Abstract
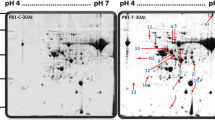
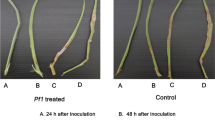
The legume root rot disease caused by the oomycete pathogen Aphanomyces euteiches is one major yield reducing factor in legume crop production. A comparative proteomic approach was carried out in order to identify proteins of the model legume Medicago truncatula which are regulated after an infection with A. euteiches. Several proteins were identified by two dimensional gel electrophoresis to be differentially expressed after pathogen challenge. Densitometric evaluation of expression values showed different regulation during the time-course analysed. Proteins regulated during the infection were identified by matrix assisted laser desorption/ionization-time of flight-mass spectrometry (MALDI-TOF-MS). Among the differentially expressed proteins, two encoded putative cell wall proteins and two were designated as small heat shock proteins. Furthermore, an isoform of the chalcone-O-methyltransferase was found to be increased in infected roots. The majority of induced proteins belonged to the family of class 10 of pathogenesis related proteins (PR10). Previously, various PR10-like proteins have been shown to be regulated by general stress or abscisic acid (ABA). Therefore, these proteins were further investigated concerning their regulation in response to drought stress and exogenous ABA-application. Complex regulation patterns were identified: three of the A. euteiches-induced PR10-like proteins were also induced by exogenous ABA- but none of them is induced after drought stress. In contrast, three of these proteins are down-regulated by drought stress. Hence, the strong expression of different PR10-family members and their regulation profiles indicates that this set of proteins plays a major role during root adaptations to various stress conditions.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Audenaert, K., De Meyer, G.B. and Höfte, M.M. 2002. Abscisic acid determines basal susceptibility of tomato to Botrytis cinerea and suppresses salicylic acid-dependant signaling mechanisms. Plant Physiol. 128: 491–501.
Bestel-Corre, G., Dumas-Gaudot, E., Poinsot, V., Dieu, M., Dierick, J.F., van, T. D., Remacle, J., Gianinazzi-Pearson, V. and Gianinazzi, S. 2002. Proteome analysis and identification of symbiosis-related proteins from Medicago truncatula Gaertn. by two-dimensional electrophoresis and mass spectrometry. Electrophoresis 23: 122–137.
Djordjevic, M.A., Chen, H.C., Natera, S., Van Noorden, G., Menzel, C., Taylor S., Renard, C., Geiger, O. and Weiler, G.F. 2003. A global analysis of protein expression profiles in Sinorhizobium meliloti: Discovery of new genes for nodule occupancy and stress adaptation. Mol. Plant Microbe Interact. 16: 508–525.
Engqvist, L.G. and Ahvenniemi, P. 1997. Interactions between common root rot (Aphanomyces euteiches) and peas (Pisum sativum) in Finland. Acta Agric. Scand. Sec. B-Soil Plant Sci. 4: 242–247.
Fedorova, M., Van de Mortel, J., Matsumoto, P.A., Cho, J., Town, C.D., VandenBosch, K.A., Gantt, J.S. and Vance, C.P. 2002. Genome-wide identification of nodule-specific transcripts in the model legume Medicago truncatula. Plant Physiol. 130: 519–537.
Grau, C.R. 1990. Aphanomyces root rot. In: D.L. Stuteville and D.C. Erwin (Eds.), Compendium of Alfalfa Diseases, American Phytopathological Society Press, St. Paul, MN.
Gygi, S.P., Rochon, Y., Franza, B.R. and Aebersold, R. 1999. Correlation between protein and mRNA abundance in yeast. Mol. Cell Biol. 19: 1720–1730.
Györgyey, J., Gartner, A., Németh, K., Magyar, Z., Hirt, H., Heberle-Bors, E. and Dudits, D. 1991. Alfalfa heat shock genes are differentially expressed during somatic embryogenesis. Plant Mol. Biol. 16: 999–1007.
Hagedorn, D.J. 1989. Aphanomyces root rot. In: Compendium of Pea Diseases, American Phytopathological Society Press, St. Paul, MN.
Handberg, K. and Stougaard, J. 1992. Lotus japonicus, an autogamous, diploid legume species for classical and molecular genetics. Plant J. 2: 487–496.
Hoagland, D.R. and Arnon, D.I. 1950. The water-culture method for growing plants without soil. Calif. Agric. Exp. Stu. Circ. 347: 1–39.
Hurkman, W.J. and Tanaka, C.K. 1986. Solubilization of plant membrane proteins for analysis by two-dimensional gel electrophoresis. Plant Physiol. 81: 802–806.
