Abstract
Aims
To assess whether the yew roots, which are able to provide a very constant environment due to their long life-span, can maintain the original arbuscular mycorrhizal (AM) fungal community during yew population decline.
Methods
The diversity of AM fungi (AMF) colonizing the roots of yew was analyzed by selecting the small subunit ribosomal RNA genes to construct a database of the overall community of AMF in the experimental area. A terminal restriction fragment length polymorphism (TRFLP) approach was used to identify the AMF communities present in yew roots. Physiological and environmental variables related to topology and soil and plant characteristics were determined as markers of habitat degradation.
Results
The AMF communities within yew roots were found to be dependent on soil, plant and topological variables indicative of habitat degradation surrounding the yew. The phylogenetic diversity of AMF associated to the yews was lower in habitats more exposed to degradation than in those better conserved.
Conclusions
The target yews can be grouped into two degradation levels. AMF communities were also affected by the degradation processes affecting their hosts. This finding rules out the role of these trees as refugia for their original AMF community, a fact that should be considered in plant reintroduction programs using AMF as bioenhancers.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Mediterranean ecosystems are characterized by particular climate conditions which include a dry and hot summer and irregular, sometimes heavy, rain events concentrated in spring and autumn. In the Southeast of the Iberian Peninsula these conditions result in a semi-arid climate, causing ecophysiological constraints for a wide range of plant species. Such stress situations are exacerbated by the negative impact of anthropogenic activities which produce disturbances that fragment and decrease plant populations (Kéfi et al. 2007). The arbuscular mycorrhizal (AM) symbiosis is known to have an ecological and functional impact on plant performance in water-deficient and degraded habitats (Allen 2007), particularly in semi-arid ecosystems (Barea et al. 2011), and this impact must be considered in the restoration of these biomes.
AM fungi (AMF) associate with most (between 70 and 80 %) vascular plants (Smith and Read 2008; Brundrett 2009), improve their growth and nutrition (Barea et al. 2005) and promote ecosystem parameters such as plant diversity, productivity and soil aggregation (Grime et al. 1987; van der Heijden et al. 1998, 2006; Rillig and Mummey 2006; Barea et al. 2011). AMF can colonize individual roots as assemblages consisting of several AMF species and some host preference has been shown (Öpik et al. 2009). Since AMF can increase the success of establishment and survival of seedlings in the field and improve soil quality (Requena et al. 2001), they have been proposed as bio-enhancers in the restoration of populations of endangered plants (Barea et al. 2011). The reliance of vegetation stability with stability of AMF has been highlighted by several studies (Jeffries and Barea 2012) revealing the need to restore the original AMF populations as well as plant cover in order to regenerate degraded habitats.
Yew (Taxus baccata L.) is a plant species which typically experiences the stress situations characteristic of Mediterranean ecosystems in South-East Spain. It is widely distributed in northern and central European countries, but where the Mediterranean climate represents its southern limit, such as in Southern Iberia, it has been relegated to certain mountain areas (Thomas and Polwart 2003). This is due to the pressure of human practices (deforestation, overgrazing, increases in wildfire events, etc.) which usually have a higher impact on plants near to the boundaries of their distribution area (Pons 1981). These practices have caused yew populations to be reduced and fragmented (García et al. 2000). Since the natural regeneration of yews is limited (García et al. 2000), pro-active restoration of populations of yew is necessary to ensure their viability in Southern Spain. As this plant species has been demonstrated to form an AM symbiosis (Harley and Harley 1987), the impact of AMF on the restoration of yew populations needs consideration (Barea et al. 2011). However, information on the diversity of root-colonizing AMF associated with yew, and on potential changes in this community under anthropogenically-altered situations, is poorly understood.
Interest in the structure of AMF communities has increased recently and molecular tools are now available for a challenging dissection of AMF population dynamics (Robinson-Boyer et al. 2009) or for analyzing the distribution of AMF in ecosystems (Öpik et al. 2009). Since this group of organisms is cryptic, i.e. do not form taxonomically informative structures in roots, molecular methodologies are needed to study their presence inside plant roots. Terminal restriction fragment length polymorphism (TRFLP) of genes coding for ribosomal RNA has been widely used to assess AMF diversity (Dickie and FitzJohn 2007; Mummey and Rillig 2008; Antunes et al. 2009). This method was applied to identify the AMF community associated with individual yew trees since it provides a high throughput and reproducible profiling of samples containing complex microbial communities.
The aim of this study was to test whether or not the AMF community associated with yew trees was conserved on isolated individuals during yew population decline. Accordingly, two alternative hypotheses can be formulated: (a) the diversity of AMF is maintained in yew roots growing in degraded habitats as in those growing in better preserved conditions. In that case, it could be expected that these trees would act as refugia for a continuous and homogeneous assemblage of AMF even during their population decline. (b) Factors which drove yew populations to decline may have also altered AMF community composition, as shown by differences in the AMF communities between plants growing under different degraded conditions. In ascertaining which of these alternative hypotheses is true, different issues need to be tested: i) the existence of differences in the degradation levels of the habitats of each individual yew, as expressed by changes in soil properties, the physiological status of each tree and the composition of the surrounding vegetation and ii) the variation in the AMF community according to the conservation status of the habitat in which each individual plant lives.
The results of this study will help to the appropriate management of AMF in the restoration programs of yew in an area of particular ecological interest, the Sierra de Baza Natural Park (Granada province, Spain).
