Abstract
Purpose
Primary central nervous system lymphoma (PCNSL) is a non-Hodgkin lymphoma that affects the central nervous system (CNS). Although previous studies have reported the most common mutated genes in PCNSL, including MYD88 and CD79b, our understanding of genetic characterizations in primary CNS lymphomas is limited. The aim of this study was to perform a retrospective analysis investigating the most frequent mutation types, and their frequency, in PCNSL.
Methods
Fifteen patients with a diagnosis of PCNSL from our institution were analyzed for mutations in 406 genes and rearrangements in 31 genes by next generation sequencing (NGS).
Results
Missense mutations were identified as the most common mutation type (32%) followed by frame shift mutations (23%). The highest mutation rate was reported in the MYD88 (33.3%), CDKN2A/B (33.3%), and TP53 (26.7%) genes. Intermediate tumor mutation burden (TMB) and high TMB was detected in 13.3% and 26.7% of PCNSL, respectively. The most frequent gene rearrangement involved the IGH-BCL6 genes (20%).
Conclusions
This study shows the most common genetic alterations in PCNSL as determined by a commercial next generation sequencing assay. MYD88 and CD79b are frequently mutated in PCNSL, IGH-BCL6 is the most frequent gene rearrangement and approximately 1/4 of cases show a high TMB. Mutations in multiple genes, in addition to high TMB and gene rearrangements, highlights the complex molecular heterogeneity of PCNSL. Knowledge about genetic alterations in PCNSL can inform the development of novel targets for diagnosis and treatment.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Diffuse large B-cell lymphoma (DLBCL) is the most prevalent type of non-Hodgkin B-cell lymphoma (B-NHL), representing roughly 30–40% of all cases worldwide. Patients commonly present with fast growing tumors which can involve single or multiple, nodal or extranodal sites [1]. DLBCL can virtually develop in any primary tissue site from two major cellular subtypes: activated B cell-like (ABC) or germinal center B cell-like (GCB). The respective subtypes have different mechanisms of development, genetic alterations, and treatment response; with the ABC subtype showing an inferior prognosis [1, 2].
Primary central nervous system lymphoma (PCNSL) is a rare subtype of DLBCL that arises and is confined to the central nervous system (CNS) [3]. This non-Hodgkin aggressive B-cell lymphoma is distinguished from extra-cerebral DLBCL by its poorer prognosis. PCNSL can occur in the setting of immunosuppression (HIV/AIDS, post-transplant) or in immunocompetent individuals [4,5,6]. While treatment response rates are high, relapses are frequent and prognosis after recurrence is poor with 5-year survival rates ranging from 22 to 40% [7, 8]. The genomic alterations (GAs) underlying PCNSL have not been comprehensively studied.
Single nucleotide mutations in various genes, including MYD88, CD79b, PIM1, and BTG2, have been reported as the most prevalent genetic alterations in PCNSL [9,10,11]. Among these mutations, an MYD88 substitution mutation at c.794T > C resulting in a replacement of leucine 265 by proline (L265P) is the most common mutation in PCNSL. Myeloid differentiation factor 88 (MYD88) is an adaptor molecule in the Toll-like receptor pathway that mediates interleukin-1 receptor signaling [12]. Similar to several other common mutations in PCNSL, MYD88 mutations lead to constitutive activation of the nuclear factor NF-κB signaling pathway [13, 14]. Translocation of NF-κB into the nucleus subsequently initiates activation of other target genes [13, 14].
From a molecular perspective, GAs are of great interest as they can serve as diagnostic biomarkers or targets for personalized therapies. Despite considerable progress in the understanding of CNS lymphomas, the majority of existing molecular data is derived from locus specific approaches targeting single candidate genes for point mutations, like MYD88 [15]. Over the past decade, the development and affordability of next generation sequencing (NGS) has facilitated several studies identifying GAs in CNS lymphomas through targeted and whole-exome sequencing [15,16,17,18]. Nonetheless, the rarity of the disease and the restricted availability of affected brain tissue hinder the study of molecular and GAs in PCNSL. Therefore, our understanding of this disease remains limited [19]. In addition, discrimination of PCNSL and secondary CNS lymphomas can be very challenging by conventional microscopic examination and magnetic resonance imaging (MRI) alone [20]. Better understanding of molecular alterations in PCNSL can be of clinical utility by facilitating the distinction of PCNSL from secondary CNS lymphoma. To increase our understanding of GAs in PCNSL, we retrospectively investigated the results of a comprehensive NGS assay in a series of 15 PCNSL.
Methods
Patients and tumor samples
This retrospective study was approved by the institutional review board of the University of Texas Health Science Center at Houston and Memorial Hermann Hospital, Houston, TX. From January 2010 to December 2017, 50 consecutive patients diagnosed with PCNSL were identified in the clinical records of the University of Texas Health Science Center at Houston and Memorial Hermann Hospital, Houston TX. The results of molecular testing were available for 15 patients.
