Abstract
To investigate the radiomic features associated with Ki-67 expression in lower grade gliomas and assess the prognostic values of these features. Patients with lower grade gliomas (n = 117) were randomly assigned into the training (n = 78) and validation (n = 39) sets. A total of 431 radiological features were extracted from each patient. Differential radiological features between the low and high Ki-67 expression groups were screened by significance analysis of microarrays. Then, generalized linear analysis was performed to select features that could predict the Ki-67 expression level. Predictive efficiencies were further evaluated in the validation set. Cox regression analysis was performed to investigate the prognostic values of Ki-67 expression level and Ki-67-related radiological features. A group of nine radiological features were screened for prediction of Ki-67 expression status; these achieved accuracies of 83.3% and 88.6% (areas under the curves, 0.91 and 0.93) in the training and validation sets, respectively. Of these features, only spherical disproportion (SD) was found to be a prognostic factor. Patients in the high SD group exhibited worse outcomes in the whole cohort (overall survival, p < 0.0001; progression-free survival, p < 0.0001). Ki-67 expression level and SD were independent prognostic factors in the multivariate Cox regression analysis. This study identified a radiomic signature for prediction of Ki-67 expression level as well as a prognostic radiological feature in patients with lower grade gliomas.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Lower grade gliomas (World Health Organization [WHO] grade II and III gliomas [1]) are primary brain malignancies. These gliomas vary significantly in terms of clinical manifestation—some lower grade gliomas rapidly progress to glioblastomas, while others are sensitive to treatment and remain stable for several years [2]. Molecular biomarkers are playing an increasing role in tumor diagnosis and guidance of customized treatment [3].
Proliferative index is a potent biomarker, which can quantitatively assess tumor growth and thus help stratify prognosis in patients with gliomas [4]. Ki-67, the most reliable marker of cell proliferation [5], recognizes a core antigen present in proliferating cells and absent in quiescent cells [6]. It has been employed in routine clinical investigations, with high Ki-67 expression levels indicating worse prognosis in patients with gliomas [7]. The conventional method for detection of Ki-67 expression status is immunohistochemistry [8], which is an invasive test involving craniotomy.
Radiogenomics, a novel high throughput method of linking radiological features to genomic data, has the potential to non-invasively retrieve comprehensive intra-, inter-, and peri-tumoral information [9]. Radiogenomics is more efficient and effective than conventional imaging analysis [10]. Several studies have focused on Ki-67-related radiological features in gliomas. Entropy, derived from apparent diffusion coefficient (ADC) maps of gliomas, was found to be associated with Ki-67 expression [11]. Additionally, in a small sample study (n = 21), Ki-67-related features were identified from among 86 radiological features extracted from conventional structural magnetic resonance (MR) images of glioblastomas [12]. However, these studies evaluated relatively few radiological features and did not explore the prognostic significances of Ki-67-related features.
In our study, a number of radiological features were extracted from a large sample of patients, and a subset of features associated with Ki-67 expression level was screened. The efficiency of prediction was evaluated in a validation set, and the prognostic values of Ki-67 expression status and Ki-67-related features were further investigated. Radiological features reflect the biological behavior of brain tumors. Ki-67 is one of a key molecular biomarker that influence the proliferation of gliomas. We here hypothesized that the association is potentially exist of radiological features with Ki-67 expression level in gliomas.
Methods
Patients
This study included 117 patients with lower grade gliomas, who received treatment at Beijing Tiantan Hospital (Beijing, China) between January 2006 and December 2008 and met the following criteria: (a) histopathologically confirmed primary WHO grade II/III gliomas; (b) preoperative T2 MR images available; (c) Ki-67 expression status available; and (d) clinical characteristics and survival data available (Supplementary Fig. 1).
The patients were randomly assigned to the training (n = 78) and validation (n = 39) sets. The training set was used to establish a radiomic signature for prediction of Ki-67 expression status, and the independent validation set was used to assess the predictive efficiency of the signature. Data for this study were retrieved from the Chinese Glioma Genome Atlas (CGGA) [13]. This study was approved by the institutional review board, and all patients was notified that their tumor samples would be used for academic investigations, and the informed consent was obtained.
