Abstract
Cytokinins (CKs) are a complex group of compounds. The complexity of CKs is not just present at the level of chemical structures, but as a family of compounds. CKs occur in plants as nucleotides, nucleosides (ribosides) that are produced during de novo biosynthesis, free bases, and conjugates that are in stored/inactivated forms. Almost all organisms make cytokinins. CKs are structural components of the tRNA, and they are located next to the anticodon loop beginning with U of a subset of tRNAs in most eukaryotes and bacteria. The biosynthesis of CKs uses several pathways, and there is evidence that its biosynthesis is also regulated by other growth regulators such as auxins, strigolactones and abscisic acid. Its signaling pathway involves a phosphotransfer signal cascade. Despite advances made so far in the knowledge of CKs, there is still a long way to go. In this review, we summarize the most up-to-date knowledge on the biosynthesis of CKs and the signaling pathway that leads to the response to the presence of CKs in plant tissue, and we identify areas that require more research to complete our understanding of the role of the CKs in plants.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Since kinetin and its effect on the differentiation of tobacco calli were discovered by researchers at the University of Wisconsin (Miller et al. 1955b; Skoog and Miller 1957), cytokinins (CKs) have been the focus of many plant development and differentiation studies. CKs serve as plant growth signals in a variety of biological processes, such as cell division and differentiation, vascular and flower development, nutrient homeostasis, leaf expansion, circadian clock, abiotic stress responses, stomata aperture, seed germination, shoot apical meristem maintenance, branching and nodulation, leaf senescence, lateral root development, regulation of polar auxin transport, and root growth (Bishopp et al. 2011; Fusconi 2013; Han et al. 2014; Kieber and Schaller 2018; Liu et al. 2019; Mok and Mok 2001; Nitschke et al. 2016; Sakakibara 2006; Sasaki et al. 2014; Werner and Schmülling 2009; Wybouw and De Rybel 2019).
Unlike auxins, CKs are a very complex group of compounds, both in terms of chemical structures and of compound mixtures. Eudicotyledons and monocotyledons use different sets of CKs, at concentrations around 1% that tend to be higher in monocotyledons (Osugi and Sakakibara 2015). Most of the CKs are conjugated, mainly as glucosides (Bielach et al. 2012; Jiskrová et al. 2016; Osugi and Sakakibara 2015) with, among others, sugars and sugar phosphates, either on side-chain hydroxyl oxygens or on purine ring nitrogens (Mok and Mok 2001; Sakakibara 2006). In another type of complexity, the trans and cis zeatin isomers are synthesized by different pathways (Kasahara et al. 2004; Sakakibara 2006), and they play different roles in regulating growth (LaCuesta et al. 2018). Finally, due to their chemical diversity, CKs can also establish complex relationships with other growth regulators, such as auxins, strigolactones, gibberellins, abscisic acid (ABA) and nitrate (Noriega and Pérez 2017; Xu et al. 2015).
Structure and conjugates of cytokinins
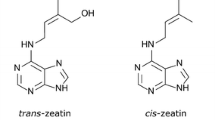
CKs are derivatives of purine bases with an isoprenoid or aromatic side chain at the N6 position (Fig. 1). The most common class of isoprenoid CKs comprises zeatin (Z), dihydrozeatin (DZ) and N6-(∆2-isopentenyl) adenine (iP). The first natural CK identified was zeatin, present in the immature endosperm of Zea mays (Miller 1961) in cis- or trans-configuration (depending on which of the two methyl groups is hydroxylated). Previously, it was shown that trans-zeatin (tZ) is the predominant CK in plants and that cis-zeatin (cZ) is less active, however the recent development of sensitive, rapid, high-throughput analytical methods has advanced our knowledge of the distribution and possible physiological functions of cZ (High et al. 2019; Schäfer et al. 2015; Sheflin et al. 2019; Wang et al. 2019).
Chemical structures of cytokinins. a Adenine structure and the numbering system for the nomenclature of CKs. b Representative Isoprenoid CKs and aromatic CKs. Only common names are given with commonly used abbreviations in parentheses. Note that the isoprenoid side chain is inside a blue oval and the aromatic chain inside the pink oval. c Structure of CK conjugates. The molecule conjugated with the CK is inside the gray oval
Aromatic CKs have an aromatic benzyl or hydroxybenzyl group at the N6 position. kinetin (KIN; 6-furfurylaminopurine) was the first synthetic aromatic CK discovered through bioassay of coconut milk concentrates in tobacco pith tissue culture (Miller et al. 1955a). The main natural aromatic CKs present in plant tissues are meta-topolin (mT; 6-[3-hydroxybenzylamino]purine), ortho-topolin (Ot; 6-[2-hydroxybenzylamino]purine), 6-benzyladenine (6-BA) and 6-BA derivatives (Aremu et al. 2014; Sáenz et al. 2003; Strnad, 1997), although these CKs are less abundant than the isoprenoid CKs (Sáenz et al. 2003). These structural variations of the side chain change the physicochemical properties of the CKs, in particular their polarity.
CKs occur in plants as nucleotides, nucleosides (ribosides), free bases and their respective conjugates. The CK bases can be conjugated with sugars (glycosylation, xylosylation) or amino acids (alanylation) (Fig. 1). The CKs are deactivated when conjugated with a sugar moiety, mostly glucose. The glycosylation at the O- or N-position is performed by the enzyme uridine diphosphate glycosyltransferase (UGT) (Entsch and Letham 1979; Hou et al. 2004). CK N-glucosides are formed irreversibly, in route to CK degradation (Hou et al. 2004), while CK-O glucosides can be reversibly de-glycosylated and represent a storage form (Brzobohaty et al. 1993; Falk and Rask 1995). Recent studies have shown that CK glycosylation occurs in the cytosol and serves as a rapid mechanism maintaining intracellular homeostasis, because glycosylation causes the CKs to be more hydrophilic and easily recognized by transmembrane transporters, effecting the release of CK glycosides from the cytosol or their storage in hydrophilic organelles such as vacuoles (Jiskrová et al. 2016; Lim and Bowles 2004; Šmehilová et al. 2016). The CKs can also be irreversibly inactivated by conjugation to alanine in the N9 position (Entsch et al. 1983). The hydroxyl group of the CK side chain can also be conjugated with xylose, and the resulting O-xylosylzeatin shows higher activity than zeatin in bioassays with Phaseolus vulgaris calli and a little conversion into zeatin. These findings suggest a possible biological function of O-xylosylzeatin, however its precise role is still unknown (Mok et al. 1987; Turner et al. 1987).
CKs are present in plant tissues at very low concentrations, in most cases at the level of pmol g−1 FW, making their isolation and quantification difficult. However, new procedures of extraction, purification and detection have allowed the detection of CKs and conjugates even at very low concentrations in minimal amounts of plant material from diverse species (Albacete 2017; Cao et al. 2017; Dobrev et al. 2017; High et al. 2019; Sheflin et al. 2019; Svacinova et al. 2012; Wang et al. 2019). The spatial distribution of CKs and their derivatives varies according to the physiological age of the plant and its interaction with abiotic factors such as stress or availability of nitrate.
Cytokinin biosynthesis pathways
Almost all organisms make CKs, as they are structural components of tRNA, located next to the anticodon loop beginning with the U of a subset of tRNAs in most eukaryotes and bacteria (Persson et al. 1994). This fact led to the initial proposition that the primary source of CKs could come from the degradation of tRNA (Swaminathan and Bock 1977). However, subsequent studies showed that there is a de novo biosynthesis pathway that does not involve the degradation of tRNA (Taya et al. 1978).
It was initially thought that the primary site of CK biosynthesis was the root. However, further research suggested CK biosynthesis in various locations of active cell proliferation, such as shoots and young leaves (Blackwell and Horgan 1994; Koda and Okasawa 1980; Sakakibara 2006, 2010). Studies of the chemical profile of the xylem and phloem sap showed that tZ-type CKs such as tZ and tZR predominate in the xylem sap, while iP-type CKs such as iPR and iP‐ribotides predominate in leaf exudates (Corbesier et al. 2003; Hirose et al. 2008).
Eukaryotic cells have the capacity to organize metabolic and signaling processes using subcellular compartmentation (Skalicky et al. 2018). This feature is of particular importance in the regulation of CK biosynthesis and homeostasis. The biosynthesis, conjugation, and storage of CKs take place in different compartments of the cell (Table 1) (Skalicky et al. 2018).
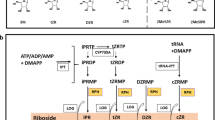
The first step in the biosynthesis of isoprenoid CKs is the N-prenylation of AMP, ADP or ATP, isoprenoid moieties donated by dimethylallyl pyrophosphate (DMAPP) or 4-hydroxy-3-methyl-2-(E)-butenyl diphosphate (HMBDP). DMAPP is a metabolic intermediate of the cytosolic mevalonate (MVA) and the plastidial methylerythritol phosphate (MEP) pathways, while HMBDP is an intermediate of the MEP pathway (Krall et al. 2002; Sakakibara et al. 2005, 2006) (Fig. 2). Isotopic marker studies in Arabidopsis thaliana seedlings showed that the prenyl side chains of tZ- and iP-type CKs are mainly produced through the MEP pathway, whereas a significant fraction of cZ derivatives are synthesized through the MVA pathway (Fig. 3) (Kasahara et al. 2004). The initial prenylation produces isopentenyl adenosine-5′-monophosphate (iPMP), isopentenyladenosine-5′-diphosphate (iPRDP) or isopentenyladenosine-5′-triphosphate (iPRTP). It is catalyzed by adenosine phosphate-isopentenyltransferase (EC 2.5.1.27; IPT) (Fig. 2), which was identified for the first time in the slime mold Dictyostelium discoideum (Taya et al. 1978). Later, the Agrobacterium tumefaciens gene tmr (subsequently termed ipt) was shown to encode an enzyme with similar activity (Akiyoshi et al. 1984; Barry et al. 1984), and overexpression of IPT genes in a variety of plants was found to cause an increase in CK content (Sun et al. 2003).
Model for cytokinin biosynthesis in plants. CK biosynthesis starts by transferring an isopentenyl moiety from dimethylallyl diphosphate (DMAPP) to AMP/ ADP/ATP. In A. thaliana the isopentenyl moiety is transferred mainly to ATP/ADP. This reaction is catalyzed by the phosphate-isopentenyltransferase (IPT) enzymes (pink). The initial products iPRMP, iPRDP, and iPRTP are subsequently hydroxylated in the isoprenoid side chain by cytochrome P450 monooxygenases CYP735A (red) for the formation of the corresponding tZ-nucleotides. The ribotides (purple box) can be converted to their free bases by two pathways: the two-step activation is catalyzed by 5′-ribonucleotide phosphohydrolase for the formation of ribonucleosides (pink box) and by adenosine nucleosidase for the formation of the active form (blue box). So far only one has been identified, a nucleoside N-ribohydrolase (NRH), which catalyzes the hydrolysis of iPR to iP (green). The direct activation pathway is catalyzed by 5′-monophosphate phosphoribohydrolases (LOGs) (blue). The formation of CKs of the cis-Zeatin-type begins with the prenylation of adenine 37 on specific (UNN-) tRNAs by tRNA-isopentenyltransferase (tRNA-IPT) (orange) and subsequent release of CK nucleotides by tRNA degradation. (Sakakibara 2006). (Color figure online)
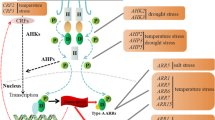
Cytokinin biosynthesis, transport, and homeostasis in plants. a Sites of biosynthesis and distribution. tZ and its inactive precursor tZR are synthesized mainly in the root; both are transported through the xylem towards the shoot. Later tZR is activated by LOG enzymes; tZ acts directly on leaf size, and root-derived tZR controls both leaf development and SAM activity. On the other hand, iPR and iP are synthesized mainly in the shoot and are distributed to the active site through the phloem ( modified from Davière and Achard P 2017). b Model of cellular homeostasis of cytokinins. The first reaction of the de novo biosynthesis of CKs is catalyzed by the IPT enzymes (blue), which are located mainly in plastids and produced trans-zeatin-type and iP-type CKs through the MEP pathway; nevertheless, IPT7 is localized in mitochondria and IPT3 undergoes a process of farnesylation that causes its translocation in the cytosol and nucleus. The cis-zeatin-type CKs are mainly synthesized through the MVA pathway in the cytosol. The LOGs enzymes (purple), which transform CK nucleotides into an active form, are located in the cytosol and nucleus. CKX enzymes (green), which degrade CKs, are preferably located in the apoplast, although they are also present in the cytosol and vacuole. The conjugation of free bases occurs mainly with glucose, a reaction that is catalyzed by uridine diphosphate glycosyltransferases (UGTs), the CK-O glucosides localized primarily in the vacuole; nevertheless, they can be reversibly deglycosylated by a β-glucosidase enzyme in the cytosol. So far three intracellular CK transporters have been identified: ABCG14 is an exporter of tZ-type CKs, while purine permeases (PUPs) and equilibrative nucleoside transporters (ENTs) have been proposed for import of apoplastic CKs to the cytosol. It is possible that there is an intracellular transport system of the plastids, mitochondria, and vacuole to the cytosol; however, there is still no evidence for this. CK receptors ARABIDOPSIS HISTIDINE KINASES (AHKs) are localized at the plasma membrane, and the endoplasmic reticulum initiates the signaling cascade. CKs are perceived by the AHK receptors, which autophosphorylate and subsequently transfer the phosphate (blue arrows) to proteins (AHP1 to AHP5), which enter the nucleus and phosphorylate type-B ARABIDOPSIS RESPONSE REGULATORS (B-ARRs) that activate transcription of CK response genes or B-ARR negative regulators of CK signaling. (Color figure online)
Plants have two classes of IPT enzymes differing in the substrates they use (Miyawaki et al. 2004). Adenosine-phosphate IPTs (AP-IPTs) use ATP, ADP or AMP as a substrate, and studies of various IPT mutants indicate that they participate in the biosynthesis of iP or tZ (Kakimoto, 2001; Takei et al. 2001a). In contrast, tRNA-IPTs catalyze the isopentenylation of UNN-codon tRNAs at adenine (A37), causing the subsequent release of CK nucleotides by the degradation of the tRNA (Murai 1994) and formation of cZ (Fig. 2) (Miyawaki et al. 2006).
