial communities at a given point in time (Wilmes and Bond 2006; Keller and Hettich 2009). Compared to other cell molecules such as lipids and nucleic acids, protein biomarkers are more reliable and provide a clearer picture of metabolic functions than functional genes or even the corresponding mRNA transcripts of microbial communities (Wilmes and Bond 2006). Proteins involved in the degradation of phenanthrene regulated proteins of strain proteome analysis of Sphe3 cells grown on phenanthrene or glucose by two-dimensional gel electrophoresis (2DE) in combination with mass spectrometry (MALDITOF MS and MS/MS). Several PAH-degrading proteins were identified including 1-hydroxy-2-naphthoate dioxygenase and protocatechuate dioxygenase (Koukkou and Drainas 2008). Proteomic profiling of microbial communities provides critical information on protein abundances and protein–protein interactions, which could not be achieved by DNA/RNA molecular techniques such as metatranscriptomics and metagenomics (Keller and Hettich 2009). Once the proteins are identified, they could be linked to corresponding metagenomic sequences to link metabolic functions to individual microbial species. The limitation of metaproteomic and metagenomic has been overcome by combining these two approaches together under the name of “proteogenomics”. In community proteogenomics, total DNA and proteins are extracted from the same sample, which allows linking of biological functions to phylogenetic identity with greater confidence. The metagenomic part of the proteogenomic approach plays a very significant role and increases the identification of protein sequences by metagenomic analysis of the same sample from which the proteins were extracted (Rastogi and Sani 2011). The proteogenomics approach was applied to decipher phyllosphere bacterial communities in a study by Delmotte et al. (2009). Metatranscriptomics (or environmental transcriptomics) allows monitoring of microbial gene expression profiles in natural environments by studying global transcription of genes by random sequencing of mRNA transcripts pooled from microbial communities at a particular time and place (Moran 2009). Metatranscriptomics is particularly suitable for measuring changes in gene expression and their regulation with respect to changing environmental conditions. A method for selectively enriching mRNA by subtractive hybridization of rRNA has been developed and evaluated for the gene transcript analysis of marine and freshwater bacterioplankton communities, which revealed the presence of many transcripts that were linked to biogeochemical processes such as sulfur oxidation (soxA), assimilation of C1 compounds (fdh1B), and acquisition of nitrogen via polyamine degradation (aphA) (Poretsky et al. 2005).
7 Conclusion
Removal of persistent organic pollutants from the environment is a real world problem. Various research studies have been proven that bioremediation is a promising tool for degradation of PAHs and many more compounds. But this field needs better understanding of the mechanism of biodegradation pathways has a high ecological significance that depends on the indigenous microorganisms to transform or mineralize the organic contaminants. There is a urgent need to address this issue for future research on expanding our knowledge on the practical application of cometabolic processes, bioaugmentation and bacterial–fungal co-cultures. As more specific gene probes are developed, improved DNA extraction techniques could provide a much more in depth image of microbial metabolic function, diversity and interdependence on each other. A combination of several techniques should be applied to investigate the diversity, metabolic function, and ecology of microorganisms. Therefore based on the present review, it may be concluded that along with cleaning strategy of bioremediation, enzyme play vital role in enhancing the microbial degradation and at genomic level molecular tools provide specific gene location in quantitative manner which deals with microbial diversity and function analysis.
7.1 Future challenges
Studies about PAHs biodegradation by microbes is a most focused branch in environmental research field and research emphasis has been changed from finding PAHs degrading microorganisms into metabolic pathways of microbes, genetic regulation and construction of high efficiency engineering microorganisms. The efficiency of PAHs degradation can be significantly improved by addressing key issues as tolerance to various compounds of this family, constitutive expression of the catabolic genes and substrate-specificity; kinetics and the stability of the encoded enzyme. Apart from quantitative assessment qualitative analysis of microbial communities is the greatest challenge due to significant biases associated with nucleic acid isolation. It requires more advanced DNA/RNA extraction techniques for environmental samples such as soil and water. All of the molecular approaches available for analysis of microbial community structure and functions have advantages and limitations associated with them. Biomolecular engineering can be successfully used to improve the capabilities of the enzymes or microorganisms in bioremediation systems. However, there are several limitations i.e. creation of enzymes with novel functions represents an overwhelming challenge in biomolecular engineering. Research is usually focused on altering enzymes that can perform a reaction similar to the desired one. Thus, it might be difficult to apply biomolecular engineering to the bioremediation of xenobiotic compounds, which are not known to be biodegradable. An effort can be made with rational design in the future when our knowledge of the protein structure–function, folding, mechanism and dynamics is significantly improved. However, the utility of constructed organisms in dealing with problems related to environmental pollution in yet to be tested.
