Abstract
Reactive oxygen species (ROS) production is an unavoidable byproduct of electron transport under aerobic conditions. Photosystem II (PS II), the cytochrome b6/f complex and Photosystem I (PS I) are all demonstrated sources of ROS. It has been proposed that PS I produces substantial levels of a variety of ROS including O .−2 , 1O2, H2O2 and, possibly, •OH; however, the site(s) of ROS production within PS I has been the subject of significant debate. We hypothesize that amino acid residues close to the sites of ROS generation will be more susceptible to oxidative modification than distant residues. In this study, we have identified oxidized amino acid residues in spinach PS I which was isolated from field-grown spinach. The modified residues were identified by high-resolution tandem mass spectrometry. As expected, many of the modified residues lie on the surface of the complex. However, a well-defined group of oxidized residues, both buried and surface-exposed, lead from the chl a’ of P700 to the surface of PS I. These residues (PsaB: 609F, 611E, 617M, 619W, 620L, and PsaF: 139L, 142A,143D) may identify a preferred route for ROS, probably 1O2, to egress the complex from the vicinity of P700. Additionally, two buried residues located in close proximity to A1B (PsaB:712H and 714S) were modified, which appears consistent with A1B being a source of O .−2 . Surprisingly, no oxidatively modified residues were identified in close proximity to the 4Fe–FS clusters FX, FA or FB. These cofactors had been identified as principal targets for ROS damage in the photosystem. Finally, a large number of residues located in the hydrophobic cores of Lhca1–Lhca4 are oxidatively modified. These appear to be the result of 1O2 production by the distal antennae for the photosystem.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In higher plants, the Photosystem I—Light-Harvesting Complex I supercomplex (PS I-LHC I) acts as a light-energy driven plastocyanin—ferredoxin oxidoreductase. Crystal structures are available for PS I-LHC I from red algae (Cyanidioschyzon merolae strain 10D, 4.0 Å (Antoshvili et al. 2019; Pi et al. 2018), green algae (Chlamydomonas reinhardtii, 2.9 Å, (Suga et al. 2019)) and angiosperms (Pisum sativum, 2.8 Å (Mazor et al. 2017; Qin et al. 2015).
Higher plant PS I-LHC I is a 650 kDa monomeric complex consisting of at least 16 subunits (PS I: 12 core subunits (PsaA–PsaO) and LHC I: 4 subunits (Lhca1-Lhca4)). Numerous prosthetic groups are associated with these proteins including 156 chlorophylls, 37 carotenoids, 2 phylloquinones, and 3 iron–sulfur clusters (4Fe–4S) (Caspy and Nelson 2018). During in vivo linear electron transport, an initial light-induced charge separation occurs between P700 (a chl a•chl a’ dimer) and A0, forming P700+ and A0−, followed by sequential electron transfer from A0− to A1 (a phylloquinone), to the iron–sulfur clusters FX and FA/FB. Subsequently, oxidized ferredoxin is reduced on the stromal side of the thylakoid membrane and P700+ is reduced by plastocyanin on the lumenal side of the membrane. It has been shown that electron transfer from P700 to FX in PS I-LHC I can occur down both pseudosymmetrical cofactor branches associated with the PsaA/PsaB core dimer (Guergova-Kuras et al. 2001).
The production of reactive oxygen species (ROS) is unavoidable, being a consequence of electron transport under oxic conditions being produced by both respiratory electron transport membrane protein complexes and those of oxygenic photosynthesis. ROS are generated by all of the photosynthetic membrane protein complexes (PS II, the b6/f complex, PS I, and, probably, the chloroplast NAD(P) dehydrogenase-like complex) and these ROS species can oxidatively damage proteins and other biomolecules (Das and Roychoudhury 2014). ROS are formed by the excitation of molecular oxygen (singlet oxygen, 1O2), the partial reduction of molecular oxygen (O .−2 ), and the partial oxidation of water (H2O2 and •OH). Additionally, dismutation of O .−2 can form H2O2 that subsequently may be converted to •OH via Fenton chemistry. It has been estimated that ROS damage to PS II, alone, could account for a 10% loss in photosynthetic productivity (Long et al. 1994). While ROS species are typically thought to be agents of damage, it should be noted that ROS also serve as important signaling molecules (Foyer and Noctor 2013; Mignolet-Spruyt et al. 2016).
ROS generation by PS II has been extensively studied (Kale et al. 2017; Pospíšil 2009, 2016), however, far fewer studies have been performed on PS I (Sonoike 2011). Under most laboratory conditions (constant moderate temperature and constant, relatively low intensity illumination), PS I appears to be significantly less susceptible to photodamage than PS II (Sonoike 2011). However, in the field environment, plants are exposed to a variety of abiotic stressors which can lead to enhanced ROS production (drought, low or high temperatures, high light intensities, nutrient limitations, etc.) (Choudhry et al. 2016; You and Chan 2015). Under such field conditions, PS I can undergo significant photoinhibition (Sejima et al. 2014; Sonoike et al. 1995; Terashima et al. 1994; Zivcak et al. 2015).
In our study, we have used high-resolution tandem mass spectrometry to identify the location of oxidized residues within PS I-LHC I supercomplex isolated from field-grown spinach. Earlier, we have used these methods to identify natively oxidized residues in spinach PS II (Frankel et al. 2012, 2013), results which have been largely verified and extended for cyanobacterial PS II (Weisz et al. 2017). We have also used these methods to examine natively oxidized residues in the cytochrome b6/f complex (Taylor et al. 2018).
