Abstract
Rauvolfia serpentina has been known to produce therapeutically important indole alkaloids used in treatment of various diseases. Despite its medicinal importance, complete understanding of its secondary metabolism is challenging due to complex interplay among various transcription factors (TFs) and genes. However, weighted co-expression analysis of transcriptome along with integration of metabolomics data has proficiency to elucidate topological properties of complex regulatory interactions in secondary metabolism. We aimed to implement an integrative strategy using “-omics” data to identify TFs of “unknown function” and exemplify their role in regulation of valuable metabolites as well as metabolic traits. A total of 69 TFs were identified through significant thresholds and removal of false positives based on cis-regulatory motif analysis. Network-biology inspired analysis of co-expression network lead to generation of four statistically significant and biologically robust modules. Similar to known regulatory roles of WRKY and AP2-EREBP TF families in Catharanthus roseus, this study presented them to regulate synthesis of alkaloids in R. serpentina as well. Moreover, TFs in module 4 were observed to be regulating connecting steps between primary and secondary metabolic pathways in the synthesis of terpenoid indole alkaloids. Integration of metabolomics data further highlight the significance of module 1 since it was statistically predicted to be involved in synthesis of specialized metabolites, and associated genes may physically clustered on genome. Importantly, putative TFs in module 1 may modulate the major indole alkaloids synthesis in response to various environmental stimuli. The methodology implemented herein may provide a better reference to identify and explore functions of transcriptional regulators.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Gene expression is a complex phenomenon which is regulated by set of proteins, called transcription factors (TFs), that activate or repress various genes (Mitsuda and Ohme-Takagi 2009). In plants, TFs regulate genes involved in many important biological processes such as growth and development (Ramachandran et al. 1994), defense against pathogens and environmental stress (Singh et al. 2002), seed maturation and flower development (Jakoby et al. 2002), light-regulated mechanisms (Jiao et al. 2007), and secondary metabolism (Vom Endt et al. 2002). These TFs regulate genes by a set of highly coordinated internal/external signals, and some even interact with other TFs (Yang et al. 2012a). Secondary metabolites have long been used in pharmaceuticals, agrochemicals, fragrance ingredients, food additives, and pesticides, and are therefore of industrial importance. These metabolites do not participate directly in plant growth, development, and reproduction (Fraenkel 1959; Chae et al. 2014). However, they often play an important role in various processes including defense against pathogens, herbivores, and other interspecies defenses (Samuni-Blank et al. 2012). Also, these specialized metabolites and their associated pathways provide unique adaptive strategies for various organisms under harsh and dynamic environmental conditions (Weng and Noel 2012). Transcriptional regulation of secondary metabolite synthesis is highly controlled by a complex network of multiple TFs. Several TFs involved in the regulation of metabolic pathway genes have been studied in model plants like Arabidopsis thaliana (D’Auria and Gershenzon 2005). With advancements in developmental and molecular biology techniques for analyzing alkaloid biosynthesis, various genes have been identified to be involved in formation of metabolites like tropane (Shoji et al. 2000; Herbert 2003), benzylisoquinoline (Herbert 2003; Ziegler and Facchini 2008), and terpenoid indole alkaloids (TIAs) (Eichinger 1999; Ruppert et al. 2005). Since various studies have reported genes related to metabolite synthesis, further attempt should also be made to identify key regulators associated with their complex pathways (De Luca and St Pierre 2000; Oudin et al. 2007). Since gene expression is highly regulated by specific TFs, various biological functions get influenced by any modification in their activity, which in turn dynamically alters transcriptome profile leading to metabolic and/or phenotypic changes (Mitsuda and Ohme-Takagi 2009). Therefore, to elucidate molecular mechanisms associated to plant secondary metabolism, a prerequisite is to identify candidate regulatory TFs through a productive and precise searching procedure (Haynes et al. 2013).
Rauvolfia serpentina is an important medicinal plant of family Apocynaceae, commonly known as “Indian snakeroot,” endemic to Indian subcontinent and South-East Asian countries. It is also found in the Himalayan mountain ranges distributed over the foothills up to elevations of 1300–1400 m (Dey and De 2011). The major phytochemical constituents of this plant are indole alkaloids (Pathania et al. 2013). Its roots are also known to produce therapeutically important alkaloids which are used in the treatment of various diseases such as ventricular arrhythmias, hypertension, and diabetes (Vakil 1955; Locket 1955; Dey and De 2011; Azmi and Qureshi 2012). Reserpine is the principle component of R. serpentina used to treat hypertension (Hutcheon 1971; Nammi et al. 2005), tachycardia (Jerie 2007), and allergy (Lelek and Furedi Szabo 1961). Other compounds such as ajmaline (Bazika 1969; Köppel et al. 1989), serpentine (Beljanski and Beljanski 1982), rescinnamine (Hutcheon 1971; Nammi et al. 2005), and yohimbine (Singh et al. 2004) have also been reported in treatment of various diseases. Despite their broad pharmacological properties, limited information is available about regulatory factors involved in the synthesis of secondary metabolites (Chou and Kutchan 1998; Wink 2010) which further affect the yield and spectrum of such pharmacologically important compounds (De Luca and St Pierre 2000; Vom Endt et al. 2002). Catharanthus roseus has been considered as model plant to understand TIA pathway (Facchini and De Luca 2008; Pani and Mahapatra 2013), and TF families such as WRKY (Suttipanta et al 2011; Schluttenhofer et al 2014) and AP2EREBP have been reported to regulate metabolite synthesis (van der Fits and Memelink 2000; Li et al. 2013). Also, bHLH and MYB-like TFs are reported to be highly expressed under jasmonate response but needs to be confirmed for their regulatory role in TIA pathway (Memelink and Gantet 2007). Therefore, identification of transcriptional regulators and their target genes, involved in the synthesis of such metabolites, is crucial to understand associated metabolic pathways.
Advances in molecular and bioinformatics techniques have already made the availability of high-throughput biological data for various model plants. Fortunately, the large-scale transcriptomics data for various medicinally important plants, including R. serpentina, is also available at Medicinal Plant Genomics resource (MPGR, http://medicinalplantgenomics.msu.edu/) (Gongora-Castillo et al. 2012). Availability of this high-throughput data allowed us to conceive a hypothesis for complex regulatory insights in R. serpentina, specifically associated to secondary metabolism. Various traditional and recently developed methods are available to perform functional annotation of TFs (Mitsuda and Ohme-Takagi 2009), but as far as our knowledge, no reports are available for omics- and network-based analysis in R. serpentina. Integration of graph theoretical based methods and “-omics” data facilitates systems-level analysis of underlying molecular mechanisms and unravel various complex biological processes (Aoki et al. 2007; Mao et al. 2009; Naika et al. 2013). Moreover, network analyses using gene co-expression data has been proposed to explain biological systems since it models and decipher interactions of real biological networks (Aoki et al. 2007; Ma et al. 2013). Such analyses assist to identify functionally important regulators from a large set of genes with unknown function, and may provide information of associated pathways (Lee and Tzou 2009; Srivastava et al. 2010). Instead of performing analysis for tens of thousands of gene comparisons, a module-based analysis limits the analysis on only few orders of magnitudes for comparisons. The module represents a set of genes with very similar expression patterns that corresponds to specific pathway, therefore focusing on this amounts to a biologically meaningful data reduction scheme. Utility of such analyses has uncovered gene regulatory mechanisms based on their expression profiles (Hartwell et al. 1999; Ihmels et al. 2004). Hence, advances in network biology have made it possible to fill the gap of understanding association of individual genes by systematically investigating their association.
In this study, we have aimed to establish an integrative genome- and systems-wide framework to identify TFs, as well as to elucidate their complex regulatory behavior, associated with synthesis of secondary metabolites in R. serpentina. Gene co-expression network analysis was implemented to identify functionality of TFs, which were not annotated through simple annotation methods. Modules were obtained from co-expression network and enrichment analysis was performed to identify highly significant modules. Genes present in each of significant module was enriched with similar ontological terms and tissue-specific expression emphasizing their robustness. These modules were associated with hormone- or elicitor-mediated synthesis of indole alkaloids, epigenetic regulations, reproductive system, and upstream of TIA pathway, respectively. Interestingly, one module was found to regulate synthesis of specialized indole alkaloids known to be present in roots under hormone- or elicitor-mediated response, and do not have any shared metabolite with the remaining three modules. The identification of such regulatory factors may aid to improve quantity and diversity of secondary metabolites, and methodology presented herein can possibly assist their experimental studies. This graph theoretical approach may provide a better reference for other non-model plants to improve the established strategies for TF identification.
Materials and Methods
Toward our goal of identification of TFs and associated modules in secondary metabolism, the following procedures were implemented: (1) annotation of R. serpentina transcriptome, (2) calculation of Pearson correlation coefficient, (3) transcription factor binding site analysis, (4) construction of weighted co-expression network, (5) enrichment analysis and module detection, (6) reference-based enrichment of significant modules, and (7) integration of metabolomics data.
Annotation of R. serpentina Transcriptome
Transcriptomic sequences and expression profile data of various R. serpentina tissues were retrieved from the Medicinal Plant Genomics Resource database (MPGR, http://medicinalplantgenomics.msu.edu/) (Gongora-Castillo et al. 2012). All transcripts were annotated by performing BLASTX (Altschul et al. 1997) search against the reference Arabidopsis proteome (TAIR10, http://Arabidopsis.org). An e value cutoff of 1e-05 was considered to identify homologous genes, and annotations of associated top hits were preserved for further analysis. In order to identify TFs, transcripts with unknown primary annotation were also searched independently against Plant Transcription Factor Database (PlnTFDB; http://plntfdb.bio.uni-potsdam.de/v3.0/) (Riaño-Pachón et al. 2007) using BLASTX with the same specifications of hit selection. To describe the functional annotation of transcripts, gene ontology (GO) (Dimmer et al 2008) analysis for all three biological domains (biological processes (BP), molecular functions (MF), and cellular components (CC)) was performed. Annot8r (Schmid and Blaxter 2008) was used to obtain GO and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway annotation for assembled transcripts based on similarity search against annotated subsets of the UniProt database (The UniProt Consortium 2007). This annotation was performed with an e value threshold of 10-01, and top hits were retained based on highest significant e value as well as bit score.
Calculation of Pearson Correlation Coefficient
In order to construct TF-gene regulatory network, Pearson correlation coefficient (PCC) (Eisen et al. 1998) between identified TFs and all transcripts were calculated from expression data in the form of FPKM (fragments per kilobase of transcript per million fragments mapped), available for different tissues (mature leaf, young leaves, upper stem, young roots, mature roots, red stem, flower, and woody stem), using customized in-house written PERL script. PERL code has been provided at GitHub (https://github.com/ShivalikaP/Perl-script-to-calculate-Pearson-correlation-coefficient). PCC was computed based on the following equation:
Where \( x \) and \( y \) denotes the corresponding expression profiles of TF and transcripts, respectively, and \( \overline{x} \) and \( \overline{y} \) represents their mean. Mean of expression profile depends upon sample size which is different for both TFs and transcripts.