In A. thaliana, the IPT family comprises seven AP-IPTs (IPT1, IPT3, IPT4, IPT5, IPT6, IPT7, and IPT8) and two tRNA-IPTs (IPT2 and IPT9) (Kakimoto 2001; Takei et al. 2001a). Phylogenetic studies of IPT genes in several angiosperms, such as A. thaliana, Oryza sativa, and Solanum lycopersicum, revealed a multigene family with complex evolutionary history marked with duplication events which may be associated with functional redundancies (Ghosh et al. 2018; Matsuo et al. 2012; Nishii et al. 2018). Furthermore, AP-IPTs showed differential expression in different organs and, thus, homolog-specific control of the local CK concentration (Miyawaki et al. 2004). For example, the A. thaliana IPT3 had high level of expression in the phloem, roots, hypocotyl, cotyledons and leaves (Miyawaki et al. 2004; Takei et al. 2004a), while several O. sativa IPTs were highly expressed in the shoot apical meristem, roots and leaf primordia (Ghosh et al. 2018). These data suggest that the precursor of iP-type CKs can be synthesized in a wide range of plant parts.
The enzymes AtIPT1, AtIPT3, AtIPT5 and AtIPT8 are located in plastids, and an Allium cepa epidermal cell assay showed that AtIPT3 contains a C-terminal CaaX motif that is farnesylated, directing the modified protein to be located in the nucleus and the cytoplasm, while the nonfarnesylated protein is located in the plastids (Galichet et al. 2008). In contrast, AtIPT4 and AtIPT2 are restricted to the cytosol and AtIPT7 to the mitochondria (Kasahara et al. 2004) (Fig. 3). Similarly, in O. sativa OsIPT2, OsIPT71, OsIPT72, and OsIPT8 proteins were also predicted to be localized in the cytoplasm, and the proteins OsIPT1, OsIPT3, OsIPT5 in the chloroplast (Ghosh et al. 2018).
The initial products in the biosynthesis of CKs, iPRMP, iPRDP and iPRTP, are subsequently hydroxylated in the isoprenoid side chain by a cytochrome P450 enzyme, forming tZ-riboside 5′-phosphates (tZRPs) (Fig. 2). A. thaliana has two cytochrome P450 monooxygenases, CYP735A1 and CYP735A2, functioning in this step of CK biosynthesis (Takei et al. 2004b). Double mutants affected in both genes showed a lower concentration of tZ and its conjugates, causing a delay in the shoot growth and underlining that they play a role in CK biosynthesis. Recent studies in Jatropha curcas showed that JcCYP735A mutants have decreased in amounts of tZ and tZR and reduced growth (Cai et al. 2018). Finally, diverse other reports confirmed the existence of CYP735A orthologs in such diverse plant species as Brassica oleracea, O. sativa, Glycine max and S. lycopersicum, suggesting that the function of CK trans-hydroxylation may be highly conserved (Guttikonda et al. 2010; Nelson and Werck-Reichhart 2011; Simm et al. 2016; Yu et al. 2017). Interestingly, CYP735A genes is predominantly expressed in root vasculature of Arabidopsis (Kiba et al. 2013; Matsumoto-Kitano et al. 2008), alongside other CK biosynthetic enzymes like IPT5 and LOG3. Together, these findings match previous reports that tZ-type cytokinins are mainly synthesized in roots (while iP-type cytokinins are mostly formed in shoots) (Fig. 3), thus further confirming the tissue- and organ-specific regulation of CK biosynthesis.
The CK ribotides can be converted into the corresponding free bases by two pathways, the two-step activation pathway and the direct activation pathway (Chen, 1997; Kurakawa et al. 2007). In the two-step activation pathway, CK ribotides are hydrolyzed to CK ribosides by 5-ribonucleotide phosphohydrolases. Subsequently, CK ribosides are hydrolyzed to free bases by adenosine nucleosidase (EC 3.2.2.7) (Chen and Kristopeit 1981) (Fig. 2). To date, there are few reports on the functionality of the two-step activation pathway. The first study on the role of the two pathways came from studies in A. thaliana (Jung et al. 2009). The enzyme uridine-ribohydrolase 1 (URH1; EC 3.2.2.3) can hydrolyze the CK derivative isopentenyladenine-riboside, as well as uridine and inosine ribosides, and changes in the expression of this gene retard germination (Jung et al. 2009). More recently, two NRH subclasses have been identified in plants (Kopecná et al. 2013), one hydrolyzing the purine ribosides inosine and xanthosine and the other uridine and xanthosine, while both can hydrolyze CKs ribosides. Their substrate specificities depend on the amino acid at position 249. A tyrosine residue in this position is associated with preferential hydrolysis of purine ribosides, while the presence of aspartate confers high activity on uridine (Kopecná et al. 2013). However, it appears that the hydrolysis of CK ribosides is not the primary function of these enzymes.
There is a direct CK biosynthesis pathway producing free bases in a reaction catalyzed by phosphoribohydrolases encoded by members of the LONELY GUY (LOG) gene family. The protein cytokinin riboside 5′-monophosphate phosphoribohydrolase (LOG, EC:3.2.2.n1) was discovered through the analysis of O. sativa mutants (Kurakawa et al. 2007) characterized by premature termination of the shoot meristem. LOG encodes an enzyme able to produce CKs from their ribosides in one step, by hydrolyzing the bond between the N6-substituted bases and the ribose 5′-monophosphate (Kuroha et al. 2009; Seo and Kim 2017). The direct hydrolysis of the CK ribotides into free CKs has significant biological implications, since it provides a direct mechanism for regulation of CK biosynthesis, where CKs ribotides can be transported to the site of action and then converted into free CKs. Studies with mutants of A. thaliana and Medicago truncatula showed that LOG genes participate in the nodule primordium development and lateral root formation (Kuroha et al. 2009; Mortier et al. 2014; Tokunaga et al. 2012), while LOG overexpression reduced apical dominance, retarded leaf senescence and increased cell division rates in root vasculature and embryos (Kuroha et al. 2009).
Although the initial step of CK de novo biosynthesis mainly occurs in plastids, further processes occur in the cytosol. Accordingly, the CK-activating LOG enzymes are localized mainly in the cytosol (Kuroha et al. 2009), like the antagonistic adenine phosphoribosyltransferases (APT1, APT2, APT3; EC: 2.4.2.7) catalyzing the conversion of free CKs into corresponding nucleotides (Allen et al. 2002; Moffatt et al. 1991; Zhang et al. 2013). Together, these findings imply a transport system that moves precursors from the plastids to the cytosol, however evidence for this transport system is lacking to date (Fig. 3).
In A. thaliana, tZR is the most abundant CK detected in the xylem, comprising approximately 80% of the total CKs, while tZ is second-most abundant (Osugi et al. 2017). tZR and tZ are transported through the vascular system from the root to the shoot using ABCG14, an ATP-binding cassette transporter (ABC) (Ko et al. 2014). Recent studies have shown that tZR and tZ are the primary long-distance signaling forms in the xylem which, together, control leaf size and meristem activity-related traits (Osugi et al. 2017). On the other hand, studies on the expression of the LOG family genes in A. thaliana and O. sativa suggest that the production of free CKs occurs in virtually all parts of the plant (Kurakawa et al. 2007; Kuroha et al. 2009; Yamburenko et al. 2017). Therefore, the plants use tZ and tZR as messengers, which are transported from the root to the shoot, and iP‐type CKs as systemic messengers distributed through the phloem to the active site (Fig. 3). Although the general features of terpenoid CK biosynthesis have been established, there are still several aspects of the process that are not fully understood. Also, we have almost no information on the biosynthesis of aromatic CKs.
Regulation of cytokinin biosynthesis
Ever since the discovery of CKs, their interaction with other growth regulators has been studied, in many cases focusing on the relative hormone concentrations in various plant developmental processes. Early on, it was shown that endogenous CK accumulation was inhibited when the tissue was cultured on auxin-containing medium (Beinsberger et al. 1991; Hansen et al. 1985; 1987; Zhang et al. 1995). Further studies into the underlying mechanisms proved that auxin reduces the expression level of IPT genes (Song et al. 1995; Zhang et al. 1996). Later, in vivo deuterium incorporation experiments in A. thaliana showed that exogenous auxin exercises a rapid negative control over the CK biosynthesis pathway (Nordström et al. 2004). Auxin also represses local CK biosynthesis in the nodal stem of Pisum sativum through the expression of the PSIPT1 and PSIPT2 genes (Tanaka et al. 2006). However, in actively growing tissues, such as developing roots and shoots, the CKs induce auxin biosynthesis, suggesting that a regulatory feedback loop maintains auxin and CK concentrations optimal for cell- and tissue-specific developmental processes (Jones et al. 2010). Subsequent studies demonstrated that auxin controlled the spatiotemporal distribution of CK biosynthesis through the negative regulation of AtIPT expression by AUXIN RESPONSE FACTOR3 (ARF3) during de novo SAM formation (Cheng et al. 2013). The ARF3 protein is a member of the transcription factor family that activates or represses auxin response gene expression (Leyser 2018). Recent studies have shown that ARF3 also regulates CK biosynthesis directly by repressing the expression of IPT genes and indirectly repressing LOG genes during floral meristem formation (Zhang et al. 2018). On the other hand, auxin upregulates CYP735A genes in A. thaliana and O. sativa; however, the molecular mechanism involved is still unknown (Takei et al. 2004b; Tsai et al. 2012).
There is evidence that CK biosynthesis is also affected by other growth regulators such as strigolactones (SLs) and abscisic acid (ABA). In roots of SL-deficient or SL-insensitve mutants of pea and Arabidopsis, CK amounts are dramatically reduced, although there are no differences in the CK levels in the shoot apices (Foo et al. 2007; Morris et al. 2001). Subsequent studies showed that in P. sativum the expression of the CK biosynthesis gene PsIPT1 is increased in the nodes and internodes of the shoot of the SL mutant (Dun et al. 2012); however, the expression of PsIPT1 and PsIPT2 genes in the stem is not affected by SL defects. Furthermore, the addition of synthetic SL did not affect transcript levels for either IPT genes, similar to the effect in O. sativa (Dun et al. 2012; Xu et al. 2015; Young et al. 2014). Recent studies in Zantedeschia have suggested an antagonistic interaction between SLs and CKs during adventitious bud development, since high SL levels were detected in a little branched cultivar while, conversely, the CK concentration was low compared in a more highly branched cultivar (Manandhar et al. 2018). Further research is needed to determine how SLs and CKs interact, as it is possible that SLs affect the CK metabolism through additional undiscovered members of gene families involved in their homeostasis or by regulation at the post-translational level.
Regarding the effect of ABA in CK biosynthesis, it is known that ABA treatments cause downregulation of essential CK biosynthetic genes such as IPT and CYP735A, which triggers a reduction in CK concentration (Nishiyama et al. 2011; Takei et al. 2004b; Tsai et al. 2012). In addition, a decrease in ABA content causes an increase in CK levels during bud sprouting in grapevine (Noriega and Pérez 2017), suggesting an antagonistic role between both growth regulators.
The biosynthesis of CKs is also modulated by the availability of mineral nutrients and environmental changes. Nitrate promotes the accumulation of CKs in the roots of Hordeum vulgare, Z. mays and A. thaliana (Takei et al. 2001b), similar to its function in tobacco and sunflower leaves (Salama et al. 1979; Singh et al. 1992). In Arabidopsis, the expression of AtIPT3 and AtCYP735A2 is rapidly induced by nitrate, leading to the accumulation of tZR and the tZRMP in roots. However, the amounts of AtIPT5 transcript in lateral root primordia increases with the availability of NO3− and NH4+ in long-term treatments (Kiba et al. 2010; Miyawaki et al. 2004; Takei et al. 2004a; Wang et al. 2004). Furthermore, in monocotyledonous plants like O. sativa, nitrogen increased CK amounts by enhancing the expression of OsIPTs (Xu et al. 2015).