References
Abu Laban N, Selesi D, Rattei T, Tischler P, Meckenstock RU (2010) Identification of enzymes involved in anaerobic benzene degradation by a strictly anaerobic iron-reducing enrichment culture. Environ Microbiol 12:2783–2796
Aitken MD, Stringfellow WT, Nagel RD, Kazunga C, Chen SH (1998) Characteristics of phenanthrene-degrading bacteria isolated from soils contaminated with polycyclic aromatic hydrocarbons. Can J Microbiol 44:743–752
Alcade M, Bulter T, Arnold FH (2002) Colorimetric assays for biodegradation of polycyclic aromatic hydrocarbons by fungal laccases. J Biomol Screen 7(6):547–553
Alexander M (1977) Introduction to soil microbiology, 2nd edn. Wiley, NewYork, p 207
Allen CCR, Boyd DR, Hempenstall F, Larkin MJ, Sharma ND (1999) Contrasting effects of nonionic surfactant on the biotransformation of polycyclic aromatic hydrocarbons to cis-dihydrodiols by soil bacteria. Appl Environ Microbiol 65:1335–1339
Amann RI, Ludwig W, Schleifer KH (1995) Identification and in situ detection of individual microbial cells without cultivation. Microbiol Rev 59:143–169
Amatya PL, Hettiaratchi JPA, Joshi RC (2002) Biotreatment of flare pit waste. J Can Petrol Technol 41:30–36
Arun A, Raja PP, Arth R, Ananthi M, Kumar KS, Eyin M (2008) Polycyclic aromatic hydrocarbons (PAHs) biodegradation by Basidiomycetes Fungi, PseudomonasIsolate and their cocultures: comparative in vivo and in silico approach. Appl Biochem Biotechnol 151:13–142
Atagana HI, Haynes RJ, Wallis FM (2003) Optimization of soil physical and chemical conditions for the bioremediation of creosote-contaminated soil. Biodegradation 14:297–307
Baastiaens L, Springael D, Dejonghe W, Wattiau P, Verachtert H, Diels L (2001) A transcriptional luxAB reporter fusion responding to fluorene in 305 Sphingomonas sp. LB126 and its initial characterization for whole-cell bioreporter purposes. Res Microbiol 15:849–859
Baldrian P, Der Wiesche IN, Gabriel J, Nerud F, Zadražil F (2000) Influence of cadmium and mercury on activities of ligninolytic enzymes and degradation of polycyclic aromatic hydrocarbons by Pleurotusostreatusin soil. Appl Environ Microbiol 66(6):2471–2478
Bamforth SM, Manning DAC, Singleton I (2005) Naphthalene transformation by the Pseudomonas at an elevated pH. J Chem Technol Biotechnol 80:723–736. doi:10.1002/jctb.1276
Banerjee DK, Fedorak PM, Hashimoto A, Masliyah JH, Pickard MA, Gray MR (1995) Monitoring the biological treatment of anthracene-contaminated soil in a rotating-drum bioreactor. Appl Microbiol Biotechnol 43:521–528
Beam HW, Perry JJ (1973) Co-metabolism as a factor in microbial degradation of cycloparaffinic hydrocarbons. Arch Microbiol 91:87–90
Bergmann F, Selesi D, Weinmaier T, Tischler P, Rattei T, Meckenstock RU (2011) Genomic insights into the metabolic potential of the polycyclic aromatic hydrocarbon degrading sulfate-reducing Deltaproteobacterium N47. Environ Microbiol 13:1125–1137
Bezalel Y, Hadar P, Fu P, Freeman JP, Cerniglia CE (1996) Initial oxidation products in the metabolism of pyrene, anthracene, fluorene, and dibenzothiophene by the white rot fungus Pleurotusostreatus. Appl Environ Microbiol 62(7):2554–2559
Bezalel L, Hadar Y, Cerniglia CE (1997) Enzymatic mechanisms involved in phenanthrene degradation by the white rot fungus Pleurotusostreatus. Appl Environ Microbiol 63:2495–2501
Boldrin B, Andreas T, Fritzche C (1993) Degradation of phenanthrene, fluorene, fluoranthene and pyrene by a Mycobacterium spp. Appl Environ Microbiol 59:1927–1930
Boonchan S (1998) Biodegradation of polycyclic aromatic hydrocarbons: application of fungal–bacterial cocultures and surfactants. Thesis, Victoria University of Technology, Melbourne Victoria
Boonchan S, Britz FL, Stanley GA (2000) Degradation and mineralization of high-molecular-weight polycyclic aromatic hydrocarbons by defined fungal-bacterial cocultures. Appl. Environ. Microbiol. 66:1007–1019
Børresen MH, Rike AG (2007) Effects of nutrient content, moisture content and salinity on mineralization of hexadecane in an Arctic soil. Cold Regions Sci Technol 48:129–138
Bulter T, Alcalde M, Sieber V, Meinhold P, Schlachtbauer C, Arnold FH (2003) Functional expression of a fungal laccase in Saccharomyces cerevisiae by directed evolution. Appl Environ Microbiol 69:987–995
Bustin SA, Benes V, Nolan T, Pfaffl MW (2005) Quantitative real-time RT-PCR—a perspective. J Mol Endocrinol 34:597–601
Caracciolo AB, Bottoni P, Grenni P (2010) Fluorescence in situ hybridization in soil and water ecosystems: a useful method for studying the effect of xenobiotics on bacterial community structure. Toxicol Environ Chem 92:567–579
Carmichael AB, Wong LL (2001) Protein engineering of Bacillus megaterium CYP102—the oxidation of polycyclic aromatic hydrocarbons. Eur J Biochem 268:3117–3125
Casillas RP, Crow SA, Heinze TM, Deck J, Cerniglia CE (1996) Initial oxidation and subsequent conjugative metabolites produced during the metabolism of phenanthrene by fungi. J Ind Microbiol 16:205–215
Cerniglia CE (1984) Microbial degradation of polycyclic aromatic hydrocarbons. Adv Appl Microbiol 30:31–71
Cerniglia CE (1992) Biodegradation of polycyclic aromatic hydrocarbons. Biodegradation 3:351–368
Cerniglia CE (1997) Fungal metabolism of polycyclic aromatic hydrocarbons: past, present and future applications in bioremediation. J Ind Microbiol Biotechnol 19:324–333
