Abstract
N6-methyladenosine (m6A), the methylation targeting the N6 position of adenosine, is the most common internal modification of mRNA in eukaryotes. Considering the roles of m6A in regulating gene expression, the investigation of m6A roles in the biological processes including cell renewal, differentiation, apoptosis, and invasion of cancer cells has become a hot research topic. There are three kinds of protein involved in m6A regulation. The methyltransferases and demethylases cooperatively regulate the m6A levels, while the m6A reading proteins recognize the m6A sites and mediate multiple m6A-dependent biological functions including mRNA splicing, transfer, translation, and degradation. At present, a large number of studies have found that the changes of m6A levels in tumor cells play a very important role in the occurrence and development of tumors, as well as metastasis and invasion of tumor cells. This review summarizes the different roles of m6A modification in the occurrence and development of various cancers, and discusses the possibility of choosing the m6A related proteins as potential therapeutic targets.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The posttranscriptional modifications of mRNA play important roles in regulating a series of physiological processes and disease development, and thus the studies about their function have become the hot topics. Currently, more than 100 RNA modifications have been identified [1]. N6-methyladenosine (m6A) is the most common methylation modification in eukaryotic mRNA and long non-coding RNA, and is found in ribosome-related mRNA as well [2]. In addition, m6A is identified in more than 25% of human mRNAs [3]. As the most common internal modification in mRNA, m6A modification is generally enriched in 3′-untranslated terminal regions (3′-UTRs) and near stop codons [4]. Zhang et al., developed a new FunDMDeep-m6A algorithm to detect the dynamic m6A levels in cells, and confirmed that m6A modifications could target many important genes involved in biological processes including embryonic development, stem cell differentiation, cell apoptosis, as well as proliferation and metastasis of cancer cells [3].
There are three types of regulatory proteins involved in m6A occurrence, elimination and function exertion; the methyltransferases (writers) are involved in catalyzing methyl transfer; the demethylase (erasers) can remove m6A modification; various m6A reading proteins can recognize and bind to m6A-modified mRNAs to mediate corresponding functions [5]. It is noted that three kinds of regulatory proteins constitute the molecular basis for m6A regulation in multiple metabolic processes of RNA.
The molecular basis for the occurrence, elimination and function of m6A
The occurrence of m6A: m6A writers
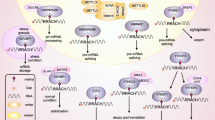
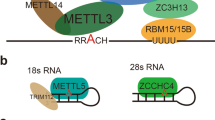
The m6A methyltransferase complex is composed of METTL3, METTL14, WTAP, and other auxiliary subunits including VIRMA [6, 7], RBM15 [8], and ZC3H13 [9] (Fig. 1). METTL3 is the catalytic part of the methyltransferase complex, and catalyzes the modification of m6A by cross-linking with S-adenosylmethionine [10]. METTL14, one of the most important auxiliary subunits in the methyltransferase complex, combines with METTL3 to form a heterodimer to activate the catalytic function of METTL3. In addition, METTL14 can recognize RNA substrates and then mediate the binding of methyltransferase complex to RNA substrates [6]. WTAP acts as a regulatory subunit to recruit METTL3/14 to the mRNA for subsequent m6A methylation [11]. Previous studies have reported that VIRMA mediates preferential methylation of mRNA near the 3’-UTR and stop codons [7]; RBM15 and RBM15B mediate m6A modification in long non-coding RNA X-inactive specific transcripts (XIST) [8]. Moreover, ZC3H13 has been proved to guide the binding of RNA-binding protein Rbm15 to mammalian WTAP [9]. In addition to the METTL3/14 complex, METTL16 is also one kind of m6A methyltransferases, which mainly functions on a large number of pre-mRNAs and various non-coding RNAs [12].
Elimination of m6A: m6A erasers
m6A modification is actually a dynamically reversible process, as the modification on mRNA can be eliminated via m6A demethylases. There are two known common demethylases of m6A: fat mass and obesity-associated protein (FTO) and AlkB homolog 5 (ALKBH5) (Fig. 1). As a member of the AlkB family, FTO has a highly conserved catalytic domain, which mainly acts on 3′-untranslated region of transcripts to regulate the m6A level. FTO has been found to be of great significance to the m6A modification of whole transcriptome mRNA [13], and closely related to metabolic diseases such as diabetes [14], obesity [15], and ischemic heart failure [16]. ALKBH5, a member of the AlkB family, has been proved to be another one mammalian demethylase [17]. By regulating the m6A levels, ALKBH5 performs biological functions including regulating cell proliferation, migration, invasion, and ossification [18]. Another demethylase ALKBH3, which is relatively less reported, usually preferentially exhibits demethylation activity in ssDNA, and there are also recent studies reporting that it exhibits demethylation activity in tRNA [19].
Recognition of m6A modification: m6A readers
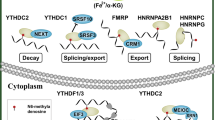
After specifically recognizing the m6A sites of RNA, m6A reading proteins mediate a series of biological functions by modulating splicing, translation, transport, and enhancing or reducing the stability of target mRNAs. The YTH protein family proteins, including YTHDF1, YTHDF2, YTHDF3, YTHDC1 and YTHDC2, have been confirmed as m6A reading proteins. YTHDF1 promotes translation of target mRNAs, while YTHDF2 reduces the stability of the target transcripts (Fig. 1) [20]. YTHDF3 assists YTHDF1 and YTHDF2 to promote RNA translation and modulate RNA degradation, respectively [21, 22]. YTHDC1, the m6A reading protein in nucleus, promotes the transport of mRNA from the nucleus to the cytoplasm [23], and also mediates RNA splicing in a concentration-dependent manner [24]. YTHDC2 is also the member of the YTH protein family, which can improve the translation efficiency or reduce the mRNA abundance of target genes [25].
