Abstract
Adventitious rooting (AR) is an obligatory step for vegetative propagation of commercial woody species. Paper industries have interest in Eucalyptus globulus Labill and its hybrids due to low lignin and lipid contents, which facilitate cellulose extraction. However, this species and some of its hybrids are recalcitrant to rooting, often requiring exogenous auxin supply. Here we performed a comparative analysis of proteome changes during AR of E. globulus and the easy-to-root species Eucalyptus grandis W. Hill and the effects of exogenous auxin in different phases of the process (induction and formation), using a label-free quantification method. We identified 398 differentially abundant proteins, which were predicted to be involved in different biological pathways, mainly oxidative stress, energy metabolism and photosynthesis. Notable differences between species included proteins involved in oxidative stress, carbon and secondary metabolism. Exogenous auxin appeared to affect the availability of carbon sources and other phytohormones, besides cell cycle and microtubule—related proteins. Important players were also identified in each phase of AR. This is the first in depth analysis of protein changes during AR in Eucalyptus. The findings can be used in future studies to evaluate rooting competence in different genotypes and provide leads for AR improvement.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Eucalyptus is planted worldwide due to its fast growth, high quality of fibers and high adaptability to different kinds of soils. The great overall diversity and adaptability of species of eucalypts make viable their cultivation for energy purposes even in relatively cooler areas, such as the UK (Leslie et al. 2019). Genetic transformation of eucalypts is also feasible, expanding their potential for adaptation to harsher environments (Thanananta et al. 2018). Eucalyptus globulus Labill is of particular interest due to the low lignin and lipid contents, which make cellulose extraction easier, decreasing production costs (Rencoret et al. 2007). However, E. globulus and its hybrids are often recalcitrant to rooting, making vegetative propagation difficult and causing loss of productivity (Serrano et al. 1996; Fett-Neto et al. 2001).
Propagation of economically important woody, horticultural and ornamental plants is essentially performed through rooting of leafy stem cuttings (adventitious rooting) and establishment of clonal gardens. Adventitious rooting (AR) is a complex process known to be affected by several factors such as phytohormone levels, phenolic compounds, light, wounding, genetic traits and nutritional conditions (da Costa et al. 2013). AR can be divided in two main phases: (1) induction, comprising the first molecular and biochemical events; and (2) formation, where the first cellular divisions occur, originating the root primordium, followed by root elongation (Fett-Neto et al. 2001).
Auxins are effective inducers of adventitious roots in different woody species (de Klerk et al. 1999). High concentrations of auxin are needed at the rooting zone during induction, but a decrease in concentration should occur during the formation phase, so roots can elongate. Local concentration of auxin at the base of cuttings seems to be important for triggering AR (da Costa et al. 2013). Endogenous auxin concentration was found to be lower in root recalcitrant species and clones than in their prone to root counterparts (Negishi et al. 2014; de Almeida et al. 2015).
Extensive work has been done to overcome rooting recalcitrance in Eucalyptus globulus and several improvements were obtained, allowing a better understanding of different factors controlling AR. When comparing E. globulus with the easy-to-root species E. saligna, Fett-Neto et al. (2001) found a positive effect of exposure of microcuttings to IBA (indole butyric acid), followed by culture in darkness. An optimized mineral nutrition can also improve AR, resulting in microcuttings better adapted to stresses and ex vitro conditions (Schwambach et al. 2005). Concerning donor plants environment and its effects on microcuttings they produce, it was found that, although mother plant exposure to long dark periods was detrimental to rooting (Corrêa et al. 2005), treatment with far-red enriched light increased rooting in absence of exogenous auxin in cuttings (Ruedell et al. 2013). However, this response was not observed in the easy-to-root E. grandis. The age of donor plants is also very important in E. globulus, as there is a significant early decline in rooting competence in microcuttings originated from in vitro aged individuals (Aumond et al. 2017).
Despite the findings related to morphological and physiological aspects of AR, molecular mechanisms controlling this process are still poorly known in woody plants. Positive or negative gene regulators of AR have been reported (Ramirez-Carvajal et al. 2009; Trupiano et al. 2013; Abu-Abied et al. 2014; de Almeida et al. 2015; Ruedell et al. 2015), but little is known about the involved proteins (Liu et al. 2013; Han et al. 2014; Wang et al. 2019). Challenges such as difficult extraction decrease data acquisition speed and availability. Protein-related research in trees focuses primarily on descriptive and differential proteomics (Abril et al. 2011).
In Eucalyptus, findings included proteins related to lignification (Celedon et al. 2007), osmotic (Bedon et al. 2011) and water stresses (Bedon et al. 2012; Valdés et al. 2013), heavy metal contamination (Guarino et al. 2014), diversification (Kersting et al. 2015), pathogen infection (Chen et al. 2015; Quecine et al. 2016), temperature (De Almeida Leonardi et al. 2015; De Santana Costa et al. 2017), seasonality (Budzinski et al. 2016a, b), and carbon assimilation (Santos and Balbuena 2017). Only a few studies have addressed proteome changes during AR, but none in Eucalyptus, representing an important information gap to be filled (Vilasboa et al. 2018).
To better understand AR in Eucalyptus, label-free qualitative and quantitative analysis was carried out identifying a large number of proteins differentially regulated during induction and formation phases in E. globulus (treated or not with auxin) and in the easy-to-root species E. grandis. These results may aid in identifying genotypes and protein patterns important for AR improvement strategies for Eucalyptus globulus.
Materials and methods
Cutting source
Seeds from E. globulus and E. grandis were prepared and germinated as previously described (Fett-Neto et al. 2001). After 3.5 months, tip microcuttings were severed and used for AR experiments.
AR conditions
Tip microcuttings were placed in rooting induction medium—MS salts (Murashige and Skoog 1962) 0.3X, 0.4 mg l−1 thiamin, 100 mg l−1 inositol and 30 g l−1 sucrose, presence (auxin treatment) or absence (control) of 10 mg l−1 indolyl-3-butyric acid (IBA), pH adjusted to 5.8 ± 1, and 0.6% (w/v) agar (Schwambach et al. 2005). As E. grandis is an easy-to-root species, these plants were submitted to control condition only. Rooting percentage was recorded after 20 days in formation media to confirm phenotypic differences between species and treatments. Plants were harvested after 1, 2 and 4 days in induction medium and mixed for protein extraction. For analysis during the formation phase of AR, plants were kept 4 days in induction medium and then transferred to formation medium (same medium as induction, but without auxin and containing 1 g l−1 activated charcoal). Plants were harvested after 1, 2 and 4 days in formation medium and mixed for protein extraction. The experiments were repeated three times with similar results. Six whole plants were used for each time-point (days of harvesting), totalizing 18 plants in induction and 18 plants in formation phase, for both control and auxin-treatment conditions. Although there is a dilution effect of using whole plants for rooting studies, this approach was chosen to achieve the needed yield for downstream protein analyses. Moreover, potentially relevant correlative interactions of shoots and the AR zone could be included in the analyses.
Protein extraction and quantification
Protein extraction was from three different biological samples (weighing 0.3 g FM each) using the 10% trichloroacetic acid (TCA)/acetone precipitation method (Damerval et al. 1986) with some modifications. Samples (whole plants) were ground under liquid nitrogen and suspended in 1 mL of cold extraction buffer with 10% (w/v) TCA (Sigma-Aldrich, St. Louis, USA) in acetone with 20 mM dithiothreitol (DTT; GE Healthcare, Little Chalfont, U.K.). Mixtures were kept at − 20 °C for 1 h and then centrifuged at 16,000 g for 30 min at 4 °C. Pellets were washed three times with cold acetone containing 20 mM DTT and air dried. Next, pellets were resuspended in buffer containing 7 M urea, 2 M thiourea, 1% DTT, 1 mM phenylmethylsulfonide, and 2% Triton X-100 (Sigma-Aldrich, St. Louis, USA), vortexed and incubated on ice for 30 min, and then centrifuged at 16,000 g for 20 min at 4 °C. Supernatants were recovered and their protein concentrations determined using a 2-D Quant Kit (GE Healthcare, Little Chalfont, U.K.).
Protein digestion
Samples were desalted using Amicon Ultra-0.5 3 kDa centrifugal filters (Merck Millipore, Germany). Filters were filled to maximum capacity with buffers and centrifuged at 15,000 g for 10 min at 25 °C. Washing was done three times with 50 mM ammonium bicarbonate (Sigma-Aldrich, St. Louis, MO, USA) pH 8.5; approximately 50 μL remained per sample after the last wash. Protein digestion followed Calderan-Rodrigues et al. (2014). For each sample, 25 μL of 0.2% (v/v) RapiGest® (Waters, Milford, CT, USA) were added, samples were quickly vortexed and incubated in an Eppendorf Thermomixer® at 80 °C for 15 min. Next, 2.5 μL of 100 mM DTT (GE Healthcare, Little Chalfont, UK) were added, followed by vortexing and incubation at 60 °C for 30 min under agitation. Then, 2.5 μL of 300 mM iodoacetamide (GE Healthcare, Little Chalfont, UK) were added, samples were vortexed and incubated in the dark for 30 min at RT. Twenty μL of trypsin solution (50 ng/μL; V5111, Promega, Madison, WI, USA) prepared in 50 mM ammonium bicarbonate was added, followed by sample incubation at 37 °C for 15 h. RapiGest® precipitation and trypsin inhibition were carried out by addition of 10 μL of 5% (v/v) trifluoroacetic acid (TFA, Sigma-Aldrich, St. Louis, MO, USA) to samples, followed by maintenance at 37 °C for 30 min, and subsequent centrifugation for 20 min at 16,000 g. Next, samples were transferred to Total Recovery Vials (Waters, Milford, MA, USA).