MRI data acquisition and tumor segmentation
Preoperative clinical structural images of the majority of patients (n = 97) were acquired with a Magnetom Trio 3T (Siemens Healthcare GmbH, Erlangen, Germany) MR scanner. The acquisition parameters for T2-weighted images were as follows: echo time, 110 ms; repetition time, 5800 ms; flip angle, 150°; matrix, 384 × 300; and voxel size, 0.6 × 0.6 × 5 mm3. Other images were acquired on a Magnetom Verio 3T (Siemens, Erlangen, Germany, n = 12) or HD 1.5T (GE Medical System, Waukesha, USA, n = 8) scanner. Tumor volumes were segmented by two experienced neurosurgeons who were blinded to the clinical information of the patients using the MRIcron software (http://www.mccauslandcenter.sc.edu/mricro). Abnormal hyperintense signals on the T2-weighted images were identified as tumor volumes, and the cerebrospinal fluid signals should not be involved in. A senior neuroradiologist subsequently reevaluated the segmented lesions. In case of discrepancies >5% between the two neurosurgeons, the senior neuroradiologist made the final decision.
Textural features
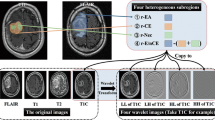
In order to avoid of the bias from data heterogeneity, all MRI data collected in the current study were underwent imaging normalization (Z score transformation) before feature extraction with using an in-house MATLAB process. Quantitative radiological features were extracted as previously reported [14]. and detailed descriptions of each feature was listed in the supplementary material of the previous study. A total of 431 radiological features were extracted, which could be divided into four groups. Features in group 1 (n = 14; first-order statistics-derived features) quantitatively described the signal intensity distribution of the image, while those in group 2 (n = 8; shape- and size-based features) quantified the shape and size data of tumors. Features in group 3 (n = 33; textural features), calculated from gray-level co-occurrence and run-length matrix, reflected intratumoral heterogeneity. Group 4 included 376 features (wavelet features) that were derived from the features of groups 1 and 3 through wavelet transformation. These 431 features are listed in Supplementary Table 1.
Immunohistochemistry
The Ki-67 expression level in the current study was routinely evaluated by experienced pathologists using typical tumor samples collected from the glioma patients. Immunostaining was performed in accordance with the manufacturer’s protocol. Briefly, formalin-fixed, paraffin-embedded tissue sections were cut into 4-µm sections, which were then dried, dewaxed in xylene, rinsed in graded ethanol, and rehydrated in double-distilled water. Immunostaining was performed with the Ki-67 protein antibody (Santa Cruz Biotechnology, Santa Cruz, CA), at a dilution of 1:100. Two pathologists blinded to the clinical data scored the degree of staining. In this study, low Ki-67 expression level was defined as ≤10% positive staining, and high Ki-67 expression level was defined as >10% positive staining [15].
Texture-based prediction of Ki-67 expression
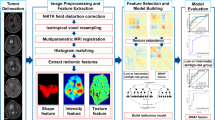
The pipeline in the radiomic analysis include imaging acquisition, tumor segmentation, feature extraction, feature selection and model construction. For the purpose of feature selection, we only screened significantly differential radiological features between the low and high Ki-67 expression groups (false discovery rate [FDR] <0.05) using the significance analysis of microarrays (SAM) algorithm. In order to further develop a Ki-67-related radiological signature, a generalized linear model (Y = β0 + β1X1 + β2X2 + … + βnXn) was established using the MATLAB (2014a) software. In this model, Y represented the estimated value of Ki-67 expression status (low and high expression levels were defined as 0 and 1, respectively); β0 indicated the unknown constant; βi (i = 1, 2, …, n) represented the unknown coefficients; and Xi indicated the values of radiological features. A cutoff value was chosen in order to classify the estimated values into the low and high Ki-67 expression groups. Subsequently, a receiver operating characteristic (ROC) curve was delineated to evaluate the predictive efficiency, and the optimal cutoff value was determined when specificity and sensitivity were maximal. This process was repeated in the training set, with the feature bearing the lowest predictive value being removed each time. A Ki-67 predictive model was constructed when the number of features was the lowest, and the AUC was greater than 0.90. The Ki-67 predictive model constructed from the training set was further evaluated in the validation set.
Statistical analysis
The SAM algorithm was applied using the “samr” package of the R software (version 3.3.2). Intergroup comparison of clinical characteristics was performed with T-statistics and Chi square test. Intergroup differences in survival were determined by log-rank and Kaplan–Meier analyses. Prognostic values of Ki-67 and Ki-67-associated radiological features were determined by Cox regression analysis. A p value <0.05 was considered statistically significant.
