Abstract
Although nonspecific lipid transfer proteins (nsLTPs) are widely present in plants, their functions are not fully understood. Here, we isolated and characterized a putative nsLTP gene, BplLTP1, from Betula platyphylla. The full-length cDNA of BplLTP1 is 638 bp long, including a 363-bp open reading frame (GenBank accession no. JQ409562). The putative protein BplLTP1 contains an N-terminal signal sequence and possesses the characteristic features of nsLTPs. An amino acid sequence alignment revealed that BplLTP1 shares a high level of similarity with other known nsLTPs. A 3D model of BplLTP1 was also constructed based on the homology model. Quantitative real time-PCR analysis showed that there were no obvious differences in the expression levels of BplLTP1 among different tissues. BplLTP1 displayed distinctly higher expression levels in young tissues than in older tissues. Moreover, BplLTP1 was upregulated at the mononuclear microspore developmental stages in male inflorescences. Expression analysis was performed using 3-month-old cultured seedlings, and the results revealed that the expression of BplLTP1 was upregulated by exogenous abscisic acid and salicylic acid, downregulated by exogenous methyl jasmonate, and not significantly altered by exogenous gibberellin A. In addition, a prokaryotic expression system was constructed with pET32a-BplLTP1 and Escherichia coli strain BL21 and subjected to abiotic stress resistance analysis. The results indicated that the expression of BplLTP1 improved the resistance of the recombinant strain to salt (NaCl) and drought (polyethylene glycol) stress, but not to alkali (NaHCO3) stress.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plants are frequently subjected to a plethora of abiotic stress conditions, such as drought, flood, salt, alkali, low temperature, heat, and heavy metal toxicity. Plants are also subjected to various biotic challenges from pathogens and herbivores. All of these abiotic and biotic stress factors have adverse effects on plant growth and productivity. In response to these stress factors, various genes are regulated to mitigate the effects of stress and to enhance plant tolerance (Mahajan and Tuteja 2005; Knight and Knight 2001).
Lipid transfer proteins (LTPs) are widely present in animals, plants, and microorganisms (Yamada 1992) and account for approximately 4 % of the total soluble protein content in higher plants (Kader 1996). Plant nonspecific lipid transfer proteins (nsLTPs), which were first identified in 1984, are characterized by their capacity to facilitate the transfer of phospholipids between liposomes in vitro (Kader et al. 1984). To date, plant nsLTPs have been identified in 50 different species, and there are more than 100 potential plant nsLTPs (José-Estanyol et al. 2004). Plant nsLTPs are cysteine-rich peptides with low molecular masses encoded by a multigene family. Plant nsLTPs are mainly grouped into two subfamilies based on their primary structures (Carvalho and Gomes 2007; Yeats and Rose 2008; Arondel et al. 2000). nsLTP1 members have molecular masses of approximately 9 kDa and contain 90–95 amino acid residues, while nsLTP2 members have molecular masses of approximately 7 kDa and possess, on average, 70 amino acids. Both nsLTP1 and nsLTP2 family members are abundant, basic proteins with isoelectric points (pI) between 8.5 and 12, with eight strictly conserved cysteine residues (Kader 1997; Carvalho and Gomes 2007). Both nsLTP1 and nsLTP2 family members contain a signal peptide at the amino terminal region, generally comprising 21–27 amino acids for the nsLTP1 family and 27–35 amino acids for the nsLTP2 family, allowing mature LTP peptides to be exported to the apoplast accurately (Carvalho and Gomes 2007). The 3D structure of the plant nsLTP family comprises a globular molecule stabilized by four disulfide bridges (Shin et al. 1995; Samuel et al. 2002). The most remarkable structural feature of the LTP family is the presence of a hydrophobic cavity that has the capacity to bind fatty acids, acyl-coenzyme A (acyl-CoA), and phospholipids (Carvalho and Gomes 2007).
Several plant nsLTP1 members have been localized to the cell wall (Thoma et al. 1993; Pyee et al. 1994; De O. Carvalho et al. 2004) and have been extracted from plant surfaces (Pyee et al. 1994). Tissue-specific expression experiments examining nsLTPs have demonstrated that nsLTP genes are expressed at different levels in various tissues during diverse developmental stages (Pyee et al. 1994; Thoma et al. 1994). Plant nsLTP genes are responsive to abiotic stresses such as drought, cold and salt stress, and biotic stresses such as bacterial and fungal pathogens (Jang et al. 2004; Jung et al. 2003; Wu et al. 2004; Wang et al. 2009; Sarowar et al. 2008). In addition, several signaling molecules such as abscisic acid (ABA), salicylic acid (SA), ethylene, and methyl jasmonate (MJ) are also able to regulate the expression of nsLTP genes (Yubero-Serrano et al. 2003; Wu et al. 2004; Jung et al. 2003; Jung et al. 2006; Wang et al. 2009). Furthermore, a variety of potential biological functions for nsLTPs have also been proposed, such as surface cutin biosynthesis (Kader 1996; DeBono et al. 2009; Lee et al. 2009), liquid secretion (Choi et al. 2012), embryogenesis (Kader 1996), anther development (Kader 1996; Ariizumi et al. 2002; Zhang et al. 2010; Chae et al. 2009; Chen et al. 2011), pollen tube tip growth (Chae et al. 2009), seed maturation (Thoma et al. 1994), and plant signaling (Gao et al. 2009; Sarowar et al. 2008; Blein et al. 2002), although the exact biological role of nsLTPs in vivo remains unclear.
