Abstract
Aims
While the effects of destocking on soil nutrient and plant productivity are known, the effect on functional groups of fungi has received less attention. The objective of this study was to evaluate the effect of long-term destocking on fungal functional guilds and their association with plants and soil.
Methods
We characterized the changes in five fungal functional guilds, including plant pathogens, animal pathogens, wood saprotrophs, dung saprotrophs, and arbuscular mycorrhizal fungi (AMF), along a 35-year chronosequence following destocking at 0–60 cm soil depths on the Chinese Loess Plateau. Fungal community composition was assigned by comparing with the FUNGuild database.
Results
After 35 years of destocking, diversities of plant pathogens, wood saprotrophs, and AMF increased, while that of animal pathogens and dung saprotrophs decreased. Destocking had a greater effect in the near-surface layers (0–10 and 10–20 cm) owing to the greater influence of plant biomass and soil nutrients. Among the above- and belowground drivers, plant pathogen diversity was largely associated with plant diversity, while animal pathogens, dung saprotrophs, and wood saprotrophs were associated with aboveground biomass, AMF were responsive to soil conditions (e.g., organic C, NO3−-N, C:N ratio, and moisture).
Conclusions
Long-term destocking can be considered to be an important predictor of fungal functional guilds, and changes in these microorganisms were associated with cessation-induced shifts in plants and soil by grazing. Our findings provide insights into the duration of destocking necessary to benefit fungal communities, and how this varies according to soil depth.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Globally, grazing is considered to be a widespread land-use strategy with far-reaching social and environmental effects (Ren et al. 2018b). It is also one of the key biotic factors influencing grassland ecosystems, which cover about 20% of Earth’s terrestrial surface and provide ecosystem services, including land for animal husbandry and carbon sequestration (Zhang et al. 2018). Grazing affects grassland ecosystems through vegetation removal, manure deposition, and trampling. Vegetation removal changes vegetation composition (e.g., different grazing intensities lead to changes in the relative proportion of slow- and fast-growing plant species), reduces vegetation productivity, and increases root exudates (Eldridge et al. 2017; Yang et al. 2019). Manure deposition can increase nutrient availability (e.g., C and N) and accelerate N cycling and reduce soil pH (Sun et al. 2018b). Moreover, trampling from large herbivores can compact soil, leading to changes in water-holding capacity and aeration (Boschi and Baur 2007). These grazing-induced changes in soil properties and vegetation may have a considerable effect on soil microbial diversity and community structure directly or indirectly, especially for soil fungi because they have more symbiotic relationships with plants, and they are dispersal-limited compared to bacteria (Schmidt et al. 2014; Yang et al. 2017).
Fungi play important roles in ecosystem processes, such as degrading recalcitrant organic matter compounds, thereby releasing mineral nutrients and carbon dioxide (Grau et al. 2017). Fungal hyphae increase soil aggregate stability, thereby reducing erosion and soil nutrient loss (de Menezes et al. 2017). Mycorrhizal fungi support mineral nutrient uptake in plants, thereby increasing the net primary productivity of an ecosystem (McGuire et al. 2010). Moreover, compared to bacteria, fungal species are well adapted to soil acidity and alkalinity and can usually survive under pH 5–9 without significant inhibition of fungal growth (Rousk et al. 2010). Many known functions of fungi are mediated by specific guilds, such as saprotrophs (decomposers related to recalcitrant organic matter), symbionts (mycorrhizal fungi), or pathogens (Eldridge and Delgado-Baquerizo 2018). Although the importance of fungi in terrestrial ecosystem processes is recognized, relatively little is known about the response of soil fungi in grasslands to disturbance from grazing, especially in arid and semiarid grassland ecosystems.
Vegetation in the Yunwushan National Nature Grassland Reserve, in the middle of the Loess Plateau, has experienced serious degradation due to overgrazing (Zhang et al. 2018). Methods such as the destocking (Jing et al. 2014), fertilization (Smits et al. 2008), and reseeding (Leff et al. 2015; Wang et al. 2006) have been used to restore degraded grasslands. Among them, destocking is considered to be the most effective approach for restoring grasslands (Cheng et al. 2016; Jing et al. 2014). Previous research has already established that the plant community composition in this area is determined by nitrogen and phosphorus content (Su et al. 2017). It was also shown that the composition of soil bacteria shifts from oligotrophic to copiotrophic groups as the intensity of destocking increases, owing to the accumulation of soil nutrients (Wang et al. 2019; Zeng et al. 2017). Dynamics of plant communities, soil properties, and microbes have been extensively reported; however, exploring the relationships among them, especially for functional group of microbes interacting with plants, has received less attention. Such information is, however, fundamental for our understanding of how to restore degraded grassland ecosystems and how to utilize grasslands efficiently and sustainably.
In the present study, the effect of long-term destocking on the fungal community was determined by investigating the diversity of fungal OTUs assigned to functional guild, including plant pathogens, animal pathogens, dung saprotrophs, wood saprotrophs, and arbuscular mycorrhizal fungi (AMF), in different soil layers (0–10, 10–20, 20–40, and 40–60 cm) of semiarid grasslands with a chronosequence since destocking (0, 10, 15, 25, and 35 years) on the Loess Plateau of China. These fungal guilds are considered to be important in biogeochemical processes because they function in the acquisition and decomposition of nutrients and plant growth (Eldridge and Delgado-Baquerizo 2018; Nguyen et al. 2016). We hypothesized that destocking would increase the diversity of plant pathogens, wood saprotrophs, and AMF, while decreasing the diversity of animal pathogens and dung saprotrophs owing to the reduction in livestock grazing. We also hypothesized that the effect of destocking on fungal communities would differ according to soil depth, with a greater effect in the near-surface layer owing to the greater influence from plant biomass and soil nutrients.