The insulin-like growth factor 2 mRNA-binding proteins (IGF2BPs) is a newly discovered m6A reading proteins. Different from the YTH protein family, IGF2BPs promote expression of target genes by increasing the stability of target mRNAs [26]. Studies have confirmed that eukaryotic initiation factor 3eIF3 can get rid of the assistance of cap complex and independently recruit the 43S ribosomal complex to start translation after binding to the m6A sites in 5'-UTR of mRNAs, indicating that eIF3 regulates mRNA cap-independent translation after recognizing and binding to m6A-modified mRNAs [27]. Moreover, METTL3 was also observed to enhance the translation of target mRNAs by activating translation process in human cancer cells [28].
m6A regulation in the tumors
Acute myeloid leukemia (AML)
Acute myeloid leukemia (AML) is an aggressive and fatal hematological malignancy, characterized by abnormal proliferation of primitive and naive myeloid cells in the bone marrow and the periphery. Epigenetic modifications including DNA methylation and histone modifications play important roles in phenotype maintenance of leukemia cells [29]. METTL3 and METTL14 have been reported to be highly expressed in AML cells relative to hematopoietic progenitor cells, and both exhibit carcinogenic effects [30, 31]. METTL3 induces enhanced expression of SP1 by upregulating the m6A methylation levels on the oncogenes SP1 and SP2 mRNAs, and SP1 promotes the differentiation of hematopoietic stem cells into AML cells [32]. In addition, METTL3 can promote the expression of c-MYC, BCL-2 and PTEN in AML cells to increase intracellular p-AKT level, thereby promoting proliferation of cancer cells and exerting carcinogenic effects [30]. Weng et al. identified that the SP1-METTL14-MYB/MYC signal axis regulates myeloid differentiation of normal cells and participates in malignant hematopoiesis [31]. In addition, Bansal et al., identified that WTAP plays an important role in the proliferation and differentiation of leukemia cells, and thus WTAP is expected to become a new target for the treatment of AML [33].
Similar to m6A writers, m6A erasers also play a role in regulating the occurrence and development of AML, among which FTO and ALKBH5 are abnormally active in various karyotypes of AML [34, 35]. Mechanism studies have found that FTO reduces the m6A levels of target mRNAs, such as tumor suppressor genes ASB-2 and RARA in the untranslated region, resulting in downregulation of their mRNA stability and thus promoting the AML development [34]. Shen et al. found that ALKBH5, overexpressed in AML, functions as an oncoprotein, and its upregulated expression is associated with poor prognosis of AML [35]. By contrast, Kwok’s study indicated that ALKBH5 is reduced in AML cells and suggested that it exhibits a tumor suppressor effect [36]. In another one study, ALKBH5 and YTHDF2 cooperatively regulate the mRNA stability of the receptor tyrosine kinase AXL in a m6A-dependent manner [37]; AXL overexpression promotes cancer cell proliferation, survival, migration, and invasion by activating PI3K/Akt and MAPK/Erk pathways [38].
Glioblastoma (GBM)
Glioblastoma, originating from poorly differentiated glial cells, is one of the most common malignant tumors in the central nervous system. Cai et al., showed that m6A methylation modification is involved in the self-renewal of glioblastoma stem cells (GSCs), and pathogenesis and development of tumors. Moreover, Cui et al., further proved that METTL3 and METTL14 can regulate growth of GSCs and GBM progression by down-regulating the expression of oncogenes, such as ADAM19, EPHA3, and KLF4 [39]. However, contradictory results about METTL3 in GBM progression were obtained, which suggested that METTL3 promotes the growth of GSCs by up-regulating SOX2 expression when these cells are exposed to radiation [40]. In addition, METTL3 has also been found to function as a regulator of nonsense-mediated mRNA decay (NMD) to maintain the aggressiveness of tumor [41]. These results suggest that the effects of METTL3 on the pathogenesis and development of GBM are diverse and complex, thus further investigation about METTL3 roles in GBM progression is needed. In glioblastoma, overexpressed ALKBH5 is associated with enhanced ability of tumor stem cells to resist radiation and invasion [42]; mechanistically, ALKBH5 reduces the level of intracellular m6A methylation and thus promotes the expression of the oncogene FOXM1 to enhance the self-renewal ability and tumorigenicity of GSCs [43]. Moreover, the m6A reading protein IGF2BP2 is observed to be upregulated in GBM tissues, which promotes the proliferation, migration, invasion and epithelial-mesenchymal transition of GBM cells by upregulating insulin-like growth factor 2 (IGF2) and then activating PI3K/AKT signaling pathway [44]. As predicted, inhibition of PI3K/AKT pathway can increase the sensitivity of GBM to temozolomide (TMZ) treatment [45]. Collectively, these results suggest that m6A modification mediates the occurrence and migration of GBM, which provides insight into therapeutic strategies by exploiting m6A RNA methylation as targets for treating GBM.
Hepatocellular carcinoma (HCC)
Hepatocellular carcinoma (HCC), a primary liver cancer with high mortality, is the third most common cause of cancer-related high mortality in the world [46]. Increasing evidences show that HCC is closely related to m6A. The TCGA analyses indicate the overall survival and disease-free survival rate of HCC patients with high METTL3 expression are poor. RNA-Seq assays revealed that METTL3 is significantly upregulated in HBV-related liver cancer tissues relative to corresponding non-tumor (NT) liver tissues. Further studies demonstrated that METTL3 reduces the mRNA stability of the tumor suppressor SOCS2 through the YTHDF2-dependent pathways in HCC, bringing about the pathogenesis and development of HCC [47]. Moreover, Chen et al. determined that Snail, an important transcription factor related to EMT, is a downstream target of METTL3; METTL3 and YTHDF1 jointly promote the transfer of HCC by enhancing translation of Snail protein [48].
METTL14 was examined to be downregulated in HCC [49]. Mechanism studies revealed that METTL14 mediates the recognition and binding of DiGeorge syndrome chromosomal region 8 (DGCR8) to pri-miR-126 through m6A-dependent pathways, resulting in miR-126 maturation [50]. It is noted that miR-126 functions as a tumor suppressor and is downregulated in a variety of tumors [51]. In addition, Li et al. identified that METTL14 might participate in malignant progression of HCC by regulating the m6A modification levels of cysteine sulfinic acid decarboxylase (CSAD), SOCS2, and glutamic-oxaloacetic transaminase 2 (GOT2) [52].
The m6A methyltransferase WTAP, KIAA1429 and the demethylase FTO have also been reported to influence the HCC growth and invasion through different mechanisms [53,54,55,56]. FTO has been reported as one tumor-promoting effector or tumor suppressor for the pathogenesis and development of HCC, respectively [55, 56]. Thus, the influences of FTO on the proliferation ability of different HCC cells were controversial, which needs further research and verification. In addition, ALKBH5 was observed to be reduced in HCC. Functional studies further confirmed that ALKBH5 could inhibit the growth and invasion of liver cancer cells in vivo and in vitro. Decreased ALKBH5 increases the m6A modification of LYPD1, which hinders the recognition of m6A-modified LYPD1 by the m6A reading protein IGF2BP1 and thus enhances LYPD1 mRNA stability, resulting in tumor-inducing effect in HCC [57]. The above studies have enriched the understanding of m6A roles in HCC development, and provided different perspectives and insights for the developing effective treatment strategies.