Iturriaga, E.A., Leech, M.J., Barratt, D.H. and Wang, T.L. 1994. Two ABA-responsive proteins from pea (Pisum sativum L.) are closely related to intracellular pathogenesis-related proteins. Plant Mol. Biol. 24: 235–240.
Journet, E.P., van Tuinen, D., Gouzy, J., Crespeau, H., Carreau, V., Farmer, M.J., Niebel, A., Schiex, T., Jaillon, O., Chatagnier, O., Godiard, L., Micheli, F., Kahn, D., Gianinazzi-Pearson, V. and Gamas, P. 2002. Exploring root symbiotic programs in the model legume Medicago truncatula using EST analysis. Nucleic Acids Res. 30: 5579–5592.
Keller, B., Sauer, N. and Lamb, C.J. 1988. Glycine-rich cell wall proteins in bean: Gene structure and association of the protein with the vascular system. EMBO. J. 7: 3625–3633.
Koller, A., Washburn, M.P., Lange, B.M., Andon, N.L., Deciu, C., Haynes, P.A., Hays, L., Schieltz, D., Ulaszek, R., Wei, J., Wolters, D. and Yates III, J.R. 2002. Proteomic survey of metabolic pathways in rice. Proc. Natl. Acad. Sci. 99: 11969–11974.
Kjoller, R. and Rosendahl, S. 1998. Enzymatic activity of the mycelium compared with oospore development during infection of pea roots by Aphanomyces euteiches. Phytopathology 88: 992–996.
Luo, M., Liu, J.H., Mohapatra, S., Hill, R.D. and Mohapatra, S.S. 1992. Characterization of a gene family encoding abscisic acid-and environmental stress-inducible proteins of alfalfa. J. Biol. Chem. 267: 15367–15374.
Mathesius, U., Keijzers, G., Natera, S.H., Weinman, J.J., Djordjevic, M.A. and Rolfe, B.G. 2001. Establishment of a root proteome reference map for the model legume Medicago truncatula using the expressed sequence tag database for peptide mass fingerprinting. Proteomics 1: 1424–1440.
Mitchel, J.E. and Yang, C.Y. 1966. Factor affecting growth and development of Aphanomyces euteiches. Phytopathology 56: 917–922.
Nyamsuren, O., Colditz, F., Rosendahl, S., Tamasloukht, M., Bekel, T., Meyer, F., Küster, H., Franken, P. and Krajinski, F. 2003. Transcriptional profiling of Medicago truncatula roots after infection with Aphanomyces euteiches (oomycota) identifies novel genes upregulated during this pathogenic interaction. Physiol. Mol. Plant Pathol. 63: 17–26.
Rock, C. D. 2000. Pathways to abscisic acid-regulated gene expression. New Phytol. 148: 357–396.
Timothy, J.C., Kortt, A.A. and Chandler, P.M. 1989. A cDNAbased comparison of dehydration-induced proteins (dehydrins) in barley and corn. Plant Mol. Biol. 13: 95–108.
Van Loon, L.C. and Van Strien, E.A. 1999. The families of pathogenesis-related proteins, their activities, and comparative analyses of PR-1 type proteins. Physiol. Mol. Plant Pathol. 55: 85–97.
Watson, B.S., Asirvatham, V.S., Wang, L. and Sumner, L.W. 2003. Mapping the proteome of barrel medic (Medicago truncatula). Plant Physiol. 131: 1104–1123.
Williams, K.R., LoPresti, M. and Stone, K. 1997. Internal protein sequencing of SDS-PAGE-separated proteins: Optimization of an in gel digest protocol. In: D. Marshak (Ed.), Techiques in Protein Chemistry, Academic Press, pp. 97–190.
Wulf, A., Manthey, K., Doll, J., Perlick, A.M., Linke, B., Bekel, T., Meyer, F., Franken, P., Küster, H. and Krajinski, F. 2003. Transcriptional changes in response to arbuscular mycorrhiza development in the model plant Medicago truncatula. Mol. Plant Microbe Interact. 16: 306–314.
Zeevaart, J.A.D. and Creelman, R.A. 1999. Metabolism and physiology of abscisic acid. Ann. Rev. Plant Physiol. Plant Mol. Biol. 39: 439–473.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Colditz, F., Nyamsuren, O., Niehaus, K. et al. Proteomic approach: Identification of Medicago truncatula proteins induced in roots after infection with the pathogenic oomycete Aphanomyces euteiches . Plant Mol Biol 55, 109–120 (2004). https://doi.org/10.1007/s11103-004-0499-1
Issue Date:
DOI: https://doi.org/10.1007/s11103-004-0499-1