Materials and methods
Study area
The experiment was conducted in the Yunwushan National Nature Grassland Reserve (106°21′–106°27′E, 36°10′–36°17′N), in Guyuan City, Ningxia Hui Autonomous Region, China. The reserve has a total area of 6660 hm2 and consists of a core conservation zone (1100 hm2), a buffer conservation zone (1300 hm2), and an experimental zone (4360 hm2). The climate is semiarid with 425 mm of mean annual precipitation, 7.01 °C mean annual temperature, and 1020–1750 mm of annual evaporation. The study area has a montane grey-cinnamon soil (Haplic Calcisol in the FAO/UNESCO classification) (Wang et al. 2019), and there are approximately 300 plant species recorded. Stipa grandis, S. bungeana, Artemisia sacrorum, S. przewalskyi, and Thymus mongolicus dominate the grasslands (Cheng et al. 2016).
Study design and sampling
In August 2017, soil samples from four grasslands along a chronosequence of grassland restoration were collected at a time corresponding to peak aboveground biomass. Before fencing, these sites had been grazed at a density of 50 sheep ha−1y− since 1952, 1962, 1972, and 1977, and thus these sites had the same grazing time (30 years). Destocking had been conducted by fencing since 1982, 1992, 2002, and 2007. Therefore, the years since destocking were 35 (D35), 25 (D25), 15 (D15), and 10 (D10), respectively. As a reference for sustainable grazing, a grassland subjected to continuous grazing at 50 sheep ha−1y−1 was selected and is referred to as the continuous grazed site. The effects of site conditions on the study outcome were minimized by ensuring that all sites were at similar elevations, and that they had similar slopes, and soil type (Table 1).
Three replicated plots (100 m × 50 m) were established randomly in the investigated sites, and the distance between adjacent plots was large enough (80–100 m) that there was no spatial dependence (< 13.5 m) for most soil variables (Marriott et al. 1997). Soil samples were collected from 0 to 10-cm, 10–20-cm, 20–40-cm, and 40–60-cm layers from each plot using a stainless steel corer with a diameter of 5 cm. Nine samples were collected along an S line in each plot and combined to form a composite sample. After the roots, litter, debris, and stones were removed, the collected soil was divided into two parts. One part was frozen at −80 °C for DNA extraction. The other part was air-dried for physicochemical analysis. In each plot, three subplots (1 m × 1 m) were randomly established to determine the above- and belowground biomass and the number of plant species. The diversity of the plant community was estimated using the Shannon-Wiener index (Zhang et al. 2016). Aboveground biomass was measured by drying all aboveground parts (shoots, leaves, and litter) at 60 °C for 36 h. A root auger (inside diameter, 10 cm) was used to determine the root biomass at different soil layers (Li et al. 2019).
Soil characteristics
Soil organic carbon (C) was measured using the potassium dichromate oxidation, and pH was determined in a soil-to-water suspension of 2.5:1 (v/w). Total nitrogen (N) was measured using the Kjeldahl method (Bremner and Mulvaney 1982). Soil NH4+ and NO3− were extracted with 2 M KCl and thereafter determined by a flow auto-analyzer. Available phosphorus (P) was measured using the Olsen method (Olsen and Sommers 1982). Bulk density was calculated based on the inner diameter of the soil corer (stainless steel cylinders with a diameter and height of 5 cm). Soil moisture was determined by oven drying and calculating the percentage of dry weight (Zhang et al. 2016).
Molecular analyses of the soil fungal communities
Microbial DNA was extracted from 0.5 g of soil using the FastDNA spin kit (MP Biomedicals, Cleveland, USA) according to the manufacturer’s instructions. The DNA concentration and purity were determined using a NanoDrop ND-1000 spectrophotometer (NanoDrop Technologies, Wilmington, USA) and by 2% agarose gel electrophoresis. The primers ITS1F (5′-CTTGGTCATTTAGAGGAAGTAA-3′) and ITS2R (5′-GCTGCGTTCTTCATCGATGC-3′) were used to amplify the ITS1 region of the fungi. Polymerase chain reactions (PCR) were performed in a volume of 20 μL containing 10 ng template DNA, 4 μL 5× FastPfu Buffer, 0.4 μL FastPfu Polymerase, 0.8 μL forward and reverse primers (5 μM), and 2 μL dNTPs (2.5 mM). The thermal cycling protocol was as follows: denaturation at 98 °C for 60 s; 30 cycles of denaturation at 98 °C for 15 s, annealing at 50 °C for 30 s, and extension at 70 °C for 50 s; and a final extension at 70 °C for 6 min. The DNA was amplified in triplicate and purified with an xyPrep DNA Gel Extraction Kit (Axygen Biosciences, Union City, CA, US). Purified DNA was collected for sequencing on an Illumina HiSeq2500 PE250 platform (Illumina Inc., USA), and the sequences were uploaded to the NCBI Sequence Read Archive (accession number: SRP149460).
