Abstract
Key message
Ethanol priming induces heat stress tolerance by the stimulation of unfolded protein response.
Abstract
Global warming increases the risk of heat stress-related yield losses in agricultural crops. Chemical priming, using safe agents, that can flexibly activate adaptive regulatory responses to adverse conditions, is a complementary approach to genetic improvement for stress adaptation. In the present study, we demonstrated that pretreatment of Arabidopsis with a low concentration of ethanol enhances heat tolerance without suppressing plant growth. We also demonstrated that ethanol pretreatment improved leaf growth in lettuce (Lactuca sativa L.) plants grown in the field conditions under high temperatures. Transcriptome analysis revealed a set of genes that were up-regulated in ethanol-pretreated plants, relative to water-pretreated controls. Binding Protein 3 (BIP3), an endoplasmic reticulum (ER)-stress marker chaperone gene, was among the identified up-regulated genes. The expression levels of BIP3 were confirmed by RT-qPCR. Root-uptake of ethanol was metabolized to organic acids, nucleic acids, amines and other molecules, followed by an increase in putrescine content, which substantially promoted unfolded protein response (UPR) signaling and high-temperature acclimation. We also showed that inhibition of polyamine production and UPR signaling negated the heat stress tolerance induced by ethanol pretreatment. These findings collectively indicate that ethanol priming activates UPR signaling via putrescine accumulation, leading to enhanced heat stress tolerance. The information gained from this study will be useful for establishing ethanol-mediated chemical priming strategies that can be used to help maintain crop production under heat stress conditions.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Agriculture is one of the sectors that is highly likely to be compromised by global warming. Based on the recent projected crop and climate models, it is anticipated that crop production, for example in maize, will decrease by −6 to −24% at the end-of-century (Jägermeyr et al. 2021). A crop management system that can flexibly induce stress adaptation by the pretreatment of growing crops with safe, chemical compounds represents an important approach that can complement genetic improvements in stress tolerance (Savvides et al. 2016; Sako et al. 2021b). Such a strategy is referred to as ‘chemical priming’. Various types of molecules have been found to function as chemical priming agents (Savvides et al. 2016; Sako et al. 2021b) and include amino acids, such as proline (Ambreen et al. 2021), phytohormones, such as salicylic acid (Li et al. 2014; Khan et al. 2015; Qamar et al. 2022), methyl jasmonate (Mageroy et al. 2020; Valenzuela-Riffo et al. 2020) and strigolactone (Ha et al. 2014; Omoarelojie et al. 2020), and other molecules, such as hydrogen sulfide (H2S) (Liu et al. 2021), nitric oxide (NO)(Dong et al. 2014), sodium nitroprusside (Hameed et al. 2021), zinc oxide nanoparticles (Waqas Mazhar et al. 2022), acetic acid (Kim et al. 2017; Utsumi et al. 2019), ascorbic acid (Qamar et al. 2022), nicotinic acid (Ahmad et al. 2021), and polyamines (PAs) (Shu et al. 2015; Chen et al. 2019; Collado-González et al. 2021; Islam et al. 2021; Jahan et al. 2022). Importantly, chemical compounds used as priming agents must be effective at low concentrations to be cost-effective, as well as environment-friendly. We previously reported that the chemical priming of plants with low concentrations of ethanol, an inexpensive compound (ex. $0.36 for 10 gallons of 10 mM ethanol, Wako) and widely used as a disinfectant, improves salt and high-light stress tolerance (Nguyen et al. 2017; Sako et al. 2021a). In these literatures, the effective concentration of ethanol was 0.3% (51 mM). Therefore, chemical priming by ethanol represents an attractive approach for the development of a crop management system that can increase stress tolerance.
Under heat stress conditions, proteins in the endoplasmic reticulum (ER) can be denatured or aggregated (Ohama et al. 2017; Nishad and Nandi 2021). The damaged proteins cannot be appropriately folded and accumulate in the ER. The condition that the unfolded or misfolded proteins excessively accumulate in the ER is defined as ER stress (Walter and Ron 2011). Significant efforts have been made to elucidate the molecular mechanisms that can restore net protein-folding capacity in the ER under stress conditions. This process has been shown to be regulated by the unfolded protein response (UPR) system, which is broadly conserved in eukaryotes (Angelos et al. 2017; Pastor-Cantizano et al. 2020). The UPR alleviates the damage of ER stress, which leads to the restoration of ER function (Iwata and Koizumi 2012; Pastor-Cantizano et al. 2020). On the other hand, when ER stress is prolonged or severe, programmed cell death occurs in the cells (Iwata and Koizumi 2012; Pastor-Cantizano et al. 2020). The UPR activates the synthesis of ER-resident molecular chaperones to alleviate the development of ER stress. Binding Protein 3 (BIP3), an ER stress marker gene, encodes an ER-resident cognate of heat shock protein 70, and its role in UPR has been characterized (Martínez and Chrispeels 2003; Kamauchi et al. 2005; Iwata et al. 2018). BIP1 to BIP3 are found in the Arabidopsis genome as genes encoding a chaperone family protein. Notably, the expression of BIP3 is highly induced under ER stress conditions (Iwata et al. 2009). Moreover, it has been shown that BIP3 expression is regulated by specific basic leucine zipper (bZIP) transcription factors, such as bZIP60, which is an important component of UPR activation (Humbert et al. 2012; Pastor-Cantizano et al. 2020).
Polyamines are metabolites that are widely distributed in organisms. Putrescine, spermidine, spermine, and thermospermine are recognized in plants as polyamines (Chen et al. 2019). They play a functional role in a variety of developmental processes and biotic and abiotic stress responses (Mustafavi et al. 2018). Notably, spermine has been reported to activate the UPR through its regulation of bZIP60 expression (Sagor et al. 2015).
It is important to elucidate the regulatory mechanism of the ethanol-pretreatment-induced enhancement of heat stress tolerance in plants. To tackle this, in the present study, we investigated the effects of the ethanol pretreatment on physiological response to heat stress and analyzed the changes in the profiles of gene expression and metabolite accumulation in response to ethanol treatment.
Materials and methods
Plant growth conditions
Arabidopsis wild-type plants (ecotype Col-0) and two bzip60 mutants (SAIL_838_C12 and SALK_050203 obtained from ABRC) and lettuce plants (“Leaf Lettuce Green” commercially purchased from Sakata Seed Corp., Yokohama, Japan) were used in the experiments. The plants were grown on petri plates containing 1 × MS, 1% sucrose and 0.8% agar medium. Petri plates containing 30 Arabidopsis plants per plate were placed in a growth incubator for 14 days at 22℃ under a 18 h light / 6 h dark condition. For soil-based experiments, seeds were sown into pots containing either Professional Potting Soil (Dio Chemicals Ltd., Tokyo, Japan) for Arabidopsis or 1:1 Professional Potting Soil and humus mix soil (Kurotsuchi A-No.2, Tachikawa Heiwa Nouen Co. Ltd., Kanuma, Japan) for the lettuce plants. After sowing, plants were grown for 3 weeks at 22 °C under a 18 h light / 6 h dark condition in a growth room.
Ethanol treatment and heat stress tolerance test
Heat stress treatment for MS-agar-grown Arabidopsis plants was performed according to the procedure reported in Nguyen et al. (2015). After 5 h of exposure to the initial light conditions (ZT5), MS-agar-grown Arabidopsis plants were placed in an air incubator (SANYO incubator MIR-153, Sanyo Electric Co. Ltd, Osaka, Japan) set at 43.5 °C with dim light. MS plates containing ethanol were prepared as follows: (1) MS medium was autoclaved, (2) the medium was cooled down, (3) before the medium was solidified, ethanol was added into the medium and mixed, (4) the medium was poured into plates and solidified. We checked the surface temperature of the MS agar petri plates placed in the air incubator. The temperatures gradually approached 43.5 °C over a period of 100 min and then remained stable at 43.5 °C (Fig. 1A). After the indicated time of exposure to the heat stress conditions, the plants were returned to the growth incubator set at 22 °C and were grown for an additional 7 days. The viability of each plant was assessed based on the emergence of new leaves. The heat stress tests for soil-grown Arabidopsis plants were performed as follows: The pots with 3-week-old plants were placed onto paper towels to remove excess water. Then the bottoms of the pots were dipped into 20 mM ethanol solution for a period of 3 days in a tray. The pots placed in the tray were covered with a vinyl film to prevent them from drying out. The tray was then exposed to a heat stress of 50 °C for 3 h in the air incubator. The tray was returned to the growth room after the heat stress treatment, where the temperature was maintained at 22 °C and tap water was put in the tray, although the vinyl film cover was retained for 24 h. The survival rate of Arabidopsis plants was assessed 10 days subsequent to the heat stress.