In this communication, we have mapped the identified natively oxidized residues in the spinach PSI-LHCI supercomplex and mapped these onto the corresponding residues of the Pisum sativum PSI-LHCI supercomplex structure (Mazor et al. 2017). We observed many oxidized residues on the surface of the complex. This is expected since ROS produced by any source, either on the stromal or lumenal sided of the thylakoid, could potentially modify surface residues. Additionally, a group of largely buried oxidized residues leads from the chl a’ of the P700 special pair to the surface of the complex. Two buried oxidized residues were also in relatively close proximity to A1B. These findings support the hypothesis that the P700 and, possibly, A1B are sources of ROS production by the PS I-LHC I supercomplex, probably 1O2 and O .−2 , respectively. Finally, a large number of buried residues located in the hydrophobic cores of Lhca1-Lhca4, the distal antenna for PS I, are oxidatively modified. These appear to be the result of 1O2 production by the distal antennae for the photosystem. Interestingly, no oxidatively modified residues were identified in close proximity to the 4Fe–4S clusters FX, FA, or FB. These metal centers have been proposed to be either a source of O .−2 or a target for O .−2 damage (Asada 1999; Caffarri et al. 2014; Tjus et al. 1999).
Materials and methods
PS I-LHC I-LHC II-enriched membranes were prepared essentially as described previously (Bell et al. 2015). The PS I-LHC I-LHC II subunits were resolved by LiDS-PAGE (Delepelaire and Chua 1979), using a non-oxidizing gel system (Rabilloud et al. 1995). It is known that standard SDS-PAGE can introduce numerous protein oxidation artifacts (Sun and Anderson 2004) due to the generation of •OH during the polymerization process. In this system, the acrylamide solution is degassed and the gels are polymerized with riboflavin (in the presence of diphenyliodonium chloride and toluensulfinate) with overnight exposure to long-wavelength UV light. During electrophoresis, the upper reservoir buffer contained thioglycolate. It has been demonstrated that proteins resolved in this system exhibit significantly lower levels of electrophoresis-induced protein oxidation than proteins resolved by standard PAGE using standard polymerization conditions, confirming earlier reports (Frankel et al. 2012; Rabilloud et al. 1995; Sun and Anderson 2004). Electrophoresis was stopped when the proteins entered the resolving gel ~ 1.0 cm. The gel was then stained with Coomassie blue, destained, and two protein bands that contained (1) principally PsaA/PsaB and (2) the low molecular weight PS I core subunits and LHCs were excised and analyzed separately. These were digested using either trypsin or chymotrypsin using standard procedures for “in-gel” proteolysis. After protease digestion and peptide isolation, the peptides were resolved by HPLC on a C:18 reversed-phase column and ionized via electrospray into a Thermo Scientific Orbitrap Fusion Lumos mass spectrometer. The samples were analyzed in a data-dependent mode with one Orbitrap MS1 scan acquired simultaneously with up to ten linear ion trap MS2 scans. The MassMatrix Program (Xu and Freitas 2009) was used for protein identification and the identification of peptides containing oxidative modifications. A FASTA library containing the sequences of 19 proposed subunits of PS I-LHC I of the spinach supercomplex was searched, as was a decoy library containing the reversed amino acid sequences of the 19 proposed subunits. If during a search of these libraries any putative peptides for a particular protein were identified from the decoy library (i.e., decoy hit% > 0.00%), the protein was excluded from further analysis. It should be noted that for standard proteomic analysis in MassMatrix, a decoy hit rate of up to 10% is tolerated. Twelve different oxidative modifications were considered during peptide screening with a maximum of two modifications allowed per peptide. For a putative positive identification of an oxidized residue by MassMatrix, the peptide containing the modification must exhibit a pp-tag value of 10−5 or smaller and either pp or pp2 must exhibit a value of 10−5 or smaller (Bricker et al. 2015; Xu and Freitas 2009). Peptides meeting these extremely rigorous p value thresholds were then examined manually and the quality of the MS2, collision-induced dissociation spectra, were confirmed. It should be noted that typically, during peptide identification by mass spectrometry for proteomic experiments, p values of 5 × 10−2 are typically used (Diz et al. 2011). We feel that this is inadequate when identifying post-translational modifications (Bricker et al. 2015). Additionally, only peptides with charge states of +3 or lower were considered. Finally, the mass error of the precursor ion was required to be ≤ 5.0 ppm and identified as the product of specific proteolytic cleavage. The identified oxidized amino acid residues were mapped onto the crystal structure of the Pisum sativum PS I-LHC I supercomplex (PDB: 5L8R, (Mazor et al. 2017)) using PYMOL (DeLano 2002).
Results and discussion
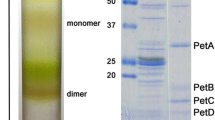
The isolation of the spinach PS I-LHC I-LHC II membranes was performed as previously described and the protein composition of these membranes (Fig. 1) was basically indistinguishable from our previous report (Bell et al. 2015). The PS I-LHC I-LHC II membranes were highly enriched in PS I core components and various light-harvesting chlorophyll proteins (Lhcas and Lhcbs) and were highly depleted of PS II core components. We had earlier shown that these membranes were also depleted of cytochrome b6/f and CF1–CFo subunits (Bell et al. 2015). Additionally, we had demonstrated that the LHC II associated with PS I was functionally coupled to the photosystem (Bos et al. 2017; Bell et al. 2015). All available evidence indicates that these membranes contain large quantities of PS I-LHC I supercomplexes that are functionally coupled to LHC II trimers.