Cerniglia CE, Heitkamp MA (1989) In: Varanasi U (ed) Metabolism of Polycyclic aromatic hydrocarbon in Aquatic Environment. CRC Press Inc., Boca Raton FL
Cerniglia CE, Kelly DW, Freeman JP, Miller DW (1986) Microbial metabolism of pyrene. Chem Biol Interact 57:203–216
Chauhan A, Fazlurrahman Oakeshott JG, Jain RK (2008) Bacterial metabolism of polycyclic aromatic hydrocarbons: strategies for bioremediation. J Ind Microbiol 48:95–113
Chen SH, Aitken MD (1999) Salicylate stimulates the degradation of high molecular weight polycyclic aromatic hydrocarbons by Pseudomonas saccharophila P15. Environ Sci Technol 33:435–439
Collins PJ, Dobson ADW (1996) Oxidation of fluorene and phenanthrene by Mn(II) dependent peroxidase activity in whole cultures of Trametes (coriolus) versicolor. Biotechnol Lett 18:801–804
Collins JF, Brown JP, Dawson SV, Marty MA (1991) Risk assessment for benzo[a] pyrene. Regul Toxicol Pharmacol 13:170–184
Cooper CS, Grover PL, Sims P (1983) The metabolism and activation of benzo(a)pyrene. Progress Drug Metabol 7:295–396
Crameri A, Stemmer WPC (1995) Combinatorial multiple cassette mutagenesis creates all the permutations of mutant and wildtype sequences. Biotechniques 18:194–196
Crameri A, Whitehorn EA, Tate E, Stemmer WPC (1996) Improved green fluorescent protein by molecular evolution using DNA shuffling. Nat Biotechnol 14:315–319
Cunningham CJ, Ivshina IB, Lozinsky VI, Kuyukina MS, Philp JC (2004) Bioremediation of diesel contaminated soil by microorganisms immobilised in polyvinyl alcohol. Int Biodeterior Biodegrad 54(2–3):167–174
Das N, Chandran P (2011) Microbial degradation of petroleum hydrocarbon contaminants: an overview biotechnology research international volume. Article ID 941810. p 13. doi:10.4061/2011/941810
Da-Silva M, Cerniglia CE, Pothuluri JV, Canhos VO, Esposito E (2003) Screening filamentous fungi isolated from estuarine sediments for the ability to oxidise polycyclic aromatic hydrocarbons. World J Microbiol Biotechnol 19:399–405
Delmotte N, Knief C, Chaffron S, Innerebner G, Roschitzki B, Schlapbach R, Von Mering C, Vorholt JA (2009) Community proteogenomics reveals insights into the physiology of phyllosphere bacteria. Proc Natl Acad Sci USA 106:16428–16433
Denome SA, Stanley DC, Olson ES, Young KD (1993) Metabolism of dibenzothiophene and naphthalene in Pseudomonas strains: complete DNA sequence of an upper naphthalene catabolic pathway. J Bacteriol 175:6890–6901
Derz K, Klinner U, Schupan I, Stackebrandt E, Kroppenstedt RM (2005) Mycobacterium pyrenivorans sp. nov., a novel polycyclic-aromatichydrocarbon-degrading species. Int J Syst Evolut Microbiol 54:2313–2317
DeSantis TZ, Brodie EL, Moberg JP, Zubieta IX, Piceno YM, Andersen GL (2007) High-density universal 16S rRNA microarray analysis reveals broader diversity than typical clone library when sampling the environment. Microb Ecol 53:371–383
Díaz MP, Boyd KG, Grigson SJW, Burgess JG (2002) Biodegradation of crude oil across a wide range of salinities by an extremely halotolerant bacterial consortium MPD-M, immobilized onto polypropylene fibers. Biotechnol Bioeng 79(2):145–153
Ding A, Sun Y, Dou J, Cheng L, Jiang L, Zhang D, Zhao X (2013) Characterizing microbial activity and diversity of hydrocarbon-contaminated sites 137-160.http://dx.doi.org/10.5772/50480
Dodor DE, Hwang HM, Ekunwe SIN (2004) Oxidation of anthracene and benzo[a]pyrene by immobilized laccase from Trametesversicolor. Enzyme Microb Technol 35:210–217
Dubey SK, Tripathi AK, Upadhyay SN (2006) Exploration of soil bacterial communities for their potential as bioresource. Bioresour Technol 97(17):2217–2224
Erickson DC, Loehr RC, Neuhauser EF (1993) PAH loss during bioremediation of manufactured gas plant site soil. Water Res 27:911–919
Fan CE, Reinfelder JR (2003) Phenanthrene accumulation kinetics in marine diatoms. Environ Sci Technol 37:3405–3412
Ferris MJ, Ward DM (1997) Seasonal distributions of dominant 16S rRNA-defined populations in a hot spring microbial mat examined by denaturing gradient gel electrophoresis. Appl Environ Microbiol 63:1375–1381
Fisher MM, Triplett EW (1999) Automated approach for ribosomal intergenic spacer analysis of microbial diversity and its application to freshwater bacterial communities. Appl Environ Microbiol 65:4630–4636
Freeman WM, Walker SJ, Vrana KE (1999) Quantitative RT-PCR: pitfalls and potential. BioTechniques 26(1):112–122, 1245
Furono S, Pazolt K, Rabe C, Neutr TR, Harm H, Wickly LY (2009) Fungal mycelia allow chemotactic dispersal of polycyclic aromatic hydrocarbon degrading bacteria in water unsaturated system. Environ Microbiol 12(6):1391–1398
Gentry TJ, Wickham GS, Schadt CW, He Z, Zhou J (2006) Microarray applications in microbial ecology research. Microb Ecol 52:159–175
Gianfreda L, Bollag JM (2002) Isolated enzymes for the transformation and detoxification of organic pollutants. In: Burns RG, Dick RP (eds) Enzymes in the environment: activity, ecology, and applications. Marcel Dekker Inc, New York, pp 495–538
Gianfreda L, Rao MA (2004) Potential of extra cellular enzymes in remediation of polluted soils: a review. Enzyme Microb Technol 35:339–354
Gibson DT, Subranian V (1984) Microbial degradation of aromatic hydrocarbons. In: Gibson DT (ed) Microbial degradation of organic compounds. Marcel Dekker Inc, New York, pp 181–252
