Abstract
Stress is usually considered an important factor resulting in plant injury. In this study, we identified a novel transcription factor gene, CanTF (Capsicum annuum transcription factor IIB), and characterized its role in the response to biotic and abiotic stresses. The full-length CanTF cDNA consists of 1488 bp, with a 1281-nucleotide open reading frame (ORF), and encodes a protein containing 426 amino acids with a theoretical molecular weight of 46.84 kDa. Real-time quantitative PCR revealed that CanTF is a stress-induced gene, with increased expression levels under both biotic and abiotic stresses. The expression of CanTF in pepper organs, especially roots, was highly induced by inoculation with an avirulent strain of Phytophthora capsici. Additionally, the differential expression of CanTF was observed under abiotic stresses, i.e., earlier expression was detected after cold, drought, and SA treatments than after salt and H2O2 treatment, suggesting its role in responses to various abiotic stresses. Furthermore, the silencing of CanTF by virus-induced gene silencing reduced the expression of defense-related genes (CaPR1, CaDEF1, and CaSAR82) under P. capsici inoculation. POD and root activity levels were lower after gene silencing than in controls, demonstrating the positive regulatory effect of CanTF against P. capsici. These results suggested that CanTF is a stress-induced gene involved in strengthening the pepper defense against biotic and abiotic stresses.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Pepper (Capsicum annuum L.) is one of the most important vegetable crops worldwide (Pimenta et al. 2016). Pepper is challenged by various biotic and abiotic stresses, such as cold stress (Sánchez-Bel et al. 2012), salinity (Penella et al. 2016), drought (Park et al. 2015), and pathogens (Lim et al. 2015). For example, chilling injury can cause lipid peroxidation and severely impair plant tolerance to low temperatures (Padma et al. 1997), while drought and salt stresses result in abnormal changes in physiology, such as photosynthesis, and low resistance to osmotic stress (Delfine et al. 2002; Park et al. 2016). Similarly, biotic stress can cause injury during pepper growth. For example, Phytophthora capsici can alleviate plant defense by secreting various pectin methyl-esterases during all stages of infection (Li et al. 2011; Zhang et al. 2016). Development is often disrupted by interactions between pepper plants and P. capsici. These biotic and abiotic stresses solely or jointly cause disorders of physiological metabolism during plant growth, resulting in plant injury. To combat these injuries, pepper has evolved several protective mechanisms, including the regulation by transcription factors (TFs) and hormones that enhance resistance to different stresses (Kim et al. 2009; Guo et al. 2015).
The importance of TFs in plant defense is evidenced by their ubiquity in plants (Jones and Dangl 2006). Regulation by TFs in plant development can activate the complex adaptive mechanisms and hence improve the resistance of plants to different stresses (Khong et al. 2015). For example, AP2/ERF TF binding to the promoter regions of pathogenesis-related genes can modulate their expression levels in response to pathogens (Ohme-Takagi and Shinshi 1995; Amorim et al. 2017). WRKY TFs are implicated in the regulation of plant defense mechanisms against pathogens by recognizing the W-box in promoter regions of some genes (Rushton et al. 2010). For example, SpWRKY3 can enhance resistance to Phytophthora infestans by inducing PR gene expression and reducing ROS accumulation (Cui et al. 2017). Most plant bZIP proteins binding to cis-elements can play important roles in plant immunity against pathogens (Alves et al. 2013; Noman et al. 2017). For example, StbZIP61, a potato transcription factor, regulates the dynamic biosynthesis of salicylic acid in defense against P. infestans infection (Zhou et al. 2018). Furthermore, GmPIB1, a bHLH TF, enhance resistance to Phytophthora sojae in soybean by repressing the expression of GmSPOD1 (Cheng et al. 2018).
The transcription factor IIB (TFIIB) family is well characterized, since it acts as a bridge between RNA polymerase II and the pre-initiation complex (PIC) and serves as the interaction target for multiple classes of trans-activator proteins that regulate gene expression in a specific manner (Lagrange et al. 2003). TFB-like proteins are characterized by an N-terminal zinc ribbon, a variable linker segment, and a cyclin fold domain, and there are a number of van der Waals contacts, salt bridges, and hydrogen bonds in the DNA–TBP–TFIIB interaction (Nikolov et al. 1995). Although the TFIIB protein does not exhibit sequence-specific DNA binding, a close association with DNA was discovered based on interactions with the phosphodiester backbone of DNA at the TATA element and its extension of the DNase footprint after joining a TBP–DNA complex (Malik et al. 1993; Nikolov et al. 1995). In Arabidopsis thaliana, 14 different TFIIB-like proteins encoding proteins in three major TFIIB subfamilies have been discovered (Knutson 2013). AtTFIIB1 has positive effects on pollen tube growth and endosperm development (Zhou et al. 2013), demonstrating its key role in the regulation of plant development. TFIIB provides protection against oxidative stress imposed by the extracellular environment or generated by cellular respiratory activity (de Faria and Fernandes 2006). However, very little information is available regarding the role of the TFIIB1 family in pepper defense against different stresses.
In the current study, a novel gene belonging to the TFIIB family, CanTF, was isolated from the pepper cultivar A3, and its characteristics under different stresses were evaluated. The objective of our study was to reveal the biological role of CanTF in the responses to biotic and abiotic stresses. The results provide insights in the mechanisms by which CanTF regulates pepper development under environmental stress and provides a basis for resistance breeding.
Materials and Methods
Plant Materials and Growth Conditions
Seeds of the pepper (Capsicum annuum L.) cultivar A3 (susceptible to HX-9 and resistant to PC of P. capsici) were obtained from the Capsicum Research Group, College of Horticulture, Northwest A&F University, China. Seeds were germinated, and seedlings were grown under the conditions described by Wang et al. (2013a).
Pathogen Preparation and Inoculation Procedures
P. capsici zoospore suspension was prepared according to the protocol used by Wang et al. (2013a) as follows: compatible (HX-9) and incompatible (PC) P. capsici strains were grown on PDA medium for 7 days, cut into pieces, and placed into sterile distilled water, followed by incubation in the light for 4 days, incubation for 1 h at 4 °C, and shaking for 1 h at 28 °C to release zoospores. The zoospores were collected by filtering through four layers of cheesecloth and were adjusted to 1 × 104 per millimeter. Pepper seedlings with six true leaves were inoculated by adding 2.5 mL of virulent/avirulent strains of P. capsici zoospore suspensions to each pot using the root drench method.