Asian white birch (Betula platyphylla) is an important economic forest tree species in northern China. In this study, we isolated and characterized a putative nsLTP gene, BplLTP1, from B. platyphylla based on the expressed sequence tag (EST) method, which was obtained from a cDNA-AFLP library created from the early and late developmental stages of male inflorescences (Xing and Liu 2011). We investigated the expression patterns of BplLTP1 in various tissues and the diverse developmental stages of male inflorescences. The BplLTP1 expression patterns under ABA, SA, MJ, and gibberellin A (GA3) treatment were also examined. In addition, a prokaryotic expression system was constructed using pET32a-BplLTP1 and Escherichia coli strain BL21 and subjected to NaCl, PEG 6000 (polyethylene glycol), and NaHCO3 stress resistance analyses. Taken together, the results of this study suggest that BplLTP1 plays an important role in the defense responses of B. platyphylla.
Materials and Methods
Plant Material and Treatment
Plant materials, which were used to clone BplLTP1 and analyze tissue-specific expression, were sampled from mature B. platyphylla in the birch forest yard of Northeast Forestry University, Heilongjiang, China. Male inflorescences were sampled every 2–3 days from July 15 to September 1, 2011. From July 15 to August 14, the male inflorescences were in the meiosis stages, and from August 23 to September 1, they were in the mononuclear microspore developmental stages (Liu and Yang 2006). Mature pollen and seeds were sampled on May 6 and August 15, respectively. Female inflorescences, male inflorescences, young leaves, older leaves, young petioles, older petioles, young stems, and older stems were sampled on July 15. The samples were quickly frozen in liquid nitrogen and stored at −80 °C before total RNA extraction.
Birch twigs with buds were cultivated in vitro in a rooting medium consisting of woody plant medium (Lloyd and McCown 1980) supplemented with 2 μM indole-3-butytric acid. The cultures were maintained under a 16-h light/8-h dark photoperiod. When the plants were 3 months old, they were sprayed with GA3 (500 μM), SA (1 mM), ABA (100 μM), and MJ (100 μM). The plants were sampled 0 (control), 4, 8, 12, or 24 h after treatment. The leaves were harvested and frozen in liquid nitrogen.
Total RNA Extraction and Purification
Total RNA was extracted from different tissues using the modified CTAB method (Chang et al. 1993), and the potential contaminating genomic DNA was digested at 37 °C for 30 min using DNase I (Takara, Japan). The integrity of the total RNA samples was verified using 1 % agarose gel electrophoresis.
Isolation of BplLTP1
To obtain the full-length cDNA sequence of BplLTP1, 5′ rapid amplification of cDNA ends (RACE) and 3′ RACE reactions were performed with a 5′ and 3′ Full RACE Kit (Takara) according to the manufacturer’s protocol. Briefly, for 5′ RACE, the 5′ cap structure was removed from purified RNA using tobacco acid pyrophosphatase, ligated to the 5′ RACE adaptor using T4 DNA ligase, and used as a template to synthesize first-strand cDNA using M-MLV reverse transcriptase (Takara). Two specific primers (5′GSP1 and 5′GSP2; Table 1) were designed. Primer 5′GSP1, along with the 5′ RACE Outer Primer, was used for the primary PCR. The primary PCR products were then diluted 100-fold with deionized water to be used as the template for the nested PCR. Primer 5′GSP2 and the 5′ RACE Inner Primer were used for the nested PCR. The PCR reactions were performed as follows: 94 °C for 5 min, 30 cycles of 94 °C for 30 s, 61 °C (for the primary PCR) or 65 °C (for the nested PCR) for 30 s and 72 °C for 1 min, followed by 72 °C for 10 min. The PCR products were purified and subcloned into the pMD18-T vector (Takara) and sequenced by the Beijing Genomic Institute (Beijing, China). For 3′ RACE, the purified RNA was used to synthesize first-strand cDNA using M-MLV reverse transcriptase (Takara) with the oligo-dT-3 site adaptor primer. The primary PCR was performed with 3′GSP1 and the 3′ RACE Outer Primer (Table 1). The PCR products were then diluted and used as templates for the nested PCR with 3’GSP2 and 3’RACE Inner Primer (Table 1). The PCR reactions were performed as follows: 94 °C for 5 min, 30 cycles of 94 °C for 30 s, 63 °C (for the primary PCR) or 66 °C (for the nested PCR) for 30 s and 72 °C for 1 min, followed by 72 °C for 10 min. The 3′ RACE PCR products were purified, subcloned into the pMD18-T vector (Takara), and sequenced.
Sequence Analysis of BplLTP1
The open reading frame (ORF) of the BplLTP1 cDNA sequence was analyzed using the ORF Finder (http://www.ncbi.nlm.nih.gov/gorf/). The pI (isoelectric point) and M w (molecular weight) predictions were performed using the Compute pI/Mw tool (http://web.expasy.org/compute_pi/; Bjellqvist et al. 1993). Conserved domains were identified using the NCBI Conserved Domain Database (http://www.ncbi.nlm.nih.gov/Structure/cdd/wrpsb.cgi/; Marchler-Bauer et al. 2011). The signal peptide was predicted using online SignalP, version 4.0 (http://www.cbs.dtu.dk/services/SignalP/; Petersen et al. 2011). Multiple sequence alignments were carried out with ClustalX, version 1.83 (Thompson et al. 1997). The phylogenetic tree was constructed based on MEGA 5 (Tamura et al. 2011). Homology modeling of the 3D structure of BplLTP1 was generated using the SWISS-MODEL server (http://swissmodel.expasy.org/; Arnold et al. 2006).