All 15 tumor samples were examined by H&E (Supplementary Fig. S1) and immunohistochemistry, and confirmed as DLBCLs. Two patients had a history of immunosuppression (HIV positive). The patients’ ages ranged from 22 to 80 years, average age of 58 years. There were seven men and eight women. Clinical and treatment information was available for all patients (Table 1). All patients underwent either biopsy or tumor resection. Although treatments were variable, high-dose methotrexate (HD-MTX) was the most prevalent therapy used in combination with other treatment modalities.
Immunohistochemistry
Paraffin-embedded tissue sections were de-paraffinized and rehydrated using xylene and graded alcohols. Immunohistochemical staining was performed in a Dako Omnis autostainer (Dako North America, Inc. Carpinteria, CA, USA). The following primary antibodies were used: CD20 (L26), CD79a (12E7), CD10 (56C6), CD23 (DAK-CD23), BCL2 (124), BCL6 (PG-B6p), MUM1 (MUM1p), cyclin D1 (EP12), and Ki67 (MIB-1). The immunohistochemical staining was interpreted as positive or negative by a board-certified pathologist in all cases.
Targeted sequencing and tumor mutation burden
Formalin-fixed paraffin-embedded tumor samples were analyzed for genetic alterations by targeted NGS (FoundationOne™Heme, Foundation Medicine Inc., Cambridge, MA, USA). The FoundationOne® Heme assay was performed in a clinical laboratory improvement amendments (CLIA)-certified laboratory, as previously described [21]. Adaptor-ligated sequencing libraries were captured by solution hybridization with two custom bait-sets targeting 406 cancer-related genes, 31 genes frequently rearranged by DNA-seq, and 265 genes frequently rearranged by RNA-seq (Supplementary Table 1). The captured products were sequenced on HiSeq2500, Illumina. Sequenced data was evaluated for four classes of GAs: base substitution, copy number alterations, fusion/rearrangements, and insertion and deletions. Final NGS results were acquired from the patients’ FoundationOne Heme reports.
Tumor mutation burden (TMB) was calculated based on the number of somatic mutations in sequenced genes and extrapolating the value to the genome as a whole using a validated algorithm [22, 23]. TMB was reported as a number of mutations per megabase (mb) of genome. Based on the FoundationOne™Heme reports, TMB results were also categorized into three groups: low (1–5 mutations/mb), intermediate (6–19 mutations/mb), and high (≥ 20 mutations/mb). Values were rounded to the nearest integer.
Gene ontology
To do the gene ontology (GO) analysis, we downloaded GO gene sets from Molecular Signatures Database (MSigDB) [24]. A total of 5917 gene sets, including 4436 biological process, 580 cellular components, and 901 molecular function gene sets, were used in this study. The size of each gene set (N size) was calculated and the number of mutant genes in our dataset (50 genes) within each gene set was determined. To evaluate the enrichment of mutant genes in GO dataset, we computed the fraction of mutant genes in each gene set (50 genes/N size). All the analysis was done in R (version 3.4.2).
Results
Patient characteristics
Clinicopathologic characteristics for all patients are summarized in Tables 1 and 2. Four patients underwent resection; all others underwent needle biopsy. All tumors were positive for CD20 and CD79a by immunohistochemical staining. Employing the immunohistochemical algorithm of Hans et al. [25], all cases were sub-classified as either ABC (n = 7, 47%) or GCB subtype (n = 7, 47%), with the exception of one case for which immunohistochemical staining was not available. Both HIV positive patients were diagnosed with PCNSL of the ABC subtype. The results of cerebrospinal fluid (CSF) cytology were available for eight patients, all of them had a negative CSF-cytology result. Among nine CSFs tested by flow cytometry, four cases had a diagnosis of B-cell lymphoma, four cases were reported as non-diagnostic and one case was negative. One patient developed recurrence with diffuse osseous metastases. At the time of the study 2/15 (13.3%) patients had died of lymphoma, both patients had declined treatment and received comfort care only.