After merging the paired-end reads, sequences were quality-filtered according to previously described methods (Caporaso et al. 2010), and those with chimeras were excluded by the UCHIME algorithm (Edgar et al. 2011). The high-quality sequences were clustered into operational taxonomic units (OTUs) using complete-linkage clustering at a 97% similarity cutoff. OTUs with just one read were excluded. The taxonomic identity was determined based on the UNITE database (release 6.0, http://unite.ut.ee/index.php) using RDP Classifier (release 11.1, http://rdp.cme.msu.edu/). The observed species and Shannon Wiener index were determined in QIIME using 52,669 reads per sample.
Fungal OTUs assigned to functional guild
Fungal OTUs were assigned to functional groups by comparing with the FUNGuild database 1.0 (Nguyen et al. 2016; Yang et al. 2017). We mainly concentrated on five fungal functional guilds: plant pathogens, AMF, wood saprotrophs, animal pathogens, and dung saprotrophs. Fungal OTUs assigned to functional guild were conducted at genus level and only assignments with confidence levels of “highly probable” or “probable” were retained in the following analyses. Approximately 60% of the OTUs were matched to a functional guild in the FUNGuild database. The relative abundance of each functional group was equal to the sum of the relative abundance of all OTUs sharing a particular functional group (Eldridge and Delgado-Baquerizo 2018). Shannon diversity of fungal functional groups was calculated using the phyloseq package (McMurdie and Holmes 2013) in R v.3.6.0 software (R Development Core Team 2019).
Statistical analysis
A linear mixed model (split-plot) ANOVA was used to assess the effect of destocking and soil depth on the root biomass, soil physicochemical properties, and fungal diversity. In the model, destocking time was treated as the fixed main plot and soil depth treated as the fixed split plot (Derner et al. 2006; Mushinski et al. 2017). Significance was established at p < 0.05. Nonmetric multidimensional scaling (NMDS) was used to assess fungal community composition along the soil profile based on the Bray-Curtis distances of the sequencing data using vegan package. Permutational multivariate analysis of variance (PERMANOVA) was used to determine the effect of soil depth on fungal community composition (OTUs). An aggregated boosted tree (ABT) analysis was performed to quantify the effect of the plant and soil variables on the community composition of fungal OTUs assigned to functional guild using the gbmplus package with 500 trees for boosting (De'ath 2007) in R v.3.6.0 software (R Development Core Team 2019). Linear mixed model ANOVA, NMDS, and PERMANOVA was performed using the nlme and vegan packages in R v.3.6.0 software (R Development Core Team 2019).
Results
Plant and soil properties
Destocking increased plant diversity and plant biomass compared with the grazed site (Table S1). Plant diversity and aboveground biomass increased over time since the destocking and peaked after 25 years but decreased significantly thereafter. Root biomass in the 0–10-cm, 10–20-cm, and 20–40-cm layers increased with time since destocking, peaking after 25 years. Root biomass decreased with soil depth. Similar to root biomass, organic C, total N, and NH4+ increased after destocking, peaking after 25 years. The soil N and C content decreased as the soil depth increased. The highest NO3− level was observed in the 0–10-cm and 10–20-cm layers after 15 years and in the 20–40-cm and 40–60-cm layers after 35 years. Soil C:N ratio increased following destocking, peaked after 25 years in the topsoil (0–10 and 10–20 cm), and decreased with increasing soil depth. Soil moisture increased after destocking but did not change consistently in the soil profile. A slight change (8.19–8.51) was observed in pH during the destocking process. Bulk density decreased with time and increased with soil depth.
Fungal community diversity and composition
A total of 4,611,583 sequences were obtained, with an average of 76,860 sequences and 257 bp in length per sample. The number of OTUs increased over time since destocking, and the highest value was observed at the D35 site among the four soil layers (Table 2). The fungal Shannon diversity did not change in the initial 10 years since destocking, and thereafter increased with time across the four soil layers. No significant difference was observed in fungal richness as represented by the Chao1 metric between the grazed site and the D10 site, but it increased thereafter over time since destocking. The number of OTUs, Shannon diversity, and fungal community richness changed consistently with soil depth, and were higher at depths of 0–10 cm and 10–20 cm compared with those at 20–40 cm and 40–60 cm. A more significant change was observed in the topsoil compared with the deeper soil. The fungal community was dominated by Ascomycota (49.7 ± 5.7%, mean relative abundance), followed by Zygomycota (29.9 ± 5.4%) and Basidiomycota (10.2 ± 2.8%) (Fig. S1). The relative abundance of Ascomycota, Zygomycota, and Basidiomycota changed differently through time since destocking in different soil layers, while the relative abundance of Glomeromycota, accounting for only 7% of total fungi, increased through time. NMDS ordination indicated that fungal composition in the topsoil (0–10 cm and 10–20 cm) was more separated from that of the deep soil (20–40 cm and 40–60 cm) (Fig. S2).
Fungal OTUs assigned to functional guild
The FUNGuild database assigned 1982 OTUs (out of 3266) to fungal functional guilds: grazing (69.0%), D10 (60.3%), D15 (61.7%), D25 (53.4%), and D35 (58.6%) (Fig. 1). Most (approximately 60%) OTUs were assigned to plant pathogens, dung saprotrophs, animal pathogens, wood saprotrophs, and AMF. Among them, the relative abundance of animal pathogens decreased from 22.3% in the grazed grassland to 13.6% in the D 35 site, and that of dung saprotrophs decreased from 21.2% to 8.2%. Both functional groups exhibited a decreasing trend in all four soil layers after 35 years since destocking (Fig. 1). In contrast, the relative abundance of AMF, plant pathogens, and wood saprotrophs increased through time since destocking in the four soil layers and was significantly higher than that in the grazed grassland after 35 years.