Ethanol pretreatment-induced heat stress tolerance in Arabidopsis. A Surface temperature of MS plates placed in a 43.5 °C air incubator. B Effect of various concentration of ethanol on the heat tolerance of Arabidopsis plants grown on MS agar medium. Seeds were sown and germinated on the MS media containing 0, 2, 10, 15, 20, and 60 mM ethanol and grown for 14 days. The plants were then subjected to a heat stress treatment (43.5 °C for 3 h) followed by non-stressed conditions for 7 days. C Survival rate of ethanol-pretreated plants following heat stress treatments. Two-week-old plants were treated at 43.5 °C for 2.5, 3.5, and 4 h. Asterisks indicate a significant difference (p < 0.05) compared to control plants without an ethanol pretreatment (n = 4, 30 plants were used in each replicate) according to a Tukey’s multiple comparison test. D Index of injury (Id) values calculated from ion leakage measurements in 10 mM ethanol-pretreated and water-pretreated plants (n = 20). Asterisks indicate a significant difference (p < 0.01) compared to that of control plants based on a t-test
The heat stress tests for lettuce plants grown in pots with soil were performed in a similar manner to the Arabidopsis plants. Three-week-old plants were treated with 20 mM ethanol by dipping the bottoms of the pots in a tray containing the solution for 3 days and then the plants were exposed to heat stress at 50 °C for 3 h in the air incubator. The pots were returned to the growth room after the heat stress treatment, where the temperature was kept at 22 °C and a vinyl film cover was retained for 24 h. The survival rate of the lettuce plants was estimated on the 10th day after heat stress.
Field experiments using lettuce plants were conducted as follows: lettuce seedlings were grown in pots with soil for 3 weeks and pretreated with subirrigation of 20 mM ethanol by placing the pots in a tray containing the ethanol solution for 2 days or left untreated (control). Twenty-six water (control)-pretreated and 26 ethanol-pretreated plants were then transplanted on February 16, 2020 to the field which was covered with a rain shelter (35°24ʹ15.8ʹʹN 139°32′59.5ʹʹE) located on the Maioka campus, Yokohama City University. The diagram of the arrangement of the plants in the field is shown in Supplemental Fig. 1. The lettuce plants were subsequently treated with 100 mL of 20 mM ethanol per plant at 2-week intervals. Survival rates of lettuce plants were assessed on May 26, 2020. Temperatures in the rain shelter were monitored and recording using a Ondotori TR-72nw (T&D Corporation, Matsumoto, Japan).
Ion leakage analysis
Shoots of Arabidopsis seedlings grown on MS plates with and without an ethanol pre-treatment for 11 days were placed in a test tube containing 2 mL of distilled water. The tubes were heated in a water bath at 43 °C for 30 min, and then cooled in a water bath set at 22 °C for 30 min. Heat-stressed and control samples were slowly shaken overnight at 30 rpm, after which electrical conductivity was measured with a Conductivity meter B-173 (Horiba Ltd, Kyoto, Japan). The samples were then autoclaved at 120 °C and shaken at room temperature for several hours, after which electrical conductivity of the samples was again measured. The index of injury (Id) was calculated based on the protocol described for ion leakage analysis of Arabidopsis seedlings by Ilík et al. (2018).
Reverse transcription: quantitative PCR (RT-qPCR)
Each treatment was analyzed using three biological replicates, and a total of ten plants was sampled for each treatment. Total RNAs were extracted using purelink Plant RNA Reagent (Thermo Fisher Scientific Inc., Waltham, MA, USA). RT-qPCR analysis was performed on a StepOnePlus system (Thermo Fisher Scientific Inc., Waltham, MA, USA) as previously described (Nguyen et al., 2015; Matsui et al., 2017). Three independent biological replicates were used in each of the RT-qPCR analyses. The sequences of the utilized primer sets were as follows:
ACT2, 5ʹ-GATCTCCAAGGCCGAGTATGAT-3ʹ and 5ʹ-CCCATTCATAAAACCCCAGC-3ʹ;
18S rRNA, 5ʹ-GGAAGTTTGAGGCAATAACAGG-3ʹ and 5ʹ-ATTGCAATGATCTATCCCCATC-3ʹ,
HSP101, 5ʹ-CCAAGTGAGCTGAGATCTTGGT-3ʹ and 5ʹ-GTTCCACAACAAGCATA-3ʹ;
HSP70b, 5ʹ-TGTCTCGTCATCGTTGCATT-3ʹ and 5ʹ-TCTTCCTTTTTCAGGGCAAA-3ʹ;
HTT1, 5ʹ-CCAAGTGAGCTGAGATCTTGGT-3ʹ and 5ʹ-GTTCCACAACAAGCATACGG-3ʹ;
ZAT12, 5ʹ-TCCGATGGGACAAGCTTTGG-3ʹ and 5ʹ-AAGCCACTCTCTTCCCACTG-3ʹ;
APX1, 5ʹ-GCACTATTGGACGACCCTGT-3ʹ and 5ʹ-AGCAAACCCAAGCTCAGAAA-3ʹ;
APX2, 5ʹ-AAGTTGAGCCACCTCCTGAA-3ʹ and 5ʹ-GTGTGTCCACCAGACAATGC-3ʹ;
HSFA2, 5ʹ-GTCTTCAGCTATAAAGGGATTCAGA-3ʹ and 5ʹ-ATTCTGCAAACCCATGTTCC-3ʹ,
BIP3, 5ʹ-CGAAACGTCTGATTGGAAGAA-3ʹ and 5ʹ-GGCTTCCCATCTTTGTTCACA-3ʹ.
Microarray analysis
Arabidopsis WT seeds were sown on MS medium with and without 2 mM ethanol and grown for 2 weeks. Whole plants were then subsequently collected at ZT5. Microarray analysis was performed as previously described (Matsui et al. 2017). Raw data were extracted using the Feature Extraction program ver. 9.1. RMA normalization was performed for signals of microarray probes using the limma (Ritchie et al. 2015) and RankProd packages (Del Carratore et al. 2017) in the R 4.0.5 program (R Core Team). The microarray data were deposited in the GEO database (GSE171036).
13C-tracer analysis using nuclear magnetic resonance (NMR) spectroscopy
The 13C labeling method for plants described in several previous reports (Kikuchi et al. 2004; Kikuchi and Hirayama 2007; Tian et al. 2007) was utilized in the present study. Briefly, 2-week-old Arabidopsis seedlings assimilated either 10 mM of 13C2-ethanol or non-labeled ethanol (Guaranteed reagent, FUJIFILM Wako Pure Chemical Corporation, Osaka, Japan) via root-uptake for 24 h. The plants were harvested after the 13C-labeling and immediately freeze-dried for use in the metabolome analysis. The lyophilized plants were milled and extracted with D2O/KPi (100 mM, pH 7.0) buffer solution. Then, two-dimensional (2D) nuclear magnetic resonance spectroscopy (NMR) analysis was performed using the method that has been previously described in several reports (Sekiyama et al. 2010; Komatsu et al. 2014, 2016).
Metabolome analysis by gas chromatography-time of flight/mass spectrometry (GC-TOF/MS)
GC-TOF/MS analysis was conducted on samples that were treated with non-labeled ethanol (Guaranteed reagent, FUJIFILM Wako Pure Chemical Corporation, Osaka, Japan) using the procedures described in earlier studies (Jonsson et al. 2005; Kusano et al. 2007a, 2007b; Redestig et al. 2009). Approximately 25 mg fresh weight of root or shoot tissue per one mL of extraction medium containing ten stable isotope reference compounds was used in the extraction of metabolites.
Putrescine and ornithine treatment
Seeds were sown on MS media containing 0.2 mM putrescine or 0.2 mM ornithine. After the cultivation for 14 days, seedlings were subjected to heat stress at 43.5 °C for the indicated hours and then placed under non-stressed condition for 7 days.
Arabidopsis cultivation in MS liquid medium and treatment with LaCl3
The bottom portions (6 mm) of the disposable pipette tips (10 μl Short Tip, Fukaekasei Co. Ltd., Kobe, Japan and WATSON Co. Ltd., Tokyo, Japan) were cut and 0.8% agarose MS medium was poured into the upper portions of the cut tips. These were set in a tip case (200 μl Tip Rack, BM Equipment Co. Ltd., Tokyo, Japan) with the void holes covered with parafilm. A total of 220 mL of liquid MS medium was added to the tip case to partially submerge the tips. Arabidopsis seedlings were grown in this system at 22 °C for 2 weeks. The seedlings were then pretreated with either 10 mM ethanol, 0.5 mM LaCl3, or 10 mM ethanol supplemented with 0.5 mM LaCl3 for 3 days and then exposed to a heat stress at 45 °C for 3.5 h.
Difluoromethylornithine (DFMO) and difluoromethylarginine (DFMA) treatment
Arabidopsis seedlings were grown in MS liquid culture for 1 week according to the method of Nguyen et al. (2017). The seedlings were treated with 2 mM DFMO, 2 mM DFMA and 10 mM ethanol for 12 h, and then subjected to heat treatment at 45 °C for 2.5 h and then placed under non-stressed condition for 7 days.