Stress Treatments
Various abiotic stresses, including cold, salt, and drought, along with hormone treatments were applied. Pepper seedlings were exposed to 4 °C for cold stress. For salt and drought stress treatments, the seedlings were immersed in 0.4 M sodium chloride (NaCl) and 0.4 M mannitol solutions, while water was used for the control-treated plants. Leaf samples from the treated and control seedlings were collected at 0, 2, 4, 8, 12, and 24 h post-stress treatment. For pathogen treatment, pepper seedlings with six-true leaves were inoculated by adding 2.5 mL of the virulent/avirulent P. capsici zoospore suspension to each pot using the root drench method. Leaf and root samples from treated and control (mock) plants were collected at 0, 2, 4, 8, 12, 24, 36, 48, and 72 h after inoculation. For hormonal treatments, each hormone (5 mM salicylic acid (SA), 50 μM methyl jasmonate (MeJA), and 10 μM H2O2) was supplemented with 0.5% Tween-20 and sprayed on pepper seedlings. Leaves from mock and treated plants were collected at 0, 2, 4, 8, 12, 24, and 48 h after treatment. All collected samples were instantly frozen in liquid nitrogen and stored at − 80 °C. All treatments were performed and analyzed thrice in separate experiments.
Sequence Analysis of the CanTF Gene
A multiple sequence alignment of CanTF amino acid sequences was obtained using DNAMAN (Version 5.0). The amino acid sequence of CanTF and homologs in other plant species were aligned using CLUSTALW as described by Guo et al. (2016), and MEGA6.0 was used to construct a phylogenetic tree by the neighbor-joining method (Benson et al. 2000).
Tissue-Specific Expression of CanTF in Pepper
To evaluate the tissue-specific expression of CanTF, samples were collected from roots, stems, leaves, flowers, green fruits, and mature fruits of the pepper cultivar A3, immediately frozen in liquid nitrogen, and stored at − 80 °C until RNA extraction.
RNA Isolation and Real-Time RT-PCR Analysis
RNA was extracted from samples at different time points, as mentioned previously, using TRIzol reagent (Invitrogen, Carlsbad, CA, USA). For reverse transcription, 0.5 μg of total RNA was used for first-strand cDNA synthesis using a PrimeScript™ Kit (TaKaRa Bio Inc., Dalian, China). Real-time quantitative PCR (qRT-PCR) was performed as described by Guo et al. (2016). Ubi3 (AY486137.1) was used as internal control (reference gene) (Wan et al. 2011). The primer sequences used in the study are shown in Table 1. Relative CanTF expression levels were determined by the comparative 2−∆∆CT method (Litvak and Schmittgen 2001).
Cloning and Sequence Analysis of CanTF
The rapid amplification of cDNA ends (RACE) technique was used to obtain the full-length cDNA designated CanTF. According to the partial cDNA fragment of the reported sequence (accession number GD094649.1), a set of gene-specific primers (CanTFF, CanTFR) for 5′ and 3’ RACE was designed. Contig Express software and BLAST were used to assemble the full-length cDNA sequence. The full-length cDNA sequence was obtained from the leaves of the pepper cultivar A3 using the primer pair (CanTFQF and CanTFQR).
The pI and MW of CanTF were analyzed using the online tool (https://www.expasy.org/), and conserved domains were identified using an NCBI tool (available online: http://www.ncbi.nlm.nih.gov/cdd/). The secondary structure was predicted using ScratchProtein Predictor (https://www.ics.uci.edu/~baldig/scratch/). The transmembrane and signal peptides were predicted using the CBS prediction server (http://www.cbs.dtu.dk/services/).
VIGS of CanTF in Pepper
For gene silencing in pepper plants, the virus-induced gene silencing (VIGS) system was used (Choi et al. 2007). CaPDS was used for the phenotypic effectiveness of VIGS. A 256-bp fragment harboring a conserved and a non-conserved region of CanTF with primers containing restriction sites for EcoRI and BamHI enzymes was cloned into the TRV2 vector. Then, the freeze-thaw method was used to transform the TRV1, TRV2, TRV2-CaPDS, and TRV2-CanTF vectors into the Agrobacterium tumefaciens strain GV3101. The A. tumefaciens strain (GV3101) harboring TRV1 was mixed at a 1:1 ratio with TRV2, TRV2-CaPDS, and TRV2-CanTF. Then, a sterilized syringe without a needle was used to inoculate the Agrobacterium suspensions to the cotyledons of pepper plants. Plants were placed in a growth chamber with the same growth conditions described by Wang et al. (2013b).
After 5 weeks of infiltration, the upper 4th leaves from TRV2:00 (control) and TRV2: CanTF (silenced) plants were injected with the P. capsici suspension. In addition, control (TRV2:00) and silenced (TRV2: CanTF) plants were subjected to 4 °C for cold stress treatment, 0.4 M sodium chloride (NaCl) and mannitol for salt and drought stress treatments. Leaf samples were collected for the measurement of peroxidase activity assay and conductivity.
Measurements of Peroxidase Activity Assay and Conductivity
Peroxidase (POD) enzyme activity was quantified using the technique of Beffa et al. (1990). To evaluate the permeability of the membrane, conductivity was determined as described by Dionisio-Sese and Tobita (1998).
Determination of Root Activity
The triphenyltetrazolium chloride (TTC) was used to measure root activity according to the methods of Wang et al. (2013a). P. capsici post-inoculation root samples of about 0.5 g were collected from control and CanTF-silenced plants. A modified TTC method was used to measure root activity as described by Jin et al. (2016). The treatments were evaluated in three biological replications, and measurements were repeated thrice.
Statistical Analysis
Data were evaluated by an analysis of variance (ANOVA) using SPSS (version 16.0, SPSS Inc., Chicago, IL, USA). Data are expressed as the means ± standard deviation (SD) of three replicates for all parameters. A least significant difference (P ≤ 0.05) test was used to identify significant differences among the treatments.