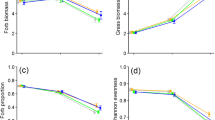
Destocking time and soil depth had a significant effect (P < 0.05) on the diversity of fungal OTUs assigned to functional guild (Table 3), and diversities of those functional guilds changed inconsistently through time at different soil depths (Fig. 2). Significant changes in the Shannon diversity of plant pathogens, AMF, and animal pathogens were observed in the topsoil (0–10 cm and 10–20 cm). In the topsoil (0–10 cm), the diversity of plant pathogens and AMF increased through time since destocking, peaked after 25 years, and thereafter significantly decreased. The diversity of animal pathogens and dung saprotrophs decreased through time, while that of wood saprotrophs increased.
Link between change in composition of fungal functional guild and environmental factors
Owing to the narrow pH range (pH 8.19–8.51) recorded during the destocking process, pH was not considered to be an environmental driver of fungal community change. NMDS ordination and PERMANOVA verified the influence of plants and soil properties on the composition of fungal communities within each guild along the soil profile, but with little evidence of the effect of destocking differing among soil depths (Fig. 3a–e; Table 4). However, the relationships between environmental variables and fungal community composition did vary with depth and among functional guilds of fungi. Plant diversity, plant biomass (including aboveground and root), organic C, NO3−-N, and moisture were significantly and positively associated with the fungal OTUs assigned to each functional guild at depths of 0–10 cm and 10–20 cm, while bulk density was significantly and positively associated with each functional guild at depths of 20–40 and 40–60 cm.
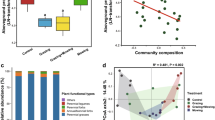
Nonmetric multidimensional scaling (NMDS) ordinations of soil fungal communities based upon their OTU composition derived from Bray-Curtis distances matrices. OC: soil organic carbon; TN: total nitrogen; AP: available phosphorus; HP: Shannon diversity of plant community; AB: aboveground biomass; RB: root biomass; SM: soil moisture; BD: bulk density. Significant factors are shown by black solid lines, while the non-significant factors are indicated using dotted lines (p < .05). D: Destocking
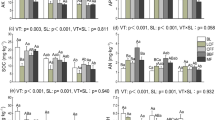
ABT models were employed to interpret the relative importance of plant characteristics and soil properties on the community composition of fungal OTUs assigned to functional guild (Fig. 4a–e). Plant characteristics, including Shannon diversity, aboveground biomass, and coverage were identified as primary factors associated with the changes in plant pathogens and wood saprotrophs, while the animal pathogens and dung saprotrophs were largely associated with by aboveground biomass (Fig. 4a–d). Soil conditions (e.g., organic C, NO3−-N, C:N ratio, and moisture) were the most accurate predictors of change in community composition of arbuscular mycorrhizal fungi (Fig. 4e).
Aggregated boosted tree (ABT) analysis showed the relative effect of plants and soil properties on the community composition of fungal OTUs assigned to functional guild. Hp: Shannon diversity index of plant community; SM: soil moisture, OC: soil organic carbon, AB: aboveground biomass, RB: root biomass, TN: total nitrogen, BD: bulk density, AP: available phosphorus. D: Destocking
Discussion
Effect of destocking on plant and soil properties
Our results showed an improvement of destocking on grassland productivity, characterized by the higher plant diversity and aboveground biomass in D35 than those in the grazed grassland (Table S1). The highest diversity and biomass were observed in the D25 site, and after 25 years, they decreased. A possible reason is the elimination of less competitive species caused by increased competition in the later restoration period (Odum 1969; Zhang et al. 2018). For example, A. capillaris and P. australis, predominated the community at the D25 and D35 sites, but the coverage of other species, such as C. squarrosa, P. heterophylla, and O. bicolor, significantly decreased (Table S2). In the continuously grazed site, herbivores ingested the ground plants and compressed the soil with their hooves (Wang et al. 2019), resulting in lower plant productivity and denser bulk soil (Jing et al. 2014). As the time since destocking increased, biomass increased, which led to improvements in soil nutrient level (e.g., C and N) through the decomposition of biomass and release of root exudates (Zhang et al. 2019). As expected, soil nutrient content (e.g., organic C, total N, and NH4+) decreased as soil depth increased. Higher nutrient content in the surface soil conformed to higher biomass accumulation and higher soil moisture. Bulk density, an indicator of soil aeration, decreased with the destocking time, suggesting a better soil aeration status caused by long-term destocking. Higher bulk density in the deep soil layer than the surface indicated that the deeper soil is prone to be more anoxic, probably because there are fewer roots in the deep layers.
Effect of destocking on fungal diversity and functional guild
Long-term destocking resulted in increased fungal diversity and richness. A previous study also reported that greater duration of destocking led to greater fungal diversity when compared with short-term destocking (14 and 19 years) in semiarid grasslands (Wang et al. 2019). Different from the humped changes in plant diversity (Table S1), fungal diversity significantly increased over time (Table 2). Our findings are consistent with those of a study by Chen et al. (2018), in which no significant relationship existed in plant diversity and fungal diversity in a semiarid grassland. These results suggested that coupling between plant and whole-fungal community probably does not occur in the grasslands of the Loess Plateau.