Gastric cancer (GC)
Although the incidence of gastric cancer (GC) has decreased in the past few decades, GC is still the fifth most common malignant tumor in the world [58]. In GC tissues, the expression of METTL3 was significantly increased. Wang et al., found that METTL3 increases the m6A modification levels of HDGF mRNA, which are then recognized and bound by IGF2BP3 to upregulate HDGF protein level, promoting tumor angiogenesis and glycolysis and thus accelerating tumor growth [59]. Moreover, a previous study showed that METTL3 downregulation can activate the apoptotic pathway and inactivate the AKT signaling pathway, thereby inhibiting the proliferation and migration of human GC cells [60]. Taken together, it is speculated that METTL3 may be a potential target for the treatment of human GC.
Zhang et al. also confirmed that ALKBH5 can promote the invasion and metastasis of GC by reducing m6A modification levels of the lncRNA NEAT1 [60], suggesting that ALKBH5 and NEAT1 may be potential therapeutic targets for GC.
Compared with the benign gastric disease patients and healthy groups, the m6A levels in the peripheral blood of GC patients were significantly increased with the progress and metastasis of GC; while the m6A levels in the GC patients decreased after surgery. Compared with healthy groups, the expression of ALKBH5 and FTO in the GC patients was significantly down-regulated. These results suggest that m6A modification of peripheral blood can be used as a new non-invasive biomarker for GC prediction and diagnosis [61]. Moreover, He et al., demonstrated that m6A modification plays a regulatory role in miR-660-mediated inhibition of GC cells, providing a novel perspective for the m6A regulatory mechanism in GC development [62].
However, Zhang et al., obtained contradictory results and found that the m6A methyltransferases are potential tumor suppressors, while demethylases are potential cancer promoters. Further studies revealed that METTL14 knockdown could promote the proliferation and invasion of GC cells by activating the oncogenic Wnt/PI3K-AKT signaling, while FTO knockdown exhibits the opposite effects [63]. These results indicate that the molecular mechanism of m6A involved in the occurrence and development of GC is still complicated and worthy of further exploration.
Breast cancer (BC)
Breast cancer (BC) is the most common cause of death among women in the world. Although early treatment of BC is effective, 30% of patients still face the risk of tumor recurrence or metastasis [64, 65]. Under hypoxic conditions in the tumor microenvironment, the expression of ALKBH5 in Breast cancer stem cells (BCSCs) is increased, which reduces the m6A methylation levels in tumor tissues and ultimately promotes the enrichment of BCSCs in the hypoxic microenvironment. Mechanistically, ALKBH5 reduces the m6A methylation level of NANOG, a totipotent or pluripotent stem cell transcription factor, to upregulate its mRNA stability [66, 67]. Furthermore, hypoxia can also induce expression of zinc finger protein 217 (ZNF217) in BC cells in a HIF-dependent manner. As an inhibitor of m6A methyltransferase, ZNF217 reduces METTL3 but upregulates ALKBH5 level to negatively regulate m6A levels of downstream genes, resulting in promoting BC tumorigenesis by increasing expression of KLF4 and NANOG [68].
As a pro-apoptotic gene, BNIP3 is a tumor suppressor in BC. In BC, the expression of FTO is upregulated. FTO targets BNIP3 and mediates its mRNA demethylation, which induce degradation of BNIP3 mRNA in YTHDF2-dependent manner, resulting in BC cell proliferation, metastasis and colony formation [69].
The methyltransferase METTL3 is highly expressed in BC, and has a strong positive correlation with expression of oncoprotein hepatitis B virus x-interacting protein (HBXIP) in tumor development. Moreover, METTL3 increases HBXIP expression through methylation modification, and HBXIP in turn promotes the expression of METTL3 by inhibiting the METTL3 inhibitor miRNA let-7 g expression. These results identify that HBXIP/let-7 g/METTL3/HBXIP forms a positive feedback pathway, accelerating the development of BC [70]. Furthermore, Wang et al., found that METTL3 targets Bcl-2 and increases its transcription, thereby inhibiting cell apoptosis and promoting the progress of BC [71]. Collectively, these results provide new potential therapeutic targets for BC.
Non-small-cell lung carcinoma (NSCLC)
In China, lung cancer is one of the malignant tumors with the highest morbidity and mortality, and it is the greatest threat to the health and life of the population [72, 73]. According to histopathological criteria, lung cancer can be divided into small-cell lung carcinoma (SCLC) and NSCLC, among which common non-small-cell carcinomas include lung adenocarcinoma (LUAD) and squamous cell carcinoma. Lin et al., found that METTL3 mRNA levels in LUAD are significantly increased; further studies identified that METTL3 regulates LUAD growth and invasion [28]. Moreover, other studies revealed that METTL3 induces brain metastasis of lung cancer by regulating expression of miR-143-3p and vascular tissue protein 1 (vasohibin-1, VASH1) [74]. METTL3 also participates in the Transforming Growth Factor-beta (TGF-β)-mediated EMT process of lung cancer cells via modulating the expression of JUNB, a key transcriptional regulator of EMT [75]. In another study on NSCLC, miR-33a was observed to directly binds to the 3′ non-coding region of METTL3 mRNA to reduce the expression of METTL3, which in turn inhibits cancer cell proliferation by decreasing the expression of target genes EGFR, TAZ and DNMT3A [76]. These results suggest that METTL3 may become one of the targets of lung cancer treatment.
Liu et al., confirmed that FTO promotes the progression of lung squamous cell carcinoma via reducing the m6A methylation level of zinc finger 1 (MZF1) and in turn enhancing its protein levels [77, 78]. Li et al., also found that FTO can accelerate the growth of lung cancer cells by targeting the m6A level of ubiquitin-specific protease 7 (USP7) [79].