Mass spectrometry analysis
A nanoAcquity UPLC (Waters, Milford, MA, USA) connected to a Synapt G2-Si (Waters, Milford, MA, USA) mass spectrometer (Waters, Milford, MA, USA) was employed for ESI-LC–MS/MS analysis. Chromatography was done by injecting 1 μL of digested samples (500 ng/μL) in order to normalize them before relative protein quantification. To ensure standardized molar values for all experimental conditions, normalization among samples was supported by stoichiometric measurements of total ion counts of MSE scouting runs before the analyses using the ProteinLynx Global Server v. 3.0 platform (PLGS; Waters, Milford, MA, USA). After sample normalization, the HDMSE runs consisted of three biological replicates per treatment. During separation, samples were applied on the nanoAcquity UPLC 5 μm C18 trap column (180 μm × 20 mm) at 5 μL/min for 3 min and then on the nanoAcquity HSS T3 1.8 μm analytical reversed phase column (75 μm × 150 mm) at 400 nL/min, at 45 °C column temperature. For peptide elution, a binary gradient was used, with mobile phase A (water - Tedia, Fairfield, OH, USA and 0.1% formic acid - Sigma-Aldrich) and mobile phase B (acetonitrile and 0.1% formic acid, both from Sigma-Aldrich, St. Louis, MO, USA). Gradient elution started at 7% B, then increased from 7% B to 40% B up to 91.12 min, and from 40% B to 99.9% B up to 92.72 min, being kept at 99.9% until 106.00 min, then decreasing to 7% B until 106.1 min and kept at 7% B until run completion at 120.00 min. Mass spectrometry was conducted in positive and resolution mode (V mode), 35,000 FWHM, with ion mobility, and in mode of data-independent acquisition (DIA). Ion mobility separation (HDMSE) was done using IMS wave velocity of 600 m/s, and helium and IMS gas flow of 180 and 90 mL/min respectively. Transfer collision energy was raised from 19 V to 55 V in high-energy mode; cone and capillary voltages were 30 V and 2750 V, respectively; and temperature source was 70 °C. In TOF parameters, the scan time was set to 0.5 s in continuum mode and mass range of 50–2000 Da. The human [Glu1]-fibrinopeptide B (Sigma-Aldrich, St. Louis, MO, USA) at 100 fmol/μL was used as an external calibrator, whereas lock mass acquisition was obtained every 30 s. Mass spectra acquisition was analyzed with MassLynx v4.0 software.
The mass spectrometry proteomic data have been deposited in the ProteomeXchange Consortium (Vizcaíno et al. 2014) via the PRIDE (Vizcaíno et al. 2016) partner repository with the dataset identifiers PXD013924 and PXD013925.
Proteomics and rooting data analysis
Proteins that were regulated during AR in control and auxin-treated plants of E. globulus and in control plants of E. grandis were identified using specific algorithms (Silva et al. 2005). Proteins obtained from each comparison group were organized into a statistically significant list corresponding to regulated (increased and decreased ratios) and unchanged ratios. Analyses were made separately with data from AR induction phase and AR formation phase and data normalization was performed as previously described (Heringer et al. 2015).
Spectra processing and database searching conditions were carried out with Progenesis QI for Proteomics Software V.2.0 (Nonlinear Dynamics, Newcastle, UK) according to Heringer et al. (2015). The following parameters were set: Apex3D of 150 counts for low energy threshold, 50 counts for elevated energy threshold, and 750 counts for intensity threshold; 1 missed cleavage, 2 minimum fragment ion per peptide, 5 minimum fragment ion per protein, 2 minimum peptide per protein, fixed modifications of carbamidomethyl (C) and variable modifications of oxidation (M) and phosphoryl (STY), and a default false discovery rate (FDR) value at four percent maximum, peptide score higher than four, and maximum mass errors of 10 ppm. The confidence score in the Progenesis QI for Proteomics Software V.2.0 is the mean score of the following properties: Mass error, Isotope distribution similarity, Retention time error, CCS (collisional cross section) error, and Fragmentation score. Samples with confidence scores lower than four are considered too noisy and are removed from the analyses (Progenesis QI for Proteomics User Guide).
Since at present there is no available complete annotated data for Eucalyptus globulus, the analysis used the Eucalyptus grandis v 2.0 protein databank of Phytozome (Goodstein et al. 2012). Label-free relative quantitative analyses were done based on the ratio of protein ion counts among contrasting samples. Proteomic experiments had three biological replicates of 6 plants each. After data processing, in order to ensure the quality of results, only proteins present or absent (for unique proteins) in all biological replicates of each treatment were considered. Based on the results of ANOVA (p < 0.05), differentially abundant proteins were regarded as upregulated when fold change (FC) was higher than 1.5 and downregulated if FC was lower than 0.67. Functional annotation used Blast2Go software v. 3.4 (Conesa et al. 2005). Rooting percentages were obtained from two independent experiments, each with 4 replicates of 5 microcuttings, analyzed by Welch ANOVA followed by Dunnet-C test (p < 0.05).
Results
Rooting profile
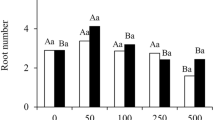
Rooting responses matched the expected differences between E. grandis (easy-to-root) and E. globulus (hard-to-root) (Fig. 1). The former species had over fivefold more rooting of cuttings than the latter. In addition, exposure of E. globulus microcuttings to exogenous auxin yielded rooting percentage equivalent to that of E. grandis control condition (Fig. 1).
Adventitious rooting phenotypes of E. globulus and E. grandis. AE. globulus in control condition, showing no roots. B Phenotype of E. globulus treated with auxin or E. grandis in control condition, showing presence of adventitious roots (arrows). Scale bar in A and B: approximately 1 cm. C Rooting percentage in E. globulus and E. grandis. Bars sharing the same letters are not different according to Dunnet-C test with p ≤ 0.05. Error bars correspond to standard deviation. Control and auxin labels indicate absence and presence of exogenous auxin treatment, respectively
Protein identification during induction and formation phases of AR
A total of 1424 protein hits were found, with an average of eight peptides per protein. Of these hits, 427 were common (unchanged) to all induction samples (Online Resource 1) and 395 were common to all formation samples (Online Resource 2). On the other hand, there were 288 differentially abundant protein hits during induction phase, four of them being unique to E. globulus samples when compared to E. grandis (Table 1 and Online Resource 3). Out of the 314 differentially abundant protein hits identified during the formation phase, there were two unique proteins to E. globulus and one to E. grandis (Table 1 and Online Resource 4). After analysis of all regulated hits, 84 proteins were regulated only in induction and 110 regulated only in formation samples, being 204 proteins common to both phases, totalizing 398 differentially abundant proteins.
The unique E. globulus induction phase proteins included a Bark storage protein A-like (Eucgr.C02912.1.p), heat shock protein (Eucgr.C02653.1.p), alkenal reductase with oxidoreductase activity (Eucgr.G01686.1.p; EC 1.3.1.74) and a protein of unknown function (Eucgr.K01872.1.p) (Table 1). During the formation phase of AR, a unique E. grandis protein was identified as aquaporin (Eucgr.J00237.2.p) and two E. globulus unique proteins were the same heat shock and alkenal reductase proteins found during the induction phase (Table 1).
Analysis of the differentially abundant proteins between each group of comparison during the induction phase revealed 8 downregulated and 26 upregulated proteins in E. globulus auxin-treated plants when compared to E. globulus control plants (Online Resource 3). When comparing E. globulus auxin-treated plants to E. grandis control plants, we found 125 down-regulated and 74 upregulated proteins (Online Resource 3). Comparing control conditions in both species, 172 down-regulated and 81 upregulated proteins in E. globulus when compared to E. grandis were found (Online Resource 3). During the formation phase of AR, E. globulus auxin-treated plants had 56 downregulated proteins and 83 upregulated proteins when compared to E. globulus control plants (Online Resource 4). When comparing E. globulus auxin-treated plants with E. grandis control plants, there were 112 down-regulated and 99 upregulated proteins in the former (Online Resource 4). Finally, the comparison between control plants during formation phase revealed that E. globulus had 128 down and 77 upregulated proteins when compared to E. grandis (Online Resource 4). Overall more proteins were identified during the formation phase compared to induction (314 vs 288, respectively).
Functional classification of identified proteins
The 602 identified hits belong to different biological pathways, with 2% (13 hits) corresponding to unknown proteins. During both induction and formation phases of AR, most of the identified proteins were related to oxidation–reduction processes (22%—130 hits), followed by energy metabolism (16%—99 hits) and photosynthesis (11%—65 hits) (Fig. 2, Online Resources 5 and 6).
During induction, among the proteins involved in oxidative stress, the Cu–Zn isoform of superoxide dismutase (SOD; EC 1.15.1.1) and peroxidase 5-like (EC 1.11.1.7) were downregulated in E. grandis when compared to E. globulus samples. On the other hand, during the same AR phase, Fe isoform of SOD (EC 1.15.1.1), peroxidase 12-like and 3-like (EC 1.11.1.7) were downregulated in both control and auxin-treated E. globulus when compared to E. grandis (Online Resource 5). At the root formation step, SOD isoforms and peroxidase 12-like were downregulated, whereas peroxidase 5-like remained upregulated in E. globulus relative to E. grandis (Online Resource 6). Isoflavone-reductases (EC 1.3.1.45) were differentially abundant in all samples and species, with most hits showing upregulation in E. globulus compared to E. grandis in both phases of AR (Online Resources 5 and 6).