LiDS-PAGE of Thylakoids, PSI-LHCI-LHCII membranes, and PS II membranes. Individual lanes were loaded with 4 µg chl. The PSI-LHCI-LHCII membranes are highly enriched in PSI core proteins, contain significant amounts of LHCs (both Lhcas and Lhcbs), and are highly depleted of PS II components (BBY) as previously reported (Bell et al. 2015). THY, thylakoids; PSI, PSI-LHCI-LHCII membranes; BBY, PS II membranes. Standard proteins (SP) are shown to the left
Tandem mass spectrometry analysis of the tryptic and chymotryptic peptides of the PS I-LHC I supercomplex allowed the identification of 166 oxidatively modified residues present on the subunits of the supercomplex. In this study, over 200,000 peptides were examined and screened for the presence of oxidative modifications. The mass spectrometry sequence coverage and the number of oxidatively modified residues observed for each PS I-LHC I subunit are shown in Table 1. One should note that most of the subunits of the PS I-LHC I supercomplex are intrinsic membrane subunits. Intrinsic membrane proteins are often difficult to examine by high-resolution tandem mass spectrometry, with typically only low sequence coverage being reported in standard experiments (Kar et al. 2017; Souda et al. 2011; Weisz et al. 2017). In our experiments, high sequence coverage was obtained for most of the subunits of the supercomplex, ranging from 100% for PsaA, PsaJ, and Lhca4 to 56% for PsaG. The use of chymotrypsin greatly expanded the portfolio of observed peptides and dramatically increases the sequence coverage since the chymotryptic peptides are, in large measure, complementary in sequence to the tryptic peptides. Peptides were detected for PsaI, PsaK, PsaO, and PsaP; however, the protein identification based on these peptides exhibited a significant number of hits to the decoy library. Since we set very rigorous criteria for protein identifications (i.e., 0.00% hits to the decoy library were allowed, see Materials and Methods), these proteins were excluded from subsequent analysis. No peptides were observed for PsaN. The number of oxidized amino acid residues observed was quite variable, ranging from 42 for PsaB (5.7% of the PsaB residues being modified) to 0 for PsaG (0% of the PsaG residues being modified), indicating that some supercomplex subunits are significantly more susceptible to oxidative modification than others.
Figure 2 shows the quality of the data used for the identification of oxidized amino acid residues within the PS I–LHC I supercomplex. The tandem mass spectrometry data collected for the 303D-314R tryptic peptide of PsaB are illustrated. This peptide bears a general oxidation modification (mass change +16) at 308M. The observed mass accuracy for the parent ion was 2.6 ppm. The pp, pp2, and pptag values for this peptide were 10−6.3, 10−6.3, and 10−5.5 (Xu and Freitas 2009), respectively, and is consequently among the lowest quality peptides used in this study (Bricker et al. 2015). Even this peptide, however, exhibited nearly complete y- and b-ion series, allowing unequivocal identification and localization of the oxidative mass modification. This result indicates that the use of p values ≤ 10−5 provided high-quality peptide identifications for the localization of post-translational modifications.
Quality of the Mass Spectrometry. Shown is the mass spectrometry result for the tryptic peptide PsaB: 303D-314R, which contains a modified 308H + go (+16 amu) modification. Top, spectrum of the CID dissociation of the unmodified peptide PsaB:303DLLEAH (+16)IPPGG314R. Selected identified ions are labeled. Bottom, table of all predicted masses for the y- and b-ions generated from this peptide sequence. Ions identified in the CID spectrum (top) are shown in red. The b’++, b’+ y’++, and y’+ ions are generated by the neutral loss of water while the b*++, b*+ y*++, and y*+ ions are generated from the loss of ammonia. For our analysis, only the yX and bX ions were considered. The ions y7+-y9+ and y10++-y12++exhibit the +16 amu mass modification as does the b6+-b7+ and b10++-b11++ ion. This verifies that 308H contains an oxidative modification which adds 16 amu to the histidyl residue. The modified b6+ and y7+ histidyl residue is highlighted in magenta. The p values for this peptide were pp = 10−6.3, pp2 = 10−6.3, and pptag = 10−5.5
The identity of these oxidized residues and the types of modifications observed are presented in Table 2. It should be noted that it is extremely unlikely that all of the observed modifications would be present on every copy of the complex. ROS modification of a particular amino acid residue is a stochastic rather than mechanistic process. The probability that a residue will be modified depends on the ROS type, the susceptibility of a residue to a particular ROS type, the residency time of the ROS in proximity to the residue, and the lifetime of the ROS species. No quantification of the frequency of modification of individual residues was attempted. Indeed, such quantification is quite problematic since peptides containing oxidative modifications are more polar and, consequently, more readily resolved by reversed-phase HPLC and more easily fragmented during CID, making quantitative comparison to the unmodified, more hydrophobic, parent peptide difficult.
Our working hypothesis is that residues located near a site of ROS production (or along an ROS exit pathway) have a higher probability of oxidative modification than those distant from an ROS production site (or not along an ROS exit pathway). Consequently, these detectable modifications are present within the full population of PS I-LHC I complexes present in our biological samples. The supercomplex was isolated from field-grown market spinach, consequently, the exact growth parameters are unknown. What is virtually certain, however, is that the plant material used in our study has been exposed to biologically relevant stress conditions (transient drought, low/high temperatures, high light intensities, fluctuating light intensity, nutrient limitations, etc.) typical of plants grown in the field (Choudhry et al. 2016; You and Chan 2015).
No high-resolution structure is available for the spinach PS I-LHC I supercomplex. However, structures are available for a variety of other eukaryotic organisms: the red alga Cyanidioschyzon merolae, (Antoshvili et al. 2019); the green alga Chlamydomonas reinhardtii, (Su et al. 2019; Suga et al. 2019); the moss Physcomitrella patens (Iwai et al. 2018); and the angiosperm Pisum sativum (PDB: 5L8R, (Mazor et al. 2017)). The sequence identity between the spinach subunits and the Pisum subunits is quite high, ranging from 98% for PsaC and PsaD to 80% for PsaF (Table 1). This high degree of identity allowed us to map the oxidized amino acids that we observed in the spinach supercomplex onto the crystal structure of the Pisum supercomplex, with 94% of the observed oxidatively modified residues being identical in both spinach and Pisum.