Network density (ND) helps in determining biologically important PCC threshold (Aoki et al. 2007). ND was measured as follows:
Where \( E \) and \( \frac{K\left(K-1\right)}{2} \) specify the number of actual links and possible links of non-singleton nodes, respectively.
The “t test” was implemented to select significant TF-gene pairs with p value ≤0.05 (McCluskey and Lalkhen 2007), and further subjected to multiple testing correction using false discovery rate (FDR) (Benjamini and Hochberg 1995) method through “stats” library of R statistical package (http://www.r-project.org/). Shortlisted pairs were compared against TF-gene connectivity data (Misra and Sriram 2013), downloaded from the Arabidopsis Gene Regulatory Information Server (AGRIS; http://Arabidopsis.med.ohio-state.edu/) (Davuluri et al. 2003), to verify their regulatory relationship based on prior knowledge available for corresponding orthologs in A. thaliana.
Transcription Factor Binding Site Analysis
Transcription factor binding sites (TFBS) were identified using pPromotif tool (Jha and Shankar 2014) for those TF-gene pairs which did not have any match to already verified TF-gene connectivity data. In order to identify TFBS, 5 kb upstream region of orthologous Arabidopsis genes of corresponding transcripts were retrieved from BioMart of Ensembl Plants database (http://plants.ensembl.org/biomart/martview) (Kinsella et al. 2011). These upstream regions were further analyzed to identify TFBS based on a position weight matrix generated using the Gibbs Motif Sampler. Furthermore, pairs were selected based on comparison of TF families assigned to TFs against all possible binding sites obtained. These selected pairs were further complemented by manual literature search. Accuracy of identified TFBS was statistically validated using Z-score for orthologous Arabidopsis genes. For calculating Z-score, pPromotif was implemented on 5 kb upstream sequences of Arabidopsis (as null model/control), and mean as well as standard deviation was calculated from score obtained from TFBS hits of 13 TF families. Z-score (Sundar et al. 2008) was computed using the following equation:
Where x is the pPromotif score for each TF hit obtained for orthologous Arabidopsis genes. Mean and SD are the average and standard deviation of pPromotif scores, respectively, for all control genes of 13 TF families.
A two-tailed p value statistics, corresponding to Z-score, was further calculated using “stats” library of R package.
Construction of Weighted Co-expression Network
Weighted co-expression network was constructed from TF-gene pairs obtained during successive filtering steps. An assessment-based co-expression network was attained since edges connecting nodes at significant PCC threshold and p values were kept. Cytoscape v 3.1.0 (Shannon et al 2003) program was used for network construction and visualization purpose. TFs obtained in resultant co-expression network were classified into different families based on BLAST search against PlnTFDB. PCC values were used as edge weights to construct network, where weights represent the strength of connections or co-expression. Random network with same number of nodes and edges, as of weighted co-expression network, was generated using “igraph” (Csardi and Nepusz 2006) library in R package. Network topology parameters of both networks were determined to compare their topological properties.
Enrichment Analysis and Module Detection
Enrichment analysis of all transcripts corresponding to each TF family was carried out with singular enrichment analysis (SEA) of agriGO web-based tool (http://bioinfo.cau.edu.cn/agriGO/index.php) (Du et al. 2010), and significantly enriched GO terms were determined in all comparative conditions by comparing it against background reference. Hypergeometric test (Shimizu et al. 2014) with Bonferroni correction (Simes 1986) was applied for selecting these statistically significant terms. In search of comparatively smaller and functionally important gene modules, Markov cluster (MCL) (Van Dongen 2008) algorithm, a systematic graph clustering method, was used to fragmentize the network. This algorithm is based on the “random walks” which states that starting with a node randomly travel to connected nodes will make to stay within a cluster than travel between, i.e., flow is easier within dense regions than across sparse boundaries. MCL algorithm, with default inflation value of 2.5, was implemented using clusterMaker2 (Morris et al. 2011) plugin of Cytoscape. Enrichment analysis of all modules was also performed independently. Expression data was used to generate heat maps using “gplots” library of R package. “Vennerable” library from same package was used to obtain venn diagram of common target genes among TFs in each significant module.
Reference-Based Enrichment of Significant Modules
In order to perform reference-based enrichment analysis, clueGO (Bindea et al. 2009) plugin of Cytoscape was used against model plant A. thaliana. Enrichment analysis was performed using a two-sided (enrichment/depletion) tests based on hypergeometric distribution with Bonferroni (Simes 1986) p value correction method. Besides functional enrichment, it also creates an annotation network of functionally grouped non-redundant biological pathways associated with enriched GO terms for visualization. Kappa statistics (Sim and Wright 2005) is used by clueGO to link similar terms in a network that provides a measure of accordance from a range between 0 and 1, where 0 and 1 indicates no and almost perfect accordance, respectively. In this analysis, a kappaScore threshold of 0.4 was considered.
Integration of Metabolomics Data
To assess quality of significant modules and to determine the involvement of TFs in regulation of secondary metabolism, metabolome data of R. serpentina was included. Correlation data of metabolome, available at the Plant and Microbial Metabolomics Resource (PMR; http://metnetdb.org/PMR/), of corresponding transcriptome was obtained using PMR metabolomic-transcriptomic co-analysis tool at selected PCC threshold and significant p value (≤0.05). TFs present in significant modules were searched against correlation data to determine module-specific regulation of metabolite synthesis. “Vennerable” library was used to obtain venn diagram of shared metabolites in different modules. Significance of modules associated with synthesis of specialized metabolites was also statistically verified using “Mann-Whitney test” from “stats” library of R package.
Results
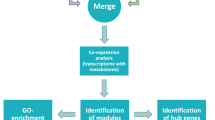
An integrative approach was established to identify TFs and their role in regulation of valuable metabolites and metabolic traits (Fig. 1) in R. serpentina. Initially, TFs from the transcripts of unknown function were identified which were further analyzed to determine their complex regulatory mechanism using graph theoretical approach.
Identification of Transcription Factors
A. thaliana has been widely used as a reference model for annotation of various plants including C. roseus, a member of Apocynaceae family (Schluttenhofer et al. 2014). Similarly, annotation of complete R. serpentina transcripts was also carried out against Arabidopsis proteome that returned 80,829 significant hits. A total of 12,670 transcript entries without primary annotation were retrieved, and 342 hits were considered as putative TFs while annotating against PlnTFDB.
GO- and KEGG-based functional analysis was performed to functionally annotate complete transcriptome using annot8r, and output has been provided as separate spreadsheets at GitHub (https://github.com/ShivalikaP/R_serpentina_transcriptome_annotation). Annot8r has been widely used to functionally annotate transcriptomes of various plants including Citrus sinensis (Yang et al. 2013) and Euphorbia fischeriana (Barrero et al. 2011). This analysis returned functional annotation for ∼71 and ∼43 % transcripts covering a broad range of GO and KEGG categories, respectively, of R. serpentina transcriptome. The most abundant BP categories were metabolic process (∼26 %), response to stimulus (∼14 %), and regulation of biological process (∼13 %) revealing that the majority of transcripts were participating in metabolic activities under response to stimulus (Fig. S1a). Most abundant GO categories such as binding (∼44 %), transferase activity (∼15 %), and catalytic activity (∼13 %) terms were determined under MF category (Fig. S1b). Similarly, major GO categories identified under CC categories included intracellular (∼68 %) and membrane (∼27 %) (Fig. S2a) terms. A total of 260 KEGG pathways were retrieved for whole trancriptome sequences, and the top 20 of them were associated with plant hormone signal transduction (∼5 %), plant-pathogen interaction (∼4 %), ribosome (∼3 %), and spliceosome (∼3 %) (Fig. S2b).
Threshold Selection Based on Network Topology
Since simple annotation was unable to annotate biological processes associated with 190 TFs (Table S1), these TFs were annotated based on their interacting transcripts. In order to determine congruity of putative TFs and associated transcripts, expression profiles among eight different tissues were used to compute PCC. This number of samples is sufficient to carry out co-expression analysis as evident from various studies to identify new candidate genes (Gongora-Castillo et al. 2012; Paul et al. 2014; Schluttenhofer et al. 2014). PCC is used to construct a correlation matrix of gene expression from which weighted co-expression network is generated (Eisen et al. 1998). A total of 4,504,588 TF-gene pairs were obtained for unique 342 TFs and 19,333 transcripts, and pairs with positive correlation were retained to analyze biological processes up-regulated by these TFs. ND demonstrates how sparse or dense a graph is according to the number of connections per node set, and has been used for choosing a statistically relevant threshold since at this cutoff, biologically significant modules are expected (Aoki et al. 2007; Mao et al. 2009). To characterize the PCC threshold, change in number of nodes, edges, and network density, as a function of various PCC cutoff values, were examined. As the cutoff value increased, node and edge number, as well as the actual number of edges and all possible edges were decreased. However, as the cutoff reached at a relatively high value, decreasing rate of edges became slower than that of all possible edges (Fig. S3a) and nodes (Fig. S3b), which leads to an increase in ND. ND was at its minimum at 0.84 cutoff (Fig. 2a), and PCC corresponding to this minimal ND was considered as threshold. TF-gene pairs at and above this threshold were analyzed to obtain biologically significant modules. Conversely, threshold was adjusted at 0.80, instead of 0.84, in order to retain a relatively large number of connections and prevent the loss of significant information in network construction. Although, PCC can be used to measure the congruity, yet it is difficult to determine its statistical significance more than expected by chance. All TF-gene pairs (151,504) at and above the threshold of 0.80 were found to be statistically significant (p value ≤0.05) (Table S2), and the number of these pairs were further reduced to 82,635 based on availability of Arabidopsis orthologs for corresponding transcript of R. serpentina in TF-gene connectivity data from AGRIS (Davuluri et al 2003). Regulatory information of these TF-gene pairs was further complemented by comparing them against this TF-gene connectivity data, and 7066 connections among all these pairs were retained, whereas the remaining 75,569 were discarded due to unavailability of the TF-gene interactions. This selection criterion helps to strengthen TF-gene pairs regularity information based on prior validated information that ensure minimum number of false positives.