Gibson DT, Venkatanarayana D, Jerina M, Yagi H, Yeh H (1975) Oxidation of carcinogens benzo(a)pyrene and benzo(a) anthracene to dihydrodiols by a bacterium. Science 189:295–297
Goyal AK, Zylstra GJ (1997) Genetics of naphthalene and phenanthrene degradation by Comamonas testosteroni. J Ind Microbiol Biotechnol 19:401–407
Grimm AC, Harwood CS (1997) Chemotaxis of pseudomonas putida to the polyaromatic hydrocarbon napthalene. Appl Envion Microb 63:4111–4115
Guillen MD, Sopelana P, Partearroyo MA (1997) Food as a source of polycyclic aromatic carcinogens. Rev Environ Health 12:133–146
Habe H, Omori T (2003) Genetics of polycyclic aromatic hydrocarbon degradation by diverse aerobic bacteria. Biosci Biotechnol Biochem 67:225–243
Hahn D, Amann RI, Ludwig W, Akkermans ADL, Schleifer KH (1992) Detection of microorganisms in soil after in situ hybridization with rRNA-targeted, fluorescently labeled oligonucleotides. J Gen Microbiol 138:879–887
Hammel KE, Kalyanaraman B, Kirk TK (1986) Oxidation of polycyclic aromatic hydrocarbons and dibenzo(p)dioxins by Phanerochaete chrysosporiumligninase. J Biol Chem 261:16948–16952
Hammel KE, Gai WZ, Green B, Moen MA (1992) Oxidative degradation of phenanthrene by the ligninolytic fungus Phanerochaete chrysosporium. Appl Environ Microbiol 58:1832–1838
Han M-J, Lee SY (2006) The Escherichia coli proteome: past, present, and future prospects. Microbiol Mol Biol Rev 70:362–439
Harayama S (1997) polycyclic aromatic hydrocarbon bioremediation design. Curr Opin Biotechnol 8:268–273
Harayama S, Timmis KN (1992) Aerobic biodegradation of aromatic hydrocarbons by bacteria. In: Sigel H, Sigel A (eds) Metal ions in biological systems, vol 28. Marcel Dekker, New York, pp 99–156
Harford-Cross CF, Carmichael AB, Allan FK, England PA, Rouch DA, Wong LL (2000) Protein engineering of cytochrome P450(cam) (CYP101) for the oxidation of polycyclic aromatic hydrocarbons. Protein Eng 13:1218
Haritash AK, Kaushik CP (2009) Biodegradation aspects of polycyclic aromatic hydrocarbons (PAHs): a review. J Hazard Mater 169(1–3):1–15. doi:10.1016/j.jhazmat.2009.03.137
Harvey RG (1996) Mechanisms of carcinogenesis of polycyclic aromatic hydrocarbons. Polycycl Aromat Compd 9:1–23
Heitkamp MA, Cerniglia CE (1988) Mineralization of polycyclic aromatic hydrocarbons by a bacterium isolated from sediment below an oil field. Appl Environ Microbiol 54:1612–1614
Heitkamp MA, Cerniglia CE (1989) Polycyclic aromatic hydrocarbon degradation by a Mycobacterium sp. in microcosms containing sediment and water from a pristine ecosystem. Appl Environ Microbiol 55:1968–1973
Heuer H, Krsek M, Baker P, Smalla K, Wellington EMH (1997) Analysis of actinomycete communities by specific amplification of genes encoding 16S rRNA and gel-electrophoretic separation in denaturing gradients. Appl Environ Microbiol 63:3233–3241
Hill GT, Mitkowski NA, Aldrich-Wolfe L, Emele LR, Jurkonie DD, Ficke A, Maldonado-Ramireza S, Lyncha ST, Nelson EB (2000) Methods for assessing the composition and diversity of soil microbial communities. Appl Soil Ecol 15:25–36
Horvath RS (1972) Microbial co-metabolism and the degradation of organic compounds in nature. Bacteriol Rev 36(2):146–155
Jain RK, Kapur M, Labana S, Lal B, Sarma PM, Bhattacharya D, Thakur S (2005) Microbial diversity: application of microorganisms for the biodegradation of xenobiotics. Curr Sci 89(1):101–112
Janke D, Fritsche W (1985) Nature and significance of microbial cometabolism of xenobiotics. J Basic Microbiol 25:603–619
Joyce C (2002) Quantitative RT-PCR. A review of current methodologies. Methods Mol Biol 193(83):92. doi:10.1385/1-59259-283-X:083
Juhasz A, Naidu R (2000) Enrichment and isolation of non-specific aromatic degraders from unique uncontaminated (Plant and Fecal Material) sources and contaminated coils. J Appl Microbiol 89:642–650
Kallimanis A, Frillingos S, Drainas C, Koukkou AI (2007) Taxonomic identification, phenanthrene uptake activity and membrane lipid alterations of the PAH degrading Arthrobacter sp. strain Sphe3. Appl Microbiol Biotechnol 76:709–717
Kanaly RA, Harayama S (2000) Biodegradation of high molecular weight polycyclic aromatic hydrocarbons by bacteria. J Bacteriol 182(8):2059–2067
Kanaly R, Bartha R, Watanabe K, Harayama S (2000) Rapid mineralisation of benzo(a)pyrene by a microbial consortium growing on diesel fuel. Appl Environ Microbiol 66(10):4205–4211
Kang XP, Jiang T, Li YQ (2010) A duplex real-time RT-PCR assay for detecting H5N1 avian influenza virus and pandemic H1N1 influenza virus. Virol J 7:113. doi:10.1186/1743-422X-7-113
Kapoor M, Lin W (1984) Studies on the induction of aryl hydrocarbon (benzo(a)pyrene) hydroxylase in Neurosporacrassa, and itssuppression by sodium selenite. Xenobiotica 14:903–915
Kastner M, Mahro B (1996) Microbial degradation of polycyclic aromatic hydrocarbons in soils affected by the organic matrix of compost. Appl Microbiol Biotechnol 44:668–675
Kastner M, Breuer-Jammali M, Mahro B (1998) Impact of inoculation protocol, salinity and pH on the degradation of polycyclic aromatic hydrocarbons and survival of PAH-degrading bacteria introduced into soil. Appl Environ Microbiol 64(1):359–362
Keller M, Hettich R (2009) Environmental proteomics: a paradigm shift in characterizing microbial community. Microbiol Mol Biol Rev 73(1):62–70. doi:10.1128/MMBR.00028-08
