Abstract
Little is known about the community dynamics of fungi on decomposing fine roots, despite the importance of fine roots as a source of carbon to detrital systems in forests. We examined fungal communities on dead roots in a sugar-maple dominated northern hardwood forest to test the hypothesis that community development is sensitive to rhizosphere disruption. We generated cohorts of dead fine roots in root windows and disturbed the rhizosphere microbial community in half of the windows by moving roots into sieved bulk soil. We sampled root fragments repeatedly over time and cultured fungi from these fragments to explore temporal patterns of fungal species composition. Disturbing the root rhizosphere prior to initiating decomposition changed the dominant fungal taxa, the distribution of dominant species within the community, and the temporal development in the culturable fungal community. Dominance in control roots shifted from Neonectria in early decay to Umbelopsis in later decay. Disturbance roots were more evenly dominated over time by Trichoderma, Neonectria, another species of Umbelopsis, and Pochonia. Our results suggest that species interactions are important in the ecology of fine root decay fungi, with the rhizosphere community of the living root influencing development of the decay community.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Litter inputs supplying detrital food webs in forest ecosystems include foliage, fine roots and woody debris (stems and woody roots). In comparison with leaf and woody litter, the organisms involved in decay of fine roots have received limited attention in part because of challenges of measurement. Fundamental differences in the nature of these detrital types, including substrate chemistry (Harmon et al. 1986; Hughes and Fahey 1994; Hobbie et al. 2010), the environment surrounding the substrates (Lodge and Cantrell 1995), and the timing and spatial arrangement of deposited litter, could influence microbial communities mediating the decay process. For example, whereas leaf litter is arrayed in a relatively uniform layer on the soil surface (Gessner et al. 2010), woody litter (Boddy 1992; Jönsson et al. 2008) and fine root litter inputs occur sporadically on and in the soil. In addition, the environment surrounding roots changes little following root death whereas aboveground litter moves from the atmospheric to the soil environment. These contrasts are likely to influence the dynamics of the microbial communities involved in litter decay. In the present study, we examined culturable fungi on decomposing sugar maple (Acer saccharum Marsh.) fine roots in a northern hardwood forest to gain new insights into decomposer communities in this little-studied detrital substrate.
Examining community development on fine roots can extend our understanding of the ecology of plant litter communities. It is not clear whether a distinct functional group of “root decomposer” fungi exists, or whether any heterotrophic fungi in soil are potentially involved in fine root decay. This should depend on colonizing dynamics and on factors influencing subsequent community development in fine root substrates. Colonization of root litter compared to aboveground litter will likely differ because of the relative importance of colonization by mycelia vs airborne spores in aboveground vs belowground substrates. The discrete nature and small size of fine roots within the soil medium is another fundamental difference that should contribute to variation in colonizing dynamics. While the pool of fungal mycelia in soil is diverse (Domsch et al. 2007), the scale of its distribution relative to the individual fine roots resources must be considered (Huston 1999), and suggests that only a relatively small subset of species have access to any one root substrate. Hence, colonization could be highly stochastic, depending upon the chance encounter between a root and a hypha growing through the soil. On the other hand, if species interactions play a role, community development could be a more structured process in which the fungal community in the rhizosphere or the living root influences the decay community. If some of the potential colonizing species differ in persistence or competitive abilities, as seen for instance in wood (Coates and Rayner 1985; Holmer and Stenlid 1996), or ectomycorrhizal fungi (Kennedy et al. 2009), then variable “priority effects” could lead communities to develop in different trajectories (Fukami et al. 2010).
Timing of community development may be especially important in fine roots because of the continuity of substrate environment as the root transitions from living tissue to detritus. The rhizosphere that develops over the lifetime of the root remains intact as the root dies, in contrast to the environments surrounding wood and leaf litter. Living roots are colonized by a wide diversity of fungi in addition to mycorrhizal fungi (Vandenkoornhuyse et al. 2002), including many culturable genera such as Cylindrocarpon, Mortierella, Nectria, Phialocephala, Pochonia, and Trichoderma (Addy et al. 2005; Germino et al. 2006; Menkis et al. 2006; Kwaśna et al. 2008). Many of these persist after root death, and the spectrum of culturable fungi in living roots overlaps with that in dead roots (Halmschlager and Kowalski 2004). Furthermore, some of the fungi that enter the rhizosphere may be connected to broader hyphal networks that have developed from the bulk soil over time. Hence, the species pool from which colonization takes place and the species that are potentially involved in community interactions are likely to depend upon the timing of rhizosphere formation.
Processes of community development, including the relative importance of stochastic and interactive processes, have received recent attention in microbial communities. The importance of chance or random occurrences varies among studies (Sloan et al. 2006; Horner-Devine et al. 2007; Woodcock et al. 2007; Fischer et al. 2009), and are of general interest for comparing communities among litter types. The net outcome of these processes is also of functional relevance in fine root substrates, if potential members of the fungal community vary widely in their ability to metabolize plant structural carbon. By analogy, in woody substrates substantial variation in decay can arise from differences in fungal communities and the effectiveness among species of wood breakdown. Heterogeneity in the community and also subsequent decay rate can arise from the stochastic component of colonization (Boddy et al. 1989). However, species interactions are also crucial to community dynamics over time in these woody substrates, and contribute to differences in decay (Boddy et al. 1989; Boddy 2000; Fukami et al. 2010). For example, Fukami et al. (2010) found that mass loss from decaying wood varied from about 10% to 40% depending on the initial colonizing fungal species. In fine roots stochastic processes might be expected to contribute to diverse communities that are highly variable among individual roots, which, in turn, could contribute to the high variation in decay found among individual roots (Fahey et al. 1988; Hendrick and Pregitzer 1992; Fahey and Hughes 1994; Ruess et al. 1998).