Figure 3 presents a broad overview of the location of the oxidized residues that we identified within the context of the PS I-LHC I supercomplex. 49% of the oxidized residues were surface-localized. This was not unexpected, as surface domains are particularly susceptible to oxidative modification. ROS produced by a variety of different sources including PS II (•OH, H2O2, 1O2 and O .−2 ), the cytochrome b6f complex (O .−2 and 1O2), and PS I, itself (1O2, O .−2 and, indirectly, H2O2 and •OH) are released to the stromal and lumenal compartments and may contribute to the oxidative modification of surface-exposed domains of the PS I-LHC I supercomplex, the exposed domains of other membrane protein complexes, and soluble proteins. Many ROS detoxification systems are present in both the stromal (Das and Roychoudhury 2014; Tripathy and Oelmuller 2012) and lumenal compartments (Bermudez et al. 2012; Levesque-Tremblay et al. 2009). However, it is highly unlikely, given the possibly large amounts of ROS produced during photosynthetic electron transport, that these would offer complete protection. Additionally, the ROS detoxification systems would be ineffective against any ROS produced within the protein matrix (at least until the ROS was released to the bulk solvent). Indeed, we observe numerous oxidized residues (51%) buried within the protein matrix. We hypothesized that a subset of these modified residues would be in close proximity to the site(s) of ROS production within the photosystem,
Overview of Natively Oxidized Amino Acid Residues in the Spinach PS I—LHC I Supercomplex. A Illustration of the entire supercomplex as viewed from the stroma (top), as viewed within the plane of the membrane (middle), and as viewed from the lumen (bottom). B Exploded view of the supercomplex as viewed within the membrane plane. Individual subunits are color-coded and labeled in B. Oxidatively modified residues are shown as spheres and are colored in darker shades for each subunit. The observed oxidatively modified residues in spinach were mapped onto their corresponding locations on the Pisum sativum PS I—LHC I supercomplex structure (PDB: 5L8R, (Mazor et al. 2017))
1O2 has been implicated in PS I photoinhibition in PS I submembrane preparations (Rajagopal et al. 2005), in vitro, and under repetitive light pulse conditions, in vivo (Takagi et al. 2016). Within the photosystem, several cofactors have been proposed to be the sources of this ROS. These include 3P700 (Rutherford et al. 2012) and the antennae chls (Alboresi et al. 2009; Cazzaniga et al. 2012; Krieger-Liszkay 2004) as possible sources of 1O2. It has also been shown that O .−2 contributed to PS I photoinhibition under both chilling (Sonoike and Terashima 1994; Sonoike et al. 1995; Terashima et al. 1994) and repetitive light pulse conditions (Takagi et al. 2016). Additionally, dismutation of O .−2 by superoxide dismutase leads to the formation of H2O2 which, in the presence of reduced iron–sulfur clusters or free iron, may generate the highly reactive •OH (Sonoike et al. 1995; Takahashi and Asada 1988) via Fenton chemistry. The sources of this ROS are under some debate. The phylloquinone A1B (Takagi et al. 2016) and the iron–sulfur clusters FX, FA, and FB have been suggested as a possible source of O .−2 (Asada 1999; Tjus et al. 1999).
The core subunits of PS I, PsaA, PsaB, and PsaC contain all of the redox-active cofactors of the photosystem (Srinivasan and Golbeck 2009). These include P700, the primary electron donor, which is a chl a•chl a’ dimer (the chl a is bound to PsaB and the chl a’ is bound to PsaA); AA and AB, two “accessory” chl a bound to PsaA and PsaB, respectively; A0A and A0B, chl a monomers bound to PsaA and PsaB, respectively, which serve as primary electron acceptors from P700; A1A and A1B, phylloquinones bound to PsaA and PsaB, respectively; FX, a 4Fe–4S iron–sulfur cluster bound to both PsaA and PsaB; and FA and FB, both 4Fe–4S iron–sulfur clusters bound to PsaC. These cofactors are arranged in two pseudosymmetric branches along the PsaA/PsaB axis with electron transfer occurring down either branch (Guergova-Kuras et al. 2001). A number of oxidatively modified residues are present closely associated with these cofactors, as illustrated in Figs. 4 and 5. It should also be noted that PsaB exhibited a greater degree of oxidative modification than PsaA. 5.7% of the PsaB residues were modified, whereas 4.2% of PsaA residues were modified (Table 1). This greater degree of oxidative modification of PsaB is fully consistent with earlier reports that PsaB appears more susceptible to oxidative damage, leading to its preferential loss during the PS I photoinhibition timecourse (Sonoike 1995, 1996; Sonoike et al. 1997).