Genomic alterations
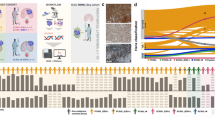
A total of 79 GAs were detected in 50 genes (Fig. 1). The median number of GAs detected by the assay per patient was 5 (range 1–10). In 8 (10.1%) events, single genes harbored more than one GA. In the two HIV positive cases, only a single mutation was detected in MLL3 (KMT2C) and TUSC3 tumor suppressor genes. The rate of mutation frequency in HIV positive patients was lower than other PCNSLs. The most prevalent mutations were missense (n = 25, 32%) followed by frame shift (n = 18, 23%) (Fig. 1a). The most commonly mutated gene was MYD88 (n = 5, 33.3%), followed by alterations in CDKN2A/B (n = 5, 33.3%), and TP53 (n = 4, 26.7%). Fusion or rearrangement of the IGH gene with Bcl2, Bcl6, and MALT1 genes was also identified in 33.3% of cases (n = 5). It is worth to mention that all gene loss mutation was reported in CDKN2A/B gene (Fig. 1b). All cases with alterations in MYD88 (33.3%) had a missense mutation, the p.L265P substitution affecting exon 5, except one case that showed the less common p.V217F mutation. MYD88 mutations occurred more commonly in DLBCL of the ABC subtype (42.9%) versus the GCB subtype (28.6%). Of the 15 patients, 4 cases (26.7%) had high TMB (≥ 20 mutations/mb), 2 cases (13.3%) had an intermediate mutation burden (8–12 mutations/mb), and 9 cases (60%) had a low TMB (1–5 mutations/mb). Both HIV positive patients demonstrated a low TMB. The GO analysis revealed that genes mutated in PCNSL were enriched in pathways involved in “Cell Death”, “Positive Regulation of Transcription from RNA Polymerase II promoter”, “Mitotic DNA Integrity Checkpoint”, “Chromosome Organization”, “Cellular Response to Endogenous Stimulus”, and “Regulation of Smooth Muscle Cell Proliferation” (Fig. 1c).
Types of mutations observed in PCNSL. a Pie chart showing the percentages of different types of somatic mutations in PCNSL. Approximately, 50% of the mutations in PCNSL are missense or frameshift mutations. Gene rearrangements were detected in 10% of cases. b Genes mutated in PCNSL. Mutations in MYD88, CDKN2A/B, TP53 and IGH-BCL6 were the most frequent alterations detected. Asterisk indicates information from various sources (e.g., My Cancer Genome, COSMIC, Pubmed, and the Foundation Medicine report) were used to determine the predicted effect of the mutations on the function of each gene. c GO enrichment analysis of 50 mutated genes. The mutant genes were enriched in pathways of “Cell Death”, “Positive Regulation of Transcription from RNA Polymerase II promoter”, “Mitotic DNA Integrity Checkpoint” and “Chromosome Organization”
Discussion
In the current study, we report the results of a comprehensive analysis of GAs in PCNSL, using a targeted NGS assay that evaluates more than 400 cancer-related genes. Among DLBCLs from different sites, PCNSL has the highest frequency of MYD88 mutations [26], which is in agreement with our results showing a high MYD88 mutation rate (33.3%) in PCNSLs. We also detected a higher frequency of MYD88 mutations in PCNSL of the ABC subtype, compared to the GCB subtype, in accordance with previous studies [12, 27]. Nonetheless, Fukumura et al. [3] reported no significant differences in MYD88 mutations between ABC- and GCB-lymphomas. Due to the low prevalence of PCNSLs, the majority of studies include a small cohort of patients, which can be partially responsible for variations in the reported frequencies of GAs. Although we report genomic mutations in a limited number of patients, our study includes evaluation for TMB and gene rearrangements, which have been neglected in the majority of previous studies on PCNSL. Previous studies have reported the absence of concurrent CXCR4 and MYD88 mutations in PCNSLs, in contrast to Waldenstrom macroglobulinemia, in which a significant number of patients show coexisting CXCR4 and MYD88 mutations [28, 29]. In our study, we identified one case of PCNSL with CXCR4 and MYD88 mutations. A recent study on genomic characterization of lymphomas also established the presence of the respective mutations in PCNSLs [30]. Waldenstrom macroglobulinemia patients who have MYD88 and CXCR4 mutations have been reported to be more resistant to ibrutinib treatment due to the activation of the AKT and ERK pathways [31,32,33]. The effects of CXCR4 mutations in PCNSL are currently unknown. However, an FDA-approved CXCR4 antagonist AMD3100 (Plerixafor/Mozobil) could potentially be a candidate therapy for patients with CXCR4 mutated malignancies [34, 35].
MYD88 mutations have been reported in other hematologic malignancies, including Waldenstrom macroglobulinemia, chronic lymphocytic leukemia (CLL) and mucosa-associated lymphoid tissue (MALT) lymphomas [12, 36]. In PCNSL, different studies have reported a wide range of mutation frequencies in the MYD88 gene ranging from 33 to 100% (Table 3). This wide spectrum is partially due to variations in sensitivity of the various assays utilized and differences in patient population. For instance, Hattori et al., reported the MYD88 p.L265P mutation was detected in 100% of cases (14/14) using ddPCR. In contrast, they could detect the mutation by targeted deep next generation sequencing (TDS) in 13 out of 14 cases [37]. In another study, whole exome sequencing revealed an MYD88 mutation in 75.6% of cases. However, after manual inspection of the negative cases, followed by sanger sequencing confirmation, the percentage or MYD88 mutant cases reached 85.4% [3]. Given the high prevalence of MYD88 mutations in PCNSLs, treatment with ibrutinib, which inhibits Bruton’s Tyrosine Kinase and further suppresses NF-κB and STAT3 activation and tumor growth, could be considered in patients with PCNSL [38, 39]. However, although there are ongoing clinical trials (NCT02315326), the efficacy of ibrutinib for the treatment of PCNSL remains to be determined. In addition to the therapeutic implication of the MYD88 mutation in PCNSL, its detection in cerebrospinal fluid has been recently introduced as a potential minimally-invasive approach for diagnosis [40].