Kelley I, Freeman JP, Evans FE, Cerniglia CE (1993) Identification of metabolites from the degradation of fluoranthene by Mycobacterium sp. strain PYR-1. Appl Environ Microbiol 59:800–806
Khan AA, Wang RF, Cao WW, Doerge DR, Wennerstrom D, Cerniglia CE (2001) Molecular cloning, nucleotide sequence and expression of genes encoding a polycyclic ring dioxygenase from Mycobacterium sp. strain PYR-1. Appl Environ Microbiol 67:3577–3585
Kim YH, Cho K, Yun SH, Kim JY, Kwon KH, Yoo JS, Kim SI (2006a) Analysis of aromatic catabolic pathways in Pseudomonas putida KT 2440 using a combined proteomic approach: 2DE/MS and cleavable isotope-coded affinity tag analysis. Proteomics 6:1301–1318
Kim SJ, Kweon O, Freeman JP, Jones RC, Adjei MD, Jhoo JW, Edmondson RD, Cerniglia CE (2006b) Molecular cloning and expression of genes encoding a novel dioxygenase involved in low- and high-molecular-weight polycyclic aromatic hydrocarbon degradation in Mycobacterium vanbaalenii PYR-1. Appl Environ Microbiol 72(2):1045–1054
Kirchhof G, Schloter M, Assmus B, Hartmann A (1997) Molecular microbial ecology approaches applied to diazotrophs associated with non-legumes. Soil Biol Biochem 29:853–862
Kiyohara H, Torigoe S, Kaida N, Asaki T, Iida T, Hayashi H, Takizawa N (1994) Cloning and characterization of a chromosomal gene cluster, pah, that encodes the upper pathway for phenanthrene and naphthalene utilization by Pseudomonas putida OUS82. J Bacteriol 176:2439–2443
Koukkou AI, Drainas C (2008) Addressing PAH biodegradation in Greece: biochemical and molecular approaches. IUBMB Life 60(5):275–280
Krivobok S, Kuony S, Meyer C, Louwagie M, Willison JC, Jouanneau Y (2003) Identification of pyrene-induced proteins in Mycobacterium sp. strain 6PY1: evidence for two ring hydroxylating dioxygenase. J Bacteriol 185:3828–3841
Lafortune I, Juteau P, Déziel E, Lépine F, Beaudet R, Villemur R (2009) Bacterial diversity of a consortium degrading high-molecular-weight polycyclic aromatic hydrocarbons in a two-liquid phase biosystem. Microb Ecol 57:455–468
Laha S, Tansel B, Ussawarujikulchai A (2009) Surfactant–soil interactions during surfactant amended remediation of contaminated soils by hydrophobic organic compounds: a review. J Environ Manag 90:95–100
Larsson B, Sahlberg G (1982) Polycyclic aromatic hydrocarbons in lettuce. Influence of a highway and an aluminium smelter. In: Cooke M, Denis AJ, Fisher GL (eds) Polynuclear aromatic hydrocarbons: physical and biological chemistry. Battelle Press, Colombus, pp 417–426
Lau KL, Tsang YY, Chiu S (2003) Use of spentmushroom compost to bioremediate PAH-contaminated samples. Chemosphere 52:1539–1546
Launen L, Pinto LJ, Wiebe C, Kiehlmann E, Moore MM (1995) The oxidation of pyrene and benzo[a]pyrene by nonbasidiomycete soil fungi. Canad J Microbiol 41:477–488
Lee HS, Lee K (2001) Bioremediation of diesel-contaminated soil by bacterial cells transported by electrokinetics. J Microbiol Biotechnol 11:1038–1045
Legge R (2012) Analysis of microbial diversity by amplicon pyrosequencing. Dissertations and Theses in Food Science and Technology. Paper 25
Li T, Wu TD, Mazéas L, Toffin L, Guerquin-Kern JL, Leblon G, Bouchez T (2008a) Simultaneous analysis of microbial identity and function using NanoSIMS. Environ Microbiol 10:580–588
Li X, Li P, Lin X, Zhang C, Li Q, Gong Z (2008b) Biodegradation of aged polycyclic aromatic hydrocarbons (PAHs) by microbial consortia in soil and slurry phases. J Hazard Mater 150(1):21–26
Liang Y, Gardener D, Miller CD, Chen D, Anderson AJ, Weimer BC, Sims RC (2006) Study of biochemical pathways and enzymes involved in pyrene degradation by Mycobacterium sp. strainKMS KMS. Appl Environ Microbiol 72:7821–7828
Lijinsky W (1991) The formation and occurence of polynucleararo- matic hydrocarbons associated with food. Mutat Res 259:251–262
Liu Y, Zhang J, Zhang Z (2004) Isolation and characterisation of polycyclicaromatic hydrocarbons-degrading Sphingomonas sp. Strain ZL5. Biodegradation 15:205–212
Lopez de Victoria G, Lovell CR (1993) Chemotaxis of azospirillum species to aromatic compounds. Appl Environ Microb 59:2951–2955
Low JYS, Abdullah N, Vikineswary S (2009) Evaluation of support materials for immobilization of Pycnoporus sanguinues mycelia for laccase production and biodegradation of polycyclic aromatic hydrocarbons. Res J Environ Sci 3(3):357–366. ISSN 1819-3412
Majcherczyk A, Johannes C, Huttermann A (1998) Oxidation of polycyclic aromatic hydrocarbons (PAH) by laccase of Trametesversicolor. Enzyme Microbiol Technol 22:335–341
Mallick S, Chatterjee S, Dutta TK (2007) A novel degradation pathway in the assimilation of phenanthrene by Staphylococcus sp. strain PN/Y via meta-cleavage of 2-hydroxy-1-naphthoic acid: formation of trans-2,3-dioxo-5-(2′-hydroxyphenyl)-pent-4-enoic acid. Microbiology 153:2104–2115
Mapelli V, Olsson L, Nielsen J (2009) Metabolicfootprinting in microbiology: methods and applications in functional genomics and biotechnology. Trends Biotechnol 26:490–497
Margesin R, Schinner F (2001) Biodegradation and bioremediation of hydrocarbons in extreme environments. Appl Microbiol Biotechnol 56:650–663
Mastral AM, Callen MS (2000) A review on polycyclic aromatic hydrocarbon (PAH) emissions from energy generation. Environ Sci Technol 34:3051–3057
