Abstract
Within a decade after its first description, the multidrug-resistant yeast Candida auris has emerged globally as a nosocomial pathogen causing difficult to control outbreaks. This, together with the alarmingly high mortality rate of up to 66% associated with C. auris candidemia, calls for a better understanding of its virulence traits and routes of transmission. Unlike other clinically relevant Candida species, C. auris seems to have the unique ability to be easily transmitted between patients. Although initially thought to express fewer virulence traits than Candida albicans, recent genomic insights suggest C. auris to possess these traits to a much more similar extent. This review highlights the virulence traits C. auris expresses to attack the host, defend itself against antimicrobial agents and to persist within the healthcare environment.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Annually, an estimated 1.5 million people die from invasive fungal infections [1]. The advance of life expectancy, the rise of immunosuppressive treatments, higher survival of patients living with cancer or chronic disease and the use of catheters are all factors that attributed to the emergence of opportunistic fungal pathogens over the last decades [2, 3].
Candida species are considered the most frequent fungi encountered in hospital settings accounting for more than 400,000 cases of bloodstream infections each year, making them the third to fourth most common cause of invasive fungal infections worldwide [1, 4,5,6]. Candida albicans is recognized as the main causative pathogen of candidiasis [1]. However, new species are on the rise, with the globally emerging multidrug-resistant Candida auris as one of the most concerning examples. Treatment options of C. auris are limited due to antifungal resistance, misidentification and its ability to persistently colonize hospital environments.
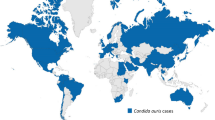
Since its first description in 2009, C. auris has been reported in over 25 countries on five continents (Fig. 1), causing fungemia outbreaks with crude mortality rates ranging from 32 to 66% [7,8,9]. Because the Candida genus is composed of a highly heterogeneous group of species, C. auris differs markedly from common well-studied pathogenic Candida species such as C. albicans and C. glabrata [10, 11]. For example, where other Candida infections are thought to result from autoinfection from host flora, C. auris seems to have the unique ability to persistently colonize the host skin, making it easily transmissible between patients [10, 12]. This transmissibility together with its multidrug resistance and high mortality rates makes C. auris a serious threat to public health.
Global epidemiology of Candida auris. Updated map of the C. auris epidemiology based on data presented by the Centers of Disease Control (Atlanta, GA, USA; https://www.cdc.gov/fungal/candida-auris/tracking-c-auris.html; [18]). Single (imported) cases were reported from Austria, Belgium, Chile, Iran, Norway, Switzerland, Taiwan and the United Arab Emirates. Multiple cases of C. auris infections were reported from Australia, Bangladesh, Canada, China, France, Germany, Israel, Japan, Kenya, Kuwait, Malaysia, the Netherlands, Oman, Panama, Russia, Saudi Arabia, Singapore and South Korea. From Colombia, India, Pakistan, South Africa, Spain, the UK, the USA and Venezuela extensive (inter)hospital transmissions were reported
A large number of papers have addressed the epidemiology of C. auris (reviewed by [8, 13, 14]). However, its mechanisms of pathogenicity and virulence have only recently been discussed [15]. Here, we will further elaborate on the current knowledge of virulence traits of C. auris, highlighting the differences and similarities with the most common pathogenic Candida species helping to understand how C. auris was able to emerge rapidly as a new global nosocomial pathogen.
What the Genome of C. auris is Telling Us
In 2009, Candida auris was described as a novel species belonging to the C. haemulonii species complex (Metschnikowiaceae clade), after isolation from the ear canal of a 70-year-old Japanese woman [16]. Unfortunately, C. auris is often misidentified as C. haemulonii, Candida famata and Rhodotorula glutinis by commercial biochemical identification systems such as Microscan (Beckman Coulter, Pasadena, CA) and API-20C AUX (BioMérieux, Marcy L’Etoile, France). Therefore, systems using matrix-assisted laser desorption ionization–time of flight mass spectrometry (MALDI-TOF MS) or identification by sequencing the internal transcribed spacer region (ITS) are needed for rapid and reliable identification of this yeast [17, 18]. Previously, lack of C. auris entries in the libraries made it impossible for both the Bruker biotyper (Bruker Daltonics, Bremen, Germany) and VITEK-MS (BioMérieux) MALDI-TOF MS systems to identify C. auris. However, when queried on an updated database, containing C. auris, accurate identification is possible [19].
A retrospective review of Candida isolates from three reported cases of nosocomial fungemia in South Korea showed that all cases were caused by C. auris, with the earliest report dating back to 1996 [20]. However, it remains unclear why C. auris has only recently emerged globally as a nosocomial pathogen. Whole genome sequencing (WGS) of clinical isolates led to the description of four unique clades of C. auris divided by geographic region (East Asian, South Asian, South African and South American). WGS analysis has shown that thousands of single-nucleotide polymorphisms (SNPs) separated clades, but within clades isolates are clonal. This analysis indicates the independent, nearly simultaneous emergence of clonal populations of C. auris on three continents [7].
The genomes of many pathogenic Candida species, such as C. albicans and C. glabrata, have extensively been studied, while the high-quality genome of C. auris was only very recently investigated [21,22,23,24]. Despite the recent efforts made in sequencing the C. auris genome, detailed information regarding the genome architecture, virulence and multidrug resistance of C. auris is lacking. Muñoz and colleagues (2018) used high-quality genome data to investigate the interspecies relationships and observed that within the C. auris clade the average pairwise nucleotide identity was 98.7% and that the nucleotide genome identity between C. auris and each of the siblings C. haemulonii, C. duobushaemulonii and C. pseudohaemulonii was 88% [24]. In addition, 40% of the predicted proteins were found to be orthologous to those of C. lusitaniae [23]. Nonetheless, the majority of these proteins are uncharacterized and hypothetical proteins. Therefore, it remains unclear whether these proteins are involved in the highly virulent and pathogenic behavior of C. auris.
Comparison of the C. auris genome to the more well-annotated and well-studied genome, yet distantly related, of C. albicans suggested the presence of a significant number of orthologues that may contribute to the virulence of C. auris. This set of orthologues included transporters belonging to the major facilitator superfamily and ABC (ATP binding cassette) superfamily, secreted proteinases, lipases, phospholipases and aspartyl proteases [21, 23]. However, the roles of specific genes need to be further investigated. The C. auris genome was also found to encode for kinases like Hog1, Protein Kinase A (PKA) and two-component histidine kinase which have been reported to be involved in the regulation of stress signaling pathways to enhance tolerance of pathogenic fungi to chemical fungicides [25]. Hog1 has recently been shown to play a role in regulating stress resistance, cell morphology, aggregation and virulence in C. auris [26]. The function of other kinases in C. auris still needs to be experimentally verified.