Results
Sequence Analysis of CanTF
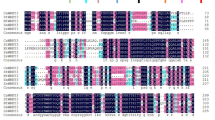
The full-length cDNA of CanTF was obtained by RACE. CanTF consisted of 1488 nucleotides, including a 5′-untranslated region (UTR) of 100 bp, an open reading frame (ORF) of 1281 bp and a 3′-UTR of 107 bp with a poly (A+) tail of 26 bp (GenBank accession number FJ617518). The ORF encoded a predicted protein of 426 amino acids with a theoretical molecular weight (MW) of 46.84 kDa and isoelectric point (pI) of 5.39. A structural analysis revealed that CanTF belongs to the transcription factor IIB (TFIIB) superfamily, containing a zinc-binding region (C3-X2-C6-X18-C25-X2-C28) and two binding sites (130-174, 226-312), as shown in Fig. 1, while TMpred showed that CanTF does not contain a trans-membrane helix. Therefore, CanTF probably lacks a trans-membrane domain. Further sequence analysis indicated that the deduced CanTF protein does not contain a signal peptide. In addition, high amino acid sequence homology was observed between CanTF and other TFIIB proteins based on multiple alignments obtained using ClustalW. The percent identities of CanTF relative to Solanum tuberosum (XP_006340594.1), Solanum lycopersicum (NP_001308007.1), Nicotiana tomentosiformis (XP_009601399.1), Nicotiana sylvestris (XP_009791322.1), Prunus persica (XP_007203626.1), Vigna radiata var. radiate (XP_014519199.1), Glycine soja (KHN36117.1), Morus notabilis (XP_010095267.1), Gossypium arboretum (KHG10594.1), Theobroma cacao (XP_007046979.1), and Prunus mume (XP_008241653.1) were 94%, 93%, 93%, 93%, 75%, 73%, 71%, 73%, 73%, 74%, and 68%, respectively (Fig. 1). The phylogenetic analysis was carried out using MEGA6.0. Two clusters were observed using the amino acid sequences of 14 TFIIB genes from Solanum lycopersicum (NP_001308007.1), Solanum pennellii (XP_015064013.1), Solanum tuberosum (XP_006340594.1), Capsicum annuum (NP_001311673.1), Nicotiana sylvestris (XP_009791324.1), Theobroma cacao (XP_007046979.2), Brassica oleracea var. oleracea (XP_013597636.1), Arabidopsis thaliana (NP_195383.1 and NP_001078502.4), Arabidopsis lyrata subsp. lyrata (XP_002867001.1), Vigna radiata var. radiate (XP_014519199.1), Cicer arietinum (XP_004512022.1), Morus notabilis (XP_010095267.1), and Prunus mume (XP_008241653.1) (Fig. 2). CanTF (FJ617518) of pepper and other Solanaceae plants contained smaller subunit TFIIB sequences.
Multiple sequence alignment of the amino acid sequence of CanTF with amino acid sequences of StTFIIB (Solanum tuberosum, XP_006340594.1), SlTFIIB (Solanum lycopersicum, NP_001308007.1), NtTFIIB (Nicotiana tomentosiformis, XP_009601399.1), NsTFIIB (Nicotiana sylvestris, XP_009791322.1), PpTFIIB (Prunus persica, XP_007203626.1), VrTFIIB (Vigna radiata var. radiata, XP_014519199.1), GsTFIIB (Glycine soja, KHN36117.1), MnTFIIB (Morus notabilis, XP_010095267.1), GaTFIIB (Gossypium arboretum, KHG10594.1), TcTFIIB (Theobroma cacao, XP_007046979.1), and PmTFIIB (Prunus mume, XP_008241653.1). Dark-blue shading and pink shading reflect 100% and 75% conservation, respectively. Gaps (−) were introduced to maximize the alignment. The zinc-binding region is indicated by a solid line and the two-binding sites are indicated by the dashed and the single short lines
Phylogenetic tree of proteins homologous to CanTF and TFIIB proteins from other species. The rooted gene tree (majority-rule consensus from 1000 bootstrap replicates) was obtained by a heuristic search using MEGA6.0. Bootstrap values are indicated at each branch node. GenBank accession numbers are in parentheses after each species and gene name. The red point represents CanTF (NP_001311673.1). Scale bar indicates the similarity coefficient
Tissue-Specific Expression of CanTF in Pepper
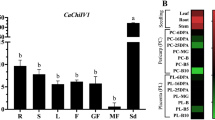
To investigate the tissue-specific expression level of CanTF, RNA was extracted from leaves, stems, roots, flowers, and both green and red fruits (Fig. 3). The results indicated that the expression levels of CanTF were significantly higher in both green and red fruits than in other tissues, with no significant difference between these fruits. Among other tissues, the expression levels were highest in leaves, followed by stems, flowers, and roots, but the differences were not significant.
Expression of CanTF in Response to P. capsici Infection
CanTF expression levels were quantified in the roots and leaves of post-inoculation P. capsici strains (HX-9 and PC). Based on qRT-PCR, CanTF was strongly up-regulated by the incompatible strain of P. capsici (Fig. 4). In roots, the expression level of CanTF was significantly increased by inoculation with the PC strain at 72 h post-inoculation, followed by 48 h and 12 h. For the HX-9 strain, the expression level was slightly increased at 2, 48, and 72 h after inoculation, without any significant differences among these and other time points (Fig. 4a). In leaves, CanTF expression levels exhibited almost the same trend as those in roots for both strains (HX-9 and PC), as shown in Fig. 4b. Generally, the transcript level of CanTF in leaf was greater for the PC strain than the HX-9 strain. The inoculation of plants with the HX-9 strain increased the expression in leaves to peak levels (3.325 ± 1.19) at 2 h post-inoculation, followed by 72 h (2.41 ± 1.499), while the expression level at 24 h ranked third. For inoculation with the PC strain, the expression level increased at three different time points and reached the highest levels at 72, 24, and 48 h after inoculation.
Expression of CanTF in Response to Abiotic Stresses and Hormonal Treatments
To evaluate the role of CanTF in protection against abiotic stresses, pepper plants were exposed to cold, salt, and drought stresses (Fig. 4c–e). Compared to the control, the expression level of CanTF was about 3-fold higher (3.82) at 4 h post-cold stress (4 °C) (Fig. 4c), while exposure to salt stress significantly increased the expression level by 3-fold or more at 12 h (3.08) and 24 h (3.32) (Fig. 4d). CanTF also respond to drought stress; its expression increased 9-fold (9.44) at 2 h post-treatment (Fig. 4e).