Quantitative and Semiquantitative RT-PCR
The cDNA sample was synthesized from 1 μg of total RNA using M-MLV reverse transcriptase (Takara). After first-strand cDNA synthesis, the cDNA sample was used as a template for quantitative RT-PCR (qRT-PCR) or semiquantitative RT-PCR analysis. qRT-PCR was performed using an MJ OpticonTM2 System (Bio-Rad, Hercules, CA) and QuantiTect SYBR-green PCR Master Mix (Toyobo, Japan). The qRT-PCR reactions were performed as follows: 95 °C for 5 min followed by 40 cycles at 95 °C for 30 s, 58 °C for 30 s, 72 °C for 30 s, and 79 °C for 1 s. Semiquantitative RT-PCR was performed at 94 °C for 5 min followed by 26 cycles at 94 °C for 30 s, 58 °C for 30 s, and 72 °C for 30 s. The Actin gene (GenBank accession no. EU588981) was used as the standard control in quantitative and semiquantitative RT-PCR (Qi et al. 2010; Li et al. 2010). The primer sequences are shown in Table 1. These experiments were performed in triplicate.
Response of Recombinant BL21 (DE3) to Abiotic Stresses
Primers PF (with a BamHI site) and PR (with a SacI site; Table 1) were used to amplify the ORF of BplLTP1 without the N-terminal signal peptide sequence. The PCR reactions were performed under the following conditions: 95 °C for 5 min, 30 cycles of 94 °C for 30 s, 58 °C for 30 s, and 72 °C for 1 min, followed by 72 °C for 10 min. The PCR products were subcloned into the pMD18-T vector (Takara) and transferred to E. coli DH5α competent cells. After sequencing identification, the recombinant plasmids were digested with the enzymes BamHI and SacI and subcloned into the bacterial expression vector pET-32a(+) (Novagen) that had been digested with the same enzymes, which resulted in a recombinant plasmid named pET32a-BplLTP1. Both pET-32a (+; blank vector) and pET32a-BplLTP1 were transformed into E. coli strain BL21 (DE3). The bacterial cultures were grown in Luria–Bertani (LB) medium (100 μg ml−1 ampicillin) at 37 °C until the culture optical density values at 600 nm (OD600) reached 0.6, as determined with an ultraviolet spectrophotometer (Shimadzu, China). One millimolar isopropyl-thiogalactoside (IPTG) was used to induce protein expression. After induction for 4 h, both pET-32a and pET32a-BplLTP1 bacterial cultures were added to fresh liquid LB medium (100 μg ml−1 ampicillin) supplemented with 0.4 M NaCl, 0.2 M NaHCO3, and 20 % PEG 6000 at a dilution of 1:100 and incubated at 37 °C with shaking (250 rpm). The OD600 values of the cultures were recorded every 1.5 h. Untreated bacterial strains were used as controls. Each experiment was performed with three parallel replicates.
Statistical Analysis
Statistical analyses were carried out using the IBM SPSS-19 statistical software package (IBM, USA). Results are represented as the mean ± SE. The data were analyzed statistically using ANOVA, and the differences between different samples were analyzed using the LSD test at a probability level of 0.05.
Results
Clone and Sequence Analysis of BplLTP1
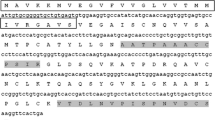
To obtain the full-length cDNA of BplLTP1, 5′ and 3′ RACE-PCRs were performed using nested gene-specific primers. The full-length cDNA of BplLTP1 (GenBank accession no. JQ409562) was 638 bp in length and contained a 363-bp ORF with a 67-bp 5′ UTR (untranslated region) and a 208-bp 3′ downstream UTR, based on the sequences of two overlapping frames obtained by the 5′ and 3′ RACE-PCR procedures. The BplLTP1 ORF is predicted to encode a polypeptide of 120 amino acid residues. The putative polypeptide contains a typical signal peptide of 26 amino acid residues at the N terminus (Fig. 1), as determined using SignalP, version 4.0. After removing the signal peptide sequence, the mature peptide has a predicted molecular mass of 9.54 kDa, with a theoretical pI of 9.18.
Alignment of the deduced BplLTP1 sequence with other known plant nsLTP sequences. The amino acid sequences used in the alignment were from Arabidopsis thaliana (AAF76927), Brassica oleracea (AAA73948), Dimocarpus longan (AEC04836), Gossypium hirsutum (AAF35185), Populus trichocarpa (XP_002305877), Castanea sativa (ADK60918), Nicotiana tabacum (BAK19150), Oryza sativa (ACA50499) and Zea mays (ABA33846). The probable signal peptide sequence is underlined. The eight strictly conserved cysteine residues are shaded in gray. The two highly conserved regions (DRQ and CGV) in BplLTP1 are boxed. Asterisks indicate identical amino acids, colon indicates strongly similar, and a period indicates weakly similar amino acids
Multiple amino acid sequence alignments showed that BplLTP1 shares identities with several other known nsLTP1s. BplLTP1 shares 64.2 % amino acid identity with Gossypium hirsutum LTP (GenBank accession no. AAF35185), and it shares 40.9 % amino acid identity with Oryza sativa LTP (GenBank accession no. ACA50499; Table 2). In addition, we also compared the homology between BplLTP1 and the LTP (GenBank accession no. EIL54273) of E. coli, which contains 392 amino acid residues and is not an nsLTP. E. coli LTP shares only 18.3 % homology with BplLTP1 (Table 2).
Further analysis showed that BplLTP1 contains the conserved in vitro lipid-binding motifs DRQ and CGV (Botton et al. 2002), and the eight strictly conserved cysteine residues form four disulfide bridges (Kader 1997; Fig. 1). To investigate the relationship between BplLTP1 and other plant nsLTP1 family proteins, a phylogenetic tree was constructed based on multiple amino acid sequence alignments. The phylogenetic tree shows that BplLTP1 is closely related to nsLTP1 from Castanea sativa (Fig. 2).