The Many ‘Faces’ of C. auris
Phenotypic switching and morphogenesis are key features for virulence of C. albicans and therefore extensively studied in this species (as reviewed by others [27,28,29]). C. albicans deploys two typical switching systems, namely the morphological yeast-filament transition and phenotypic white-opaque switching. The ability to produce hyphae is thought to promote virulence by giving the fungus the ability to invade epithelial cell layers by exerting mechanical force, breaching and damaging endothelial cells, and causing lysis of macrophages and neutrophils following phagocytoses [27].
The first micromorphological studies of C. auris suggested that this pathogen was not capable of forming germ tubes, pseudohyphae or hyphae [20, 30, 31]. However, pseudohyphae-like forms seem to be induced under high-salt stress and when forming biofilms. These pseudohyphae are characterized by rudimentary growth, elongated shape and incomplete cell division [31, 32]. The presence of a pseudohyphal phenotype in C. auris suggests the potential to undergo filamentation under certain unidentified conditions. Although well-known environmental factors of filamentation in C. albicans could not induce filamentous growth in C. auris in vitro [31], Yue et al. [33] showed that passage of C. auris through a mammalian host could trigger a heritable switch between the typical yeast and a filamentous phenotype, in which hyphae are formed. C. auris cells recovered from mice kidney and liver tissues displayed very elongated filaments after culturing on YPD medium for 24 h. More recently, a regulative role of the essential molecular chaperone Hsp90 in the morphogenesis of C. auris was found. Depletion of Hsp90 in C. auris resulted in filamentous growth similar to C. albicans, showing Hsp90 to be a key regulator of morphogenesis [34].
Comparative genomics showed genes encoding homologs of C. albicans hyphal regulators to be upregulated in filamentous cells of C. auris [33]. On the other hand, a number of conserved genes that govern filamentous growth in C. albicans are differentially expressed in C. auris. For example, EFG1 known to be required for filamentous growth in C. albicans is down regulated in filamentous C. auris cells [33, 35]. Moreover, genome analysis has shown that C. auris lacks the gene encoding candidalysin (ECE1) and hyphal cell wall protein (HWP1), both of which are strongly associated with hyphal formation [24, 34]. This suggests a partially conserved mechanism for filamentation between C. auris and C. albicans, but also the presence of other transcriptional regulators and signaling molecules, capable of inducing filamentation in C. auris, activated by yet to be explored environmental factors.
Phenotypic switching has been described in many Candida species such as C. albicans [36], C. glabrata [37], C. tropicalis [38, 39] and C. dubliniensis [40]. The reversible and heritable switching between white and opaque cellular phenotypes is a well-known virulence attribute of C. albicans, but also plays a role in sexual reproduction [36]. This phenotypic plasticity allows the fungus to colonize distinct host niches. For example, opaque cells show reduced virulence compared to white cells, but are excellent colonizers of the skin. Moreover, opaque cells are phagocytosed less efficiently [41]. Recently, Bentz and co-workers [42] reported phenotypic switching in C. auris when cultured onto CHROMagar Candida, on which C. albicans and C. tropicalis can be relatively reliably identified via a colony color change (C. albicans—green, C. tropicalis—navy blue). Other clinically relevant Candida species, including C. auris, will have a pale appearance. Further culturing of C. auris on CHROMagar Candida led to the description of three predominant colony types: white, pale and sectored (dark purple). No texture changes were observed, as all colonies displayed a smooth and glossy phenotype [42].
The identification of a transition between the three C. auris phenotypes shows similarities to the white-opaque transition in C. albicans, which was proposed to be tri-stable, due to the presence of an intermediate gray phenotype [36]. In addition, phenotypic switching in C. albicans is regulated by the master regulator Wor1 [41]. C. auris possesses three potential genes homologous with WOR1 that could control phenotypic switching in C. auris, similar to that of C. albicans [42]. The more closely related species C. lusitaniae also seems to possess a tri-stable switching system, which is highly associated with antifungal resistance and filamentation [43, 44], indicating that phenotypic switching could contribute to antifungal resistance in C. auris as well. Taken together, phenotypic switching in C. auris could have important implications for its pathogenicity, virulence, sexual reproduction and ability to colonize distinct host niches, which warrants further investigation.
Attacking the Host: Lytic Enzyme Production and Secretion
The production of extracellular hydrolytic enzymes has been recognized as an important virulence trait contributing to the pathogenicity of Candida species. Proteinases are by far the most commonly virulence-associated enzymes. In addition, also hemolysins, lipases and phospholipases seem to play a crucial role.
Secreted aspartyl proteinases (SAPs) are one of the most significant extracellular enzymes produced in C. albicans. Classically, these enzymes were considered to play a role in the degradation of host tissue to provide nutrients for pathogen propagation. However, in recent decades, proteinases have also been associated with cell wall maintenance, the formation of polymicrobial biofilms, adhesin to external protective barriers of the host, deregulation of the complement system, inactivation of host antimicrobial peptides, evasion of the immune responses and the induction of inflammatory mediator release from host cells (as reviewed by others [45, 46]). In C. albicans, SAPs are encoded by a family of ten genes. Especially SAP4, SAP5 and SAP6 seem to play an important role in virulence, since inhibition of the production of these proteins greatly attenuates C. albicans pathogenicity [47].
Many other pathogenic Candida species possess SAP genes, including C. dubliniensis, C. parapsilosis and C. tropicalis [45, 46]. Genomic analysis showed C. tropicalis to have at least four SAP genes. C. parapsilosis possesses as many as 14 potential SAP genes, and the genome of C. dubliniensis encodes eight members of the aspartyl proteinase family. In the C. auris genome (strain Ci 6684), hydrolases are the largest group of enzymes (42%), in which four orthologues of SAPs have been found [21]. In vitro studies confirmed that C. auris is able to secrete proteinases, although in a strain-dependent manner [31, 48, 49]. Comparing SAP activity of C. auris and C. albicans demonstrated high activities at 25 °C, 37 °C and 40 °C. Interestingly, C. auris SAP activity at 42 °C was much higher than that of C. albicans, indicating C. auris may be able to maintain its pathogenicity at higher temperatures [31].