Identification of Transcription Factor Binding Sites
Since PCC of discarded TF-gene pairs were found to be statistically significant, transcripts were analyzed for possible TFBS to match coinciding TF in each pair. Promoter sequences are often conserved between diverse plant species (Fauteux and Strömvik 2009); (Koch et al. 2001) that allow the use of A. thaliana as reference to perform TFBS analysis. Upstream sequences of orthologous Arabidopsis genes, corresponding to 9676 transcripts, were processed to identify TFBS. A total of 13,439 TF-gene pairs were selected by comparing them against pPromotif output, and the rest were discarded in further studies. In order to predict accuracy of TFBS prediction, hits obtained for orthologous Arabidopsis genes were compared against TFs from AGRIS (control) to compute Z-score. Z-score explains the direction as well as measure of deviation of number of TFBS occurrences from background distribution’s mean. Null hypothesis states that there was no significant difference in scores between orthologous Arabidopsis genes and control. It was found that for ABI3VP1, 10,829 out of 10,829 (100 %) number of sites had insignificant p value (>0.05). Similarly, the percentage of hits obtained for other TF families such as AP2EREBP, ARF, bHLH, bZIP, CCAAT, C2C2-GATA, HB, MYB, MYB-related, MADS, SBP, and WRKY were 93.59 % (9385/10,028), 79.24 % (8067/10,180), 100 % (10,758/10,758), 96.11 % (10,408/10,829), 99.88 % (10,816/10,829), 99.90 % (10,819/10,829), 94.99 % (6632/6982), 92.85 % (10,055/10,829), 98.29 % (10,644/10,829), 96.63 % (9651/9987), 100 % (1087/1087), and 94.78 % (10,132/10,690), respectively. Since p value was found to be insignificant, null hypothesis was accepted which further signifies the accuracy of TFBS prediction. These selected pairs were further augmented by critical manual literature search for TFs regulating either corresponding orthologous Arabidopsis genes or orthologs of orthologous Arabidopsis genes in some other plant species. The search was carried out for 290 TF-gene pairs randomly selected at different cutoffs in order to cover a wide range of PCC (0.8–1.0). Interestingly, ∼94 % of TF-gene pairs were observed to be supported in different literature studies (Table S3) which further enhanced the confidence of output obtained. Importantly, this analysis enhances number of connections by identifying TF-gene pair regulatory relationship left in earlier step which was supported by expression data.
Construction of Weighted Co-expression Network
Weighted co-expression network was constructed for deciphering systematic involvement of identified TFs in regulation of various biological processes. Such co-expression networks often utilizes measured PCC as weights, and has been widely used to determine biologically important modules in a variety of organisms (Aoki et al. 2007; Mao et al. 2009; Ruan et al. 2010; Gongora-Castillo et al. 2012; Fukushima et al. 2012). Co-expression network constituted of nodes and edges representing genes and connections between them, respectively. The network had 7304 nodes (including unique 69 TFs and 7235 target genes) with 20,505 edges. All identified 69 TF genes were classified into 13 TF families (Table S4, Fig. 3) based on the BLAST search against PlnTFDB. PCC values obtained in former step were used as edge attribute where these numeric attributes quantifies strength of connections among nodes, which again enhanced the method of module detection. Real networks are often compared with random networks to decipher scale-free nature of networks and to distinguish their topological properties (Alon 2006; Raval and Ray 2013). A weighted random network was constructed with same number of nodes, as of actual co-expression network, by creating random connection among edges for 2000 iterations. Compared to bell-shaped distribution of random network (Fig. 2b), the degree distribution (DD) of co-expression network was highly skewed indicating that majority of nodes had a degree, close to average degree in random graphs and thereby satisfies properties of biological networks (Albert 2005). DD of co-expression network also satisfied the power law (Fig. S4) since there was no peak around average degree. Graph rather sloped steeply downwards with increase in degree demonstrating the presence of a large number of nodes with very few connections dominated by some extremely well-connected ones. This revealed that co-expression network had scale-free behavior and robust against random perturbations since such networks are most likely to hit a node with only few neighbors, and therefore disrupt only a small portion (Emmert-streib and Dehmer 2008). Negative value of assortativity (−0.68) also augmented the scale-free nature of co-expression network as compared to random network (0.0014) (Johnson et al 2010).
Functional Annotation and Enrichment (Over-Representation) Analysis
GO terms of target transcripts, corresponding to each putative TF family, were used as background to perform enrichment analysis through agriGO. Analysis of each TF family revealed that several transcripts were observed to be significantly over-represented (p value ≤0.05) to participate in more than one biological process. TF families such as ABI3VP1, bHLH, HB, MYB, MYB-related, and WRKY (Figs. S5–S10) were significantly enriched with terms related to plant reproductive system. Major significant terms obtained among these TFs included floral whorl development, stamen development, floral organ development, pollination, pollen tube growth, and androecium development. TF families (ABI3VP1, AP2-EREBP, MYB-related, and WRKY) (Figs. S5, S11, S9, and S10) were enriched for terms such as response to hormone stimulus, response to endogenous stimulus, response to organic substance, and response to defense response presented their direct involvement in response to environmental conditions. HB (Fig. S7) and bZIP (Fig. S12) were observed to be significantly enriched with terms related to cell cycle such as cell cycle process and microtubule-based process, while cell division AP2-EREBP (Fig. S11) and bHLH (Fig. S6) were significantly over-represented with terms photosynthesis and photosynthesis, light reaction. Interestingly, some TF families were perceived to be enriched with terms associated with specific process such as ABI3VP1 (Fig. S5) (polarity specification of adaxial/abaxial axis and specification of axis polarity) to axis or pattern formation, AP2-EREBP (Fig. S11) (pentose-phosphate shunt, NADPH regeneration, and NADP metabolic process) to primary metabolic pathway as well as upstream of TIA pathway (Dubey et al. 2003), HB (Fig. S7) (chloroplast RNA processing, DNA alkylation, and DNA methylation) to post-translational modifications via ubiquitin-dependent proteolytic pathway, and MADS (Fig. S13) (multicellular organismal development) to organ development. TF families ARF, CCAAT, C2C2-GATA, and SBP were not found to be enriched with any significant term.
Detection of Functional Modules
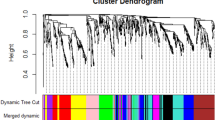
Gene-enrichment analysis demonstrated that multiple biological pathways were under regulation of one TF family that did not specify pathway regulated by each TF of given family. Therefore, module-based analysis was performed to identify the TFs that may function in an integrative manner to regulate specific pathway. The identification of modules is a better choice to understand the coordinated behavior of TFs of different families since each module occupies certain set of genes shared by TFs present in it. Top-down approach (or non-targeted approach) (Aoki et al. 2007) can be applied to fragmentize network into modular structures through graph clustering algorithms based on the topology of connections (Mao et al. 2009). MCL algorithm has successfully been implemented for module detection in many biological networks (Mao et al. 2009). It sub-divided the large network, to simplify the analysis of weighted co-expression network, into 42 modules (Fig. S14) ranging from 1251 (largest) to 4 (smallest) nodes at default inflation value. These modules were referred to by their number henceforth, as a standard practice. The first five modules (1–5) were largest in size with 1,251, 995, 762, 750, and 417 nodes including 9, 5, 7, 5, and 4 regulatory TF genes, respectively. Modules from 6–10, 11–16, 17–28, 29–37, and 38–42 fragmented in ranges from 201 to 290, 101 to 150, 51 to 100, 20 to 45, and 4 to 15 nodes, respectively, whereas modules 6–42, except 7 and 18 (2 TFs), had only 1 TF.
Enrichment Analysis of Modules
Enrichment analysis was performed for all 42 modules, and only few (1–4, 6, 8–11, 13–15, 20, 21) were characterized with at least one significant enriched GO term. The first four modules (Fig. 4) were considered as significant modules since each of them presented maximum number of enriched terms with similar biological processes, and implicated in downstream analysis. A number of earlier reports have presented considering only significant modules (Childs et al. 2011; Chou et al. 2014). Transcript identifiers of all TF families in significant modules are provided in Table 1. Most transcripts present in each module were enriched with similar ontological terms indicating the robustness of modules. GO and KEGG pathways annotation for all transcripts present in these significant modules (1–4) is provided in Tables S5–S8. Details of significant modules are provided as follows:
-
Module 1
Module 1 was associated with defense response and indole alkaloid synthesis (Fig. 5) as majority of significant terms included response to stimulus (GO:0050896; 1.02e-08), response to endogenous stimulus (GO:0009719; 7.45e-07), defense response (GO:0006952; 2.52e-06), abscisic acid-mediated signaling pathway (GO:0009738; 0.00306), indole derivative biosynthetic process (GO:0042435; 0.0208), indole derivative metabolic process (GO:0042434; 0.0239), and response to jasmonic acid stimulus (GO:0009753; 0.0427). Intentness of abscisic acid and jasmonate-mediated signaling response for major indole alkaloids synthesis has already been reported in R. verticillata (Fernandez and de Luca 1994) and C. roseus (Schluttenhofer et al. 2014), closely related plants to R. serpentina, respectively. It has already been reported that diverse metabolic pathways are induced under various stress responses (Naika et al. 2013). Therefore, all these major ontological terms may reveal possible role of module 1 in defense response (Wei et al. 2014; Zarate et al. 2007) and indole alkaloid synthesis (Fernandez and de Luca 1994; Schluttenhofer et al. 2014). Also, TF families (ABI3VP1, AP2-EREBP, MYB-related, SBP, and WRKY) (Fig. 6a) and their targets (Fig. S15a) had comparatively higher expression in roots further complementing earlier reports of synthesis of major alkaloids in roots. Most significant pathways, like plant-pathogen interaction and plant hormone signal transduction, were also found to be associated with defense response and response to stimulus (Fig. S16a).
-
Module 2
Module 2 was observed to be involved in epigenetic regulations as majority of enriched terms (Fig. S17) were related to chloroplast RNA processing (GO:0031425; 5.85e-07), cell cycle process (GO:0022402; 0.000235), embryonic development ending in seed dormancy (GO:0009793; 0.00182), fruit development (GO:0010154; 0.00237), fruit development (GO:0010154; 0.00237), seed development (GO:0048316; 0.00158), and post-embryonic development (GO:0009791; 0.0247). Most significant chloroplast RNA processing term present an important regulatory step of post-translational modification via ubiquitin-dependent proteolytic pathway, and involves in stabilization of primary chloroplast transcripts (Schuster and Gruissem 1991). This ontology term also emphasized on localization of upstream of TIA pathway in chloroplast (Ershov et al. 2002; Flügge and Gao 2005; Herrmann and Weaver 1999; Schmid and Amrhein 1995). TIA pathway (Fernandez and de Luca 1994; Shukla et al. 2006) and jasmonic acid-mediated activation of ORCA (AP2-like) TF (Vom Endt et al. 2002) have already been reported to be regulated by post-translational modifications under ubiquitin-dependent proteolysis. Also, TF families (ABI3VP1, bHLH, HB, and AP2-EREBP) presented were found to be highly expressed in young leaves (Fig. 6b), and may necessitate chloroplast localized synthesis of precursors under epigenetic regulation. Interestingly, enrichment analysis of HB TF family was also enriched with few similar enriched terms signified its involvement in epigenetic regulations. Moreover, HB TF present in flowering wageningen (FWA) gene has already been reported to be involved in RNA-directed DNA methylation in A. thaliana (Matzke et al. 2007). This highlighted the involvement of HB TF family in epigenetic regulations and may contribute to growth, development, and consequently plant evolution (Grafi and Ohad 2013). Significance of module was also augmented by the involvement of cell cycle process (Perianez-Rodriguez et al. 2014), chromatin modifications/remodeling in seed development (Baroux et al. 2007), embryonic development ending in seed dormancy (Ríos et al. 2014), and post-embryonic development (de Vega-Bartol et al. 2013; Perianez-Rodriguez et al. 2014). Similarly, most significant pathways for this module comprised of processes such as cell cycle, spliceosome, mismatch repair, and RNA degradation complementing GO terms assigned, and its involvement in epigenetic regulations (Fig. S16b).