Disturbance experiments can be used to differentiate among processes of community development (Ellwood et al. 2009). Studies of fine root decay in litterbags, representing rather extreme disturbances to the rhizosphere, have yielded intriguing results, including unexpectedly low decomposition rates (McClaugherty et al. 1982; Fahey 1992; Fahey and Arthur 1994; Majdi and Nylund 1997; Dornbush et al. 2002). Disturbance might not be expected to alter decay rates if the decomposer communities develop via stochastic processes. An alternative explanation is that an intact rhizosphere community influences the community of decay fungi in roots, through a combination of initial colonization and subsequent species interactions. No previous research to evaluate this idea has been reported. Therefore, in this study we used a disturbance experiment to examine fungal community change over time on decomposing fine roots at the Hubbard Brook Experimental Forest in NH. We hypothesized that the disruption of the rhizosphere alters fungal community development. Our specific objectives were to: 1) describe the culturable fungal community on decomposing fine roots, 2) quantify changes in the community over time, and 3) test effects of rhizosphere disturbance on community composition. We hoped to gain a better understanding of the overall process of community development on fine root substrates, especially in relation to changes in the rhizosphere.
Methods and materials
Study site
This study was conducted at the Hubbard Brook Experimental Forest (HBEF) in north-central New Hampshire, USA (43° 56′N, 71° 45′W). The forest is second-growth northern hardwoods, which arose following clearcut harvest from 1910 to 1920 (Likens et al. 1985). Our study sites were located in the mid-elevation zone (approximately 590 m elevation) on a south-facing slope immediately west of Watershed 6 and just above the long-term litterfall collection sites described in Fahey and Hughes (1994) and Bohlen et al. (2001). We selected four plots, approximately 30 × 30 m and within 300 m distance of one another, with sugar maple (Acer saccharum Marsh.) in the overstory and some American beech (Fagus grandifolia Ehrh.) in the understory. Plots were subjectively chosen to maximize similarity in tree species composition, and to ensure that all were on moderate slopes with good drainage. Soils are mostly acidic spodosols (typic and aquic Haplorthods) of sandy loam texture with a surface organic horizon of approximately 5 cm depth.
Root windows
We installed two root windows at 3–5 m distance in each site in fall 2000. Each window consisted of a 30 × 30 cm sheet of Lexan glass fitted against a vertical undisturbed soil face that had been exposed by cutting with a 0.4 m-wide sharpened steel plate. We excavated a hole in front of each soil face to allow access, and we secured the window by constructing a tight-fitting insulated box (40 cm × 50 cm × 40 cm depth) in the hole. Visual evidence indicated that we were successful in installing these windows without significantly disrupting the soil adjacent to the windows. Root growth against windows was traced on acetate sheets through spring 2002, at which time we identified six 1-year-old root networks per window. These root networks were located approximately 5–15 cm depth from the forest floor surface and, while the main axis roots were often larger, all of the first and second order root branches that were sampled were < 0.5 mm diameter. Roots were most likely sugar maple, based on overstory dominance by this tree species and the absence of any obvious ectomycorrhizal associations. One window of each site was randomly designated as a control and the other was designated as a disturbance treatment window. To establish disturbance, we opened the window, removed the six mapped root networks and gently cleaned the roots of adhering rhizosphere soil. These roots were then placed back on the inside wall of the window, between 5 and 15 cm depth, and the window was returned to its place and backfilled with a 1–2 cm thick layer of sieved and uniform mineral soil mixture collected from a depth of 5–20 cm in the study sites. Disturbance roots were in close contact with both soil and Lexan windows, similar to control roots; hence, the principal difference between disturbance and control roots was the removal of rhizosphere soil and disruption of soil mycelial networks.
At the same time as the disturbance treatment was established, we killed the six mapped root networks in the control windows by drilling through the window and cutting the main root axis. We sampled each root network in control and disturbance windows 4 times, at 1, 3, 6, and 13 months after cutting the axis roots. To collect a sample we drilled next to a root branch (<0.5 mm diameter) and clipped a fragment (approximately 2 cm) from the branch. Subsequent samples were taken from branches in close proximity to each other (Fig. 1). The same procedure was used to collect roots in both control and disturbance treatments. Root fragments were stored in sterile vials on ice in a cooler until we processed them in the laboratory, within 12 hours.
Soil core collection
In June 2002 we collected dead fine roots from soil cores to evaluate the composition of root decay fungal communities without root-window effects. We established a 50-m transect between sites 1 and 2 (above) and randomly selected 15 sample points along the transect. At each sample point we extracted 3 soil cores (5-cm diameter including 10-cm depth in mineral soil; approximately 15 cm depth from the surface of the forest floor) and divided each core into organic (forest floor), A, and B horizons. For each sample point, soil from multiple cores was composited into one individual sample per horizon. Fine roots were removed from these samples in the laboratory and separated into living and dead categories based on visual criteria, including color, condition of the cortex, and elasticity. Up to 6 dead roots were taken per horizon per sample point and from each of these we processed a fragment approximately 2 cm long. This yields a maximum total of 36 fragments per sample point; however, numbers of root fragments was substantially less because we could not always find 6 dead roots in a composited horizon sample, and because A horizons were present at only about half of the sample points.
Root processing and fungal culturing
We removed soil particles and organic debris from root fragments of window and core samples under a dissecting microscope using sterile tweezers. Cleaned fragments were shaken vigorously (60 s) in 3 subsequent 10-mL aliquots of sterile water to remove surface fungi. We cut each washed fragment from root windows into six c. 1/3 cm long sub-fragments and placed three of these on a malt agar plate containing benomyl and the other 3 on a malt agar plate without benomyl. Each washed fragment from soil cores was cut into 3 sub-fragments and placed on a malt agar plate containing benomyl. Benomyl inhibits the growth of some ascomycete fungi, and we included it to aid selection for basidiomycete fungi (Thorn et al. 1996).