Details of the Oxidatively Modified Residues Identified in the Vicinity of the Cofactors Bound to PsaA and PsaB. A View within the plane of the membrane. B View from the lumenal face of the membrane. Cofactors are shown as sticks and are individually labeled. Oxidatively modified residues are shown as spheres. The residues PsaB: 609F, 617M, 619W, 620L, 712H, and 714S are buried and shown in blue. The residues PsaB:611E and PsaF: 139L, 142A, and 143D are surface-exposed and shown in black. These residues form a near continuous path of modified residues leading from the chl a’ of P700 to the surface of the supercomplex. Please note that no PsaA residues were observed to be oxidatively modified in the vicinity of these cofactors. Additionally, the residue PsaB: 712H (and 714S) is in relatively close proximity to A0B (8.2 Å), A1B (7.0 Å) and in close proximity to chl a 839 (4.6 Å), which is the closest chl to A1B
Details of the Oxidatively Modified Residues Identified in PsaC, PsaD, and PsaE. A View within the plane of the membrane. B View from the lumenal face of the membrane. The three iron–sulfur clusters, FX, FA, and FB, are shown as spheres with FX being included for orientation purposes. Oxidatively modified residues are shown as spheres. The residues PsaD: 116M, 117P, 124M, and 192S and PsaE:116Y are buried and shown as orange spheres and green spheres, respectively. The residues PsaC: 71H, 72E, and 75R; PsaD:104T, 105W, 125R, and 190M; and PsaE:119D are all surface-exposed residues and are shown as black spheres. PsaC: 75R is relatively close to FA (8.0 Å). All of the other oxidized residues present in these three subunits are > 10 Å from the iron–sulfur clusters
Figure 4A, B illustrates the oxidized residues identified to be in close proximity to the redox-active cofactors associated with PsaA and PsaB. No oxidized residues were found in close proximity to FX; the nearest, PsaB:547M, is 10.6 Å distant. PsaB:712H and 714S are in relatively close proximity to A1B, with 712H (5.9 Å from A1B) being coordinated to chl a 839, which spatially is the closest chl to A1B. While this observation is consistent with the hypothesis that A1B is a source of ROS, it should be noted that the chlorin ring of chl a 839 lies between A1B and these oxidized residues. This complicates the interpretation, as the ROS responsible for the modification of PsaB:712H and 714S could have been generated at A1B, chl a 839, or other location.
Most interestingly, a chain of eight oxidized residues (PsaB:620L, 617M, 619W, 609F, and 611E and PsaF: 143D, 142A, 139L) lead from the chl a’ of P700 to the lumenal surface of the complex. No similar distribution of oxidized residues was identified associated with the chl a of P700. This does not appear to be due to sampling error since 100% mass spectrometry coverage was obtained for both PsaA and PsaB residues in this region. This chain of oxidized residues may define a preferred ROS exit pathway. Several studies have suggested that the triplet state of P700 (3P700) is preferentially localized to the chl a’ (Breton et al. 1999; Li et al. 2004; Rutherford and Setif 1990). Importantly, the nearest carotenoid to the chl a’ is nearly 11 Å distant and it is unlikley to efficiently quench 3P700. The interaction of 3P700 with dioxygen may lead to the formation of 1O2, and this may be a source of this ROS in the photosystem. Consequently, within the core of PS I, the production of 1O2 may center on the chl a’ of P700. The presence of a defined set of oxidized residues leading away from the chl a’ to the surface of the complex strongly supports this hypothesis.
Figure 5 illustrates the locations of oxidatively modified amino acid residues in the vicinity of FX, FA, and FB. Interestingly, no residues were found in the immediate vicinity of any of these iron–sulfur clusters. Specifically, the nearest modified residue to FX was PsaB:547M (10.6 Å), the nearest modified residue to FB was PsaD:192S (11.6 Å), and the nearest modified residue to FA was PsaC:75R (8.0 Å). The absence of proximal oxidative modifications associated with these cofactors is intriguing since the iron–sulfur clusters had been implicated in the formation of O .−2 and possibly •OH from H2O2 (Sonoike et al. 1995; Takahashi and Asada 1988). Damage to the 4Fe-4S clusters is also recognized as an early event during PS I photoinhibition (Sonoike et al. 1995; Takahashi and Asada 1988; Tjus et al. 1998). While FX is buried within the PsaA/PsaB protein matrix and would be expected to have only limited accessibility to either O .−2 or H2O2, FB is much more exposed, being 3–4 Å from the surface of PsaC. Interestingly, the PsaD and PsaE components exhibited a significant number of oxidative modifications. While the majority of these were localized on the surface of the complex, several were buried and proximally located near PsaC (PsaD: 192S, 116M, 124M; PsaE:116Y). Given the close proximity of PsaD and PsaE to PsaC, a working hypothesis is that ROS produced at the redox-active center(s) FB, and possibly FA, lead to the oxidative modifications observed on PsaD and PsaE. So the question arises, “why are no oxidative modifications observed in close proximity to FB or FA in the PsaC subunit?” Oxidative modifications may accumulate in the majority of PS I subunits with relatively limited effects on PS I function. If, however, oxidative damage occurs to FB or FA (or FX), electron flow to ferredoxin would be abolished, possibly triggering turnover and replacement of the entire PS I complex. This is the generally accepted model for PS I photoinhibition. Interestingly, it has been shown that under in vivo photoinhibitory conditions, PsaC was lost much more rapidly than other PS I core components in Brassica (Jiao et al. 2004), presumably due to damage of FA and FB (Sonoike 1996; Sonoike et al. 1995). In this study, plants were grown under conditions similar to what the plants would experience in the field. A similar result was observed in vitro in spinach (Yu et al. 2000), with PsaC being among the most labile subunits during a photoinhibition timecourse (along with Lhca1–Lhca4, see below). This raises the intriguing possibility that under some conditions, damage to FA or FB leads to a more selective loss (and possibly replacement of PsaC). It should be noted that, in this hypothetical scenario, loss of PsaC would also lead to the release of PsaD and PsaE since PsaC is required for the binding of these subunits (at least in cyanobacteria (Yu et al. 1995)). If newly synthesized PsaC (or possibly PsaC bearing repaired 4Fe-4S clusters (Djaman et al. 2004)) was incorporated into PS I, released PsaD and PsaE, possibly bearing oxidized modifications, could be re-incorporated into the photosystem.