CDKN2A and CDKN2B encode p14ARF and p16INK4a, and p15INK4b tumor suppressor proteins, respectively. In accordance with our study, loss of CDKN2A/B genes have been commonly reported in DLBCL, which has been associated with reduced mRNA expression and higher tumor grades [45,46,47,48]. The p16INK4a and p15INK4b proteins maintain Rb tumor suppressor activity through suppression of CDK4 and CDK6 [49]. Therefore, using CDK4/6 inhibitors, including LY2835219, LEE011, and the FDA-approved palbociclib has been suggested as a potentially helpful therapy for CDKN2A/B mutated tumors [50]. However, initial clinical results did not provide promising results and further investigations for PCNSL is required [51,52,53,54]. The p14ARF protein is responsible for TP53 activation and induced cell death [55, 56]. TP53 was also among the commonly mutated genes in our study. A research study on 506 primary DLBCL patients treated with R-CHOP reported that TP53 mutation significantly correlate with worse survival in either ABC- or GCB-DLBCL [57].
There is a growing body of clinical and experimental evidence that TMB could be used as a biomarker for tumor prognosis and predicting response to immunotherapy [58, 59]. Various factors, including microsatellite instability [60], cigarette smoke [61, 62], and exposure to mutagens like UV light [63] have been associated with TMB in other tumors. High-TMB was reported in PCNSLs recently with a frequency of 33% [30], which is similar to our results (26.7%). From the therapeutic point of view, a high TMB was suggested to be associated with better prognosis in different tumors and susceptibility to nivolumab treatment in non-small cell lung cancer [58, 59]. A previous study analyzing 100,00 human cancer genomes showed that TMB determination, by comprehensive genomic profiling like the one reported in our study, correlated with TMB determined by whole exome sequencing [64]. This is clinically useful in regards to lymphoma, since it has been shown that high TMB and high PD-L1 expression in DLBCL may be linked to greater responsiveness to immunotherapeutic agents and checkpoint inhibitor therapies like anti-PDL1 [30, 65].
The presence of IGH-Bcl6 rearrangements in 20% of PCNSL is in accordance with prior studies (17–47%) [66,67,68,69]. Overexpression of Bcl6 due to the chromosomal translocation with impaired immunoglobulin IGH and further arrest in B-cell differentiation was reported to be one of the primary contributing factors to PCNSL pathogenesis. On the other hand, IGH-Bcl2 rearrangement, resulting from the reciprocal chromosomal t(14;18)(q31;q21) translocation was detected in one case (6.7%). Multiple studies have tried to define a prognostic role for Bcl2 and Bcl6 rearrangement in patients with PCNSL, however, the results are conflicting [67, 70,71,72]. A recent FDA approved anti-CD19 CAR T-cell therapy, Axicabtagene ciloleucel (KTE-C19) has been recommended for high-grade B-cell lymphoma with MYC, Bcl2 and/or Bcl6 rearrangement [73, 74]. However, its utility for the treatment of PCNSL remains to be determined. Characterization of GAs in PCNSL is critical for the development of non-invasive methods for diagnosis and targeted therapies [40]. In addition to point mutations, our study highlights other genomic events, including gene deletions, rearrangements or fusions in PCNSLs. While the number of patients in the study is limited, our findings broadens our understanding of the molecular heterogeneity of PCNSL.