While evaluating the expression levels of CanTF in response to signaling molecules, leaf samples from plants treated with SA, MeJA, and H2O2 were collected, and their cDNA was used for qRT-PCR to analyze expression levels. Plants treated with H2O2 showed a slight increase in the expression of CanTF at 2 h (10.79) post-treatment, followed by a slight decrease and an increase to the maximum level at 48 h post-treatment, which was almost 56-fold increase compared with the expression level in the control (Fig. 5a). In response to SA treatment, the expression level of CanTF increased dramatically by 77-fold (77.36) at 2 h post-treatment as compared to the control, but showed no significant effect at other time points (Fig. 5b). Plants treated with MeJA showed initially downregulated expression, reaching the lowest level at 4 h post-treatment, followed by an upregulation, reaching a maximum at 12 h, and downregulation after 12 h, but the differences in the expression level of CanTF were not significant (Fig. 5c).
Silencing of CanTF Weakened the Defense Response of Pepper Against P. capsici
Five weeks after infiltration, the visible symptoms of photobleaching due to a loss of chlorophyll were observed in CaPDS-silenced plant leaves (Fig. 6a), confirming that VIGS was successful. Furthermore, a qRT-PCR analysis of RNA extracted from the leaves of CanTF-silenced plants (TRV2:CanTF) and empty vector control plants (TRV2:00) was performed to clarify the efficiency of CanTF silencing by VIGS. As shown in Fig. 6b, the levels of CanTF transcripts were significantly reduced to different extents in CanTF-silenced plants compared to those in the empty vector control. These results indicated that CanTF was partially silenced. Thus, to verify the role of CanTF in defense response, the fourth to fifth leaves from the top of the silenced (TRV2:CanTF) and control (TRV2:00) pepper plants were removed and inoculated with HX-9 strain of P. capsici. On the 4th day of inoculation, more lesions were detected on the leaves of the CanTF-silenced plants (Fig. 7b) than on the control (TRV2:00) plant leaves (Fig. 7a). In addition, POD activity was measured in the leaves of control (TRV2:00) and silenced plants after HX-9 strain infection. The results indicated that POD activity was increased in both silenced and control plants, but the silenced plants had the lowest POD activity as compared to that of the control (TRV2:00).
Effect of CanTF gene silencing on pepper plants. Images were obtained 35 days after inoculation. a Phenotypes of gene-silenced pepper plants. Left: control plant (TRV2:00); middle: CaPDS-silenced plant (TRV2:CaPDS); right: CanTF-silenced plant (TRV2:CanTF). b Real-time RT-PCR analysis of CanTF expression levels in TRV2:00 line and TRV2:CanTF lines (#1, #2, and #3 indicate three biological levels of efficiency of CanTF gene silencing) 35 days after inoculation
Silencing of CanTF weakened the tolerance of pepper plants to P. capsici inoculation. a, b Disease symptoms in detached-leaves after inoculation with P. capsici. a Control plant (TRV2:00). b Silenced pepper plant (TRV2:CanTF). c Activity of POD after inoculation with P. capsici. d Root activity in CanTF-silenced and control plants after inoculation with the PC strain. e Root activity in CanTF-silenced and control plants after inoculation with the HX-9 strain
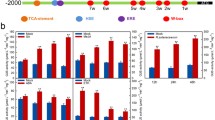
After inoculate with the PC strain, the expression levels of CanTF, CaPR1, CaDEF1, and CaSAR82 were elevated, but the increases in TRV2:00 plants were greater than those in CanTF-silenced plants. Significant increases in the expression of CanTF, CaPR1, and CaDEF1 were detected at 24 h post-inoculation in the non-silenced plants, while no significant increases in expression were observed in silenced plants (Fig. 8a–d). Similarly, after the inoculation of the HX-9 strain, the expression levels of CanTF and defense-related genes (CaPR1, CaPR1, CaDEF1, and CaSAR8.2) were upregulated at 24 h post-inoculation (Fig. 8e–h). The upregulation was significantly higher in non-silenced plants than in silenced plants. Furthermore, after P. capsici inoculation, we measured root activity in the silenced and control plants by the TTC method. Root activity in the silenced plants was lower than that in the control, and in the case of the HX-9 strain, a significant difference was observed at 3 and 7 days post-inoculation between silenced and control plants (Fig. 7d, e).
Silencing of CanTF reduced the expression of defense-related genes (CaPR1, CaDEF1, and CaSAR82) in pepper leaves at 1 d post-inoculation with PC (a–d) and HX-9 (e–h) strains of P. capsici. Values are means ± SD from three independent experiments. Small letters represent significant differences (p < 0.05)
Silencing of CanTF Weakened Tolerance to Abiotic Stress
Silencing of CanTF resulted in more severely bleached leaf discs compared to those of empty vector-treated plants after exposure to salt stress for 4 days (Fig. 9a). The same results were obtained for POD activity (Fig. 9b). There was no significant difference in POD activity at 0 h between the TRV2:CanTF and TRV2:00 lines, but a significant difference was observed at 6 h, 24 h, and 48 h after salt stress treatment. Although POD activity in the leaves of CanTF-silenced plants was increased at 6 h, an obvious decrease was observed at 24 and 48 h. These results indicated that the silencing of CanTF results in poor plant defense under long-term salt stress.
Weakened tolerance of CanTF-silenced pepper plants to salt stress. a Phenotypes of the silenced and control plants after salt stress. Leaf discs from the gene-silenced pepper leaves and control plant leaves were floated in 400 mM NaCl solutions for 4 days at 26 °C with continuous fluorescent lighting. b Activity of POD after salt stress. Error bars represent SD for three biological replicates. Small letters represent significant differences (p < 0.05)
Exposure of the TRV2:00 and TRV2:CanTF lines to cold stress resulted in increased POD activity at 24 h post-treatment compared with that at 0 h (Fig. 10a). However, the increases were less substantial for the TRV2:CanTF line than TRV2:00. The opposite trend was observed for electrolyte leakage. Leaf conductivity increased throughout the experimental period after cold stress treatment in the TRV2:CanTF line. Furthermore, cold stress increased electrolyte leakage more significantly in leaves of the TRV2:CanTF line than the TRV2:00 line (Fig. 10b).
Weakened tolerance of CanTF-silenced pepper plants to cold and drought stress. a Activity of POD after cold stress. b Conductivity of leaf discs after cold treatment. c Activity of POD after drought stress. d Conductivity of the leaf discs after drought stress. Error bars represent SD for three biological replicates. Small letters represent significant differences (p < 0.05)
Mannitol stress resulted in lower POD activity in CanTF-silenced plants at 24 and 48 h post-treatment than in TRV2:00 (Fig. 10c). Under drought stress, electrolyte leakage increased gradually and a significant difference was observed at 6 and 24 h post-treatment (Fig. 10d).