Results
Patient demographic characteristics
Magnetic resonance images were acquired of 117 patients with primary lower grade gliomas, including 78 with grade II (male, 44; age range 18–62 years) and 39 with grade III (male, 25; age range 18–71 years) gliomas. The enhancement exists in 80% grade III gliomas and no enhancement exists in grade II gliomas. These patients were allocated to the training (n = 78) and validation (n = 39) sets. There was no significant difference in age, sex, grade, Ki-67 expression level, or extent of resection between the two sets of patients (p > 0.05). Table 1 presents the detailed demographic and clinical characteristics of the patients.
Texture-based prediction of Ki-67 expression level
In the training set a total of 52 differential radiological features (FDR <0.05) related to Ki-67 expression level were screened by SAM analysis, including first-order statistics-derived features such as energy, maximum, minimum, and range; shape- and size-based features such as surface area, maximum 3D diameter, and spherical disproportion (SD); and textural features such as correlation and gray-level non-uniformity (Supplementary Table 2).
From among these 52 differential radiological features, a subset of nine features—including six wavelet features derived from first-order statistics (energy, range, and maximum); 1 second-order feature (SD); and 2 wavelet features derived from textural features (correlation and high gray-level emphasis)—was further selected by generalized linear analysis and evaluated in the training set for efficacy of prediction of Ki-67 expression status. The distribution of estimated values could be significantly differentiated using these nine features (Fig. 1a). In ROC analysis, the area under the ROC curve (AUC) was 0.916, and at the optimal cutoff point (Y = 0.584), the sensitivity, specificity, and accuracy were 77.3, 91.2, and 83.3%, respectively (Fig. 1b).
Estimated values and prediction efficiencies of the training and validation sets. a and c Distribution of estimated values of patients. Blue dots indicate the target values of patients. Green and red dots represent the estimated values of the low and high Ki-67 expression groups, respectively. b and d Receiver operating characteristic curves for prediction of Ki-67 expression level. In the training set, the area under the curve (AUC) was 0.916, and at the optimal cutoff point (Y = 0.584), the sensitivity, specificity, and accuracy were 77.3, 91.2, and 83.3% respectively. In the validation set, the AUC was 0.900, and the optimal cutoff value (Y = 0.672) exhibited a sensitivity, specificity, and accuracy of 70.2, 92.9, and 88.6% respectively
In the validation set, the radiomic signature constituted by the nine features could also efficiently predict Ki-67 expression levels and significantly differentiate the distribution of estimated values of patients in the validation set (Fig. 1c). In ROC analysis, the AUC was 0.900, and the optimal cutoff value (Y = 0.672) exhibited a sensitivity, specificity, and accuracy of 72.0, 92.9, and 88.6%, respectively (Fig. 1d).
Supplementary Fig. 2 presents two representative cases of patients with lower grade gliomas. The first case was that of a 25-year-old female patient with low Ki-67 expression, who was assigned to the low Ki-67 group on the basis of the estimated value being low (−0.11). Case 2 was that of a 39-year-old male patient with high Ki-67 expression, who was correctly assigned to the high Ki-67 group on the basis of the estimated value being high (0.96).
Relationship between Ki-67 expression level and survival
As shown in Fig. 2a and b, progression-free (PFS) and overall survival (OS) durations were significantly longer in patients with low Ki-67 expression than in patients with high Ki-67 expression in the whole cohort (p = 0.0012 and 0.0006, respectively).
Kaplan–Meier survival curves of different Ki-67 expression groups and patients stratified based on spherical disproportion (SD). Both Ki-67 expression level and the feature SD are significantly prognostic factors in progression-free survival (p = 0.0012 and p < 0.0001, respectively; a and c) and overall survival (p = 0.0006 and p < 0.0001, respectively; b and d) in the whole cohort
Prognostic significance of Ki-67-related radiological features
In the training set, the results of univariate Cox regression analysis with the nine radiological features (Table 2) revealed only SD to be a significant prognostic factor (p = 0.024). All subjects were divided into the low and high SD groups on the basis of the median SD value, and the prognostic value of SD was further investigated in the whole cohort. Patients in the high SD group exhibited poor prognosis (OS, p < 0.0001; PFS, p < 0.0001; Fig. 2c and d), indicating that SD level can help significantly differentiate the prognosis in patients with lower grade gliomas.
Multivariate Cox regression
The results of multivariate Cox regression analysis of SD, age, sex, WHO grade, extent of resection, and Ki-67 expression levels (Table 3) of the 117 patients revealed only SD and Ki-67 expression level to be independent prognostic factors of both PFS and OS (PFS: p = 0.017 and 0.049, respectively; OS: p = 0.019 and 0.019, respectively).