Several studies have shown that CKs regulate various processes in the plants in response to nitrate, for example dictating leaf size in S. lycopersicum and N. tabacum (Rahayu et al. 2005; Walch-Liu et al. 2000), and SAM size and branch number in A. thaliana (Landrein et al. 2018; Müller et al. 2015). Furthermore, when little nitrate is available overall, then CKs enhance the number and development of lateral roots in those areas rich in NO3− (Kiba and Krapp 2016; Ruffel et al. 2011). Recent results have shown that nitrate availability controls the ratio of tZ to tZR in xylem sap (Osugi et al. 2017), further underlining that CKs act as long-distance signals mediating the shoot response to NO3− perception in roots. New studies have revealed that the CK biosynthetic and transport mutants IPT3,5,7 and ABCG14 in the presence of nitrate failed induce genes that respond rapidly to long-distance N-supply signals, indicating that N assimilation pathways in shoots are CK-dependent. Furthermore, levels of tZ in shoots of both mutants decreased, suggesting that tZ is an important element of the long-distance N-signaling network controlling root responses (Poitout et al. 2018). On the other hand, the expression of AtIPT3 is also induced by the provision of other nutrients like iron, sulfate and phosphate to the roots (Kiba et al. 2010; Séguéla et al. 2008). In addition, salicylic acid induces CK biosynthesis, increasing the level of expression of the IPT7 gene in A. thaliana and Sorghum bicolor. In both species, silicon accumulation delays dark-induced leaf senescence (Markovich et al. 2017). Therefore, to manage resource availability, IPT genes can affect overall plant development by controlling the biosynthesis of different active forms of CKs in response to either scarcity or excess of a nutrient.
Cytokinin catabolism
One of the factors controlling intracellular CK levels is their conjugation, mainly with β-D-glucose at the O- and N-positions. O-glycosylation occurs at the hydroxyl group of the Z and DHZ side chains, while N-glycosylation occurs at positions N3, N7 or N9 of the purine ring (Entsch et al. 1979; Entsch and Letham 1979). The glycosylation reaction is catalyzed by UGT enzymes (Lairson et al. 2008; Mok et al. 2000) and, for example, the O-glycosylation of Z is carried out by zeatin O-glucosyltransferase (EC: 2.4.1.203). In Arabidopsis, five genes encoding CK-specific UGT enzymes (UGT76C1, UGT76C2, UGT85A1, UGT73C5, and UGT73C1) have been identified (Hou et al. 2004; Jin et al. 2013; Li et al. (2015); Wang et al. 2011). UGT76C1 and UGT76C2 are specific cytokinin N-glucosyltransferases conjugating CKs on the N7 and N9 positions of the adenine moiety (Hou et al. 2004; Wang et al. 2011; 2013), while UGT85A1, UGT73C5, and UGT73C1 catalyze the O-glycosylation of tZ and DHT (Hou et al. 2004; Jin et al. 2013). Both loss-of-function and gain-of-function mutants of CK-specific UGTs (ugt76c2 and ugt76c1) change the expression levels of AHK2, AHK3, ARR1, CYP735A2, LOG2, IPT5, and CKX3, suggesting a tight control of CK homeostasis (Wang et al. 2011, 2013). Several studies have shown that CK-specific UGTs are mainly located in the cytosol (Jin et al. 2013; Šmehilová et al. 2016).
The CK O-glucosides can be de-glycosylated and converted into active forms by β-glucosidase enzymes (Brzobohaty et al. 1993; Kiran et al. 2006). New methods of quantification and purification in isolated organelles have shed light on the intracellular distribution of CKs, showing that O-glucosides are mainly found in the vacuole. Therefore, this organelle is an essential reservoir for storing CK conjugates, and it is possible that there are specific transporters for O-glucosides from the cytosol to the vacuole (Fig. 3) (Jiskrová et al. 2016; Kiran et al. 2012).
Further mechanisms regulating endogenous CK levels involve CK oxidase/dehydrogenase (CKX; EC: 1.5.99.12) enzymes catalyzing the irreversible oxidative breakdown of CKs into adenine and unsaturated aldehyde 3-methylbut-2-enal (Bilyeu et al. 2001; Brownlee et al. 1975; Galuszka et al. 2001, 2004; Niemann et al. 2018; Reid et al. 2016). CKXs are encoded by a multigene family, and A. thaliana has seven members located mainly in the apoplast, cytosol and vacuole. However, CKX1 is also localized in the endoplasmic reticulum (ER), from there CKX1 control the subcellular levels of CK, which triggers the signaling from the ER (Niemann et al. 2018; Werner et al. 2003).
Cytokinin transport
CK homeostasis and signal perception rely on complex patterns of intercellular movement as well as local (short-distance) transport between neighboring cells and long-distance translocation for example between roots and shoots (Table 1) (Liu et al. 2019; Skalicky et al. 2018). The transport of CKs between organs and the intracellular trafficking involved are very actively being researched, and several review papers have recently summarized progress in this area (Durán-Medina et al. 2017; Kang et al. 2017; Ko and Helariutta 2017; Liu et al. 2019; Park et al. 2017). Here, only a brief summary of our present knowledge of CK transport will be provided.
CKs are transported from their biosynthesis site to their site of action by passive diffusion and through active transport mechanisms, and different CKs are translocated using various mechanisms. For example, the tZ-type CKs synthesized in roots are transported apoplastically to shoots, where they act by promoting growth (Beveridge et al. 1997; Hirose et al. 2008). Thus, the main CKs transported in the xylem sap are tZ and tZR, accounting for 15% and 80% of CKs there, respectively (Beveridge et al. 1997; Hirose et al. 2008; Kuroha et al. 2009; Osugi et al. 2017). Conversely, the iP- and cZ-type CKs synthesized in shoots travel through the phloem to roots (Corbesier et al. 2003; Hirose et al. 2008).
Three types of transporters are involved in CK trans-membrane transport and intercellular translocation. Among these, the ENT (Hirose et al. 2005; Sun et al. 2005) and PUP (Bürkle et al. 2003; Zürcher et al. 2016) transporters import apoplastic CKs nucleoside or nucleobase CKs into the cytosol, respectively (Fig. 3). Yeast studies showed that the ENT transporters of A. thaliana and O. sativa participate in the uptake of iP-ribosides and tZ-ribosides (Qi and Xiong 2013; Sun et al. 2005), while PUPs transport tZ and iP (Bürkle et al. 2003). However, PUPs and ENTs have also been implicated in the transport of other molecules, so they must be considered non-specific for CK transport (Durán-Medina et al. 2017; Gillissen et al. 2000; Girke et al. 2014). Very interestingly, the Arabidopsis PUP14 was recently found to modulate CK signaling (Zürcher et al. 2016) by regulating the availability of CKs in the apoplast, where they are perceived by ARABIDOPSIS HISTIDINE KINASE (AHK) receptors. The Arabidopsis AHK2, AHK3 and AHK4 proteins are located in the plasma membrane and the ER, and they are involved in transcriptional activation of various target genes (Fig. 3b) (Caesar et al. 2011; Lomin et al. 2018; Pernisová et al. 2018; Wulfetange et al. 2011; Yamada et al. 2001).
The third type of CK transporters, ATP-binding cassette (ABC) transporters, G subfamily, affects the acropetal long-distance transport of CKs from the root to the shoot (Ko et al. 2014; Zhang et al. 2014). In particular, the AtABCG14 and OsABCG18 proteins participate in xylem loading and are essential for the long-distance transport of root-derived CKs and, thus, the positive regulation of shoot growth (Kang et al. 2017; Zhao et al. 2019).
Interestingly, the Arabidopsis genome encodes numerous members of the three families of transporters, including 21 PUPs, 8 ENTs and 28 half-size ABCG-type transporters (Kang et al. 2011; Li et al. 2003; Zürcher et al. 2016). The numbers of homologs in each family varies drastically between plant species (Liu et al. 2019), with only four ENTs in rice for example (Hirose et al. 2005).
Cytokinin signaling: a two-component system
The elucidation of the CK signaling pathway started two decades ago with the studies of CK effects on Arabidopsis tissues (Inoue et al. 2001; Kakimoto 1996). This pathway involves a phosphotransfer signal cascade, similar to that in bacteria and most fungi (including yeast). The plant signal cascade involves histidine kinases (HKs), histidine phosphotransfer (HPt) proteins, and a specific type of response regulators (RRs). To elucidate the different components of the CK signal cascade, experiments were performed in Arabidopsis mutants in combination with protoplast transient expression assays (Argyros et al. 2008; Ferreira and Kieber 2005; Higuchi et al. 2004).
The cytokinin signaling system found in plants is very similar to the two-component system for signal transduction occurring in bacteria (Grefen and Harter 2004; Schaller et al. 2007; Stock et al. 2000). The CK bind to the CHASE domain of HK receptors and the signal is transmitted across the membrane (Bartrina et al. 2017; Kieber and Schaller 2018; Steklov et al. 2013). The HK cytokinin receptors have a conserved cytokinin-binding extracytosolic CHASE (cyclases/histidine kinases-associated sensing extracellular) domain (Kieber and Schaller 2018).
CKs binding to the hybrid histidine kinase receptors [AHK2, AHK3, AHK4 (CRE1/WOL)], results in the autophosphorylation of a conserved histidine (His) residue in their kinase domain. The phosphate is then first transferred to a conserved aspartic acid residue of the AHKs and then to a His residue of AHP proteins (Argyros et al. 2008). The modified AHPs move to the nucleus, where they transfer the phosphate group to two classes of ARRs (Argyros et al. 2008; Gordon et al. 2009).
HK proteins are known to participate in different biological processes in various plant species, including responses to environmental changes and growth regulator signaling. The HK proteins can be divided into two groups, the ethylene and the non-ethylene receptors, and both groups are conserved between plant species with high structural similarity (Nongpiur et al. 2012). Eight transmembrane HKs belonging to the latter group have been identified in Arabidopsis, and more in other species like G. max with 17 AHKs (Müller and Sheen 2007a). Only some of the Arabidopsis AHKs function as CK receptors, with AHK2, AHK3 and AHK4 each binding to different CKs. The AHK4 receptor (also known as CRE1 or WOL1) was the first component of CK signaling identified, phosphorylating AHPs when CKs are present and, conversely, dephosphorylating AHPs in their absence (Higuchi et al. 2004). AHK single and double mutants were not affected in CK signaling, but triple mutants showed severe phenotypes, especially in roots and shoots, suggesting highly redundant functions of these receptors (Ferreira and Kieber 2005). Root elongation in particular was reduced in ahk4 single mutants, but not in ahk2 and ahk3 single and double mutants (Ferreira and Kieber 2005).
The HK receptors possess a conserved region crucial for CK binding of 200 to 230 amino acids, designated as cyclases-/histidine kinases–associated sensory extracellular (CHASE) domain (Anantharaman and Aravind 2001; Hutchison and Kieber 2002). The same domain is present in a variety of receptor-like proteins from prokaryotes and eukaryotes (Anantharaman and Aravind 2001; Mougel and Zhulin 2001). The CHASE domain in the histidine kinase PcrK sensing CKs in the bacterial pathogen Xanthomonas campestris pathovar campestris was crystallized and its structure determined at 2.55 Å resolution (Chen et al. 2019). The PcrK protein is a homodimer with overall topology likely similar to AHK4, but also with several significant differences. Among these, the bacterial protein has a ligand-binding pocket with limited size to allow only the binding of iP, whereas the plant CK receptor can interact with a broad spectrum of CKs. All bacterial PcrK homologs have a positively charged residue Arg175 in the ligand-binding cavity important for binding negatively charged and hydrophobic groups, which is replaced by a negatively charged residue Asp262 in AHK4 (Chen et al. 2019).
Although the CHASE domain appeared a long time ago, it is still not clear how land plants acquired it, but there are several theories. One of them is that the Ectocarpus siliculosus virus-1 could have infected and integrated its genome into his host, the brown algae; and the other one, is through their chloroplasts, due to a cyanobacterial ancestry (Heyl et al. 2007; Schaller et al. 2011). In A. thaliana has been reported three histidine kinases containing the CHASE domain (AHK2, AHK3, and CRE1/AHK4) in which the difference among them is the number of transmembrane segments at their N-terminus. The CHASE domain at these proteins is the recognition site in CK receptors, and also plays an essential role in the regulation of plant growth and development, such as the vascular morphogenesis of the root (Mougel and Zhulin 2001; Müller and Sheen 2007b; Schaller et al. 2011).
There are five canonical AHP genes (AHP1-5) in the Arabidopsis genome comprising the conserved domain necessary for the phosphotransfer signal cascade, along with one pseudo-AHP gene. The latter, AHP6, lacks the His residue and does not participate in phosphate group transfer, therefore interfering and competing with the other AHPs and acting as a negative regulator of CK signaling. Nonetheless, mutants devoid of this protein show abnormalities in vascular tissue patterns (Hutchison et al. 2006; Hwang et al. 2012; Müller and Sheen 2007a). Further experiments with mutants deficient in different combinations of the other AHPs confirmed that AHP1-5 all play similar roles as positive CK signaling regulators, and that loss of their function causes damage to plant growth and development.
Phosphorylation of AHP proteins leads to nuclear translocation, followed by further transfer of the phosphate group first to B-ARRs, which in turn induce further transfer to a conserved Asp residue of A-ARRs. More than 20 ARRs have been identified and classified into four groups according to their C-terminal domain and similarities in their core domain sequence, including 10 A-ARRs (ARR3-9, ARR15-17), 11 B-ARRs, 2 C-ARRs and 9 pseudo-ARRs in Arabidopsis. The A- and B-AARs are best studied (Argyros et al. 2008; To and Kieber 2008), establishing that B-ARRs have long C-termini with a Myb-like domain for DNA binding (also known as GARP), while type A-ARRs have short C-termini. Despite this difference, all ARRs share a conserved domain for phosphorylation. B-ARRs are positive regulators of CK signaling, while A-ARRs are negative CK signaling regulators (Argyros et al. 2008; To et al. 2004). A-ARRs lack a DNA-binding domain and are affected in the presence of light. It is thought that they inhibit CK signaling by competing with the B-ARRs for phosphate groups transferred through the signaling cascade (Osugi and Sakakibara 2015; To et al. 2004; To and Kieber 2008). However, they can participate in signaling by integrating the ethylene and CK signaling pathways in a plant responses to environmental stress and varying nitrogen levels, resulting in changes in the endogenous levels of CKs (Shi et al. 2012; To and Kieber 2008).