Figure 6 illustrates the locations of oxidative modifications in the Lhca1-Lhca4 subunits. These components serve as a distal antenna for PS I, delivering excitation energy to the core antenna which is associated with the PsaA and PsaB subunits. All four of the Lhcas exhibit significant oxidative modification. While many of the oxidatively modified residues are surface-exposed, many are buried near the hydrophobic cores of these subunits, along the helix A and B axis (Fig. 6A). In these locations, the modified residues are in very close proximity to the light-harvesting chls and carotenoids (Fig. 6B).
Modified residues in the Lhca1-Lhca4 distal antenna viewed within the plane of the membrane. A Without cofactors shown. B With cofactors shown. In (A) individual Lhca subunits are labeled, as are the stromal and lumenal sides of the membrane complex. Modified residues are shown as spheres, with colored residues being buried and black residues being surface-exposed. Cofactors are shown in stick representation with chl (both chl a and chl b) shown in pale green and the various carotenoids, β-carotene, lutein, and violaxanthin, shown in pale yellow
Earlier studies have indicated that the Lhcas are rapidly lost during PS I photoinhibition (Alboresi et al. 2009; Yu et al. 2000). The Lhca3 and Lhca4 contain “red” chl forms (Croce and van Amerongen 2013; Croce et al. 1996; Melkozernov and Blankenship 2005) with absorption bands of longer wavelengths than exhibited by P700. Excitation energy is concentrated in these low-energy sinks prior to transfer to the proximal antenna and the reaction center. This slows excitation energy transfer; consequently there is a higher probability of intersystem crossing and formation of 3chl, and subsequently 1O2, at these sites. It has been suggested that the Lhca antenna acts as a “fuse” for the photosystem (Alboresi et al. 2009). Under conditions which promote PS I photoinhibition (high light intensity and acceptor-side electron transport limitation), the Lhca antenna chlorophylls are degraded in parallel with the Lhca proteins, while the reaction center retains photochemical activity (Alboresi et al. 2009). In this scenario, any 3chl which is formed is directly quenched by a proximal carotenoid; if this fails and 1O2 is generated, then this is quenched either physically or chemically by the carotenoid, if this fails 1O2 either escapes the Lhca or oxidatively damages this component (either the Lhca protein or its chl cofactors), perhaps leading to Lhca turnover. This would not have an effect on PS I mechanistically, but would directly lead to a smaller optical cross-section for the photosystem. Ultimately, this would offer some degree of protection for the photosystem from a high photon flux and/or an acceptor-side limitation. Our observation that the Lhca subunits exhibit significant oxidative modification strongly supports the hypothesis that these components are a source of 1O2 in the PS I–LHC II supercomplex.
Conclusions
In this communication, we have identified oxidized residues in the PS I- LHC I supercomplex. The locations of these support our hypothesis that residues in the vicinity of ROS production sites would be particularly prone to oxidative modification. A clearly defined group of oxidatively modified residues appears to lead from the chl a’ of P700 to the surface of the complex. This result supports the hypothesis that 3P700, which is localized to the chl a’, participates in the formation of 1O2. Two oxidized residues are located in proximity to A1B (PsaB:712H and 714S). This finding is consistent with the hypothesis that A1B may be a source of ROS, probably O .−2 . Additionally, numerous oxidized residues are located in Lhca1–Lhca4. Many of these are buried in the hydrophobic domains of these subunits and are in close proximity to the chls and carotenoid cofactors. This observation is consistent with the hypothesis that the Lhcas are a source of 1O2. Finally, no oxidized residues were observed in close proximity to the iron–sulfur clusters FX, FA, or FB. This observation indicates that oxidative damage associated with these cofactors, which would interrupt electron flow to ferredoxin, may trigger turnover of the entire PS I-LHC I supercomplex or, possibly a repair pathway leading to the insertion of undamaged or repaired PsaC. We have not, at this time, determined the relative importance of the various possible sites of ROS production for photoinhibition of PS I, nor the time course for the appearance of oxidative modifications during the photoinhibition process. These important questions are the subject of future studies.
References
Alboresi A, Ballottari B, Hienerwadel R, Giacometti GM, Morosinotto T (2009) Antenna complexes protect Photosystem I from photoinhibition. BMC Plant Biol 9:71
Antoshvili M, Caspy I, Hippler M, Nelson N (2019) Structure and function of Photosystem I in Cyanidioschyzon merolae. Photosyn Res 139:499–508
Asada K (1999) The water-cycle in chloroplasts: scavenging of active oxygens and dissipation of excess photons. Annu Rev Plant Physiol Plant Mol Biol 50:601–639
Bell AJ, Frankel LK, Bricker TM (2015) High yield non-detergent Isolation of Photosystem I-Light harvesting Chlorophyll II membranes from spinach thylakoids. J Biol Chem 290:18429–18437
Bermudez MA, Galmes J, Moreno I, Mullineaux PM, Gotor C, Romero LC (2012) Photosynthetic adaptation to length of day is dependent on S-sulfocysteine synthase activity in the thylakoid lumen. Plant Physiol 160:274–288