Other molecules such as the pore-forming toxins hemolysins are needed to achieve host colonization. Secretion of hemolysins is considered to promote survival within the mammalian host by allowing assimilating iron from the hemoglobin-heme group [50]. Many common pathogenic Candida species display hemolysin activity, including C. albicans, C. dubliniensis, C. glabrata and C. tropicalis [50,51,52,53]. Hemolysin production seems to be higher in strains isolated from hospital infections compared to those from environmental sources, indicating this trait to be an important virulence factor [5]. In C. auris, hemolysin production was also observed [49]. Nonetheless, this was a single isolate only and variation in hemolysin activity between the different C. auris clades still needs to be evaluated.
Another important group of lytic enzymes, the lipases and phospholipases, have also been shown to be produced by C. auris [48, 49]. Lipases play an important role in biofilm formation, host cell damage and immune evasion [54, 55]. For, example null mutants of C. parapsilosis unable to secrete lipases were more efficiently ingested and killed by macrophage-like cells [54]. Moreover, lipase disruptant mutants of C. albicans and C. parapsilosis were shown to be significantly less virulent in a rat model of neonatal candidiasis [56]. A comparative analysis of multiple pathogenic Candida species demonstrated that C. albicans has the highest phospholipase activity [57]. Although C. auris has a similar amount of lipase encoding genes in its genome as C. albicans [24], the ability to produce phospholipases seems to be reduced and strain-dependent [48]. Production and secretion of a wide variety of enzymes is likely to contribute to rapid spread of C. auris. However, further investigation is warranted to reveal to which extend these enzymes are involved in C. auris virulence and pathogenicity.
Defense Against Antifungals: Intrinsic Resistance of Biofilms
Many microbes are found in biofilm ecosystems. The biofilm forms a structured microbial community encased in a matrix of exopolymeric material. Biofilm formation by Candida species is of particular interest, since its association with increased antifungal resistance and protecting cells within the biofilm from the host immune system [58]. This makes biofilm formation an important virulence trait for Candida species linked to excess morbidity and mortality [59, 60].
Multiple studies showed that C. albicans has the highest biofilm forming ability compared to other pathogenic Candida species [32, 57, 61]. Also C. auris strains form significantly reduced biofilms compared to C. albicans [32, 48]. On the other hand, biofilm formation by C. auris is significantly higher than that by C. glabrata [32]. Phenotypical observation also demonstrated C. auris biofilms being intermediate to that of C. albicans and C. glabrata. The C. auris biofilms mostly consists of budding yeasts and occasionally pseudohyphae embedded in a limited amount of extracellular matrix. C. albicans biofilms are formed by densely packed hyphae and yeast cells embedded in extracellular matrix, whereas C. glabrata forms a thin biofilm with yeast cells only, lacking extracellular matrix [32].
Transcriptomic analysis of temporally developing C. auris biofilms demonstrated adhesin-related glycosylphosphatidylinositol (GPI)-anchored cell wall genes (CSA1, IFF4, PGA26 and PGA52) are upregulated during every stage of biofilm production. Moreover, when the biofilm develops into more intermediate and mature stages the expression of genes encoding efflux pumps of the ABC transporter (CDR1, SNQ2 and YHD3) and major facilitator superfamily (MDR1 and RDC3) is increased [62].
Adherence is essential for biofilm formation. Agglutinin-like sequence (ALS) proteins, especially Als3, play a key role in C. albicans adherence [63, 64]. Interestingly, only two orthologues of members of the ALS proteins (Als1 and Als5) were found in the transcriptome of temporally developing C. auris biofilms [62]. Moreover, comparative genomic studies also revealed a highly reduced number of ALS and other proteins belonging to the adhesin and integrin gene families in C. auris compared to C. albicans [21, 24]. These findings seem to be in contrast with a recent study conducted by Singh and colleagues (2018) who identified three C. albicans Als3 protein homologs in C. auris through bioinformatic and structural homology modeling [65]. Even more interesting, antibodies targeting C. albicans Als3 protein were also able to bind in vitro to five C. auris strains, obtained from different clades. This showed the universal presence of Als3 homologs on the cell surface of C. auris. In addition, sera containing anti-Als3 antibodies significantly inhibit biofilm formation of C. auris, indicating an essential role of Als3 in biofilm formation similar to C. albicans [65]. However, the underrepresentation of the ALS protein family in C. auris implies that C. auris rely on additional adherence mechanisms. Likely, the GPI-anchored cell wall proteins play a role in C. auris adherence. For example, Csa1 and Iff4 have been associated with adherence in C. albicans [62, 66]. Despite Iff4 being associated with adherence to silicone catheter [66], the lack of ALS genes seems to impair adherence of C. auris, since C. albicans exhibits a significantly higher ability to adhere to silicone elastomer of catheters [48].
Candida biofilms show intrinsic resistance against antifungals. Several mechanisms were proposed to contribute to this resistance: (1) the high cell density within the biofilm; (2) decreased growth rate and nutrient limitation; (3) sequestration of drugs by the extracellular matrix (ECM); (4) the high expression of resistance genes, especially those encoding efflux pumps; and (5) the presence of ‘persister’ cells [58]. Also C. auris biofilms display lower susceptibility against antifungals, including caspofungin, micafungin and amphotericin B [32, 62]. The limited amount of ECM and reduced biomass of C. auris biofilms, compared to C. albicans, suggests other mechanisms to be more important for this reduced susceptibility [32, 48]. Nonetheless, sequestration of fluconazole by the ECM has recently been observed in C. auris biofilms [67].
High expression of efflux pumps also seems to play an important role, as a number of genes encoding these pumps are significantly upregulated in C. auris biofilms [62]. Moreover, inhibition of these pumps by an efflux pump inhibitor increases susceptibility of C. auris biofilms to fluconazole 2 to 8 times [62]. In addition, the ABC transporter Cdr1 was shown to be more highly expressed among azole-resistant isolates of C. auris [34, 68]. Deletion of this transporter dramatically decreased the minimum inhibitory concentration for all clinically available triazoles as much as 128-fold [68].
Candida auris Stress Tolerance and Persistence as a Nosocomial Pathogen
Candida auris is reported to be thermotolerant, being able to grow at temperatures up to 42 °C [16, 69, 70]. The ability of C. auris isolates to grow at elevated temperatures appears to be similar to C. albicans, but the closer related C. haemulonii seems to lose viability at temperatures above 37 °C [70]. In addition, C. auris shows excellent salt tolerance compared to other Candida species (e.g., C. albicans, C. duobushaemulonii, C. haemulonii, C. parapsilosis and C. tropicalis). When grown in saline Sabouraud broth (10% wt/vol NaCl), C. glabrata was the only species other than C. auris to have observable growth [71]. Improved tolerance to thermic and osmotic stresses compared to other Candida species is likely to contribute to the pathogenicity of C. auris. However, the molecular mechanism behind this increased tolerance remains elusive.