Other tumors
m6A has been found to be involved in the occurrence and development of human urinary system-related tumors in recent studies including renal cell carcinoma (RCC), bladder cancer (BCA), and male prostate cancer (PCA) [80,81,82,83,84,85,86]. RCC is the most fatal malignant tumor of the urinary system [58], in which METTL3 can inhibit the proliferation, migration and epithelial-mesenchymal transition of renal cancer cells by regulating the PI3K-AKT-mTOR pathway [80]. Methylenetetrahydrofolate dehydrogenase 2 (MTHFD2), overexpressed in RCC, can enhance the m6A modification of hypoxia-inducible factor-2α (HIF-2α) and thus induce translation of HIF-2α, promoting glycolytic metabolism in tumor cells and malignant phenotype of tumors [81]. In BCA, METTL3 with increased expression upregulates the methylation level of CUB domain-containing protein 1 (CUB domain-containing protein 1, CDCP1), and YTHDF1 then recognizes the m6A motifs in CDCP1 3’-UTR to facilitate translation of the oncogene CDCP1 [82]. Recent studies have demonstrated that the METTL3-YTHDF2 axis can directly target and recognize the downstream tumor suppressor genes SETD7 and KLF4 mRNA to induce their degradation, thereby promoting the pathogenesis and development of BCA [83]. In addition, Gao et al. revealed the METTL3-AFF4-SOX2/MYC regulatory axis plays a key role in the self-renewal and tumorigenicity of BCA stem cells [84]. In PCA, YTHDF2 and miR-493-3p are confirmed to be two key oncogenes to participate in the progression of PCA [85]. Ma et al. found that METTL3 increases m6A methylation levels of LEF1 mRNA, increasing its protein translation and enhancing the activity of the Wnt signaling pathway to promote the occurrence and development of PCA [86].
Among gynecological tumors, m6A is involved in cervical squamous cell carcinoma (CSCC), ovarian cancer (OC), and endometrial cancer (EC) [87,88,89]. In CSCC, Zhou et al., found that FTO reduces β-catenin expression by reducing the m6A levels in its mRNA transcripts, thereby increasing the excision repair cross-complementation group 1 (ERCC1) activity to enhance the resistance of CSCC to chemotherapy and radiotherapy [87]. Moreover, YTHDF1 increases the translation of EIF3C by recognizing the m6A modification sites in EIF3C mRNA, and elevated EIF3C ultimately promotes the progression of OC [88]. In EC, Liu et al., demonstrated that estrogen decreases the m6A methylation levels by downregulating the METTL3/METTL14 levels in cancer cells, which then reduces the expression of the negative AKT regulator PHLPP2, but induces the expression of the positive regulator mTORC2, thereby activating the AKT signaling pathway and enhancing cancer cell proliferation, migration, and invasion [89].
In uveal melanoma (UM), METTL3-mediated m6A methylation modification regulates the proliferation, migration and invasion of UM cells by targeting c-Met. As a protein product encoded by the proto-oncogene c-Met, c-Met plays a critical carcinogenic role in the development of UM [90]. Moreover, METTL3 can promote cancer cell proliferation and tumor growth in cutaneous squamous cell carcinoma (cSCC) by upregulating the expression of delta Np63, one of the subtypes of p63 gene, which plays an important role in the growth, differentiation and pathological development of normal epithelium [91].
The dual role of m6A in tumorigenesis and development
We have found that although m6A modification plays a broad role in the occurrence and development of various tumors, and the mechanisms are diverse and complex (Table 1 and Fig. 2). In many types of cancer cells, increased m6A levels are associated with upregulated expression of oncogenes, decrease apoptosis of cancer cells, and improved cellular migration and invasion abilities, which are conducive to the cancer progression. Moreover, the increased m6A levels could improve resistance of cancer cells to drugs and shorten life expectancy of patients. For example, Hua et al. found the upregulated METTL3 promotes the occurrence and invasion of OC by stimulating AXL transcription and inducing EMT progression [92]. However, in the research on colorectal cancer (CRC), the effects of m6A on tumors are diverse and contradictory. Li et al., found that METTL3 improves the stability of SOX2 mRNA and increases the expression of SOX2 through m6A-IGF2BP2-dependent mechanism in CRC, which enhances the stemness of CRC cells and finally promotes cancer progression [93]. Similarly, Wen et al., confirmed that METTL3 promotes CRC metastasis via miR-1246/SPRED2/MAPK signaling pathway [94]. However, Deng et al., found that METTL3 inhibits the proliferation and migration of CRC through the p38/ERK pathway [95]. The contradictory effects of m6A are also reflected in the researches on glioblastoma [39,40,41,42]. In addition, different studies have found that at different stages of liver cancer development, METTL3 and ALKBH5 promote the progression of HCC by up-regulating or down-regulating the m6A modification levels [48, 50], which may provide new insights into understanding the roles of m6A in regulating cancer.
Functional roles of m6A in development of various cancers. The m6A modification exerts diverse effects on various cancers. AML acute myeloid leukemia, GBM glioblastoma, HCC hepatocellular carcinoma, GC gastric cancer, BC breast cancer, NSCLC non-small-cell lung carcinoma, BCA bladder cancer, OC ovarian cancer, UM uveal melanoma, W-E-R writers, erasers, readers
Drugs and targeted therapies in cancer development
In the research of targeted therapeutics against m6A for tumors, the research on FTO targeted therapies is the most accomplished and representative. Since Han et al. discovered the crystal structure of FTO in 2010, the research on FTO inhibitors has gained more and more attention [96]. Up to date, various small molecules have been applied to inhibit FTO expression, such as Rhein [97], meclofenamic acid [98], R-2HG [99], entacapone [100], etc. However, due to the low sensitivity and specificity, the potential for clinical application of these FTO inhibitors are limited [101]. Huang et al. recently discovered two derivatives of meclofenamic acid, FB23 and FB23-2, which could remarkably reduce the survival rate of AML cells and show the potential to treat AML [102]. Su et al. also found two new high-efficiency FTO inhibitors CS1 and CS2, which show strong anti-leukemia effects in vitro [101]. Yang et al., revealed that knocking out the FTO gene could reduce the resistance of melanoma to anti-PD-1 immunotherapy [103]. In summary, the main effects of FTO inhibitors against tumors include reducing tumor cell drug resistance, inhibiting cancer stem cell proliferation and suppressing immune evasion, suggesting the possibility of FTO as a promising therapeutic target.
In addition to FTO, many studies on other m6A-related enzymes for targeted therapy of tumors have been performed in recent years. Wang et al., reported that knocking out METL3 or METL14 can enhance the responsiveness of colorectal cancer and melanoma cells to immunotherapy by regulating the IFN-γ-Stat1-Irf1 signaling [104], and Li et al., succeeded in using ALKBH5 specific inhibitor ALK-04 to improve the efficacy of immunotherapy for colorectal cancer and melanoma [105]. It is noted that Bedi RK et al., proposed potent and selective inhibitors of METTL3, which is conducive to developing target drugs [106].