A substitution of certain groups over time in fungal communities during natural restoration has been reported (Hu et al. 2019; Wang et al. 2019). However, we did not observe such a successional pattern in the present study. This finding disagreed with our previous observation of a significant change in fungal composition during the secondary succession on the abandoned cropland (Zhang et al. 2017). Given the unique characteristics of the different ecosystems, the response of fungi to succession may be ecosystem-specific, and dependent upon climate, soil conditions, and human disturbance (Hu et al. 2019; Kuramae et al. 2011; Peay et al. 2010).
In this study, destocking time drove the fungal OTUs assigned to functional guild (Figs. 1 and 2), consistent with our understanding of the grazing-induced responses of plant communities. In line with the first hypothesis, 35 years of destocking increases the diversity of wood saprotrophs, plant pathogens, and AMF, while it decreases the diversity of animal pathogens and dung saprotrophs. These findings are consistent with those of a previous study on grasslands that reported changes in fungal communities caused by grazing (Eldridge and Delgado-Baquerizo 2018). ABT analysis (Fig. 4a) indicated that increasing plant diversity and coverage was related to an increase in plant pathogens. Plant diversity has been reported to be a dominant factor affecting plant pathogens because plants provide diverse substrates for pathogens (Chen et al. 2018; Yang et al. 2017). A more diverse plant community could have a greater root biomass and diverse root exudates (Cui et al. 2018; Zhang et al. 2019), and thus provide abundant and different resource for fungi. By increasing plant diversity, destocking would likely increase the abundance and diversity of plant pathogens.
Grazing has been reported to have a positive effect on the growth of animal pathogens and dung saprotrophs because the livestock excrement offered rich resource for some animal pathogens and fungi (Eldridge and Delgado-Baquerizo 2018). The destocking decreases the number of herbivores, consequently decreasing the abundance of animal pathogens and dung saprotrophs. This was confirmed by the results of the ABT analysis, in which aboveground biomass was an important predictor of animal pathogens and dung saprotrophs (Fig. 4b-c). Our results supported the resource diversity hypothesis that plant productivity determines soil fungal diversity (Hiiesalu et al. 2017; Yang et al. 2017), and the interaction between plants and fungi depends on nutrient input.
Given the ubiquitous symbiotic relationships between AMF and plants (Koziol and Bever 2017; Ren et al. 2018a), we expected there to be a positive association between plants and AMF. However, the variation in AMF diversity was mainly explained by soil resources (soil organic C, C:N ratio, NO3−-N and moisture), and less by plant properties (Fig. 4e). This finding was confirmed by the regression analysis that showed that plant community was not a predictor of the abundance of AMF. The soil C:N ratio and moisture are usually considered to be key and limiting factors for AMF growth, and both could play important roles for fungi, especially in semiarid areas (Hu et al. 2019; Nie et al. 2018). Soil water availability influenced AMF community richness and composition in soil and roots (Deveautour et al. 2019; Li et al. 2015), while AMF stimulation of fresh residue decomposition can be greater for those residues with high C:N ratios (Wei et al. 2019). However, a positive relationship between fungal growth and N-concentration (Wei et al. 2013) or inorganic N-addition in soil has also been previously reported (Chen et al. 2018).
Effect of soil depth on fungal OTUs assigned to functional guild
We observed strong differences in fungal communities within each depth along the soil profile, as well as differences in the primary environmental variables associated with these changes (Fig. 3; Table 4). These observations agreed with those of a previous study (Wang et al. 2019), in which, decreased bacterial abundance and nutrient availability were observed with soil depth in a semiarid grassland. It is reasonable to consider that the observed differences along the soil profile could be attributed to the changes in plant diversity and biomass, and soil nutrients, which may have caused changes in the available substrates for microorganisms. Similar to other studies (Kramer and Gleixner 2008), soil C and N contents were associated with fungal composition changes, and their effects were correlated with plant biomass (above- and belowground), which was a main source of soil C and N. Furthermore, there were no significant changes in fungal diversity below 20 cm, possibly indicating that fungi in deeper soil might be more adaptable to changing environmental conditions (e.g., nutrient input, porosity, and pH) (Angst et al. 2016; Rumpel and Koegel-Knabner 2011). Decreased variability in fungal communities in the soil profile has been reported by Mushinski et al. (2018), who found that significant changes in the diversity of fungal functional groups only occurred in the surface soil (up to 30 cm) after organic matter removal. Other researchers also found differential distribution of fungal communities with soil depth in different ecosystems (Sun et al. 2018a; Wang et al. 2019). These results probably indicated the niche differentiation of fungi to adapt to changeable soil conditions and are a manifestation of the interaction between microbes and soil. Future studies concerning microbial communities should focus on the deeper soil layers.
Conclusions
Our results indicated that the destocking in semiarid grasslands increased the diversity of plant pathogens, wood saprotrophs, and AMF, while it decreases the diversity of animal pathogens and dung saprotrophs. The effect of destocking on the composition of fungal communities differs according to soil depth, with a greater effect in the near-surface layers (0–10 and 10–20 cm). Plant communities have a greater effect on the diversity of plant pathogens, animal pathogen, dung saprotrophs, and wood saprotrophs, while soil conditions (e.g., organic C, NO3−-N, C:N ratio, and moisture) exerted a greater effect on the diversity of AMF. Our results provide insights into how the lengths of destocking and soil depth influence fungal communities.