Material and methods
Study site and sampling procedure
The Sierra de Baza Natural Park has been historically managed by humans. Its use to feed cattle and the deforestation for the local use of wood were the main sources of degradation during last centuries. Since 1950, these mountains have been reforested for timber production. However the conformation of too dense areas of pines did not allow a good production of wood and their exploitation was abandoned. These wide mono-specific woodlands of pines replaced indigenous vegetation and increased the fire risk causing some fire events in recent years.
Under these circumstances, the population of Taxus baccata L. has been affected similarly as the rest of the vegetation of Sierra de Baza. However, the slow regeneration rate of yew trees makes them more vulnerable to the degradation of their natural habitats. Nowadays only more or less isolated trees persist in the highest area of the mountains in contrast with a more wide and homogeneous distribution in the past.
The studied population is located at altitudes between 1,800 and 2,000 m a.s.l., where the mean precipitation is about 600 mm per annum (Blanca and Morales 1991). The bedrock is calcareous forming litosols, regosols and cambisols. The population is situated within an area of 4 km2 (Fig. 1) with 23 trees, 13 of which were sampled. On July 2010, two subsamples of 5 g fine roots, at the north and south side of each tree, carefully tracing roots from the stem, were collected and immediately put on ice.
Spatial distribution of the sampled yews in the Sierra de Baza Natural Park. Triangles represent yews belonging to the first group (living in altered habitats) while squares represent the second group (growing in better conserved habitats). The dotted line is the road track and the continuous line represents altitude. Image source: Junta de Andalucía (2003)
Two subsamples of soil (500 g) and plant material (branches with leafs) were also collected for soil nutrient determination and Specific Leaf Area (SLA) measurements.
Vegetation analysis, plant physiological status and mycorrhizal colonization
The presence and cover of plant species in a 5 m radius surrounding the trees were determined. To assess the level of ecosystem degradation, the coverage of taxa indicating an alteration of the natural habitat of yew as well as taxa belonging to a well-conserved plant community growing with yew were determined following the description of habitat requirements by Tutin et al. (1980) and the description of the natural vegetation community of yew in García et al. (2000) (Supplementary Material Table S1). Shannon index (H′) and richness (S) were calculated to be used in subsequent analyses.
Specific leaf area (SLA) is a good indicator of potential relative growth rate, as individuals growing in resource-rich environments tend to have larger SLA than those in environments with limited resources (Cornelissen et al. 2003). Two distal branches per individual from the same aspect of each yew plant were collected and kept in a cool box. At the laboratory, ten leaves were removed, scanned with a portable scanner (Canoscan N656U, Canon) at 300 dpi, and the projected area measured (Midebmp v.4.2, Ordiales-Plaza 2000) to calculate leaf area. Due to their small size, leaves of each plant were measured together, first scanned and then weighed after drying at 72 °C for 48 h. Specific leaf area (SLA, m2 kg−1) was computed as the ratio between leaf area and mass. Average SLA was calculated for each yew.
Roots (2 g) from each subsample were cut in 1 cm pieces, mixed and stained (Phillips and Hayman 1970). The percentage of mycorrhizal colonization was determined by using the magnified intersection method (McGonigle et al. 1990) under a compound light microscope.
Soil and topological data
A subsample of 10 g of the collected soil was sieved to 2 mm and milled. Total soil nutrients (Table 1) were determined by the Servicio de Ionómica of Centro de Edafología y Biología Aplicada del Segura (CSIC) by ICP-OES (ICAP 6500 Duo/Iris Entrepid II XDL). An elemental analyzer C/N (Flash EA 1112 Series-Leco Truspec) was used to measure total and organic C and total N. Available P was also measured (Olsen et al. 1954). Soil pH was determined in a 1:2.5 (w/v) suspension of sieved soil in dH2O.
A Digital Elevation Model was provided by the Centro Nacional de Información Geográfica (2004) and processed using ArcGIS 9.2 (ESRI, 2006) to obtain topological data: latitude, longitude, altitude, slope, orientation and distances to the nearest yew and to a 3.5 m wide sandy forest road through the natural park.
Molecular methods
The remaining roots were frozen (−80 °C) in aliquots of 200 mg. DNA from 200 mg of roots per subsample was extracted using a DNeasy plant mini-kit (Qiagen Inc., Mississauga, ON, Canada) and eluted in 50 μl ddH2O.
The database TRFLP approach (see Dickie and FitzJohn 2007) was used to characterize the AMF community of yew. To construct a TRFLP database, sequences derived from AMF colonizing yew roots were produced. Partial ribosomal SSU DNA fragments were amplified using the primer set NS31 (Simon et al. 1992), a universal eukaryotic primer, and AML2 (Lee et al. 2008), a specific primer for AMF DNA amplification. Polymerase Chain Reactions (PCR) were carried out in a final volume of 25 μl using the Illustra Pure-Taq Ready-To-Go PCR beads (GE Healthcare UK Limited, Buckinghamshire, UK) and 20 μM of each primer. PCR conditions were: an initial minute at 94 °C, 30 cycles at 94 °C for 30 s, 52 °C for 30 s, and 72 °C for 40 s, followed by a final extension period at 72 °C for 5 min. As a template, 1 μl of extracted DNA was used in all reactions.