-
Module 3
Module 3 was associated with reproductive system since significantly enriched terms (Fig. S18) were found to be related to reproductive process (GO:0022414; 2.18e-06), reproduction (GO:0000003; 4.3e-06), pollen tube growth (GO:0009860; 2.4e-05), stamen development (GO:0048443; 3.73e-05), androecium development (GO:0048466; 3.73e-05), and pollination (GO:0009856; 4.18e-05). TF families ABI3VP1, bHLH, MYB, MYB-related, SBP, and WRKY were found to be expressed mainly in flower (Fig. 6c) which further complement the enrichment results and highlights robustness of module. Pathway annotation of module was significantly over-represented with terms like flavonoid biosynthesis and phenylalanine metabolism (Fig. S16c) that may be associated to synthesis of anthocyanins and thereby represent its involvement in pollination (Mol et al. 1998).
-
Module 4
Module 4 was related to primary metabolic pathway, and upstream of TIA pathway as significant GO terms (Fig. S19) presented were plastid localization (GO:0051644; 1.45e-08), chloroplast relocation (GO:0009902; 1.45e-08), photosynthesis, light reaction (GO:0019684; 3.72e-07), pentose-phosphate shunt (GO:0006098; 1.23e-06), NADPH regeneration (GO:0006740; 1.28e-06), NADPH metabolic process (GO:0006739; 1.76e-06), oxidoreduction coenzyme metabolic process (GO:0006733; 3.04e-05), and generation of precursor metabolites and energy (GO:0006091; 0.000863). Pentose-phosphate cycle is known to be a direct precursor for isoprenoid synthesis (Ershov et al. 2002; Flügge and Gao 2005) and shikimate pathway that are involved in the release of various aromatic amino acids localized inside the chloroplast (Herrmann and Weaver 1999). Similarly, isoprenoid biosynthesis utilizes NADPH which is a byproduct of methylerythritol phosphate (MEP) pathway localized in chloroplast/plastid (Dubey et al. 2003; Seemann et al. 2006). Since upstream of TIA pathway utilizes NADPH, the presence of significantly enriched terms together like NADPH regeneration, NADPH metabolic process, plastid localization, and chloroplast relocation intimated the involvement of this module in synthesis of precursor metabolites through MEP pathway (El-Sayed and Verpoorte 2007). Enrichment analysis of AP2-EREBP TF family was also enriched with similar terms like pentose-phosphate shunt (GO:0006098; 0.0179), NADPH regeneration (GO:0006740; 0.0186), NADP metabolic process (GO:0006739; 0.0251), photosynthesis (GO:0015979), and photosynthesis, light reaction (GO:0019684) (Fig. S11). Moreover, AP2EREBP TF family has also been reported to regulate primary and secondary metabolism in C. roseus (Van der Fits and Memelink 2000) that may possibly regulate identical processes in module 2. Other co-expressed TF families (bHLH, WRKY, and SBP) may correspond to unknown TFs, as revealed in the study by Oudin et al. (2007), regulating upstream of TIA pathway. Tissue-specific expression of these TF families in mature leaves also (Fig. 6d) emphasized chloroplast localized regulation of precursor synthesis. Similarly, most significant pathways such as photosynthesis and glycolysis/gluconeogenesis (Fig. S16d) also augmented its involvement in regulating primary metabolic pathways as well as synthesis of precursors in upstream of TIA pathway (Aniszewski 2007; Glenn et al. 2013). It has also been stated earlier that the initial committed steps in secondary metabolite synthesis are recruited from primary metabolic pathways (Aharoni and Galili 2011; Chu et al. 2011). Interestingly, these results revealed that this module may act as connecting link between primary and secondary metabolic pathways, and TFs presented may regulate connecting steps between them.
Putative TFs have been found to work in a highly coordinated manner to perform important biological functions since 38 % of them were highly clustered only in the first four modules. A total of 39, 123, 50, and 2 common transcripts (Fig. 7), among all TF families, were obtained for significant modules (1–4), respectively, and were associated with representative pathways. This analysis represented that TFs do not perform their functions independently but in a highly coordinated manner as composite/co-regulatory modules (CRMs) where more than one TF families work together to facilitate regulation of gene expression (Jha et al. 2011; Matys et al. 2006). This further indicated the interdependence and complex coordinated behavior among all TF families to regulate specific biological pathways.
Enrichment Analysis Against Reference Model
Networks obtained from reference-based enrichment consisted of significant GO terms as nodes and shared genes among these terms as edges. In all significant modules, various biological pathways were distinctly clustered by iterative merging of initially defined groups based on kappa score threshold. A total of 605 transcripts were enriched in module 1 with majority of significant terms being hormone-mediated signaling pathway, defense response, and secondary metabolic process (Fig. S20). All these terms were similar to agriGO enrichment which further highlights its involvement in indole alkaloids synthesis (Chang et al. 2013; Schluttenhofer et al. 2014). Similarly, a total of 472, 242, and 389 transcripts in module 2, 3, and 4, respectively, were over-represented with significant GO terms. In module 2 (Fig. S21), significant terms like chromatin organization and RNA processing represented its participation in epigenetic regulations that may be associated with TIA pathway (Fernandez and de Luca 1994; Shukla et al. 2006; Siberil 2002; Vom Endt et al. 2002). In module 3 (Fig. S22), pathways represented by significant GO terms such as external encapsulating structure organization, stamen development, and pollen exine formation were also related to reproductive system. Similarly, module 4 was represented with significant terms such as photosynthesis, thylakoid membrane organization, and plastid organization (Fig. S23). Conclusively, this reference-based enrichment analysis also complements agriGO enrichments, and thereby highlighting robustness and quality of significant modules.
Module-Specific Regulation of Secondary Metabolites
Integration of transcriptome and metabolome data has been successfully implemented to determine TFs involved in regulation of specific metabolic pathways in the earlier reports (Hirai et al. 2005; Yonekura-Sakakibara et al. 2008). TFs present in significant modules were compared against correlation data (40 metabolites), and significant modules (1–4) were found to regulate 43.59, 17.94, 17.94, and 7.69 %, respectively, of all secondary metabolites (39) except tetraphyllicine. Module 1 (Table 2) was observed to regulate major indole alkaloids known to be synthesized in roots, and did not even share any metabolite with the rest of the significant modules (Fig. 8). Interestingly, major root indole alkaloids such as ajmaline, ajmalicine, reserpine, sarpagine, and serpentine (O’Connor and Maresh 2006) were present only in module 1. Therefore, transcripts in module 1 may indicate a complex interaction among them to regulate synthesis of these important indole alkaloids. Similarly, our findings were also supported by an earlier study that presented hormonal- or elicitor-mediated signaling pathways (e.g., WRKY TF family) to regulate major indole alkaloids, such as ajmalicine and serpentine, in C. roseus (Schluttenhofer et al. 2014). All these important findings, along with specifically high expression of module 1 transcripts in roots (Fig. S15a), determined the quality of identified modules as well as the protocol implemented for identification of key regulators. Modules 2–4 (Fig. 8, Table S9) shared few common as well as similar isomers of precursor metabolites (tryptamine, vellosimine, strictosidine, kaempferol hexoside dirhamnoside isomer 2, and kaempferol hexoside dirhamnoside isomer 1) that may reveal their intra- and inter-cellular trafficking of biosynthetic intermediates throughout the course of associated biosynthetic pathways (Kutchan 2005; O’Connor and Maresh 2006). Based on functional annotation and enrichment analysis, these modules were represented with significant representative processes like epigenetic regulations, reproductive system, and precursor’s biosynthesis with majority of expression in young roots, young leaves, flower, and mature leaves, respectively (Fig. S15b–d). Modules 2 and 4 were found to be expressed in leaves (young and mature, respectively), and may be grouped according to their representative functions viz., epigenetic regulations and photosynthesis associated synthesis of precursors. Module 3 was found to be regulating metabolites, mainly flavonoids and its derivatives, involved in pollination which also complemented enrichment results (Mol et al. 1998; Borevitz 2000). Importantly, integration of metabolomics and transcriptomics data had strengthened the reliability of significant modules due to module-specific regulation of various secondary metabolites (Fig. 8, Tables 2 and S9).
Significance of Gene Clustering in Module 1
Module 1 was observed to be involved in the synthesis of ajmaline/sarpagine-type alkaloids (O’Connor and Maresh 2006) of the last few steps of TIA pathway, under response of hormone- or elicitor-mediated signals (Leménager et al. 2005). Genes regulating synthesis of specialized molecules, under various environmental stresses, have been reported to physically clustered (Ehrlich and Raven 1964; Nützmann and Osbourn 2014) and represents biased clustering against non-specialized metabolites (Chae et al. 2014). In order to statistically verify biased nature of modules related to synthesis of specialized metabolites, Mann-Whitney test was implemented on co-expression values of all modules which had at least one specific alkaloid expressed only in roots (18) against remaining modules (24). Null hypothesis states that there was no significant preference of metabolic pathways genes in modules of specialized metabolites as compared to remaining modules. Mean co-expression value of specialized metabolites containing modules (0.88) was significantly higher than the remaining modules (0.87), and it was found that modules of specialized metabolites had significant preference for their synthesis over non-specialized metabolites (p value = 4.651e-16). It has been concluded that module 1 is statistically found to be most significant, and mainly under the control of diverse environmental conditions associated with specialized metabolites synthesis. Similarly, TFs present in this module may contribute into the diversity of major indole alkaloids in R. serpentina in response to various internal/external stimuli.
Discussion
R. serpentina has been widely used in Ayurvedic medicinal preparations and tribal ethnomedical systems for thousands of years (Dey and De 2011), and roots of this plant are well known to produce medicinally important alkaloids used in treatment of various diseases (Azmi and Qureshi 2012; Dey and De 2011; Locket 1955; Vakil 1955). In a previous study, we have been able to computationally identify inhibitors against aldose reductase from a repository of natural compounds compiled from the same plant (Pathania et al. 2013). Despite its medicinal importance, few studies have reported regulation of secondary metabolite synthesis in R. serpentina (W. M. Chou and Kutchan 1998; Wink 2010). The limited information of regulators that control the production of these molecules inhibit biotechnological application which may affect yield and spectrum of these pharmacologically important compounds (De Luca and St Pierre 2000; Vom Endt et al. 2002). Therefore, identification of transcriptional regulators as well as their target genes is crucial to understand metabolic pathways that lead to synthesis of such molecules. In recent years, gene co-expression networks have been used to unravel important biological processes and their regulatory mechanisms. There has been a growing trend in using large-scale protein-protein interaction (PPI) networks in plant systems (Yang et al. 2012b); however, the datasets cover only a limited range of PPI data. Furthermore, PPI networks are not static but dynamic entity and expression of protein is intrinsically controlled by different regulatory mechanisms (Liang and Li 2007). With an exponential growth of gene expression data, most of the current research has been shifted toward systems biology that largely include co-expression networks of transcriptome data (Yonekura-Sakakibara et al. 2008; Zhang et al. 2012).