Tunicamycin (Tm) and azetidine-2-carboxylic acid (AZC) treatment
Seeds were sown on MS media containing 0.015 mg / L Tm or 15 μM AZC. After the cultivation for 14 days, seedlings were subjected to heat stress at 43.5 °C for 3 h and then placed under non-stressed condition for 7 days.
Results
Ethanol pretreatment enhances the heat stress tolerance of plants grown on MS agar medium and in soil
A heat stress test was conducted on Arabidopsis plants to determine the effect of an ethanol pretreatment on heat stress tolerance. The heat stress was administered in an air incubator as previously described (Nguyen et al. 2015). Arabidopsis plants grown on MS solid medium supplemented with either 0, 2, 10, 15, 20, or 60 mM ethanol for 2 weeks were subjected to a heat stress and then returned to control conditions. The plants subjected to a heat stress were photographed after 1 week (Fig. 1B) and survival rates were assessed (Fig. 1C). The viability of plants grown on MS solid medium with 0 mM ethanol decreased to approximately 30% and 10% by exposure to a heat stress treatment of 2.5 h and 3 h, respectively (Fig. 1C). In contrast, the survival rates of plants grown on MS solid medium containing 10 mM ethanol or greater exhibited a viability of 70% or more after exposure to a 2.5 h heat stress. After exposure to a 3 h heat stress, the survival rates of plants grown on MS solid medium containing 10 mM, 15 mM, 20 mM, and 60 mM ethanol showed 30%, 51%, 86%, and 60%, respectively (Fig. 1C). These results indicate that plants grown on MS medium supplemented with ethanol have an increasing level of viability as the concentration of ethanol increased.
Ion leakage assays were conducted to estimate the degree of damage caused by the heat stress treatment. An index of damage (Id) was calculated as reported previously (Ilík et al. 2018). Arabidopsis plants grown on MS medium containing 10 mM ethanol had a significantly lower Id value than plants grown on MS medium without ethanol (Fig. 1D), suggesting that the ethanol pretreatment alleviates the damage caused by the heat stress.
In some cases, a high concentration of a chemical priming agent can adversely affect growth (Kim et al. 2017). Therefore, we evaluated the growth of Arabidopsis plants grown on agar plates supplemented with various concentrations of ethanol. Results indicated that no significant differences in growth were observed between untreated control plants and plants treated with < 10 mM ethanol, however, concentrations > 15 mM ethanol inhibited plant growth in a dose-dependent manner (Supplemental Fig. 2A, B).
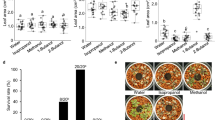
We next investigated the heat stress tolerance of Arabidopsis plants pretreated with ethanol and grown on soil. Ethanol was administered to 3-week-old plants from the bottom of pots for 3 days, and then subjected to a heat treatment. Soil-grown plants not treated with ethanol exhibited about a 10% survival rate after exposure to a 3 h heat stress treatment (Fig. 2A). In contrast, soil-grown plants pretreated with ethanol exhibited a survival rate of about 70% (Fig. 2A), indicating that the ethanol pretreatment was also effective on soil-grown Arabidopsis plants. We also conducted the test on soil-grown lettuce plants. Three-week-old lettuce seedlings grown in pots containing soil were treated with ethanol for 3 days, in the same manner used to treat soil-grown Arabidopsis plants, and subjected to a heat stress treatment. Lettuce leaf growth in the ethanol-pretreated plants following the heat stress treatment exhibited less inhibition than the leaf growth in non-ethanol-treated control plants (Fig. 2B). These results indicate that ethanol pretreatment exerts a beneficial effect on leaf growth in lettuce plants subjected to heat stress. We also investigated the effect of the ethanol pretreatment on the heat tolerance of lettuce plants grown in the field. Notably, the ethanol pretreatment improved the survival rates and leaf growth of plants exposed to high temperature conditions under field conditions (Fig. 2C).
Effect of an ethanol pretreatment on heat stress tolerance in Arabidopsis and lettuce plants grown in pots with soil. A Appearance and survival rate of Arabidopsis Col-0 plants. Ethanol was administered to 3-week-old plants from the bottom of pots for three days, and then plants were subjected to a heat stress treatment. Plants were returned to 22 °C after the heat stress treatment. Photos were taken on the 10th day after the heat stress treatment and the survival rate of the Arabidopsis plants was determined. Asterisks indicate a significant difference (p < 0.01) compared to that of control plants based on a t-test. B Appearance of lettuce plants tested using pots with soil. Three-week-old lettuce seedlings grown in pots with soil were treated with ethanol for 3 days and subjected to a heat stress treatment (at 50 °C for 3 h). Plants were returned to 22 °C after the heat stress treatment. Photos of the lettuce plants were taken on the 10th day after the heat stress treatment and the fresh weight of the lettuce plants was recorded. Asterisks indicate a significant difference (p < 0.01) compared to that of control plants based on a t-test. C Appearance of lettuce plants tested in the field and changes in air temperature under the rain shelter. Lettuce seedlings were grown in pots with soil for 3 weeks and pretreated with ethanol for 2 days. The plants were then transplanted to a field covered by a vinyl rain shelter and grown for approximately 100 days from Feb. 16, 2020 to May 26, 2020. The lettuce plants were treated with ethanol at 2-week intervals during the period of field cultivation. Photos were taken on the last day of field cultivation
Heat stress treatment following ethanol pretreatment of Arabidopsis plants did not result in an increase in the expression of reactive oxygen species (ROS) detoxification-related and heat stress-inducible genes
A previous study reported that salt stress tolerance was enhanced by an ethanol pretreatment by the effect of ethanol on the expression of genes encoding the antioxidant enzymes, ascorbate peroxidase (APX) 1 and APX2, and a gene encoding the ZAT12 transcription factor (Nguyen et al. 2017). Expression of APX1 is regulated by the transcriptional regulator, ZAT12 (Rizhsky et al. 2004). Therefore, we investigated the expression of these genes using RT-qPCR in ethanol-treated and non-treated Arabidopsis plants during exposure to a heat stress treatment. ROS generation is known to function as a stress signal and ROS detoxification is an essential aspect of heat stress tolerance in plants (Choudhury et al. 2017). The expression level of APX1, APX2 and ZAT12 in both ethanol- pretreated and non-treated plants increased in response to the heat stress treatment, however, the inducibility tended to be unexpectedly lower in the ethanol-treated plants than in the untreated plants (Fig. 3A). We also analyzed the expression level of representative heat-inducible genes, including HSFA2, HSP101, HTT1, and HSP70B (Fig. 3B). Similar to the results obtained for APX1, APX2, and ZAT12, plants grown on ethanol-containing medium also exhibited a weaker induction in the expression of these heat-inducible genes, relative to their expression in non-treated plants. These results suggest that ethanol-mediated heat stress tolerance may potentially function through a molecular mechanisms that is different from the mechanism associated with ethanol-mediated salt stress tolerance (Nguyen et al. 2017).
RT-qPCR analysis of gene expression in response to heat stress in ethanol-treated and untreated Arabidopsis plants. A APX1 and APX2 are ROS-related genes. ZAT12 is a gene that has been reported as a transcriptional regulator of APX1 (Rizhsky et al. 2004). B HSFA2, HSP101, HTT1, and HSP70b are representative heat stress-inducible genes. A and B The bar graphs illustrate the relative expression level of the indicated genes relative to the expression of ACT2 (n = 3), a housekeeping gene. Asterisks indicate a significant difference (p < 0.01) compared to that of control plants based on a t-test
RT-qPCR analysis of the expression of genes found to be up-regulated in a transcriptome analysis of ethanol-pretreated Arabidopsis plants
A microarray analysis was conducted to comprehensively examine changes in gene expression in Arabidopsis plants grown on MS medium supplemented with ethanol. Results indicated that the expression of 11 genes was higher in ethanol-pretreated plants than in non-ethanol-pretreated plants (FC > 1.5, p < 0.05, FDR < 0.16) (Supplemental Table 1). Notably, Binding Protein 3 (BIP3), an ER stress marker gene encoding an ER-resident cognate of heat shock protein 70 (Nagashima et al. 2011; Sagor et al. 2015; Iwata et al. 2018), was among the group of up-regulated genes. The fold change value of BIP3 was 1.88 (Supplemental Table 1). The change in BIP3 expression identified in the microarray analysis was further confirmed by RT-qPCR analysis, which showed 1.83-times higher in the ethanol-pretreated plants than in the non-ethanol-pretreated plants (Supplemental Fig. 3).