McKenna EJ, Heath RD (1976) Biodegradation of polynuclear aromatic hydrocarbon pollutants by soil and water microorganisms. University of Illinois (Urbana-Champaign) Research Report no. 113
Mersch-Sundermann V, Mochayedi S, Kevekordes S (1992) Genotoxicity of polycyclic aromatic hydrocarbons in Escherichia coli PQ37. Mutat Res 278:1–9
Mester T, Tien M (2000) Oxidation mechanism of ligninolytic enzymes involved in the degradation of environmental pollutants. Int Biodeterior Biodegrad 46:51–59
Miyata N, Iwahori K, Foght JM, Gray MR (2004) Saturable, energy dependent uptake of phenanthrene in aqueous phase by Mycobacterium sp. strain RJGII-135. Appl Environ Microbiol 70(1):363–369
Molina MC, González N, Bautista LF, Sanz R, Simarro R, Sánchez I, Sanz JL (2009) Isolation and genetic identification of PAH degrading bacteria from a microbial consortium. Biodegradation 20:789–800
Moody J, Freeman J, Doerge D, Cerniglia C (2001) Degradation of phenanthrene and anthracene by cell suspensions of Mycobacterium sp. PYR-1. Appl Environ Microbiol 67(4):1476–1483
Moran MA (2009) Metatranscriptomics: eavesdropping on complex microbial communities. Microbe 4:329–335
Mori T, Kitano S, Kondo R (2003) Biodegradation of chloronaphthalenes and polycyclic aromatic hydrocarbons by the white-rot fungus Phlebialindtneri. Appl Microbiol Biotechnol 61(4):380–383
Mueller JG, Lantz SE, Blattmann BO, Chapman PJ (1991) Bench-scale evaluation of alternative biological treatment process for the remediation of pentachlorophenol and creosote contaminated materials: solid phase bioremediation. Environ Sci Technol 25:1045–1055
Musat F, Galushko A, Jacob J, Widdel F, Kube M, Reinhardt R (2009) Anaerobic degradation of naphthalene and 2-methylnaphthalene by strains of marine sulfate-reducing bacteria. Environ Microbiol 11:209–219
Muyzer G, Smalla K (1998) Application of denaturing gradient gel electrophoresis (DGGE) and temperature gradient gel electrophoresis (TGGE) in microbial ecology. Antonie Van Leeuwenhoek 73:127–141
Muyzer GE, de Waal EC, Uitterlinden AG (1993) Profiling of complex microbial populations by denaturing gradient gel electrophoresis analysis of polymerase chain reaction-amplified genes coding for 16S rRNA. Appl Environ Microbiol 59:695–700. PMID 7683183. www.ncbi.nlm.nih.gov/pubmed/7683183
Nakagaki T, Kobayashi R, Nishiura Y, Ueda T (2004) Obtaining multiple separate food sources: behavioural intelligence in the Physarium plasmodium. Proc R Soc B 271:2305–2310
Nichols NN, Lunde TA, Graden KC, Hallock KA, Kowalchyk CK, Southern RM, Soskin EJ, Ditty JL (2012) Chemotaxis to furan compounds by furan-degrading Pseudomonas strains. Appl Environ Microbiol 78(17):6365. doi:10.1128/AEM.01104-12
Nylund L, Heikkila P, Hameila M, Pyy L, Linnainmaa K, Sorsa M (1992) Genotoxic e€ects and chemical composition of four creosotes. Mutat Res 265:223–236
Ortegocalvo JJ, Marchenko AI, Votobyow AV, Borovick RV (2003) Chemotaxis in polycyclic aromatic hydrocarbon-degrading bacteria isolated from coal-tar and oil polluted rhizospheres. FEMS Microb Ecol 44:373–381
Park KS, Sims RC, Dupont R (1990) Transformations of PAHs in soil systems. J Environ Eng (ASCE) 116:32–640
Peltola R (2010) Bioavailability aspects of hydrophobic contaminant degradation in soils. ISSN 1795-7079. ISBN 978-952-10-4683-4
Phillips DH (1983) Fifty years of benzo(a)pyrene. Nature 303:472–486
Pinyakong O, Habe H, Supaka N, Pinpanichkarn P, Juntongjin K, Yoshida T (2000) Identification of novel metabolites in the degradation of phenanthrene by Sphingomonassp. strain P2. FEMS Microbiol Lett 191:115–121
Poretsky RS, Bano N, Buchan A, LeCleir G, Kleikemper J, Pickering M, Pate WM, Moran MA, Hollibaugh JT (2005) Analysis of microbial gene transcripts in environmental samples. Appl Environ Microbiol 71:4121–4126
Pothuluri JV, Freeman JP, Evans FE, Cerniglia CE (1990) Fungal transformation of fluoranthene. Appl Environ Microbiol 56:2974–2983
Pozdnyakova NN (2012) Involvement of the ligninolytic system of white-rot and litter-decomposing fungi in the degradation of polycyclic aromatic hydrocarbons. Biotechnol Res Int. Article ID 243217, p 20. doi:10.1155/2012/243217
Prabhu Y, Phale PS (2003) Biodegradation of phenanthrene by Pseudomonas sp. strain PP2: novel metabolic pathway, role of biosurfactant and cell surface hydrophobicity in hydrocarbon assimilation. Appl Microbiol Biotechnol 61:342–351
Qiu YL, Sekiguchi Y, Imachi H, Kamagata Y, Tseng IC, Cheng SS, Ohashi A, Harada H (2004) Identification and isolation of anaerobic, syntropic phthalate isomer degrading microbes from methanogenic sludges treating wastewater from terepthalate manufacturing. Appl Environ Microbiol 70:1617–1626
Rahman RNZA, Ghazali FM, Salleh AB, Basri M (2006) Biodegradation of hydrocarbon contamination by immobilized bacterial cells. J Microbiol 44(3):354–359
Rama R, Mougin C, Boyer FD, Kollmann A, Malosse C, Sigoillot JC (1998) Biotransformation of benzo(a)pyrene in bench scale reactor using laccase of Pycnoporus cinnabarinus. Biotechnol Lett 20:1101–1104
Rastogi G, Sani R (2011) Molecular techniques to assess microbial community structure, function, and dynamics in the environment. Microbes Microb Technol Agric Environ Appl. doi:10.1007/978-1-4419-7931-5_2
Renner R (1999) EPA to strengthen persistent, bioaccumulative and toxic pollutant controls—mercury first to be targeted. Environ Sci Technol 33:62