Candida auris not only displays excellent stress tolerance, it also has the alarming ability to persistently colonize the human host and the hospital environment. Nosocomial C. auris outbreaks have been reported in hospitals all around the world, some of them persisting up to 16 months [12, 72, 73]. Moreover, C. auris supposedly has the ability to cause low-grade disease years after colonization. Heath et al. [69] described a case of C. auris sternal osteomyelitis in a patient who was colonized by C. auris 3 years prior clinical disease manifestation. In addition, a case report from Belgium reported about a patient who was persistently colonized by echinocandin-resistant C. auris up to 18 months after its first detection [74]. This demonstrates that C. auris infection can progress slowly for > 12 months.
Although the ecological niches of C. auris remain unidentified, environmental sampling within the hospital environment has demonstrated C. auris to colonize and persist on abiotic surfaces such as bedding material, floors, sinks, as well as human skin, ears and nasal cavities [12, 71, 73, 75]. After a period of 7 days, C. auris recovery from dry or moist surfaces was shown to be similar to that of other clinically relevant Candida species, including C. albicans, C. glabrata and C. parapsilosis [75]. Additionally, it was shown that C. auris is able to remain viable for at least 14 days on a plastic health care surface, as measured by colony forming units (CFU) [71]. However, compared to C. parapsilosis, a species known to colonize plastic and skin, this was strongly reduced, since C. parapsilosis remains viable for at least 28 days on this kind of surfaces. On the other hand, using an esterase activity assay to measure individual cells for viability demonstrated that C. auris cells were viable 2 weeks longer than initially measured by CFU. This suggests that C. auris cells enter a viable but non-culturable state after 14 days [71]. Even though C. auris seems to be able to colonize plastic health care surfaces, this fungus shows a weak adherence ability to catheter surfaces made of silicone elastomer, compared to C. albicans [48]. Implying a reduced number of catheter-associated candidiasis caused by C. auris relative to C. albicans.
Several (review) articles report suboptimal efficacy of commonly used hospital environment disinfectants against C. auris as one of the factors contributing to its persistence within the hospital environment. However, multiple original studies show high efficacy of a plethora of commercially available disinfectants against C. auris compared to C. albicans [76,77,78]. Only quaternary ammonium-based disinfectants seem to be significantly less effective against C. auris, but also against C. albicans and C. glabrata [78]. However, most studies did not assess the efficacy of disinfectants on hospital surfaces and other materials commonly found in hospital settings, such as fabrics and polymer. To establish effective infection prevention protocols preventing the transmission of C. auris via contaminated surfaces, examination of C. auris disinfection on this kind of surfaces is needed. To this end, several qPCR assays that have recently been developed for the direct detection of C. auris can be applied, either by using DNA extracted directly from environmental swabs, overnight enrichment broth cultures or colonies [71, 79,80,81,82,83].
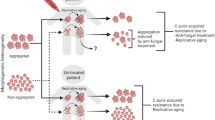
As mentioned before, the integrin and adhesin gene families seem to be underrepresented in the C. auris genome and transcriptome. Therefore, C. auris likely employs different strategies to adhere and persist on abiotic surfaces. The aggregation of cells into large and difficult to disperse clusters may be one of these strategies that promote persistence in the hospital environment. Aggregation seems to be strain-dependent and caused by the failure to release daughter cells after budding. This results in large aggregates that cannot be physically disrupted [84]. Although aggregating strains display significantly less virulence compared to non-aggregating strains, it might be that this morphology could have a role in protecting C. auris from detergents used to clean hospital environments. Further studies are required to unravel the molecular mechanism of persistence and adhesin in C. auris.
Candida auris Virulence In Vivo
Various experimental in vivo models have been developed to test virulence exhibited by Candida species [85]. The mouse model is widely used as a representative of the mammalian model. Despite mice being an excellent model to study pathogenicity and virulence of fungi, use of this model is facing ethical conflict and economic issues [86]. Since the innate immune system is evolutionary conserved in insects, invertebrate organisms such as Drosophila melanogaster [85], Caenorhabditis elegans [87, 88] Tenebrio molitor [89] and Galleria mellonella [86] are gaining interest as models for studying virulence traits of Candida species and host response against Candida infections.
Comparative analysis of virulence exhibited by C. auris strains and most other common pathogenic Candida species in the G. mellonella model demonstrated C. auris to be significantly more virulent than most of the tested species, with non-aggregative strains showing even similar pathogenicity as C. albicans (in terms of kinetics of larval death and number of larvae killed) [30]. This is remarkable, since pathogenicity of Candida species in G. mellonella was previously reported to be directly related to the development of hyphal filaments or pseudohyphae [84, 90]. Dissection of larvae infected with C. auris showed no hyphal or pseudohyphal formation by C. auris, indicating a different mechanism of pathogenicity in G. mellonella [30]. Sherry et al. [32] also confirmed non-aggregative C. auris strains to exhibit similar or even higher virulence than C. albicans in the G. mellonella model. Since biofilm formation is a key driver of C. albicans pathogenicity [59] and non-aggregative C. auris strains were shown to have better biofilm forming capacity compared to aggregative strains, this could be one of the factors explaining the difference in virulence between these strain types of C. auris [32].
Invertebrate models lack an adaptive immune system and are not suitable for organ colonization assessments. Moreover, some models (e.g., C. elegans and D. melanogaster) are not functional at the mammalian body temperature of 37 °C. Therefore, the validity of these models in reference to the human host is negotiable. To this end, a mouse model is likely to more closely represent the human situation.