The Arabidopsis genome contains eleven B-ARRs (ARR1, ARR2, ARR10–14 and ARR18–21), of which seven belong to subfamily I and two each to subfamilies II and III. Characterization of B-ARR mutants confirmed that especially ARR1, ARR2, ARR10, ARR11 and ARR12 play pivotal, partially redundant roles (Argueso et al. 2010; Ishida et al. 2008; Yokoyama et al. 2007). It has recently been found that the CK signaling pathway is mediated by S-PHASE KINASE-ASSOCIATED PROTEIN1 (SKP1)/Cullin/F-box (SCF) E3-ubiquitin ligase complex and also includes a proteasome-mediated degradation, similar to other plant growth regulators like auxins (Kim et al. 2013a, b; Osugi and Sakakibara 2015). The F-box family protein named KISS ME DEADLY (KMD) regulates interacts with B-ARRs in Arabidopsis and rice, thus affecting their ubiquitination-dependent degradation via the 26S proteasome and regulating their local levels and activity. KMD is interacting most strongly with ARR1, ARR2 and ARR12, which have been identified as the most important ARRs in CK response (Kim et al. 2013a, b; Osugi and Sakakibara 2015).
ARRs can also interact with nuclear Cytokinin Response Factors (CRF), a family of APETALA2/ERF transcription factors comprising six members in Arabidopsis (Argueso et al. 2010; Müller and Sheen 2007a; Nongpiur et al. 2012). Mutants deficient in CRFs have altered expression of several (but not all) CK-related genes similar to B-ARR mutants. However, the mechanisms of CRF regulation and signal transduction are not yet known (Argueso et al. 2010; Hwang et al. 2012; To and Kieber 2008).
There are reports of other transcription factors involved in CK signaling, such as the GLABROUS1 enhancer-binding protein (GeBP/GPL) and splindy (SPY). The inactivation of GeBP/GPL transcription factors causes a reduction in the sensitivity to exogenous CKs and an increase in the expression of A-ARRs, antagonizing the negative regulation of CK signaling (Argueso et al. 2010). In Arabidopsis, loss-of-function mutants in SPY cause CK insensitivity, suggesting that SPY may be acting as a positive regulator of the CK signal cascade and also as a regulator of CK/gibberellin homeostasis (Greenboim-Wainberg et al. 2005).
The mechanistic understanding of the two-component CK signaling system has enabled the development of interesting molecular tools, most notably a synthetic CK reporter coupled with GFP (TCS::GFP) for in vivo tracking of the CK signaling output (Brenner et al. 2012; Hwang et al. 2012). This reporter was designed based on the conserved DNA-binging domain of the B-ARRs (A/G)GAT(T/C), with six direct repeats of this motif. Interestingly, the CK signaling output was first detected in the hypophysis and, thus, coinciding with relatively low auxin activity, showing that auxin is an antagonist to CKs (Brenner et al. 2012; Müller and Sheen 2008). However, the signal of the synthetic CK reporter was weak in some developmental tissues where high CK activity had previously been reported, and GFP expression was found to decrease over generation, suggesting that the repetitive DNA-binding motif may have led to silencing. To test this, a novel TCS reporter was developed that exhibited strong expression patterns that are stable from one generation to another (Zürcher et al. 2013).
Despite all the efforts made to elucidate the complete CK signaling pathway, questions remain. In particular, it is not known how the different components of the CK signaling pathway work, which combinations of proteins participate in a given biological event and, specifically, which CK forms trigger certain response mechanisms in different plant models.
Cytokinin crosstalk with ABA
Under normal conditions, plant cells receive signals and stimuli from adjoining tissues, and they respond to these signals such that physiological conditions are stabilized in the whole plant. However, this status can be disrupted if isolated cells or tissues are subjected to stress, due to changes in nutrient concentrations or environmental conditions, and affected cells initiate alternative regulatory mechanisms for acclimatization (Nic-Can et al. 2016). Those mechanisms include the reprogramming of gene expression and physiological as well as metabolic changes that let the cells survive and differentiate (Tardieu et al. 2018; VanWallendael et al. 2019). Stress tolerance involves mechanisms operating at different spatial and temporal scales that, together, greatly affect plant growth. Several PGRs play central roles in this response, and crosstalk among them is crucial to maintain the health of the plants. Crosstalk between CKs and ABA has been investigated in some detail, as it occurs in several processes both in response to stress and independent of it.
During the artificial bud sprouting induced in Vitis vinefera, ABA biosynthesis decreased while expression of CK-related genes increased, including the CK biosynthesis genes ISOPENTENYL TRANSFERASE (VvIPTs) and LONELY GUY (VvLOG1), the CK catabolism gene CYTOKININ OXIDASE (VvCKX3), and the key auxin biosynthesis gene VvYUC3. Together, these findings suggest that bud sprouting is preceded by a decrease in ABA content and an increase in CKs as well as auxin levels (Noriega and Pérez 2017). Bud outgrowth inhibition by low photosynthetic photon flux density (PPFD) was associated with lower CK and sugar contents and higher ABA content in the stem of Rosa hybrida ‘Radrazz’. CKs exogenously supplied to the stem restored bud outgrowth under low PPFD, and ABA supply antagonized the effect of CKs, further underlining that CKs and ABA play opposite roles (Corot et al. 2017).
The complexity of crosstalk among PGRs is manifested in the maintenance of stem cell and vascular differentiation during adventitious root formation in Marubakaido apple rootstock, where crosstalk among CKs, auxins, plant peptide hormones and ABA occurs (Saito et al. 2019). A similar degree of complexity was also reported for the regulation of root growth under osmotic stress conditions, where ABA functions in interaction with a PGR network of CKs, ethylene and auxin (Rowe et al. 2016).
CKs regulate several aspects of plant growth and development, including plant response to abiotic stress (Albacete 2017), e.g., CKs negatively regulate salt (Aldesuquy et al. 2014; Tounekti et al. 2011) and drought stress signaling (Davies and Zhang 1991; Rivero et al. 2010). Consequently, CK-deficient plants are stress-tolerant, showing increased cell membrane integrity and ABA hypersensitivity (Nishiyama et al. 2011). Conversely, wild-type Arabidopsis plants show a decline in CKs both under stress and after ABA treatment, as a consequence of decreased expression of ISOPENTENYL-TRANSFERASE and CYTOKININ OXIDASES/DEHYDROGENASES genes (Nishiyama et al. 2011). Interestingly, also the uridine diphosphate glycosyltransferase 76C2, a CK glycosyltransferase, is also suppressed under these conditions, revealing novel cues in abiotic stress adaptation (Cai et al. 2018). These results suggest that a reciprocal regulation mechanism exists between the CKs and ABA metabolism in response to stress, plant growth and development (Nishiyama et al. 2011). The consequence of the stress-induced changes in CKs and ABA is leaf abscission, likely to reduce water loss and to enhance the survival of perennial plants, albeit at the expense of reduced growth (Rivero et al. 2009). On the other hand, the addition of kinetin induces tolerance against various abiotic stresses such as drought in Nicotiana tabacum (Rivero et al. 2010), and salinity in Triticum aestivum (Aldesuquy et al. 2014) as well as Salvia officinalis (Tounekti et al. 2011).
The presence of heavy metal in the soil is another source of stress for plants (Kumar et al. 2019), and both Cadmium and Zinc are important contaminants in shoreline areas. Presence of the latter metal reduces the endogenous CKs concentration in Kosteletzkya pentacarpos whereas, conversely, the addition of exogenous CKs improves plant growth, stomatal conductance, net photosynthesis, total ascorbate and reduced oxidative stress in Zn-treated plants maintained in the absence of NaCl. The presence of Cd in addition to Zn induces specific physiological responses involving, in particular, increased levels of ABA (Zhou et al. 2019). The presence of salt has severe impact on senescing PGR, independent of heavy metal pollution (Albacete et al. 2009; Lutts and Lefèvre 2015; Rivero et al. 2009; Singh et al. 2018). Finally, the addition of exogenous CKs also increases plant resistance to heavy metals in Lupinus termis (Gadallah and El-Enany 1999), Pisum sativum (Al-Hakimi 2007), Zea mays (Lukatkin et al. 2007), Alyssum murale (Cassina et al. 2011), Helianthus annuus (Cassina et al. 2012), Solanum melongena (Singh and Prasad 2014) and S. lycopersicum (Singh et al. 2018).
Conclusions and perspectives
Our understanding of the CK biosynthesis and signaling pathways has improved substantially over the last several years. However, some of the roles CKs play, especially during the early stages of cell fate reprogramming and the induction of somatic embryogenesis, are still elusive (Pernisová et al. 2018). This is due, in part, to the complexity of the set of structures with CK activity, where in particular the biosynthesis of aromatic CKs is largely unknown. It will be important to investigate how CK biosynthetic intermediates are intracellularly mobilized and whether specific transporters are required for this. Important questions to be addressed are whether multiple copies of PUPs and ENTs are involved in transport, whether are other transporters are involved, and in how far the CK transporters are regulated independently? (Kang et al. 2017).
Another important subject that needs more attention is the occurrence of CK isomers on monocotyledons and eudicotyledons, and their potentially different functions in various plant systems. Furthermore, CK distributions in individual cell compartments should be mapped in more detail, including in the apoplastic space (Zürcher et al. 2016), to understand the physiological role of these molecules. The use of new techniques, such as matrix-assisted laser desorption/ionization time-of-flight mass spectrometry imaging, could help to localize and quantify PGRs (Shiono et al. 2017). Since the vacuole appears to be a reservoir for storing CK conjugates, import and export from this organelle likely involve active transport systems; however no such transporters have been found to date.
Even though it is known that different copies of IPT genes are expressed throughout the Arabidopsis plant (Miyawaki et al. 2004, 2006; Takei et al. 2004b), the biosynthesis of the iP-ribosides is restricted to seedling roots (Gelová et al. 2017; Žd'árská et al. 2013). Together, these findings suggest active transport toward the shoot, but the physiological functions of this transport system remain to be explored.
Another emerging area of research addresses the potential crosstalk between different growth regulators. Elucidating the interaction between CKs, auxins, ABA, strigolactones and gibberellins, and how they affect the homeostasis of each other, will lead to a better comprehension of plant development. Since auxins control the biosynthesis of CKs and SL in the stem, it must be clarified if these growth regulators serve as second messengers for auxins (Domagalska and Leyser 2011).