Malt agar was prepared by autoclaving, in 1 L distilled water, 0.5 g KH2PO4, 0.2 g MgSO4 .7H2O, 0.1 g NH4NO3, 0.1 g KCl, 0.2 g FeSO4 .7H2O, 0.05 g Ca(NO3)2 .4H2O, 2 g Difco malt extract (Difco Laboratories Inc, Detroit MI), and 15 g Difco agar. After cooling agar to approximately 55°C, we aesceptically added to each liter the following filter-sterilized solutions: 5 mL 1 M KOH, and 1 mL each of 60 mg/mL chlorotetracycline-HCl, 30 mg/mL streptomycin sulfate, and 30 mg/mL penicillin G (Na salt), and mixed gently with an autoclaved stirbar prior to pouring plates. For plates with benomyl, we also added to each liter a filter-sterilized solution of 4 mg benomyl suspended in 2 mL of 1:1 acetone and 70% ethanol.
Fungi that grew from root sub-fragments were subcultured onto the same medium, usually within 1 week of plating. Cultures were subsequently transferred to slants of malt-yeast extract agar, made as above with the addition of 1 g/L yeast extract, grown at room temperature for approximately 1 week, and stored at 4°C prior to transfer back to agar plates for DNA extraction and analysis.
We extracted DNA from 0.5 cm diameter plugs of fungal cultures using an alkaline lysis and chloroform extraction procedure (Miller et al. 1999). We PCR-amplified the ITS region of the nuclear rDNA genes using primers ITS1f and ITS4 (White et al. 1990; Gardes and Bruns; 1993). Amplicons were sorted into restriction-fragment length polymorphism (RFLP) groups by digesting at 37°C for 12h with the restriction enzymes AluI, HinfI, and HaeIII and visualizing products of restriction digests in 3% NuSieve agaraose gels stained with ethidium bromide. RFLP patterns were grouped by the sizes of fragments (±8%) exceeding 70 bases long.
We sequenced the ITS region for a subset of RFLP groups, including the 26 groups that were most common in the whole dataset (cultures from window and cores, with and without benomyl). Amplicons from 3 to 6 representatives of each RFLP group were sequenced at Cornell University’s Biotechnology Resource Center. We assembled forward and reverse sequences using Contig Express software in Vector NTI (Invitrogen, Carlsbad CA), manually checked each sequence, and aligned sequences from replicate representatives to verify similarity within each group using AlignX software in Vector NTI. Sequences were BLAST matched (Altschul et al. 1997) to the Genbank nonredundant database and were submitted to the Genbank database under accession numbers HQ392591–HQ392617. The fungal ITS region is often used to determine species identities in fungi (Bruns and Shefferson 2004). A 1%–3% sequence dissimilarity is on average appropriate for distinguishing fungal species, but there are numerous exceptions in which within species differences are higher (Nilsson et al. 2008). There are also instances in which species cannot be distinguished by ITS comparisons (i.e., Trichoderma; Lieckfeldt et al. 1999; Jaklitsch et al. 2006). Therefore we present the closest matches as very tentative identifications that should be interpreted with caution.
Data analyses
RFLP groups were recorded as presence/absence for an individual root system, and their abundance was estimated as a relative frequency (number of roots present/total roots) for each of our 4 study plots. Shannon-diversity and evenness were estimated within sites. We used Sørensen’s index of community similarity (van Tongeren 1995) to compare species composition between treatments and among sites. We also used non-metric multidimensional scaling (NMS; McCune and Grace 2002) to compare communities between treatments.
We used paired t-tests (SAS Institute, Cary, North Carolina USA) to test effects of disturbance on diversity (H’), evenness, and number of RFLP groups at the site level. For RFLP groups present on 10% or more of the root samples, we analyzed the probability of occurrence as a correlated bionomial response variable in a repeated measures design with the factors treatment (between-subjects) and sampling time (within-subjects). We used generalized estimating equations (Myers et al. 2002) in Proc GENMOD in SAS to analyze correlated binomial responses and to model probability of occurrence with the logit link function. We tested treatment and time effects with pairwise contrasts and Wald’s chi-square test.
Results
Fungi cultured from dead roots were separated into 78 RFLP groups, recovered from 186 (no-benomyl cultures) or 152 (benomyl cultures) root fragments from windows and 189 root fragments from soil cores. Root fragments from windows originated from 48 total root systems (4 plots, 2 windows per plot, and 6 root systems per window), each sampled 4 times. Thirty-one of these RFLP groups accounted for 90% of total frequency on roots from the windows. Twenty-eight RFLP groups were in common between windows and soil cores, and 23 groups were found only in cores. ITS sequences were obtained for representatives of 18 of the groups found in windows and cores, and for representatives of 8 additional groups that were common on dead roots from soil cores but were not found on window roots (Table 1). Many of the ITS sequences from our fungal cultures matched Genbank sequences of common soil, saprotrophic and semi-pathogenic fungi (Table 1).
Richness and diversity of fungi from the windows did not respond to disturbance treatment (P ≥ 0.15 for richness, diversity, and evenness at the site level). Using benomyl in our culture medium improved the detection of a number of species to give us a second, complementary view of the community, with almost twice as many RFLP groups cultured from window roots using benomyl compared to no benomyl (P ≤ 0.0015 for richness, diversity, and evenness at the site level; Table 2). Most RFLP groups were not common, especially for benomyl cultures, in which 42 groups were detected on five or fewer root samples and 21 groups were detected on only one root sample. In cultures without benomyl, 19 RFLP groups were detected on five or fewer roots and ten on only one root. In no-benomyl cultures, 13 groups occurred in common between control and disturbance treatments, accounting for 85% of total occurrences. In benomyl cultures, 9 groups occurred in common between treatments, accounting for 71% of the total.
In contrast to the lack of diversity response, disturbance strongly affected fungal species composition. Community similarity between control and disturbance roots was low, averaging 0.37 across sites in no-benomyl and 0.38 in benomyl cultures. Community similarity between each pair of disturbance and control windows varied among sites, more widely in no-benomyl (0.18–0.50) compared to benomyl cultures (0.23–0.41). NMS ordination of no-benomyl cultures using 2 axes accounted for 76.2% of variation (stress = 21.16; P = 0.016), and separated control and disturbance communities (Fig. 2a). NMS ordination of benomyl cultures was less straightforward, requiring 4 axes and accounting for 69.4% of variation (stress = 16.16, P = 0.028). Control and disturbance communities separated most clearly along axes 2 and 3 (Fig. 2b). NMS ordinations provided no clear separations of communities by time or by site, for either culture type.