A comparative study of C. auris virulence and its sibling C. haemulonii was performed in an immunosuppressed mouse model [70]. Mice infected with C. auris rapidly died with only 20% surviving after 5 days. In contrast, C. haemulonii isolates showed no virulence at all, with 100% of the mice surviving 12 days post-inoculation. Despite similar virulence observed in the G. mellonella model, death of mice infected with C. albicans was significantly faster than that of mice infected with C. auris [70]. Another comparative study of C. auris virulence, using an immunocompetent mouse model, showed similar results, with high virulence of C. auris, C. albicans and C. glabrata isolates. Although C. haemulonii did exhibit virulence, this was still significantly lower compared to all other Candida species tested. On the other hand, no significant differences in mice survival were found between C. auris with C. albicans and C. glabrata [91]. Nonetheless, a single C. auris isolate, obtained from a Chinese fungemic patient, was shown to have significant reduced virulence compared to C. albicans in both a mouse model and in G. mellonella. In the mouse model, all mice survived 14 days after infection with this C. auris isolate, where mice infected with C. albicans died by the 6th day post-infection [31]. This indicates the outcome of in vivo virulence assays with C. auris is highly strain specific. Fungal burden assays showed highest fungal load of C. auris and C. albicans to be detected in kidneys, spleen, liver and lungs, respectively [70, 91]. Interestingly, histopathological analysis showed yeast cell aggregates in kidneys of C. auris-infected mice, distinct from tissue invasive hyphae observed in kidneys of C. albicans-infected mice [70]. Altogether, compared to C. albicans, C. auris seem to be less virulent in the mouse model. This could be caused by the inability of C. auris to produce invasive hyphal filaments.
The immune response to C. auris was investigated using a zebrafish model of invasive candidiasis [92]. First in vitro experiments, using human neutrophils showed co-culturing of C. auris with human neutrophils had no effect on the initial fungal burden and C. auris even replicated beyond the initial inoculum. In contrast, neutrophils inhibited C. albicans growth with 75%. Moreover, in mixed cultures, neutrophils preferentially engulfed and killed C. albicans over C. auris. The strong antifungal response to C. albicans results in only 5% survival, while C. auris was strikingly resistant to neutrophil killing. The zebrafish model revealed that approximately 50% less neutrophils were recruited in response to C. auris infection when compared to C. albicans. Fluorescence microscopy revealed neutrophils failed to form neutrophil extracellular traps (NETs) in C. auris-infected zebrafish. In contrast, these antimicrobial structures were readily formed when zebrafish were challenged with C. albicans [92].
Taken together, the data provided by both mammalian and invertebrate in vivo models suggest C. auris is significantly more virulent than most other non-albicans Candida species. Even compared to C. albicans, the virulence of C. auris was not always significantly lower [91] and in the G. mellonella model non-aggregating strains were even shown to have higher virulence [30, 32]. The higher virulence of C. auris in invertebrate models could be explained by its ability to evade neutrophil attack [92]. Invertebrates only have an innate immune system and thus are relying mainly on granulocytes (neutrophil-like cells) as their defense against fungal infection [89], suggesting this system fails to kill C. auris effectively.
Conclusions and Research Outlooks
C. auris recently emerged as a global nosocomial pathogen associated with multidrug resistance and high mortality rates. However, the origin of this unprecedented emergence remains unclear. Genomic analyses revealed C. auris possesses many genes associated with virulence and reduced antifungal susceptibility, including genes encoding secreted aspartyl proteases, lipases, phospholipases, hemolysins and drugs efflux pumps [7, 21, 24]. Nonetheless, many genes are still uncharacterized and further investigation is required to understand the molecular mechanism responsible for the high pathogenicity and antifungal resistance of this pathogen.
Although the molecular mechanisms remain mostly unknown, this review shows that C. auris expresses many important virulence traits, such as biofilm formation, phenotypic switching, secretion of lytic enzymes and high stress tolerance, that are likely to have contributed to its emergence as a nosocomial pathogen. Moreover, C. auris seems to have the alarming ability to persistently colonize health care environments and human host, despite the reduced amount of adhesins in its genome.
An important limitation of virulence analysis in C. auris seems to be the high variability between strains, especially between strains of different clades [7]. For example, secretion of lytic enzymes was demonstrated to be strain specific [48]. More interestingly, antifungal susceptibility also seems to be highly heterogeneous, since multidrug-resistant strains [7] as well as totally susceptible strains have been isolated [31]. This shows that future studies should be careful when extrapolating findings to all isolates of the species and preferably incorporate multiple strains of different clades in their experimental setup.
Animal studies demonstrated C. auris is highly virulent and capable of inducing systemic infection and mortality to a much higher extent than other non-albicans Candida species, such as the closely related C. haemulonii or other potential multidrug-resistant yeast C. glabrata. Although strain specific, C. auris approaches the pathogenicity of C. albicans, which could be partially explained by the observation made that C. auris is able to evade the innate immune response and production of NETs by human neutrophils. In contrast, C. albicans is highly susceptible to killing by the innate immune system [30, 92].
In conclusion, C. auris expresses many important virulence traits, including traits that are well characterized in other Candida species, and seemingly unique traits, such as the ability to evade the innate immune system and persistently colonize the skin of human host. This together with the high propensity to develop resistance to multiple antifungals likely contributed to its emergence as a nosocomial pathogen.
References
Brown GD, Denning DW, Gow NA, Levitz SM, Netea MG, White TC. Hidden killers: human fungal infections. Sci Transl Med. 2012;4:165rv13.
Pfaller MA. Epidemiology of nosocomial candidiasis: the importance of molecular typing. Braz J Infect Dis. 2000;4:161–7.
Pappas PG, Lionakis MS, Arendrup MC, Ostrosky-Zeichner L, Kullberg BJ. Invasive candidiasis. Nat Rev Dis Primers. 2018;4:18026.
Cortegiani A, Misseri G, Chowdhary A. What’s new on emerging resistant Candida species. Intensive Care Med. 2019;45:512–5.
Ferreira A, Prado C, Carvalho R, Dias K, Dias A. Candida albicans and non-C. albicans Candida species: comparison of biofilm production and metabolic activity in biofilms, and putative virulence properties of isolates from hospital environments and infections. Mycopathologia. 2013;175:265–72.
Chowdhary A, Sharma C, Meis JF. Candida auris: a rapidly emerging cause of hospital-acquired multidrug-resistant fungal infections globally. PLoS Pathog. 2017;13:e1006290.
Lockhart SR, Etienne KA, Vallabhaneni S, Farooqi J, Chowdhary A, Govender NP, et al. Simultaneous emergence of multidrug-resistant Candida auris on 3 continents confirmed by whole-genome sequencing and epidemiological analyses. Clin Infect Dis. 2016;64:134–40.
Bidaud A, Chowdhary A, Dannaoui E. Candida auris: an emerging drug resistant yeast—a mini-review. J Myc Med. 2018;28:568–73.
Chowdhary A, Kumar VA, Sharma C, Prakash A, Agarwal K, Babu R, et al. Multidrug-resistant endemic clonal strain of Candida auris in India. Eur J Clin Microbiol Infect Dis. 2014;33:919–26.