Rockne KJ, Strand SE (1998) Biodegradation of bicyclic and polycyclic aromatic hydrocarbons in anaerobic enrichments. Environ Sci Technol 32:2962–2967
Ruggaber TP, Talley JW (2006) Enhancing bioremediation with enzymatic processes: a review. Pract Period Hazard Toxic Radioact Waste Manag 10:73–85
Sack U, Fritsche W (1997) Enhancement of pyrene mineralization in soil by wood-decaying fungi. FEMS Microbiol Ecol 22(1):77–83
Sack U, Gunther T (1993) Metabolism of PAH by fungi and correlation with extracellular enzymatic activities. J. Basic Microbiol. 33:269–277
Salicis F, Krivobok MJS, Benoit-Guyod JL (1999) Biodegradation of fluoranthene by soil fungi. Chemosphere 38:3031–3039
Schloss PD, Handelsman J (2003) Biotechnological prospects from metagenomics. Curr Opin Microbiol 14:303–310
Schneider J, Grosser R, Jayasimhulu K, Xue W, Warshawsky D (1996) Degradation of pyrene, benz[a]anthracene, and benzo[a]pyrene byMycobacterium sp. strain RJGII-135, isolated from a former coal gasificationsite. Appl Environ Microbiol 62:13–19
Schutzendubel A, Majcherczyk A, Johannes C, Huttermann A (1999) Degradation of fluorene, anthracene, phenanthrene, fluoranthene and pyrene lacks connection to the production of extracellular enzymes by Pleurotusostreatus and Bjerkanderaadjusta. Int Biodeterior Biodegrad 43:93–100
Schwieger F, Tebbe CC (1998) A new approach to utilize PCR-single-strand conformation polymorphism for 16S rRNA gene-based microbial community analysis. Appl Environ Microbiol 64:4870–4876
Semple KT, Cain RB, Schmidt S (1999) Biodegradation of aromatic compounds by microalgae. FEMS Microbiol Lett 170:291–300
Semple KT, Morriss WJ, Paton GI (2003) Bioavailability of hydrophobic organic contaminants in soils: fundamental concepts and techniques for analysis. Eur J Soil Sci 54:809–818
Seo JS, Keum YS, Hu Y, Lee SE, Li QX (2006) Phenanthrene degradation in Arthrobacter sp. P1-1: initial 1,2-, 3,4- and 9,10-dioxygenation, and meta- and ortho-cleavages of naphthalene-1,2-diol after its formation from naphthalene-1,2-dicarboxylic acid and hydroxyl naphthoic acids. Chemosphere 65:2388–2394
Seo JS, Keum YS, Hu Y, Lee SE, Li QX (2007) Degradation of phenanthrene by Burkholderia sp. C3: initial 1,2- and 3,4-dioxygenation and meta- and ortho-cleavages of naphthalene-1,2-diol. Biodegradation 18:123–131
Seo J-S, Keum Y-S, Li QX (2009) Bacterial degradation of aromatic compounds. Int J Environ Res Pub Health 6:278–309. doi:10.3390/ijerph6010278
Sharma PM, Bhattacharya D, Krishnan S, Lal B (2004) Degradation of polycyclic aromatic hydrocarbons by a newly discovered enteric bacterium Leclercia adecarboxylata. Appl Environ Microbiol 70(5):3163–3166
Shi Z, Tian L, Zhang Y (2010) Molecular biology approaches for understanding microbial polycyclic aromatic hydrocarbons (PAHs) degradation. Acta Ecol Sin 30:292–295
Sims RC, Overcash MR (1983) Fate of polynuclear aromatic compounds (PNAs) in soil-plant systems. Residue Rev 88:1–68
Sinha S, Chattopadhyay P, Pan I, Chatterjee S, Chanda P, Bandyopadhya D, Das K, Sen SK (2009) Microbial transformation of xenobiotics for environmental bioremediation. Afr J Biotechnol 8(22):6016–6027
Smith CJ, Osborn AM (2009) Advantages and limitations of quantitative PCR (Q-PCR)-based approaches in microbial ecology. FEMS Microbiol Ecol 67:6–20
Steffen K, Hatakka A, Hofrichter M (2003) Removal and mineralization of polycyclic aromatic hydrocarbons by litter decomposing basidiomycetous fungi. Appl Microbiol Biotechnol 60(1–2):212–217
Stemmer WPC (1994a) DNA shuffling by random fragmentation and reassembly: in vitro recombination for molecular evolution. Proc Natl Acad Sci USA 91:10747–10751
Stemmer WPC (1994b) Rapid evolution of a protein in vitro by DNA shuffling. Nature 370:389–391
Su D, Li PJ, Frank S, Xiong XZ (2006) Biodegradation of benzo[a]pyrene in soil by Mucor sp. SF06 and Bacillus sp. SB02 co-immobilized on vermiculite. J Environ Sci 18(6):1204–1209
Sutherland JB, Selby AL, Freeman JP, Evans FE, Cerniglia CE (1991) Metabolism of phenanthrene by Phanerochaete chrysosporium. Appl Environ Microbiol 57:3310–3316
Sutherland JB, Fu PP, Yang SK, Vontungeln LS, Casillas RP, Crow SA, Cerniglia CE (1993) Enantiomeric composition of the trans-dihydrodiols produced from phenanthrene by fungi. Appl Environ Microbiol 59:2145–2149
Takata N, Sakata M (2002) Effect of photooxidation on delta C-13 of benzo(a)pyrene and benzo(e)pyrene in the atmosphere. Geochem J 36(3):235–245
Tao X-Q, Lu G-N, Liu J-P, Li T, Yang L-N (2009) Rapid degradation of phenanthrene by using Sphingomonas sp. GY2B immobilized in calcium alginate gel beads. Int J Environ Res Publ Health 6:2470–2480
Thies JE (2007) Soil microbial community analysis using terminal restriction fragment length polymorphisms. Soil Sci Soc Am J 71:579–591
Tomotada I, NaSu M (2001) Current bioremediation practice and perspective. J Biosci Bioeng 92(1):1–8
University of Minnesota Biocatalysis/Biodegradation Database (UMBBD) (2004) University of Minnesota. http://umbbd.ahc.umn.edu
Vanrooij JGM, Bodelierbade MM, Jongeneelen FJ (1993) Estimation of individual dermal and respiratory uptake of polycyclic aromatic-hydrocarbons in 12 coke-oven workers. Brit Jn Ind Med 50:623–632
Varkonyi-Gasic E, Hellens RP (2010) qRT-PCR of small RNAs. Methods Mol Biol 631(109):22. doi:10.1007/978-1-60761-646-7_10