Root uptake of 13C2-ethanol analyzed by 2D-NMR
13C-tracer analysis using 13C-heteronuclear single quantum coherence (HSQC) NMR spectroscopy was conducted to characterize changes in the metabolism of Arabidopsis plants caused by the exogenous application of ethanol to roots. Labeled 13C-acetic acid, which was apparently metabolized relatively early from the labeled ethanol, and other metabolites including sugars, alcohols, amines, and nucleic acids were detected in the plants treated with the 13C2-ethanol, even though the time allotted for root-uptake of 13C2-ethanol was only 24 h (Supplemental Fig. 5; Supplemental Table 2). These data indicate that the molecules of ethanol supplied exogenously were incorporated and widely metabolized within the cells and that a variety of metabolic pathways were activated by the root-uptake of 13C2-ethanol. 13C-labeled organic acids, amino acids, as well as sugars metabolized from 13C2-ethanol were detected. This indicates carbon central metabolisms through TCA were totally activated.
Metabolomic analysis of ethanol-treated Arabidopsis plants
The level of 88 metabolites in Arabidopsis plants treated with ethanol applied exogenously to the root system was investigated by GC-TOF/MS to characterize changes in the metabolome induced by the ethanol. The results revealed that the ethanol treatment affected the abundance of certain metabolites (Supplemental Table 3). For example, in the roots and/or shoots of ethanol-treated plants, amino acids (lysine and threonine), ornithine, 3-aminopiperidin-2-one, glycolic acid, glutaric acid, suberic acid, sugars (isomaltose and raffinose), and terpenoid-related metabolites (phytol, sitosterol and stigmasterol) were more than 2.5-times greater than they were in control-treated (water-treated) (non-ethanol) roots and/or shoots. The activated pathways appeared to include the putrescine biosynthesis-related pathway because the level of both putrescine and arginine, which serve as a source of putrescine biosynthesis, were elevated in the ethanol-treated plants compared to the control plants (Supplemental Table 3).
Increased levels of putrescine in ethanol-treated Arabidopsis plants is associated with increased heat tolerance
Recent studies have reported that polyamines, especially putrescine, is involved in the increased abiotic stress tolerance (Shu et al. 2015; Collado-González et al. 2021; Islam et al. 2021; Jahan et al. 2022). Considering that the contents of putrescine and ornithine were 1.7 and 5.2 times higher in the ethanol-pretreated shoots than in the control-treated ones (Supplemental Fig. 4), it is possible that ethanol increases heat tolerance through the accumulation of these polyamines. Therefore, we investigated the effect of polyamine pretreatment on heat stress tolerance. Arabidopsis plants that received a putrescine pretreatment had a significantly higher survival rate (approximately 45%) when subjected to a heat stress for 3.5 h, compared to non-treated control plants, which had a survival rate of about 20% when subjected to the same heat stress treatment (Fig. 4A). The survival rate of ornithine-pretreated plants; however was similar to that of the untreated control plants (Fig. 4B).
Putrescine pretreatment enhances heat stress tolerance in Arabidopsis. A and B Photos and survival rate of Arabidopsis plants grown on MS media containing A 0.2 mM putrescine or B 0.2 mM ornithine, subjected to a heat stress at 43.5 °C for the indicated hours and then placed under non-stressed conditions for another 7 days. Asterisks indicate a significant difference (p < 0.05) compared to MS plate without ethanol as determined by a t-test (n = 4, 30 plants were used for each replicate)
Ethanol treatment promotes heat stress tolerance in Arabidopsis through UPR signaling
We subsequently examined if the heat stress tolerance induced by ethanol priming was associated with UPR signaling brought about by the increase of endogenous polyamines caused by the ethanol pretreatment. UPR excitation by polyamines has been reported to be inhibited by the Ca2+-channel blocker, La3+ (Sagor et al. 2015). Therefore, we determined if combined ethanol and LaCl3 pretreatments could affect ethanol-induced heat stress tolerance. Arabidopsis plants were grown in liquid MS medium for 2 weeks and then pretreated with ethanol and/or LaCl3 for 3 days prior to being subjected to a heat stress treatment. Results indicated that the fresh weight of plants pretreated with ethanol and LaCl3 was significantly lower than the fresh weight of plants pretreated with ethanol alone (Fig. 5A). A similar trend was also observed on the expression of BIP3 in response to the same treatments (Fig. 5B). We also analyzed the level of BIP3 expression in plants treated with ethanol and the polyamine biosynthetic inhibitors, DFMO and DFMA (Tiburcio et al. 1987). Results indicated that inhibiting polyamine biosynthesis completely suppressed ethanol-induced expression of BIP3 (Fig. 5C). These data suggest that BIP3 is involved in the ethanol-induced heat stress tolerance via the UPR signaling mediated by the increase in endogenous polyamines.
Effect of LaCl3, a Ca2+-channel blocker, on ethanol-induced heat stress tolerance. A Photos and fresh weight of Arabidopsis plants after plants grown on MS liquid media with LaCl3, ethanol, or both LaCl3 and ethanol were subjected to a heat stress treatment (45 °C for 2.5 h) and then placed under non-stressed condition for another seven days. Sixteen plants were used for each replicate. B RT-qPCR analysis of BIP3 expression in control (without ethanol and LaCl3), ethanol, ethanol and LaCl3, and LaCl3 treated plants. C RT-qPCR analysis of BIP3 expression in control (without ethanol, DFMO, or DFMA), ethanol, all three chemicals (ethanol, DFMO and DFMA), and both DFMO and DFMA treated plants. A–C Different letters indicate a significant difference (p < 0.05) between treatment groups (n = 3) as determined by a Tukey’s multiple comparison test
We also investigated the effect of the UPR inducers tunicamycin (Breckenridge et al. 2003) and azetidine-2-carboxylic acid (Rubenstein 2008) on ethanol-mediated heat stress tolerance. Arabidopsis plants grown on MS medium containing the UPR inducers had a higher rate of survival after a heat stress treatment compared to control plants (Fig. 6A). We then analyzed survival rates in the UPR signaling component mutants, bzip60-1 and bzip60-2, exposed to a heat stress treatment with and without an ethanol pretreatment. The survival rate of bzip60 mutants pretreated with ethanol and subjected to a heat stress treatment was about 20%, whereas survival rate was approximately 40% in wild-type plants pretreated with ethanol and subjected to the same heat stress treatment (Fig. 6B). These results indicate that the UPR signaling component bZIP60 plays a role in the mechanism responsible for ethanol-induced heat stress tolerance in Arabidopsis.
Involvement of UPR signaling in ethanol-induced heat stress tolerance. A Photos and survival rate of Arabidopsis plants pretreated with tunicamycin (Tm) or azetidine-2-carboxylic acid (AZC) and treated with heat stress. Seeds were sown on MS media containing 0.015 mg / L Tm or 15 μM AZC. After the cultivation for 14 days, seedlings were subjected to heat stress at 43.5 ˚C for 3 h and then placed under non-stressed condition for seven days. Asterisks indicate a significant difference (p < 0.05) compared to control plants as determined by a Tukey’s multiple comparison test (n = 4, 30 plants were used in each replicate). B Photos and survival rate of bzip60-1 (SAIL_838_C12) and bzip60-2 (SALK_050203). Arabidopsis plants after plants grown on MS media containing ethanol were subjected to a heat stress treatment (43.5 °C for 3 h) and then placed under non-stressed conditions for another 7 days. Red arrows in pictures indicate viable plants. Asterisks indicate a significant difference (p < 0.05) compared to untreated, control plants as determined by a t-test
Discussion
In the present study, we demonstrated that an ethanol pretreatment enhances heat stress tolerance in Arabidopsis and lettuce plants. Dosage concentrations in chemical priming are important for minimizing unfavorable effects, such as growth inhibition (Savvides et al. 2016). Therefore, we evaluated the level of heat stress tolerance and growth in plants pretreated with various concentrations of ethanol. Results indicated that pretreatments of ethanol at concentrations of 10 mM to 60 mM increased heat stress tolerance (Fig. 1). Importantly, however, pretreatments of ethanol at concentrations of 15–60 mM resulted in growth inhibition (Supplemental Fig. 2). These results indicate that a pretreatment of 10 mM ethanol can increase heat stress tolerance without suppressing plant growth in Arabidopsis. Plants reduce their growth in response to mild abiotic stress conditions and activate specific molecular and metabolic pathways that contribute to stress tolerance (Skirycz and Inzé 2010; Todaka et al. 2017; Urano et al. 2017). Therefore, studies utilizing chemical priming with ethanol will be useful for understanding the link between stress tolerance and growth regulation.