Abbreviations
- HMBDP:
-
4-Hydroxy-3-methyl-2-(E)-butenyl diphosphate
- 6-BA:
-
6-Benciladenine
- ABA:
-
Abscisic acid
- APT:
-
Adenine phosphoribosyltransferase
- iPMP:
-
Adenosine-5′-monophosphate
- AHK:
-
Arabidopsis histidine kinase
- AHP:
-
Arabidopsis histidine-containing phosphotransfer protein
- ARR:
-
Arabidopsis response regulators
- ABC:
-
ATP-binding cassette
- ARF3:
-
AUXIN RESPONSE FACTOR3
- CHASE:
-
Cyclases/histidine kinases associated sensory extracellular
- CKX:
-
CYTOKININ OXIDASE/DEHYDROGENASE
- CRE1:
-
Cytokinin response1
- DZ:
-
Dihydrozeatin
- DMAPP:
-
Dimethylallyl pyrophosphate
- ER:
-
Endoplasmic reticulum
- ENTs:
-
Equilibrative nucleoside transporters
- GeBP/GPL:
-
Glabrous1 enhancer-binding protein
- HPt:
-
Histidine phosphotransfer
- AHK:
-
Hybrid his kinases
- iP:
-
Hydroxymethylbutenyl diphosphate isopentenyladenine
- iPRDP:
-
Isopentenyladenosine-5′-diphosphate
- iPRTP:
-
Isopentenyladenosine-5′-triphosphate
- IPT:
-
ISOPENTENYL TRANSFERASE
- KIN:
-
Kinetin
- LOG:
-
LONELY GUY
- mT:
-
meta-Topolin
- MEP:
-
Methylerythritol phosphate
- MVA:
-
Mevalonate
- Ot:
-
ortho-Topolin
- PUPs:
-
Purine permeases
- SPY:
-
Spindly
- SL:
-
Strigolactone
- UGT:
-
Uridine diphosphate glycosyltransferase
- WOL:
-
Wooden leg1
- Z:
-
Zeatin
- cZ:
-
cis-Zeatin
- tZ:
-
trans-Zeatin
- tZRPs:
-
tZ-Riboside 5′-phosphates
References
Akiyoshi DE, Klee H, Amasino RM et al (1984) T-DNA of Agrobacterium tumefaciens encodes an enzyme of cytokinin biosynthesis. Proc Natl Acad Sci (USA) 81:5994–5998. https://doi.org/10.1073/pnas.81.19.5994
Albacete A (2017) Quantification of cytokinin levels and responses in abiotic stresses. In: Dandekar T, Naseem M (eds) Auxins and cytokinins in plant biology: methods and protocols. Springer, New York, pp 101–111. https://doi.org/10.1007/978-1-4939-6831-2_8
Albacete A, Martínez-Aldújar C, Ghanem ME et al (2009) Rootstock-mediated changes in xylem ionic and hormonal status are correlated with delayed leaf senescence, and increased leaf area and crop productivity in salinized tomato. Plant Cell Environ 32:928–938. https://doi.org/10.1111/j.1365-3040.2009.01973.x
Aldesuquy H, Baka Z, Mickky B (2014) Kinetin and spermine mediated induction of salt tolerance in wheat plants: leaf area, photosynthesis and chloroplast ultrastructure of flag leaf at ear emergence. Egypt J Basic Appl Sci 1:77–87. https://doi.org/10.1016/j.ejbas.2014.03.002
Al-Hakimi AMA (2007) Modification of cadmium toxicity in pea seedlings by kinetin. Plant Soil Environ 53:129–135
Allen M, Qin W, Moreau F, Moffatt B (2002) Adenine phosphoribosyltransferase isoforms of Arabidopsis and their potential contributions to adenine and cytokinin metabolism. Physiol Plant 115:56–68. https://doi.org/10.1034/j.1399-3054.2002.1150106.x
Anantharaman V, Aravind L (2001) The CHASE domain: a predicted ligand-binding module in plant cytokinin receptors and other eukaryotic and bacterial receptors. Trends Biochem Sci 26:579–582. https://doi.org/10.1016/S0968-0004(01)01968-5
Aremu A, Placková L, Bairu M et al (2014) Endogenous cytokinin profiles of tissue-cultured and acclimatized ´Williams´ bananas subjected to different aromatic cytokinin treatments. Plant Sci 214:88–98. https://doi.org/10.1016/j.plantsci.2013.09.012
Argueso CT, Raines T, Kieber JJ (2010) Cytokinin signaling and transcriptional networks. Curr Opin Plant Biol 13:533–539. https://doi.org/10.1016/j.pbi.2010.08.006
Argyros RD, Mathews DE, Chiang YH et al (2008) Type B response regulators of Arabidopsis play key roles in cytokinin signaling and plant development. Plant Cell 20:2102–2116. https://doi.org/10.1105/tpc.108.059584
Barry GF, Rogers SG, Fraley RT, Brand L (1984) Identification of a cloned cytokinin biosynthesis gene. Proc Natl Acad Sci (USA) 81:4776–4780. https://doi.org/10.1073/pnas.81.15.4776
Bartrina I, Jensen H, Novák O et al (2017) Gain-of-function mutants of the cytokinin receptors AHK2 and AHK3 regulate plant organ size, flowering time and plant longevity. Plant Physiol 173:1783–1797. https://doi.org/10.1104/pp.16.01903
Beinsberger SEI, Valcke RLM, Deblaere RY et al (1991) Effects of the introduction of Agrobacterium tumefaciens T-DNA ipt gene in Nicotiana tabacum L. cv. Petit havana SR1 plant cells. Plant Cell Physiol 32:489–496. https://doi.org/10.1093/oxfordjournals.pcp.a078106
Beveridge CA, Murfet IC, Kerhoas L et al (1997) The shoot controls zeatin riboside export from pea roots. Evidence from the branching mutant rms4. Plant J 11:339–345. https://doi.org/10.1046/j.1365-313X.1997.11020339.x
Bielach A, Podlesáková K, Marhavy P et al (2012) Spatiotemporal regulation of lateral root organogenesis in Arabidopsis by cytokinin. Plant Cell 24:3967–3981. https://doi.org/10.1105/tpc.112.103044
Bilyeu KD, Cole JL, Laskey JG et al (2001) Molecular and biochemical characterization of a cytokinin oxidase from maize. Plant Physiol 125:378–386. https://doi.org/10.1104/pp.125.1.378
Bishopp A, Lehesranta S, Vatén A et al (2011) Phloem-transported cytokinin regulates polar auxin transport and maintains vascular pattern in the root meristem. Curr Biol 21:927–932. https://doi.org/10.1016/j.cub.2011.04.049
Blackwell JR, Horgan R (1994) Cytokinin biosynthesis by extracts of Zea mays. Phytochemistry 35:339–342. https://doi.org/10.1016/S0031-9422(00)94760-5
Brenner WG, Ramireddy E, Heyl A, Schmülling T (2012) Gene regulation by cytokinin in Arabidopsis. Front Plant Sci 3:8. https://doi.org/10.3389/fpls.2012.00008
Brownlee BG, Hall RH, Whitty CD (1975) 3-methyl-2-butenal: an enzymatic degradation product of the cytokinin, N6-(∆2-isopentenyl) adenine. Can J Biochem 53:37–41
Brzobohaty B, Moore I, Kristoffersen P et al (1993) Release of active cytokinin by a ∆-glucosidase localized to the maize root meristem. Science 262:1051–1054. https://doi.org/10.1126/science.8235622
Bürkle L, Cedzich A, Döpke C et al (2003) Transport of cytokinins mediated by purine transporters of the PUP family expressed in phloem, hydathodes, and pollen of Arabidopsis. Plant J 34:13–26. https://doi.org/10.1046/j.1365-313X.2003.01700.x
Caesar K, Thamm AM, Witthöft J et al (2011) Evidence for the localization of the Arabidopsis cytokinin receptors AHK3 and AHK4 in the endoplasmic reticulum. J Exp Bot 62:5571–5580. https://doi.org/10.1093/jxb/err238
Cai L, Zhang L, Fu Q, Xu Z-F (2018) Identification and expression analysis of cytokinin metabolic genes IPTs, CYP735A and CKXs in the biofuel plant Jatropha curcas. PeerJ 6:e4812. https://doi.org/10.7717/peerj.4812
Cao ZY, Ma YN, Sun LH et al (2017) Direct determination of six cytokinin nucleotide monophosphates in coconut flesh by reversed-phase liquid chromatography-tandem mass spectrometry. J Agric Food Chem 65:9909–9915. https://doi.org/10.1021/acs.jafc.7b03798
Cassina L, Tassi E, Morelli E et al (2011) Exogenous cytokinin treatments of an Ni hyper-accumulator, Alyssum murale, grown in a serpentine soil: implications for phytoextraction. Int J Phytoremediation 13:90–101. https://doi.org/10.1080/15226514.2011.568538
Cassina L, Tassi E, Pedron F et al (2012) Using a plant hormone and a thioligand to improve phytoremediation of Hg-contaminated soil from a petrochemical plant. J Hazard Mater 231:36–42. https://doi.org/10.1016/j.jhazmat.2012.06.031
Chen C (1997) Cytokinin biosynthesis and interconversion. Physiol Plant 101:665–673. https://doi.org/10.1111/j.1399-3054.1997.tb01051.x
Chen CM, Kristopeit SM (1981) Metabolism of cytokinin: deribosylation of cytokinin ribonucleoside by adenosine nucleosidase from wheat germ cells. Plant Physiol 68:1020–1023. https://doi.org/10.1104/pp.68.5.1020
Chen P, Jiao X, Zhang Y et al (2019) The crystal structure of the phytopathogenic bacterial sensor PcrK reveals different cytokinin recognition mechanism from the plant sensor AHK4. J Struct Biol 208:69–76. https://doi.org/10.1016/j.jsb.2019.08.001
Cheng ZJ, Wang L, Sun W et al (2013) Pattern of auxin and cytokinin responses for shoot meristem induction results from the regulation of cytokinin biosynthesis by AUXIN RESPONSE FACTOR3. Plant Physiol 161:240–251. https://doi.org/10.1104/pp.112.203166
Corbesier L, Prinsen E, Jacqmard A et al (2003) Cytokinin levels in leaves, leaf exudate and shoot apical meristem of Arabidopsis thaliana during floral transition. J Exp Bot 54:2511–2517. https://doi.org/10.1093/jxb/erg276
Corot A, Roman H, Douillet O et al (2017) Cytokinins and abscisic acid act antagonistically in the regulation of the bud outgrowth pattern by light intensity. Front Plant Sci 8:1724
Davière J-M, Achard P (2017) Organ communication: cytokinins on the move. Nat Plants 17:116. https://doi.org/10.1038/nplants.2017.116
Davies WJ, Zhang J (1991) Root signals and the regulation of growth and development of plants in drying soil. Annu Rev Plant Physiol Plant Mol Biol 42:55–76. https://doi.org/10.1146/annurev.pp.42.060191.000415
Dobrev PI, Hoyerová K, Petrášek J (2017) Analytical determination of auxins and cytokinins. In: Dandekar T, Naseem M (eds) Auxins and cytokinins in plant biology: methods and protocols. Springer, New York, pp 31–39. https://doi.org/10.1007/978-1-4939-6831-2_2
Domagalska MA, Leyser O (2011) Signal integration in the control of shoot branching. Nat Rev Mol Cell Bio 12:211. https://doi.org/10.1038/nrm3088
Dun EA, de Saint GA, Rameau C, Beveridge CA (2012) Antagonistic action of strigolactone and cytokinin in bud outgrowth control. Plant Physiol 158:487–498. https://doi.org/10.1104/pp.111.186783
Durán-Medina Y, Díaz-Ramírez D, Marsch-Martínez N (2017) Cytokinins on the move. Front Plant Sci 8:146. https://doi.org/10.3389/fpls.2017.00146
Entsch B, Letham D (1979) Enzymic glucosylation of the cytokinin, 6-benzylaminopurine. Plant Sci Lett 14:205–212. https://doi.org/10.1016/0304-4211(79)90061-0
Entsch B, Parker CW, Letham DS, Summons RE (1979) Preparation and characterization, using high-performance liquid chromatography of an enzyme forming glucosides of cytokinins. Biochim Biophys Acta Enzymol 570:124–139. https://doi.org/10.1016/0005-2744(79)90207-9
Entsch B, Parker C, Letham D (1983) An enzyme from lupin seeds forming alanine derivatives of cytokinins. Phytochemistry 22:375–381. https://doi.org/10.1016/0031-9422(83)83008-8
Falk A, Rask L (1995) Expression of a zeatin-O-glucoside-degrading [beta]-glucosidase in Brassica napus. Plant Physiol 108:1369–1377. https://doi.org/10.1104/pp.108.4.1369
Ferreira FJ, Kieber JJ (2005) Cytokinin signaling. Curr Opin Plant Biol 8:518–525. https://doi.org/10.1016/j.pbi.2005.07.013
Foo E, Morris SE, Parmenter K et al (2007) Feedback regulation of xylem cytokinin content Is conserved in pea and Arabidopsis. Plant Physiol 143:1418–1428. https://doi.org/10.1104/pp.106.093708
Fusconi A (2013) Regulation of root morphogenesis in arbuscular mycorrhizae: what role do fungal exudates, phosphate, sugars and hormones play in lateral root formation? Ann Bot 113:19–33. https://doi.org/10.1093/aob/mct258
Gadallah MAA, El-Enany AE (1999) Role of kinetin in alleviation of copper and zinc toxicity in Lupinus termis plants. Plant Growth Regul 29:151–160. https://doi.org/10.1023/A:1006245628188
Galichet A, Hoyerová K, Kamínek M, Gruissem W (2008) Farnesylation directs AtIPT3 subcellular localization and modulates cytokinin biosynthesis in Arabidopsis. Plant Physiol 146:1155. https://doi.org/10.1104/pp.107.107425
Galuszka P, Frébort I, Šebela M et al (2001) Cytokinin oxidase or dehydrogenase? Mechanism of cytokinin degradation in cereals. Eur J Biochem 268:450–461. https://doi.org/10.1046/j.1432-1033.2001.01910.x
Galuszka P, Frébortová J, Werner T et al (2004) Cytokinin oxidase/dehydrogenase genes in barley and wheat: cloning and heterologous expression. Eur J Biochem 271:3990–4002. https://doi.org/10.1111/j.1432-1033.2004.04334.x
Gelová Z, ten Hoopen P, Novák O et al (2017) Antibody-mediated modulation of cytokinins in tobacco: organ-specific changes in cytokinin homeostasis. J Exp Bot 69:441–454. https://doi.org/10.1093/jxb/erx426