Identification and annotation of TFs at genome scale are the first step toward understanding the mechanism of gene expression and its regulation. Though various genome-wide experimental studies have reported candidate TFs hitherto (Hu et al. 2014; Wang et al. 2010; Zhu et al. 2012), not much systems-level work has been done to assimilate the data. With increase in high-throughput proteomic/genomic studies and computational facilities, identification and annotation of TFs at genome scale are now being used to understand the regulation of biological mechanism (He et al. 2010; Pani and Mahapatra 2013). Although TFs are initially identified by computational approaches, they have also been validated by experimental approaches (Van Dijk et al. 2010) presenting the strength of computational protocols. In this study, an integrative approach has been implemented that involves computational identification of TFs, and their regulatory mechanisms in tissue-specific synthesis of secondary metabolite in R. serpentina. Co-expression network was constructed, with relatively stringent PCC threshold, from statistically significant TF-gene pairs. It has been augmented that genes of interest are co-expressed if their PCC is more than 0.80 and is generally preferred (Chien et al. 2014). However, choosing a strict threshold may lead to loss of significant gene pairs. Therefore, ND offers a reliable measure for choosing calibrated thresholds since at this cutoff, biologically significant modules are expected to be found (Aoki et al. 2007; Mao et al. 2009). Moreover, our approach includes identification of possible TFBS that enhanced the quality and enlarged the dataset used for network analysis. These stringent criteria were used to reduce weak connections followed by false positives among TF-gene pairs by comparing against TF-gene connectivity data and TFBS.
Gene co-expression network analysis has been successfully applied for identifying functional clusters of highly correlated genes in a variety of organisms (Aoki et al. 2007; Mao et al. 2009; Ruan et al. 2010; Gongora-Castillo et al. 2012; Fukushima et al. 2012). Gene-enrichment analysis demonstrated that various biological pathways were under regulation of multiple TF families. To determine functions of TFs, those were not annotated by simple annotation (Table S1), a module-based analysis was performed to annotate them based on interacting transcripts. Identification and characterization of modules is a better choice to understand the coordinated behavior of TFs since each module may occupy certain genes representing specific pathways with similar GO terms. Four statistically and biologically significant modules (1–4) were obtained with similar enrichments that were broadly categorized into functional classes viz., defense response, epigenetic regulations, reproductive system, and upstream of TIA pathway, respectively. Moreover, these modules have shown tissue-specific expression (Fig. S15) that further signifies tissue-specific synthesis and accumulation of metabolites. Functional annotation has identified AP2-EREBP TF family to be involved in regulation of primary metabolic pathway and precursors of TIA pathway, similar to ORCA3, which has been previously reported to regulate these pathways in C. roseus (Van der Fits and Memelink 2000). Additionally, WRKY TF family was also observed to regulate ajmalicine and serpentine synthesis that has been previously reported in C. roseus (Schluttenhofer et al. 2014). Few putative TFs (e.g., SBP) identified, which were not functionally annotated through simple annotation and TF family enrichment analysis, have been inferred based on high degree of transcriptional connectivity using module-based analysis. Therefore, graph theoretical approach implemented presented utility to functionally annotate TFs based on their interacting partners.
TFs were found to work in a highly coordinated manner to perform important biological functions since 38 % of them were highly clustered only in the first four modules. Shared genes among significant modules were found to be associated with representative pathways also highlight coordinated and complex behavior of TF families in each module. This analysis shows that TFs do not perform their functions independently but act as composite regulatory modules (CRMs) where more than one TF family work together to facilitate regulation of gene expression (Matys et al. 2006; Jha et al. 2011). TFs present in modules 1 and 4 were found to be associated with regulation of major indole alkaloids and primary metabolic pathway as well as upstream of TIA pathway with high expression in roots and mature leaves, respectively (Figs. 6 and S15). Similar to known regulatory roles of WRKY and AP2-EREBP TF families in C. roseus, this study has also been proposed to regulate the synthesis of indole alkaloids in R. serpentina.
Various studies have reported the integration of gene expression data with metabolomics data that contributes to identifying significant biological processes and molecular mechanisms more precisely (Hirai et al 2005; Yonekura-Sakakibara et al 2008; Meyer et al 2014). In our analysis, inclusion of metabolomics data has also strengthened the reliability of significant modules because of module-specific regulation of various secondary metabolites (Fig. 8). Statistical analysis inferred that transcripts present in module 1 may be close enough on chromosome to form small physical clusters to regulate synthesis of specialized metabolites as supported from previous studies (Frey et al. 2003; Nützmann and Osbourn 2014). Formation of such functional gene set has already proven to be involved in mainly defense response and expedite parallel activation of genes from physically linked genes (Chu et al. 2011) through genomic rearrangements that accelerate the expression of genes in specific pathway. Similarly, TFs present in module 1 may contribute into the diversity of major indole alkaloids in R. serpentina in response to various internal/external stimuli. Significant modules obtained were biologically conceivable with similar enrichments for transcripts in a particular functional classes and their tissue-specific expression may correlate to synthesis as well as accumulation of secondary metabolites. While limited gene expression data was available for this analysis, further experimental studies would be required to ascertain specific role(s) of putative TFs in R. serpentina.
Conclusion
The study presented herein provides an integrative approach constituting of genome- and system-wide analysis to identify regulatory TFs and associated modules involved in R. serpentina secondary metabolism. Integration of metabolomics data provided an advantage to further filter out the putative TFs involved in module-specific regulation of secondary metabolites. Of all four significant modules, module 1 was the most significant since it was found to be statistically associated with synthesis of specialized indole alkaloids, and transcripts contributing to module may be physically clustered on the genome to involve in their synthesis. TF families in modules 1 and 4 may coordinately regulate synthesis of specialized metabolites under response to stimulus and precursors of TIA pathway, respectively. Moreover, module 4 also acts as connecting link between primary and secondary metabolic pathways. Similar to known regulatory roles of WRKY and AP2-EREBP TF families in C. roseus, the present study also proposed them to regulate the synthesis of major indole alkaloids in R. serpentina. As far as our knowledge, we present the first in silico study to identify TFs and their regulatory role in R. serpentina secondary metabolism. Conclusively, the methodology implemented could help to better understand TF-mediated regulation of genes related to TIAs synthesis. Furthermore, this study may assist to identify and explore functions of transcriptional regulators which in turn may aid to improve quantity and diversity of secondary metabolites, and also provide a reference to other non-model plant systems.
Abbreviations
- TFs:
-
Transcription factors
- PCC:
-
Pearson correlation coefficient
- MPGR:
-
Medicinal plant genomics resource
- TAIR10:
-
The Arabidopsis Information Resource 10
- PlnTFDB:
-
Plant Transcription Factor Database
- ND:
-
Network density
- AGRIS:
-
Arabidopsis gene regulatory information server
- TFBS:
-
Transcription factor binding sites
- MCL:
-
Markov cluster
- GO:
-
Gene ontology
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- PMR:
-
Plant and microbial metabolomics resource
- TIA:
-
Terpenoid indole alkaloid
References
Aharoni A, Galili G (2011) Metabolic engineering of the plant primary-secondary metabolism interface. Curr Opin Biotechnol 22:239–244. doi:10.1016/j.copbio.2010.11.004
Albert R (2005) Scale-free networks in cell biology. J Cell Sci 118:4947–57. doi:10.1242/jcs.02714
Alon U (2006) An introduction to systems biology: design principles of biological circuits, 1st edn. 320
Altschul SF, Madden TL, Schäffer AA et al (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402. doi:10.1093/nar/25.17.3389
Aniszewski T (2007) Alkaloids-secrets of life: alkaloid chemistry, biological significance, applications and ecological role, 1st edn. 334
Aoki K, Ogata Y, Shibata D (2007) Approaches for extracting practical information from gene co-expression networks in plant biology. Plant Cell Physiol 48:381–390. doi:10.1093/pcp/pcm013
Azmi M, Qureshi A (2012) Methanolic root extract of Rauwolfia serpentina improves the glucose tolerance in wister mice. J Food Drug Anal 20:484–488. doi:10.6227/jfda.2012200208
Baroux C, Pien S, Grossniklaus U (2007) Chromatin modification and remodeling during early seed development. Curr Opin Genet Dev 17:473–479. doi:10.1016/j.gde.2007.09.004
Barrero RA, Chapman B, Yang Y et al (2011) De novo assembly of Euphorbia fischeriana root transcriptome identifies prostratin pathway related genes. BMC Genomics 12:600. doi:10.1186/1471-2164-12-600
Bazika V (1969) Ajmaline, an alkaloid of Rauwolfia serpentina, in the treatment of cardiac arrhythmias. Vnitr Lek 15:662–670
Beljanski M, Beljanski MS (1982) Selective inhibition of in vitro synthesis of cancer DNA by alkaloids of beta-carboline class. Exp Cell Biol 50:79–87
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc Ser B 57:289–300. doi:10.2307/2346101
Bindea G, Mlecnik B, Hackl H et al (2009) ClueGO: a Cytoscape plug-in to decipher functionally grouped gene ontology and pathway annotation networks. Bioinformatics 25:1091–1093. doi:10.1093/bioinformatics/btp101