Forsberg K, Woodworth K, Walters M, Berkow EL, Jackson B, Chiller T, et al. Candida auris: the recent emergence of a multidrug-resistant fungal pathogen. Med Myc. 2019;57:1–12.
de Cássia Orlandi Sardi J, Silva DR, Soares Mendes-Giannini MJ, Rosalen PL. Candida auris: Epidemiology, risk factors, virulence, resistance, and therapeutic options. Microb Pathog. 2018;125:116–21.
Schelenz S, Hagen F, Rhodes JL, Abdolrasouli A, Chowdhary A, Hall A, et al. First hospital outbreak of the globally emerging Candida auris in a European hospital. Antimicrob Resist Infect Control. 2016;5:35.
Cortegiani A, Misseri G, Fasciana T, Giammanco A, Giarratano A, Chowdhary A. Epidemiology, clinical characteristics, resistance, and treatment of infections by Candida auris. J Intensive Care. 2018;6:69.
Osei Sekyere J. Candida auris: a systematic review and meta-analysis of current updates on an emerging multidrug-resistant pathogen. MicrobiologyOpen. 2018;7:e00578.
Rossato L, Lopes Colombo A. Candida auris: what have we learned about its mechanisms of pathogenicity? Front Microbiol. 2018;9:3081.
Satoh K, Makimura K, Hasumi Y, Nishiyama Y, Uchida K, Yamaguchi H. Candida auris sp. nov., a novel ascomycetous yeast isolated from the external ear canal of an inpatient in a Japanese hospital. Microbiol Immunol. 2009;53:41–4.
Kathuria S, Singh PK, Sharma C, Prakash A, Masih A, Kumar A, et al. Multidrug-resistant Candida auris misidentified as Candida haemulonii: characterization by Matrix-Assisted Laser Desorption Ionization-Time of Flight Mass Spectrometry and DNA sequencing and its antifungal susceptibility profile variability by Vitek 2, CLSI broth microdilution, and Etest method. J Clin Microbiol. 2015;53:1823–30.
Mizusawa M, Miller H, Green R, Lee R, Durante M, Perkins R, et al. Can multidrug-resistant Candida auris be reliably identified in clinical microbiology laboratories? J Clin Microbiol. 2017;55:638–40.
Bao JR, Master RN, Azad KN, Schwab DA, Clark RB, Jones RS, et al. Rapid, accurate identification of Candida auris by using a novel Matrix-Assisted Laser Desorption Ionization-Time of Flight Mass Spectrometry (MALDI-TOF MS) database (library). J Clin Microbiol. 2018;56(4). pii: e01700-17.
Lee WG, Shin JH, Young U, Kang MG, Kim SH, Park KH, et al. The first three reported cases of nosocomial fungemia caused by Candida auris. J Clin Microbiol. 2011;49:3139–42.
Chatterjee S, Alampalli SV, Nageshan RK, Chettiar ST, Joshi S, Tatu US. Draft genome of a commonly misdiagnosed multidrug resistant pathogen Candida auris. BMC Genom. 2015;16:686.
Sharma C, Kumar N, Meis JF, Pandey R, Chowdhary A. Draft genome sequence of a fluconazole-resistant Candida auris strain from a candidemia patient in India. Genome Announc. 2015;3: pii: e00722-15.
Sharma C, Kumar N, Pandey R, Meis J, Chowdhary A. Whole genome sequencing of emerging multidrug resistant Candida auris isolates in India demonstrates low genetic variation. New Microbes New Infect. 2016;13:77–82.
Munoz JF, Gade L, Chow NA, Loparev VN, Juieng P, Farrer RA, et al. Genomic basis of multidrug-resistance, mating, and virulence in Candida auris and related emerging species. Nat Commun. 2018;9:5346.
Hayes BM, Anderson MA, Traven A, van der Weerden NL, Bleackley MR. Activation of stress signalling pathways enhances tolerance of fungi to chemical fungicides and antifungal proteins. Cell Mol Life Sci. 2014;71:2651–66.
Day AM, McNiff MM, da Silva Dantas A, Gow NAR, Quinn J. Hog1 regulates stress tolerance and virulence in the emerging fungal pathogen Candida auris. mSphere. 2018;3: pii: e00506-18.
Thompson DS, Carlisle PL, Kadosh D. Coevolution of morphology and virulence in Candida species. Eukaryot Cell. 2011;10:1173–82.
Desai J. Candida albicans hyphae: from growth initiation to invasion. J Fungi. 2018;4:10.
Whiteway M, Bachewich C. Morphogenesis in Candida albicans. Annu Rev Microbiol. 2007;61:529–53.
Borman AM, Szekely A, Johnson EM. Comparative pathogenicity of United Kingdom isolates of the emerging pathogen Candida auris and other key pathogenic Candida species. mSphere. 2016;1: pii: e00189-16.
Wang X, Bing J, Zheng Q, Zhang F, Liu J, Yue H, et al. The first isolate of Candida auris in China: clinical and biological aspects. Emerg Microbes Infect. 2018;7:93.
Sherry L, Ramage G, Kean R, Borman A, Johnson EM, Richardson MD, et al. Biofilm-forming capability of highly virulent, multidrug-resistant Candida auris. Emerg Infect Dis. 2017;23:328–31.
Yue H, Bing J, Zheng Q, Zhang Y, Hu T, Du H, et al. Filamentation in Candida auris, an emerging fungal pathogen of humans: passage through the mammalian body induces a heritable phenotypic switch. Emerg Microbes Infect. 2018;7:188.
Kim SH, Iyer KR, Pardeshi L, Munoz JF, Robbins N, Cuomo CA, et al. Genetic analysis of Candida auris implicates Hsp90 in morphogenesis and azole tolerance and Cdr1 in azole resistance. mBio. 2019;10: pii: e02529-18.
Lo H, Köhler JR, DiDomenico B, Loebenberg D, Cacciapuoti A, Fink GR. Nonfilamentous C. albicans mutants are avirulent. Cell. 1997;90:939–949.
Bommanavar SB, Gugwad S, Malik N. Phenotypic switch: the enigmatic white-gray-opaque transition system of Candida albicans. J Oral Maxillofac Pathol. 2017;21:82–6.
Lachke SA, Joly S, Daniels K, Soll DR. Phenotypic switching and filamentation in Candida glabrata. Microbiology. 2002;148:2661–74.
Porman AM, Alby K, Hirakawa MP, Bennett RJ. Discovery of a phenotypic switch regulating sexual mating in the opportunistic fungal pathogen Candida tropicalis. Proc Natl Acad Sci USA. 2011;108:21158–63.