Discussion
In this study, Ki-67-related radiological features were analyzed using data retrieved from a large-scale genetic and imaging database. A set of radiological signatures that could predict Ki-67 expression levels was constructed by extraction and screening of radiological features using statistical models. In addition, a prognostic radiological feature was identified, which could non-invasively provide crucial prognostic information in patients with lower grade gliomas in the future.
Radiogenomics, an emerging field that focuses on the associations between imaging phenotypes and genomic features [16], has extensive application [9]. A pioneer quantitative radiogenomics work revealed a correlation between MRI features and cellular invasion in glioblastomas [17]. In patients with recurrent glioblastomas, radiological signatures could be used as biomarkers to identify patients who could most benefit from antiangiogenic therapy [18]. Radiogenomics could also help predict treatment-related molecular markers in gliomas—such as mutations in the isocitrate dehydrogenase 1 gene [19] and methylation of the O6-methylguanine methyltransferase gene promoter [20]—which could be beneficial for customizing therapeutics in the era of personalized medicine.
Several studies on gliomas have described the associations between radiological features and Ki-67 expression level. Evaluation by diffusion tensor imaging has revealed a strong correlation between ADC ratio and Ki-67 expression level [21]. The findings of MR spectroscopy have revealed choline/creatine and lactate/creatine ratios to be positively correlated with Ki-67 expression [22]. These findings suggest that Ki-67 expression level is closely associated with MRI features. In the present study, nine radiological features were screened by SAM and generalized linear analyses. The equation stemming from linear regression analysis could efficiently predict the Ki-67 expression status, which indicated the ability of the selected quantitative radiological features to reflect the Ki-67 expression status in lower grade gliomas.
The correlations between the selected quantitative radiological features and Ki-67 expression status can be interpreted on the basis of the biological behavior of tumors. While “range” represents the range of voxel intensity values, “maximum” refers to the maximum voxel intensity in the region of interest [14]. In the present study, these two features were significantly higher in the high Ki-67 expression group than in the low Ki-67 expression group. This might be attributable to the great variation in intratumoral gray values, because tumors with high Ki-67 expression levels corresponded with high proliferation rates and heterogeneity [11]. In addition, the index “high gray-level run emphasis” measures the distribution of high gray-level values, which are expected to be relatively high in regions of interest containing substantial areas with high-gray values [23]. This index was also significantly higher in the high Ki-67 group than in the low Ki-67 group, indicating that tumors in the former group contained more regions with high gray-levels, which might also be attributable to high heterogeneity.
Studies on the prognostic value of Ki-67 expression level have been widely conducted in recent years. A meta-analysis of 51 studies, involving a total of 4307 patients, demonstrated that Ki-67 overexpression was predictive of poor PFS and OS in patients with gliomas, regardless of region, cutoff value, pathological type, or statistical analysis method [24]. These findings are consistent with the present findings indicating that Ki-67 expression level to be an independent prognostic factor in patients with lower grade gliomas.
Additionally, as the sole prognostic feature in this signature, we further evaluated the index SD. Higher SD values are indicative of a greater deviation of the tumor from the spherical shape [14]. In the present study, the SD value of the high Ki-67 expression group was significantly higher than that of the low Ki-67 expression group, which suggests that high proliferative status is associated with greater irregularities in tumor shape in lower grade gliomas. Additionally, in the subsequent survival analysis, SD values exhibited a potent ability to predict the prognosis of the included patients. A previous study on glioblastomas reported that patients with spherical tumors exhibited better prognosis than those with irregularly shaped tumors [25]. The present study demonstrated a similar result in lower grade gliomas. Traditionally, clinical variables such as age, Karnofsky performance score, and extent of resection have been found to be predictive to the prognosis of glioma patients [1, 2]. The inclusion of radiological features would provide additionally important information for a better prediction of the survival outcomes and potentially guide the patient-tailored treatment in clinical practice.
The SAM algorithm used in this study was originally designed to identify differential genes in bioinformatics, with the advantage of providing FDR for multiple testing [26]. Generalized linear analysis is a liner regression method that can integrate various quantitative features into one analysis to produce a significantly reduced model with only a few significant predictors [27]. This method has been employed in studies of various medical fields. A previous study employed generalized linear analysis to integrate data from the infected host and the virus in order to identify predictors for liver damage [28]. In the present study, unimportant features were excluded one by one, which enabled us to achieve data dimensional reduction simply and efficiently. This is the first instance that generalized linear analysis has been used for processing radiological features for prediction of Ki-67 expression.