In agreement with the marked presence of proteins related to oxidation–reduction processes, 3% of proteins detected were involved in defense responses (Fig. 2, Online Resources 5 and 6). Unlike the profile of expression of defense-related proteins, photosynthesis-associated proteins had lower expression in E. globulus when compared to E. grandis, mostly during the induction phase (Online Resource 5), but also for the majority of proteins in this class of function at the formation step (Online Resource 6).
In the energy metabolism pathway category, we found proteins involved in the homeostasis of carbon such as soluble sugars and starch metabolism and enzymes involved in glycolytic reactions (Table 2). Two hits were identified as fructose-1,6-bisphosphatase chloroplastic (Eucgr.I02616.1 and Eucgr.B02755.1; EC 3.1.3.11), both of which have shown similar expression patterns, being more abundant in conditions relatively less favorable to rooting. Eucgr.I02616.1 was more abundant in control and auxin-treated E. globulus when compared to E. grandis. In contrast, Eucgr.B02755.1 had higher accumulation in E. globulus control samples when compared to auxin-treated ones (Table 2). In starch-related pathways, three plastidic proteins were upregulated in E. grandis control compared to the same treatment in E. globulus: glucose-1-phosphate adenylyltransferase (Eucgr.K01372.1; EC 2.7.7.27), glucose-6-phosphate isomerase (Eucgr.F02133.1; EC 5.3.1.9) and a fructokinase (probable fructokinase-chloroplastic—Eucgr.F02835.1; EC 2.7.1.4). The first protein was upregulated only during induction and the others were upregulated in both induction and formation phases (Table 2). Concerning sucrose metabolism, we detected upregulation of a sucrose synthase (Eucgr.C03205.1; EC 2.4.1.13) in E. globulus during the formation phase, in response to exogenous auxin treatment. As far as glycolysis, five hits were identified for fructose-bisphosphate aldolase (EC 4.1.2.13), being three cytoplasmic (Eucgr.B03891.1, Eucgr.K02073.1, Eucgr.A01538.1) and two plastidic isoforms (Eucgr.I01326.1, Eucgr.B02864.1), which, for the most part, remained unchanged (Table 2).
Galactose metabolism was represented by two UDP-glucose 4-epimerases GEPI48 (Eucgr.D01750.1 and Eucgr.E00872.1; EC 5.1.3.2). Eucgr.D01750.1 was upregulated in E. globulus and Eucgr.E00872.1 in E. grandis (Table 2). Another protein induced in E. globulus was UDP-glucose 6-dehydrogenase 1 (Eucgr.J01372.1; EC 1.1.1.22), which was upregulated in control samples during the induction phase (Table 2).
Although quantitatively less represented (53 hits in total, corresponding to 9% of identified hits) (Fig. 2), several differentially regulated proteins related to biological pathways important for rooting competence and root architecture establishment were observed, including phytohormone signaling (Table 3), microtubule and cell wall organization (Table 4) and secondary metabolism (Table 5). As far as phytohormones, the findings include proteins involved in biosynthesis and signaling of auxin, abscisic acid (ABA), salicylic acid (SA), and jasmonic acid (JA) (Table 3). The auxin-binding protein ABP19a-like (Eucgr.C03537.1) was upregulated in E. globulus samples treated with exogenous auxin when compared to E. globulus control (induction phase) or E. grandis samples (formation phase) (Table 3). Exogenous auxin was associated with downregulation of the Indole-Acetic Acid (IAA) biosynthesis-related protein amidase 1 (Eucgr.B01994.1; EC 3.5.1.4) during formation phase and upregulation of an aspartate aminotransferase (AAT; EC 2.6.1.1) (Eucgr.F02571.1) during induction phase compared to E. globulus control. AAT was also upregulated in E. grandis, but in both phases of AR (Table 3). Two hits were identified as the same salicylic acid-binding 2-like protein (Eucgr.I01013.1 and Eucgr.I01014.1). Eucgr.I01013.1 was upregulated in E. globulus auxin treated samples during formation phase and Eucgr.I01014.1 was upregulated in E. grandis samples in both induction and formation phases (Table 3). The ABA-related proteins zeaxanthin epoxidase (Eucgr.I00922.1; EC 1.14.15.21) and annexin D1 (Eucgr.F02411.1) were also upregulated in E. grandis. On the other hand, the abscisic stress-ripening 1-like protein (Eucgr.F04219.1) was downregulated in response to auxin (Table 3). JA biosynthetic-related protein peroxisomal fatty acid beta-oxidation multifunctional AIM1 (Eucgr.C01261.1) was downregulated during induction phase in the easy-to-root species E. grandis (Table 3).
A cell division protein (Eucgr.K01258.1) was upregulated during formation phase of AR in E. globulus auxin-treated samples (Table 4). The cell wall biogenesis protein xylosidase (EC 3.2.1.37) was found three times during formation phase (Eucgr.F02441.1, Eucgr.G00649.1, Eucgr.G00651.1), being regulated in different ways, for the most part downregulated by auxin treatment and upregulated in E. grandis (Table 4). The same profile was observed for microtubule-related proteins actin (Eucgr.G02932.1 and Eucgr.J00015.2) and alpha (Eucgr.B03604.1) and beta tubulin (Eucgr.K01666.1 and Eucgr.K00264.1), which were also differently abundant depending on phase of AR (Table 4). Overall, these proteins were downregulated in presence of auxin and upregulated in E. grandis. A kinesin protein (kinesin KIF22 isoform X2-Eucgr.K00277.1) was upregulated in auxin-treated samples during formation phase (Table 4).
Secondary metabolism was represented by the differential expression of proteins involved in lignin, flavonoid and terpene metabolism (Table 5). The enzyme laccase-9 (Eucgr.F04160.1; EC 1.10.3.2), involved in the biosynthesis of lignin, was upregulated in E. grandis when compared to E. globulus control samples, during both induction and formation phases (Table 5). The flavonoid metabolism enzyme anthocyanidin reductase (Eucgr.D00858.1; EC 1.3.1.77) was downregulated in E. grandis (Table 5). On the other hand, the application of exogenous auxin induced the upregulation of a chalcone isomerase (Eucgr.F03816.1; EC 5.5.1.6) during formation phase in E. globulus auxin-treated samples when compared to E. globulus control samples, whereas the same enzyme was upregulated in E. grandis (Table 5).
Discussion
The overall higher number of identified proteins during the formation phase (314 vs 288 during induction) probably reflects the important cellular changes that occur during this phase leading to the development and elongation of adventitious roots. In the comparison between both E. globulus control and auxin-treated conditions with E. grandis, more proteins were downregulated, possibly as a result of the attenuation or shut down of production of AR repressing-related proteins in the easy-to-root species, whose corresponding transcripts were shown to be more expressed in the hard-to-root counterpart in tissue specific evaluations (de Almeida et al. 2015). This condition would allow higher expression of proteins related to AR development in the easy-to-root species.
Finding of proteins involved in stress response during AR is expected, mainly due to the wounding caused by cutting severance (Ahkami et al. 2009). Wounding increases reactive oxygen species and polyphenols, which can modulate AR (Steffens and Rasmussen 2016).
Superoxide dismutase expression indicates dismutation of superoxide and production of hydrogen peroxide (H2O2) during AR (Zelko et al. 2002). H2O2 may act as AR signal (Santos Macedo et al. 2012; Li et al. 2017) but it also needs to be controlled, both as signal and oxidative agent. This could explain the overall upregulation of these H2O2 generating enzymes in E. grandis samples when compared to E. globulus ones, affording finer regulation of superoxide and H2O2 pools in the former. The presence of defense response proteins in both phases of AR is also relevant after wounding.
Peroxidase activity can be a biochemical marker of adventitious root development in trees (Fett-Neto et al. 1992), including E. globulus (Schwambach et al. 2008; Aumond et al. 2017). Peroxidase 5-like is involved in H2O2 and perhaps auxin catabolism at cutting bases, decreasing rooting capacity, consistent with its upregulation during induction and formation in E. globulus, when compared to E. grandis. This concurs with lower H2O2 production in the former species, which has lower amounts of SOD. Two other peroxidases (Peroxidase 3-like and Peroxidase 12-like), involved in the biosynthesis of polyphenols (e.g. matairesinol), were upregulated in E. grandis, possibly favoring AR by avoiding auxin degradation (De Klerk et al. 2011; Steffens and Rasmussen 2016).
Isoflavone reductase is involved in the biosynthesis of phenylpropanoid-derived defense compounds, sharing similarities with lignan biosynthetic enzymes (Min et al. 2003). The accumulation of phenolic acids and flavonoids was correlated with in vitro cutting rooting, as these compounds can modulate peroxidase activity and prevent auxin degradation (de Klerk et al. 1999). Isoflavone reductase was found in the cambium region of E. grandis (Celedon et al. 2007), an important location for the origin of adventitious roots (de Almeida et al. 2015).
The increased expression of defense proteins (including isoflavone reductase-like proteins) throughout rooting in E. globulus relative to E. grandis may reflect a slower and/or limited healing response to wounding. This could restrict rooting capacity in the former also as a result of energy, carbon and nitrogen diversion to defense. This is in line with the predominantly lower expression of photosynthesis related-proteins in E. globulus compared to E. grandis along the rooting phases.