Ghosh A, Shah Md, Jui ZS et al (2018) Evolutionary variation and expression profiling of isopentenyl transferase gene family in Arabidopsis thaliana L. and Oryza sativa L. Plant Gene 15:15–27. https://doi.org/10.1016/j.plgene.2018.06.002
Gillissen B, Bürkle L, André B et al (2000) A new family of high-affinity transporters for adenine, cytosine, and purine derivatives in Arabidopsis. Plant Cell 12:291–300. https://doi.org/10.1105/tpc.12.2.291
Girke C, Daumann M, Niopek-Witz S, Möhlmann T (2014) Nucleobase and nucleoside transport and integration into plant metabolism. Front Plant Sci 5:443. https://doi.org/10.3389/fpls.2014.00443
Gordon SP, Chickarmane VS, Ohno C, Meyerowitz EM (2009) Multiple feedback loops through cytokinin signaling control stem cell number within the Arabidopsis shoot meristem. Proc Natl Acad Sci (USA) 106:16529–16534. https://doi.org/10.1073/pnas.0908122106
Greenboim-Wainberg Y, Maymon I, Borochov R et al (2005) Cross talk between gibberellin and cytokinin: the Arabidopsis GA response inhibitor SPINDLY plays a positive role in cytokinin signaling. Plant Cell 17:92–102. https://doi.org/10.1105/tpc.104.028472
Grefen C, Harter K (2004) Plant two-component systems: principles, functions, complexity and cross talk. Planta 219:733–742. https://doi.org/10.1007/s00425-004-1316-4
Guttikonda SK, Trupti J, Bisht NC et al (2010) Whole genome co-expression analysis of soybean cytochrome P450 genes identifies nodulation-specific P450 monooxygenases. BMC Plant Biol 10:243. https://doi.org/10.1186/1471-2229-10-243
Han Y, Zhang C, Yang H, Jiao Y (2014) Cytokinin pathway mediates APETALA1 function in the establishment of determinate floral meristems in Arabidopsis. Proc Natl Acad Sci (USA) 111:6840. https://doi.org/10.1073/pnas.1318532111
Hansen CE, Meins F, Milani A (1985) Clonal and physiological variation in the cytokinin content of tobacco-cell lines differing in cytokinin requirement and capacity for neoplastic growth. Differentiation 29:1–6. https://doi.org/10.1111/j.1432-0436.1985.tb00284.x
Hansen CE, Meins F, Aebi R (1987) Hormonal regulation of zeatin-riboside accumulation by cultured tobacco cells. Planta 172:520–525. https://doi.org/10.1007/BF00393869
Heyl A, Wulfetange K, Pils B et al (2007) Evolutionary proteomics identifies amino acids essential for ligand-binding of the cytokinin receptor CHASE domain. BMC Evol Biol 7:62–72. https://doi.org/10.1186/1471-2148-7-62
High KE, Ashton P, Nelson M et al (2019) New approaches using mass spectrometry to investigate changes to cytokinin and abscisic acid (ABA) concentrations in soil. Soil Biol Biochem 135:108–116. https://doi.org/10.1016/j.soilbio.2019.04.017
Higuchi M, Pischke MS, Mähönen AP et al (2004) In planta functions of the Arabidopsis cytokinin receptor family. Proc Natl Acad Sci (USA) 101:8821–8826. https://doi.org/10.1073/pnas.0402887101
Hirose N, Makita N, Yamaya T, Sakakibara H (2005) Functional characterization and expression analysis of a gene, OsENT2, encoding an equilibrative nucleoside transporter in rice suggest a function in cytokinin transport. Plant Physiol 138:196–206. https://doi.org/10.1104/pp.105.060137
Hirose N, Takei K, Kuroha T et al (2008) Regulation of cytokinin biosynthesis, compartmentalization and translocation. J Exp Bot 59:75–83. https://doi.org/10.1093/jxb/erm157
Hou B, Lim EK, Higgins GS, Bowles DJ (2004) N-glucosylation of cytokinins by glycosyltransferases of Arabidopsis thaliana. J Biol Chem 279:47822–47832. https://doi.org/10.1074/jbc.M409569200
Hutchison CE, Kieber JJ (2002) Cytokinin signaling in Arabidopsis. Plant Cell 14:S47–S59. https://doi.org/10.1105/tpc.010444
Hutchison CE, Li J, Argueso C et al (2006) The Arabidopsis histidine phosphotransfer proteins are redundant positive regulators of cytokinin signaling. Plant Cell 18:3073–3087. https://doi.org/10.1105/tpc.106.045674
Hwang I, Sheen J, Müller B (2012) Cytokinin signaling networks. Annu Rev Plant Biol 63:353–380. https://doi.org/10.1146/annurev-arplant-042811-105503
Inoue T, Higuchi M, Hashimoto Y et al (2001) Identification of CRE1 as a cytokinin receptor from Arabidopsis. Nature 409:1060–1063. https://doi.org/10.1038/35059117
Ishida K, Yamashino T, Yokoyama A, Mizuno T (2008) Three type-B response regulators, ARR1, ARR10 and ARR12, play essential but redundant roles in cytokinin signal transduction throughout the life cycle of Arabidopsis thaliana. Plant Cell Physiol 49:47–57. https://doi.org/10.1093/pcp/pcm165
Jin SH, Ma XM, Kojima M et al (2013) Overexpression of glucosyltransferase UGT85A1 influences trans-zeatin homeostasis and trans-zeatin responses likely through O-glucosylation. Planta 237:991–999. https://doi.org/10.1007/s00425-012-1818-4
Jiskrová E, Novák O, Pospísilová H et al (2016) Extra-and intracellular distribution of cytokinins in the leaves of monocots and dicots. New Biotechnol 33:735–742. https://doi.org/10.1016/j.nbt.2015.12.010
Jones B, Gunneras SA, Petersson SV et al (2010) Cytokinin regulation of auxin synthesis in Arabidopsis involves a homeostatic feedback loop regulated via auxin and cytokinin signal transduction. Plant Cell 22:2956–2969. https://doi.org/10.1105/tpc.110.074856
Jung B, Flörchinger M, Traub M et al (2009) Uridine-ribohydrolase is a key regulator in the uridine degradation pathway of Arabidopsis. Plant Cell 21:876–891. https://doi.org/10.1105/tpc.108.062612
Kakimoto T (1996) CKI1, a histidine kinase homolog implicated in cytokinin signal transduction. Science 274:982–985. https://doi.org/10.1126/science.274.5289.982
Kakimoto T (2001) Identification of plant cytokinin biosynthetic enzymes as dimethylallyl diphosphate: ATP/ADP isopentenyltransferases. Plant Cell Physiol 42:677–685. https://doi.org/10.1093/pcp/pce112
Kang J, Park J, Choi H et al (2011) Plant ABC transporters. Arabidopsis Book 9:e0153. https://doi.org/10.1199/tab.0153
Kang J, Lee Y, Sakakibara H, Martinoia E (2017) Cytokinin transporters: GO and STOP in signaling. Trends Plant Sci 22:455–461. https://doi.org/10.1016/j.tplants.2017.03.003
Kasahara H, Takei K, Ueda N et al (2004) Distinct isoprenoid origins of cis-and trans-zeatin biosyntheses in Arabidopsis. J Biol Chem 279:14049–14054. https://doi.org/10.1074/jbc.M314195200
Kiba T, Krapp A (2016) Plant nitrogen acquisition under low availability: regulation of uptake and root architecture. Plant Cell Physiol 57:707–714. https://doi.org/10.1093/pcp/pcw052
Kiba T, Kudo T, Kojima M, Sakakibara H (2010) Hormonal control of nitrogen acquisition: roles of auxin, abscisic acid, and cytokinin. J Exp Bot 62:1399–1409. https://doi.org/10.1093/jxb/erq410
Kiba T, Takei K, Kojima M, Sakakibara H (2013) Side-chain modification of cytokinins controls shoot growth in Arabidopsis. Dev Cell 27:452–461. https://doi.org/10.1016/j.devcel.2013.10.004
Kieber JJ, Schaller GE (2018) Cytokinin signaling in plant development. Development 145:dev149344. https://doi.org/10.1242/dev.149344
Kim HJ, Chiang YH, Kieber JJ, Schaller GE (2013a) SCFKMD controls cytokinin signaling by regulating the degradation of type-B response regulators. Proc Natl Acad Sci (USA) 110:10028–10033. https://doi.org/10.1073/pnas.1300403110
Kim HJ, Kieber JJ, Schaller GE (2013b) The rice F-box protein KISS ME DEADLY2 functions as a negative regulator of cytokinin signalling. Plant Signal Behav 8:e26434. https://doi.org/10.4161/psb.26434
Kiran NS, Polanská L, Fohlerová R et al (2006) Ectopic over-expression of the maize ∆-glucosidase Zm-p60.1 perturbs cytokinin homeostasis in transgenic tobacco. J Exp Bot 57:985–996. https://doi.org/10.1093/jxb/erj084
Kiran N, Benková E, Reková A et al (2012) Retargeting a maize ∆-glucosidase to the vacuole-evidence from intact plants that zeatin-O-glucoside is stored in the vacuole. Phytochemistry 79:67–77. https://doi.org/10.1016/j.phytochem.2012.03.012
Ko D, Helariutta Y (2017) Shoot-root communication in flowering plants. Curr Biol 27:R973–R978. https://doi.org/10.1016/j.cub.2017.06.054
Ko D, Kang J, Kiba T et al (2014) Arabidopsis ABCG14 is essential for the root-to-shoot translocation of cytokinin. Proc Natl Acad Sci (USA) 111:7150–7155. https://doi.org/10.1073/pnas.1321519111
Koda Y, Okasawa Y (1980) Cytokinin production by Asparagus shoot apex cultured in vitro. Physiol Plant 49:193–197. https://doi.org/10.1111/j.1399-3054.1980.tb02651.x
Köllmer I, Novák O, Strnad M et al (2014) Overexpression of the cytosolic cytokinin oxidase/dehydrogenase (CKX7) from Arabidopsis causes specific changes in root growth and xylem differentiation. Plant J 78:359–371. https://doi.org/10.1111/tpj.12477
Kopecná M, Blaschke H, Kopecný D et al (2013) Structure and function of nucleoside hydrolases from Physcomitrella patens and maize catalyzing the hydrolysis of purine, pyrimidine, and cytokinin ribosides. Plant Physiol 163:1568–1583. https://doi.org/10.1104/pp.113.228775
Krall L, Raschke M, Zenk MH, Baron C (2002) The Tzs protein from Agrobacterium tumefaciens C58 produces zeatin riboside 5'-phosphate from 4-hydroxy-3-methyl-2-(E)-butenyl diphosphate and AMP. FEBS Lett 527:315–318. https://doi.org/10.1016/S0014-5793(02)03258-1
Kumar V, Sharma A, Kaur P et al (2019) Pollution assessment of heavy metals in soils of India and ecological risk assessment: a state-of-the-art. Chemosphere 216:449–462. https://doi.org/10.1016/j.chemosphere.2018.10.066
Kurakawa T, Ueda N, Maekawa M et al (2007) Direct control of shoot meristem activity by a cytokinin-activating enzyme. Nature 445:652–655. https://doi.org/10.1038/nature05504
Kuroha T, Tokunaga H, Kojima M et al (2009) Functional analyses of LONELY GUY cytokinin-activating enzymes reveal the importance of the direct activation pathway in Arabidopsis. Plant Cell 21:3152–3169. https://doi.org/10.1105/tpc.109.068676
LaCuesta M, Saiz-Fernández I, Podlesáková K et al (2018) The trans and cis zeatin isomers play different roles in regulating growth inhibition induced by high nitrate concentrations in maize. Plant Growth Regul 85:199–209. https://doi.org/10.1007/s10725-018-0383-7
Lairson LL, Henrissat B, Davies GJ, Withers SG (2008) Glycosyltransferases: structures, functions, and mechanisms. Annu Rev Biochem 77:521–555. https://doi.org/10.1146/annurev.biochem.76.061005.092322
Landrein B, Formosa-Jordan P, Malivert A et al (2018) Nitrate modulates stem cell dynamics in Arabidopsis shoot meristems through cytokinins. Proc Natl Acad Sci (USA) 15:1382–1387. https://doi.org/10.1073/pnas.1718670115
Leyser O (2018) Auxin signaling. Plant Physiol 176:465–479. https://doi.org/10.1104/pp.17.00765
Li G, Liu K, Baldwin SA, Wang D (2003) Equilibrative nucleoside transporters of Arabidopsis thaliana cDNA cloning, expression pattern, and analysis of transport activities. J Biol Chem 278:35732–35742. https://doi.org/10.1074/jbc.M304768200
Li YJ, Wang B, Dong RR, Hou BK (2015) AtUGT76C2, an Arabidopsis cytokinin glycosyltransferase is involved in drought stress adaptation. Plant Sci 236:157–167. https://doi.org/10.1016/j.plantsci.2015.04.002
Lim E-K, Bowles D (2004) A class of plant glycosyltransferases involved in cellular homeostasis. EMBO J 23:2915–2922. https://doi.org/10.1038/sj.emboj.7600295
Liu CJ, Zhao Y, Zhang K (2019) Cytokinin transporters: multisite players in cytokinin homeostasis and signal distribution. Front Plant Sci 10:693. https://doi.org/10.3389/fpls.2019.00693
Lomin SN, Myakushina YA, Arkhipov DV et al (2018) Studies of cytokinin receptor–phosphotransmitter interaction provide evidences for the initiation of cytokinin signalling in the endoplasmic reticulum. Funct Plant Biol 45:192–202. https://doi.org/10.1071/FP16292
Lukatkin AS, Gracheva NV, Grishenkova NN et al (2007) Cytokinin-like growth regulators mitigate toxic action of zinc and nickel ions on maize seedlings. Russ J Plant Physiol 54:381–387. https://doi.org/10.1134/S1021443707030132
Lutts S, Lefèvre I (2015) How can we take advantage of halophyte properties to cope with heavy metal toxicity in salt-affected areas? Ann Bot 115:509–528. https://doi.org/10.1093/aob/mcu264
Manandhar S, Funnell KA, Woolley DJ, Cooney J (2018) Interaction between strigolactone and cytokinin on axillary and adventitious bud development in Zantedeschia. J Plant Physiol Pathol 6:1. https://doi.org/10.4172/2329-955X.1000172