References
Li S, Young KH, Medeiros LJ (2018) Diffuse large B-cell lymphoma. Pathology 50:74–87. https://doi.org/10.1016/j.pathol.2017.09.006
Rosenwald A, Wright G, Chan WC et al (2002) The use of molecular profiling to predict survival after chemotherapy for diffuse large-B-cell lymphoma. N Engl J Med 346:1937–1947. https://doi.org/10.1056/NEJMoa012914
Fukumura K, Kawazu M, Kojima S et al (2016) Genomic characterization of primary central nervous system lymphoma. Acta Neuropathol 131:865–875. https://doi.org/10.1007/s00401-016-1536-2
Diamond C, Taylor TH, Aboumrad T, Anton Culver H (2006) Changes in acquired immunodeficiency syndrome-related non-Hodgkin lymphoma in the era of highly active antiretroviral therapy. Cancer 106:128–135. https://doi.org/10.1002/cncr.21562
Besson C, Goubar A, Gabarre J et al (2001) Changes in AIDS-related lymphoma since the era of highly active antiretroviral therapy. Blood 98:2339–2344
Shiels MS, Pfeiffer RM, Besson C et al (2016) Trends in primary central nervous system lymphoma incidence and survival in the U.S. Br J Haematol 174:417–424. https://doi.org/10.1111/bjh.14073
Gavrilovic IT, Hormigo A, Yahalom J et al (2006) Long-term follow-up of high-dose methotrexate-based therapy with and without whole brain irradiation for newly diagnosed primary CNS lymphoma. J Clin Oncol 24:4570–4574. https://doi.org/10.1200/JCO.2006.06.6910
Nayak L, Hedvat C, Rosenblum MK et al (2011) Late relapse in primary central nervous system lymphoma: clonal persistence. Neuro-oncology 13:525–529. https://doi.org/10.1093/neuonc/nor014
Zheng M, Perry AM, Bierman P et al (2017) Frequency of MYD88 and CD79B mutations, and MGMT methylation in primary central nervous system diffuse large B-cell lymphoma. Neuropathology 37:509–516. https://doi.org/10.1111/neup.12405
Yamada S, Ishida Y, Matsuno A, Yamazaki K (2015) Primary diffuse large B-cell lymphomas of central nervous system exhibit remarkably high prevalence of oncogenic MYD88 and CD79B mutations. Leuk Lymphoma 56:2141–2145. https://doi.org/10.3109/10428194.2014.979413
Nakamura T, Tateishi K, Niwa T et al (2016) Recurrent mutations of CD79B and MYD88 are the hallmark of primary central nervous system lymphomas. Neuropathol Appl Neurobiol 42:279–290. https://doi.org/10.1111/nan.12259
Ngo VN, Young RM, Schmitz R et al (2011) Oncogenically active MYD88 mutations in human lymphoma. Nature 470:115–119. https://doi.org/10.1038/nature09671
Lim K-H, Yang Y, Staudt LM (2012) Pathogenetic importance and therapeutic implications of NF-κB in lymphoid malignancies. Immunol Rev 246:359–378. https://doi.org/10.1111/j.1600-065X.2012.01105.x
Fitzgerald KA, Palsson-McDermott EM, Bowie AG et al (2001) Mal (MyD88-adapter-like) is required for Toll-like receptor-4 signal transduction. Nature 413:78–83. https://doi.org/10.1038/35092578
Montesinos-Rongen M, Godlewska E, Brunn A et al (2011) Activating L265P mutations of the MYD88 gene are common in primary central nervous system lymphoma. Acta Neuropathol 122:791–792. https://doi.org/10.1007/s00401-011-0891-2
Braggio E, Van Wier S, Ojha J et al (2015) Genome-wide analysis uncovers novel recurrent alterations in primary central nervous system lymphomas. Clin Cancer Res 21:3986–3994. https://doi.org/10.1158/1078-0432.CCR-14-2116
Vater I, Montesinos-Rongen M, Schlesner M et al (2015) The mutational pattern of primary lymphoma of the central nervous system determined by whole-exome sequencing. Leukemia 29:677–685. https://doi.org/10.1038/leu.2014.264
Gonzalez-Aguilar A, Idbaih A, Boisselier B et al (2012) Recurrent mutations of MYD88 and TBL1XR1 in primary central nervous system lymphomas. Clin Cancer Res 18:5203–5211. https://doi.org/10.1158/1078-0432.CCR-12-0845
Ostrom QT, Gittleman H, Liao P et al (2017) CBTRUS statistical report: primary brain and other central nervous system tumors diagnosed in the United States in 2010–2014. Neuro-oncology 19:v1–v88. https://doi.org/10.1093/neuonc/nox158
Lu S, Gao Q, Yu J et al (2016) Utility of dynamic contrast-enhanced magnetic resonance imaging for differentiating glioblastoma, primary central nervous system lymphoma and brain metastatic tumor. Eur J Radiol 85:1722–1727. https://doi.org/10.1016/j.ejrad.2016.07.005