References
Angst G, John S, Mueller CW, Koegel-Knabner I, Rethemeyer J (2016) Tracing the sources and spatial distribution of organic carbon in subsoils using a multi-biomarker approach. Sci Rep 6:1–12. https://doi.org/10.1038/srep29478
Boschi C, Baur B (2007) The effect of horse, cattle and sheep grazing on the diversity and abundance of land snails in nutrient-poor calcareous grasslands. Basic Appl Ecol 8:55–65. https://doi.org/10.1016/j.baae.2006.02.003
Bremner JM, Mulvaney CS (1982) Nitrogen-total. In: Page AL, Miller RH, Keeney DR (eds) Methods of soil analysis, part 2, chemical and microbial properties. Agronomy Society of America, Agronomy Monograph 9, Madison, pp 595–624
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, Fierer N, Pena AG, Goodrich JK, Gordon JI, Huttley GA, Kelley ST, Knights D, Koenig JE, Ley RE, Lozupone CA, McDonald D, Muegge BD, Pirrung M, Reeder J, Sevinsky JR, Tumbaugh PJ, Walters WA, Widmann J, Yatsunenko T, Zaneveld J, Knight R (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7:335–336. https://doi.org/10.1038/nmeth.f.303
Chen W, Xu R, Wu Y, Chen J, Zhang Y, Hu T, Yuan X, Zhou L, Tan T, Fan J (2018) Plant diversity is coupled with beta not alpha diversity of soil fungal communities following N enrichment in a semi-arid grassland. Soil Biol Biochem 116:388–398. https://doi.org/10.1016/j.soilbio.2017.10.039
Cheng J, Jing G, Wei L, Jing Z (2016) Long-term grazing exclusion effects on vegetation characteristics, soil properties and bacterial communities in the semi-arid grasslands of China. Ecol Eng 97:170–178. https://doi.org/10.1016/j.ecoleng.2016.09.003
Cui Y, Fang L, Guo X, Wang X, Zhang Y, Li P, Zhang X (2018) Ecoenzymatic stoichiometry and microbial nutrient limitation in rhizosphere soil in the arid area of the northern loess plateau, China. Soil Biol Biochem 116:11–21. https://doi.org/10.1016/j.soilbio.2017.09.025
de Menezes AB, Richardson AE, Thrall PH (2017) Linking fungal-bacterial co-occurrences to soil ecosystem function. Curr Opin Microbiol 37:135–141. https://doi.org/10.1016/j.mib.2017.06.006
De'ath G (2007) Boosted trees for ecological modeling and prediction. Ecology 88:243–251. https://doi.org/10.1890/0012-9658
Derner JD, Boutton TW, Briske DD (2006) Grazing and ecosystem carbon storage in the north American Great Plains. Plant Soil 280:77–90. https://doi.org/10.1007/s11104-005-2554-3
Deveautour C, Power SA, Barnett KL, Ochoa-Hueso R, Donn S, Bennett AE, Powell JR (2019) Temporal dynamics of mycorrhizal fungal communities and co-associations with grassland plant communities following experimental manipulation of rainfall. J Ecol. https://doi.org/10.1111/1365-2745.13267
Edgar RC, Haas BJ, Clemente JC, Quince C, Knight R (2011) UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27:2194–2200. https://doi.org/10.1093/bioinformatics/btr381
Eldridge DJ, Delgado-Baquerizo M (2018) Functional groups of soil fungi decline under grazing. Plant Soil 426:51–60. https://doi.org/10.1007/s11104-018-3617-6
Eldridge DJ, Delgado-Baquerizo M, Travers SK, Val J, Oliver I (2017) Do grazing intensity and herbivore type affect soil health? Insights from a semi-arid productivity gradient. J Appl Ecol 54:976–985. https://doi.org/10.1111/1365-2664.12834
Grau O, Geml J, Perez-Haase A, Ninot JM, Semenova-Nelsen TA, Penuelas J (2017) Abrupt changes in the composition and function of fungal communities along an environmental gradient in the high Arctic. Mol Ecol 26:4798–4810. https://doi.org/10.1111/mec.14227
Hiiesalu I, Bahram M, Tedersoo L (2017) Plant species richness and productivity determine the diversity of soil fungal guilds in temperate coniferous forest and bog habitats. Mol Ecol 26:4846–4858. https://doi.org/10.1111/mec.14246
Hu Y, Veresoglou SD, Tedersoo L, Xu T, Ge T, Liu L, Chen Y, Hao Z, Su Y, Rillig MC, Chen B (2019) Contrasting latitudinal diversity and co-occurrence patterns of soil fungi and plants in forest ecosystems. Soil Biol Biochem 131:100–110. https://doi.org/10.1016/j.soilbio.2019.01.001
Jing Z, Cheng J, Su J, Bai Y, Jin J (2014) Changes in plant community composition and soil properties under 3-decade grazing exclusion in semiarid grassland. Ecol Eng 64:171–178. https://doi.org/10.1016/j.ecoleng.2013.12.023
Koziol L, Bever JD (2017) The missing link in grassland restoration: arbuscular mycorrhizal fungi inoculation increases plant diversity and accelerates succession. J Appl Ecol 54:1301–1309. https://doi.org/10.1111/1365-2664.12843