Phylogenetic tree of BplLTP1 and homologous nsLTPs from other plants. The phylogenetic tree was generated with MEGA 5 software using the neighbor-joining method (Tamura et al. 2011). Scale bar represents evolutionary distance
Homology modeling of the 3D structure of BplLTP1 was performed based on the template Pru p 3 (PDB code 2algB, resolution 2.30 Å) from Prunus persica (Pasquato et al. 2006). BplLTP1 shows 67.7 % amino acid sequence identity with P. persica Pru p 3 at an E value cutoff of 2.2e−21 (Arnold et al. 2006). The hypothetical 3D structural model of BplLTP1 shows a protein that contains four α-helices, with a secondary structure compacted by four disulfide bridges and a C-terminal tail. The four bridges are formed by Cys4 and Cys53, Cys14 and Cys30, Cys31 and Cys76, and Cys51 and Cys90 (Fig. 3).
Expression Patterns of BplLTP1 in Various B. platyphylla Tissues
The results of qRT-PCR analysis showed that the expression level of BplLTP1 was significantly different in different tissues. BplLTP1 expression was high in the leaves and male inflorescences, with the highest expression level in young leaves, while the transcripts were almost undetectable in the older stems and mature pollen. Interestingly, the results also showed that the expression level of BplLTP1 was significantly higher in young tissues than in older tissues (Fig. 4a). We also examined the expression patterns of BplLTP1 at different stages of male inflorescence development. The results (Fig. 4b) showed that BplLTP1 expression was reduced by 57 % (P < 0.05) at the meiosis stages, and the expression of this gene was significantly (P < 0.05) increased (threefold) at the mononuclear microspore developmental stages (Liu and Yang 2006).
qRT-PCR expression patterns of BplLTP1 in B. platyphylla. a Expression analysis of BplLTP1 in female inflorescences (FI), male inflorescences (MI), young leaves (YL), older leaves (OL), young petioles (YP), older petioles (OP), young stems (YS), older stems (OS), seeds (SE), and mature pollen (PO) of B. platyphylla. b Expression analysis of BplLTP1 in different developmental stages of male inflorescences. The sampling dates of male inflorescences are indicated in the figure. Values represent the means of three replicate experiments with standard error. Lowercase letters indicate statistically significant differences (P < 0.05)
Response to Different Exogenous Plant Hormones
To elucidate the possible role of BplLTP1 in B. platyphylla, the expression patterns of BplLTP1 in response to exogenous ABA, SA, MJ, and GA3 were analyzed using semiquantitative RT-PCR. The results revealed that BplLTP1 transcript levels increased significantly after 4 h of exogenous ABA treatment, and this elevated level was maintained (Fig. 5a). Under exogenous SA treatment, BplLTP1 transcript levels gradually increased (Fig. 5b). In contrast, the levels of the BplLTP1 transcripts decreased significantly after 4 h of exogenous MJ treatment, followed by an increase after 16 h of treatment (Fig. 5c). Compared with ABA, SA, and MJ, exogenous GA3 did not significantly alter the expression level of BplLTP1 (Fig. 5d).
Expression patterns of BplLTP1 in response to exogenous hormone treatments. Expression analyses of BplLTP1 were performed with total RNA extracted from leaves treated with abscisic acid (ABA) (a), salicylic acid (SA) (b), methyl jasmonate (MJ) (c), or gibberellin A (GA 3 ) (d). The Actin gene was used as an amplification and loading control. The number of PCR cycles in each experiment was 26. Experiments were repeated three times. A representative experiment is shown in the figure
Response of Recombinant BL21 (DE3) to Abiotic Stresses
After 1 mM IPTG induction for 4 h, transformed E. coli BL21 (DE3) strain pET32a-BplLTP1 and the control strain pET32a containing an empty vector were cultured at 37 °C in LB medium. The OD600 values of the cultures were measured every 1.5 h. As shown in Fig. 6a, both bacterial strains presented typical “S” growth curves, which were substantially overlapping. The maximum OD600 values were both close to 2 during the stationary phase. These results suggest that the expression of BplLTP1 rarely affects the normal growth of recombinant E. coli BL21 (DE3).
Growth curves of E. coli BL21 (DE3) strains harboring recombinant plasmid pET32a-BplLTP1 or empty vector pET32a(+). Growth curves of E. coli BL21 (DE3) strains in liquid LB medium (100 μg ml−1 ampicillin) with no supplements (control) (a) or supplemented with 0.4 M NaCl (b), 20 % (w/v) PEG 6000 (c), or 0.2 M NaHCO3 (d). Experiments were repeated three times. Data represent the mean ± SE
To determine the function of the fusion protein under salt (NaCl), alkali (NaHCO3), and drought (PEG 6000) stress, the growth curves of E. coli BL21 (DE3) cultures harboring pET32a-BplLTP1 or pET32a were examined. Under salt stress, E. coli strain pET32a-BplLTP1 displayed an obviously faster growth than the control strain; the control strain pET32a barely survived (Fig. 6b). Under drought stress, E. coli strain pET32a showed a slightly reduced growth rate compared with E. coli strain pET32a-BplLTP1 (Fig. 6c). However, under alkali stress, the growth curve assay showed no significant difference between E. coli strains pET32a-BplLTP1 and pET32a, with maximum OD600 values of <0.3 for both strains (Fig. 6d).
Discussion
RACE is a simple but effective technique that has recently been employed to obtain full-length c-DNA sequences based on the sequences of known ESTs (Li et al. 2012; Phillips et al. 2013; Yang et al. 2012; Qin et al. 2013). In the present study, BplLTP1 was successfully isolated from B. platyphylla using the RACE method. Similar to other nsLTPs, BplLTP1 encodes a small protein with eight strictly conserved cysteine residues predicted to form four disulphide bridges, as well as a 26-amino acid N-terminal signal peptide predicted to target proteins to the secretory pathway (Carvalho and Gomes 2007). The molecular mass of mature BplLTP1 is estimated to be approximately 9.5 kDa, suggesting that this protein belongs to the type I LTP family (Carvalho and Gomes 2007).