In short, although the current targeted therapeutics against m6A for treating cancer is still in its infancy, it is promising to develop clinically effective drugs in the near future with the increasing knowledge of related molecular mechanisms of m6A modification being explored.
Conclusion and perspectives
This review summarizes the different roles of m6A modification and the underlying mechanism in cancer development, and reveals that m6A modification plays a critical role in the occurrence and development of various cancers. But our knowledge about m6A is far from complete, and current research results on m6A cannot provide accurate and universal guidance for the clinical diagnosis and therapeutic strategies of cancer yet. Little is known about whether the regulation of tumors by m6A-related proteins is associated with tumor types and tumor progression stages, whether the inhibition of m6A regulatory factors for a certain tumor leads to the occurrence of another tumor or other diseases. The challenges need to be explored and solved urgently. Under these circumstances, clearer research directions have emerged: the first is to clarify the phenotype, staging and prognosis of m6A-related cancers; the second is an anti-tumor therapy research related to m6A; the third is a new immunotherapy strategy targeting m6A [107].
References
Boccaletto P, Machnicka MA, Purta E, Piatkowski P, Baginski B, Wirecki TK, de Crecy-Lagard V, Ross R, Limbach PA, Kotter A, Helm M, Bujnicki JM (2018) MODOMICS: a database of RNA modification pathways. Nucleic Acids Res 46(D1):D303–D307. https://doi.org/10.1093/nar/gkx1030
Bodi Z, Bottley A, Archer N, May ST, Fray RG (2015) Yeast m6A methylated mRNAs are enriched on translating ribosomes during meiosis, and under rapamycin treatment. PLoS ONE 10(7):e0132090. https://doi.org/10.1371/journal.pone.0132090
Zhang SY, Zhang SW, Fan XN, Zhang T, Meng J, Huang Y (2019) FunDMDeep-m6A: identification and prioritization of functional differential m6A methylation genes. Bioinformatics 35(14):i90–i98. https://doi.org/10.1093/bioinformatics/btz316
Meyer KD, Saletore Y, Zumbo P, Elemento O, Mason CE, Jaffrey SR (2012) Comprehensive analysis of mRNA methylation reveals enrichment in 3’ UTRs and near stop codons. Cell 149(7):1635–1646. https://doi.org/10.1016/j.cell.2012.05.003
Deng X, Su R, Weng H, Huang H, Li Z, Chen J (2018) RNA N(6)-methyladenosine modification in cancers: current status and perspectives. Cell Res 28(5):507–517. https://doi.org/10.1038/s41422-018-0034-6
Schwartz S, Mumbach MR, Jovanovic M, Wang T, Maciag K, Bushkin GG, Mertins P, Ter-Ovanesyan D, Habib N, Cacchiarelli D, Sanjana NE, Freinkman E, Pacold ME, Satija R, Mikkelsen TS, Hacohen N, Zhang F, Carr SA, Lander ES, Regev A (2014) Perturbation of m6A writers reveals two distinct classes of mRNA methylation at internal and 5’ sites. Cell Rep 8(1):284–296. https://doi.org/10.1016/j.celrep.2014.05.048
Yue Y, Liu J, Cui X, Cao J, Luo G, Zhang Z, Cheng T, Gao M, Shu X, Ma H, Wang F, Wang X, Shen B, Wang Y, Feng X, He C, Liu J (2018) VIRMA mediates preferential m(6)A mRNA methylation in 3’UTR and near stop codon and associates with alternative polyadenylation. Cell Discov 4:10. https://doi.org/10.1038/s41421-018-0019-0
Patil DP, Chen CK, Pickering BF, Chow A, Jackson C, Guttman M, Jaffrey SR (2016) m(6)A RNA methylation promotes XIST-mediated transcriptional repression. Nature 537(7620):369–373. https://doi.org/10.1038/nature19342
Knuckles P, Lence T, Haussmann IU, Jacob D, Kreim N, Carl SH, Masiello I, Hares T, Villasenor R, Hess D, Andrade-Navarro MA, Biggiogera M, Helm M, Soller M, Buhler M, Roignant JY (2018) Zc3h13/Flacc is required for adenosine methylation by bridging the mRNA-binding factor Rbm15/Spenito to the m(6)A machinery component Wtap/Fl(2)d. Genes Dev 32(5–6):415–429. https://doi.org/10.1101/gad.309146.117
Bokar JA, Shambaugh ME, Polayes D, Matera AG, Rottman FM (1997) Purification and cDNA cloning of the AdoMet-binding subunit of the human mRNA (N6-adenosine)-methyltransferase. RNA 3(11):1233–1247
Ping XL, Sun BF, Wang L, Xiao W, Yang X, Wang WJ, Adhikari S, Shi Y, Lv Y, Chen YS, Zhao X, Li A, Yang Y, Dahal U, Lou XM, Liu X, Huang J, Yuan WP, Zhu XF, Cheng T, Zhao YL, Wang X, Rendtlew Danielsen JM, Liu F, Yang YG (2014) Mammalian WTAP is a regulatory subunit of the RNA N6-methyladenosine methyltransferase. Cell Res 24(2):177–189. https://doi.org/10.1038/cr.2014.3
Warda AS, Kretschmer J, Hackert P, Lenz C, Urlaub H, Hobartner C, Sloan KE, Bohnsack MT (2017) Human METTL16 is a N(6)-methyladenosine (m(6)A) methyltransferase that targets pre-mRNAs and various non-coding RNAs. EMBO Rep 18(11):2004–2014. https://doi.org/10.15252/embr.201744940