Markovich O, Steiner E, Kouril S et al (2017) Silicon promotes cytokinin biosynthesis and delays senescence in Arabidopsis and sorghum. Plant Cell Environ 40:1189–1196. https://doi.org/10.1111/pce.12913
Matsumoto-Kitano M, Kusumoto T, Tarkowski P et al (2008) Cytokinins are central regulators of cambial activity. Proc Natl Acad Sci (USA) 105:20027–20031. https://doi.org/10.1073/pnas.0805619105
Matsuo S, Kikuchi K, Fukuda M et al (2012) Roles and regulation of cytokinins in tomato fruit development. J Exp Bot 63:5569–5579. https://doi.org/10.1093/jxb/ers207
Miller CO (1961) A kinetin-like compund in maize. Proc Natl Acad Sci (USA) 47:170–174. https://doi.org/10.1073/pnas.47.2.170
Miller CO, Skoog F, Okumura FS et al (1955a) Structure and synthesis of kinetin. J Am Chem Soc 77:2662–2663. https://doi.org/10.1021/ja01614a108
Miller CO, Skoog F, Von Saltza MH, Strong FM (1955b) Kinetin, a cell division factor from deoxyribonucleic acid. J Am Chem Soc 77:1392. https://doi.org/10.1021/ja01610a105
Miyawaki K, Matsumoto-Kitano M, Kakimoto T (2004) Expression of cytokinin biosynthetic isopentenyltransferase genes in Arabidopsis: tissue specificity and regulation by auxin, cytokinin, and nitrate. Plant J 37:128–138. https://doi.org/10.1046/j.1345-313X.2003.01945.x
Miyawaki K, Tarkowski P, Matsumoto-Kitano M et al (2006) Roles of Arabidopsis ATP/ADP isopentenyltransferases and tRNA isopentenyltransferases in cytokinin biosynthesis. Proc Natl Acad Sci (USA) 103:16598–16603. https://doi.org/10.1073/pnas.0603522103
Moffatt B, Pethe C, Laloue M (1991) Metabolism of benzyladenine is impaired in a mutant of Arabidopsis thaliana lacking adenine phosphoribosyltransferase activity. Plant Physiol 95:900–908. https://doi.org/10.1104/pp.95.3.900
Mok DWS, Mok MC (2001) Cytokinin metabolism and action. Annu Rev Plant Physiol Plant Mol Biol 52:89–118. https://doi.org/10.1146/annurev.arplant.52.1.89
Mok M, Mok D, Arsden KE, Shaw G (1987) The biological activity and metabolism of a novel cytokinin metabolite, O-xylosylzeatin, in callus tissue of Phaseolus vulgaris and P. lunatus. J Plant Physiol 130:423–431. https://doi.org/10.1016/S0176-1617(87)80207-9
Mok DWS, Martin RC, Shan X, Mok MC (2000) Genes encoding zeatin O-glycosyltransferases. Plant Growth Regul 32:285–287. https://doi.org/10.1023/A:1010712102890
Morris SE, Turnbull CGN, Murfet IC, Beveridge CA (2001) Mutational analysis of branching in pea. Evidence that Rms1 and Rms5 regulate the same novel signal. Plant Physiol 126:1205–1213. https://doi.org/10.1104/pp.126.3.1205
Mortier V, Wasson A, Jaworek P et al (2014) Role of LONELY GUY genes in indeterminate nodulation on Medicago truncatula. New Phytol 202:582–593. https://doi.org/10.1111/nph.12681
Mougel C, Zhulin IB (2001) CHASE: an extracellular sensing domain common to transmembrane receptors from prokaryotes, lower eukaryotes and plants. Trends Biochem Sci 26:582–584. https://doi.org/10.1016/S0968-0004(01)01969-7
Müller B, Sheen J (2007a) Advances in cytokinin signaling. Science 318:68–69. https://doi.org/10.1126/science.1145461
Müller B, Sheen J (2007b) Arabidopsis cytokinin signaling pathway. Science's STKE 2007:5
Müller B, Sheen J (2008) Cytokinin and auxin interaction in root stem-cell specification during early embryogenesis. Nature 453:1094–1097. https://doi.org/10.1038/nature06943
Müller D, Waldie T, Miyawaki K et al (2015) Cytokinin is required for escape but not release from auxin mediated apical dominance. Plant J 82:874–886. https://doi.org/10.1111/tpj.12862
Murai M (1994) Cytokinin biosynthesis in tRNA and cytokinin incorporation into plant RNA. In: Mok DWS, Mok MC (eds) Cytokinins, chemistry, activity, and function. CRC Press, Boca Raton, Fl, pp 87–99
Nelson D, Werck-Reichhart D (2011) A P450-centric view of plant evolution. Plant J 66:194–211. https://doi.org/10.1111/j.1365-313X.2011.04529.x
Nic-Can GI, Avilez-Montalvo JR, Avilez-Montalvo RN et al. (2016) The relationship between stress and somatic embryogenesis. In: Loyola-Vargas VM, Ochoa-Alejo N (eds) Somatic embryogenesis Fundamental aspects and applications. Springer, Switzerland, pp 151–170. https://doi.org/10.1007/978-3-319-33705-0_9
Niemann MCE, Weber H, Hluska T et al (2018) The cytokinin oxidase/dehydrogenase CKX1 Is a membrane-bound protein requiring homooligomerization in the endoplasmic reticulum for its cellular activity. Plant Physiol 176:2024–2039. https://doi.org/10.1104/pp.17.00925
Nishii K, Wright F, Chen YYu, Möller M (2018) Tangled history of a multigene family: the evolution of ISOPENTENYL TRANSFERASE genes. PLoS ONE 13:e0201198. https://doi.org/10.1371/journal.pone.0201198
Nishiyama R, Watanabe Y, Fujita Y et al (2011) Analysis of cytokinin mutants and regulation of cytokinin metabolic genes reveals important regulatory roles of cytokinins in drought, salt and abscisic acid responses, and abscisic acid biosynthesis. Plant Cell 23:2169–2183. https://doi.org/10.1105/tpc.111.087395
Nitschke S, Cortleven A, Iven T et al (2016) Circadian stress regimes affect the circadian clock and cause jasmonic acid-dependent cell death in cytokinin-deficient Arabidopsis plants. Plant Cell 28:1616–1639. https://doi.org/10.1105/tpc.16.00016
Nongpiur R, Soni P, Karan R et al (2012) Histidine kinases in plants: cross talk between hormone and stress responses. Plant Signal Behav 7:1230–1237. https://doi.org/10.4161/psb.21516
Nordström A, Tarkowski P, Tarkowska D et al (2004) Auxin regulation of cytokinin biosynthesis in Arabidopsis thaliana: a factor of potential importance for auxin–cytokinin-regulated development. Proc Natl Acad Sci (USA) 101:8039–8044. https://doi.org/10.1073/pnas.0402504101
Noriega X, Pérez FJ (2017) ABA biosynthesis genes are down-regulated while auxin and cytokinin biosynthesis genes are up-regulated during the release of grapevine buds from endodormancy. J Plant Growth Regul 36:814–823. https://doi.org/10.1007/s00344-017-9685-7
Osugi A, Sakakibara H (2015) How do plants respond to cytokinins and what is their importance? BMC Biol 13:1–10. https://doi.org/10.1186/s12915-015-0214-5
Osugi A, Kojima M, Takebayashi Y et al (2017) Systemic transport of trans-zeatin and its precursor have differing roles in Arabidopsis shoots. Nat Plants 3:17112. https://doi.org/10.1038/nplants.2017.112
Park J, Lee Y, Martinoia E, Geisler M (2017) Plant hormone transporters: what we know and what we would like to know. BMC Biol 15:93. https://doi.org/10.1186/s12915-017-0443-x
Pernisová M, Grochova M, Konecny T et al (2018) Cytokinin signalling regulates organ identity via the AHK4 receptor in Arabidopsis. Development 145:dev163907. https://doi.org/10.1242/dev.163907
Persson BC, Esberg B, Ólafsson Ó, Björk GR (1994) Synthesis and function of isopentenyl adenosine derivatives in tRNA. Biochimie 76:1152–1160. https://doi.org/10.1016/0300-9084(94)90044-2
Poitout A, Crabos A, Petrick I et al (2018) Responses to systemic nitrogen signaling in Arabidopsis roots involve trans-zeatin in shoots. Plant Cell 30:1243–1257. https://doi.org/10.1105/tpc.18.00011
Qi Z, Xiong L (2013) Characterization of a Purine Permease Family Gene OsPUP7 Involved in Growth and Development Control in Rice. J Int Plant Biol 55:1119–1135. https://doi.org/10.1111/jipb.12101
Rahayu YS, Walch-Liu P, Neumann G et al (2005) Root-derived cytokinins as long-distance signals for NO3-induced stimulation of leaf growth. J Exp Bot 56:1143–1152. https://doi.org/10.1093/jxb/eri107
Reid DE, Heckmann AB, Novak O et al (2016) CYTOKININ OXIDASE/DEHYDROGENASE3 maintains cytokinin homeostasis during root and nodule development in Lotus japonicus. Plant Physiol 170:1060–1074. https://doi.org/10.1104/pp.15.00650
Rivero RM, Shulaev V, Blumwald E (2009) Cytokinin-dependent photorespiration and the protection of photosynthesis during water deficit. Plant Physiol 150:1530. https://doi.org/10.1104/pp.109.139378
Rivero RM, Gimeno J, van Deynze A et al (2010) Enhanced cytokinin synthesis in tobacco plants expressing PSARK: IPT prevents the degradation of photosynthetic protein complexes during drought. Plant Cell Physiol 51:1929–1941. https://doi.org/10.1093/pcp/pcq143
Rowe JH, Topping JF, Liu J, Lindsey K (2016) Abscisic acid regulates root growth under osmotic stress conditions via an interacting hormonal network with cytokinin, ethylene and auxin. New Phytol 211:225–239. https://doi.org/10.1111/nph.13882
Ruffel S, Krouk G, Ristova D et al (2011) Nitrogen economics of root foraging: transitive closure of the nitrate-cytokinin relay and distinct systemic signaling for N supply vs. demand. Proc Natl Acad Sci (USA) 108:18524–18529. https://doi.org/10.1073/pnas.1108684108
Sáenz L, Jones LH, Oropeza C et al (2003) Endogenous isoprenoid and aromatic cytokinins in different plant parts of Cocos nucifera (L.). Plant Growth Regul 39:205–215. https://doi.org/10.1023/A:1022851012878
Saito T, Opio P, Wang S et al. (2019) Association of auxin, cytokinin, abscisic acid, and plant peptide response genes during adventitious root formation in Marubakaido apple rootstock (Malus prunifolia Borkh. var. ringo Asami). Acta Physiol Plant 41:41. https://doi.org/10.1007/s11738-019-2827-8
Sakakibara H (2006) Cytokinins: activity, biosynthesis, and translocation. Annu Rev Plant Biol 57:431–449. https://doi.org/10.1146/annurev.arplant.57.032905.105231
Sakakibara H (2010) Cytokinin biosynthesis and metabolism. In: Davies PJ (ed) Plant hormones. Springer, Amsterdam, pp 95–114. https://doi.org/10.1007/978-1-4020-2686-7_5
Sakakibara H, Kasahara H, Ueda N et al (2005) Agrobacterium tumefaciens increases cytokinin production in plastids by modifying the biosynthetic pathway in the host plant. Proc Natl Acad Sci (USA) 102:9972–9977. https://doi.org/10.1073/pnas.0500793102
Salama AMS, El-D A, Wareing PF (1979) Effects of mineral nutrition on endogenous cytokinins in plant of sunflower (Helianthus annuus L.). J Exp Bot 30:971–981. https://doi.org/10.1093/jxb/30.5.971
Sasaki T, Suzaki T, Soyano T et al (2014) Shoot-derived cytokinins systemically regulate root nodulation. Nat Commun 5:4983. https://doi.org/10.1038/ncomms5983
Schäfer M, Brütting C, Meza-Canales ID et al (2015) The role of cis-zeatin-type cytokinins in plant growth regulation and mediating responses to environmental interactions. J Exp Bot 66:4873–4884. https://doi.org/10.1093/jxb/erv214
Schaller GE, Doi K, Hwang I et al. (2007) Nomenclature for two-component signaling elements of rice. Plant Physiol 143:555–557
Schaller GE, Shiu SH, Armitage JP (2011) Two-component systems and their co-option for eukaryotic signal transduction. Curr Biol 21:320–330. https://doi.org/10.1016/j.cub.2011.02.045
Schmülling T, Werner T, Riefler M et al (2003) Structure and function of cytokinin oxidase/dehydrogenase genes of maize, rice, Arabidopsis and other species. J Plant Res 116:241–252. https://doi.org/10.1007/s10265-003-0096-4
Séguéla M, Briat JF, Vert G, Curie C (2008) Cytokinins negatively regulate the root iron uptake machinery in Arabidopsis through a growth-dependent pathway. Plant J 55:289–300. https://doi.org/10.1111/j.1365-313X.2008.03502.x
Seo H, Kim KJ (2017) Structural basis for a novel type of cytokinin-activating protein. Sci Rep UK 7:45985. https://doi.org/10.1038/srep45985
Sheflin AM, Kirkwood JS, Wolfe LM et al (2019) High-throughput quantitative analysis of phytohormones in sorghum leaf and root tissue by ultra-performance liquid chromatography-mass spectrometry. Anal Bioanal Chem 411:4839–4848. https://doi.org/10.1007/s00216-019-01658-9
Shi Y, Tian S, Hou L et al (2012) Ethylene signaling negatively regulates freezing tolerance by repressing expression of CBF and type-A ARR genes in Arabidopsis. Plant Cell 24:2578–2595. https://doi.org/10.1105/tpc.112.098640
Shiono K, Hashizaki R, Nakanishi T et al (2017) Multi-imaging of cytokinin and abscisic acid on the roots of rice (Oryza sativa) using matrix-assisted laser desorption/ionization mass spectrometry. J Agric Food Chem 65:7624–7628. https://doi.org/10.1021/acs.jafc.7b02255