Abundance of RFLP groups from roots in control and disturbance windows, under two culture conditions (a: no benomyl; and b: with benomyl). Groups are ranked according to relative frequency in benomyl cultures and include those groups that accounted for 90% of total occurrence in benomyl cultures. Bars are means of 4 sites and 4 sample times
Neonectria sp 1 was the most widespread taxon, followed by Trichoderma, in control window roots when cultured without benomyl (Fig. 3). Disturbance significantly increased frequency of Trichoderma and reduced the frequency of Neonectria sp 1 relative to control roots. Trichoderma was common at 1 month on both control and disturbance roots (Fig. 4a). Neonectria sp 1 rapidly increased in frequency on control roots, but not on disturbance roots, which continued to be dominated by Trichoderma (Fig. 4a). Several other fungi, found in about 5 to 10% of roots when cultured without benomyl (Umbelopsis sp 1 and sp 2, Mortierella sp 1), did not respond to disturbance (Fig. 3; Table 3). Umbelopsis sp 1 and sp 2 increased in frequency over time, especially between 6 and 13 months (Fig. 4a; Table 3).
Treatment responses were also detected for some of the most common species when cultured with benomyl, including one (Umbelopsis sp 2) for which a response was not detected in cultures without benomyl (Table 3, Fig. 3). The frequency of Trichoderma and Neonectria sp 1 again responded clearly to disturbance in benomyl cultures (Figs. 3 and 5; Table 3), but their overall frequency was reduced. The species that compensated for that reduction depended upon treatment. Umbelopsis sp 1 was common on control roots and Umbelopsis sp 2 was much more common on disturbance roots (Table 3). Pochonia sp 1 and Mortierella sp 1 were common and again did not appear to respond to treatment (Fig. 3). The disturbance-related shift in the fungal community included a change in timing of colonization by Umbelopsis sp 1 and sp 2 and by Trichoderma. In the absence of disturbance, frequency of common species was low at first and then increased over time, especially for Umbelopsis sp 1. In contrast, disturbance promoted high initial frequencies of Umbelopsis sp 2 and Trichoderma, and slowed the increase of the most common control species, Umbelopsis sp 1 (Fig. 4b).
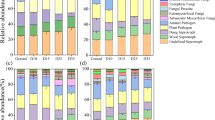
Frequency of fungi cultured (with benomyl) from dead roots in soil cores, presented in the rank-order of RFLP groups from window roots. Roots originated from the O horizon (forest floor) and from the A and B horizons in the surface 10 cm of mineral soil. Bars are averages of roots from 12 soil cores
Most of the fungal taxa that were present in root windows were also present in soil core samples, but some taxa were found only in soil cores (Table 1). Only a few of the fungal taxa that were common on dead roots were clearly associated with a particular soil horizon. For example, Trichoderma occurred frequently in both windows and cores and was generally ubiquitous, but also was clearly favored in the forest floor horizon (Fig. 6). Frequency of most other taxa showed no patterns of depth distribution in cores, and NMS ordination did not separate communities (data not shown). Frequency of 3 taxa appeared to differ between windows and cores. Umbelopsis sp 2 and Paecilomyces were common on window roots but were rare in soil core roots (Fig. 6). Pochonia sp 2 was common on roots from soil cores and was most widespread on roots collected from the A horizon, but was rare on roots from windows (Fig. 6).
Discussion
The culturable fungal community on dead fine roots included very common taxa, some with frequencies up to 0.65 (Fig. 4a). Sequences of the most common taxa were most similar to Umbelopsis, Trichoderma, Neonectria, and Pochonia (Table 1). These genera are among the dominant fungi in the rhizosphere of forest trees and on living and dead roots in other culture studies. Sequences of other RFLP groups that we found in moderate frequency most closely matched those of Geomyces, Mortierella, Mucor, Trichosporon, and Tolypocladium (Table 1), genera that have also been reported in roots and rhizosphere (Halmschlager and Kowalski 2004; Kwaśna 2004; Summerbell 2005; Menkis et al. 2006; Vandegrift et al. 2007; Kwaśna et al. 2008). Most of these taxa, such as Umbelopsis, Trichoderma, Mortierella, Mucor, Pochonia, and Geomyces are common soil and litter fungi in forest ecosystems (Hudson 1968; Bridge and Spooner 2001; DeBellis et al. 2007; Domsch et al. 2007; Osono and Takeda 2007). Umbelopsis spp appear to have an especially high affinity for fine roots (Summerbell 2005; Vandegrift et al. 2007), and Cylindrocarpon destructans (anamorph of Neonectria) is known for its potential as a mild pathogen (Rahman and Punja 2005; Götz et al. 2006; Domsch et al. 2007; Kwaśna and Bateman 2009). Trichosporon porosum is also common in rotting wood (Middelhoven et al. 2001; Vasiliauskas et al. 2005). Two other sequences from our study match fungal taxa that might not be expected on roots: Isaria farinosa is known as an insect parasite and Pochonia suchlasporia is known as a nematode cyst parasite (Domsch et al. 2007). Our results suggest that these rather specialized taxa can grow in fine roots but it is also possible that we have instead cultured species that are not in the sequence database. Other taxa that were important in previous studies of rhizospheres and dead fine roots of trees (Halmschlager and Kowalski 2004; Kwaśna 2004) were notably absent (Absidia, Phialocephala, Phialophora) or infrequent (Geomyces, Penicillium) in our study. The difference between ectomycorrhizal (EM) and arbuscular mycorrhizal (AM) tree species is likely to have great impact on fungal communities; our study focused on sugar maple roots (AM) whereas most others dealt with oak and conifer forests (EM). In fact, it seems remarkable that the root decay communities in various cited studies shared as many dominant species as they did in widely contrasting settings and with different mycorrhizal types.