This study has a few limitations. Our study was designed on the basis of retrospectively collected single-institution data. The findings of this study need to be further evaluated in a prospective study with multicenter data. In addition, correlations between observed imaging features and biological function were not investigated in the current study. Lastly, as the most suitable and available sequence for the lesion evaluation of lower-grade gliomas, only T2-weighted MR images were used in the radiomic analysis. Multi-model imaging data (such as arterial spin-labeling, perfusion-weighted imaging, etc.) need to be integrated into our model in order to optimize the performance of the present model.
In conclusion, this study has demonstrated a correlation between radiomic features and Ki-67 phenotype in lower grade gliomas. By means of SAM and generalized linear analyses, we have established a radiomic signature that can efficiently and non-invasively predict Ki-67 expression levels. Furthermore, we found SD and Ki-67 expression level to be independent prognostic factors in patients with lower grade gliomas.
References
Cancer Genome Atlas Research N, Brat DJ, Verhaak RG et al (2015) Comprehensive, integrative genomic analysis of diffuse lower-grade gliomas. N Engl J Med 372:2481–2498. doi:10.1056/NEJMoa1402121
van den Bent MJ (2014) Practice changing mature results of RTOG study 9802: another positive PCV trial makes adjuvant chemotherapy part of standard of care in low-grade glioma. Neuro Oncol 16:1570–1574. doi:10.1093/neuonc/nou297
Kogiku M, Ohsawa I, Matsumoto K et al (2008) Prognosis of glioma patients by combined immunostaining for survivin, Ki-67 and epidermal growth factor receptor. J Clin Neurosci 15:1198–1203. doi:10.1016/j.jocn.2007.11.012
Wakimoto H, Aoyagi M, Nakayama T et al (1996) Prognostic significance of Ki-67 labeling indices obtained using MIB-1 monoclonal antibody in patients with supratentorial astrocytomas. Cancer 77:373–380
Duregon E, Bertero L, Pittaro A et al (2016) Ki-67 proliferation index but not mitotic thresholds integrates the molecular prognostic stratification of lower grade gliomas. Oncotarget 7:21190–21198. doi:10.18632/oncotarget.8498
Johannessen AL, Torp SH (2006) The clinical value of Ki-67/MIB-1 labeling index in human astrocytomas. Pathol Oncol Res 12:143–147
Zeng A, Hu Q, Liu Y et al (2015) IDH1/2 mutation status combined with Ki-67 labeling index defines distinct prognostic groups in glioma. Oncotarget 6:30232–30238. doi:10.18632/oncotarget.4920
Liu Y, Tang K, Yan W et al (2013) Identifying Ki-67 specific miRNA–mRNA interactions in malignant astrocytomas. Neurosci Lett 546:36–41. doi:10.1016/j.neulet.2013.04.030
Bai HX, Lee AM, Yang L et al (2016) Imaging genomics in cancer research: limitations and promises. Br J Radiol 89:20151030. doi:10.1259/bjr.20151030
Kuo MD, Jamshidi N (2014) Behind the numbers: Decoding molecular phenotypes with radiogenomics–guiding principles and technical considerations. Radiology 270:320–325. doi:10.1148/radiol.13132195
Ryu YJ, Choi SH, Park SJ et al (2014) Glioma: application of whole-tumor texture analysis of diffusion-weighted imaging for the evaluation of tumor heterogeneity. PLoS One 9:e108335. doi:10.1371/journal.pone.0108335
Liu C, Zhang H, Pan Y et al (2012) Towards MIB-1 and p53 detection in glioma magnetic resonance image: a novel computational image analysis method. Phys Med Biol 57:8393–8404. doi:10.1088/0031-9155/57/24/8393
Yan W, Zhang W, You G et al (2012) Molecular classification of gliomas based on whole genome gene expression: a systematic report of 225 samples from the Chinese Glioma Cooperative Group. Neuro Oncol 14:1432–1440. doi:10.1093/neuonc/nos263
Aerts HJ, Velazquez ER, Leijenaar RT et al (2014) Decoding tumour phenotype by noninvasive imaging using a quantitative radiomics approach. Nat Commun 5:4006. doi:10.1038/ncomms5006
Cai J, Zhang C, Zhang W et al (2016) ATRX, IDH1-R132H and Ki-67 immunohistochemistry as a classification scheme for astrocytic tumors. Oncoscience 3:258–265 doi:10.18632/oncoscience.317