Our previous study demonstrated that an ethanol pretreatment enhances salt stress tolerance through its impact on the regulation of the antioxidant enzymes, APX1 and APX2, as well as their transcriptional regulators, ZAT12 and ZAT10 (Nguyen et al. 2017). The expression levels of these genes in response to the heat stress treatment used in the present study did not increase in ethanol-pretreated plants, relative to their level in ethanol-untreated control plants (Fig. 3). This might be due to different concentration of the ethanol used. In the previous study (Nguyen et al. 2017) for high-salinity stress tolerance, the ethanol concentration used was 51 mM. In the present study, the transcriptome analysis was done at the concentration of 2 mM. The results of the expression levels of these genes suggest that these genes are not involved in the mechanism responsible for ethanol-induced heat stress tolerance, although our data do not rule out the possibility that ethanol treatment changes the ROS levels under heat stress condition because the enzymatic activities have not yet been investigated. Instead, we propose that under heat stress conditions, UPR induction by an ethanol pretreatment has an important function in the enhancement of stress tolerance. This premise is supported by the finding that heat stress activates components of the UPR signaling network (Gao et al. 2008; Deng et al. 2011). In some situations, materials such as fallen leaves can be degraded by soil microorganisms. These microorganisms emit heat through respiration and produce alcohol (Kästner and Miltner 2016), thus, creating soil conditions with elevated temperature and the presence of alcohol. The elevated temperature can be reached to 45–70 °C (Kästner and Miltner 2016). This aspect of soil biology may be related to the induction of the UPR signaling network that occurs in the response to exogenous ethanol under heat stress conditions.
Our transcriptome analysis revealed a set of differentially expressed genes in ethanol-pretreated Arabidopsis plants (Supplemental Table 1). BIP3, a gene encoding an ER-resident cognate of heat shock protein 70, was among the identified, up-regulated genes, (Supplemental Table 1; Supplemental Fig. 2). BIP3 expression has been reported to be induced in response to ER-stress conditions and has been utilized as a UPR marker gene (Iwata and Koizumi 2012; Iwata et al. 2018). GC–MS analysis indicated that the level of polyamines, including putrescine and ornithine, increased in ethanol-pretreated plants (Supplemental Table 3). Polyamines have been reported to increase the expression of BIP3 (Sagor et al. 2015). Therefore, we hypothesized that BIP3 could function as one of the important components in the mechanisms underlying ethanol-induced heat stress tolerance. Notably, experiments using La3+, an inhibitor of polyamine-induced UPR excitation (Fig. 5A, B), polyamine biosynthetic inhibitors, such as DFMO and DFMA (Fig. 5C), UPR inducers, such as tunicamycin and azetidine-2-carboxylic acid (Fig. 6A), and mutants for UPR signaling components, such as bzip60 (Fig. 6B) further validated the involvement of UPR in ethanol-induced heat stress tolerance.
GO analysis using the set of up-regulated genes in plants pretreated with ethanol clarified that they were enriched in the GO terms of oxidation–reduction process (GO:0,055,114, p value = 0.0015) in the biological process category and oxidoreductase activity (GO:0,016,491, p value = 6.8e-05) in the molecular function category (Supplemental Fig. 4). Although further investigation is needed to elucidate whether the GO-term genes are involved in these designated regulatory mechanisms, changes in intracellular oxidation/reduction status or in the level of the relevant molecules may play an important role in ethanol-induced heat stress tolerance. FERRIC REDUCTASE OXIDASE5 (FRO5), one of the upregulated genes in our transcriptome analysis (Supplemental Table 1), has been shown to function in root-to-shoot Fe translocation and homeostasis of Cu and Fe (Bernal et al. 2012). Thus, there is a possibility that plants perceive the presence of ethanol in soils by changes in the inter/intracellular distribution of these metals. We also investigated the presence of cis-elements responsible for ER stress response in the promoter regions of the up-regulated genes identified in our transcriptome analysis, including ER stress-responsive element-I (ERSE-I, 5ʹ-CCAAT-N10-CACG-3ʹ) and unfolded protein response element-1 (UPRE, 5ʹ-TGACGT-GR-3ʹ) (Liu and Howell, 2010). These cis-elements were not observed in the promoter regions (data not shown), suggesting that other unknown cascades are involved in this regulatory mechanism.
Polyamines affect environmental stress response and growth in plants by their impact on various molecular processes, such as nucleic acid folding, translation activation, and membrane fluidity (Chen et al. 2019). In the present study, exogenous putrescine treatment increased heat stress tolerance in Arabidopsis plants. Consistent with this finding, it has been reported that exogenous putrescine treatment enhanced heat tolerance in tomato (Jahan et al. 2022) and in cauliflower (Collado-González et al. 2021). We investigated the expression profiles of the polyamine biosynthesis-related genes in the water- and ethanol-pretreated plants under heat stress condition because the polyamine contents were increased in the ethanol-pretreated plants (Supplemental Table 3). The results did not show the significant differences between the water- and ethanol-pretreated plants (data not shown). Although further analysis is necessary, the polyamine pathways associated with the ethanol-mediated heat tolerance might be regulated not by transcriptional regulatory mechanisms but by other regulatory mechanisms including enzymatic activity and protein stability. Previous papers have speculated that changes in the membrane by polyamines induce UPR through leakage in Ca2+ channels (Sagor et al. 2015). Polyamines in maize regulate the major transcription factor of UPR, bZIP60, which regulates the expression of heat shock factor at high temperatures (Li et al. 2020). Transgenic lines of Arabidopsis overexpressing bZIP60 exhibit enhanced salt stress tolerance along with the up-regulation of a number of stress-inducible genes (Fujita et al. 2007). In the present study, UPR and bZIP60 were found to be involved in ethanol induced heat tolerance. Our results suggest that specific bZIP proteins, including bZIP60, play an important role in the UPR activated by an ethanol pretreatment. Another UPR transcription factor, bZIP28, contributes to the upregulation of BiP2 and HSP26.5-P, in response to heat stress (Gao et al. 2008).
Although our study found that the activation of UPR plays an important role in ethanol-induced heat stress tolerance, the involvement of other factors in this process cannot be excluded. Liquid–liquid phase separation (LLPS) has been recently recognized as an important regulatory system in biological processes. LLPS has been shown to drive the formation of organelles not bound by membranes, such as the nucleolus, Cajal bodies, nuclear speckles, stress granules, and P-bodies (Fang et al. 2019; Jung et al. 2020; Henninger et al. 2021; Huang et al. 2021). LLPS has been reported to be associated with the physicochemical dynamics of aqueous co-solvent systems composed of small, polar organic molecules, including alcohols (Oh and Baiz 2020). These findings raise the possibility that the application of exogenous ethanol may affect LLPS in vivo and cause specific membraneless organellar compartmentation with changes in gene expression, leading to the enhancement of stress tolerance. Dorone et al. (2021) revealed that a prion-like protein named FLOE1 can function as a water potential sensor and regulate seed germination through a reversible hydration-dependent phase separation system in Arabidopsis. Although direct evidence is not yet available, similar mechanisms in the ethanol-pretreated plants in the present study may be regulated by prion-like or other uncharacterized proteins. Our transcriptome analysis identified a set of up-regulated genes in ethanol-pretreated plants (Supplemental Table 1). Regarding the proteins encoded by these genes, we utilized the python-based intrinsic disorder prediction application MobiDB-lite (Necci et al. 2017) to determine if these proteins have an intrinsically disordered (ID) region, which has been recognized as a factor driving phase separation (Nott et al. 2015; Shin and Brangwynne 2017). The analysis revealed that SUC6 (1 to 26 amino acid sequence), BIP3 (656 to 675 amino acid sequence), and FRO5 (558 to 580 amino acid sequence) have an ID region, suggesting that these proteins might function as ID proteins in cells of ethanol-pretreated Arabidopsis plants. Further analysis is necessary to reveal any relationship of LLPS with these proteins.
Several studies have shown that external ethanol application has physiological effects on plants. Ethanol treatment changes the localization of PIN1 by weakening the tension of the cell membrane and enhancing endocytosis (Ackermann and Stanislas 2020). The changes of membrane fluidity activate the specific MAPK signaling under cold and heat stress conditions (Sangwan et al. 2002). Ethanol and acetaldehyde are also known to be involved in the process of flower aging (Podd and Van Staden 1998). These studies imply the existence of ethanol-mediated signaling pathways under heat stress condition.
In conclusion, we found that ethanol-pretreatment increased heat stress tolerance in Arabidopsis and lettuce plants. Ethanol-pretreated plants exhibited a lower level of ion leakage, relative to water-pretreated, control plants. Our data suggest that plants grown on the ethanol-containing MS plates are primed for increased heat stress tolerance under normal conditions and is activated when plants are subjected to heat stress. Transcriptome and metabolome analyses raised a possibility that activated UPR contributed to enhanced heat stress tolerance in ethanol-pretreated plants. Evidence obtained by the experiments using UPR inducers, UPR inhibitors, and UPR-related mutants supported the contribution of UPR to the enhancement of heat stress tolerance in the ethanol-pretreated plants. Chemical priming technology has significant potential to improve various important agronomic traits (Savvides et al. 2016; Sako et al. 2021b).Therefore, chemical priming using ethanol represents an attractive strategy for developing elite plants, such as “climate-smart”crops. Our findings has provided a molecular framework for understanding the enhancement of heat stress tolerance by ethanol priming.
Data availability
Enquiries about data availability should be directed to the authors.