Kramer C, Gleixner G (2008) Soil organic matter in soil depth profiles: distinct carbon preferences of microbial groups during carbon transformation. Soil Biol Biochem 40:425–433. https://doi.org/10.1016/j.soilbio.2007.09.016
Kuramae E, Gamper H, van Veen J, Kowalchuk G (2011) Soil and plant factors driving the community of soil-borne microorganisms across chronosequences of secondary succession of chalk grasslands with a neutral pH. FEMS Microbiol Ecol 77:285–294. https://doi.org/10.1111/j.1574-6941.2011.01110.x
Leff JW, Jones SE, Prober SM, Barberan A, Borer ET, Firn JL, Harpole WS, Hobbie SE, Hofmockel KS, Knops JMH, McCulley RL, La Pierre K, Risch AC, Seabloom EW, Schuetz M, Steenbock C, Stevens CJ, Fierer N (2015) Consistent responses of soil microbial communities to elevated nutrient inputs in grasslands across the globe. P Natl Acad Sci USA 112:10967–10972. https://doi.org/10.1073/pnas.1508382112
Li X, Zhu T, Peng F, Chen Q, Lin S, Christie P, Zhang J (2015) Inner Mongolian steppe arbuscular mycorrhizal fungal communities respond more strongly to water availability than to nitrogen fertilization. Environ Microbiol 17:3051–3068. https://doi.org/10.1111/1462-2920.12931
Li JW, Liu Y, Hai X, Shangguan Z, Deng L (2019) Dynamics of soil microbial C:N:P stoichiometry and its driving mechanisms following natural vegetation restoration after farmland abandonment. Sci Total Environ 693:133613–133613. https://doi.org/10.1016/j.scitotenv.2019.133613
Marriott CA, Hudson G, Hamilton D, Neilson R, Boag B, Handley LL, Wishart J, Scrimgeour CM, Robinson D (1997) Spatial variability of soil total C and N and their stable isotopes in an upland Scottish grassland. Plant Soil 196:151–162
McGuire KL, Bent E, Borneman J, Majumder A, Allison SD, Treseder KK (2010) Functional diversity in resource use by fungi. Ecology 91:2324–2332. https://doi.org/10.1890/09-0654.1
McMurdie PJ, Holmes S (2013) Phyloseq: An R package for reproducible interactive analysis and graphics of microbiome census data. Plos One 8. https://doi.org/10.1371/journal.pone.0061217
Mushinski RM, Gentry TJ, Dorosky RJ, Boutton TW (2017) Forest harvest intensity and soil depth alter inorganic nitrogen pool sizes and ammonia oxidizer community composition. Soil Biol Biochem 112:216–227. https://doi.org/10.1016/j.soilbio.2017.05.015
Mushinski RM, Gentry TJ, Boutton TW (2018) Organic matter removal associated with forest harvest leads to decade scale alterations in soil fungal communities and functional guilds. Soil Biol Biochem 127:127–136. https://doi.org/10.1016/j.soilbio.2018.09.019
Nguyen NH, Song Z, Bates ST, Branco S, Tedersoo L, Menke J, Schilling JS, Kennedy PG (2016) FUNGuild: An open annotation tool for parsing fungal community datasets by ecological guild. Fungal Ecol 20:241–248. https://doi.org/10.1016/j.funeco.2015.06.006
Nie S, Lei X, Zhao L, Brookes PC, Wang F, Chen C, Yang W, Xing S (2018) Fungal communities and functions response to long-term fertilization in paddy soils. Appl Soil Ecol 130:251–258. https://doi.org/10.1016/j.apsoil.2018.06.008
Odum EP (1969) The strategy of ecosystem development. Science (New York, NY) 164:262–270. https://doi.org/10.1126/science.164.3877.262
Olsen SR, Sommers LE (1982) Phosphorous. In: Page AL, Miller RH, Keeney DR (eds) Methods of soil analysis, part 2, chemical and microbial properties. Agronomy monograph 9. Agronomy Society of America, Madison, pp 403–430
Peay KG, Kennedy PG, Davies SJ, Tan S, Bruns TD (2010) Potential link between plant and fungal distributions in a dipterocarp rainforest: community and phylogenetic structure of tropical ectomycorrhizal fungi across a plant and soil ecotone. New Phytol 185:529–542. https://doi.org/10.1111/j.1469-8137.2009.03075.x
R Development Core Team (2019) R version 3.6.0: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna
Ren GH, Wang CX, Dong KH, Zhu HS, Wang YC, Zhao X (2018a) Effects of grazing exclusion on soil-vegetation relationships in a semiarid grassland on the loess plateau, China. Land Degrad Dev 29:4071–4079. https://doi.org/10.1002/ldr.3164
Ren H, Eviner VT, Gui W, Wilson GWT, Cobb AB, Yang G, Zhang Y, Hu S, Bai Y, Gallery R (2018b) Livestock grazing regulates ecosystem multifunctionality in semi-arid grassland. Funct Ecol 32:2790–2800. https://doi.org/10.1111/1365-2435.13215
Rousk J, Baath E, Brookes PC, Lauber CL, Lozupone C, Caporaso JG, Knight R, Fierer N (2010) Soil bacterial and fungal communities across a pH gradient in an arable soil. ISME J 4:1340–1351. https://doi.org/10.1038/ismej.2010.58