A equimolecular mix of all PCR products was purified using QIAquick Gel Extraction Kit (QIAGEN Iberia, Madrid, Spain) and cloned into the pCR®2.1 vector following the protocol recommended by the manufacturer of the TA Cloning® Kit (Invitrogen Life Technologies, Karlsruhe, Germany) and transformed into One Shot® TOP10F’ Chemically Competent Escherichia coli cells (Invitrogen Life Technologies). Ninety six clones were sequenced. The sequencing was done by the sequencing service of the Estación Experimental del Zaidín (Granada, Spain) using the set of vector binding primers M13F-M13R. Rarefaction analysis of the sequences was done using the Analytic Rarefaction 1.3 software (Holland 2008).
To obtain a good coverage of the AMF diversity present in the Sierra de Baza Natural Park and to complete the sequence database produced by cloning AMF sequences from the yew roots, additional sequences of AMF derived from roots, soil and spore samples from the Sierra de Baza Natural Park were added to the database (Sánchez-Castro et al. 2012a, b; Palenzuela et al. unpublished).
Sequences were aligned with MAFFT version 6 and similarities excluding the primer sequence were determined using BioEdit software. Phylogenetic analysis was carried out using the sequences obtained in the study and in the previous database of the Sierra de Baza Natural Park as well as a representative group of sequences of all major Glomeromycota groups from GenBank. The outgroup used was Morteriella polycephala X89436. The phylogenetic analysis was computed in MEGA4 (Tamura et al. 2007) using the neighbour-joining algorithm with Kimura-2 parameters as the model of substitution and 1,000 replications to obtain bootstrap values.
Phylotypes were determined using a minimum of 97 % sequence similarity and a high bootstrap value as indicators (Öpik et al. 2010). A blast search in the MaarjAM (Öpik et al. 2010) database was used to name the sequences with the number code of the closest virtual taxon when showing similarities higher than 97 %.
REPK online software (Collins and Rocap 2007) was used to select a set of four enzymes which allowed discrimination between the phylotypes found in the database. These enzymes were: MboI, VspI, TaqI and Bme1390I (New England Biolabs, Ipswich, USA). Sequences were subjected to in-silico T-RFLP analysis using TRiFLe (Junier et al. 2008).
PCR reactions were redone as described above with the exception of a fluorescent label (6-FAM) attached to the forward primer NS31. After purification of the PCR products using NucleoSpin Extract II (Macherey-Nagel GmbH, Düren, Germany), 80 ng of each amplified product were individually digested for 2 h at 37 °C with 1.5, 1.5, 3 and 6 units of MboI, Bme1390I, VspI and TaqI, respectively. Fragment lengths were determined at the SMB Service GmbH (Berlin, Germany) using a customized ROX size standard as marker size on a ABI 3100 capillary sequencer (Applied Biosystems Carlsbad, CA, USA). Data were processed using GeneMapper software version 3.7 (Applied Biosystems 2004) excluding fragments shorter than 50 bp and longer than 600 bp. The amplitude threshold was 20 RFU and the calling method “Local Southern Method”. The R (R Development Core Team 2010) package, TRAMPR (FitzJohn and Dickie 2007), was used with three nucleotides mismatch as threshold level for matching TRFLP profiles with the T-RFLP fragment sizes obtained during the in-silico analysis of the sequences of the constructed database.
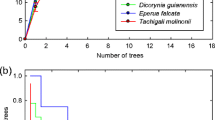
Since the data obtained from the TRFLP database approach consisted of a presence/absence matrix, only richness (S) was calculated. In order to assess the differences in phylotype composition in the two groups of yews identified in the study, the frequency of detection was calculated as the percentage of trees containing certain phylotypes. Species accumulation curves were also calculated by group to ensure that a similar coverage was obtained in the TRFLP approach.
Statistical analyses
In order to reduce the dimensionality of predictor variables, one PCA (Principal Component Analysis) per group of variables (plant, soil and topological related variables, see Table 1) was carried out. Since they represent the major part of variation, the first two axes of each PCA were selected to be used in the subsequent analyses. To ensure that collinearity between PCA axes of each variable set did not confuse the results, Pearson correlations were used to evaluate this possibility.
A cluster based on Euclidean distance was used to set up classes in terms of environmental conditions affecting the yews. Since the separation by this method was not totally clear, k-means was used to check the possibility of having two or three groups and to define them. Both clustering processes included the six PCA axes and were carried out using PAST 2.08 (Hammer et al. 2001). Mean values of the most relevant variables were also calculated to characterize the groups of yews found in the clustering and a t-test was carried out to check for significant differences between groups.
Effects of the six environmental axes on the AMF community were investigated by permutational multivariate analysis of variance (permanova, McArdle and Anderson 2001) embedded in the function adonis from vegan package in R (Oksanen et al. 2011), using Jaccard distance as a measure of community dissimilarity. This methodology was also applied to study differences in AMF community between both groups of yews.
In order to analyze the partial influence of each type of variable on the AMF community, a variance partitioning analysis (varpart function from vegan package) followed by a redundancy analysis (RDA) test for significance was used. Again, we only applied the first two axes of each explanatory PCA.