References
Ackermann F, Stanislas T (2020) The plasma membrane: an integrating compartment for mechano-signaling. Plants. https://doi.org/10.3390/plants9040505
Ahmad Z, Bashir K, Matsui A, Tanaka M, Sasaki R, Oikawa A, Hirai MY, Chaomurilege ZuY, Kawai-Yamada M, Rashid B, Husnain T, Seki M (2021) Overexpression of nicotinamidase 3 (NIC3) gene and the exogenous application of nicotinic acid (NA) enhance drought tolerance and increase biomass in Arabidopsis. Plant Mol Biol 107:63–84. https://doi.org/10.1007/s11103-021-01179-z
AmbreenAtharUrKhanZafarAyyazKalaji SHRAZUAHM (2021) Seed priming with proline improved photosystem II efficiency and growth of wheat (Triticum aestivum L.). BMC Plant Biol 21:1–12. https://doi.org/10.1186/s12870-021-03273-2
Angelos E, Ruberti C, Kim SJ, Brandizzi F (2017) Maintaining the factory: the roles of the unfolded protein response in cellular homeostasis in plants. Plant J 90:671–682. https://doi.org/10.1111/tpj.13449
Bernal M, Casero D, Singh V, Wilson GT, Grande A, Yang H, Dodani SC, Pellegrini M, Huijser P, Connolly EL, Merchant SS, Krämer U (2012) Transcriptome sequencing identifies SPL7-regulated copper acquisition genes FRO4/FRO5 and the copper dependence of iron homeostasis in Arabidopsis. Plant Cell 24:738–761. https://doi.org/10.1105/tpc.111.090431
Breckenridge DG, Germain M, Mathai JP, Nguyen M, Shore GC (2003) Regulation of apoptosis by endoplasmic reticulum pathways. Oncogene 22:8608–8618. https://doi.org/10.1038/sj.onc.1207108
Chen D, Shao Q, Yin L, Younis A, Zheng B (2019) Polyamine function in plants: metabolism, regulation on development, and roles in abiotic stress responses. Front Plant Sci 9:1–13. https://doi.org/10.3389/fpls.2018.01945
Choudhury FK, Rivero RM, Blumwald E, Mittler R (2017) Reactive oxygen species, abiotic stress and stress combination. Plant J 90:856–867. https://doi.org/10.1111/tpj.13299
Collado-González J, Piñero MC, Otálora G, López-Marín J, Del Amor FM (2021) Effects of different nitrogen forms and exogenous application of putrescine on heat stress of cauliflower: photosynthetic gas exchange, mineral concentration and lipid peroxidation. Plants 10:1–19. https://doi.org/10.3390/plants10010152
Del Carratore F, Jankevics A, Eisinga R, Heskes T, Hong F, Breitling R (2017) RankProd 2.0: a refactored bioconductor package for detecting differentially expressed features in molecular profiling datasets. Bioinformatics 33:2774–2775. https://doi.org/10.1093/bioinformatics/btx292
Deng Y, Humbert S, Liu JX, Srivastava R, Rothstein SJ, Howell SH (2011) Heat induces the splicing by IRE1 of a mRNA encoding a transcription factor involved in the unfolded protein response in Arabidopsis. Proc Natl Acad Sci USA 108:7247–7252. https://doi.org/10.1073/pnas.1102117108
Dong Y, Xu L, Wang Q, Fan Z, Kong J, Bai X (2014) Effects of exogenous nitric oxide on photosynthesis, antioxidative ability, and mineral element contents of perennial ryegrass under copper stress. J Plant Interact 9:402–411. https://doi.org/10.1080/17429145.2013.845917
Dorone Y, Boeynaems S, Flores E, Jin B, Hateley S, Bossi F, Lazarus E, Pennington JG, Michiels E, De Decker M, Vints K, Baatsen P, Bassel GW, Otegui MS, Holehouse AS, Exposito-Alonso M, Sukenik S, Gitler AD, Rhee SY (2021) A prion-like protein regulator of seed germination undergoes hydration-dependent phase separation. Cell 184:4284-4298.e27. https://doi.org/10.1016/j.cell.2021.06.009
Fang X, Wang L, Ishikawa R, Li Y, Fiedler M, Liu F, Calder G, Rowan B, Weigel D, Li P, Dean C (2019) Arabidopsis FLL2 promotes liquid–liquid phase separation of polyadenylation complexes. Nature 569:265–269. https://doi.org/10.1038/s41586-019-1165-8
Fujita M, Mizukado S, Fujita Y, Ichikawa T, Nakazawa M, Seki M, Matsui M, Yamaguchi-Shinozaki K, Shinozaki K (2007) Identification of stress-tolerance-related transcription-factor genes via mini-scale full-length cDNA Over-eXpressor (FOX) gene hunting system. Biochem Biophys Res Commun 364:250–257. https://doi.org/10.1016/j.bbrc.2007.09.124
Gao H, Brandizzi F, Benning C, Larkin RM (2008) A membrane-tethered transcription factor defines a branch of the heat stress response in Arabidopsis thaliana. Proc Natl Acad Sci USA 105:16397–16403. https://doi.org/10.1073/pnas.0808463105
Ha CV, Leyva-Gonzalez MA, Osakabe Y, Tran UT, Nishiyama R, Watanabe Y, Tanaka M, Seki M, Yamaguchi S, Van DN, Yamaguchi-Shinozaki K, Shinozaki K, Herrera-Estrella L, Tran LSP (2014) Positive regulatory role of strigolactone in plant responses to drought and salt stress. Proc Natl Acad Sci USA 111:851–856. https://doi.org/10.1073/pnas.1322135111
Hameed A, Farooq T, Hameed A, Sheikh MA (2021) Sodium nitroprusside mediated priming memory invokes water-deficit stress acclimation in wheat plants through physio-biochemical alterations. Plant Physiol Biochem 160:329–340. https://doi.org/10.1016/j.plaphy.2021.01.037
Henninger JE, Oksuz O, Shrinivas K, Sagi I, LeRoy G, Zheng MM, Andrews JO, Zamudio AV, Lazaris C, Hannett NM, Lee TI, Sharp PA, Cissé II, Chakraborty AK, Young RA (2021) RNA-mediated feedback control of transcriptional condensates. Cell 184:207-225.e24. https://doi.org/10.1016/j.cell.2020.11.030
Huang S, Zhu S, Kumar P, MacMicking JD (2021) A phase-separated nuclear GBPL circuit controls immunity in plants. Nature 594:424–429. https://doi.org/10.1038/s41586-021-03572-6
Humbert S, Zhong S, Deng Y, Howell SH, Rothstein SJ (2012) Alteration of the bZIP60/IRE1 pathway affects plant response to ER stress in Arabidopsis Thaliana. PLoS ONE 7:1–8. https://doi.org/10.1371/journal.pone.0039023
Ilík P, Špundová M, Šicner M, Melkovičová H, Kučerová Z, Krchňák P, Fürst T, Večeřová K, Panzarová K, Benediktyová Z, Trtílek M (2018) Estimating heat tolerance of plants by ion leakage: a new method based on gradual heating. New Phytol 218:1278–1287. https://doi.org/10.1111/nph.15097
Islam MJ, Ryu BR, Azad MOK, Rahman MH, Rana MS, Lim JD, Lim YS (2021) Exogenous putrescine enhances salt tolerance and ginsenosides content in Korean ginseng (Panax ginseng meyer) sprouts. Plants. https://doi.org/10.3390/plants10071313
Iwata Y, Koizumi N (2012) Plant transducers of the endoplasmic reticulum unfolded protein response. Trends Plant Sci 17:720–727. https://doi.org/10.1016/j.tplants.2012.06.014
Iwata Y, Yoneda M, Yanagawa Y, Koizumi N (2009) Characteristics of the nuclear form of the Arabidopsis transcription factor AtbZIP60 during the endoplasmic reticulum stress response. Biosci Biotechnol Biochem 73:865–869. https://doi.org/10.1271/bbb.80779
Iwata Y, Iida T, Matsunami T, Yamada Y, Mishiba KI, Ogawa T, Kurata T, Koizumi N (2018) Constitutive BiP protein accumulation in Arabidopsis mutants defective in a gene encoding chloroplast-resident stearoyl-acyl carrier protein desaturase. Genes Cells 23:456–465. https://doi.org/10.1111/gtc.12585
Jägermeyr J, Müller C, Ruane AC, Elliott J, Balkovic J, Castillo O, Faye B, Foster I, Folberth C, Franke JA, Fuchs K, Guarin JR, Heinke J, Hoogenboom G, Iizumi T, Jain AK, Kelly D, Khabarov N, Lange S, Lin TS, Liu W, Mialyk O, Minoli S, Moyer EJ, Okada M, Phillips M, Porter C, Rabin SS, Scheer C, Schneider JM, Schyns JF, Skalsky R, Smerald A, Stella T, Stephens H, Webber H, Zabel F, Rosenzweig C (2021) Climate impacts on global agriculture emerge earlier in new generation of climate and crop models. Nat Food 2:873–885. https://doi.org/10.1038/s43016-021-00400-y