Borevitz JO (2000) Activation tagging identifies a conserved MYB regulator of phenylpropanoid biosynthesis. Plant Cell Online 12:2383–2394. doi:10.1105/tpc.12.12.2383
Chae L, Kim T, Nilo-Poyanco R, Rhee SY (2014) Genomic signatures of specialized metabolism in plants. Science (80) 344:510–513. doi: 10.1126/science.1252076
Chang K, Chen M, Zeng L et al (2013) Abscisic acid enhanced ajmalicine biosynthesis in hairy roots of Rauvolfia verticillata by upregulating expression of the MEP pathway genes. Russ J Plant Physiol 61:136–140. doi:10.1134/S102144371401004X
Chien C-H, Chiang-Hsieh Y-F, Tsou A-P et al (2014) Large-scale investigation of human TF-miRNA relations based on coexpression profiles. Biomed Res Int. doi:10.1155/2014/623078
Childs KL, Davidson RM, Buell CR (2011) Gene coexpression network analysis as a source of functional annotation for rice genes. PLoS One 6:e22196. doi:10.1371/journal.pone.0022196
Chou WM, Kutchan TM (1998) Enzymatic oxidations in the biosynthesis of complex alkaloids. Plant J 15:289–300. doi:10.1046/j.1365-313X.1998.00220.x
Chou W-C, Cheng A-L, Brotto M, Chuang C-Y (2014) Visual gene-network analysis reveals the cancer gene co-expression in human endometrial cancer. BMC Genomics 15:300. doi:10.1186/1471-2164-15-300
Chu HY, Wegel E, Osbourn A (2011) From hormones to secondary metabolism: the emergence of metabolic gene clusters in plants. Plant J 66:66–79. doi:10.1111/j.1365-313X.2011.04503.x
Csardi G, Nepusz T (2006) The igraph software package for complex network research. InterJournal Complex Sy:1695
D’Auria JC, Gershenzon J (2005) The secondary metabolism of Arabidopsis thaliana: growing like a weed. Curr Opin Plant Biol 8:308–316. doi:10.1016/j.pbi.2005.03.012
Davuluri RV, Sun H, Palaniswamy SK et al (2003) AGRIS:Arabidopsis gene regulatory information server, an information resource of Arabidopsis cis-regulatory elements and transcription factors. BMC Bioinf 4:25. doi:10.1186/1471-2105-4-25
De Luca V, St Pierre B (2000) The cell and developmental biology of alkaloid biosynthesis. Trends Plant Sci 5:168–173. doi: 10.1016/S1360-1385(00)01575-2
De Vega-Bartol JJ, Simões M, Lorenz WW et al (2013) Transcriptomic analysis highlights epigenetic and transcriptional regulation during zygotic embryo development of Pinus pinaster. BMC Plant Biol. doi:10.1186/1471-2229-13-123
Dey A, De J (2011) Ethnobotanical aspects of Rauvolfia serpentina (L). Benth. ex Kurz. in India, Nepal and Bangladesh. J Med Plants Res 5:144–150
Dimmer EC, Huntley RP, Barrell DG, et al (2008) The Gene Ontology - providing a functional role in proteomic studies. Proteomics 8:n/a. doi: 10.1002/pmic.200800002
Du Z, Zhou X, Ling Y et al (2010) agriGO: a GO analysis toolkit for the agricultural community. Nucleic Acids Res 38:W64–W70. doi:10.1093/nar/gkq310
Dubey VS, Bhalla R, Luthra R (2003) An overview of the non-mevalonate pathway for terpenoid biosynthesis in plants. J Biosci 28:637–646. doi:10.1007/BF02703339
Ehrlich PR, Raven PH (1964) Butterflies and plants: a study in coevolution. Evol (N Y) 18:586–608
Eichinger D (1999) Analysis of metabolic pathways via quantitative prediction of isotope labeling patterns: a retrobiosynthetic 13C NMR study on the monoterpene loganin. Phytochemistry 51:223–236. doi:10.1016/S0031-9422(98)00733-X
Eisen MB, Spellman PT, Brown PO, Botstein D (1998) Cluster analysis and display of genome-wide expression patterns. Proc Natl Acad Sci 95:14863–14868. doi:10.1073/pnas.95.25.14863
El-Sayed M, Verpoorte R (2007) Catharanthus terpenoid indole alkaloids: biosynthesis and regulation. Phytochem Rev 6:277–305. doi:10.1007/s11101-006-9047-8
Emmert-streib F, Dehmer M (2008) Robustness in scale-free networks: comparing directed and undirected networks. Int J Mod Phys C 19:717–726. doi:10.1142/S0129183108012510
Ershov YV, Gantt RR, Cunningham FX, Gantt E (2002) Isoprenoid biosynthesis in Synechocystis sp. strain PCC6803 is stimulated by compounds of the pentose phosphate cycle but Not by pyruvate or deoxyxylulose-5-phosphate. J Bacteriol 184:5045–5051. doi:10.1128/JB.184.18.5045-5051.2002
Facchini PJ, De Luca V (2008) Opium poppy and Madagascar periwinkle: model non-model systems to investigate alkaloid biosynthesis in plants. Plant J 54:763–784. doi:10.1111/j.1365-313X.2008.03438.x
Fauteux F, Strömvik MV (2009) Seed storage protein gene promoters contain conserved DNA motifs in Brassicaceae, Fabaceae and Poaceae. BMC Pant Biol 9:126. doi:10.1186/1471-2229-9-126
Fernandez JA, de Luca V (1994) Ubiquitin-mediated degradation of tryptophan decarboxylase from Catharanthus roseus. Phytochemistry 36:1123–1128. doi:10.1016/S0031-9422(00)89624-7
Flügge U-I, Gao W (2005) Transport of isoprenoid intermediates across chloroplast envelope membranes. Plant Biol 7:91–97. doi:10.1055/s-2004-830446
Fraenkel GS (1959) The Raison d’Etre of Secondary Plant Substances: These odd chemicals arose as a means of protecting plants from insects and now guide insects to food. Science (80- ) 129:1466–1470. doi: 10.1126/science.129.3361.1466
Frey M, Huber K, Park WJ et al (2003) A 2-oxoglutarate-dependent dioxygenase is integrated in DIMBOA-biosynthesis. Phytochemistry 62:371–376. doi:10.1016/S0031-9422(02)00556-3
Fukushima A, Nishizawa T, Hayakumo M et al (2012) Exploring tomato gene functions based on coexpression modules using graph clustering and differential coexpression approaches. Plant Physiol 158:1487–1502. doi:10.1104/pp. 111.188367
Glenn WS, Runguphan W, O’Connor SE (2013) Recent progress in the metabolic engineering of alkaloids in plant systems. Curr Opin Biotechnol 24:354–365. doi:10.1016/j.copbio.2012.08.003
Gongora-Castillo E, Childs KL, Fedewa G et al (2012) Development of transcriptomic resources for interrogating the biosynthesis of monoterpene indole alkaloids in medicinal plant species. PLoS One 7:e52506. doi:10.1371/journal.pone.0052506
Grafi G, Ohad N (2013) Epigenetic Memory and Control in Plants. 270
Hartwell LH, Hopfield JJ, Leibler S, Murray AW (1999) From molecular to modular cell biology. Nature 402:C47–C52. doi:10.1038/35011540
Haynes BC, Maier EJ, Kramer MH et al (2013) Mapping functional transcription factor networks from gene expression data. Genome Res 23:1319–1328. doi:10.1101/gr.150904.112
He K, Guo A-Y, Gao G et al (2010) Computational identification of plant transcription factors and the construction of the PlantTFDB database. Methods Mol Biol 674:351–368. doi:10.1007/978-1-60761-854-6_21
Herbert RB (2003) The biosynthesis of plant alkaloids and nitrogenous microbial metabolites. Nat Prod Rep 20:494–508. doi:10.1039/b006522f
Herrmann KM, Weaver LM (1999) The shikimate pathway. Annu Rev Plant Physiol Plant Mol Biol 50:473–503. doi:10.1146/annurev.arplant.50.1.473
Hirai MY, Klein M, Fujikawa Y et al (2005) Elucidation of gene-to-gene and metabolite-to-gene networks in Arabidopsis by integration of metabolomics and transcriptomics. J Biol Chem 280:25590–255595. doi:10.1074/jbc.M502332200
Hu J, Wang D, Li J et al (2014) Genome-wide identification of transcription factors and transcription-factor binding sites in oleaginous microalgae Nannochloropsis. Sci Rep 4:5454. doi:10.1038/srep05454
Hutcheon DE (1971) The pharmacological basis of therapeutics, 4th Edition. Louis S. Goodman and Alfred Gilman. J Clin Pharmacol New Drugs 11:388. doi:10.1177/009127007101100511
Ihmels J, Levy R, Barkai N (2004) Principles of transcriptional control in the metabolic network of Saccharomyces cerevisiae. Nat Biotechnol 22:86–92. doi:10.1038/nbt918
Jakoby M, Weisshaar B, Dröge-Laser W et al (2002) bZIP transcription factors in Arabidopsis. Trends Plant Sci 7:106–111. doi:10.1016/S1360-1385(01)02223-3
Jerie P (2007) Milestones of cardiovascular therapy. IV. Reserpine. Cas Lek Cesk 146:573–577
Jha A, Shankar R (2014) MiRNAting control of DNA methylation. J Biosci 39:365–380. doi:10.1007/s12038-014-9437-9
Jha A, Mehra M, Shankar R (2011) The regulatory epicenter of miRNAs. J Biosci 36:621–638. doi:10.1093/nar/gkj143
Jiao Y, Lau OS, Deng XW (2007) Light-regulated transcriptional networks in higher plants. Nat Rev Genet 8:217–230. doi:10.1038/nrg2049
Johnson S, Torres JJ, Marro J, Muñoz MA (2010) Entropic origin of disassortativity in complex networks. Phys Rev Lett 104:108702. doi:10.1103/PhysRevLett.104.108702
Kinsella RJ, Kähäri A, Haider S, et al (2011) Ensembl BioMarts: a hub for data retrieval across taxonomic space. Database (Oxford) 2011:bar030. doi: 10.1093/database/bar030
Koch MA, Weisshaar B, Kroymann J et al (2001) Comparative genomics and regulatory evolution: conservation and function of the Chs and Apetala3 promoters. Mol Biol Evol 18:1882–1891
Köppel C, Wagemann A, Martens F (1989) Pharmacokinetics and antiarrhythmic efficacy of intravenous ajmaline in ventricular arrhythmia of acute onset. Eur J Drug Metab Pharmacokinet 14:161–167. doi:10.1007/BF03190857
Kutchan TM (2005) A role for intra- and intercellular translocation in natural product biosynthesis. Curr Opin Plant Biol 8:292–300. doi:10.1016/j.pbi.2005.03.009
Lee WP, Tzou W-S (2009) Computational methods for discovering gene networks from expression data. Brief Bioinform 10:408–423. doi:10.1093/bib/bbp028
Lelek I, Furedi Szabo M (1961) On the antiallergic effect of the reserpine alkaloid of Rauwolfia serpentina. Allerg Asthma (Leipz) 7:142–144
Leménager D, Ouelhazi L, Mahroug S et al (2005) Purification, molecular cloning, and cell-specific gene expression of the alkaloid-accumulation associated protein CrPS in Catharanthus roseus. J Exp Bot 56:1221–1228. doi:10.1093/jxb/eri116
Li CY, Leopold AL, Sander GW et al (2013) The ORCA2 transcription factor plays a key role in regulation of the terpenoid indole alkaloid pathway. Plant Biol 13:155. doi:10.1186/1471-2229-13-155
Liang H, Li W-H (2007) MicroRNA regulation of human protein protein interaction network. RNA 13:1402–1408. doi:10.1261/rna.634607
Locket S (1955) Oral preparations of Rauwolfia serpentina in treatment of essential hypertension. Br Med J 1:809–816. doi:10.1136/bmj.1.4917.809
Ma S, Shah S, Bohnert HJ et al (2013) Incorporating motif analysis into gene co-expression networks reveals novel modular expression pattern and new signaling pathways. PLoS Genet 9:e1003840. doi:10.1371/journal.pgen.1003840
Mao L, Van Hemert JL, Dash S, Dickerson JA (2009) Arabidopsis gene co-expression network and its functional modules. BMC Bioinf 10:346. doi:10.1186/1471-2105-10-346
Matys V, Kel-Margoulis OV, Fricke E et al (2006) TRANSFAC and its module TRANSCompel: transcriptional gene regulation in eukaryotes. Nucleic Acids Res 34:D108–D110. doi:10.1093/nar/gkj143