Discussion
In response to different biotic and abiotic stresses, plants have a finely regulated and intricate defense system (Choi and Hwang 2012). The roles of TFs and proteins in controlling different biological processes, e.g., responses to biotic and abiotic stresses, development, differentiation, metabolism, and defense, are clearly established (Ambawat et al. 2013). TFs can activate complex biological process (Fishburn et al. 2015) and can be affected by stress (Singh et al. 2002; Shoji and Hashimoto 2015). In higher plants, TF genes have been investigated in several species, including petunia, rice, cotton, maize, and Arabidopsis (Ambawat et al. 2013). In the current study, a new TF gene, CanTF, was isolated and identified from pepper. TFIIB-related genes are characterized by several intriguing features. The full sequence of CanTF in pepper cultivar A3 consisted of 1488 nucleotides. CanTF did not include a trans-membrane domain, demonstrating that the deduced protein does not contain a signal peptide. The putative amino acid sequence of CanTF showed high homology to other TFIIB proteins, such as those of Solanum tuberosum and Nicotiana tomentosiformis. A tissue-specific expression analysis showed that CanTF was expressed in all the tested tissues and was highly expressed in both the green and red fruits of pepper plants. In agreement with our results, previous studies have reported that AtTFB genes are widely expressed in different plant tissues, including vegetative nuclei, generative cells of pollen grains, pollen tubes, endosperm, and embryos. The results suggest that AtTFIIBs play important roles in the reproductive phase of a plant (Knutson 2013), and members of TFIIB exhibit sensitivity to oxidative stress (de Faria and Fernandes 2006).
CanTF showed early expression under cold, drought, and SA stresses as well as late expression under salt and H2O2. These results suggested that CanTF can be induced by abiotic stress and signaling molecules, and this may be related to the characteristics of the TFIIB family. Previous studies have reported that TFIIB genes have potential roles in controlling the cell cycle (Gibson et al. 1994), and the response of the cell cycle to stress is usually regulated in a time-dependent manner (Reichheld et al. 1999; Solé et al. 2015). Our results clearly indicate that CanTF expression increased in pepper plants exposed to abiotic stresses. Cold stress significantly increased CanTF gene expression at 4 h post-treatment, salt stress at 12 and 24 h, and drought stress at 2 h post-treatment. As a result of exposure to various abiotic stresses, the expression level of CanTF increased, suggesting that this gene is directly related to abiotic stress tolerance. Our results are in agreement with those of Denekamp and Smeekens (2003), who found that AtMYB in Arabidopsis is up-regulated by drought stress. Furthermore, AtMYB2 was induced by drought and salt stresses, which indicates that AtMYB2 is responsive to water deficiency at the transcriptional level (Abe et al. 2003). Yi et al. (2004) also revealed that the transcript level of CaPF1 is stimulated by several treatments, including cold stress, ethephon, and methyl jasmonate.
Various plant hormones are important signaling molecules with a vital role in the resistance of plants to biotic and abiotic stresses (Choi and Hwang 2011). In the present study, SA and H2O2 signaling molecules were used to treat pepper plants, resulting in significantly increased CanTF expression. These results suggest that CanTF has a potential role in the hormonal response to biotic and abiotic stress. In Arabidopsis, various TFs are stimulated by salt and dehydration stress (Shinozaki et al. 1992). Moreover, Lim et al. (2015) have found that the CabZIP2 gene is induced by defense-related hormones, such as SA and MeJA.
The results of the current study revealed that infecting pepper plants with two strains of P. capsici (HX-9 and PC) affected CanTF expression. These results suggested that CanTF is involved in pathogen defense. Our results are also in line with the findings of Yi et al. (2004). In Arabidopsis, MYB-TF genes play roles in the defense response against pathogen infection (Li et al. 2009). Furthermore, functional analyses of pathogen-induced TFs have revealed key roles in plant immunity (Reddy et al. 2011). In response to X. campestris pv. vesicatoria infection, CabZIP2 transcripts accumulate more rapidly and strongly for the incompatible interaction compared with the compatible interaction (Lim et al. 2015). Thus, the difference in stress response by CanTF, a member of the TFIIB family, may be closely related to the cell cycle, which is affected by the TFIIB structure. Higher expression levels of CanTF were observed when pepper plants were exposed to P. capsici, especially the PC strain, suggesting that CanTF also enhances the pepper defense response against P. capsici. Generally, the PC strain post-inoculation exhibits higher expression levels of CanTF in roots than in leaves, while lower expression of CanTF was observed in both roots and leaves of the HX-9 strain post-inoculation.
Some genes in pepper showed low expression under PC strain inoculation than HX-9 strain inoculation (Jin et al. 2016; Zhang et al. 2016). However, in our study, CanTF showed the opposite response to P. capsici strain inoculation, i.e., higher expression of CanTF was observed under PC strain inoculation as compared to HX-9 strain inoculation. However, the mechanisms underlying the interaction between CanTF and the pepper defense response remain elusive. Thus, we speculate that CanTF responses to both biotic and abiotic stress may regulate the plant defense response. Therefore, we further investigated the function of CanTF by VIGS.
In the current study, photobleaching symptoms were observed on CaPDS-silenced plants leaves, indicating that the VIGS assay was successful. Silencing of CanTF revealed that the transcript levels of CanTF in silenced plants (TRV2:CanTF) were drastically reduced compared to the levels in control plants (TRV:00), indicating that CanTF was effectively silenced in pepper plants.
Furthermore, after inoculation with compatible and incompatible strains of P. capsici, the expression levels of defense-related CaDEF1 (JA-dependent), CaPR1 (SA-dependent), and CaSAR82 (systemic acquired resistance) genes in the leaves of CanTF-silenced plants were lower than those in control plants. Additionally, root activity in the silenced plants was reduced as compared with that of the control plants, indicating its role in P. capsici resistance. Similarly, the silencing of CaPTI1 also compromised the expression levels of CaPR1, CaDEF1, and CaSAR82 as well as the root activity of silenced plants as compared to those in control plants (Jin et al. 2016). The transcript levels of defense-related CaPR1 and CaDEF1 were significantly lower in leaves of R. solanacearum-inoculated CabZIP63-silenced than in the control pepper plants (Shen et al. 2016). Another study supporting our results showed that the silencing of CaPAL1 suppresses the expression of CaPR1 in silenced plants as compared to control plants during Xcv infection (Kim and Hwang 2014). The silencing of CaMLO2 significantly suppresses the induction of CaDEF1 transcripts in the leaves of CaMLO2-silenced plants as compared to the control plants after inoculation with virulent Xcv Ds1 (EV) or avirulent Xcv Ds1 (avrBsT) (Kim et al. 2014). For infection with Xcv (especially the virulent strain), CaPO2-silenced plants exhibited significantly less induction of CaDEF1 and CaSAR82 (Choi et al. 2007). Yeom et al. (2011) used the cell vitality of the root as an indicator of the degree of injury in response to P. capsici infection and measured TTC reductase activity in the roots of pepper cultivars after P. capsici inoculation; they observed significant differences in root activity from 24 to 48 hpi, and concluded that the activity of CM334 is 2–3 times greater than that in Chilsungcho.