Jia G, Fu Y, Zhao X, Dai Q, Zheng G, Yang Y, Yi C, Lindahl T, Pan T, Yang YG, He C (2011) N6-methyladenosine in nuclear RNA is a major substrate of the obesity-associated FTO. Nat Chem Biol 7(12):885–887. https://doi.org/10.1038/nchembio.687
Shen F, Huang W, Huang JT, Xiong J, Yang Y, Wu K, Jia GF, Chen J, Feng YQ, Yuan BF, Liu SM (2015) Decreased N(6)-methyladenosine in peripheral blood RNA from diabetic patients is associated with FTO expression rather than ALKBH5. J Clin Endocrinol Metab 100(1):E148-154. https://doi.org/10.1210/jc.2014-1893
Church C, Moir L, McMurray F, Girard C, Banks GT, Teboul L, Wells S, Bruning JC, Nolan PM, Ashcroft FM, Cox RD (2010) Overexpression of Fto leads to increased food intake and results in obesity. Nat Genet 42(12):1086–1092. https://doi.org/10.1038/ng.713
Mathiyalagan P, Adamiak M, Mayourian J, Sassi Y, Liang Y, Agarwal N, Jha D, Zhang S, Kohlbrenner E, Chepurko E, Chen J, Trivieri MG, Singh R, Bouchareb R, Fish K, Ishikawa K, Lebeche D, Hajjar RJ, Sahoo S (2019) FTO-dependent N(6)-methyladenosine regulates cardiac function during remodeling and repair. Circulation 139(4):518–532. https://doi.org/10.1161/CIRCULATIONAHA.118.033794
Zheng G, Dahl JA, Niu Y, Fedorcsak P, Huang CM, Li CJ, Vagbo CB, Shi Y, Wang WL, Song SH, Lu Z, Bosmans RP, Dai Q, Hao YJ, Yang X, Zhao WM, Tong WM, Wang XJ, Bogdan F, Furu K, Fu Y, Jia G, Zhao X, Liu J, Krokan HE, Klungland A, Yang YG, He C (2013) ALKBH5 is a mammalian RNA demethylase that impacts RNA metabolism and mouse fertility. Mol Cell 49(1):18–29. https://doi.org/10.1016/j.molcel.2012.10.015
Wang J, Wang J, Gu Q, Ma Y, Yang Y, Zhu J, Zhang Q (2020) The biological function of m6A demethylase ALKBH5 and its role in human disease. Cancer Cell Int 20:347. https://doi.org/10.1186/s12935-020-01450-1
Ueda Y, Ooshio I, Fusamae Y, Kitae K, Kawaguchi M, Jingushi K, Hase H, Harada K, Hirata K, Tsujikawa K (2017) AlkB homolog 3-mediated tRNA demethylation promotes protein synthesis in cancer cells. Sci Rep 7:42271. https://doi.org/10.1038/srep42271
Wang X, Zhao BS, Roundtree IA, Lu Z, Han D, Ma H, Weng X, Chen K, Shi H, He C (2015) N(6)-methyladenosine modulates messenger RNA translation efficiency. Cell 161(6):1388–1399. https://doi.org/10.1016/j.cell.2015.05.014
Shi H, Wang X, Lu Z, Zhao BS, Ma H, Hsu PJ, Liu C, He C (2017) YTHDF3 facilitates translation and decay of N(6)-methyladenosine-modified RNA. Cell Res 27(3):315–328. https://doi.org/10.1038/cr.2017.15
Li A, Chen YS, Ping XL, Yang X, Xiao W, Yang Y, Sun HY, Zhu Q, Baidya P, Wang X, Bhattarai DP, Zhao YL, Sun BF, Yang YG (2017) Cytoplasmic m(6)A reader YTHDF3 promotes mRNA translation. Cell Res 27(3):444–447. https://doi.org/10.1038/cr.2017.10
Roundtree IA, Luo GZ, Zhang Z, Wang X, Zhou T, Cui Y, Sha J, Huang X, Guerrero L, Xie P, He E, Shen B, He C (2017) YTHDC1 mediates nuclear export of N(6)-methyladenosine methylated mRNAs. Elife. https://doi.org/10.7554/eLife.31311
Nayler O, Hartmann AM, Stamm S (2000) The ER repeat protein YT521-B localizes to a novel subnuclear compartment. J Cell Biol 150(5):949–962. https://doi.org/10.1083/jcb.150.5.949
Hsu PJ, Zhu Y, Ma H, Guo Y, Shi X, Liu Y, Qi M, Lu Z, Shi H, Wang J, Cheng Y, Luo G, Dai Q, Liu M, Guo X, Sha J, Shen B, He C (2017) Ythdc2 is an N(6)-methyladenosine binding protein that regulates mammalian spermatogenesis. Cell Res 27(9):1115–1127. https://doi.org/10.1038/cr.2017.99
Huang H, Weng H, Sun W, Qin X, Shi H, Wu H, Zhao BS, Mesquita A, Liu C, Yuan CL, Hu YC, Huttelmaier S, Skibbe JR, Su R, Deng X, Dong L, Sun M, Li C, Nachtergaele S, Wang Y, Hu C, Ferchen K, Greis KD, Jiang X, Wei M, Qu L, Guan JL, He C, Yang J, Chen J (2018) Recognition of RNA N(6)-methyladenosine by IGF2BP proteins enhances mRNA stability and translation. Nat Cell Biol 20(3):285–295. https://doi.org/10.1038/s41556-018-0045-z
Meyer KD, Patil DP, Zhou J, Zinoviev A, Skabkin MA, Elemento O, Pestova TV, Qian SB, Jaffrey SR (2015) 5’ UTR m(6)A promotes cap-independent translation. Cell 163(4):999–1010. https://doi.org/10.1016/j.cell.2015.10.012
Lin S, Choe J, Du P, Triboulet R, Gregory RI (2016) The m(6)A methyltransferase METTL3 promotes translation in human cancer cells. Mol Cell 62(3):335–345. https://doi.org/10.1016/j.molcel.2016.03.021
Chen J, Odenike O, Rowley JD (2010) Leukaemogenesis: more than mutant genes. Nat Rev Cancer 10(1):23–36. https://doi.org/10.1038/nrc2765
Vu LP, Pickering BF, Cheng Y, Zaccara S, Nguyen D, Minuesa G, Chou T, Chow A, Saletore Y, MacKay M, Schulman J, Famulare C, Patel M, Klimek VM, Garrett-Bakelman FE, Melnick A, Carroll M, Mason CE, Jaffrey SR, Kharas MG (2017) The N(6)-methyladenosine (m(6)A)-forming enzyme METTL3 controls myeloid differentiation of normal hematopoietic and leukemia cells. Nat Med 23(11):1369–1376. https://doi.org/10.1038/nm.4416