Overall presence of photosynthesis related proteins was probably increased by the use of whole plants for protein extraction. Most photosynthesis-related proteins were less abundant in E. globulus when compared to E. grandis, contrasting with findings in Arabidopsis, where several plastid-encoded proteins were accumulated in poor rooting genotypes (Sorin et al. 2006). However, microarray analyses in Pinus contorta showed down-regulation of plastid protein transcripts, which was tentatively explained by the loss of photosynthetic capacity of hypocotyl cells during AR (Brinker et al. 2004).
Unique proteins
Bark storage proteins are involved in nitrogen dynamics in temperate hardwoods, decreasing during cambium activation and increasing during dormancy (Druart et al. 2007), wounding, and methyl jasmonate treatment (Beardmore et al. 2000). Presence of this protein in E. globulus probably reflects a reduced or slower N mobilization capacity during AR. Lower abundance of JA biosynthetic-related protein peroxisomal fatty acid beta-oxidation multifunctional AIM1 (Eucgr.C01261.1) in E. grandis during root induction is in agreement with less bark storage protein (Table 3).
Heat shock proteins (HSPs) are essential in cells. Some can act as chaperones in protein folding, assembly, translocation, and degradation (Park and Seo 2015). HSP90 is related to plant development (Xu et al. 2012), involved in responses to auxins and brassinosteroids (Amzallag and Goloubinoff 2003; Wang et al. 2016). The finding of this protein in E. globulus suggests a role for HSP90 in hormonal homeostasis during AR.
Alkenal reductases are involved in detoxifying lipid peroxide-derived reactive carbonyls (RCs), preserving respiration and growth under oxidative stress (Mano et al. 2005; Yamauchi et al. 2012). Detection of these proteins in E. globulus highlights the need to control oxidative stress associated with severance wounding.
The only protein found exclusively in E. grandis was an aquaporin (Probable aquaporin PIP-type 7a—Eucgr.J00237.2). Aquaporins are involved in water uptake and are induced by auxin during lateral root formation to facilitate emergence (Péret et al. 2012). In poplar AR, transcripts encoding PIP aquaporins were highly expressed in root primordium (Kohler et al. 2003), which agrees with the protein accumulation in the easy-to-root eucalypt during the formation phase.
Carbon metabolism
Proteins involved in energy metabolism, encompassing carbon biochemistry-related enzymes among several others, were highly represented. This is consistent with AR proteome analyses in Arabidopsis, Chrysanthemum, hybrid larch and apple (Han et al. 2014; Lei et al. 2018; Liu et al. 2013; Sorin et al. 2006) and rooting in Petunia hybrida and mung bean (Ahkami et al. 2014; Li et al. 2017), highlighting the important roles of energy metabolism-related proteins and enzymes in resources accumulation and distribution. Carbon metabolism during rooting is paramount, as it provides necessary energy and carbon skeletons to support cell division, establishment of the new root meristems and root formation (da Costa et al. 2013). Soluble sugars can positively regulate the development of adventitious roots (Steffens and Rasmussen 2016).
Fructose-1,6-bisphosphatase converts fructose-1,6-bisphosphate to fructose-6-phosphate and Pi in an irreversible way. The cytosolic form is involved in sucrose synthesis whereas the plastidic form plays a regulatory role in CO2 assimilation (Chueca et al. 2002) and starch synthesis (Rojas-González et al. 2015). In IBA-exposed apple cuttings, upregulation of four proteins identified as fructose-1,6-bisphosphatase was reported (Lei et al. 2018). During AR promotion by etiolation in black locust, genes encoding fructose-1,6-bisphosphatase had increased expression, presumably leading to induction of sucrose biosynthesis and improved formation of adventitious roots (Lu et al. 2017). Decrease of cytosolic fructose-1,6-bisphosphatase activity was observed over time in Petunia hybrida cuttings and starch accumulation was seen later in AR (Ahkami et al. 2009). Here two proteins identified as chloroplastic fructose-1,6-bisphosphatase showed similar expression patterns (increased in the recalcitrant species). This corroborates previous findings, as the content of soluble sugars and starch was higher in E. globulus than in E. grandis microcuttings (Ruedell et al. 2013). In contrast, auxin treated E. globulus showed lower accumulation of chloroplastic fructose-1,6-bisphosphatase (Eucgr.B02755.1) than its untreated control. Increased chloroplastic fructose-1,6-bisphosphatase was observed in both induction and formation steps, often the latter (Table 2).
Two starch-related proteins from chloroplast were upregulated in E. grandis. Eucgr.K01372.1 is involved in starch biosynthesis and was more abundant in the easy-to-root species only during induction phase in control condition compared to E. globulus. This enzyme was induced in apple cuttings by exogenous auxin (Lei et al. 2018). In turn, Eucgr.F02133.1 was upregulated in both induction and formation phases, being involved in starch accumulation in plastids (Grauvogel et al. 2007).
A fructokinase was also upregulated in E. grandis in both rooting phases. Transcription of the gene encoding this enzyme was induced in microcuttings of E. globulus derived from donor plants exposed to far-red light (a condition that promotes adventitious root development) during early induction step of AR (Ruedell et al. 2015). Fructokinase was also upregulated in etiolated cuttings of black locust (Lu et al. 2014) and in IBA-treated apple cuttings (Lei et al. 2018).
Considering the induction of fructokinases in samples prone to rooting, these results suggest a positive role of these enzymes in the homeostasis of carbohydrates in AR. Results suggest that starch is acting as the main source of energy during AR, its availability being directly correlated with rooting. Other factors could affect starch use in E. globulus contributing to recalcitrance.
Transcripts of sucrose synthase, which converts sucrose into UDP-glucose and fructose (Rolland et al. 2006), were upregulated in carnation cuttings early in AR (Villacorta-Martín et al. 2015). In E. globulus microcuttings from donor plants exposed to far-red light, sucrose synthase gene expression increased during formation phase (Ruedell et al. 2015). Herein, the enzyme was upregulated in E. globulus in response to exogenous auxin treatment during formation. Positive interactions between glucose and auxin in controlling development of roots in Arabidopsis (Mishra et al. 2009) and of adventitious roots in eucalypts (Corrêa et al. 2005) have been reported. Proteome data further support a role for this interaction in AR regulation of Eucalyptus.
Fructose-bisphosphate aldolase correlated negatively with adventitious root number and free IAA content in Arabidopsis and Chrysanthemum (Sorin et al. 2006; Liu et al. 2013). This profile appeared to be matched by the cytosolic isoform Eucgr.B03891.1, which was overall less abundant in E. grandis. However, transcript of a plastidic isoform was upregulated during in AR of Petunia hybrida at the time of formation of the first new meristematic cells, probably involving activation of glycolysis in the chloroplast to support starch biosynthesis (Ahkami et al. 2014). Five hits for fructose-bisphosphate aldolase were detected, being three cytoplasmic (Eucgr.B03891.1, Eucgr.K02073.1, Eucgr.A01538.1) and two plastidic isoforms (Eucgr.I01326.1, Eucgr.B02864.1).
UDP-glucose 4-epimerase is involved in galactose metabolism, important for cell wall synthesis. Transcriptome analysis during AR in P. contorta, revealed induction of a UDP-glucose 4-epimerase at the stage of new root meristem formation (Brinker et al. 2004). In Arabidopsis, the enzyme was found to be important for root epidermal cells (Wang et al. 2015b). Considering the evidence for main action of this enzyme in root meristem formation, our results suggest that Eucgr.D01750.1 acts preferentially in E. globulus and Eucgr.E00872.1 in E. grandis. UDP-glucose 6-dehydrogenase is responsible for the conversion of UDP-glucose in UDP-glucuronic acid, from which monomers for pectic cell-wall polysaccharides derive (Samac et al. 2004). Transcriptional studies support a role for this enzyme in providing precursors for cell wall biosynthesis (Seitz et al. 2000). Considering that during the induction phase of AR cell walls should be weakened and extensively remodeled, allowing cell growth and expansion, the upregulation of UDP-glucose 6-dehydrogenase 1 in E. globulus control samples during induction phase suggests that the activation of cell wall synthesis at this stage could contribute for the hard-to-root phenotype of this species. In addition, the exogenous supply of auxin seems to reverse this trend.
Phytohormones
The role of phytohormones in the control of AR can be considerably variable, depending mainly on species, system (cuttings, hypocotyls, petioles), phase, and culture conditions (da Costa et al. 2013; Druege et al. 2016).
Auxins are known as major regulators of rooting (Pacurar et al. 2014). Three auxin-related proteins were found in the present study, one involved in signaling and two involved in biosynthesis. Auxin-binding protein ABP19a-like was upregulated in E. globulus in response to exogenous auxin. In mung bean cuttings (easy-to-root) treated with water or H2O2, increased expression of ABP19 gene from 6 h to 24 h after treatment was observed (Li et al. 2015, 2017). ABP19a-like probably acts as an auxin binding molecule similar to the well known ABP1, which could modulate fast responses to auxin (Tromas et al. 2009). Transcriptome profiles during AR in E. globulus and E. grandis revealed higher expression of ABP1 in E. grandis cuttings treated with exogenous auxin during the induction phase (de Almeida et al. 2015). Considering that ABP1 and ABP19 are from the same family and could have overlapping functions, these results suggest that exogenous auxin could induce the accumulation of auxin binding proteins, potentially affecting water uptake and cellular expansion (Tromas et al. 2009).
Exogenous auxin led to down-regulation of amidase 1 during the formation phase. This protein is involved in IAA biosynthesis via IAM (indole-3-acetamide) (Zhao 2010), a process that would be less required upon exogenous supply of auxin. Repression of auxin production during the formation phase is expected to allow root elongation (de Klerk et al. 1999).