Disease symptoms were observed on leaves detached from the CanTF-silenced plants on the 4th day of P. capsici inoculation, while no disease symptoms were observed on the detached leaves of the control (TRV2:00) plants. This indicates that CanTF has an important role in the defense response to stress (Choi et al. 2007). In agreement with the present results, Wang et al. (2013b) also found that control plants were more resistant to P. capsici infection than TF-silenced plants. Our results are similar to those of Lim et al. (2015) who found that CabZIP2-silenced pepper plants are susceptible to infection by a virulent strain of pathogen. In addition, it was reported in the same study that TF genes are involve in the pepper plant defense response against different stresses. These results demonstrate that CanTF has an important role in the pepper defense response against different types of stresses.
Conclusion
A novel TF gene belonging to the TFIIB family, CanTF, was identified in pepper by a bio-informatics analysis, with a potential role in plant development. CanTF is induced by abiotic and biotic stresses and may be responsible for plant defense. The expression of CanTF in pepper organs, especially roots, exhibits high sensitivity to avirulent strains compared with virulent strains, while differences in responses to abiotic stresses may be explained by differences in the expression of CanTF. Early expression under cold, drought, and SA stresses and late expression under salt and H2O2 were observed in our study. VIGS technology showed that CanTF may positively regulate the responses to biotic and abiotic stresses, which indicates that CanTF plays a key role in plant defense. Our results provide important information regarding the regulatory role of TFIIB genes for plant defense, and additional work should focus on the mechanisms underlying the interaction between the CanTF gene and environmental stresses.
References
Abe H, Urao T, Ito T, Seki M, Shinozaki K, Yamaguchi-Shinozaki K (2003) Arabidopsis AtMYC2 (bHLH) and AtMYB2 (MYB) function as transcriptional activators in abscisic acid signaling. Plant Cell 15:63–78. https://doi.org/10.1105/tpc.006130
Alves MS, Dadalto SP, Gonçalves AB, De Souza GB, Barros VA, Fietto LG (2013) Plant bZIP transcription factors responsive to pathogens: a review. Int J Mol Sci 14:7815–7828
Ambawat S, Sharma P, Yadav NR, Yadav RC (2013) MYB transcription factor genes as regulators for plant responses: an overview. Physiol Mol Biol Plants 19:307–321. https://doi.org/10.1007/s12298-013-0179-1
Amorim LL, da Fonseca-Dos-Santos R, Guida-Santos M, Crovella S, Benko-Iseppon AM (2017) Transcription factors involved in plant resistance to pathogens. Curr Protein Pept Sci 18:335–351
Beffa R, Martin HV, Pilet P (1990) In vitro oxidation of indoleacetic acid by soluble auxin-oxidases and peroxidases from maize roots. Plant Physiol 94:485–491. https://doi.org/10.1104/pp.94.2.485
Benson DA, Karsch-Mizrachi I, Lipman DJ, Ostell J, Rapp BA, Wheeler DL (2000) GenBank. Nucleic Acids Res 28:15–18
Cheng Q, Dong L, Gao T, Liu T, Li N, Wang L, … Zhang S (2018). The bHLH transcription factor GmPIB1 facilitates resistance to Phytophthora sojae in Glycine max. J Exp Botany, 69(10), 2527–2541. Doi:https://doi.org/10.1093/jxb/ery103
Choi DS, Hwang BK (2011) Proteomics and functional analyses of pepper abscisic acid-responsive 1 (ABR1), which is involved in cell death and defense signaling. Plant Cell 23:823–842. https://doi.org/10.1105/tpc.110.082081
Choi HW, Hwang BK (2012) The pepper extracellular peroxidase CaPO2 is required for salt, drought and oxidative stress tolerance as well as resistance to fungal pathogens. Planta 235:1369–1382. https://doi.org/10.1007/s00425-011-1580-z
Choi HW, Kim YJ, Lee SC, Hong JK, Hwang BK (2007) Hydrogen peroxide generation by the pepper extracellular peroxidase CaPO2 activates local and systemic cell death and defense response to bacterial pathogens. Plant Physiol 145:890–904
Cui J, Xu P, Meng J, Li J, Jiang N, Luan Y (2017) Transcriptome signatures of tomato leaf induced by Phytophthora infestans and functional identification of transcription factor SpWRKY3. Theor Appl Genet 131(4):787–800. https://doi.org/10.1007/s00122-017-3035-9
de Faria J, Fernandes L (2006) Protection against oxidative stress through SUA7/TFIIB regulation in Saccharomyces cerevisiae. Free Radic Biol Med 41:1684–1693. https://doi.org/10.1016/j.freeradbiomed.2006.09.003
Delfine S, Tognetti R, Loreto F, Alvino A (2002) Physiological and growth responses to water stress in field-grown bell pepper (Capsicum annuum L.). J Hortic Sci Biotechnol 77:697–704. https://doi.org/10.1080/14620316.2002.11511559
Denekamp M, Smeekens SC (2003) Integration of wounding and osmotic stress signals determines the expression of the AtMYB102 transcription factor gene. Plant Physiol 132:1415–1423. https://doi.org/10.1104/pp.102.019273
Dionisio-Sese ML, Tobita S (1998) Antioxidant responses of rice seedlings to salinity stress. Plant Sci 135:1–9. https://doi.org/10.1016/S0168-9452(98)00025-9
Fishburn J, Tomko E, Galburt E, Hahn S (2015) Double-stranded DNA translocase activity of transcription factor TFIIH and the mechanism of RNA polymerase II open complex formation. Proc Natl Acad Sci U S A 112:3961–3966. https://doi.org/10.1073/pnas.1417709112