Bos I et al (2017) Multiple LHCII antennae can transfer energy efficiently to a single Photosystem I. Biochim Biophys Acta 1858:371–378
Breton J, Nabedryk E, Leibl W (1999) FTIR study of the primary electron donor of photosystem I (P700) revealing delocalization of the charge in P700+ and localization of the triplet character in 3P700. Biochemistry 38:11585–11592
Bricker TM, Mummadisetti MP, Frankel LK (2015) Recent advances in the use of mass spectrometry to examine structure/function relationships in Photosystem II. J Photochem Photobiol B 152:227–246. https://doi.org/10.1016/j.jphotobiol.2015.08.031
Caffarri S, Tibiletti T, Jennings RC, Santabarbara S (2014) A Comparison between plant Photosystem I and Photosystem II architecture and functioning. Curr Prot Pep Sci 14:296–331
Caspy I, Nelson N (2018) Structure of plant Photosystem I. Biochem Soc Trans 46:285–294
Cazzaniga S, Li Z, Niyogi KK, Bassi R, Dall’Osto L (2012) The Arabidopsis szl1 mutant reveals a critical role of β-carotene in photosystem I photoprotection. Plant Physiol 159:1745–1758
Choudhry FK, Rivero RM, Blumwald E, Mittler R (2016) Reactive oxygen species, abiotic stress and stress combination. Plant J 90:856–867
Croce R, van Amerongen H (2013) Light-harvesting in Photosystem I. Photosyn Res 116:153–166
Croce R, Zucchelli G, Garlaschi FM, Bassi R, Jennings RC (1996) Excited state equilibration in the photosystem I-light-harvesting I complex: P700 is almost isoenergetic with its antenna. Biochemistry 35:8572–8579
Das K, Roychoudhury A (2014) Reactive oxygen species (ROS) and response of antioxidants as ROS-scavengers during environmental stress in plants. Front Environ Sci 2:53. https://doi.org/10.3389/fenvs.2014.00053
DeLano WL (2002) The PyMOL molecular graphics system Software
Delepelaire P, Chua NH (1979) Lithium dodecyl sulfate/polyacrylamide gel electrophoresis of thylakoid membranes at 4 degrees C: characterizations of two additional chlorophyll a-protein complexes. Proc Natl Acad Sci USA 76:111–115
Diz AP, Carvajal-Rodriguez A, Skibinski DOF (2011) Multiple hypothesis testing in proteomics: a strategy for experimental work. Mol Cell Proteomics. https://doi.org/10.1074/mcp.m110.004374
Djaman O, Outten FW, Imlay JA (2004) Repair of oxidized iron-sulfur clusters in Escherichia coli. J Biol Chem 279:44590–44599
Foyer CH, Noctor G (2013) Redox signaling in plants. Antiox Redox Signal 18:2087–2090
Frankel LK, Sallans L, Limbach PA, Bricker TM (2012) Identification of oxidized amino acid residues in the vicinity of the Mn4CaO5 cluster of Photosystem II: implications for the identification of oxygen channels within the photosystem. Biochemistry 51:6371–6377. https://doi.org/10.1021/bi300650n
Frankel LK, Sallans L, Limbach PA, Bricker TM (2013) Oxidized amino acid residues in the vicinity of QA and PheoD1 of the Photosystem II reaction center: putative generation sites of reducing-side reactive oxygen species. PLoS ONE 8:e58042
Guergova-Kuras M, Brent Boudreaux MB, Joliot A, Joliot P, Redding K (2001) Evidence for two active branches for electron transfer in Photosystem I. Proc Natl Acad Sci USA 98:4437–4442
Iwai MPG, Iavarone AT, Nogales E, Niyogi KK (2018) A unique supramolecular organization of Photosystem I in the moss Physcomitrella patens. Nat Plants 4:904–909
Jiao S, Emmanuel H, Guikema JA (2004) High light stress inducing photoinhibition and protein degradation of Photosystem I in Brassica rapa. Plant Sci 167:733–741
Kale R, Hebert AE, Frankel LK, Sallans L, Bricker TM, Pospíšil P (2017) Amino acid oxidation of the D1 and D2 proteins by oxygen radicals during photoinhibition of Photosystem II. Proc Natl Acad Sci USA 114:2988–2993
Kar UK, Simonian M, Whitelegge JP (2017) Integral membrane proteins: bottom-up, top-down and structural proteomics. Expert Rev Proteomics 14:715–723
Krieger-Liszkay A (2004) Singlet oxygen production in photosynthesis. J Exp Bot 56:337–346
Levesque-Tremblay G, Havaux M, Ouellet F (2009) The chloroplastic lipocalin AtCHL prevents lipid peroxidation and protects Arabidopsis against oxidative stress. Plant J 60:691–702
Li Y et al (2004) Mutation of the putative hydrogen-bond donor to P700 of Photosystem I. Biochemistry 43:12634–12647
Long SP, Humphries S, Falkowski PG (1994) Photoinhibition of photosynthesis in nature. Ann Rev Plant Mol Biol 45:633–662
Mazor Y, Borovikova A, Caspy I, Nelson N (2017) Structure of the plant Photosystem I supercomplex at 2.6 angstrom resolution. Nat Plants 3:17014
Melkozernov AN, Blankenship RE (2005) Structural and functional organization of the peripheral light-harvesting system in Photosystem I. Photosyn Res 85:33–50
Mignolet-Spruyt L et al (2016) Spreading the news: subcellular and organellar reactive oxygen species production and signalling. J Exp Bot 67:3831–3844
Pi X et al (2018) Unique organization of photosystem I-light-harvesting supercomplex revealed by cryo-EM from a red alga. Proc Natl Acad Sci USA 115:4423–4428
Pospíšil P (2009) Production of reactive oxygen species by Photosystem II. Biochim Biophys Acta 1787:1151–1160
Pospíšil P (2016) Production of reactive oxygen species by Photosystem II as a response to light and temperature stress. Front Plant Sci 7:1950. https://doi.org/10.3389/fpls.2016.01950
Qin X, Suga M, Kuang T, Shen JR (2015) Structural basis for energy transfer pathways in the plant PSI-LHCI supercomplex. Science 348:989–995
Rabilloud T, Vincon M, Garin J (1995) Micropreparative one- and two-dimensional electrophoresis: improvement with new photopolymerization systems. Electrophoresis 16:1414–1422