Aspartate aminotransferase (AAT) is involved in the regulation of biosynthesis of auxin and ethylene. Genetic data showed that AAT acts downstream of TAA1 (conversion of Trp to indole-3-pyruvic acid, IPA) and upstream of YUCCA (conversion of IPA to IAA), negatively modulating IAA biosynthesis by altering IPA pool (Zheng et al. 2013). Upregulation of an AAT in E. globulus auxin-treated samples during induction phase and in E. grandis in both phases of AR was recorded, suggesting decrease in auxin biosynthesis. Similar result was found in IBA-treated cuttings of black locust, where an AAT was upregulated at early stages of AR (Quan et al. 2017). Taking into account that E. grandis has significantly higher concentration of total IAA compared to E. globulus (de Almeida et al. 2015), it is possible that ATT acts on the modulation of an auxin pool favorable to rooting. AAT also converts oxaloacetate in aspartate (Melzer and O’Leary 1987). The gene encoding AAT was induced at early AR stages in petunia cuttings, along with aspartate level, indicating its importance during rooting (Ahkami et al. 2014). Further studies are necessary to better understand the specifics of aspartate and ATTs in the regulation of AR.
SA may have positive roles in AR (Pacurar et al. 2014). The upregulation of two salicylic acid-binding 2-like proteins in auxin-treated E. globulus and in untreated E. grandis microcuttings reinforces the idea of a crosstalk between auxin and SA (Agulló-Antón et al. 2014) in AR, as both proteins were induced in samples showing rooting competent phenotypes.
Most studies have shown ABA negative effect on AR (da Costa et al. 2013). Surprisingly, proteomic analyses of IBA-induced AR in apple cuttings showed increase in ABA content and ABA-related proteins in IBA-treated samples (Lei et al. 2018). Our results are in agreement with these findings, as most ABA response, signaling and biosynthesis-related proteins identified were predominantly upregulated in easy-to-root E. grandis. However, one of the proteins involved in response to ABA (Eucgr.F04219.1) was downregulated in response to auxin. ABA has multiple roles, ranging from cell division inhibition to water stress adaptive responses both of them relevant for cutting rooting. Its participation in root architecture and lateral root development is consensus and shown to feature plant species variation and dependence on concentration and environmental stress effects (Harris 2015). It is feasible that a potential ABA-driven repression of auxin signaling during root formation step could help root elongation in Eucalyptus.
JA has been pointed out as a positive regulator of AR, mainly during the induction phase (Ravnikar et al. 1992; Rasmussen et al. 2015; Lischweski et al. 2015). This is in contrast to the negative role of JA observed in AR formation in intact hypocotyls of Arabidopsis (Gutierrez et al. 2012). Herein, a JA biosynthesis protein was downregulated during root induction phase in E. grandis. This may reflect a faster response and/or higher sensitivity of the easy-to-root species to JA in early stages of AR relative to the recalcitrant one. Several proteins could be involved in the JA biosynthesis and response pathway, some of them having overlapping functions.
Microtubule and cell wall modifications
Corroborating our findings, proteome and transcriptome studies during AR in cuttings of different plants including pine, poplar, carnation, E. grandis and Robinia report expression of proteins involved in cell wall remodeling and biosynthesis, cytoskeleton and microtubules (Álvarez et al. 2016; Díaz-Sala 2014; Druege et al. 2016). Exogenous auxin supply to cuttings acts predominantly at the wound site and lower region of basal stems, promoting cell dedifferentiation and new root meristem formation (Acosta et al. 2009). Accumulation of endogenous auxin at the base of cuttings can also lead to cell division and expansion (da Costa et al. 2013). Upregulation of a cell cycle protein in response to auxin during the formation phase of AR in E. globulus is in line with these changes. Beta-xylosidases have been identified as players during AR in black locust cuttings (Quan et al. 2017), and upregulation of a β-d-xylosidase during AR in IBA-treated samples was observed when compared to samples treated with cytokinin, a known AR inhibitor. We found three beta xylosidases during the formation phase, one of which (Eucgr.G00649.1) was more abundant upon auxin-treatment. Overall, beta xylosidases identified were distinctly regulated depending on the treatment contrast, showing that these proteins could have specific roles in regulating cell wall biogenesis during AR in Eucalyptus.
Microtubule-related proteins changed depending on phase of AR and treatment. A kinesin protein was consistently upregulated in auxin-treated samples during the formation phase, in agreement with findings of Abu-Abied et al. (2014), who reported a highly expressed gene coding for a kinesin-like protein in juvenile auxin-treated cuttings of E. grandis when compared to juvenile control, mature control and mature auxin-treated. Expression of kinesin-coding genes increased at the stage of cell division activation of carnation (Villacorta-Martín et al. 2015). Considering the role of microtubules in plant morphogenesis and organogenesis (Landrein and Hamant 2013) and the crosstalk between auxin, microtubule and cell wall remodeling (Abu-Abied et al. 2015), these results corroborate the participation of microtubule-related proteins in AR regulation.
Secondary metabolism
Plant laccases act in lignin biosynthesis, wound healing and maintenance of cell wall structure and integrity (Wang et al. 2015a). Transcript profile of E. grandis cambium showed that laccases are involved in lignification of juvenile wood-forming tissues (De Carvalho et al. 2008). Lignin metabolism is related to the regulation of cell division and differentiation (Santos Macedo et al. 2012). During AR in mung bean cuttings, upregulation of laccase-7, possibly involved in lignin catabolism and cell wall loosening, was observed (Li et al. 2017). In the present study, upregulation of laccase-9 in E. grandis when compared to E. globulus control samples was recorded, particularly during the induction phase, implicating lignin dynamics under conditions prone to adventitious roots development.
Anthocyanidin reductase, involved in the negative regulation of flavonoid biosynthesis, was downregulated in E. grandis. Consistently, flavonoid biosynthesis enzymes naringenin-2-oxoglutarate 3-dioxygenase and chalcone isomerase were upregulated. Flavonoids are a major class of phenolic compounds that can modulate auxin transport (Peer and Murphy 2007) by the interaction with PIN proteins (Buer et al. 2010) and act as auxin-protectors due to their antioxidant activity. In Eucalyptus gunnii, rooting was associated with high concentrations of some flavonoids (Curir et al. 1990), which was also reported during root formation phase in eucalypt (Schwambach et al. 2008).
Exogenous auxin treatment induced upregulation of a chalcone isomerase, also involved in flavonoid biosynthesis, during formation phase in E. globulus. This corroborates findings in cuttings of Camellia sinensis treated with IBA, which showed increased expression of flavonoid biosynthesis genes during AR (Wei et al. 2013, 2014). Chalcone isomerase was also upregulated in E. grandis during both AR phases. These results, together with the repression of the negative regulator of flavonoid biosynthesis anthocyanidin reductase in E. grandis, implicate flavonoids in AR capacity.
Conclusion
Proteins regulated along the different phases of eucalypt AR were identified and links between exogenous auxin, rooting recalcitrance and proteome were detected. Proteins related to positive regulators of AR, such as hydrogen peroxide and polyphenols, starch metabolism, lignin dynamics and increased flavonoid content were upregulated in E. grandis when compared to E. globulus. Also associated with the easy-to-root phenotype were proteins involved in higher auxin signaling, salicylic acid and, unexpectedly, ABA-related proteins. Auxin proved determinant in reversing recalcitrant rooting phenotypes in crosstalk with carbon sources, other phytohormones, cell cycle, and microtubule-related proteins. Cell cycle and microtubule-related proteins were apparently more important during the formation step. Lower expression of flavonoid metabolism-related proteins in E. globulus and increased UDP-glucose 4-epimerase involved in galactose metabolism and cell wall synthesis during the root induction phase could contribute to its hard-to-root phenotype. Taken together, data support an initial working model of key proteins regulating Eucalyptus AR (Fig. 3). To the best of our knowledge, this is the first study concerning protein changes during AR in Eucalyptus. These results represent a relevant step in understanding this important process at the protein level and may help advance its control in eucalypts.