Jahan M, Hasan M, Alotaibi F, Alabdallah N, Alharbi B, Ramadan K, Bendary E, Alshehri D, Jabborova D, Al-Balawi D, Dessoky E, Ibrahim M, Guo S (2022) Exogenous putrescine increases heat tolerance in tomato seedlings by regulating chlorophyll metabolism. Plants 11:1038. https://doi.org/10.3390/plants11081038
Jonsson P, Johansson AI, Gullberg J, Trygg J, A J, Grung B, Marklund S, Sjöström M, Antti H, Moritz T, (2005) High-throughput data analysis for detecting and identifying differences between samples in GC/MS-based metabolomic analyses. Anal Chem 77:5635–5642. https://doi.org/10.1021/ac050601e
Jung JH, Barbosa AD, Hutin S, Kumita JR, Gao M, Derwort D, Silva CS, Lai X, Pierre E, Geng F, Kim SB, Baek S, Zubieta C, Jaeger KE, Wigge PA (2020) A prion-like domain in ELF3 functions as a thermosensor in Arabidopsis. Nature 585:256–260. https://doi.org/10.1038/s41586-020-2644-7
Kamauchi S, Nakatani H, Nakano C, Urade R (2005) Gene expression in response to endoplasmic reticulum stress in Arabidopsis thaliana. FEBS J 272:3461–3476. https://doi.org/10.1111/j.1742-4658.2005.04770.x
Kästner M, Miltner A (2016) Application of compost for effective bioremediation of organic contaminants and pollutants in soil. Appl Microbiol Biotechnol 100:3433–3449. https://doi.org/10.1007/s00253-016-7378-y
Khan MIR, Fatma M, Per TS, Anjum NA, Khan NA (2015) Salicylic acid-induced abiotic stress tolerance and underlying mechanisms in plants. Front Plant Sci 6:1–17. https://doi.org/10.3389/fpls.2015.00462
Kikuchi J, Hirayama T (2007) Practical aspects of uniform stable isotope labeling of higher plants for heteronuclear NMR-based metabolomics. Methods Mol Biol 358:273–286. https://doi.org/10.1007/978-1-59745-244-1_15
Kikuchi J, Shinozaki K, Hirayama T (2004) Stable isotope labeling of Arabidopsis thaliana for an NMR-based metabolomics approach. Plant Cell Physiol 45:1099–1104. https://doi.org/10.1093/pcp/pch117
Kim JM, To TK, Matsui A, Tanoi K, Kobayashi NI, Matsuda F, Habu Y, Ogawa D, Sakamoto T, Matsunaga S, Bashir K, Rasheed S, Ando M, Takeda H, Kawaura K, Kusano M, Fukushima A, Endo TA, Kuromori T, Ishida J, Morosawa T, Tanaka M, Torii C, Takebayashi Y, Sakakibara H, Ogihara Y, Saito K, Shinozaki K, Devoto A, Seki M (2017) Acetate-mediated novel survival strategy against drought in plants. Nat Plants 3:17097. https://doi.org/10.1038/nplants.2017.97
Komatsu T, Ohishi R, Shino A, Akashi K, Kikuchi J (2014) Multi-spectroscopic analysis of seed quality and 13C-stable-iotopologue monitoring in initial growth metabolism of Jatropha curcas L. Metabolites 4:1018–1033. https://doi.org/10.3390/metabo4041018
Komatsu T, Ohishi R, Shino A, Kikuchi J (2016) Structure and metabolic-flow analysis of molecular complexity in a 13C-labeled tree by 2D and 3D NMR. Angew Chemie 55:6000–6003. https://doi.org/10.1002/anie.201600334
Kusano M, Fukushima A, Arita M, Jonsson P, Moritz T, Kobayashi M, Hayashi N, Tohge T, Saito K (2007a) Unbiased characterization of genotype-dependent metabolic regulations by metabolomic approach in Arabidopsis thaliana. BMC Syst Biol 1:1–17. https://doi.org/10.1186/1752-0509-1-53
Kusano M, Fukushima A, Kobayashi M, Hayashi N, Jonsson P, Moritz T, Ebana K, Saito K (2007b) Application of a metabolomic method combining one-dimensional and two-dimensional gas chromatography-time-of-flight/mass spectrometry to metabolic phenotyping of natural variants in rice. J Chromatogr B 855:71–79. https://doi.org/10.1016/j.jchromb.2007.05.002
Li T, Hu Y, Du X, Tang H, Shen C, Wu J (2014) Salicylic acid alleviates the adverse effects of salt stress in Torreya grandis cv. merrillii seedlings by activating photosynthesis and enhancing antioxidant systems. PLoS ONE 9:1–9. https://doi.org/10.1371/journal.pone.0109492
Li Z, Tang J, Srivastava R, Bassham DC, Howell SH (2020) The transcription factor bZIP60 links the unfolded protein response to the heat stress response in maize. Plant Cell 32:3559–3575. https://doi.org/10.1105/TPC.20.00260
Liu H, Wang J, Liu J, Liu T, Xue S (2021) Hydrogen sulfide (H2S) signaling in plant development and stress responses. aBIOTECH 2:32–63. https://doi.org/10.1007/s42994-021-00035-4
Liu JX, Howell SH (2010) bZIP28 and NF-Y transcription factors are activated by ER stress and assemble into a transcriptional complex to regulate stress response genes in Arabidopsis. Plant Cell 22:782–796. https://doi.org/10.1105/tpc.109.072173
Mageroy MH, Wilkinson SW, Tengs T, Cross H, Almvik M, Pétriacq P, Vivian-Smith A, Zhao T, Fossdal CG, Krokene P (2020) Molecular underpinnings of methyl jasmonate-induced resistance in Norway spruce. Plant Cell Environ 43:1827–1843. https://doi.org/10.1111/pce.13774
Martínez IM, Chrispeels MJ (2003) Genomic analysis of the unfolded protein response in Arabidopsis shows its connection to important cellular processes. Plant Cell 15:561–576. https://doi.org/10.1105/tpc.007609
Matsui A, Iida K, Tanaka M, Yamaguchi K, Mizuhashi K, Kim JM, Takahashi S, Kobayashi N, Shigenobu S, Shinozaki K, Seki M (2017) Novel stress-inducible antisense RNAs of protein-coding loci are synthesized by RNA-dependent RNA polymerase. Plant Physiol 175:457–472. https://doi.org/10.1104/pp.17.00787
Mustafavi SH, Naghdi Badi H, Sękara A, Mehrafarin A, Janda T, Ghorbanpour M, Rafiee H (2018) Polyamines and their possible mechanisms involved in plant physiological processes and elicitation of secondary metabolites. Acta Physiol Plant 40:1–19. https://doi.org/10.1007/s11738-018-2671-2
Nagashima Y, Mishiba KI, Suzuki E, Shimada Y, Iwata Y, Koizumi N (2011) Arabidopsis IRE1 catalyses unconventional splicing of bZIP60 mRNA to produce the active transcription factor. Sci Rep. https://doi.org/10.1038/srep00029
Necci M, Piovesan D, Dosztanyi Z, Tosatto SCE (2017) MobiDB-lite: fast and highly specific consensus prediction of intrinsic disorder in proteins. Bioinformatics 33:1402–1404. https://doi.org/10.1093/bioinformatics/btx015
Nguyen AH, Matsui A, Tanaka M, Mizunashi K, Nakaminami K, Hayashi M, Iida K, Toyoda T, Van ND, Seki M (2015) Loss of Arabidopsis 5′-3′ exoribonuclease AtXRN4 function enhances heat stress tolerance of plants subjected to severe heat stress. Plant Cell Physiol 56:1762–1772. https://doi.org/10.1093/pcp/pcv096
Nguyen HM, Sako K, Matsui A, Suzuki Y, Mostofa MG, Van Ha C, Tanaka M, Tran LSP, Habu Y, Seki M (2017) Ethanol enhances high-salinity stress tolerance by detoxifying reactive oxygen species in Arabidopsis thaliana and rice. Front Plant Sci 8:1–10. https://doi.org/10.3389/fpls.2017.01001
Nishad A, Nandi AK (2021) Recent advances in plant thermomemory. Plant Cell Rep 40:19–27. https://doi.org/10.1007/s00299-020-02604-1
Nott TJ, Petsalaki E, Farber P, Jervis D, Fussner E, Plochowietz A, Craggs TD, Bazett-Jones DP, Pawson T, Forman-Kay JD, Baldwin AJ (2015) Phase transition of a disordered nuage protein generates environmentally responsive membraneless organelles. Mol Cell 57:936–947. https://doi.org/10.1016/j.molcel.2015.01.013
Oh KI, Baiz CR (2020) Molecular heterogeneity in aqueous cosolvent systems. J Chem Phys 152:1–7. https://doi.org/10.1063/5.0007647
Ohama N, Sato H, Shinozaki K, Yamaguchi-Shinozaki K (2017) Transcriptional regulatory network of plant heat stress response. Trends Plant Sci 22:53–65. https://doi.org/10.1016/j.tplants.2016.08.015
Omoarelojie LO, Kulkarni MG, Finnie JF, Pospíšil T, Strnad M, Van Staden J (2020) Synthetic strigolactone (rac-GR24) alleviates the adverse effects of heat stress on seed germination and photosystem II function in lupine seedlings. Plant Physiol Biochem 155:965–979. https://doi.org/10.1016/j.plaphy.2020.07.043