Weng H, Huang H, Wu H, Qin X, Zhao BS, Dong L, Shi H, Skibbe J, Shen C, Hu C, Sheng Y, Wang Y, Wunderlich M, Zhang B, Dore LC, Su R, Deng X, Ferchen K, Li C, Sun M, Lu Z, Jiang X, Marcucci G, Mulloy JC, Yang J, Qian Z, Wei M, He C, Chen J (2018) METTL14 Inhibits hematopoietic stem/progenitor differentiation and promotes leukemogenesis via mRNA m(6)A modification. Cell Stem Cell 22(2):191–205. https://doi.org/10.1016/j.stem.2017.11.016
Barbieri I, Tzelepis K, Pandolfini L, Shi J, Millan-Zambrano G, Robson SC, Aspris D, Migliori V, Bannister AJ, Han N, De Braekeleer E, Ponstingl H, Hendrick A, Vakoc CR, Vassiliou GS, Kouzarides T (2017) Promoter-bound METTL3 maintains myeloid leukaemia by m(6)A-dependent translation control. Nature 552(7683):126–131. https://doi.org/10.1038/nature24678
Bansal H, Yihua Q, Iyer SP, Ganapathy S, Proia DA, Penalva LO, Uren PJ, Suresh U, Carew JS, Karnad AB, Weitman S, Tomlinson GE, Rao MK, Kornblau SM, Bansal S (2014) WTAP is a novel oncogenic protein in acute myeloid leukemia. Leukemia 28(5):1171–1174. https://doi.org/10.1038/leu.2014.16
Li Z, Weng H, Su R, Weng X, Zuo Z, Li C, Huang H, Nachtergaele S, Dong L, Hu C, Qin X, Tang L, Wang Y, Hong GM, Huang H, Wang X, Chen P, Gurbuxani S, Arnovitz S, Li Y, Li S, Strong J, Neilly MB, Larson RA, Jiang X, Zhang P, Jin J, He C, Chen J (2017) FTO plays an oncogenic role in acute myeloid leukemia as a N(6)-methyladenosine RNA demethylase. Cancer Cell 31(1):127–141. https://doi.org/10.1016/j.ccell.2016.11.017
Shen C, Sheng Y, Zhu AC, Robinson S, Jiang X, Dong L, Chen H, Su R, Yin Z, Li W, Deng X, Chen Y, Hu YC, Weng H, Huang H, Prince E, Cogle CR, Sun M, Zhang B, Chen CW, Marcucci G, He C, Qian Z, Chen J (2020) RNA demethylase ALKBH5 selectively promotes tumorigenesis and cancer stem cell self-renewal in acute myeloid leukemia. Cell Stem Cell 27(1):64–80. https://doi.org/10.1016/j.stem.2020.04.009
Kwok CT, Marshall AD, Rasko JE, Wong JJ (2017) Genetic alterations of m(6)A regulators predict poorer survival in acute myeloid leukemia. J Hematol Oncol 10(1):39. https://doi.org/10.1186/s13045-017-0410-6
Wang J, Li Y, Wang P, Han G, Zhang T, Chang J, Yin R, Shan Y, Wen J, Xie X, Feng M, Wang Q, Hu J, Cheng Y, Zhang T, Li Y, Gao Z, Guo C, Wang J, Liang J, Cui M, Gao K, Chai J, Liu W, Cheng H, Li L, Zhou F, Liu L, Luo Y, Li S, Zhang H (2020) Leukemogenic chromatin alterations promote AML leukemia stem cells via a KDM4C-ALKBH5-AXL signaling axis. Cell Stem Cell 27(1):81–97. https://doi.org/10.1016/j.stem.2020.04.001
Shen Y, Chen X, He J, Liao D, Zu X (2018) Axl inhibitors as novel cancer therapeutic agents. Life Sci 198:99–111. https://doi.org/10.1016/j.lfs.2018.02.033
Cui Q, Shi H, Ye P, Li L, Qu Q, Sun G, Sun G, Lu Z, Huang Y, Yang CG, Riggs AD, He C, Shi Y (2017) m(6)A RNA methylation regulates the self-renewal and tumorigenesis of glioblastoma stem cells. Cell Rep 18(11):2622–2634. https://doi.org/10.1016/j.celrep.2017.02.059
Visvanathan A, Patil V, Arora A, Hegde AS, Arivazhagan A, Santosh V, Somasundaram K (2018) Essential role of METTL3-mediated m(6)A modification in glioma stem-like cells maintenance and radioresistance. Oncogene 37(4):522–533. https://doi.org/10.1038/onc.2017.351
Li F, Yi Y, Miao Y, Long W, Long T, Chen S, Cheng W, Zou C, Zheng Y, Wu X, Ding J, Zhu K, Chen D, Xu Q, Wang J, Liu Q, Zhi F, Ren J, Cao Q, Zhao W (2019) N(6)-methyladenosine modulates nonsense-mediated mRNA decay in human glioblastoma. Cancer Res 79(22):5785–5798. https://doi.org/10.1158/0008-5472.CAN-18-2868
Kowalski-Chauvel A, Lacore MG, Arnauduc F, Delmas C, Toulas C, Cohen-Jonathan-Moyal E, Seva C (2020) The m6A RNA demethylase ALKBH5 promotes radioresistance and invasion capability of glioma stem cells. Cancers (Basel). https://doi.org/10.3390/cancers13010040
Zhang S, Zhao BS, Zhou A, Lin K, Zheng S, Lu Z, Chen Y, Sulman EP, Xie K, Bogler O, Majumder S, He C, Huang S (2017) m(6)A demethylase ALKBH5 maintains tumorigenicity of glioblastoma stem-like cells by sustaining FOXM1 expression and cell proliferation program. Cancer Cell 31(4):591–606. https://doi.org/10.1016/j.ccell.2017.02.013
Mu Q, Wang L, Yu F, Gao H, Lei T, Li P, Liu P, Zheng X, Hu X, Chen Y, Jiang Z, Sayari AJ, Shen J, Huang H (2015) Imp2 regulates GBM progression by activating IGF2/PI3K/Akt pathway. Cancer Biol Ther 16(4):623–633. https://doi.org/10.1080/15384047.2015.1019185
Prasad G, Sottero T, Yang X, Mueller S, James CD, Weiss WA, Polley MY, Ozawa T, Berger MS, Aftab DT, Prados MD, Haas-Kogan DA (2011) Inhibition of PI3K/mTOR pathways in glioblastoma and implications for combination therapy with temozolomide. Neuro Oncol 13(4):384–392. https://doi.org/10.1093/neuonc/noq193
Qu M, Kong Y, Yuan YJ, Wang DY (2019) Neuronal damage induced by nanopolystyrene particles in nematode Caenorhabditis elegans. Environ Sci-Nano 6(8):2591–2601. https://doi.org/10.1039/c9en00473d