The hypothetical 3D structure of BplLTP1 is compact and globular, comprising four α-helices and a C-terminal tail (Fig. 3). This structure forms a flexible hydrophobic pocket with a unique capacity to accommodate fatty acids, acyl-CoA, and phospholipids (Carvalho and Gomes 2007). In addition, the conserved in vitro lipid-binding motifs DRQ and CGV (Botton et al. 2002) were also observed in BplLTP1 and homologs from other plants (Fig. 1). These results indicate that BplLTP1 may possess lipid transfer activity. Although the cavities of nsLTPs are thought to be vitally important for the biological functions of these proteins, no direct evidence has revealed the function of these cavities in vivo (Yeats and Rose 2008). Blein et al. (2002) found that nsLTP from wheat (Triticum aestivum) cannot load lipids from intact membranes under normal physiological conditions.
In a variety of plants, nsLTP genes have different transcription patterns in diverse tissues during various developmental stages and under different physiological conditions (Carvalho and Gomes 2007). For example, nsLTPs have been found in vascular tissues and epidermal cells during the formative stage of Arabidopsis development (Thoma et al. 1994) as well as in the cuticular waxy layer of broccoli (Pyee et al. 1994). The expression patterns of BplLTP1 demonstrated that this gene is expressed at higher levels in young tissues (including young leaves, young stems, and young petioles) than in older tissues (including older leaves, older stems, and older petioles; Fig. 4a). This expression pattern suggests that BplLTP1 plays a necessary role in the formation and deposition of cuticular material (Pyee et al. 1994; Thoma et al. 1994). Similar results were also obtained in previous studies of plant nsLTPs. For instance, immunological measurements showed that nsLTP from broccoli is highly expressed in young leaves, constituting 50 % of leafy proteins, while the level of expression drops to only 4 % when the leaves age (Pyee et al. 1994). The nsltp1 gene from Arabidopsis is expressed at high levels in young developing tissues, and its expression declines in fully expanded tissues; this pattern is also observed in the petal and sepal abscission zones (Thoma et al. 1994).
It is noteworthy that the expression level of BplLTP1 in male inflorescences was up to 12-fold that in female inflorescences (Fig. 4a). We measured the expression levels of BplLTP1 in male inflorescences at different developmental stages. Our results demonstrated that the expression of BplLTP1 decreased during the meiosis stages, while the expression of this gene was significantly increased during the mononuclear microspore developmental stages (Fig. 4b; Liu and Yang 2006). These results suggest that BplLTP1 plays an important role in male inflorescences. Unfortunately, we cannot determine the direct link between BplLTP1 and pollen development from these data because the male inflorescences contained abundant bracts. Nevertheless, our results indirectly suggest that BplLTP1 plays a role during pollen development (Zhang et al. 2010; Chen et al. 2011). A previous qRT-PCR analysis showed that OsC6 expression is increased in anthers, beginning at the early mononuclear microspore development stage, while OsC6 transcripts are barely detectable during the formation of mature trinucleate pollen (Zhang et al. 2010). This expression pattern, which is similar to the expression pattern of BplLTP1, helps explain why BplLTP1 was almost undetectable in mature pollen (Fig. 4a).
NsLTPs are responsive to various plant hormones (Wu et al. 2004; Wang et al. 2009; Jung et al. 2006; Yubero-Serrano et al. 2003). ABA is considered to be a stress hormone because of its important role in plant stress responses. There is much overlap in the expression patterns of stress-related genes after plants are exposed to drought, high salt, or exogenous ABA (Zhang et al. 2006; Leung and Giraudat 1998). Previous studies have shown that several putative cis-acting elements, such as ABRE, DRE, G-box, and coupling elements, which are required for ABA-induced gene expression, are not found in the promoter region of LTP (Jung et al. 2006; George and Parida 2010). However, in this study, the transcription of BplLTP1 was strongly induced by exogenous ABA treatment (Fig. 5a), suggesting that ABA upregulates LTP gene expression through the action of other elements rather than by direct contact with the promoter of this gene. SA and MJ are also plant signaling molecules involved in stress responses. Many studies have shown that the SA- and JA-mediated signaling pathways interact in an antagonistic manner (Niki et al. 1998; Kachroo et al. 2001). BplLTP1 expression was induced in leaves treated with exogenous SA (Fig. 5b), but suppressed by exogenous MJ treatment (Fig. 5c). These results suggest that MJ might antagonize the induction of BplLTP1 by the stress signaling molecule SA. In this study, we found that exogenous GA3 treatment did not increase BplLTP1 transcription in the leaves of B. platyphylla. This result suggests that GA3 is not an effective signaling molecule for the induction of BplLTP1 transcription.