Gibson TJ, Thompson JD, Blocker A, Kouzarides T (1994) Evidence for a protein domain superfamily shared by the cyclins, TFIIB and RB/p107. Nucleic Acids Res 22:946–952. https://doi.org/10.1093/nar/22.6.946
Guo W-L, Wang S-B, Chen R-G, Chen B-H, Du X-H, Yin Y-X, Gong Z-H, Zhang Y-Y (2015) Characterization and expression profile of CaNAC2 pepper gene. Front Plant Sci 6:1–9. https://doi.org/10.3389/fpls.2015.00755
Guo M, Liu J-H, Ma X, Zhai Y-F, Gong Z-H, Lu M-H (2016) Genome-wide analysis of the Hsp70 family genes in pepper (Capsicum annuum L.) and functional identification of CaHsp70-2 involvement in heat stress. Plant Sci 252:246–256. https://doi.org/10.1016/j.plantsci.2016.07.001
Jin J-H, Zhang H-X, Tan J-Y, Yan M-J, Li D-W, Khan A, Gong Z-H (2016) A new ethylene-responsive factor CaPTI1 gene of pepper (Capsicum annuum L.) involved in the regulation of defense response to Phytophthora capsici. Front Plant Sci 6:1–12. https://doi.org/10.3389/fpls.2015.01217
Jones J, Dangl J (2006) The plant immune system. Nature 444:323–329. https://doi.org/10.1038/nature05286
Khong GN, Richaud F, Coudert Y, Pati PK, Périn C, Breitler J, Meynard D, Do N, Guiderdoni E, Gantet P (2015) Modulating rice stress tolerance by transcription factors modulating rice stress tolerance by transcription factors. Biotechnol Genet Eng Rev 25:381–404. https://doi.org/10.5661/bger-25-381
Kim DS, Choi HW, Hwang BK (2014) Pepper mildew resistance locus O interacts with pepper calmodulin and suppresses Xanthomonas AvrBsT-triggered cell death and defense responses. Planta 240:827–839. https://doi.org/10.1007/s00425-014-2134-y
Kim DS, Hwang BK (2014) An important role of the pepper phenylalanine ammonia-lyase gene (PAL1) in salicylic acid-dependent signalling of the defence response to microbial pathogens. J Exp Bot 65:2295–2306. https://doi.org/10.1093/jxb/eru109
Kim T-H, Ok SH, Kim D, Suh S-C, Ok Byun M, Shin JS (2009) Molecular characterization of a biotic and abiotic stress resistance-related gene RelA/SpoT homologue (PepRSH) from pepper. Plant Sci 176(5):635–642. https://doi.org/10.1016/j.plantsci.2009.02.004
Knutson BA (2013) Emergence and expansion of TFIIB-like factors in the plant kingdom. Gene 526:30–38. https://doi.org/10.1016/j.gene.2013.04.022
Lagrange T, Hakimi M-A, Pontier D, Courtois F, Alcaraz JP, Grunwald D, Lam E, Lerbs-Mache S (2003) Transcription factor IIB (TFIIB)-related protein (pBrp), a plant-specific member of the TFIIB-related protein family. Mol Cell Biol 23:3274–3286. https://doi.org/10.1128/MCB.23.9.3274-3286.2003
Li P, Feng B, Wang H, Tooley PW, Zhang X (2011) Isolation of nine Phytophthora capsici pectin methylesterase genes which are differentially expressed in various plant species. J Basic Microbiol 51:61–70. https://doi.org/10.1002/jobm.201000317
Li L, Yu X, Thompson A, Guo M, Yoshida S, Asami T, Chory J, Yin Y (2009) Arabidopsis MYB30 is a direct target of BES1 and cooperates with BES1 to regulate brassinosteroid-induced gene expression. Plant J 58:275–286. https://doi.org/10.1111/j.1365-313X.2008.03778.x
Lim CW, Baek W, Lim S, Han S, Lee SC (2015) Expression and functional roles of the pepper pathogen–induced bZIP transcription factor CabZIP2 in enhanced disease resistance to bacterial pathogen infection. Mol Plant-Microbe Interact 28:825–833. https://doi.org/10.1094/MPMI-10-14-0313-R
Litvak K, Schmittgen T (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2 (Delta Delta C (T)) method. Methods 25:402–408. https://doi.org/10.1006/meth.2001.1262
Malik S, Lee KUN, Roeder RG (1993) Potential RNA polymerase II-induced interactions of transcription factor TFIIB. Mol Cell Biol 13:6253–6259. https://doi.org/10.1128/MCB.13.10.6253
Nikolov DB, Chen H, Halay ED, Usheva AA, Hisatake K, Lee DK, Roeder RG, Burley SK (1995) Crystal structure of a TFIIB-TBP-TATA-element ternary complex. Nature 377:119–128. https://doi.org/10.1038/377119a0
Noman A, Liu Z, Aqeel M, Zainab M, Khan MI, Hussain A, Ashraf MF, Li X, Weng Y, He S (2017) Basic leucine zipper domain transcription factors: the vanguards in plant immunity. Biotechnol Lett 39:1779–1791
Ohme-Takagi M, Shinshi H (1995) Ethylene-inducible DNA binding proteins that interact with an ethylene-responsive element. Plant Cell 7:173–182
Padma P, Chansouria JPN, Khosa RL (1997) Effect of alcohol extract of Annona muricata on cold immobilization stress induced tissue lipid peroxidation. Phytother Res 11:326–327. https://doi.org/10.1002/(SICI)1099-1573(199706)11:4<326::AID-PTR94>3.0.CO;2-B
Park C, Lim CW, Baek W, Lee SC (2015) RING type E3 ligase CaAIR1 in pepper acts in the regulation of ABA signaling and drought stress response. Plant Cell Physiol 56:1808–1819. https://doi.org/10.1093/pcp/pcv103
Park C, Lim CW, Lee SC (2016) The pepper ring-type E3 ligase, CaAIP1, functions as a positive regulator of drought and high salinity stress responses. Plant Cell Physiol 57:2202–2212. https://doi.org/10.1093/pcp/pcw139
Penella C, Landi M, Guidi L, Nebauer SG, Pellegrini E, San BA, Remorini D, Nali C, López-Galarza S, Calatayud A (2016) Salt-tolerant rootstock increases yield of pepper under salinity through maintenance of photosynthetic performance and sinks strength. J Plant Physiol 193:1–11. https://doi.org/10.1016/j.jplph.2016.02.007
Pimenta S, Menezes D, Neder DG, Melo RA (2016) Adaptability and stability of pepper hybrids under conventional and organic production systems. Hortic Bras 34:168–174. https://doi.org/10.1590/S0102-053620160000200004