Simm S, Scharf KD, Jegadeesan S et al (2016) Survey of genes involved in biosynthesis, transport, and signaling of phytohormones with focus on Solanum lycopersicum. Bioinform Biol Insights 10:185–207. https://doi.org/10.4137/BBI.S38425
Singh S, Prasad SM (2014) Growth, photosynthesis and oxidative responses of Solanum melongena L. seedlings to cadmium stress: mechanism of toxicity amelioration by kinetin. Sci Hortic 176:1–10. https://doi.org/10.1016/j.scienta.2014.06.022
Singh S, Letham S, Zhang X, Alni L (1992) Cytokinin biochemistry in relation to leaf senescence VI. Effect of nitrogenous nutrients on cytokinin levels and senescence of tobacco leaves. Physiol Plant 84:262–268. https://doi.org/10.1111/j.1399-3054.1992.tb04663.x
Singh S, Singh A, Srivastava PK, Prasad SM (2018) Cadmium toxicity and its amelioration by kinetin in tomato seedlings vis-à-vis ascorbate-glutathione cycle. J Photochem Photobiol Biol 178:76–84. https://doi.org/10.1016/j.jphotobiol.2017.10.025
Skalicky V, Kubes M, Napier R, Novák O (2018) Auxins and cytokinins-the role of subcellular organization on homeostasis. Int J Mol Sci 19:3115. https://doi.org/10.3390/ijms19103115
Skoog F, Miller CO (1957) Chemical regulation of growth and organ formation in plant tissues cultured in vitro. Symp Soc Exp Biol 11:118–131
Šmehilová M, Dobrusková J, Novák O et al (2016) Cytokinin-specific glycosyltransferases possess different roles in cytokinin homeostasis maintenance. Front Plant Sci 7:1264. https://doi.org/10.3389/fpls.2016.01264
Song JY, Choi EY, Lee HS et al (1995) Effect of auxin on expression of the isopentenyl transferase gene (ipt) in transformed bean (Phaseolus vulgaris L.) single-cell clones induced by Agrobacterium tumefaciens C58. J Plant Physiol 146:148–154. https://doi.org/10.1016/S0176-1617(11)81981-4
Steklov M, Lomin S, Osolodkin D, Romanov G (2013) Structural basis for cytokinin receptor signaling: an evolutionary approach. Plant Cell Rep 32:781–793. https://doi.org/10.1007/s00299-013-1408-3
Stock AM, Robinson VL, Goudreau PN (2000) Two-component signal transduction. Annu Rev Biochem 69:183–215
Strnad M (1997) The aromatic cytokinins. Physiol Plant 101:674–688. https://doi.org/10.1111/j.1399-3054.1997.tb01052.x
Sun J, Niu QW, Tarkowski P et al (2003) The Arabidopsis AtIPT8/PGA22 gene encodes an isopentenyl transferase that is involved in de novo cytokinin biosynthesis. Plant Physiol 131:167–176. https://doi.org/10.1104/pp.011494
Sun J, Hirose N, Wang X et al (2005) Arabidopsis SOI33/AtENT8 gene encodes a putative equilibrative nucleoside transporter that is involved in cytokinin transport in planta. J Int Plant Biol 47:588–603. https://doi.org/10.1111/j.1744-7909.2005.00104.x
Svacinova J, Novak O, Plackova L et al (2012) A new approach for cytokinin isolation from Arabidopsis tissues using miniaturized purification: pipette tip solid-phase extraction. Plant Meth 8:17. https://doi.org/10.1186/1746-4811-8-17
Swaminathan S, Bock RM (1977) Isolation and identification of cytokinins from Euglena gracilis transfer ribonucleic acid. Biochemistry 16:1355–1360. https://doi.org/10.1021/bi00626a018
Takei K, Sakakibara H, Sugiyama T (2001a) Identification of genes encoding adenylate isopentenyltransferase, a cytokinin biosynthesis enzyme, in Arabidopsis thaliana. J Biol Chem 276:26405–26410. https://doi.org/10.1074/jbc.M102130200
Takei K, Sakakibara H, Taniguchi M, Sugiyama T (2001b) Nitrogen-dependent accumulation of cytokinins in root and the translocation to leaf: implication of cytokinin species that induces gene expression of maize response regulator. Plant Cell Physiol 42:85–93. https://doi.org/10.1093/pcp/pce009
Takei K, Ueda N, Aoki K et al (2004a) AtIPT3 is a key determinant of nitrate-dependent cytokinin biosynthesis in Arabidopsis. Plant Cell Physiol 45:1053–1062. https://doi.org/10.1093/pcp/pch119
Takei K, Yamaya T, Sakakibara H (2004b) Arabidopsis CYP735A1 and CYP735A2 encode cytokinin hydroxylases that catalyze the biosynthesis of trans-zeatin. J Biol Chem 279:41866–41872. https://doi.org/10.1074/jbc.M406337200
Tanaka H, Dhonukshe P, Brewer PB, Friml J (2006) Spatiotemporal asymmetric auxin distribution: a means to coordinate plant development. Cell Mol Life Sci 63:2738–2754. https://doi.org/10.1007/s00018-006-6116-5
Tardieu F, Simonneau T, Muller B (2018) The physiological basis of drought tolerance in crop plants: a scenario-dependent probabilistic approach. Annu Rev Plant Biol 69:733–759. https://doi.org/10.1146/annurev-arplant-042817-040218
Taya Y, Tanaka Y, Nishimura S (1978) 5′-AMP is a direct precursor of cytokinin in Dictyostelium discoideum. Nature 271:545–547. https://doi.org/10.1038/271545a0
To JP, Kieber JJ (2008) Cytokinin signaling: two-components and more. Trends Plant Sci 13:85–92. https://doi.org/10.1016/j.tplants.2007.11.005
To JP, Haberer G, Ferreira FJ et al (2004) Type-A Arabidopsis response regulators are partially redundant negative regulators of cytokinin signaling. Plant Cell 16:658–671. https://doi.org/10.1105/tpc.018978
Tokunaga H, Kojima M, Kuroha T et al (2012) Arabidopsis lonely guy (LOG) multiple mutants reveal a central role of the LOG-dependent pathway in cytokinin activation. Plant J 69:355–365. https://doi.org/10.1111/j.1365-313X.2011.04795.x
Tounekti T, Hernández I, Müller M et al (2011) Kinetin applications alleviate salt stress and improve the antioxidant composition of leaf extracts in Salvia officinalis. Plant Physiol Biochem 49:1165–1176. https://doi.org/10.1016/j.plaphy.2011.07.011
Tsai YC, Weir NR, Hill K et al (2012) Characterization of genes involved in cytokinin signaling and metabolism from rice. Plant Physiol 158:1666–1684. https://doi.org/10.1104/pp.111.192765
Turner J, Mok D, Mok M, Shaw G (1987) Isolation and partial purification of an enzyme catalyzing the formation of O-xylosylzeatin in Phaseolus vulgaris embryos. Proc Natl Acad Sci (USA) 84:3714–3717. https://doi.org/10.1073/pnas.84.11.3714
VanWallendael A, Soltani A, Emery NC et al (2019) A molecular view of plant local adaptation: incorporating stress-response networks. Annu Rev Plant Biol 70:559–583. https://doi.org/10.1146/annurev-arplant-050718-100114
Walch-Liu P, Neumann G, Bangerth F, Engels C (2000) Rapid effects of nitrogen form on leaf morphogenesis in tobacco. J Exp Bot 51:227–237. https://doi.org/10.1093/jexbot/51.343.227
Wang R, Tischner R, Gutierrez RA et al (2004) Genomic analysis of the nitrate response using a nitrate reductase-null mutant of Arabidopsis. Plant Physiol 136:2512–2522. https://doi.org/10.1104/pp.104.044610
Wang J, Ma XM, Kojima M et al (2011) N-glucosyltransferase UGT76C2 is involved in cytokinin homeostasis and cytokinin response in Arabidopsis thaliana. Plant Cell Physiol 52:2200–2213. https://doi.org/10.1093/pcp/pcr152
Wang J, Ma XM, Kojima M et al (2013) Glucosyltransferase UGT76C1 finely modulates cytokinin responses via cytokinin N-glucosylation in Arabidopsis thaliana. Plant Physiol Biochem 65:9–16. https://doi.org/10.1016/j.plaphy.2013.01.012
Wang Hx, Wang Ml, Wang Xz, Ding Yl (2019) Detection of seven phytohormones in peanut tissues by ultra-high-performance liquid chromatography-triple quadrupole tandem mass spectrometry. J Integr Agric 18:2–10. https://doi.org/10.1016/S2095-3119(19)62640-7
Werner T, Schmülling T (2009) Cytokinin action in plant development. Curr Opin Plant Biol 12:527–538. https://doi.org/10.1016/j.pbi.2009.07.002
Werner T, Motyka V, Laucou V et al (2003) Cytokinin-deficient transgenic Arabidopsis plants show multiple developmental alterations indicating opposite functions of cytokinins in the regulation of shoot and root meristem activity. Plant Cell 15:2532–2550. https://doi.org/10.1105/tpc.014928
Wulfetange K, Lomin SN, Romanov GA et al (2011) The cytokinin receptors of Arabidopsis are located mainly to the endoplasmic reticulum. Plant Physiol 156:1808–1818. https://doi.org/10.1104/pp.111.180539
Wybouw B, De Rybel B (2019) Cytokinin: a developing story. Trends Plant Sci 24:177–185. https://doi.org/10.1016/j.tplants.2018.10.012
Xu J, Zha M, Li Y et al (2015) The interaction between nitrogen availability and auxin, cytokinin, and strigolactone in the control of shoot branching in rice (Oryza sativa L.). Plant Cell Rep 34:1647–1662. https://doi.org/10.1007/s00299-015-1815-8
Yamada H, Suzuki T, Terada K et al (2001) The Arabidopsis AHK4 histidine kinase is a cytokinin-binding receptor that transduces cytokinin signals across the membrane. Plant Cell Physiol 42:1017–1023. https://doi.org/10.1093/pcp/pce127
Yamburenko MV, Kieber JJ, Schaller GE (2017) Dynamic patterns of expression for genes regulating cytokinin metabolism and signaling during rice inflorescence development. PLoS ONE 12:e0176060. https://doi.org/10.1371/journal.pone.0176060
Yokoyama A, Yamashino T, Amano YI et al (2007) Type-B ARR transcription factors, ARR10 and ARR12, are implicated in cytokinin-mediated regulation of protoxylem differentiation in roots of Arabidopsis thaliana. Plant Cell Physiol 48:84–96. https://doi.org/10.1093/pcp/pcl040
Young NF, Ferguson BJ, Antoniadi I et al (2014) Conditional auxin response and differential cytokinin profiles in shoot branching mutants. Plant Physiol 165:1723–1736. https://doi.org/10.1104/pp.114.239996
Yu J, Tehrim S, Wang L et al (2017) Evolutionary history and functional divergence of the cytochrome P450 gene superfamily between Arabidopsis thaliana and Brassica species uncover effects of whole genome and tandem duplications. BMC Genom 18:733. https://doi.org/10.1186/s12864-017-4094-7
Žd'árská M, Zatloukalová P, Benítez M et al (2013) Proteome analysis in Arabidopsis reveals shoot- and root-specific targets of cytokinin action and differential regulation of hormonal homeostasis. Plant Physiol 161:918. https://doi.org/10.1104/pp.112.202853
Zhang R, Zhang X, Wang J et al (1995) The effect of auxin on cytokinin levels and metabolism in transgenic tobacco tissue expressing an ipt gene. Planta 196:84–94. https://doi.org/10.1007/BF00193221
Zhang XD, Letham DS, Zhang R, Higgins TJV (1996) Expression of the isopentenyl transferase gene is regulated by auxin in transgenic tobacco tissues. Transg Res 5:57–65. https://doi.org/10.1007/BF01979922
Zhang X, Chen Y, Lin X et al (2013) Adenine phosphoribosyl transferase 1 is a key enzyme catalyzing cytokinin conversion from nucleobases to nucleotides in Arabidopsis. Mol Plant 6:1661–1672. https://doi.org/10.1093/mp/sst071
Zhang K, Novak O, Wei Z et al (2014) Arabidopsis ABCG14 protein controls the acropetal translocation of root-synthesized cytokinins. Nat Commun 5:3274. https://doi.org/10.1038/ncomms4274
Zhang K, Wang R, Zi H et al (2018) AUXIN RESPONSE FACTOR3 regulates floral meristem determinacy by repressing cytokinin biosynthesis and signaling. Plant Cell 30:324–346. https://doi.org/10.1105/tpc.17.00705
Zhao J, Yu N, Ju M et al (2019) ABC transporter OsABCG18 controls the shootward transport of cytokinins and grain yield in rice. J Exp Bot (In press). https://doi.org/10.1093/jxb/erz382
Zhou M, Ghnaya T, Dailly H et al (2019) The cytokinin trans-zeatine riboside increased resistance to heavy metals in the halophyte plant species Kosteletzkya pentacarpos in the absence but not in the presence of NaCl. Chemosphere 233:954–965. https://doi.org/10.1016/j.chemosphere.2019.06.023
Zürcher E, Tavor-Deslex D, Lituiev D et al (2013) A robust and sensitive synthetic sensor to monitor the transcriptional output of the cytokinin signaling network in planta. Plant Physiol 161:1066–1075. https://doi.org/10.1104/pp.112.211763
Zürcher E, Liu J, di Donato M et al (2016) Plant development regulated by cytokinin sinks. Science 353:1027–1030. https://doi.org/10.1126/science.aaf7254
Funding
This work was supported by the National Council of Science and Technology (FS-1515 and 257436 to VML-V).
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Márquez-López, R.E., Quintana-Escobar, A.O. & Loyola-Vargas, V.M. Cytokinins, the Cinderella of plant growth regulators. Phytochem Rev 18, 1387–1408 (2019). https://doi.org/10.1007/s11101-019-09656-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11101-019-09656-6