Prokaryotic expression systems are an effective way to produce exogenous protein. Previous studies have shown that the expression of exogenous plant genes can directly contribute to increasing stress tolerance in the host bacterium (Yamada et al. 2002; Miyasaka et al. 2000; Lan et al. 2005). In this study, to elucidate the contribution of BplLTP1 to abiotic stress responses, recombinant E. coli harboring BplLTP1 was subjected to abiotic stresses including salt, alkali, and drought stress. Previous studies have shown that nsLTP expression is induced under salt stress conditions (Jang et al. 2004; Jung et al. 2003; Wu et al. 2004; Wang et al. 2009). Furthermore, overexpression of the CALTP gene enhances salt stress resistance in transgenic Arabidopsis (Jung et al. 2005). These results indicate that nsLTPs play an important role in salt stress response. In this study, our results demonstrated that the expression of BplLTP1 increases salt stress tolerance in the host E. coli, suggesting that BplLTP1 plays a role in plant salt tolerance response. The induction of nsLTP by drought stress occurs in wheat (T. aestivum; Jang et al. 2004), pepper (Capsicum annuum; Jung et al. 2003), bromegrass (Bromus inermis; Wu et al. 2004), and tamarix (Tamarix hispida; Wang et al. 2009). In addition, the overexpression of the nsLTP gene increases drought tolerance in transgenic Arabidopsis (Jung et al. 2005). The expression patterns of BplLTP1 suggest that this gene plays a role in the formation and deposition of cuticular material (Pyee et al. 1994; Thoma et al. 1994). Epicuticular wax plays a significant physiological role in maintaining water balance in plants (Lemieux 1996). Our data also indicate that the expression of BplLTP1 in host E. coli can increase drought stress tolerance. Taken together, these results suggest that BplLTP1 plays a role in plant drought tolerance. Moreover, the upregulation of BplLTP1 expression in response to the exogenous application of the plant hormones ABA and SA may contribute to plant survival by increasing cuticle thickness in response to stress (Jung et al. 2003). Liu et al. (2008) found that the expression of Polygonum sibiricum nsLTPs (PsnsLTPs) is significantly increased under NaHCO3 stress, which suggests that PsnsLTPs play an important role in saline stress resistance. However, our results indicate that the alkali resistance of host E. coli is not increased by the expression of BplLTP1. Whether this is also the case in B. platyphylla remains to be determined.
References
Ariizumi TA, Amagai MA, Shibata DS, Hatakeyama KH, Watanabe MW, Toriyama KT (2002) Comparative study of promoter activity of three anther-specific genes encoding lipid transfer protein, xyloglucan endotransglucosylase/hydrolase and polygalacturonase in transgenic Arabidopsis thaliana. Plant Cell Rep 21:90–96. doi:10.1007/s00299-002-0487-3
Arnold K, Bordoli L, Kopp J, Schwede T (2006) The SWISS-MODEL workspace: a web-based environment for protein structure homology modelling. Bioinformatics 22:195–201. doi:10.1093/bioinformatics/bti770
Arondel V, Vergnolle C, Cantrel C, Kader J-C (2000) Lipid transfer proteins are encoded by a small multigene family in Arabidopsis thaliana. Plant Sci 157:1–12. doi:10.1016/s0168-9452(00)00232-6
Bjellqvist B, Hughes GJ, Pasquali C, Paquet N, Ravier F, Sanchez J-C, Frutiger S, Hochstrasser D (1993) The focusing positions of polypeptides in immobilized pH gradients can be predicted from their amino acid sequences. Electrophoresis 14:1023–1031. doi:10.1002/elps.11501401163
Blein JP, Coutos-Thévenot P, Marion D, Ponchet M (2002) From elicitins to lipid-transfer proteins: a new insight in cell signalling involved in plant defence mechanisms. Trends Plant Sci 7:293–296. doi:10.1016/s1360-1385(02)02284-7
Botton A, Begheldo M, Rasori A, Bonghi C, Tonutti P (2002) Differential expression of two lipid transfer protein genes in reproductive organs of peach (Prunus persica L. Batsch). Plant Sci 163:993–1000. doi:10.1016/s0168-9452(02)00271-6
Carvalho AO, Gomes VM (2007) Role of plant lipid transfer proteins in plant cell physiology—a concise review. Peptides 28:1144–1153. doi:10.1016/j.peptides.2007.03.004
Chae K, Kieslich CA, Morikis D, Kim S-C, Lord EM (2009) A gain-of-function mutation of Arabidopsis lipid transfer protein 5 disturbs pollen tube tip growth and fertilization. Plant Cell 21:3902–3914. doi:10.1105/tpc.109.070854
Chang S, Puryear J, Cairney J (1993) A simple and efficient method for isolating RNA from pine trees. Plant Mol Biol Rep 11:113–116. doi:10.1007/bf02670468
Chen C, Chen G, Hao X, Cao B, Chen Q, Liu S, Lei J (2011) CaMF2, an anther-specific lipid transfer protein (LTP) gene, affects pollen development in Capsicum annuum L. Plant Sci 181:439–448. doi:10.1016/j.plantsci.2011.07.003
Choi YE, Lim S, Kim H-J, Han JY, Lee M-H, Yang Y, Kim J-A, Kim Y-S (2012) Tobacco NtLTP1, a glandular-specific lipid transfer protein, is required for lipid secretion from glandular trichomes. Plant J 70:480–491. doi:10.1111/j.1365-313X.2011.04886.x
de Oliveira Carvalho A, Teodoro CES, Da Cunha M, Okorokova-Façanha AL, Okorokov LA, Fernandes KVS, Gomes VM (2004) Intracellular localization of a lipid transfer protein in Vigna unguiculata seeds. Physiol Plant 122:328–336. doi:10.1111/j.1399-3054.2004.00413.x