Mycorrhizal and soil fungal communities tend to change with depth and differ between forest floor and mineral soil (Dickie et al. 2002; Lindahl et al. 2007; Hartmann et al. 2009). It was thus somewhat surprising not to find a more pronounced pattern with depth on dead roots. Instead, our soil cores suggest a fairly generalized community on dead roots throughout the surface 20 cm in our study sites. Trichoderma showed the strongest affinity for the forest floor, yet even this taxon was common in A and B horizons, and most of the taxa that were detected on more than one root were detected in multiple horizons (Fig. 6). The lack of clear depth-related patterns in our cores suggests that root depth did not contribute substantial variation to fungal community composition within windows, and it is unlikely that sampling only in the mineral soil in our windows caused us to miss any key taxa from the forest floor.
Our experimental approach allowed us to characterize fungal communities over time following root death with minimal disturbance to the surrounding soil and root systems. Communities of culturable fungi overlap between living and dead root systems (Halmschlager and Kowalski 2004; Menkis et al. 2006; Kwaśna et al. 2008) but the possibility of successional patterns in roots of forest trees has not previously received much attention. We found significant changes over time in frequency of several of the most common fungal taxa on dead roots, adding temporal resolution to the general descriptions that exist for root fungal communities. For example, Trichoderma and Neonectria sp 1 were primary colonizers that persisted in the community throughout the experiment. Both of these genera contain species that are common on living roots (Germino et al. 2006; Götz et al. 2006; Rahman and Punja 2005; Kwaśna et al. 2008; Kwaśna and Bateman 2009). Sugar maple fine roots in our study sites can continue respiring for several weeks following trenching (Littell 2007), suggesting that our experimentally killed roots did not die right away. Hence, the abundance of Trichoderma and Neonectria only one month after roots were cut suggests that they colonized roots prior to the time at which we killed the roots. Several other species that we found in root windows were secondary colonizers, appearing only after the first sampling time and persisting on root systems after that. For example, Umbelopsis sp 1 was absent from window communities at month one in three of our four study sites, after which it colonized roots in all sites and increased in frequency over time. Two other common taxa (Mortierella sp 1 and sp 2) also appeared to be late-stage members of the root decay community; they were patchily distributed among sites, but once they colonized roots in a particular window, they proliferated within that window and persisted throughout the experiment (data not shown). Frequency of Pochonia sp 1, another common member of the community, was more stable over time.
The fine root community that we measured also had many transient members that colonized at various times and were not detected again. These taxa appear to colonize sporadically but either are unable to grow and expand into adjacent root tissue, or are unable to persist at all. Representing a true successional time course in decaying fine roots is challenging, and more intensive and smaller-scale sampling of single root systems is needed to distinguish fungi that are truly transient members of the community from those that are not detected by sequential sampling because of very small scales of distribution.
While frequency of some fungal taxa changed over time, we did not find the marked temporal change in the community that has been noted for wood or leaf litter. NMS ordination, for instance, did not separate samples by time. This lack of community-level pattern may be related to high diversity and the transience of many taxa, but it is also noteworthy that we found no clear decline over time in common species, with the exception of Trichoderma in benomyl cultures. Nor did we observe obvious species replacements over time. This lack of marked change is consistent with the remarkable similarity that we found between communities in root windows and in soil cores, despite the differences in age of root substrates. Dead roots in soil cores senesced without any disturbance and presumably died at varying times prior to core collection, whereas roots in windows were experimentally killed all at one time. Only the absence of Pochonia sp 2 from windows compared to cores represents an obvious effect of the windows. These patterns in decaying roots contrast somewhat with patterns in wood (Coates and Rayner 1985; Boddy 1992) and litter (Hudson 1968; Frankland 1998; Frankland 1992; Osono 2005), in which some primary colonizing species do not persist after arrival of secondary colonizers. The phyllosphere and leaf endophyte community of leaves encounters some change in environment when litter falls to the ground after senescence, and while some phyllosphere taxa can persist and degrade structural C compounds (Osono 2006), the phyllosphere community has not been found to exert much influence on the subsequent fungal community in leaf litter (Osono 2005). In contrast, fungi colonize fine roots over the entire lifespan of the root and the transition from living tissue to detritus is more continuous. Taxa such as Trichoderma, Mortierella, Umbelopsis, and Mucor were detected right away in roots in this study, whereas they tend to be later-stage secondary colonizers of litter (Frankland 1998; Osono 2005). Prior colonization of wood can be important but also can be highly variable (Rayner and Boddy 1988), and temporal effects may differ because fine roots are smaller and also such ephemeral substrates compared to wood.
The fungal community response to disturbance supported our hypothesis that disruption of the rhizosphere would alter the dynamics of decay fungi. Disturbance altered the dynamic between two species that had probably colonized living roots: Trichoderma was able to maintain dominance over time following disturbance, while Neonectria was suppressed. Disturbance also favored colonization and growth by Umbelopsis sp 2 and suppressed that of Umbelopsis sp 1. Although the mechanism underlying the disturbance responses is uncertain, a potential contributor could have been a change in the pool of species available to colonize dead roots; i.e. species in the immediate rhizosphere were replaced with those in bulk soil. This changes temporal dynamics for species that colonize the rhizosphere from the bulk soil. However, this explanation would not account for the shift between Trichoderma and Neonectria sp 1, which must have resulted primarily from interactions with other members of the rhizosphere community. Moreover, the most common species were found in both treatments, indicating that the potential pool of colonizing species was not substantially altered and is not the only mechanism by which our treatment influenced the fungal community composition.
The response to disturbance that we found would not be expected if processes of colonization and persistence were entirely stochastic. Whereas recent work in microbial communities suggests that stochastic factors can strongly influence community development (Sloan et al. 2006; Woodcock et al. 2007), our results indicate that species interactions are an important part of community development in fine roots. There are good reasons to expect stochasticity in fungal communities that decompose fine roots, such as the diversity of potential colonizing species in litter and soil and the small and discreet nature of the fine root substrate. The potential for these traits to lead to randomly assembled communities could be compounded by the relatively short-term existence of the habitat caused by microbial consumption of the fine root substrate. Stochasticity of colonization is at least partly supported by the high fungal diversity, large number of rare taxa, and variation among individual roots in our study. However, several common species emerged in communities, even across multiple study sites, indicating that deterministic factors are of greater importance in the ecology of fine root decay fungi. Fischer et al. (2009) showed that initially stochastic processes can give way over time to deterministic effects. Our results differ somewhat from those of Fischer et al. (2009), because we did not find evidence for notable changes in community processes over time.