Matzke M, Kanno T, Huettel B et al (2007) Targets of RNA-directed DNA methylation. Curr Opin Plant Biol 10:512–519. doi:10.1016/j.pbi.2007.06.007
McCluskey A, Lalkhen AG (2007) Statistics IV: interpreting the results of statistical tests. Contin Educ Anaesthesia, Crit Care Pain 7:208–212. doi:10.1093/bjaceaccp/mkm042
Memelink J, Gantet P (2007) Transcription factors involved in terpenoid indole alkaloid biosynthesis in Catharanthus roseus. Phytochem Rev 6:353–362. doi:10.1007/s11101-006-9051-z
Meyer E, Aspinwall MJ, Lowry DB et al (2014) Integrating transcriptional, metabolomic, and physiological responses to drought stress and recovery in switchgrass (Panicum virgatum L.). BMC Genomics 15:527. doi:10.1186/1471-2164-15-527
Misra A, Sriram G (2013) Network component analysis provides quantitative insights on an Arabidopsis transcription factor-gene regulatory network. BMC Syst Biol 7:126. doi:10.1186/1752-0509-7-126
Mitsuda N, Ohme-Takagi M (2009) Functional analysis of transcription factors in Arabidopsis. Plant Cell Physiol 50:1232–1248. doi:10.1093/pcp/pcp075
Mol J, Grotewold E, Koes R (1998) How genes paint flowers and seeds. Trends Plant Sci 3:212–217. doi:10.1016/S1360-1385(98)01242-4
Morris JH, Apeltsin L, Newman AM et al (2011) clusterMaker: a multi-algorithm clustering plugin for Cytoscape. BMC Bioinf 12:436. doi:10.1186/1471-2105-12-436
Naika M, Shameer K, Sowdhamini R (2013) Comparative analyses of stress-responsive genes in Arabidopsis thaliana: insight from genomic data mining, functional enrichment, pathway analysis and phenomics. Mol Biosyst 9:1888–1908. doi:10.1039/C3MB70072K
Nammi S, Boini KM, Koppula S, Sreemantula S (2005) Reserpine-induced central effects: pharmacological evidence for the lack of central effects of reserpine methiodide. Can J Physiol Pharmacol 83:509–515. doi:10.1139/y05-039
Nützmann H-W, Osbourn A (2014) Gene clustering in plant specialized metabolism. Curr Opin Biotechnol 26:91–99. doi:10.1016/j.copbio.2013.10.009
O’Connor SE, Maresh JJ (2006) Chemistry and biology of monoterpene indole alkaloid biosynthesis. Nat Prod Rep 23:532–547. doi:10.1039/B512615K
Oudin A, Courtois M, Rideau M, Clastre M (2007) The iridoid pathway in Catharanthus roseus alkaloid biosynthesis. Phytochem Rev 6:259–276. doi:10.1007/s11101-006-9054-9
Pani A, Mahapatra RK (2013) Computational identification of microRNAs and their targets in Catharanthus roseus expressed sequence tags. Genomics Data 1:2–6. doi:10.1016/j.gdata.2013.06.001
Pathania S, Randhawa V, Bagler G (2013) Prospecting for novel plant-derived molecules of Rauvolfia serpentina as inhibitors of Aldose Reductase, a potent drug target for diabetes and its complications. PLoS One 8:e61327. doi:10.1371/journal.pone.0061327
Paul A, Jha A, Bhardwaj S et al (2014) RNA-seq-mediated transcriptome analysis of actively growing and winter dormant shoots identifies non-deciduous habit of evergreen tree tea during winters. Sci Rep 4:5932. doi:10.1038/srep05932
Perianez-Rodriguez J, Manzano C, Moreno-Risueno MA (2014) Post-embryonic organogenesis and plant regeneration from tissues: two sides of the same coin. Front Plant Sci. doi:10.3389/fpls.2014.00219
Ramachandran S, Hiratsuka K, Chua NH (1994) Transcription factors in plant growth and development. Curr Opin Genet Dev 4:642–646. doi:10.1016/0959-437X(94)90129-Q
Raval A, Ray A (2013) Introduction to biological networks, 1st edn. 335.
Riaño-Pachón DM, Ruzicic S, Dreyer I, Mueller-Roeber B (2007) PlnTFDB: an integrative plant transcription factor database. BMC Bioinf 8:42. doi:10.1186/1471-2105-8-42
Ríos G, Leida C, Conejero A, Badenes ML (2014) Epigenetic regulation of bud dormancy events in perennial plants. Front Plant Sci 5:247. doi:10.3389/fpls.2014.00247
Ruan J, Dean AK, Zhang W (2010) A general co-expression network-based approach to gene expression analysis: comparison and applications. BMC Syst Biol. doi:10.1186/1752-0509-4-8
Ruppert M, Woll J, Giritch A et al (2005) Functional expression of an ajmaline pathway-specific esterase from Rauvolfia in a novel plant-virus expression system. Planta 222:888–98. doi:10.1007/s00425-005-0031-0
Samuni-Blank M, Izhaki I, Dearing MD et al (2012) Intraspecific directed deterrence by the mustard oil bomb in a desert plant. Curr Biol 22:1218–1220. doi:10.1016/j.cub.2012.04.051
Schluttenhofer C, Pattanaik S, Patra B, Yuan L (2014) Analyses of Catharanthus roseus and Arabidopsis thaliana WRKY transcription factors reveal involvement in jasmonate signaling. BMC Genomics 15:502. doi:10.1186/1471-2164-15-502
Schmid J, Amrhein N (1995) Molecular organization of the shikimate pathway in higher plants. Phytochemistry 39:737–749. doi:10.1016/0031-9422(94)00962-S
Schmid R, Blaxter ML (2008) annot8r: GO, EC and KEGG annotation of EST datasets. BMC Bioinf 9:180. doi:10.1186/1471-2105-9-180
Schuster G, Gruissem W (1991) Chloroplast mRNA 3’ end processing requires a nuclear-encoded RNA-binding protein. EMBO J 10:1493–1502
Seemann M, Tse Sum Bui B, Wolff M et al (2006) Isoprenoid biosynthesis in plant chloroplasts via the MEP pathway: direct thylakoid/ferredoxin-dependent photoreduction of GcpE/IspG. FEBS Lett 580:1547–1552. doi:10.1016/j.febslet.2006.01.082
Shannon P, Markiel A, Ozier O et al (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13:2498–2504. doi:10.1101/gr.1239303
Shimizu M, Fujimoto R, Ying H et al (2014) Identification of candidate genes for Fusarium yellows resistance in Chinese cabbage by differential expression analysis. Plant Mol Biol 85:247–57. doi:10.1007/s11103-014-0182-0
Shoji T, Yamada Y, Hashimoto T (2000) Jasmonate induction of putrescine N-methyltransferase genes in the root of Nicotiana sylvestris. Plant Cell Physiol 41:831–839
Shukla AK, Shasany AK, Gupta MM, Khanuja SPS (2006) Transcriptome analysis in Catharanthus roseus leaves and roots for comparative terpenoid indole alkaloid profiles. J Exp Bot 57:3921–3932. doi:10.1093/jxb/erl146
Siberil Y (2002) Cloning of a cDNA encoding an E2 ubiquitin-conjugating enzyme from Catharanthus roseus: expression analysis in plant organs and in response to hormones in cell suspensions. J Exp Bot 53:149–150. doi:10.1093/jexbot/53.366.149
Sim J, Wright CC (2005) The kappa statistic in reliability studies: use, interpretation, and sample size requirements. Phys Ther 85:257–268
Simes RJ (1986) An improved Bonferroni procedure for multiple tests of significance. Biometrika 73:751–754. doi:10.1093/biomet/73.3.751
Singh K, Foley RC, Oñate-Sánchez L (2002) Transcription factors in plant defense and stress responses. Curr Opin Plant Biol 5:430–436. doi:10.1016/S1369-5266(02)00289-3
Singh DK, Srivastava B, Sahu A (2004) Spectrophotometric determination of Rauwolfia alkaloids: estimation of reserpine in pharmaceuticals. Anal Sci 20:571–573. doi:10.2116/analsci.20.571
Srivastava GP, Li P, Liu J, Xu D (2010) Identification of transcription factor’s targets using tissue-specific transcriptomic data in Arabidopsis thaliana. BMC Syst Biol. doi:10.1186/1752-0509-4-S2-S2
Sundar AS, Varghese SM, Shameer K et al (2008) STIF: Identification of stress-upregulated transcription factor binding sites in Arabidopsis thaliana. Bioinformation 2:431–437
Suttipanta N, Pattanaik S, Kulshrestha M et al (2011) The transcription factor CrWRKY1 positively regulates the terpenoid indole alkaloid biosynthesis in Catharanthus roseus. Plant Physiol 157:2081–2093. doi:10.1104/pp.111.181834
The UniProt Consortium (2007) The Universal Protein Resource (UniProt). Nucleic Acids Res 35:D193–D197. doi:10.1093/nar/gkl929
Vakil RJ (1955) Rauwolfia serpentina in the treatment of high blood pressure: a review of the literature. Circulation 12:220–229. doi:10.1161/01.CIR.12.2.220
Van der Fits L, Memelink J (2000) ORCA3, a jasmonate-responsive transcriptional regulator of plant primary and secondary metabolism. Science (80- ) 289:295–297. doi: 10.1126/science.289.5477.295
Van Dijk ADJ, Morabito G, Fiers M et al (2010) Sequence motifs in MADS transcription factors responsible for specificity and diversification of protein-protein interaction. PLoS Comput Biol 6:e1001017. doi:10.1371/journal.pcbi.1001017
Van Dongen S (2008) Graph clustering via a discrete uncoupling process. SIAM J Matrix Anal Appl 30:121–141. doi:10.1137/040608635
Vom Endt D, Kijne JW, Memelink J (2002) Transcription factors controlling plant secondary metabolism: what regulates the regulators. Phytochemistry 61:107–114. doi:10.1016/S0031-9422(02)00185-1
Wang D, Zhang C, Hearn DJ et al (2010) Identification of transcription-factor genes expressed in the Arabidopsis female gametophyte. BMC Plant Biol 10:110. doi:10.1186/1471-2229-10-110
Wei J, van Loon JJA, Gols R et al (2014) Reciprocal crosstalk between jasmonate and salicylate defence-signalling pathways modulates plant volatile emission and herbivore host-selection behaviour. J Exp Bot 65:3289–3298. doi:10.1093/jxb/eru181
Weng J-K, Noel JP (2012) The remarkable pliability and promiscuity of specialized metabolism. Cold Spring Harb Symp Quant Biol 77:309–320. doi:10.1101/sqb.2012.77.014787
Wink M (2010) Annual plant reviews, functions and biotechnology of plant secondary metabolites, 2nd edn. 424
Yang C-Q, Fang X, Wu X-M et al (2012a) Transcriptional regulation of plant secondary metabolism. J Integr Plant Biol 54:703–12. doi:10.1111/j.1744-7909.2012.01161.x
Yang J, Osman K, Iqbal M et al (2012b) Inferring the Brassica rapa interactome using protein-protein interaction data from Arabidopsis thaliana. Front Plant Sci 3:297. doi:10.3389/fpls.2012.00297
Yang LT, Qi YP, Bin LY et al (2013) iTRAQ protein profile analysis of Citrus sinensis roots in response to long-term boron-deficiency. J Proteomics 93:179–206. doi:10.1016/j.jprot.2013.04.025
Yonekura-Sakakibara K, Tohge T, Matsuda F et al (2008) Comprehensive flavonol profiling and transcriptome coexpression analysis leading to decoding gene-metabolite correlations in Arabidopsis. Plant Cell 20:2160–2176. doi:10.1105/tpc.108.058040
Zarate SI, Kempema LA, Walling LL (2007) Silverleaf whitefly induces salicylic acid defenses and suppresses effectual jasmonic acid defenses. Plant Physiol 143:866–875. doi: 10.1104/pp.106.090035
Zhang L, Yu S, Luo KZ et al (2012) Identification of gene modules associated with drought response in rice by network-based analysis. PLoS One 7:e33748. doi:10.1371/journal.pone.0033748
Zhu J-Y, Sun Y, Wang Z-Y (2012) Genome-wide identification of transcription factor-binding sites in plants using chromatin immunoprecipitation followed by microarray (ChIP-chip) or sequencing (ChIP-seq). Methods Mol Biol 876:173–188. doi:10.1007/978-1-61779-809-2_14