Veeken AHM, Hamelers BVM (1999) Effect of substrate-seed mixing and leachate recirculation on solid state digestion of biowaste. In: Mata-Alvarez J, Tilche A, Cecchi F (eds) Proceedings of the second international symposium on anaerobic digestion of solid wastes. Barcelona 1. Gr_A®Ques 92:15-18: 250–257
Verdin A, Sahraoui AL-H, Durand R (2003) Degradation of benzo(a)pyrene by mitosporic fungi and extracellular oxidative enzymes. Int Biodeterior Biodegrad 53:65–70
Voordouw G (1998) Reverse sample genome probing of microbial community dynamics. ASM News 64:627–633
Vyas B, Sasek V, Matucha M (1994) Degradation of anthracene by selected white rot fungi. FEMS Microbiol Ecol 14(1):65–70
Walter U, Beyer M, Klein J, Rehm HJ (1991) Degradation of pyrene by rhodococcus sp. UW1. Appl Microbiol Biotechnol 34:671–676
Wang S, Li X, Liu W, Li P, Kong L, Ren W, Wu H, Tu Y (2012) Degradation of pyrene by immobilized microorganisms in saline-alkaline soil. J Environ Sci 24(9):1662–1669
Wattiau P, Bastiaens L, van Herwijnen R, Daal L, Parsons JR, Renard ME, Springael D, Cornelis GR (2001) Fluorene degradation by Sphingomonas sp. LB126 proceeds through protocatechuic acid: a genetic analysis. Res Microbiol. 52(10):861–872
Weissenfels WD, Beyer M, Klein J, Rehm HJ (1991) Microbial metabolism of fluoranthene: isolation and identification of ring fission products. Appl Microbiol Biotechnol 34:528–535
Widdle F, Rabus R (2001) Anaerobic biodegradation of saturated and aromatic hydrocarbons. Curr Opin Biotechnol 12:259–276
Wilmes P, Bond PL (2006) Metaproteomics: studying functional gene expression in microbial ecosystems. Trends Microbiol 14:92–97
Wilmes P, Wexler M, Bond PL (2008) Metaproteomics provides functional insight into activated sludge wastewater treatment. PLoS ONE 12(e1778):1–11
Wilson NG, Bradley G (1996) The effect of immobilization on rhamnolipid production by Pseudomonas fluorescens. J Appl Bacteriol 81(5):525–530
Wilson SC, Jones KC (1993) Bioremediation of soils contaminated with polynuclear aromatic hydrocarbons (PAHs): a review. Environ Pollut 88(229):249
Wong JWC, Lai KM, Wan CK, Ma KK, Fang M (2002) Isolation and optimisation of PAH-degradative bacteria from contaminated soil for PAH bioremediation. Water Air Soil Pollut 139:1–13
Wood TK (2008) Molecular approaches in bioremediation. Curr Opin Biotechnol 19:572–578
Wu Y, Teng Y, Li Z, Liao X, Luo Y (2008a) Potential role of polycyclic aromatic hydrocarbons (PAHs) oxidation by fungal laccase in the remediation of an aged contaminated soil. Soil Biol Biochem 40:789–796
Wu Y, Luo Y, Zou D, Ni J, Liu W, Teng Y, Li Z (2008b) Bioremediation of polycyclic aromatic hydrocarbons contaminated soil with Moilinia sp.: degradation and microbial community analysis. Biodegradation 19:247–257
Wullings BA, van Beuningen AR, Janse JD, Akkermanns ADL (1998) Detection of Ralstoniasolanacearumwhich causes brown rot of potato, by fluorescent in situ hybridization with 23S rRNA-targeted probes. Appl Environ Microbiol 64:4546–4554
Wunder T, Marr J, Kremer S, Sterner O, Anke H (1997) 1- Methoxypyrene and 1,6-dimethoxypyrene: two novel metabolites in fungal metabolism of polycyclic aromatic hydrocarbons. Arch Microbiol 167:310–316
Xie S, Liu J, Li L, Qiao C (2009) Biodegradation of malathion by Acinetobacter johnsonii MA19 and optimization of cometabolism substrates. J Environ Sci 21(1):2176–2182
Xu YH, Lu M (2010) Bioremediation of crude oil-contaminated soil: comparison of different biostimulation and bioaugmentation treatments. J Hazard Mater 183(1–3):395–401
Yamazoe A, Yagi O, Oyaizu H (2004) Degradation of polycyclic aromatic hydrocarbon by a newly isolated dibenzofuran utilizing Janibacter sp strain yy 1. Appl microbial biot 65:211–218
Yuan SY, Wei SH, Chang BV (2000) Biodegradation of polycyclic aromatic hydrocarbons by a mixed culture. Chemosphere 41:1463–1468
Zhang C, Bennet GN (2005) Biodegradation of xenobiotics by anaerobic bacteria. Appl Microbiol Biotechnol 67:600–618
Zhang Y, Zhu YX, Kwon KK, Park JH, Kim SJ (2002) Detection of biodegradation of pyrene by synchronous fluorometry. China Environ Sci 22:289–292
Zhang H, Kallimanis A, Koukkou AI, Drainas C (2004a) Isolation and characterization of novel bacteria degrading polycyclic aromatic hydrocarbons from polluted Greek soils. Appl Microbiol Biotechnol 65:124–131
Zhang W, Wang H, Zhang R, Yu XZ, Qian PY, Wong MH (2004b) Bacterial communities in PAH contaminated soils at an electronic-waste processing center in China. Ecotoxicology 19:96–104
Zhong Y, Luan T, Wang X, Lan C, Tam NF (2007) Influence of growth medium on cometabolic degradation of polycyclic aromatic hydrocarbons by Sphingomonas sp. strain PheB4. Appl Microbiol Biotechnol 75(1):175–186
Zhou J (2003) Microarrays for bacterial detection and microbial community analysis. Curr Opin Microbiol 6:288–294
Zhou HW, Wong AHY, Yu RMK, Park YD, Wong YS, Tam NFY (2009) Polycyclic aromatic hydrocarbon-induced structural shift of bacterial communities in mangrove sediment. Microb Ecol 58:153–160
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Gupta, S., Pathak, B. & Fulekar, M.H. Molecular approaches for biodegradation of polycyclic aromatic hydrocarbon compounds: a review. Rev Environ Sci Biotechnol 14, 241–269 (2015). https://doi.org/10.1007/s11157-014-9353-3
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11157-014-9353-3