Reddy AS, Ali GS, Celesnik H, Day IS (2011) Coping with stresses: roles of calcium-and calcium/calmodulin-regulated gene expression. Plant Cell 23:2010–2032. https://doi.org/10.1105/tpc.111.084988
Reichheld J, Vernoux T, Lardon F, Van Montagu M, Inze D, Wilrijk B (1999) Specific checkpoints regulate plant cell cycle progression in response to oxidative stress. Science 17:647–656. https://doi.org/10.1046/j.1365-313X.1999.00413.x
Rushton P, Somssich I, Ringler P, Shen Q (2010) WRKY transcription factors. Trends Plant Sci 15:247–258
Sánchez-Bel P, Egea I, Sánchez-Ballesta MT, Martinez-Madrid C, Fernandez-Garcia N, Romojaro F, Olmos E, Estrella E, Bolarín MC, Flores FB (2012) Understanding the mechanisms of chilling injury in bell pepper fruits using the proteomic approach. J Proteome 75:5463–5478. https://doi.org/10.1016/j.jprot.2012.06.029
Shen L, Liu Z, Yang S, Yang T, Liang J, Wen J, Liu Y, Li J, Shi L, Tang Q, Shi W, Hu J, Liu C, Zhang Y, Lin W, Wang R, Yu H, Mou S, Hussain A, Cheng W, Cai H, He L, Guan D, Wu Y, He S (2016) Pepper CabZIP63 acts as a positive regulator during Ralstonia solanacearum or high temperature-high humidity challenge in a positive feedback loop with CaWRKY40. J Exp Bot 67:2439–2451. https://doi.org/10.1093/jxb/erw069
Shinozaki K, Yamaguchi-Shinozaki K, Urao T, Koizumi M (1992) Nucleotide sequence of a gene from Arabidopsis thaliana encoding a MYB homologue. Plant Mol Biol 19:493–499
Shoji T, Hashimoto T (2015) Stress-induced expression of NICOTINE2-locus genes and their homologs encoding ethylene response factor transcription factors in tobacco. Phytochemistry 113:4–49. https://doi.org/10.1016/j.phytochem.2014.05.017
Singh K, Foley RC, Oñate-Sánchez L (2002) Transcription factors in plant defense and stress responses. Curr Opin Plant Biol 5:430–436. https://doi.org/10.1016/s1369-5266(02)00289-3
Solé C, Nadal-Ribelles M, de Nadal E, Posas F (2015) A novel role for lncRNAs in cell cycle control during stress adaptation. Curr Genet 61:299–308. https://doi.org/10.1007/s00294-014-0453-y
Wan H, Yuan W, Ruan M, Ye Q, Wang R, Li Z, Zhou G, Yao Z, Zhao J, Liu S (2011) Identification of reference genes for reverse transcription quantitative real-time PCR normalization in pepper (Capsicum annuum L.). Biochem Biophys Res Commun 416:24–30. https://doi.org/10.1016/j.bbrc.2011.10.105
Wang JE, Li DW, Zhang YL, Zhao Q, He YM, Gong ZH (2013a) Defence responses of pepper (Capsicum annuum L.) infected with incompatible and compatible strains of Phytophthora capsici. Eur J Plant Pathol 136:625–638. https://doi.org/10.1007/s10658-013-0193-8
Wang JE, Liu KK, Li DW, Zhang YL, Zhao Q, He YM, Gong ZH (2013b) A novel peroxidase CanPOD gene of pepper is involved in defense responses to Phytophtora capsici infection as well as abiotic stress tolerance. Int J Mol Sci 14:3158–3177. https://doi.org/10.3390/ijms14023158
Yeom S, Baek H, Oh S, Kang W, Lee SJ, Lee JM, Seo E, Rose JKC, Kim B, Choi D (2011) Use of a secretion trap screen in pepper following Phytophthora capsici infection reveals novel functions of secreted plant proteins in modulating cell death. Mol Plant-Microbe Interact 24:671–684. https://doi.org/10.1094/MPMI-08-10-0183
Yi SY, Kim J-H, Joung Y, Lee S, Kim W, Yu SH, Choi D (2004) The pepper transcription factor CaPF1 confers pathogen and freezing tolerance in Arabidopsis. Plant Physiol 136:2862–2874. https://doi.org/10.1104/pp.104.042903
Zhang H-X, Jin J-H, He Y-M, Lu B-Y, Li D-W, Chai W-G, Khan A, Gong Z-H (2016) Genome-wide identification and analysis of the SBP-box family genes under Phytophthora capsici stress in pepper (Capsicum annuum L.). Front Plant Sci 7:1–14. https://doi.org/10.3389/fpls.2016.00504
Zhou J, Liang Y, Niu Q, Chen L, Zhang X, Ye D (2013) The Arabidopsis general transcription factor TFIIB1 ( AtTFIIB1 ) is required for pollen tube growth and endosperm development. J Exp Bot 64:2205–2218. https://doi.org/10.1093/jxb/ert078
Zhou X-T, Jia L-J, Wang H-Y, Zhao P, Wang W-Y, Liu N et al (2018) The potato transcription factor StbZIP61 regulates dynamic biosynthesis of salicylic acid in defense against Phytophthora infestans infection. Plant J 95:1055–1068. https://doi.org/10.1111/tpj.14010
Funding
This work was supported through funding from the National Key R&D Program of China (No. 2016YFD0101900), the National Natural Science Foundation of China (No. U1603102), and the Independent Innovation Fund Project of Agricultural Science and Technology in Jiangsu (NO.CX (17) 3040).
Author information
Authors and Affiliations
Contributions
YMH, DXL, and ZHG conceived the research. YMH, KKL, HXZ, AK, GXC, and XM performed the research. YMH, HXZ, and MHA performed statistical analyses. YMH and AK wrote the paper. YMH, MHA, AK, and ZHG revised the paper. DXL and ZHG provided the materials and resources for the research. YMH, KKL, and ZHG performed the integrity of the work. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
He, YM., Luo, DX., Khan, A. et al. CanTF, a Novel Transcription Factor in Pepper, Is Involved in Resistance to Phytophthora capsici as well as Abiotic Stresses. Plant Mol Biol Rep 36, 776–789 (2018). https://doi.org/10.1007/s11105-018-1121-z
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11105-018-1121-z