Xie J, Du H, Guan G, Tong Y, Kourkoumpetis TK, Zhang L, et al. N-Acetylglucosamine induces white-to-opaque switching and mating in Candida tropicalis providing new insights into adaptation and fungal sexual evolution. Eukaryot Cell. 2012;11:773–82.
Fukui K, Nakamura K, Kuwashima H, Majima T. White-to-opaque switching is involved in the phospholipase B production of Candida dubliniensis on Price’s egg yolk agar. Odontology. 2019;107:174–85.
Alkafeef SS, Yu C, Huang L, Liu H. Wor1 establishes opaque cell fate through inhibition of the general co-repressor Tup1 in Candida albicans. PLoS Genet. 2018;14:e1007176.
Bentz ML, Sexton DJ, Welsh RM, Litvintseva AP. Phenotypic switching in newly emerged multidrug-resistant pathogen Candida auris. Med Mycol. 2018. https://doi.org/10.1093/mmy/myy100.
Favel A, Michel-Nguyen A, Peyron F, Martin C, Thomachot L, Datry A, et al. Colony morphology switching of Candida lusitaniae and acquisition of multidrug resistance during treatment of a renal infection in a newborn: case report and review of the literature. Diagn Microbiol Infect Dis. 2003;47:331–9.
Miller NS, Dick JD, Merz WG. Phenotypic switching in Candida lusitaniae on copper sulfate indicator agar: association with amphotericin B resistance and filamentation. J Clin Microbiol. 2006;44:1536–9.
Rapala-Kozik M, Bochenska O, Zajac D, Karkowska-Kuleta J, Gogol M, Zawrotniak M, et al. Extracellular proteinases of Candida species pathogenic yeasts. Mol Oral Microbiol. 2018;33:113–24.
Naglik JR, Challacombe SJ, Hube B. Candida albicans secreted aspartyl proteinases in virulence and pathogenesis. Microbiol Mol Biol Rev. 2003;67:400–28.
Lee SA, Jones J, Hardison S, Kot J, Khalique Z, Bernardo SM, et al. Candida albicans VPS4 is required for secretion of aspartyl proteases and in vivo virulence. Mycopathologia. 2009;167:55–63.
Larkin E, Hager C, Chandra J, Mukherjee PK, Retuerto M, Salem I, et al. The emerging pathogen Candida auris: growth phenotype, virulence factors, activity of antifungals, and effect of SCY-078, a novel glucan synthesis inhibitor, on growth morphology and biofilm formation. Antimicrob Agents Chemother. 2017;61: pii: e02396-16.
Kumar D, Banerjee T, Pratap CB, Tilak R. Itraconazole-resistant Candida auris with phospholipase, proteinase and hemolysin activity from a case of vulvovaginitis. J Infect Dev Countries. 2015;9:435–7.
Furlaneto MC, Góes HP, Perini HF, dos Santos RC, Furlaneto-Maia L. How much do we know about hemolytic capability of pathogenic Candida species? Folia Microbiol (Praha). 2018;63:405–12.
Rossoni RD, Barbosa JO, Vilela SFG, Jorge AOC, Junqueira JC. Comparison of the hemolytic activity between C. albicans and non-albicans Candida species. Braz Oral Res. 2013;27:484–9.
Luo G, Samaranayake LP, Yau JY. Candida species exhibit differential in vitro hemolytic activities. J Clin Microbiol. 2001;39:2971–4.
Seneviratne C, Wong S, Yuen K, Meurman J, Pärnänen P, Vaara M, et al. Antifungal susceptibility and virulence attributes of bloodstream isolates of Candida from Hong Kong and Finland. Mycopathologia. 2011;172:389–95.
Gacser A, Trofa D, Schafer W, Nosanchuk JD. Targeted gene deletion in Candida parapsilosis demonstrates the role of secreted lipase in virulence. J Clin Invest. 2007;117:3049–58.
Ghannoum MA. Potential role of phospholipases in virulence and fungal pathogenesis. Clin Microbiol Rev. 2000;13:122–43.
Trofa D, Soghier L, Long C, Nosanchuk JD, Gacser A, Goldman DL. A rat model of neonatal candidiasis demonstrates the importance of lipases as virulence factors for Candida albicans and Candida parapsilosis. Mycopathologia. 2011;172:169–78.
Dabiri S, Shams-Ghahfarokhi M, Razzaghi-Abyaneh M. Comparative analysis of proteinase, phospholipase, hydrophobicity and biofilm forming ability in Candida species isolated from clinical specimens. J Myc Med. 2018;28:437–42.
Ramage G, Saville SP, Thomas DP, Lopez-Ribot JL. Candida biofilms: an update. Eukaryot Cell. 2005;4:633–8.
Rajendran R, Sherry L, Nile CJ, Sherriff A, Johnson E, Hanson M, et al. Biofilm formation is a risk factor for mortality in patients with Candida albicans bloodstream infection—Scotland, 2012–2013. Clin Microbiol Infect. 2016;22:87–93.
Tumbarello M, Fiori B, Trecarichi EM, Posteraro P, Losito AR, De Luca A, et al. Risk factors and outcomes of candidemia caused by biofilm-forming isolates in a tertiary care hospital. PLoS ONE. 2012;7:e33705.
Kuhn DM, Ghannoum MA. Candida biofilms: antifungal resistance and emerging therapeutic options. Curr Opin Investig Drugs. 2004;5:186–97.
Kean R, Delaney C, Sherry L, Borman A, Johnson EM, Richardson MD, et al. Transcriptome assembly and profiling of Candida auris reveals novel insights into biofilm-mediated resistance. mSphere. 2018;3: pii: e00334-18.
Zhao X, Daniels KJ, Oh SH, Green CB, Yeater KM, Soll DR, et al. Candida albicans Als3p is required for wild-type biofilm formation on silicone elastomer surfaces. Microbiology. 2006;152:2287–99.
Liu Y, Filler SG. Candida albicans Als3, a multifunctional adhesin and invasin. Eukaryot Cell. 2011;10:168–73.
Singh S, Uppuluri P, Alqarihi A, Elhassan H, French S, Lockhart SR, et al. The NDV-3A vaccine protects mice from multidrug resistant Candida auris infection. BioRxiv. 2018:465096.
Kempf M, Cottin J, Licznar P, Lefrançois C, Robert R, Apaire-Marchais V. Disruption of the GPI protein-encoding gene IFF4 of Candida albicans results in decreased adherence and virulence. Mycopathologia. 2009;168:73–7.