Results
Conservation status of the yew environments
Two groups of yews were defined by euclidean and k-means clustering (Fig. 1). They showed different levels of habitat degradation based on vegetation, soil characteristics and topological data (Table 2). Group 1 (six individuals) showed a greater degree of alteration with respect to group 2 (seven individuals), according to the accompanying vegetation (Table 2). Although no differences were found in the total coverage of well-conserved habitat indicators, the presence of Ononis aragonensis, a well-conserved habitat indicator, is significantly higher in the group of yews living in better conserved habitat. On the other side, Ononis pusilla and Galium verum, indicators of altered conditions, were more frequent in the group of yews with altered habitat. While no significant differences were found in the rest of indicator plant species, Santolina rosmarinifolia and Plantago lanceolata, altered conditions indicators, were absent surrounding yew plants growing in better conserved habitats.
Specific Leaf Area was found to be significantly higher in the group of yews from the better conserved habitat. Among soil data, pH and available P were found to be very similar for both groups but Ca, Mg, total N and organic C contents were higher in the soils corresponding to the better conserved habitats, however only significant differences were found for Ca (Table 2).
The distance between yews was almost ten times higher in the group developing in altered habitats (group 1). In contrast, the distance to the road track and the altitude were higher in the group of yews growing in better conserved habitats (group 2). Longitude had a similar pattern revealing the distribution of yews along a slope (Table 2).
Characterization of AMF community composition and influential environmental variables
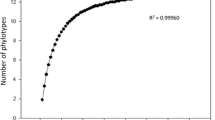
A total of 96 sequenced clones obtained from the pooled PCR products were analyzed and eight different phylotypes were found. Of these 96, 33 corresponded to non-specific amplifications of plant and saprotrophic fungi, and 63 belonged to glomeromycotan fungi. Through a rarefaction analysis, 12 phylotypes were predicted as the total in the mix of all PCR products, with 8 phylotypes representing 65.6 % (Fig. 2). The analysis also implied that it would be necessary to sequence 100 additional clones to increase the number of phylotypes by one and an overall of ~1,500 clones to cover the expected diversity.
The eight phylotypes found in the clone library were added to another 16 phylotypes from previous studies of Sierra de Baza Natural Park (Sánchez-Castro et al. 2012a, b; Palenzuela et al. unpublished). Thus, a total of 24 phylotypes composed the final TRFLP database. The TRFLP analysis found 17 of these phylotypes in the studied samples, belonging to seven glomeromycotan families: Acaulosporaceae (2 phylotypes), Claroideoglomeraceae (1), Diversisporaceae (1), Gigasporaceae (1), Glomeraceae (9), Pacisporaceae (1) and Paraglomeraceae (2) (Fig. S1).
Species accumulation curves based on the phylotypes presented by both groups of yews showed a similar coverage of the total predicted diversity (74.4 % for group 1 and 76.9 % for group 2) (Fig. 3).
The AMF community associated with yew in Sierra de Baza was significantly influenced by plant-related variables (axis 1, permanova: F = 0.23, p = 0.0149), topological variables (axis 1, F = 0.12, p = 0.0499) and soil characteristics (axis 2, F = 0.17, p = 0.033). Pearson correlations between the first two PCA axes of each variable set and the rest of PCA axes were not important (less than 0.6 or higher correlation with a PCA axis with a low percentage —less than 5 %— of explained variance). This avoids the possibility of misinterpreting the results due to the influence of PCA axes not included in the analysis.
Analysis of variance partition revealed a higher influence of plant related variables (R2 = 0.34, F = 3.08, p = 0.005), followed by topological ones (R2 = 0.16, F = 1.96, p = 0.025) and soil characteristics (R2 = 0.14, F = 1.85, p = 0.05), with a total of 64 % explained variance in the whole analysis. Mean values for the most influential variables on the AMF communities, which correspond to variables related to significant axes of each PCA with a correlation value higher than 0.5, are shown in Table 2.
Effects of habitat degradation on AMF community
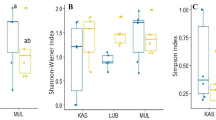
Mycorrhizal colonization data (around 20 % in average) did not show significant differences between the groups of trees (Table 2). Comparisons of the AMF community composition between the two groups of yews revealed a similar richness of phylotypes per tree. However, the number of glomeromycotan families detected in the yews growing in the better conserved habitats was significantly higher than that in those with altered habitats (Table 2).
The frequency of detection of AMF phylotypes in each group of yews was clearly different (Table 3). The most abundant phylotype was Div060, present in every tree studied. Another ten phylotypes were shared between both yew groups, although they showed very different frequencies for group 1 and 2. Two of these 10 phylotypes (Cla193 and Par336) were very common in yew group 2, appearing in 85.7 % of the trees, while in group 1 these phylotypes never had a frequency higher than 33.3 %. The other eight shared phylotypes (Aca029, Aca285, Glo065, Glo072, Glo143, Glo149, Glo177 and Par281) had a more similar distribution in both groups. Finally three phylotypes were exclusive to group 1 (Glo153, Glo222 and Glo319) and three exclusive to Group 2 (Gig052, Glo312 and Pac284). However the frequency of these was low, being present only in one or two trees of the group. Despite these differences, PERMANOVA did not show significant differences in the whole community of AMF.
At the family level, some glomeromycotan taxa were absent in the yews living in the more altered habitats. Group 1 showed the presence of members of Glomeraceae, Claroideoglomeraceae, Diversisporaceae, Acaulosporaceae and Paraglomeraceae. In contrast, group 2 was colonized by members of all detected families (Glomeraceae, Claroideoglomeraceae, Diversisporaceae, Acaulosporaceae, Paraglomeraceae, Gigasporaceae and Pacisporaceae). Moreover the frequencies of detection of some families varied widely between groups of yews (Fig. 4), e.g. Claroideoglomeraceae and Paraglomeraceae were detected only in 33.3 % of the yews in group 1 while they appeared in more than 80 % of the members of group 2.