Rumpel C, Koegel-Knabner I (2011) Deep soil organic matter-a key but poorly understood component of terrestrial C cycle. Plant Soil 338:143–158. https://doi.org/10.1007/s11104-010-0391-5
Schmidt SK, Nemergut DR, Darcy JL, Lynch R (2014) Do bacterial and fungal communities assemble differently during primary succession? Mol Ecol 23:254–258. https://doi.org/10.1111/mec.12589
Smits NAC, Willems JH, Bobbink R (2008) Long-term after-effects of fertilisation on the restoration of calcareous grasslands. Appl Veg Sci 11:279–U292. https://doi.org/10.3170/2008-7-18417
Su J, Jing G, Jin J, Wei L, Liu J, Cheng J (2017) Identifying drivers of root community compositional changes in semiarid grassland on the loess plateau after long-term grazing exclusion. Ecol Eng 99:13–21. https://doi.org/10.1016/j.ecoleng.2016.11.050
Sun RB, Li WY, Dong WX, Tian YP, Hu CS, Liu BB (2018a) Tillage changes vertical distribution of soil bacterial and fungal communities. Frontiers in microbiology 9. ARTN 699. https://doi.org/10.3389/fmicb.2018.00699
Sun Y, He XZ, Hou F, Wang Z, Chang S (2018b) Grazing increases litter decomposition rate but decreases nitrogen release rate in an alpine meadow. Biogeosciences 15:4233–4243. https://doi.org/10.5194/bg-15-4233-2018
Wang WY, Wang QJ, Wang HC (2006) The effect of land management on plant community composition, species diversity, and productivity of alpine Kobersia steppe meadow. Ecol Res 21:181–187. https://doi.org/10.1007/s11284-005-0108-z
Wang Z, Zhang Q, Staley C, Gao H, Ishii S, Wei X, Liu J, Cheng J, Hao M, Sadowsky MJ (2019) Impact of long-term grazing exclusion on soil microbial community composition and nutrient availability. Biol Fertil Soils 55:121–134. https://doi.org/10.1007/s00374-018-01336-5
Wei C, Yu Q, Bai E, Lu X, Li Q, Xia J, Kardol P, Liang W, Wang Z, Han X (2013) Nitrogen deposition weakens plant-microbe interactions in grassland ecosystems. Glob Chang Biol 19:3688–3697. https://doi.org/10.1111/gcb.12348
Wei L, Vosatka M, Cai B, Ding J, Lu C, Xu J, Yan W, Li Y, Liu C (2019) The role of Arbuscular Mycorrhiza Fungi in the decomposition of fresh residue and soil organic carbon: a mini-review. Soil Sci Soc Am J 83:511–517. https://doi.org/10.2136/sssaj2018.05.0205
Yang T, Adams JM, Shi Y, He J-s, Jing X, Chen L, Tedersoo L, Chu H (2017) Soil fungal diversity in natural grasslands of the Tibetan plateau: associations with plant diversity and productivity. New Phytol 215:756–765. https://doi.org/10.1111/nph.14606
Yang F, Niu KC, Collins CG, Yan XB, Ji YG, Ling N, Zhou XH, Du GZ, Guo H, Hu SJ (2019) Grazing practices affect the soil microbial community composition in a Tibetan alpine meadow. Land Degrad Dev 30:49–59. https://doi.org/10.1002/ldr.3189
Zeng QC, An SS, Liu Y (2017) Soil bacterial community response to vegetation succession after fencing in the grassland of China. Sci Total Environ 609:2–10. https://doi.org/10.1016/j.scitotenv.2017.07.102
Zhang C, Liu G, Xue S, Wang G (2016) Soil bacterial community dynamics reflect changes in plant community and soil properties during the secondary succession of abandoned farmland in the loess plateau. Soil Biol Biochem 97:40–49. https://doi.org/10.1016/j.soilbio.2016.02.013
Zhang C, Liu G, Song Z, Qu D, Fang L, Deng L (2017) Natural succession on abandoned cropland effectively decreases the soil erodibility and improves the fungal diversity. Ecol Appl 27:2142–2154. https://doi.org/10.1002/eap.1598/full
Zhang C, Liu G, Song Z, Wang J, Guo L (2018) Interactions of soil bacteria and fungi with plants during long-term grazing exclusion in semiarid grasslands. Soil Biol Biochem 124:47–58. https://doi.org/10.1016/j.soilbio.2018.05.026
Zhang C, Wang J, Liu G, Song Z, Fang L (2019) Impact of soil leachate on microbial biomass and diversity affected by plant diversity. Plant Soil 439:505–523. https://doi.org/10.1007/s11104-019-04032-x
Funding
This work was financially supported by the National Natural Sciences Foundation of China (41771554); Shaanxi Innovation Support Plan for Youth (2019KJXX-081) and the National Key Research and Development Program of China (2016YFC0501707).
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible Editor: Jeff R. Powell.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(DOCX 516 kb)
Rights and permissions
About this article
Cite this article
Wang, J., Liu, G., Zhang, C. et al. Effect of long-term destocking on soil fungal functional groups and interactions with plants. Plant Soil 448, 495–508 (2020). https://doi.org/10.1007/s11104-020-04452-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11104-020-04452-0