Variation in fungal communities that we found, either in response to disturbance or among individual roots, could influence the decay process. Most of the taxa that we detected can metabolize sugars and slightly more structural compounds like pectin (Domsch et al. 2007). Most are also able to metabolize cellulose, although some, including species of Mortierella, Mucor, and Sporothrix, cannot (Deacon et al. 2006; Domsch et al. 2007). Important functional differences in the root community probably exist between Trichoderma and Neonectria, likely to metabolize cellulose, and Umbelopsis isabellina and U. ramanniana, which are more likely to metabolize chitin. Trichosporon is notable in the fine root decay community for its ability to metabolize phenolic compounds that are likely produced as plant defense compounds (Middelhoven et al. 2001; Middelhoven 2006). Probably most importantly, only two or three of the taxa that we detected are known to break down lignin. These include Epicoccum and Trametes (Worrall et al. 1997; Domsch et al. 2007), and possibly RFLP group 30 which is likely Nolanea (or Entoloma) or Mycena (Osono 2002; Deacon et al. 2006). While the common species that we detected on fine roots have some potential for differences utilizing cellulose or lignin, they also have a fair amount of functional redundancy, especially among taxa that responded to disturbance. It is also important to note that functions can vary among strains of the same species (Deacon et al. 2006). Thus it is not clear from our data that changes in the fungal community related to rhizosphere disturbance would alter the decay process, as suggested by a number of studies of fine root decomposition rates (Fahey 1992; Fahey and Arthur 1994; Majdi and Nylund 1997; Dornbush et al. 2002). However, differences within the fungal community are more likely to influence the decay process in the less easily cultured components of the fungal community, including basidiomycetes and ascomycetes that can degrade lignin and whose functions can vary substantially (Boddy et al. 1989; Chapela et al. 1988; Worrall et al. 1997; Fukami et al. 2010). These taxa appear best represented in PCR-based community assessments (Vandegrift et al. 2007; Kwaśna et al. 2008), and molecular genetic studies are needed to complement what we have learned in the culturable community.
Our experimental approach was intended to represent the complexity of the relatively undisturbed soil environment to the best extent possible, to most conclusively demonstrate patterns over time and effects of rhizosphere disruption. While this approach limits our ability to confirm specific mechanisms of apparent interactions in response to disturbance, it does provide valuable evidence that such interactions are relevant in the intact soil system and are worth examining under more controlled conditions. For example, further work that directly compares fungal communities on decaying roots to those in surrounding rhizosphere and bulk soil (representing the pool of potential colonizers) would contribute to our understanding of the relative importance of chance vs species interactions. Our results also show that future attention to possible priority effects would be especially interesting in the fine root substrate because of the apparent continuity of some species between living and dead roots. Finally, while it is logistically challenging to quantify the decay rates of individual roots, our data show that this might be the most informative level of investigation based on heterogeneity among roots, and our characterization of the culturable fungal community on fine roots provides a useful basis from which to target the most common species for study.
References
Addy HD, Piercey MM, Currah RS (2005) Microfungal endophytes in roots. Can J Bot 83:1–13
Altschul SF, Madden TL, Schaffer AA, Zhang JH, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSL-BLAST: a new generation of protein database programs. Nucl Acids Res 25:2289–3402
Boddy L (1992) Development and function of fungal communities in decomposing wood. In: Carroll GC, Wicklow DT (eds) The fungal community: Its organization and role in the ecosystem. Marcel Dekker, NY, pp 749–782
Boddy L (2000) Interspecific combative interactions between wood-decaying basidiomycetes. FEMS Microbiol Ecol 31:184–194
Boddy L, Owens EM, Chapela IH (1989) Small-scale variation in decay rate within logs one year after felling: effect of fungal community structure and moisture content. FEMS Microbiol Ecol 62:173–184
Bohlen PJ, Groffman PM, Driscoll CT, Fahey TJ, Siccama TG (2001) Plant-soil-microbial interactions in a northern hardwood forest. Ecology 82:965–978
Bridge P, Spooner B (2001) Soil fungi: diversity and detection. Plant Soil 232:147–154
Bruns TD, Shefferson RP (2004) Evolutionary studies of ectomycorrhizal fungi: recent advances and future directions. Can J Bot 82:1122–1132
Chapela IH, Boddy L, Rayner ADM (1988) Structural development of fungal communities in beech logs four and a half years after felling. FEMS Microbiol Ecol 53:59–70
Coates D, Rayner ADM (1985) Fungal population and community development in cut beech logs III. Spatial dynamics, interactions and strategies. New Phytol 101:183–198
Deacon LJ, Pryce-Miller EJ, Frankland JC, Bainbridge BW, Moore PD, Robinson CH (2006) Diversity and function of decomposer fungi from a grassland soil. Soil Biol Biochem 38:7–20
DeBellis T, Kernaghan G, Widden P (2007) Plant community influences on soil microfungal assemblages in boreal mixed-wood forests. Mycologia 99:356–367