Discussion
Disturbance effects on plant and AMF assemblages
A relationship between the environmental variables (vegetation, plant and soil characteristics and topological data) and the level of habitat degradation and AMF community composition was evident. In particular, the percent coverage of plants characteristic of altered habitats relative to those of well-conserved habitats surrounding the yews indicated two degradation levels for the studied system: Group 1 includes yews living in a more altered habitat while group 2 contains yews having a better conserved habitat. The physiological state of the yews, as measured by SLA, supports this pattern. Thus, a parallel effect of degradation on both plant and AMF communities was evident. Fitzsimons et al. (2008) also found relationships between the time after disturbance, structure of the plant cover and AMF communities. Soil variables, such as mineral nutrient and organic C contents, were also closely related to the extent of plant cover and AMF community conservation, as found for other ecosystems (Requena et al. 2001; Schnoor et al. 2011). As a general trend, degradation of vegetation cover is concomitant with a reduction of AMF diversity (as reviewed by Barea et al. 2011).
The presence of a road track seems to be an important source of disturbances in the study area. In fact, the distance to the road track appeared as a very influential variable. Previous work has shown that even a low number of vehicles driving through the forest can affect the surrounding vegetation (Angold 1997; Truscott et al. 2005). Some of these studies have highlighted the increase of ruderal plant species near roads, as also found in the present study. Trampling by visitors also has an impact on plant communities and soil microbiota (Malmivaara-Lämsä et al. 2008; Lucas-Borja et al. 2011). Additionally, trampling by cattle is probably an important source of disturbance since this road track is regularly used to reach the highest grasslands of the Natural Park. The cattle also contribute to degradation by overgrazing (Blanca and Morales 1991). Other human practices provoking environmental degradation are the fire prevention measures at the edge of the roads, a labour consisting mainly in the clearing of the understory cover, which is necessary due to the dry and hot summers in Mediterranean environments. Clearing the understory is known to affect microbial activities (Mummey et al. 2010).
Altitude and spatial coordinates (longitude) appear to have a similar effect as the distance to the road. This could be due to the position of the road, which runs north–south leaving in the west the top of the mountain. Consequently, it is not possible to separate the effects of both variables. Differences in altitude are not large (about 200 m) which minimize changes in climatic conditions. In turn, the spatial effect can be ruled out as well since small distances separate both yew groups and neither geographical barriers nor changes in topological or geologic features were found between them.
In relation to AMF community composition, the absence of some glomeromycotan families in the roots of yews living in altered habitats is remarkable. The exclusive presence of Gigasporaceae and Pacisporaceae in the better conserved habitat group along with the higher frequencies of Claroideoglomeraceae and Paraglomeraceae, supports that more simple communities in disturbed environments are dominated by species of Glomeraceae, as also found in other studies (Egerton-Warburton and Allen 2000). However, no statistical differences in the AMF community could be found between groups of yew trees. This can be due to the interaction with other variables not clearly related to degradation. For example the available phosphorus, which appears as a determinant of the AMF community, was not different between the two groups of yews. As this variable is typically influencing the effectiveness of the AM symbiosis (Johnson 2010) and probably also the AMF community composition, this variable can thereby lead to blur the sole influence of disturbance factors.
Yew as inoculum source for restoration
An important objective of this study was to assess the possibility that yew trees could act as a refuge for the AMF community, and that these trees could thereby play a role in providing AMF inoculum for the surrounding vegetation. This would represent an important contribution in ecological restoration programs. This hypothesis was based on the importance of woody plants, particularly in Mediterranean ecosystems, acting as resource islands (Allen 1988; Azcón-Aguilar et al. 2003; Caravaca et al. 2005). Yew seemed a good candidate for this role due to its long lifespan (hundreds of years) and because its roots provide an extensive habitat for AMF. If this were the case, yew trees would maintain their original AMF communities while the surrounding environments were affected by degradation. When the disturbance processes were reduced, the progress towards more advanced succession stages could be driven by the presence of this preserved AMF inoculum. However, the hypothesis supporting that yew roots can act as refugia for their original AMF communities should be rejected since our results, showing that AMF phylogenetic diversity was negatively affected by degradation, are in favour of the alternative hypothesis. A similar situation was reported in studies on the effect of habitat fragmentation on different fungal groups (Peay et al. 2007; Grilli et al. 2012) and on AMF in disturbed (agricultural) environments (Jansa et al. 2003; Oehl et al. 2009). This is critical, since a higher phylogenetic diversity can imply a higher functional diversity (Powell et al. 2009), with a parallel increase in AMF life strategies (Ijdo et al. 2010). Consequently, a lack of a phylogenetically diverse AMF community can obstruct the development of late-seral plant species, since they can need the association with specific groups of AMF (Davison et al. 2011), and difficult the progress of natural succession.
Conclusion
AMF communities associated with Taxus baccata L. are affected by the environmental processes involved in its population decline. The source of disturbance is related, at least in part, to anthropogenic activities as shown by a clear effect of the road track in the study area. Vegetation, plant and soil characteristics and topological data indicated that the studied yews fell into two different degradation levels. The group of yews living in the better conserved habitats had higher AMF phylogenetic diversity. The lack of some families of AMF in the group of yews developing in the more altered habitats can imply a loss of functional diversity, highlighting the necessity of reintroducing these groups through the appropriate management of AMF in the restoration programs.