Proposed working model representing suggested important regulators of AR in Eucalyptus at the protein level. A Main differences between hard-to-root (E. globulus) and easy-to-root (E. grandis) phenotypes. During induction phase occurs the upregulation of UDP-glucose 6-dehydrogenase 1 increasing cell wall synthesis and contributing to the difficult-to-root feature of E. globulus. This phenotype is also result of low content of AR positive regulators such as H2O2, through the upregulation of peroxidase 5, low lignins (down-regulation of laccase-9) and flavonoids (down-regulation of chalcone isomerase and upregulation of the negative regulator anthocyanidin reductase). Besides, a slower N mobilization capacity and an increase in defense responses could contribute to the recalcitrant phenotype. On the other hand, in E. grandis occurs the upregulation of ABP19 during the induction phase, possibly modulating the auxin signaling pathway. The increase of positive regulators of AR such as aquaporin, H2O2 (upregulation of SOD), polyphenols (high peroxidases 3 and 12), starch synthesis and accumulation (high glucose-6-phosphate isomerase and fructokinase), SA content (upregulation of SA binding protein), lignin (high laccase-9) and flavonoid (upregulation of chalcone isomerase and down-regulation of anthocyanidin reductase) also contributes to the rooting competent phenotype of E. grandis. The increase in ABA signaling could also suggest a positive role for ABA on AR. B Effect of exogenous auxin treatment on AR. The exogenous auxin induces auxin signaling (upregulation of ABP19), starch accumulation (high glucose-6-phosphate isomerase and fructokinase), high SA content (upregulation of SA binding protein) and high flavonoid (upregulation of chalcone isomerase and down-regulation of anthocyanidin reductase). Besides, important factors are associated with the formation phase, such as increase in sucrose metabolism (upregulation of sucrose synthase), high activity of cell cycle protein and MT-related kinesin, and decrease in auxin synthesis (down-regulation of amidase). Red rectangles are related to induction phase and blue rectangle is related to formation phase. Functions written in the middle of figures indicate no specific phase. SA, salicylic acid; MT, microtubule, ABA, abscisic acid
References
Abril N, Gion J-M, Kerner R et al (2011) Proteomics research on forest trees, the most recalcitrant and orphan plant species. Phytochemistry 72:1219–1242. https://doi.org/10.1016/j.phytochem.2011.01.005
Abu-Abied M, Szwerdszarf D, Mordehaev I et al (2014) Gene expression profiling in juvenile and mature cuttings of Eucalyptus grandis reveals the importance of microtubule remodeling during adventitious root formation. BMC Genom 15:1–10. https://doi.org/10.1186/1471-2164-15-826
Abu-Abied M, Rogovoy (Stelmakh) O, Mordehaev J et al (2015) Dissecting the contribution of microtubule behaviour in adventitious root induction. J Exp Bot 66:2813–2824. https://doi.org/10.1093/jxb/erv097
Acosta M, Oliveros-Valenzuela MR, Nicolás C, Sánchez-Bravo J (2009) Rooting of carnation cuttings: the auxin signal. Plant Signal Behav 4:234–236. https://doi.org/10.1016/j.plaphy.2008.07.009
Agulló-Antón MÁ, Ferrández-Ayela A, Fernández-García N et al (2014) Early steps of adventitious rooting: morphology, hormonal profiling and carbohydrate turnover in carnation stem cuttings. Physiol Plant 150:446–462. https://doi.org/10.1111/ppl.12114
Ahkami AH, Lischewski S, Haensch KT et al (2009) Molecular physiology of adventitious root formation in Petunia hybrida cuttings: involvement of wound response and primary metabolism. New Phytol 181:613–625. https://doi.org/10.1111/j.1469-8137.2008.02704.x
Ahkami A, Scholz U, Steuernagel B et al (2014) Comprehensive transcriptome analysis unravels the existence of crucial genes regulating primary metabolism during adventitious root formation in Petunia hybrida. PLoS ONE 9:e100997. https://doi.org/10.1371/journal.pone.0100997
Álvarez C, Valledor L, Sáez P et al (2016) Proteomic analysis through adventitious rooting of Pinus radiata stem cuttings with different rooting capabilities. Am J Plant Sci 07:1888–1904. https://doi.org/10.4236/ajps.2016.714174
Amzallag GN, Goloubinoff P (2003) An Hsp90 inhibitor, geldanamycin, as a brassinosteroid antagonist: evidence from salt-exposed roots of Vigna radiata. Plant Biol 5:143–150. https://doi.org/10.1055/s-2003-40725
Aumond MLML, de Araujo AT, de Oliveira Junkes CF et al (2017) Events associated with early age-related decline in adventitious rooting competence of Eucalyptus globulus Labill. Front Plant Sci 8:1–10. https://doi.org/10.3389/fpls.2017.01734
Beardmore T, Wetzel S, Kalous M (2000) Interactions of airborne methyl jasmonate with vegetative storage protein gene and protein accumulation and biomass partitioning in Populus plants. Can J For Res 30:1106–1113. https://doi.org/10.1139/x00-046
Bedon F, Majada J, Feito I et al (2011) Interaction between environmental factors affects the accumulation of root proteins in hydroponically grown Eucalyptus globulus (Labill.). Plant Physiol Biochem 49:69–76. https://doi.org/10.1016/j.plaphy.2010.09.020
Bedon F, Villar E, Vincent D et al (2012) Proteomic plasticity of two Eucalyptus genotypes under contrasted water regimes in the field. Plant, Cell Environ 35:790–805. https://doi.org/10.1111/j.1365-3040.2011.02452.x
Brinker M, van Zyl L, Liu WB et al (2004) Microarray analyses of gene expression during adventitious root development in Pinus contorta. Plant Physiol 135:1526–1539. https://doi.org/10.1104/pp.103.032235.et
Budzinski IGF, Moon DH, Lindén P et al (2016a) Seasonal variation of carbon metabolism in the cambial zone of Eucalyptus grandis. Front Plant Sci 7:1–17. https://doi.org/10.3389/fpls.2016.00932
Budzinski IGF, Moon DH, Morosini JS et al (2016b) Integrated analysis of gene expression from carbon metabolism, proteome and metabolome, reveals altered primary metabolism in Eucalyptus grandis bark, in response to seasonal variation. BMC Plant Biol 16:1–15. https://doi.org/10.1186/s12870-016-0839-8
Buer CS, Imin N, Djordjevic MA (2010) Flavonoids: new roles for old molecules. J Integr Plant Biol 52:98–111. https://doi.org/10.1111/j.1744-7909.2010.00905.x
Calderan-Rodrigues MJ, Jamet E, Bonassi MBCR et al (2014) Cell wall proteomics of sugarcane cell suspension cultures. Proteomics 14:738–749. https://doi.org/10.1002/pmic.201300132
Celedon PAF, De Andrade A, Meireles KGX et al (2007) Proteomic analysis of the cambial region in juvenile Eucalyptus grandis at three ages. Proteomics 7:2258–2274. https://doi.org/10.1002/pmic.200600989
Chen Q, Guo W, Feng L et al (2015) Data for transcriptome and proteome analysis of Eucalyptus infected with Calonectria pseudoreteaudii. Data Br 3:24–28. https://doi.org/10.1016/j.dib.2014.12.008
Chueca A, Sahrawy M, Pagano EA, López Gorgé J (2002) Chloroplast fructose-1,6-bisphosphatase: structure and function. Photosynth Res 74:235–249. https://doi.org/10.1023/A:1021243110495
Conesa A, Gotz S, Garcia-Gomez JM et al (2005) Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21:3674–3676. https://doi.org/10.1093/bioinformatics/bti610
Corrêa LdaR, Paim DC, Schwambach J, Fett-Neto AG (2005) Carbohydrates as regulatory factors on the rooting of Eucalyptus saligna Smith and Eucalyptus globulus Labill. Plant Growth Regul 45:63–73. https://doi.org/10.1007/s10725-004-6125-z
Curir P, VanSumere CF, Termini A et al (1990) Flavonoid accumulation is correlated with adventitious roots formation in Eucalyptus gunnii Hook micropropagated through axillary bud stimulation. Plant Physiol 92:1148–1153. https://doi.org/10.1104/pp.92.4.1148
da Costa CTCT, de Almeida MR, Ruedell CMCM et al (2013) When stress and development go hand in hand: main hormonal controls of adventitious rooting in cuttings. Front Plant Sci 4:1–19. https://doi.org/10.3389/fpls.2013.00133
Damerval C, De Vienne D, Zivy M, Thiellement H (1986) Technical improvements in two-dimensional electrophoresis increase the level of genetic variation detected in wheat-seedling proteins. Electrophoresis 7:52–54. https://doi.org/10.1002/elps.1150070108
de Almeida MR, de Bastiani D, Gaeta MLML et al (2015) Comparative transcriptional analysis provides new insights into the molecular basis of adventitious rooting recalcitrance in Eucalyptus. Plant Sci 239:155–165. https://doi.org/10.1016/j.plantsci.2015.07.022
De Almeida Leonardi G, Carlos NA, Mazzafera P, Balbuena TS (2015) Eucalyptus urograndis stem proteome is responsive to short-term cold stress. Genet Mol Biol 38:191–198. https://doi.org/10.1590/S1415-475738220140235
De Carvalho MCDCG, Caldas DGG, Carneiro RT et al (2008) SAGE transcript profiling of the juvenile cambial region of Eucalyptus grandis. Tree Physiol 28:905–919. https://doi.org/10.1093/treephys/28.6.905
de Klerk G-J, van der Krieken W, de Jong JC (1999) The formation of adventitious roots: new concepts, new possibilities. Vitr Cell Dev Biol Plant 35:189–199. https://doi.org/10.1007/s11627-999-0076-z
De Klerk G-J, Guan H, Huisman P, Marinova S (2011) Effects of phenolic compounds on adventitious root formation and oxidative decarboxylation of applied indoleacetic acid in Malus ‘Jork 9’. Plant Growth Regul 63:175–185. https://doi.org/10.1007/s10725-010-9555-9