Wang K, Sanchez-Martin M, Wang X et al (2017) Patient-derived xenotransplants can recapitulate the genetic driver landscape of acute leukemias. Leukemia 31:151–158. https://doi.org/10.1038/leu.2016.166
Rosenberg JE, Hoffman-Censits J, Powles T et al (2016) Atezolizumab in patients with locally advanced and metastatic urothelial carcinoma who have progressed following treatment with platinum-based chemotherapy: a single-arm, multicentre, phase 2 trial. Lancet 387:1909–1920. https://doi.org/10.1016/S0140-6736(16)00561-4
Goodman AM, Kato S, Bazhenova L et al (2017) Tumor mutational burden as an independent predictor of response to immunotherapy in diverse cancers. Mol Cancer Ther 16:2598–2608. https://doi.org/10.1158/1535-7163.MCT-17-0386
Subramanian A, Tamayo P, Mootha VK et al (2005) Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA 102:15545–15550. https://doi.org/10.1073/pnas.0506580102
Hans CP, Weisenburger DD, Greiner TC et al (2004) Confirmation of the molecular classification of diffuse large B-cell lymphoma by immunohistochemistry using a tissue microarray. Blood 103:275–282. https://doi.org/10.1182/blood-2003-05-1545
Lee J-H, Jeong H, Choi J-W et al (2017) Clinicopathologic significance of MYD88 L265P mutation in diffuse large B-cell lymphoma: a meta-analysis. Sci Rep 7:659. https://doi.org/10.1038/s41598-017-01998-5
Kraan W, Horlings HM, van Keimpema M et al (2013) High prevalence of oncogenic MYD88 and CD79B mutations in diffuse large B-cell lymphomas presenting at immune-privileged sites. Blood Cancer J 3:e139–e139. https://doi.org/10.1038/bcj.2013.28
Treon SP, Cao Y, Xu L et al (2014) Somatic mutations in MYD88 and CXCR4 are determinants of clinical presentation and overall survival in Waldenstrom macroglobulinemia. Blood 123:2791–2796. https://doi.org/10.1182/blood-2014-01-550905
Poulain S, Boyle EM, Tricot S et al (2015) Absence of CXCR4 mutations but high incidence of double mutant in CD79A/B and MYD88 in primary central nervous system lymphoma. Br J Haematol 170:285–287. https://doi.org/10.1111/bjh.13293
Severson E, Vergilio J-A, Gay L et al (2017) PATH-20. Comprehensive genomic profiling comparing primary CNS lymphoma to systemic diffuse large B cell lymphoma reveals biomarkers indicating potential benefit from immune checkpoint inhibitors. Neuro-oncology 19:vi175–vi175. https://doi.org/10.1093/neuonc/nox168.711
Domanska UM, Kruizinga RC, Nagengast WB et al (2013) A review on CXCR4/CXCL12 axis in oncology: no place to hide. Eur J Cancer 49:219–230. https://doi.org/10.1016/j.ejca.2012.05.005
Cao Y, Hunter ZR, Liu X et al (2015) The WHIM-like CXCR4(S338X) somatic mutation activates AKT and ERK, and promotes resistance to ibrutinib and other agents used in the treatment of Waldenstrom’s Macroglobulinemia. Leukemia 29:169–176. https://doi.org/10.1038/leu.2014.187
Cao Y, Hunter ZR, Liu X et al (2015) CXCR4 WHIM-like frameshift and nonsense mutations promote ibrutinib resistance but do not supplant MYD88(L265P) -directed survival signalling in Waldenström macroglobulinaemia cells. Br J Haematol 168:701–707. https://doi.org/10.1111/bjh.13200
Pusic I, DiPersio JF (2010) Update on clinical experience with AMD3100, an SDF-1/CXCL12-CXCR4 inhibitor, in mobilization of hematopoietic stem and progenitor cells. Curr Opin Hematol 17:319–326. https://doi.org/10.1097/MOH.0b013e328338b7d5
McDermott DH, Lopez J, Deng F et al (2011) AMD3100 is a potent antagonist at CXCR4(R334X), a hyperfunctional mutant chemokine receptor and cause of WHIM syndrome. J Cell Mol Med 15:2071–2081. https://doi.org/10.1111/j.1582-4934.2010.01210.x
Martínez-Trillos A, Pinyol M, Navarro A et al (2014) Mutations in TLR/MYD88 pathway identify a subset of young chronic lymphocytic leukemia patients with favorable outcome. Blood 123:3790–3796. https://doi.org/10.1182/blood-2013-12-543306
Hattori K, Sakata-Yanagimoto M, Suehara Y et al (2018) Clinical significance of disease-specific MYD88 mutations in circulating DNA in primary central nervous system lymphoma. Cancer Sci 109:225–230. https://doi.org/10.1111/cas.13450
Treon SP, Xu L, Hunter Z (2015) MYD88 mutations and response to ibrutinib in Waldenström’s macroglobulinemia. N Engl J Med 373:584–586. https://doi.org/10.1056/NEJMc1506192