Pastor-Cantizano N, Ko DK, Angelos E, Pu Y, Brandizzi F (2020) Functional diversification of ER stress responses in Arabidopsis. Trends Biochem Sci 45:123–136. https://doi.org/10.1016/j.tibs.2019.10.008
Podd LA, Van Staden J (1998) The role of ethanol and acetaldehyde in flower senescence and fruit ripening : a review. Plant Growth Regul 26:183–189. https://doi.org/10.1023/A:1006131517539
Qamar R, Khan S, Safdar ME, Atique-ur-Rehman RA, Javeed HMR, Nadeem MA, Al-Yahyai R, Alkahtani J (2022) Seed priming with growth regulators modulates production, physiology and antioxidant defense of Indian squash (Praecitrullus fistulosus) under semi-arid conditions. PLoS ONE 17:e0265694. https://doi.org/10.1371/journal.pone.0265694
Redestig H, Fukushima A, Stenlund H, Moritz T, Arita M, Saito K, Kusano M (2009) Compensation for systematic cross-contribution improves normalization of mass spectrometry based metabolomics data. Anal Chem 81:7974–7980. https://doi.org/10.1021/ac901143w
Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W, Smyth GK (2015) Limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res 43:e47. https://doi.org/10.1093/nar/gkv007
Rizhsky L, Davletova S, Liang H, Mittler R (2004) The zinc finger protein Zat12 Is required for cytosolic ascorbate peroxidase 1 expression during oxidative stress in Arabidopsis. J Biol Chem 279:11736–11743. https://doi.org/10.1074/jbc.M313350200
Rubenstein E (2008) Misincorporation of the proline analog azetidine-2-carboxylic acid in the pathogenesis of multiple sclerosis: a hypothesis. J Neuropathol Exp Neurol 67:1035–1040. https://doi.org/10.1097/NEN.0b013e31818add4a
Sagor GHM, Chawia P, Kim DW, Berberich T, Kojima S, Niitsu M, Kusano T (2015) The polyamine spermine induces the unfolded protein response via the MAPK cascade in Arabidopsis. Front Plant Sci 6:1–11. https://doi.org/10.3389/fpls.2015.00687
Sako K, Nagashima R, Tamoi M, Seki M (2021a) Exogenous ethanol treatment alleviates oxidative damage of Arabidopsis thaliana under conditions of high-light stress. Plant Biotechnol 38:339–344. https://doi.org/10.5511/plantbiotechnology.21.0715a
Sako K, Nguyen HM, Seki M (2021b) Advances in chemical priming to enhance abiotic stress tolerance in plants. Plant Cell Physiol 61:1995–2003. https://doi.org/10.1093/pcp/pcaa119
Sangwan V, Örvar BL, Beyerly J, Hirt H, Dhindsa Rajinder S (2002) Opposite changes in membrane fluidity mimic cold and heat stress activation of distinct plant MAP kinase pathways. Plant J 31:629–638. https://doi.org/10.1046/j.1365-313X.2002.01384.x
Savvides A, Ali S, Tester M, Fotopoulos V (2016) Chemical priming of plants against multiple abiotic stresses: mission possible? Trends Plant Sci 21:329–340. https://doi.org/10.1016/j.tplants.2015.11.003
Sekiyama Y, Chikayama E, Kikuchi J (2010) Profiling polar and semipolar plant metabolites throughout extraction processes using a combined solution-state and high-resolution magic angle spinning NMR approach. Anal Chem 82:1643–1652. https://doi.org/10.1021/ac9019076
Shin Y, Brangwynne CP (2017) Liquid phase condensation in cell physiology and disease. Science. https://doi.org/10.1126/science.aaf4382
Shu S, Yuan Y, Chen J, Sun J, Zhang W, Tang Y, Zhong M, Guo S (2015) The role of putrescine in the regulation of proteins and fatty acids of thylakoid membranes under salt stress. Sci Rep 5:1–16. https://doi.org/10.1038/srep14390
Skirycz A, Inzé D (2010) More from less: plant growth under limited water. Curr Opin Biotechnol 21:197–203. https://doi.org/10.1016/j.copbio.2010.03.002
Tian C, Chikayama E, Tsuboi Y, Kuromori T, Shinozaki K, Kikuchi J, Hirayama T (2007) Top-down phenomics of Arabidopsis thaliana: metabolic profiling by one- and two-dimensional nuclear magnetic resonance spectroscopy and transcriptome analysis of albino mutants. J Biol Chem 282:18532–18541. https://doi.org/10.1074/jbc.M700549200
Tiburcio AF, Kaur-Sawhney R, Galston AW (1987) Effect of polyamine biosynthetic inhibitors on alkaloids and organogenesis in tobacco callus cultures. Plant Cell Tissue Organ Cult 9:111–120. https://doi.org/10.1007/BF00044246
Todaka D, Zhao Y, Yoshida T, Kudo M, Kidokoro S, Mizoi J, Kodaira KS, Takebayashi Y, Kojima M, Sakakibara H, Toyooka K, Sato M, Fernie AR, Shinozaki K, Yamaguchi-Shinozaki K (2017) Temporal and spatial changes in gene expression, metabolite accumulation and phytohormone content in rice seedlings grown under drought stress conditions. Plant J 90:61–78. https://doi.org/10.1111/tpj.13468
Urano K, Maruyama K, Jikumaru Y, Kamiya Y, Yamaguchi-Shinozaki K, Shinozaki K (2017) Analysis of plant hormone profiles in response to moderate dehydration stress. Plant J 90:17–36. https://doi.org/10.1111/tpj.13460
Utsumi Y, Utsumi C, Tanaka M, Van HC, Takahashi S, Matsui A, Matsunaga TM, Matsunaga S, Kanno Y, Seo M, Okamoto Y, Moriya E, Seki M (2019) Acetic acid treatment enhances drought avoidance in cassava (Manihot esculenta crantz). Front Plant Sci 10:1–12. https://doi.org/10.3389/fpls.2019.00521
Valenzuela-Riffo F, Zúñiga P, Morales-Quintana L, Lolas M, Cáceres M, Figueroa C (2020) Priming of defense systems and upregulation of MYC2 and JAZ1 genes after botrytis cinerea inoculation in methyl jasmonate-treated strawberry fruits. Plants 9:447. https://doi.org/10.3390/plants9040447
Walter P, Ron D (2011) The unfolded protein response: from stress pathway to homeostatic regulation. Science 334:1081–1086
Waqas Mazhar M, Ishtiaq M, Hussain I, Parveen A, Hayat Bhatti K, Azeem M, Thind S, Ajaib M, Maqbool M, Sardar T, Muzammil K, Nasir N (2022) Seed nano-priming with Zinc Oxide nanoparticles in rice mitigates drought and enhances agronomic profile. PLoS ONE 17:e0264967. https://doi.org/10.1371/journal.pone.0264967
Acknowledgements
The method of Arabidopsis cultivation in MS liquid medium was advised by Dr. Takatoshi Kiba at Nagoya University. We thank Imato Takeuchi from Kihara Institute for Biological Research, Yokohama City University for the technical assistance of field experiments. We also thank Prof. Nozomu Koizumi at Osaka Prefecture University, for providing the mutant seeds (SALK_050203). We thank Journal Doctors (http://www.journaldoctors.com/about.html) for editing a draft of this manuscript.
Funding
This work was supported by grants from RIKEN, Japan (to MS), such as RIKEN–AIST Joint Research Fund (Full research), Core Research for Evolutionary Science and Technology (JPMJCR13B4 to MS) and A-STEP (JPMJTM19BS to MS) of the Japan Science and Technology Agency (JST), and Grants-in-Aid for Scientific Research from the Ministry of Education, Culture, Sports, Science and Technology (MEXT), Japan (Innovative Areas 18H04791 and 18H04705 to MS).
Author information
Authors and Affiliations
Contributions
The heat stress experiments of ethanol-treated plants and transcriptome analysis were performed by AM, YS, KM, ST, MT, KK, JI and MS. NMR analysis was performed by KB, YT and JK. GC/MS analysis was performed by KB, MK and MK. The manuscript was written by AM, DT and MS. The total experiments of this study were designed by AM and MS.
Corresponding author
Ethics declarations
Conflict of interest
The authors have not disclosed any competing interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Matsui, A., Todaka, D., Tanaka, M. et al. Ethanol induces heat tolerance in plants by stimulating unfolded protein response. Plant Mol Biol 110, 131–145 (2022). https://doi.org/10.1007/s11103-022-01291-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-022-01291-8