De Santana Costa MG, Mazzafera P, Balbuena TS (2017) Insights into temperature modulation of the Eucalyptus globulus and Eucalyptus grandis antioxidant and lignification subproteomes. Phytochemistry 137:15–23. https://doi.org/10.1016/j.phytochem.2017.01.017
Díaz-Sala C (2014) Direct reprogramming of adult somatic cells toward adventitious root formation in forest tree species: the effect of the juvenile-adult transition. Front Plant Sci 5:1–8. https://doi.org/10.3389/fpls.2014.00310
Druart N, Johansson A, Baba K et al (2007) Environmental and hormonal regulation of the activity-dormancy cycle in the cambial meristem involves stage-specific modulation of transcriptional and metabolic networks. Plant J 50:557–573. https://doi.org/10.1111/j.1365-313X.2007.03077.x
Druege U, Franken P, Hajirezaei MR (2016) Plant hormone homeostasis, signaling, and function during adventitious root formation in cuttings. Front Plant Sci. https://doi.org/10.3389/fpls.2016.00381
Fett-Neto AG, Teixeira SL, Da Silva EAM, Sant’ Anna R (1992) Biochemical and morphological changes during in vitro rhizogenesis in cuttings of Sequoia sempervirens (D. Don) Endl. J Plant Physiol 140:720–728. https://doi.org/10.1016/S0176-1617(11)81029-1
Fett-Neto AG, Fett JP, Goulart LWV et al (2001) Distinct effects of auxin and light on adventitious root development in Eucalyptus saligna and Eucalyptus globulus. Tree Physiol 21:457–464. https://doi.org/10.1093/treephys/21.7.457
Goodstein DM, Shu S, Howson R et al (2012) Phytozome: a comparative platform for green plant genomics. Nucleic Acids Res 40:1178–1186. https://doi.org/10.1093/nar/gkr944
Grauvogel C, Brinkmann H, Petersen J (2007) Evolution of the glucose-6-phosphate isomerase: the plasticity of primary metabolism in photosynthetic eukaryotes. Mol Biol Evol 24:1611–1621. https://doi.org/10.1093/molbev/msm075
Guarino C, Conte B, Spada V et al (2014) Proteomic analysis of eucalyptus leaves unveils putative mechanisms involved in the plant response to a real condition of soil contamination by multiple heavy metals in the presence or absence of mycorrhizal/rhizobacterial additives. Environ Sci Technol 48:11487–11496. https://doi.org/10.1021/es502070m
Gutierrez L, Mongelard G, Floková K et al (2012) Auxin controls Arabidopsis adventitious root initiation by regulating jasmonic acid homeostasis. Plant Cell 24:2515–2527. https://doi.org/10.1105/tpc.112.099119
Han H, Sun X, Xie Y et al (2014) Transcriptome and proteome profiling of adventitious root development in hybrid larch (Larix kaempferi × Larix olgensis). BMC Plant Biol 14:1–13. https://doi.org/10.1186/s12870-014-0305-4
Harris J (2015) Abscisic acid: hidden architect of root system structure. Plants 4:548–572. https://doi.org/10.3390/plants4030548
Heringer AS, Barroso T, Macedo AF et al (2015) Label-free quantitative proteomics of embryogenic and non-embryogenic callus during sugarcane somatic embryogenesis. PLoS ONE 10:e0127803. https://doi.org/10.1371/journal.pone.0127803
Kersting AR, Mizrachi E, Bornberg-Bauer E, Myburg AA (2015) Protein domain evolution is associated with reproductive diversification and adaptive radiation in the genus Eucalyptus. New Phytol 206:1328–1336. https://doi.org/10.1111/nph.13211
Kohler A, Delaruelle C, Martin D et al (2003) The poplar root transcriptome: analysis of 7000 expressed sequence tags. FEBS Lett 542:37–41. https://doi.org/10.1016/S0014-5793(03)00334-X
Landrein B, Hamant O (2013) How mechanical stress controls microtubule behavior and morphogenesis in plants: history, experiments and revisited theories. Plant J 75:324–338. https://doi.org/10.1111/tpj.12188
Lei C, Fan S, Li K et al (2018) iTRAQ-based proteomic analysis reveals potential regulation networks of IBA-induced adventitious root formation in apple. Int J Mol Sci 19:667. https://doi.org/10.3390/ijms19030667
Leslie AD, Mencuccini M, Perks MP, Wilson ER (2019) A review of the suitability of eucalypts for short rotation forestry for energy in the UK. New For. https://doi.org/10.1007/s11056-019-09717-w
Li S-W, Shi R-F, Leng Y (2015) De novo characterization of the mung bean transcriptome and transcriptomic analysis of adventitious rooting in seedlings using RNA-Seq. PLoS ONE 10:e0132969. https://doi.org/10.1371/journal.pone.0132969
Li SW, Leng Y, Shi RF (2017) Transcriptomic profiling provides molecular insights into hydrogen peroxide-induced adventitious rooting in mung bean seedlings. BMC Genom 18:1–23. https://doi.org/10.1186/s12864-017-3576-y
Lischweski S, Muchow A, Guthörl D, Hause B (2015) Jasmonates act positively in adventitious root formation in petunia cuttings. BMC Plant Biol 15:229. https://doi.org/10.1186/s12870-015-0615-1
Liu R, Chen S, Jiang J et al (2013) Proteomic changes in the base of Chrysanthemum cuttings during adventitious root formation. BMC Genom 14:1–14. https://doi.org/10.1186/1471-2164-14-919
Lu N, Xu Z, Meng B et al (2014) Proteomic analysis of etiolated juvenile tetraploid Robinia pseudoacacia branches during different cutting periods. Int J Mol Sci 15:6674–6688. https://doi.org/10.3390/ijms15046674
Lu N, Dai L, Luo Z et al (2017) Characterization of the transcriptome and gene expression of tetraploid black locust cuttings in response to etiolation. Genes (Basel) 8:1–18. https://doi.org/10.3390/genes8120345
Mano J, Belles-Boix E, Babiychuk E, Inzé D et al (2005) Protection against photooxidative injury of tobacco leaves by 2-alkenal reductase. Detoxication of lipid peroxide-derived reactive carbonyls. Plant Physiol 139:1773–1783. https://doi.org/10.1104/pp.105.070391
Melzer E, O’Leary MH (1987) Anapleurotic CO2 fixation by phosphoenolpyruvate carboxylase in C3 plants. Plant Physiol 84:58–60. https://doi.org/10.1104/pp.84.1.58
Min T, Kasahara H, Bedgar DL et al (2003) Crystal structures of pinoresinol-lariciresinol and phenylcoumaran benzylic ether reductases and their relationship to isoflavone reductases. J Biol Chem 278:50714–50723. https://doi.org/10.1074/jbc.M308493200
Mishra BS, Singh M, Aggrawal P, Laxmi A (2009) Glucose and auxin signaling interaction in controlling Arabidopsis thaliana seedlings root growth and development. PLoS ONE 4:e4502. https://doi.org/10.1371/journal.pone.0004502
Murashige T, Skoog F (1962) A revised medium for rapid growth and bio assays with tobacco tissue cultures. Physiol Plant 15:473–497. https://doi.org/10.1111/j.1399-3054.1962.tb08052.x
Negishi N, Nakahama K, Urata N et al (2014) Hormone level analysis on adventitious root formation in Eucalyptus globulus. New For 45:577–587. https://doi.org/10.1007/s11056-014-9420-1
Pacurar DI, Perrone I, Bellini C (2014) Auxin is a central player in the hormone cross-talks that control adventitious rooting. Physiol Plant 151:83–96. https://doi.org/10.1111/ppl.12171
Park C-J, Seo Y-S (2015) Heat shock proteins: a review of the molecular chaperones for plant immunity. Plant Pathol J 31:323–333. https://doi.org/10.5423/PPJ.RW.08.2015.0150
Peer WA, Murphy AS (2007) Flavonoids and auxin transport: modulators or regulators? Trends Plant Sci 12:556–563. https://doi.org/10.1016/j.tplants.2007.10.003
Péret B, Li G, Zhao J et al (2012) Auxin regulates aquaporin function to facilitate lateral root emergence. Nat Cell Biol 14:991–998. https://doi.org/10.1038/ncb2573
Quan J, Meng S, Guo E et al (2017) De novo sequencing and comparative transcriptome analysis of adventitious root development induced by exogenous indole-3-butyric acid in cuttings of tetraploid black locust. BMC Genom 18:1–14. https://doi.org/10.1186/s12864-017-3554-4
Quecine MC, Leite TF, Bini AP et al (2016) Label-free quantitative proteomic analysis of Puccinia psidii uredospores reveals differences of fungal populations infecting Eucalyptus and guava. PLoS ONE 11:1–19. https://doi.org/10.1371/journal.pone.0145343
Ramirez-Carvajal GA, Morse AM, Dervinis C, Davis JM (2009) The cytokinin type-B response regulator PtRR13 Is a negative regulator of adventitious root development in Populus. Plant Physiol 150:759–771. https://doi.org/10.1104/pp.109.137505
Rasmussen A, Hosseini SA, Hajirezaei MR et al (2015) Adventitious rooting declines with the vegetative to reproductive switch and involves a changed auxin homeostasis. J Exp Bot 66:1437–1452. https://doi.org/10.1093/jxb/eru499
Ravnikar M, Vilhar B, Gogala N (1992) Stimulatory effects of jasmonic acid on potato stem node and protoplast culture. J Plant Growth Regul 11:29–33. https://doi.org/10.1007/BF00193840
Rencoret J, Gutiérrez A, del Río JC (2007) Lipid and lignin composition of woods from different eucalypt species. Holzforschung 61:165–174. https://doi.org/10.1515/HF.2007.030
Rojas-González JA, Soto-Súarez M, García-Díaz Á et al (2015) Disruption of both chloroplastic and cytosolic FBPase genes results in a dwarf phenotype and important starch and metabolite changes in Arabidopsis thaliana. J Exp Bot 66:2673–2689. https://doi.org/10.1093/jxb/erv062
Rolland F, Baena-Gonzalez E, Sheen J (2006) Sugar sensing and signaling in plants: conserved and novel mechanisms. Annu Rev Plant Biol 57:675–709. https://doi.org/10.1146/annurev.arplant.57.032905.105441