Abbreviations
- AM:
-
Arbuscular mycorrhizal
- AMF:
-
Arbuscular mycorrhizal fungi
- PCA:
-
Principal component analysis
- PCR:
-
Polymerase chain reaction
- RDA:
-
Redundancy analysis
- SLA:
-
Specific leaf area
- TRFLP:
-
Terminal restriction fragment length polymorphism
References
Allen MF (1988) Belowground structure: A key to reconstructing a productive arid ecosystem. In: Allen EB (ed) Reconstruction of disturbed arid ecosystems. Westview Press, Boulder, pp 113–135
Allen MF (2007) Mycorrhizal fungi: highways for water and nutrients in arid soils. Vadose Zone J 6:291–297
Angold PG (1997) The impact of a road upon adjacent heathland vegetation: effects on plant species composition. J Appl Ecol 34:409–417
Antunes PM, Koch AM, Dunfield KE, Hart MM, Downing A, Rillig MC, Klironomos JN (2009) Influence of commercial inoculation with Glomus intraradices on the structure and functioning of an AM fungal community from an agricultural site. Plant Soil 317:257–266
Azcón-Aguilar C, Palenzuela J, Roldán A, Bautista S, Vallejo R, Barea JM (2003) Analysis of the mycorrhizal potential in the rizhosphere of representative plant species from desertification-threatened Mediterranean shrublands. Appl Soil Ecol 22:29–37
Barea JM, Pozo MJ, Azcón R, Azcón-Aguilar C (2005) Microbial co-operation in the rhizosphere. J Exp Bot 417:1761–1778
Barea JM, Palenzuela J, Cornejo P, Sánchez-Castro I, Navarro-Fernández C, López-García A, Estrada B, Azcón R, Ferrol N, Azcón-Aguilar C (2011) Ecological and functional roles of mycorrhizas in semi-arid ecosystems of Southeast Spain. J Arid Environ 75:1292–1301
Blanca G, Morales C (1991) Flora del parque natural de la Sierra de Baza. University of Granada, Granada
Brundrett MC (2009) Mycorrhizal associations and other means of nutrition of vascular plants: understanding the global diversity of host plants by resolving conflicting information and developing reliable means of diagnosis. Plant Soil 320:37–77
Caravaca F, Alguacil MM, Barea JM, Roldán A (2005) Survival of inocula and native AM fungi species associated with shrubs in a degraded Mediterranean ecosistema. Soil Biol Biochem 37:227–233
Centro Nacional de Información Geográfica (2004) MDT200. Plan Nacional de Ortofotografía Aérea. Ministerio de Fomento, Spain
Collins RE, Rocap G (2007) REPK: an analytical web server to select retriction endonucleases for terminal restriction fragment length polymorphism analysis. Nucleic Acids Res 35:W58–W62. doi:10.1093/nar/gkm384
Cornelissen JHC, Lavorel S, Garnier E, Díaz S, Buchmann N, Gurvich DE, Reich PB, ter Steege H, Morgan HD, van der Heijden MGA, Pausas JG, Poorter H (2003) A handbook of protocols for standardised and easy measurement of plant functional traits worldwide. Aust J Bot 51:335–380
Davison J, Öpik M, Daniell TJ, Moora M, Zobel M (2011) Arbuscular mycorrhizal fungal communitites in plant roots are not random assemblages. FEMS Microbiol Ecol 78:103–115
Dickie IA, FitzJohn RG (2007) Using terminal restriction fragment length polymorphism (T-RFLP) to identify mycorrhizal fungi: a methods review. Mycorrhiza 17:259–270
Egerton-Warburton LM, Allen EB (2000) Shifts in arbuscular mycorrhizal communities along an anthropogenic nitrogen deposition gradient. Ecol Appl 10:484–496
FitzJohn RG, Dickie IA (2007) TRAMPR: an R package for analysis and matching of terminal-restriction fragment length polymorphism (TRFLP) profiles. Mol Ecol Notes 7:583–587
Fitzsimons MS, Miller RM, Jastrow JD (2008) Scale-dependent niche axes of arbuscular mycorrhizal fungi. Oecologia 158:117–127
García D, Zamora R, Hódar JA, Gómez JM, Castro J (2000) Yew (Taxus baccata L.) regeneration is facilitated by fleshy-fruited shrubs in Mediterranean environments. Biol Conserv 95:31–38
Grilli G, Urcelay C, Galetto L (2012) Forest fragment size and nutrient availability: complex responses of mycorrhizal fungi in native-exotic hosts. Plant Ecol 213:155–165
Grime JP, Mackey JML, Hillier SH, Read DJ (1987) Floristic diversity in a model system using experimental microcosms. Nature 328:420–422
Hammer O, Harper DAT, Ryan PD (2001) PAST: Paleontological Statistics Software Package for education and data analysis. Palaeontol Electron 4
Harley JL, Harley EL (1987) A check-list of mycorrhiza in the British flora. New Phytol (Suppl) 105:1–102
Holland SM (2008) Analytic Rarefaction 1.3. http://strata.uga.edu/software/anRareReadme.html
Ijdo M, Schtickzelle N, Cranenbrouck S, Declerk S (2010) Do arbuscular mycorrhizal fungi with contrasting life-history strategies differ in their responses to repeated defoliation? FEMS Microbiol Ecol 72:114–122
Jansa J, Mozafar A, Kuhn G, Anken T, Ruh R, Sanders IR, Frossard E (2003) Soil tillage affects the community structure of mycorrhizal fungi in maize roots. Ecol Appl 13:1164–1176