DeBono A, Yeats TH, Rose JKC, Bird D, Jetter R, Kunst L, Samuels L (2009) Arabidopsis LTPG is a glycosylphosphatidylinositol-anchored lipid transfer protein required for export of lipids to the plant surface. Plant Cell 21:1230–1238. doi:10.1105/tpc.108.064451
Gao G, Jin LP, Xie KY, Qu DY (2009) The potato StLTPa7 gene displays a complex Ca2+-associated pattern of expression during the early stage of potato–Ralstonia solanacearum interaction. Mol Plant Pathol 10:15–27. doi:10.1111/j.1364-3703.2008.00508.x
George S, Parida A (2010) Characterization of an oxidative stress inducible nonspecific lipid transfer protein coding cDNA and its promoter from drought tolerant plant Prosopis juliflora. Plant Mol Biol Rep 28:32–40. doi:10.1007/s11105-009-0127-y
Jang CS, Lee HJ, Chang SJ, Seo YW (2004) Expression and promoter analysis of the TaLTP1 gene induced by drought and salt stress in wheat (Triticum aestivum L.). Plant Sci 167:995–1001. doi:10.1016/j.plantsci.2004.05.019
José-Estanyol M, Gomis-Rüth FX, Puigdomènech P (2004) The eight-cysteine motif, a versatile structure in plant proteins. Plant Physiol Biochem 42:355–365. doi:10.1016/j.plaphy.2004.03.009
Jung HW, Kim W, Hwang BK (2003) Three pathogen-inducible genes encoding lipid transfer protein from pepper are differentially activated by pathogens, abiotic, and environmental stresses. Plant Cell Environ 26:915–928. doi:10.1046/j.1365-3040.2003.01024.x
Jung HW, Kim KD, Hwang BK (2005) Identification of pathogen-responsive regions in the promoter of a pepper lipid transfer protein gene (CALTPI) and the enhanced resistance of the CALTPI transgenic Arabidopsis against pathogen and environmental stresses. Planta 221:361–373
Jung HW, Lim CW, Hwang BK (2006) Isolation and functional analysis of a pepper lipid transfer protein III (CALTPIII) gene promoter during signaling to pathogen, abiotic and environmental stresses. Plant Sci 170:258–266. doi:10.1016/j.plantsci.2005.08.010
Kachroo P, Shanklin J, Shah J, Whittle EJ, Klessig DF (2001) A fatty acid desaturase modulates the activation of defense signaling pathways in plants. Proc Natl Acad Sci U S A 98:9448–9453
Kader JC (1996) Lipid-transfer proteins in plants. Annu Rev Plant Physiol Plant Mol Biol 47:627–654
Kader JC (1997) Lipid-transfer proteins: a puzzling family of plant proteins. Trends Plant Sci 2:66–70. doi:10.1016/s1360-1385(97)82565-4
Kader JC, Julienne M, Vergnolle C (1984) Purification and characterization of a spinach-leaf protein capable of transferring phospholipids from liposomes to mitochondria or chloroplasts. Eur J Biochem 139:411–416
Knight H, Knight MR (2001) Abiotic stress signalling pathways: specificity and cross-talk. Trends Plant Sci 6:262–267. doi:10.1016/s1360-1385(01)01946-x
Lan Y, Cai D, Zheng YZ (2005) Expression of three differen t group soybean lea genes enhanced stress tolerance in Escherichia coli. J Integr Plant Biol 47:613–621
Lee SB, Go YS, Bae H-J, Park JH, Cho SH, Cho HJ, Lee DS, Park OK, Hwang I, Suh MC (2009) Disruption of glycosylphosphatidylinositol-anchored lipid transfer protein gene altered cuticular lipid composition, increased plastoglobules, and enhanced susceptibility to infection by the fungal pathogen Alternaria brassicicola. Plant Physiol 150:42–54. doi:10.1104/pp.109.137745
Lemieux B (1996) Molecular genetics of epicuticular wax biosynthesis. Trends Plant Sci 1:312–318
Leung J, Giraudat J (1998) Abscisic acid signal transduction. Annu Rev Plant Physiol Plant Mol Biol 49:199–222
Li Q-F, Sun S, Yuan D-Y, Yu H-X, Gu M-H, Liu Q-Q (2010) Validation of candidate reference genes for the accurate normalization of real-time quantitative RT-PCR data in rice during seed development. Plant Mol Biol Rep 28:49–57. doi:10.1007/s11105-009-0124-1
Li S, Yu F, Wang M, Guo X, Li H (2012) Molecular characterization of a Nicotiana tabacum NtRDR6 gene. Plant Mol Biol Rep 30:1375–1384. doi:10.1007/s11105-012-0455-1
Liu X, Yang C (2006) Temporal characteristics of developmental cycles of female and male flowers in Betula platyphylla in northeastern China. Sci Silvae Sin 42:28–32
Liu GJ, Tian X, Liu CC, Qu CP, Liu GF, Yang CP (2008) Cloning of coding sequence of non-specific transfer protein from Polygonum sibiricum Laxm. and expression under salinity stress. Chin J Biochem Mol Biol 24:1140–1145
Lloyd G, McCown B (1980) Commercially-feasible micropropagation of mountain laurel, Kalmia latifolia, by use of shoot-tip culture. Proc Int Plant Prop Soc 3:421–427
Mahajan S, Tuteja N (2005) Cold, salinity and drought stresses: an overview. Arch Biochem Biophys 444:139–158. doi:10.1016/j.abb.2005.10.018
Marchler-Bauer A, Lu S, Anderson JB, Chitsaz F, Derbyshire MK, DeWeese-Scott C, Fong JH, Geer LY, Geer RC, Gonzales NR, Gwadz M, Hurwitz DI, Jackson JD, Ke Z, Lanczycki CJ, Lu F, Marchler GH, Mullokandov M, Omelchenko MV, Robertson CL, Song JS, Thanki N, Yamashita RA, Zhang D, Zhang N, Zheng C, Bryant SH (2011) CDD: a conserved domain database for the functional annotation of proteins. Nucleic Acids Res 39:225–229. doi:10.1093/nar/gkq1189
Miyasaka H, Kanaboshi H, Ikeda K (2000) Isolation of several anti-stress genes from the halotolerant green alga Chlamydomonas by simple functional expression screening with Escherichia coli. World J Microbiol Biotechnol 16:23–29. doi:10.1023/a:1008982332139