Rajagopal S, Joly D, Gauthier A, Beauregard M, Carpentier R (2005) Protective effect of active oxygen scavengers on protein degradation and photochemical function in Photosystem I submembrane fractions during light stress. FEBS J 272:892–902
Rutherford AW, Setif P (1990) Orientation of P700, the primary electron donor of Photosystem I. Biochim Biophys Acta 1019:128–132
Rutherford AW, Osyczka A, Rappaport F (2012) Back-reactions, short-circuits, leaks and other energy wasteful reactions in biological electron transfer: Redox tuning to survive life in O2. FEBS Lett 586:603–616
Sejima T, Takagi D, Fukayama H, Makino ACM, (2014) Repetitive short-pulse light mainly inactivates Photosystem I in sunflower leaves. Plant Cell Physiol 55:1184–1193
Sievers F et al (2011) Fast, scalable generation of high-quality protein multiple sequence alignments using Clust Omega. Mol Sys Biol 7:539. https://doi.org/10.1038/msb.2011.75
Sonoike K (1995) Selective photoinhibition of Photosystem I in isolated thylakoid membranes from cucumber and spinach. Plant Cell Physiol 36:825–830
Sonoike K (1996) Degradation of psaB gene product, the reaction center subunit of Photosystem I, is caused during photoinhibition of Photosystem I: possible involvement of active oxygen species. Plant Sci 115:157–164
Sonoike K (2011) Photoinhibition of Photosystem I. Physiol Plant 142:56–64
Sonoike K, Terashima I (1994) Mechanism of photosystem-I photoinhibition in leaves of Cucumis sativus L. Planta 194:287–293
Sonoike K, Terashima I, Iwaki M, Itoh S (1995) Destruction of photosystem I iron sulfur centers in leaves of Cucumis sativus L by weak illumination at chilling temperatures. FEBS Lett 3626(235):238
Sonoike K, Kamo M, Hihara Y, Hiyama T, Enami I (1997) The mechanism of the degradation of PsaB gene product, one of the photosynthetic reaction centre subunits of Photosystem I, upon photoinhibition. Photosyn Res 53:55–63
Souda P, Ryan CM, Cramer WA, Whitelegge JP (2011) Profiling of integral membrane proteins and their post translational modifications using high-resolution mass spectrometry. Methods 55:330–336
Srinivasan N, Golbeck JH (2009) Protein–cofactor interactions in bioenergetic complexes: the role of the A1A and A1B phylloquinones in Photosystem I. Biochim Biophys Acta 1787:1057–1088
Su X et al (2019) Antenna arrangement and energy transfer pathways of a green algal Photosystem-I-LHCI supercomplex. Nat Plants 5:273–281
Suga M, Ozawa SI, Yoshida-Motomura K, Akita F, Miyazaki N, Takahashi Y (2019) Structure of the green algal photosystem I supercomplex with a decameric light-harvesting complex I. Nat Plants 5:626–636
Sun G, Anderson VE (2004) Prevention of artifactual protein oxidation generated during sodium dodecyl sulfate-gel electrophoresis. Electrophoresis 25:959–965
Takagi D, Shigeo S, Hashiguchi M, Sejima T, Miyake C (2016) Superoxide and singlet oxygen produced within the thylakoid membranes both cause Photosystem I photoinhibition. Plant Physiol 171:1626–1634
Takahashi M, Asada K (1988) Superoxide production in aprotic interior of chloroplast thylakoids. Arch Biochem Biophys 267:714–722
Taylor RM, Sallans L, Frankel LK, Bricker TM (2018) Natively oxidized amino acid residues in the spinach cytochrome b6 /f complex. Photosyn Res 137:141–151
Terashima I, Funayama S, Sonoike K (1994) The site of photoinhibition in leaves of Cucumis sativus L. at low temperatures is Photosystem I, not Photosystem II. Planta 193:300–306
Tjus SE, Møller BL, Scheller HV (1998) Photosystem I is an early target of photoinhibition in barley illuminated at chilling temperatures. Plant Physiol 116:755–764
Tjus SE, Møller BL, Scheller HV (1999) Photoinhibition of Photosystem I damages both reaction centre proteins PSI-A and PSI-B and acceptor-side located small Photosystem I polypeptide Photosyn Res 60:75–86
Tripathy BC, Oelmuller R (2012) Reactive oxygen species generation and signaling in plants Plant Signal Behav 7:1621–1633
Weisz DA, Gross ML, Pakrasi HB (2017) Reactive oxygen species leave a damage trail that reveals water channels in Photosystem II Sci Adv 3:eaao3013
Xu H, Freitas MA (2009) MassMatrix: A database search program for rapid characterization of proteins and peptides from tandem mass spectrometry data Proteomics 9:1548-1555
You J, Chan Z (2015) ROS regulation during abiotic stress in crop plants Front. Plant Sci 6:1092
Yu Y, Smart LB, Jung Y-S, Golbeck J, Mcintosh L (1995) Absence of PsaC subunit allows assembly of Photosystem I core but prevents the binding of PsaD and PsaE in Synechocystis sp. PCC6803. Plant Mol Biol 29:331–342
Yu H, Wei J, Carpentier R (2000) Degradation of the Photosystem I complex during photoinhibition. Photochem Photobiol 72:508–512
Zivcak M, Brestic M, Kunderlikova K, Olsovska K, Allakhverdiev SI (2015) Effect of photosystem I inactivation on chlorophyll a fluorescenceinduction in wheat leaves: does activity of photosystem I play any role in OJIP rise? J Photochem Photobiol B 152:318–324
Acknowledgements
This work was solely supported by the United States Department of Energy, Office of Basic Energy Sciences grant DE-FG02-09ER20310 to TMB and LKF.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Kale, R., Sallans, L., Frankel, L.K. et al. Natively oxidized amino acid residues in the spinach PS I-LHC I supercomplex. Photosynth Res 143, 263–273 (2020). https://doi.org/10.1007/s11120-019-00698-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11120-019-00698-7