Jeffries P, Barea JM (2012) Arbuscular mycorrhiza - a key component of sustainable plant-soil ecosystems. In: Hock B (ed) The mycota, vol IX. Fungal Associations, 2nd edn. Springer-Verlag, Berlin, Heidelberg (in press)
Johnson NC (2010) Resource stoichiometry elucidates th estructure and function of arbuscular mycorrhizas across scales. New Phytol 185:631–647
Junier P, Junier T, Witzel KP (2008) TRiFLe, a program for in silico terminal restriction fragment length polymorphism analysis with user-defined sequence sets. Appl Environ Microbiol 74:6452–6456
Junta de Andalucía (2003) Ortofotografía digital de Andalucía, provincia de Granada. Junta de Andalucía, Sevilla
Kéfi S, Rietkerk M, Alados CL, Pueyo Y, ElAich A, Papanastasis V, de Ruiter PC (2007) Spatial vegetation patterns and imminent desertification in Mediterranean arid ecosystems. Nature 449:213–218
Krüger M, Krüger C, Walker C, Stockinger H, Schüssler A (2012) Phylogenetic reference data for systematics and phylotaxonomy of arbuscular mycorrhizal fungi from phylum to species level. New Phytol 193:970–984
Lee J, Lee S, Young PW (2008) Improved PCR primers for the detection and identification of arbuscular mycorrhizal fungi. FEMS Microbiol Ecol 65:339–349
Lucas-Borja ME, Bastida F, Moreno JL, Nicolás C, Andres M, López FR, del Cerro A (2011) The effects of human trampling on the microbiological properties of soil and vegetation in mediterranean mountain areas. Land Degrad Dev 22:383–394
Malmivaara-Lämsä M, Hamberg L, Haapamäki E, Liski J, Kotze DJ, Lehvävirta S, Fritze H (2008) Edge effects and trampling in boreal urban forest fragments – impacts on the soil microbial community. Soil Biol Biochem 40:1612–1621
McArdle BH, Anderson MJ (2001) Fitting multivariate models to community data: a comment on distance-based redundancy analysis. Ecology 82:290–297
McGonigle TP, Miller MH, Evans DG, Fairchild GL, Swan JA (1990) A new method which gives an objective measure of colonization of roots by vesicular-arbuscular mycorrhizal fungi. New Phytol 115:495–501
Mummey DL, Rillig MC (2008) Spatial characterization of arbuscular mycorrhizal fungal molecular diversity at the submetre scale in a temperate grassland. FEMS Microbiol Ecol 64:260–270
Mummey DL, Clarke JT, Cole A, O’Connor BG, Gannon JE, Ramsey PW (2010) Spatial análisis reveals differences in soil microbiol community interactions between adjacent coniferous forest and clearcut ecosystems. Soil Biol Biochem 42:1138–1147
Oehl F, Sieverding E, Ineichen K, Mäder P, Wiemken A, Boller T (2009) Distinct sporulation dynamics of arbuscular mycorrhizal fungal communities from different agroecosystems in long-term microcosms. Agric Ecosyst Environ 134:257–268
Oksanen J, Blanchet FG, Kindt R, Legendre P, Minchin PR, O’Hara RB, Simpson GL, Solymos P, Henry M, Stevens H, Wagner H (2011) Vegan: Community ecology package. R package version 2.0-1. http://CRAN.R-project.org/package=vegan
Olsen SR, Cole CV, Watanabe FS, Dean LA (1954) Estimation of available phosphorus in soils by extraction with sodium bicarbonate. USDA Circular 939. US gov Print Office, Washington DC
Öpik M, Metsis M, Daniell TJ, Zobel M, Moora M (2009) Large-scale parallel 454 sequencing reveals host ecological group specifity of arbuscular mycorrhizal fungi in a boreonemoral forest. New Phytol 184:424–437
Öpik M, Vanatoa E, Moora M, Davison J, Kalwij JM, Reier Ü, Zobel M (2010) The online database MarrjAM reveals global and ecosystemic distribution patterns in arbuscular mycorrhizal fungi (Glomeromycota). New Phytol 188:223–241
Ordiales-Plaza R (2000) Midebmp, Version 4.2. Estación Experimental de Zonas Áridas (CSIC), Almería
Peay KG, Bruns TD, Kennedy PG, Bergemann SE, Garbelotto M (2007) A strong species-area relationship for eukariotic soil microbes: island size matters for ectomycorrhizal fungi. Ecol Lett 10:470–480
Phillips JM, Hayman DS (1970) Improved procedures for clearing roots and staining parasitic and vesicular-arbuscular mycorrhizal fungi for rapid assessment of infection. Trans Br Mycol Soc 55:158–161
Pons A (1981) The History of the Mediterranean shrublands. In: Di Castri F, Goodall DW, Specht RL (eds) Mediterranean-type shrublands. Elsevier Academic Press, Amsterdam, pp 131–138
Powell JR, Parrent JL, Hart MM, Klironomos JN, Rillig MC, Maherali H (2009) Phylogenetic trait conservatism and the evolution of functional trade-offs in arbuscular mycorrhizal fungi. Proc R Soc B Biol Sci 276:4237–4245
R Development Core Team (2010) R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna
Requena N, Perez-Solis E, Azcón-Aguilar C, Jeffries P, Barea JM (2001) Management of indigenous plant-microbe symbioses aids restoration of desertified. Appl Environ Microbiol 67:495–498
Rillig MC, Mummey DL (2006) Mycorrhizas and soil structure. New Phytol 171:41–53