Dickie IA, Xu B, Koide RT (2002) Vertical niche differentiation of ectomycorrhizal hyphae in soil as shown by T-RFLP analysis. New Phytol 156:527–535
Domsch KH, Gams W, Anderson T-H (2007) Compendium of Soil Fungi, 2nd edn. IHW-Verlag, Eching, 672 pp
Dornbush ME, Isenhart TM, Raich JW (2002) Quantifying fine-root decomposition: an alternative to buried litterbags. Ecology 83:2985–2990
Ellwood MD, Manica A, Foster WA (2009) Stochastic and deterministic processes jointly structure tropical arthropod communities. Ecol Lett 12:277–284
Fahey TJ (1992) Mycorrhizae and forest ecosystems. Mycorrhiza 1:83–89
Fahey TJ, Arthur MA (1994) Further studies of root decomposition following harvest of northern hardwoods. For Sci 40:618–629
Fahey TJ, Hughes JW (1994) Fine root dynamics in a northern hardwood forest ecosystem, Hubbard Brook Experimental Forest, NH. J Ecol 82:533–548
Fahey TJ, Hughes JW, Pu M, Arthur MA (1988) Root decomposition and nutrient flux following whole-tree harvest of northern hardwood forest. For Sci 34:744–768
Fischer H, Bergfur J, Goedkoop W, Tranvik L (2009) Microbial leaf degraders in boreal streams: bringing together stochastic and deterministic regulators of community composition. Freshw Biol 54:2276–2289
Frankland JL (1992) Mechanisms in fungal succession. In: Carroll GC, Wicklow DT (eds) The fungal community: Its organization and role in the ecosystem. Marcel Dekker, NY, pp 383–401
Frankland JL (1998) Fungal succession—unraveling the unpredictable. Mycol Res 102:1–15
Fukami T, Dickie IA, Wilkie JP, Paulus BC, Park D, Roberts A, Buchanan PK, Allen RB (2010) Assembly history dictates ecosystem functioning: evidence from wood decomposer communities. Ecol Lett 13:675–684
Gardes M, Bruns TD (1993) ITS primers with enhanced specificity for basidiomycetes—application to the identification of mycorrhizae and rusts. Mol Ecol 2:113–118
Germino MJ, Hasselquist NJ, McGonigle T, Smith WK, Sheridan P (2006) Landscape- and age-based factors affecting fungal colonization of conifer seedling roots at the alpine tree line. Can J For Res 36:901–909
Gessner MO, Swan CM, Dang CK, McKie BG, Bardgett RD, Wall DH, Hättenschwiler S (2010) Diversity meets decomposition. Trends Ecol Evol 25:372–380
Götz M, Nirenberg H, Krause S, Wolters H, Draeger S, Buchner A, Lottmann J, Berg G, Smalla K (2006) Fungal endophytes in potato roots studied by traditional isolation and cultivation-independent DNA-based methods. FEMS Microbiol Ecol 58:404–413
Halmschlager E, Kowalski T (2004) The mycobiota in nonmycorrhizal roots of healthy and declining oaks. Can J Bot 82:1446–1458
Harmon ME, Franklin JF, Swanson FJ, Sollins P, Gregory SV, Lattin JD, Anderson NH, Cline SP, Aumen NG, Sedell JR, Lienkaemper GW, Cromack K, Cummins KW (1986) Ecology of coarse woody debris in temperate ecosystems. Adv Ecol Res 15:133–302
Hartmann M, Lee S, Hallam SJ, Mohn WW (2009) Bacterial, archaeal and eukaryal community structures throughout soil horizons of harvested and naturally disturbed forest stands. Environ Microbiol 11:3045–3062
Hendrick RL, Pregitzer KS (1992) The demography of fine roots in a northern hardwood forest. Ecology 73:1094–1104
Hobbie SE, Oleksyn J, Eissenstat DM, Reich PB (2010) Fine root decomposition rates do not mirror those of leaf litter among temperate tree species. Oecologia 162:505–513
Holmer L, Stenlid J (1996) Diffuse competition for heterogeneous substrate in soil among six species of wood-decomposing basidiomycetes. Oecologia 106:531–538
Horner-Devine MC, Silver JM, Leibold MA, Bohannan BJM, Colwell RK, Fuhrman JA, Green JL, Kuske CR, Markiny JBH, Muyzer G, Øvreas L, Reysenback AL, Smith VH (2007) A comparison of taxon co-occurrences: Patterns for macro-and microorganisms. Ecology 88:1345–1353
Hudson HJ (1968) The ecology of fungi on plant remains above the soil. New Phytol 67:837–874
Hughes JW, Fahey TJ (1994) Litterfall dynamics and ecosystem recovery during forest development. For Ecol Manag 63:181–198
Huston MA (1999) Local processes and regional patterns: appropriate scales for understanding variation in the diversity of plants and animals. Oikos 86:393–401
Jaklitsch WM, Samuels GJ, Dodd SL, Lu B-S, Druzhinina IS (2006) Hypocrea rufa/Trichoderma viride: a reassessment, and description of five closely related species with and without warted conidia. Stud Mycol 55:135–177
Jönsson MT, Edman M, Jonsson BG (2008) Colonization and extinction patterns of wood-decaying fungi in a boreal old-growth Picea abies forest. J Ecol 96:1065–1075
Kennedy PG, Peay KG, Bruns TD (2009) Root tip competition among ectomycorrhizal fungi: are priority effects a rule or an exception? Ecology 90:2098–2107
Kwaśna H (2004) Natural shifts in communities of rhizosphere fungi of common oak after felling. Plant Soil 264:209–218
Kwaśna H, Bateman GL (2009) Microbial communities in roots of Pinus sylvestris seedlings with damping-off symptoms in two forest nurseries as determined by ITS1/2 rDNA sequencing. For Pathol 39:239–248
Kwaśna H, Bateman GL, Ward E (2008) Determining species diversity of microfungal communities in forest tree roots by pure-culture isolation and DNA sequencing. Appl Soil Ecol 40:44–56
Lieckfeldt E, Samuels GJ, Nirenberg H, Petrini O (1999) A morphological and molecular perspective of Trichoderma viride: is it one or two species? Appl Environ Microbiol 65:2418–2428
Likens GE, Bormann FH, Pierce RS, Eaton JS (1985) The Hubbard Brook valley. In: Likens GE (ed) An ecosystem approach to aquatic ecology: Mirror lake and its environment. Springer, New York, pp 9–39
Lindahl BD, Ihrmark K, Boberg J, Trumbore SE, Högberg P, Stenlid J, Finlay RD (2007) Spatial separation of litter decomposition and mycorrhizal nitrogen uptake in a boreal forest. New Phytol 173:611–620