Niki T, Mitsuhara I, Seo S, Ohtsubo N, Ohashi Y (1998) Antagonistic effect of salicylic acid and jasmonic acid on the expression of pathogenesis-related (PR) protein genes in wounded mature tobacco leaves. Plant Cell Physiol 39:500–507
Pasquato N, Berni R, Folli C, Folloni S, Cianci M, Pantano S, Helliwell JR, Zanotti G (2006) Crystal structure of peach Pru p3, the prototypic member of the family of pant non-specific lipid transfer protein pan-allergens. J Mol Biol 356:684–694. doi:10.1016/j.jmb.2005.11.063
Petersen TN, Brunak S, von Heijne G, Nielsen H (2011) SignalP 4.0: discriminating signal peptides from transmembrane regions. Nat Methods 8:785–786, http://www.nature.com/nmeth/journal/v8/n10/abs/nmeth.1701.html#supplementary-information
Phillips SM, Dubery IA, Heerden H (2013) Identification and molecular characterisation of a lectin receptor-like kinase (GhLecRK-2) from cotton. Plant Mol Biol Rep 31:9–20. doi:10.1007/s11105-012-0470-2
Pyee J, Yu HS, Kolattukudy PE (1994) Identification of a lipid transfer protein as the major protein in the surface wax of broccoli (Brassica oleracea) leaves. Arch Biochem Biophys 311:460–468. doi:10.1006/abbi.1994.1263
Qi J, Yu S, Zhang F, Shen X, Zhao X, Yu Y, Zhang D (2010) Reference gene selection for real-time quantitative polymerase chain reaction of mRNA transcript levels in Chinese cabbage (Brassica rapa L. ssp. pekinensis). Plant Mol Biol Rep 28:597–604. doi:10.1007/s11105-010-0185-1
Qin Q, Kaas Q, Zhang L, Xu K, Li N, Zheng W, Lai Q (2013) Isolation and characterization of a cytosolic pyruvate kinase cDNA from loquat (Eriobotrya japonica Lindl.). Plant Mol Biol Rep 31:109–119. doi:10.1007/s11105-012-0479-6
Samuel D, Liu Y-J, Cheng C-S, Lyu P-C (2002) Solution structure of plant nonspecific lipid transfer protein-2 from rice (Oryza sativa). J Biol Chem 277:35267–35273. doi:10.1074/jbc.M203113200
Sarowar S, Kim YJ, Kim KD, Hwang BK, Ok SH, Shin JS (2008) Overexpression of lipid transfer protein (LTP) genes enhances resistance to plant pathogens and LTP functions in long-distance systemic signaling in tobacco. Plant Cell Rep 28:419–427. doi:10.1007/s00299-008-0653-3
Shin DH, Lee JY, Hwang KY, Kyu Kim K, Suh SW (1995) High-resolution crystal structure of the non-specific lipid-transfer protein from maize seedlings. Structure 3:189–199. doi:10.1016/s0969-2126(01)00149-6
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Thoma S, Kaneko Y, Somerville C (1993) A non-specific lipid transfer protein from Arabidopsis is a cell wall protein. Plant J 3:427–436. doi:10.1046/j.1365-313X.1993.t01-25-00999.x
Thoma S, Hecht U, Kippers A, Botella J, De Vries S, Somerville C (1994) Tissue-specific expression of a gene encoding a cell wall-localized lipid transfer protein from Arabidopsis. Plant Physiol 105:35–45. doi:10.1104/pp.105.1.35
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The ClustalX windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Wang C, Yang C, Gao C, Wang Y (2009) Cloning and expression analysis of 14 lipid transfer protein genes from Tamarix hispida responding to different abiotic stresses. Tree Physiol 29:1607–1619. doi:10.1093/treephys/tpp082
Wu G, Robertson AJ, Liu X, Zheng P, Wilen RW, Nesbitt NT, Gusta LV (2004) A lipid transfer protein gene BG-14 is differentially regulated by abiotic stress, ABA, anisomycin, and sphingosine in bromegrass (Bromus inermis). J Plant Physiol 161:449–458. doi:10.1078/0176-1617-01259
Xing L, Liu X-m (2011) Characterization of Betula platyphylla gene transcripts associated with early development of male inflorescence. Mol Biol Rep 39:929–935. doi:10.1007/s11033-011-0818-y
Yamada M (1992) Lipid transfer proteins in plants and microorganisms. Plant Cell Physiol 33:1–6
Yamada A, Sekiguchi M, Mimura T, Ozeki Y (2002) The role of plant CCTα in salt- and osmotic-stress tolerance. Plant Cell Physiol 43:1043–1048. doi:10.1093/pcp/pcf120
Yang Y, Xian Z, Wu Y, Li J, Deng W (2012) Molecular cloning and expression analysis of a putative 5′–3′ EXORIBONUCLEASE4 (XRN4) gene from tomato. Plant Mol Biol Rep 30:1348–1356. doi:10.1007/s11105-012-0449-z
Yeats TH, Rose JK (2008) The biochemistry and biology of extracellular plant lipid-transfer proteins (LTPs). Protein Sci 17:191–198
Yubero-Serrano EM, Moyano E, Medina-Escobar N, Muñoz-Blanco J, Caballero JL (2003) Identification of a strawberry gene encoding a non-specific lipid transfer protein that responds to ABA, wounding and cold stress. J Exp Bot 54:1865–1877. doi:10.1093/jxb/erg211
Zhang JH, Jia WS, Yang JC, Ismail AM (2006) Role of ABA in integrating plant responses to drought and salt stresses. Field Crop Res 97:111–119
Zhang D, Liang W, Yin C, Zong J, Gu F (2010) OsC6, encoding a lipid transfer protein, is required for postmeiotic anther development in rice. Plant Physiol 154:149–162. doi:10.1104/pp.110.158865
Acknowledgments
The work was supported by grants from the National Natural Science Foundation of China (31100449) and the Key Project of Chinese Ministry of Education (109053).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Guan, M., Chai, R., Kong, X. et al. Isolation and Characterization of a Lipid Transfer Protein Gene (BplLTP1) from Betula platyphylla . Plant Mol Biol Rep 31, 991–1001 (2013). https://doi.org/10.1007/s11105-013-0571-6
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11105-013-0571-6