Robinson-Boyer L, Grzyb I, Jeffries P (2009) Shifting the balance from qualitative to quantitative analysis of arbuscular mycorrhizal communities in field soils. Fungal Ecol 2:1–9
Sánchez-Castro I, Ferrol N, Barea JM (2012a) Analyzing the community composition of arbuscular mycorrhizal fungi colonizing the roots of representative shrubland species in a Mediterranean ecosystem. J Arid Environ 80:1–9
Sánchez-Castro I, Ferrol N, Cornejo P, Barea JM (2012b) Temporal dynamics of arbuscular mycorrhizal fungi colonizing roots of representative shrub species in a semi-arid Mediterranean ecosystem. Mycorrhiza 22:449–460
Schnoor TK, Lekberg Y, Rosendahl S, Olsson PA (2011) Mechanical soil disturbance as a determinant of arbuscular mycorrhizal fungal communities in semi-natural grassland. Mycorrhiza 21:211–220
Simon LM, Lalonde TD, Bruns TD (1992) Specific amplification of 18S fungal ribosomal genes from vesicular arbuscular endomycorrhizal fungi colonising roots. Appl Environ Microbiol 58:291–295
Smith SE, Read DJ (2008) Mycorrhizal symbiosis, 3rd edn. Elsevier Academic Press, New York
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Thomas PA, Polwart A (2003) Taxus baccata L. J Ecol 91:489–524
Truscott AM, Palmer SCF, McGowan GM, Cape JN, Smart S (2005) Vegetation composition of roadside verges in Scotland: the effects of nitrogen deposition, disturbance and management. Environ Pollut 136:109–118
Tutin TG, Heywood VH, Burges NA, Moore DM, Valentine DH, Walters SM, Webb DA (eds) (1980) Flora Europaea, vols. 1-5. Cambridge University Press, Cambridge
van der Heijden MGA, Klironomos JN, Ursic M, Moutoglis P, Streitwolf-Engel R, Boller T, Wiemken A, Sanders IR (1998) Mycorrhizal fungal diversity determines plant biodiversity, ecosystem variability and productivity. Nature 396:69–72
van der Heijden MGA, Streitwolf-Engel R, Riedl R, Siegrist S, Neudecker A, Ineichen K, Boller T, Wiemken A, Sanders IR (2006) The mycorrhizal contribution to plant productivity, plant nutrition and soil structure in experimental grassland. New Phytol 172:739–752
Acknowledgments
A. López-García thanks the Formación de Personal Investigador Programme (Ministerio de Ciencia e Innovación, Spain) for financial support. This research was supported by the Spanish Goverment under the Plan Nacional de I+D+I (project CGL-2009-08825). We strongly thank Dr. I. Sánchez-Castro for the permission to use the Sierra de Baza AMF database created during his PhD Thesis. We also sincerely thank Professor P. Jeffries (Univ. of Kent) for grammatical corrections to the manuscript. We thank the Consejería de Medio Ambiente, Junta de Andalucía (Spain) for permission to work in Sierra de Baza Natural Park and to Drs. F. Bruno and J. Molero for helping us to identify and interpret vegetation inventories. Additionally, we would like to thank the two anonymous reviewers and the Section Editor for their valuable comments and suggestions to improve the manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible Editor: Thom W. Kuyper.
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(DOC 19 kb)
Fig. S1
Neighbor-joining phylogenetic tree based on the NS31-AML2 fragment of the SSU rDNA gene. Sequences from single AMF spores and root samples from the Sierra de Baza Natural Park are showed together with reference sequences from GeneBank. Numbers above branches indicate the bootstrap values. Only topologies with values ≥50 % are shown (1,000 replicates). Sequences are labelled according to the data set from which it originated (Tb-root = obtained from T. baccata roots; SB-root = from roots of other plants characteristic of the site; SB-spore = from spores isolated from the Sierra de Baza Natural Park), followed by the clone identity number. Sequences having a pairwise similarity higher than 97 % were clustered as phylotypes (delimited by vertical lines). Phylotypes are named following the closest virtual taxa code of MaarjAM database (Öpik et al. 2010). The prefix corresponds to the glomeromycotan family (following Krüger et al. 2012): Aca-Acaulosporaceae, Glo-Glomeraceae, Cla-Claroideoglomeraceae, Par-Paraglomeraceae, Pac-Pacisporaceae, Div-Diversisporaceae and Gig-Gigasporaceae. Mortierella polycephala was used as out-group. To reduce the size of the tree, half of the sequences were removed (JPEG 78 kb)
Rights and permissions
About this article
Cite this article
López-García, Á., Hempel, S., de D. Miranda, J. et al. The influence of environmental degradation processes on the arbuscular mycorrhizal fungal community associated with yew (Taxus baccata L.), an endangered tree species from Mediterranean ecosystems of Southeast Spain. Plant Soil 370, 355–366 (2013). https://doi.org/10.1007/s11104-013-1625-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11104-013-1625-0