Ziegler J, Facchini PJ (2008) Alkaloid Biosynthesis: metabolism and trafficking. Annu Rev Plant Biol 59:735–769. doi:10.1146/annurev.arplant.59.032607.092730
Acknowledgments
We acknowledge the computational infrastructure provided in the form of project MLP0076 by CSIR-Institute of Himalayan Bioresource Technology (CSIR-IHBT), a constituent national laboratory of Council of Scientific and Industrial Research, India, and Department of Biotechnology, Government of India for infrastructural support in the form of Bioinformatics Infrastructure Facility (BIF) as well. The authors are thankful to Dr. Paramvir Singh Ahuja for encouragement and constant support. Shivalika Pathania is grateful to the Department of Science and Technology (DST) for INSPIRE fellowship. We are also thankful to Ashwani Jha and Vinay Randhawa for technical help in manuscript preparation. The CSIR-IHBT communication number for this article is 3777.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Table S1
(XLSX 20 kb)
Table S2
(XLSX 10090 kb)
Table S3
(DOCX 668 kb)
Table S4
(DOCX 15 kb)
Table S5
(XLSX 53 kb)
Table S6
(XLSX 43 kb)
Table S7
(XLSX 36 kb)
Table S8
(XLSX 36 kb)
Table S9
(DOCX 13 kb)
Fig. S1
Gene ontology (GO) annotation of complete R. serpentina transcriptome. Pie chart representing GO-based annotation of complete transcriptome for a biological process and b molecular function category (GIF 92 kb)
Fig. S2
Gene ontology (GO) and KEGG pathways annotation of complete R. serpentina transcriptome. Pie chart representing a GO-based annotation for cellular component and b KEGG pathways annotation of complete transcriptome (GIF 88 kb)
Fig. S3
Threshold selection. a The actual number of edges and all possible edges among non-singleton nodes as a function of PCC cutoff values. b The actual number of edges and all nodes among the non-singleton nodes as a function of PCC cutoff values. The “igraph” library of R package is used to obtain these plots (GIF 13 kb)
Fig. S4
Weighted co-expression network follows power law degree distribution. A data set from the weighted co-expression network is represented with black filled circles, and the degree distribution adheres to a power law as all these circles lie on or close to the red line which is the graph of a function of the form ax-k. The “igraph” library of R package is used to obtain this plot (GIF 12 kb)
Fig. S5
Hierarchical tree representing significantly enriched GO terms for ABI3VP1 TF family. These over-represented GO terms for biological process category are obtained using agriGO. Each GO term represented by box are labeled by their GO ID, term definition, and statistical information. Degree of color saturation of a box is positively correlated to the enrichment level of the term (GIF 48 kb)
Fig. S6
Hierarchical tree representing significantly enriched GO terms for bHLH TF family. These over-represented GO terms for biological process category are obtained using agriGO. Each GO term represented by box are labeled by their GO ID, term definition, and statistical information. Degree of color saturation of a box is positively correlated to the enrichment level of the term (GIF 43 kb)
Fig. S7
Hierarchical tree representing significantly enriched GO terms for HB TF family. These over-represented GO terms for biological process category are obtained using agriGO. Each GO term represented by box are labeled by their GO ID, term definition, and statistical information. Degree of color saturation of a box is positively correlated to the enrichment level of the term (GIF 30 kb)
Fig. S8
Hierarchical tree representing significantly enriched GO terms for MYB TF family. These over-represented GO terms for biological process category are obtained using agriGO. Each GO term represented by box are labeled by their GO ID, term definition, and statistical information. Degree of color saturation of a box is positively correlated to the enrichment level of the term (GIF 58 kb)
Fig. S9
Hierarchical tree representing significantly enriched GO terms for MYB-related TF family. These over-represented GO terms for biological process category are obtained using agriGO. Each GO term represented by box are labeled by their GO ID, term definition, and statistical information. Degree of color saturation of a box is positively correlated to the enrichment level of the term (GIF 55 kb)
Fig. S10
Hierarchical tree representing significantly enriched GO terms for WRKY TF family. These over-represented GO terms for biological process category are obtained using agriGO. Each GO term represented by box are labeled by their GO ID, term definition, and statistical information. Degree of color saturation of a box is positively correlated to the enrichment level of the term (GIF 51 kb)
Fig. S11
Hierarchical tree representing significantly enriched GO terms for -EREBP TF family. These over-represented GO terms for biological process category are obtained using agriGO. Each GO term represented by box are labeled by their GO ID, term definition, and statistical information. Degree of color saturation of a box is positively correlated to the enrichment level of the term (GIF 33 kb)
Fig. S12
Hierarchical tree representing significantly enriched GO terms in enrichment analysis of bZIP TF family. These over-represented GO terms for biological process category are obtained using agriGO. Each GO term represented by box are labeled by their GO ID, term definition, and statistical information. Degree of color saturation of a box is positively correlated to the enrichment level of the term (GIF 7 kb)
Fig. S13
Hierarchical graph representing significantly enriched GO terms for MADS TF family. These over-represented GO terms for biological process category are obtained using agriGO. Each GO term represented by box are labeled by their GO ID, term definition, and statistical information. Degree of color saturation of a box is positively correlated to the enrichment level of the term (GIF 6 kb)
Fig. S14
Clustering of weighted co-expression network. A total of 42 modules are obtained from clustering of weighted co-expression network using MCL algorithm. Colored filled circles representing TFs in each module. Cytoscape is used for visualization of the network (GIF 89 kb)
Fig. S15
Heat map of transcripts in four significant modules (1–4) using expression data of different tissues. Heat map is representing tissue-specific expression of transcripts in significant modules: a module 1 in roots, b module 2 in young leaves, c module 3 in flower, and d module 4 in mature leaves, where average expression is calculated based on normalized transcriptomics data. The “gplots” library of R package is used to plot heat map (GIF 100 kb)
Fig. S16
Top most enriched KEGG pathway annotation for four significant modules (1–4). Each pie segment is labeled with significant KEGG pathway, and the percentage fraction of annotations associated with that particular pathway. a Module 1, b module 2, c module 3, and d module 4 (GIF 65 kb)
Fig. S17
Hierarchical graph representing significantly enriched GO terms for module 2. These over-represented GO terms for biological process category are obtained using agriGO. Each GO term represented by box are labeled by their GO ID, term definition, and statistical information. Degree of color saturation of a box is positively correlated to the enrichment level of the term (GIF 20 kb)
Fig. S18
Hierarchical tree representing significantly enriched GO terms for module 3. These over-represented GO terms for biological process category are generated through agriGO. Each GO term represented by box are labeled by their GO ID, term definition, and statistical information. Degree of color saturation of a box is positively correlated to the enrichment level of the term (GIF 40 kb)
Fig. S19
Hierarchical tree representing significantly enriched GO terms for module 4. These over-represented GO terms for biological process category are obtained using agriGO. Each GO term represented by box are labeled by their GO ID, term definition, and statistical information. Degree of color saturation of a box is positively correlated to the enrichment level of the term (GIF 28 kb)
Fig. S20
Network representing the significantly enriched pathways in module 1. GO terms-based network is obtained from enrichment analysis against reference model A. thaliana for module 1. Functionally grouped pathways are mainly found to be associated to monocarboxylic acid biosynthetic process, hormone-mediated signaling pathway, and defense response. ClueGO plugin is used to generate this GO terms-based network (GIF 100 kb)
Fig. S21
Network representing the significantly enriched pathways in module 2. GO terms-based network is obtained from enrichment analysis against reference model Arabidopsis thaliana. Functionally grouped pathways are mainly found to be associated to “regulation of gene expression, epigenetic,” “RNA metabolic process,” and “macromolecular modification” which also complements agriGO enrichment result. ClueGO, plugin in cytoscape, is used to generate this GO terms-based network. (GIF 95 kb)
Fig. S22
Network representing the significantly enriched pathways in module 3. GO terms-based network is obtained from enrichment analysis against reference model Arabidopsis thaliana. Functionally grouped pathways are mainly found to be associated to “pollen exine formation,” “stamen development,” and “external encapsulating structure organization” which also complements agriGO enrichment result. ClueGO, plugin in cytoscape, is used to generate this GO terms-based network (GIF 27 kb)
Fig. S23
Network representing the significantly enriched pathways in module 4. GO terms-based network is obtained from enrichment analysis against reference model Arabidopsis thaliana. Functionally grouped pathways are mainly found to be associated to “photosynthesis”, “plastid organization,” and “monocarboxylic acid biosynthetic process” which also complements agriGO enrichment result. ClueGO, plugin in cytoscape, is used to generate this GO terms-based network (GIF 80 kb)
Rights and permissions
About this article
Cite this article
Pathania, S., Acharya, V. Computational Analysis of “-omics” Data to Identify Transcription Factors Regulating Secondary Metabolism in Rauvolfia serpentina . Plant Mol Biol Rep 34, 283–302 (2016). https://doi.org/10.1007/s11105-015-0919-1
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11105-015-0919-1