Ruedell CMCM, de Almeida MR, Schwambach J et al (2013) Pre and post-severance effects of light quality on carbohydrate dynamics and microcutting adventitious rooting of two Eucalyptus species of contrasting recalcitrance. Plant Growth Regul 69:235–245. https://doi.org/10.1007/s10725-012-9766-3
Ruedell CMCM, de Almeida MR, Fett-Neto AGAG (2015) Concerted transcription of auxin and carbohydrate homeostasis-related genes underlies improved adventitious rooting of microcuttings derived from far-red treated Eucalyptus globulus Labill mother plants. Plant Physiol Biochem 97:11–19. https://doi.org/10.1016/j.plaphy.2015.09.005
Samac DA, Litterer L, Temple G et al (2004) Expression of UDP–glucose dehydrogenase reduces cell-wall polysaccharide concentration and increases xylose content in alfalfa stems. Appl Biochem Biotechnol 116:1167–1182. https://doi.org/10.1385/ABAB:116:1-3:1167
Santos BM dos, Balbuena TS (2017) Carbon assimilation in Eucalyptus urophylla grown under high atmospheric CO2 concentrations: a proteomics perspective. J Proteom 150:252–257. https://doi.org/10.1016/j.jprot.2016.09.010
Santos Macedo E, Sircar D, Cardoso HG et al (2012) Involvement of alternative oxidase (AOX) in adventitious rooting of Olea europaea L. microshoots is linked to adaptive phenylpropanoid and lignin metabolism. Plant Cell Rep 31:1581–1590. https://doi.org/10.1007/s00299-012-1272-6
Schwambach J, Fadanelli C, Fett-Neto AG (2005) Mineral nutrition and adventitious rooting in microcuttings of Eucalyptus globulus. Tree Physiol 25:487–494. https://doi.org/10.1093/treephys/25.4.487
Schwambach J, Ruedell CMCM, De Almeida MR et al (2008) Adventitious rooting of Eucalyptus globulus × maidenii mini-cuttings derived from mini-stumps grown in sand bed and intermittent flooding trays: a comparative study. New For 36:261–271. https://doi.org/10.1007/s11056-008-9099-2
Seitz B, Klos C, Wurm M, Tenhaken R (2000) Matrix polysaccharide precursors in Arabidopsis cell walls are synthesized by alternate pathways with organ-specific expression patterns. Plant J 21:537–546. https://doi.org/10.1046/j.1365-313x.2000.00696.x
Serrano L, Rochange F, Semblat JP et al (1996) Genetic transformation of Eucalyptus globulus through biolistics: complementary development of procedures for organogenesis from zygotic embryos and stable transformation of corresponding proliferating tissue. J Exp Bot 47:285–290. https://doi.org/10.1093/jxb/47.2.285
Silva JC, Denny R, Dorschel CA et al (2005) Quantitative proteomic analysis by accurate mass retention time pairs. Anal Chem 77:2187–2200. https://doi.org/10.1021/ac048455k
Sorin C, Negroni L, Balliau T et al (2006) Proteomic analysis of different mutant genotypes of arabidopsis led to the identification of 11 proteins correlating with adventitious root development. Plant Physiol 140:349–364. https://doi.org/10.1104/pp.105.067868
Steffens B, Rasmussen A (2016) The physiology of adventitious roots. Plant Physiol 170:603–617. https://doi.org/10.1104/pp.15.01360
Thanananta N, Vuttipongchaikij S, Apisitwanich S (2018) Agrobacterium-mediated transformation of a Eucalyptus camaldulensis × E. tereticornis hybrid using peeled nodal-stem segments with yeast HAL2 for improving salt tolerance. New For 49:311–327. https://doi.org/10.1007/s11056-017-9621-5
Tromas A, Braun N, Muller P et al (2009) The AUXIN BINDING PROTEIN 1 is required for differential Auxin responses mediating root growth. PLoS ONE 4:1–11. https://doi.org/10.1371/journal.pone.0006648
Trupiano D, Yordanov Y, Regan S et al (2013) Identification, characterization of an AP2/ERF transcription factor that promotes adventitious, lateral root formation in Populus. Planta 238:271–282. https://doi.org/10.1007/s00425-013-1890-4
Valdés AE, Irar S, Majada JP et al (2013) Drought tolerance acquisition in Eucalyptus globulus (Labill.): a research on plant morphology, physiology and proteomics. J Proteom 79:263–276. https://doi.org/10.1016/j.jprot.2012.12.019
Vilasboa J, Da Costa CT, Fett-Neto AG (2018) Rooting of eucalypt cuttings as a problem-solving oriented model in plant biology. Prog Biophys Mol Biol. https://doi.org/10.1016/j.pbiomolbio.2018.12.007
Villacorta-Martín C, Sánchez-García AB, Villanova J et al (2015) Gene expression profiling during adventitious root formation in carnation stem cuttings. BMC Genom 16:789. https://doi.org/10.1186/s12864-015-2003-5
Vizcaíno JA, Deutsch EW, Wang R et al (2014) ProteomeXchange provides globally coordinated proteomics data submission and dissemination. Nat Biotechnol 32:223–226. https://doi.org/10.1038/nbt.2839
Vizcaíno JA, Csordas A, Del-Toro N et al (2016) 2016 update of the PRIDE database and its related tools. Nucleic Acids Res 44:D447–D456. https://doi.org/10.1093/nar/gkv1145
Wang J, Feng J, Jia W et al (2015a) Lignin engineering through laccase modification: a promising field for energy plant improvement. Biotechnol Biofuels 8:145. https://doi.org/10.1186/s13068-015-0331-y
Wang S, Ito T, Uehara M et al (2015b) UDP-d-galactose synthesis by UDP-glucose 4-epimerase 4 is required for organization of the trans-Golgi network/early endosome in Arabidopsis thaliana root epidermal cells. J Plant Res 128:863–873. https://doi.org/10.1007/s10265-015-0737-4
Wang R, Zhang Y, Kieffer M et al (2016) HSP90 regulates temperature-dependent seedling growth in Arabidopsis by stabilizing the auxin co-receptor F-box protein TIR1. Nat Commun 7:1–10. https://doi.org/10.1038/ncomms10269
Wang Z, Hua J, Yin Y et al (2019) An integrated transcriptome and proteome analysis reveals putative regulators of adventitious root formation in Taxodium ‘Zhongshanshan’. Int J Mol Sci 20:1225. https://doi.org/10.3390/ijms20051225
Wei K, Wang L, Cheng H et al (2013) Identification of genes involved in indole-3-butyric acid-induced adventitious root formation in nodal cuttings of Camellia sinensis (L.) by suppression subtractive hybridization. Gene 514:91–98. https://doi.org/10.1016/j.gene.2012.11.008
Wei K, Wang L-Y, Wu L-Y et al (2014) Transcriptome analysis of indole-3-butyric acid-induced adventitious root formation in nodal cuttings of Camellia sinensis (L.). PLoS ONE 9:1–9. https://doi.org/10.1371/journal.pone.0107201
Xu Z-S, Li Z-Y, Chen Y et al (2012) Heat shock protein 90 in plants: molecular mechanisms and roles in stress responses. Int J Mol Sci 13:15706–15723. https://doi.org/10.3390/ijms131215706
Yamauchi Y, Hasegawa A, Mizutani M, Sugimoto Y (2012) Chloroplastic NADPH-dependent alkenal/one oxidoreductase contributes to the detoxification of reactive carbonyls produced under oxidative stress. FEBS Lett 586:1208–1213. https://doi.org/10.1016/j.febslet.2012.03.013
Zelko IN, Mariani TJ, Folz RJ (2002) Superoxide dismutase multigene family: a comparison of the CuZn-SOD (SOD1), Mn-SOD (SOD2), and EC-SOD (SOD3) gene structures, evolution, and expression. Free Radic Biol Med 33:337–349. https://doi.org/10.1016/S0891-5849(02)00905-X
Zhao Y (2010) Auxin biosynthesis and its role in plant development. Annu Rev Plant Biol 61:49–64. https://doi.org/10.1146/annurev-arplant-042809-112308
Zheng Z, Guo Y, Novák O et al (2013) Coordination of auxin and ethylene biosynthesis by the aminotransferase VAS1. Nat Chem Biol 9:244–246. https://doi.org/10.1038/nchembio.1178
Acknowledgements
Seeds used for production of E. globulus and E. grandis microcuttings were a kind gift from Celulose Riograndense S/A (Guaíba, RS, Brazil). This work was funded by Brazilian agencies National Council for Scientific and Technological Development (CNPq-grants 402618/2016-5 and 303560/2017-7) and National Commission for Improvement of Higher Education Personnel (CAPES)-Finance Code 001.
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiments: AGFN. Performed the experiments: MRA, JS, VS. Analyzed the data: MRA, JS. Contributed reagents/materials/analysis tools: JPF, AGFN, VS, JS, ASH. Drafted the paper: MRA. Supervised the study and finalized the manuscript: AGFN. All authors contributed to the preparation, revised and agreed with the final version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Online Resource 1.
Unchanged proteins during the induction phase. (XLSX 143 kb)
Online Resource 2.
Unchanged proteins during the formation phase. (XLSX 136 kb)
Online Resource 3.
Regulated proteins during the induction phase. (XLSX 99 kb)
Online Resource 4.
Regulated proteins during the formation phase. (XLSX 110 kb)
Online Resource 5.
Identification and classification of regulated proteins during the induction phase. Respective Fold change values and ANOVA results can be accessed in Online Resource 3. (XLSX 27 kb)
Online Resource 6.
Identification and classification of regulated proteins during the formation phase. Respective Fold change values and ANOVA results can be accessed in Online Resource 4. (XLSX 29 kb)
Rights and permissions
About this article
Cite this article
de Almeida, M.R., Schwambach, J., Silveira, V. et al. Proteomic profiles during adventitious rooting of Eucalyptus species relevant to the cellulose industry. New Forests 51, 213–241 (2020). https://doi.org/10.1007/s11056-019-09728-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11056-019-09728-7