Mazurowski MA (2015) Radiogenomics: what it is and why it is important. J Am Coll Radiol 12:862–866. doi:10.1016/j.jacr.2015.04.019
Zinn PO, Mahajan B, Sathyan P et al (2011) Radiogenomic mapping of edema/cellular invasion MRI-phenotypes in glioblastoma multiforme. PLoS ONE 6:e25451. doi:10.1371/journal.pone.0025451
Kickingereder P, Gotz M, Muschelli J et al (2016) Large-scale radiomic profiling of recurrent glioblastoma identifies an imaging predictor for stratifying anti-angiogenic treatment response. Clin Cancer Res. doi:10.1158/1078-0432.CCR-16-0702
Zhang B, Chang K, Ramkissoon S et al (2016) Multimodal MRI features predict isocitrate dehydrogenase genotype in high-grade gliomas. Neuro Oncol. doi:10.1093/neuonc/now121
Korfiatis P, Kline TL, Coufalova L et al (2016) MRI texture features as biomarkers to predict MGMT methylation status in glioblastomas. Med Phys 43:2835. doi:10.1118/1.4948668
Alexiou GA, Zikou A, Tsiouris S et al (2014) Correlation of diffusion tensor, dynamic susceptibility contrast MRI and (99 m)Tc-Tetrofosmin brain SPECT with tumour grade and Ki-67 immunohistochemistry in glioma. Clin Neurol Neurosurg 116:41–45. doi:10.1016/j.clineuro.2013.11.003
Fudaba H, Shimomura T, Abe T et al (2014) Comparison of multiple parameters obtained on 3 T pulsed arterial spin-labeling, diffusion tensor imaging, and MRS and the Ki-67 labeling index in evaluating glioma grading. AJNR 35:2091–2098. doi:10.3174/ajnr.A4018
Molina D, Perez-Beteta J, Martinez-Gonzalez A et al (2016) Influence of gray level and space discretization on brain tumor heterogeneity measures obtained from magnetic resonance images. Comput Biol Med 78:49–57. doi:10.1016/j.compbiomed.2016.09.011
Chen W-J, He D-S, Tang R-X et al (2015) Ki-67 is a valuable prognostic factor in gliomas: evidence from a systematic review and meta-analysis. Asian Pac J Cancer Prev 16:411–420. doi:10.7314/apjcp.2015.16.2.411
Itakura H, Achrol AS, Mitchell LA et al (2015) Magnetic resonance image features identify glioblastoma phenotypic subtypes with distinct molecular pathway activities. Sci Transl Med 7:303ra138. doi:10.1126/scitranslmed.aaa7582
Larsson O, Wahlestedt C, Timmons JA (2005) Considerations when using the significance analysis of microarrays (SAM) algorithm. BMC Bioinformatics 6:129. doi:10.1186/1471-2105-6-129
Soch J, Haynes JD, Allefeld C (2016) How to avoid mismodelling in GLM-based fMRI data analysis: cross-validated Bayesian model selection. NeuroImage 141:469–489 doi:10.1016/j.neuroimage.2016.07.047
Matas M, Picornell A, Cifuentes C et al (2013) Generalized Linear Model (GLM) framework for the association of host variables and viral strains with liver fibrosis in HCV/HIV coinfected patients. Infect Genet Evol 13:284–291 doi:10.1016/j.meegid.2012.10.020
Funding
This study was supported by the Beijing Natural Science Foundation (No. 7174295) and the National Natural Science Foundation of China (No. 81601452).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Yiming Li and Zenghui Qian contributed equally to this work and they are both the first authors of this article.
Electronic supplementary material
Below is the link to the electronic supplementary material.
11060_2017_2576_MOESM2_ESM.tif
Supplementary Fig. 2 Magnetic resonance images of two patients with lower glioma with different Ki-67 expression levels. Case 1 was that of a 25-year-old woman with low Ki-67 expression level, who was correctly classified into the low Ki-67 group on basis of the radiomic signature. Case 2 was that of a 39-year-old man with high Ki-67 expression level, who was correctly classified into the high Ki-67 group. (TIF 3856 KB)
Rights and permissions
About this article
Cite this article
Li, Y., Qian, Z., Xu, K. et al. Radiomic features predict Ki-67 expression level and survival in lower grade gliomas. J Neurooncol 135, 317–324 (2017). https://doi.org/10.1007/s11060-017-2576-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11060-017-2576-8