Littell AM (2007) Fungal community change during fine root decomposition in a northern hardwood forest. Master’s thesis, Appalachian State University, Boone, NC, USA. 63 pp
Lodge DJ, Cantrell S (1995) Fungal communities in wet tropical forests: variation in time and space. Can J Bot 73:S1391–S1398
Majdi H, Nylund J (1997) Does liquid fertilization affect fine root dynamics and lifespan of mycorrhizal short roots? Plant Soil 185:305–309
McClaugherty CA, Aber JD, Melillo JM (1982) The role of fine roots in the organic matter and nitrogen budgets of two forest ecosystems. Ecology 63:1481–1490
McCune B, Grace JB (2002) Analysis of ecological communities. MjM Software Design, Gleneden Beach, 300 pp
Menkis A, Vasiliauskas R, Taylor AFS, Stenström E, Stenlid J, Finlay R (2006) Fungi in decaying roots of conifer seedlings in forest nurseries, afforested clear-cuts and abandoned farmland. Plant Pathol 55:117–129
Middelhoven WJ (2006) Trichosporon porosum isolated from decaying wood in forests. Antonie Van Leewenhoek Int J Gen Mol Microbiol 90:57–67
Middelhoven WJ, Scorzetti G, Fell JW (2001) Trichosporon porosum comb. Nov., an anamorphic basidiomycetous yeast inhabiting soil, related to the loubieri/laibachii group of species that assimilate hemicelluloses and phenolic compounds. FEMS Yeast Res 1:15–22
Miller DN, Bryant JE, Madsen EL, Ghiorse WC (1999) Evaluation and optimization of DNA extraction and purification procedures for soil and sediment samples. Appl Environ Microbiol 65:4715–4724
Myers RH, Montgomery DC, Vining GG (2002) Generalized linear models. Wiley, New York
Nilsson RH, Kristiansson E, Ryberg M, Hallenberg N, Larsson K-H (2008) Intraspecific ITS variability in the kingdom Fungi as expressed in the international sequence databases and its implications for molecular species identification. Evol Bioinform 4:193–201
Osono T (2002) Comparison of litter decomposing ability among diverse fungi in a cool temperate deciduous forest in Japan. Mycologia 94:421–427
Osono T (2005) Colonization and succession of fungi during decomposition of Swida controversa leaf litter. Mycologia 97:589–597
Osono T (2006) Role of phyllosphere fungi of forest trees in the development of decomposer fungal communities and decomposition processes of leaf litter. Can J Microbiol 52:701–716
Osono T, Takeda H (2007) Microfungi associated with Abies needles and Betula leaf litter in a subalpine coniferous forest. Can J Microbiol 53:1–7
Rahman M, Punja ZK (2005) Factors influencing development of root rot on Ginseng caused by Cylindrocarpon destructans. Phytopathol 95:1381–1390
Rayner ADM, Boddy L (1988) Fungal communities in the decay of wood. Adv Microb Ecol 10:115–166
Ruess RW, Hendrick RL, Bryant JP (1998) Regulation of fine root dynamics by mammalian browsing in early successional forests of interior Alaska. Ecology 79:2706–2720
Sloan T, Lunn M, Woodcock S, Head IM, Nee S, Curtis TP (2006) Quantifying the roles of migration and chance in shaping prokaryote community structure. Environ Microbiol 8:732–740
Summerbell RC (2005) From Lamarkian fertilizers to fungal castles: recapturing the pre-1985 literature on endophytic and saptrotrophic fungi associated with ectomycorrhizal root systems. Stud Mycol 53:191–265
Thorn RG, Reddy CA, Harris D, Paul EA (1996) Isolation of saprophytic basidiomycetes from soil. Appl Environ Microbiol 62:4288–4292
van Tongeren OFR (1995) Cluster analysis. In: Jongman RHG, ter Braak CJF, van Tongeren OFR (eds) Data analysis in community and landscape ecology. Cambridge University Press, New York, pp 174–212
Vandegrift EVH, Chen H, Harmon ME (2007) Fungal genetic diversity within decomposing woody conifer roots in Oregon, USA. Northwest Sci 81:125–137
Vandenkoornhuyse P, Baldauf SL, Leyval C, Straczek J, Young JPW (2002) Extensive fungal diversity in plant roots. Science 295:2051
Vasiliauskas R, Larsson E, Larsson K-H, Stenlid J (2005) Persistence and long-term impact of Rotstop biological control agent on mycodiversity in Picea abies stumps. Biol Control 32:295–304
White TJ, Bruns TD, Lee SB, Taylor JW (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ (eds) PCR protocols—a guide to methods and applications. Academic, San Diego, pp 315–322
Woodcock S, van der Gast CJ, Bell T, Lunn M, Curtis TP, Head IM, Sloan WT (2007) Neutral assembly of bacterial communities. FEMS Microbiol Ecol 62:171–180
Worrall JJ, Anagnost SE, Zabel RA (1997) Comparison of wood decay among diverse lignicolous fungi. Mycologia 89:199–219
Acknowledgements
We thank Cindy Wood and Jess Taybolt for assistance in the laboratory. We also thank two anonymous reviewers for their comments, which substantially improved the manuscript. This research was supported by grant 2002-00498 from the United States Department of Agriculture. This manuscript is a contribution of the Hubbard Brook Ecosystem Study. Hubbard Brook is part of the Long-Term Ecological Research (LTER) network, which is supported by the National Science Foundation. The Hubbard Brook Experimental Forest is operated and maintained by the USDA Forest Service, Northern Research Station, Newtown Square, PA.
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible Editor: Thom W. Kuyper.
Rights and permissions
About this article
Cite this article
Fisk, M.C., Fahey, T.J., Sobieraj, J.H. et al. Rhizosphere disturbance influences fungal colonization and community development on dead fine roots. Plant Soil 341, 279–293 (2011). https://doi.org/10.1007/s11104-010-0643-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11104-010-0643-4