Wilson WH, Young RM, Schmitz R et al (2015) Targeting B cell receptor signaling with ibrutinib in diffuse large B cell lymphoma. Nat Med 21:922–926. https://doi.org/10.1038/nm.3884
Hiemcke-Jiwa LS, Minnema MC, Radersma-van Loon JH et al (2017) The use of droplet digital PCR in liquid biopsies: a highly sensitive technique for MYD88 p.(L265P) detection in cerebrospinal fluid. Hematol Oncol 21:E7. https://doi.org/10.1002/hon.2489
Bruno A, Boisselier B, Labreche K et al (2014) Mutational analysis of primary central nervous system lymphoma. Oncotarget 5:5065–5075. https://doi.org/10.18632/oncotarget.2080
Chapuy B, Roemer MGM, Stewart C et al (2016) Targetable genetic features of primary testicular and primary central nervous system lymphomas. Blood 127:869–881. https://doi.org/10.1182/blood-2015-10-673236
Choi J-W, Kim Y, Lee J-H, Kim Y-S (2013) MYD88 expression and L265P mutation in diffuse large B-cell lymphoma. Hum Pathol 44:1375–1381. https://doi.org/10.1016/j.humpath.2012.10.026
Takano S, Hattori K, Ishikawa E et al (2018) MyD88 mutation in elderly predicts poor prognosis in primary central nervous system lymphoma: multi-institutional analysis. World Neurosurg 112:e69–e73. https://doi.org/10.1016/j.wneu.2017.12.028
Baur AS, Shaw P, Burri N et al (1999) Frequent methylation silencing of p15(INK4b) (MTS2) and p16(INK4a) (MTS1) in B-cell and T-cell lymphomas. Blood 94:1773–1781
Guney S, Jardin F, Bertrand P et al (2012) Several mechanisms lead to the inactivation of the CDKN2A (P16), P14ARF, or CDKN2B (P15) genes in the GCB and ABC molecular DLBCL subtypes. Genes Chromosomes Cancer 51:858–867. https://doi.org/10.1002/gcc.21970
Quelle DE, Zindy F, Ashmun RA, Sherr CJ (1995) Alternative reading frames of the INK4a tumor suppressor gene encode two unrelated proteins capable of inducing cell cycle arrest. Cell 83:993–1000
Sharpless NE (2005) INK4a/ARF: a multifunctional tumor suppressor locus. Mutat Res 576:22–38. https://doi.org/10.1016/j.mrfmmm.2004.08.021
Roussel MF (1999) The INK4 family of cell cycle inhibitors in cancer. Oncogene 18:5311–5317. https://doi.org/10.1038/sj.onc.1202998
Dickson MA (2014) Molecular pathways: CDK4 inhibitors for cancer therapy. Clin Cancer Res 20:3379–3383. https://doi.org/10.1158/1078-0432.CCR-13-1551
DeMichele A, Clark AS, Tan KS et al (2015) CDK 4/6 inhibitor palbociclib (PD0332991) in Rb + advanced breast cancer: phase II activity, safety, and predictive biomarker assessment. Clin Cancer Res 21:995–1001. https://doi.org/10.1158/1078-0432.CCR-14-2258
Finn RS, Crown JP, Lang I et al (2015) The cyclin-dependent kinase 4/6 inhibitor palbociclib in combination with letrozole versus letrozole alone as first-line treatment of oestrogen receptor-positive, HER2-negative, advanced breast cancer (PALOMA-1/TRIO-18): a randomised phase 2 study. Lancet Oncol 16:25–35. https://doi.org/10.1016/S1470-2045(14)71159-3
Infante JR, Cassier PA, Gerecitano JF et al (2016) A phase I study of the cyclin-dependent kinase 4/6 inhibitor ribociclib (LEE011) in patients with advanced solid tumors and lymphomas. Clin Cancer Res 22:5696–5705. https://doi.org/10.1158/1078-0432.CCR-16-1248
Johnson DB, Dahlman KH, Knol J et al (2014) Enabling a genetically informed approach to cancer medicine: a retrospective evaluation of the impact of comprehensive tumor profiling using a targeted next-generation sequencing panel. Oncologist 19:616–622. https://doi.org/10.1634/theoncologist.2014-0011
SHERR CJ, BESTEN den BERTWISTLED W, et al (2005) p53-dependent and -independent functions of the arf tumor suppressor. Cold Spring Harb Symp Quant Biol 70:129–137. https://doi.org/10.1101/sqb.2005.70.004
Ozenne P, Eymin B, Brambilla E, Gazzeri S (2010) The ARF tumor suppressor: structure, functions and status in cancer. Int J Cancer 127:2239–2247. https://doi.org/10.1002/ijc.25511
Xu-Monette ZY, Wu L, Visco C et al (2012) Mutational profile and prognostic significance of TP53 in diffuse large B-cell lymphoma patients treated with R-CHOP: report from an International DLBCL rituximab-CHOP Consortium Program Study. Blood 120:3986–3996. https://doi.org/10.1182/blood-2012-05-433334