Dominguez E, Zarnowski R, Choy H, Zhao M, Sanchez H, Nett JE, et al. Conserved role for biofilm matrix polysaccharides in Candida auris drug resistance. mSphere. 2019;4:e00680-18.
Rybak JM, Doorley LA, Nishimoto AT, Barker KS, Palmer GE, Rogers PD. Abrogation of triazole resistance upon deletion of CDR1 in a clinical isolate of Candida auris. Antimicrob Agents Chemother. 2019;63: pii: e00057-19.
Heath CH, Dyer JR, Pang S, Coombs GW, Gardam DJ. Candida auris sternal osteomyelitis in a man from Kenya visiting Australia, 2015. Emerg Infect Dis. 2019;25:192–4.
Ben-Ami R, Berman J, Novikov A, Bash E, Shachor-Meyouhas Y, Zakin S, et al. Multidrug-resistant Candida haemulonii and C. auris, Tel Aviv, Israel. Emerg Infect Dis. 2017;23:195–203.
Welsh RM, Bentz ML, Shams A, Houston H, Lyons A, Rose LJ, et al. Survival, persistence, and isolation of the emerging multidrug-resistant pathogenic yeast Candida auris on a plastic health care surface. J Clin Microbiol. 2017;55:2996–3005.
Calvo B, Melo AS, Perozo-Mena A, Hernandez M, Francisco EC, Hagen F, et al. First report of Candida auris in America: clinical and microbiological aspects of 18 episodes of candidemia. J Infect. 2016;73:369–74.
Vallabhaneni S, Kallen A, Tsay S, Chow N, Welsh R, Kerins J, et al. Investigation of the first seven reported cases of Candida auris, a globally emerging invasive, multidrug-resistant fungus—United States, May 2013–August 2016. Am J Transplant. 2017;17:296–9.
Dewaele K, Frans J, Smismans A, Ho E, Tollens T, Lagrou K. First case of Candida auris infection in Belgium in a surgical patient from Kuwait. Acta Clin Belg. 2018. https://doi.org/10.1080/17843286.2018.1555114.
Piedrahita CT, Cadnum JL, Jencson AL, Shaikh AA, Ghannoum MA, Donskey CJ. Environmental surfaces in healthcare facilities are a potential source for transmission of Candida auris and other Candida species. Infect Control Hosp Epidemiol. 2017;38:1107–9.
Rutala WA, Kanamori H, Gergen MF, Sickbert-Bennett EE, Weber DJ. Susceptibility of Candida auris and Candida albicans to 21 germicides used in healthcare facilities. Infect Control Hosp Epidemiol. 2019;40:380–2.
Abdolrasouli A, Armstrong-James D, Ryan L, Schelenz S. In vitro efficacy of disinfectants utilised for skin decolonisation and environmental decontamination during a hospital outbreak with Candida auris. Mycoses. 2017;60:758–63.
Cadnum JL, Shaikh AA, Piedrahita CT, Sankar T, Jencson AL, Larkin EL, et al. Effectiveness of disinfectants against Candida auris and other Candida species. Infect Control Hosp Epidemiol. 2017;38:1240–3.
Arastehfar A, Fang W, Daneshnia F, Al-Hatmi AM, Liao W, Pan W, et al. Novel multiplex real-time quantitative PCR detecting system approach for direct detection of Candida auris and its relatives in spiked serum samples. Future Microbiol. 2018;14:33–45.
Sexton DJ, Kordalewska M, Bentz ML, Welsh RM, Perlin DS, Litvintseva AP. Direct detection of emergent fungal pathogen Candida auris in clinical skin swabs by SYBR Green-based quantitative PCR assay. J Clin Microbiol. 2018;56: pii: e01337-18.
Martínez-Murcia A, Navarro A, Bru G, Chowdhary A, Hagen F, Meis JF. Internal validation of GPS™ MONODOSE CanAur dtec-qPCR kit following the UNE/EN ISO/IEC 17025: 2005 for detection of the emerging yeast Candida auris. Mycoses. 2018;61:877–84.
Kordalewska M, Zhao Y, Lockhart SR, Chowdhary A, Berrio I, Perlin DS. Rapid and accurate molecular identification of the emerging multidrug-resistant pathogen Candida auris. J Clin Microbiol. 2017;55:2445–52.
Leach L, Zhu Y, Chaturvedi S. Development and validation of a real-time PCR assay for rapid detection of Candida auris from surveillance samples. J Clin Microbiol. 2018;56: pii: e01223-17.
Borman AM, Szekely A, Johnson EM. Comparative pathogenicity of United Kingdom isolates of the emerging pathogen Candida auris and other key pathogenic Candida species. mSphere. 2016;1(4): pii: e00189-16.
Segal E, Frenkel M. Experimental in vivo models of candidiasis. J Fungi. 2018;4:21.
Trevijano-Contador N, Zaragoza O. Immune response of Galleria mellonella against human fungal pathogens. J Fungi. 2019;5:3.
Elkabti A, Issi L, Rao R. Caenorhabditis elegans as a model host to monitor the Candida infection processes. J Fungi. 2018;4:123.
Ortega-Riveros M, De-la-Pinta I, Marcos-Arias C, Ezpeleta G, Quindós G, Eraso E. Usefulness of the non-conventional Caenorhabditis elegans model to assess Candida virulence. Mycopathologia. 2017;182:785–95.
Canteri de Souza P, Custódio Caloni C, Wilson D, Sergio Almeida R. An invertebrate host to study fungal infections, mycotoxins and antifungal drugs: Tenebrio molitor. J Fungi. 2018;4:125.
Cotter G, Doyle S, Kavanagh K. Development of an insect model for the in vivo pathogenicity testing of yeasts. FEMS Immunol Med Microbiol. 2000;27:163–9.
Fakhim H, Vaezi A, Dannaoui E, Chowdhary A, Nasiry D, Faeli L, et al. Comparative virulence of Candida auris with Candida haemulonii, Candida glabrata and Candida albicans in a murine model. Mycoses. 2018;61:377–82.
Johnson CJ, Davis JM, Huttenlocher A, Kernien JF, Nett JE. Emerging fungal pathogen Candida auris evades neutrophil attack. mBio. 2018;9: pii: e01403-18.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Research Involving Human Participants and/or Animals
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Handling Editor: Vishnu Chaturvedi.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
de Jong, A.W., Hagen, F. Attack, Defend and Persist: How the Fungal Pathogen Candida auris was Able to Emerge Globally in Healthcare Environments. Mycopathologia 184, 353–365 (2019). https://doi.org/10.1007/s11046-019-00351-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11046-019-00351-w