Birkbak NJ, Kochupurakkal B, Izarzugaza JMG et al (2013) Tumor mutation burden forecasts outcome in ovarian cancer with BRCA1 or BRCA2 mutations. PLoS ONE 8:e80023. https://doi.org/10.1371/journal.pone.0080023
Peters S, Creelan B, Hellmann MD et al (2017) Abstract CT082: impact of tumor mutation burden on the efficacy of first-line nivolumab in stage iv or recurrent non-small cell lung cancer: an exploratory analysis of CheckMate 026. Cancer Res 77:CT082–CT082. https://doi.org/10.1158/1538-7445.AM2017-CT082
Roberts SA, Gordenin DA (2014) Hypermutation in human cancer genomes: footprints and mechanisms. Nat Rev Cancer 14:786–800. https://doi.org/10.1038/nrc3816
Rizvi NA, Hellmann MD, Snyder A et al (2015) Cancer immunology. Mutational landscape determines sensitivity to PD-1 blockade in non-small cell lung cancer. Science 348:124–128. https://doi.org/10.1126/science.aaa1348
Pfeifer GP, Denissenko MF, Olivier M et al (2002) Tobacco smoke carcinogens, DNA damage and p53 mutations in smoking-associated cancers. Oncogene 21:7435–7451. https://doi.org/10.1038/sj.onc.1205803
Pfeifer GP, You Y-H, Besaratinia A (2005) Mutations induced by ultraviolet light. Mutat Res 571:19–31. https://doi.org/10.1016/j.mrfmmm.2004.06.057
Chalmers ZR, Connelly CF, Fabrizio D et al (2017) Analysis of 100,000 human cancer genomes reveals the landscape of tumor mutational burden. Genome Med 9:34. https://doi.org/10.1186/s13073-017-0424-2
Karim LA, Wang P, de Guzman J et al (2017) Abstract 3724: PDL1 protein expression and tumor mutation burden in hematologic malignancies: correlation with Hodgkin and high grade lymphoma. Cancer Res 77:3724–3724. https://doi.org/10.1158/1538-7445.AM2017-3724
Montesinos-Rongen M, Siebert R, Deckert M (2009) Primary lymphoma of the central nervous system: just DLBCL or not? Blood 113:7–10. https://doi.org/10.1182/blood-2008-04-149005
Cady FM, O’Neill BP, Law ME et al (2008) Del(6)(q22) and BCL6 rearrangements in primary CNS lymphoma are indicators of an aggressive clinical course. J Clin Oncol 26:4814–4819. https://doi.org/10.1200/JCO.2008.16.1455
Montesinos-Rongen M, Akasaka T, Zühlke-Jenisch R et al (2003) Molecular characterization of BCL6 breakpoints in primary diffuse large B-cell lymphomas of the central nervous system identifies GAPD as novel translocation partner. Brain Pathol 13:534–538
Montesinos-Rongen M, Zühlke-Jenisch R, Gesk S et al (2002) Interphase cytogenetic analysis of lymphoma-associated chromosomal breakpoints in primary diffuse large B-cell lymphomas of the central nervous system. J Neuropathol Exp Neurol 61:926–933
Kramer MH, Hermans J, Wijburg E et al (1998) Clinical relevance of BCL2, BCL6, and MYC rearrangements in diffuse large B-cell lymphoma. Blood 92:3152–3162
Akyurek N, Uner A, Benekli M, Barista I (2012) Prognostic significance of MYC, BCL2, and BCL6 rearrangements in patients with diffuse large B-cell lymphoma treated with cyclophosphamide, doxorubicin, vincristine, and prednisone plus rituximab. Cancer 118:4173–4183. https://doi.org/10.1002/cncr.27396
Horn H, Ziepert M, Becher C et al (2013) MYC status in concert with BCL2 and BCL6 expression predicts outcome in diffuse large B-cell lymphoma. Blood 121:2253–2263. https://doi.org/10.1182/blood-2012-06-435842
Neelapu SS, Locke FL, Bartlett NL et al (2017) Axicabtagene Ciloleucel (AXI-CEL; KTE-C19) in patients with refractory aggressive non-hodgkin lymphomas (NHL): primary results of the pivotal trial ZUMA-1. Hematol Oncol 35:28–28. https://doi.org/10.1002/hon.2437_7
Locke FL, Neelapu SS, Bartlett NL et al (2017) Abstract CT019: primary results from ZUMA-1: a pivotal trial of axicabtagene ciloleucel (axicel; KTE-C19) in patients with refractory aggressive non-Hodgkin lymphoma (NHL). Cancer Res 77:CT019–CT019. https://doi.org/10.1158/1538-7445.AM2017-CT019
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Informal consent
For this type of study formal consent is not required.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zorofchian, S., El-Achi, H., Yan, Y. et al. Characterization of genomic alterations in primary central nervous system lymphomas. J Neurooncol 140, 509–517 (2018). https://doi.org/10.1007/s11060-018-2990-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11060-018-2990-6