Chen M, Wei L, Law CT, Tsang FH, Shen J, Cheng CL, Tsang LH, Ho DW, Chiu DK, Lee JM, Wong CC, Ng IO, Wong CM (2018) RNA N6-methyladenosine methyltransferase-like 3 promotes liver cancer progression through YTHDF2-dependent posttranscriptional silencing of SOCS2. Hepatology 67(6):2254–2270. https://doi.org/10.1002/hep.29683
Chen M, Wong CM (2020) The emerging roles of N6-methyladenosine (m6A) deregulation in liver carcinogenesis. Mol Cancer 19(1):44. https://doi.org/10.1186/s12943-020-01172-y
Liu X, Qin J, Gao T, Li C, Chen X, Zeng K, Xu M, He B, Pan B, Xu X, Pan Y, Sun H, Xu T, Wang S (2020) Analysis of METTL3 and METTL14 in hepatocellular carcinoma. Aging (Albany NY) 12(21):21638–21659. https://doi.org/10.18632/aging.103959
Ma JZ, Yang F, Zhou CC, Liu F, Yuan JH, Wang F, Wang TT, Xu QG, Zhou WP, Sun SH (2017) METTL14 suppresses the metastatic potential of hepatocellular carcinoma by modulating N(6) -methyladenosine-dependent primary MicroRNA processing. Hepatology 65(2):529–543. https://doi.org/10.1002/hep.28885
Ebrahimi F, Gopalan V, Smith RA, Lam AK (2014) miR-126 in human cancers: clinical roles and current perspectives. Exp Mol Pathol 96(1):98–107. https://doi.org/10.1016/j.yexmp.2013.12.004
Li Z, Li F, Peng Y, Fang J, Zhou J (2020) Identification of three m6A-related mRNAs signature and risk score for the prognostication of hepatocellular carcinoma. Cancer Med 9(5):1877–1889. https://doi.org/10.1002/cam4.2833
Chen Y, Peng C, Chen J, Chen D, Yang B, He B, Hu W, Zhang Y, Liu H, Dai L, Xie H, Zhou L, Wu J, Zheng S (2019) WTAP facilitates progression of hepatocellular carcinoma via m6A-HuR-dependent epigenetic silencing of ETS1. Mol Cancer 18(1):127. https://doi.org/10.1186/s12943-019-1053-8
Lan T, Li H, Zhang D, Xu L, Liu H, Hao X, Yan X, Liao H, Chen X, Xie K, Li J, Liao M, Huang J, Yuan K, Zeng Y, Wu H (2019) KIAA1429 contributes to liver cancer progression through N6-methyladenosine-dependent post-transcriptional modification of GATA3. Mol Cancer 18(1):186. https://doi.org/10.1186/s12943-019-1106-z
Li J, Zhu L, Shi Y, Liu J, Lin L, Chen X (2019) m6A demethylase FTO promotes hepatocellular carcinoma tumorigenesis via mediating PKM2 demethylation. Am J Transl Res 11(9):6084–6092
Liu X, Liu J, Xiao W, Zeng Q, Bo H, Zhu Y, Gong L, He D, Xing X, Li R, Zhou M, Xiong W, Zhou Y, Zhou J, Li X, Guo F, Xu C, Chen X, Wang X, Wang F, Wang Q, Cao K (2020) SIRT1 Regulates N(6) -methyladenosine RNA modification in hepatocarcinogenesis by inducing RANBP2-dependent FTO SUMOylation. Hepatology 72(6):2029–2050. https://doi.org/10.1002/hep.31222
Chen Y, Zhao Y, Chen J, Peng C, Zhang Y, Tong R, Cheng Q, Yang B, Feng X, Lu Y, Xie H, Zhou L, Wu J, Zheng S (2020) ALKBH5 suppresses malignancy of hepatocellular carcinoma via m(6)A-guided epigenetic inhibition of LYPD1. Mol Cancer 19(1):123. https://doi.org/10.1186/s12943-020-01239-w
Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A (2018) Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 68(6):394–424. https://doi.org/10.3322/caac.21492
Wang Q, Chen C, Ding Q, Zhao Y, Wang Z, Chen J, Jiang Z, Zhang Y, Xu G, Zhang J, Zhou J, Sun B, Zou X, Wang S (2020) METTL3-mediated m(6)A modification of HDGF mRNA promotes gastric cancer progression and has prognostic significance. Gut 69(7):1193–1205. https://doi.org/10.1136/gutjnl-2019-319639
Zhang J, Guo S, Piao HY, Wang Y, Wu Y, Meng XY, Yang D, Zheng ZC, Zhao Y (2019) ALKBH5 promotes invasion and metastasis of gastric cancer by decreasing methylation of the lncRNA NEAT1. J Physiol Biochem 75(3):379–389. https://doi.org/10.1007/s13105-019-00690-8
Ge L, Zhang N, Chen Z, Song J, Wu Y, Li Z, Chen F, Wu J, Li D, Li J, Wang C, Wang H, Wang J (2020) Level of N6-methyladenosine in peripheral blood RNA: a novel predictive biomarker for gastric cancer. Clin Chem 66(2):342–351. https://doi.org/10.1093/clinchem/hvz004
He X, Shu Y (2019) RNA N6-methyladenosine modification participates in miR-660/E2F3 axis-mediated inhibition of cell proliferation in gastric cancer. Pathol Res Pract 215(6):152393. https://doi.org/10.1016/j.prp.2019.03.021
Zhang C, Zhang M, Ge S, Huang W, Lin X, Gao J, Gong J, Shen L (2019) Reduced m6A modification predicts malignant phenotypes and augmented Wnt/PI3K-Akt signaling in gastric cancer. Cancer Med 8(10):4766–4781. https://doi.org/10.1002/cam4.2360
DeSantis CE, Ma J, Gaudet MM, Newman LA, Miller KD, Goding Sauer A, Jemal A (2019) Siegel RL (2019) Breast cancer statistics. CA Cancer J Clin 69(6):438–451. https://doi.org/10.3322/caac.21583
Tian T, Wang M, Lin S, Guo Y, Dai Z, Liu K, Yang P, Dai C, Zhu Y, Zheng Y, Xu P, Zhu W, Dai Z (2018) The impact of lncRNA dysregulation on clinicopathology and survival of breast cancer: a systematic review and meta-analysis. Mol Ther Nucleic Acids 12:359–369. https://doi.org/10.1016/j.omtn.2018.05.018
Zhang C, Samanta D, Lu H, Bullen JW, Zhang H, Chen I, He X, Semenza GL (2016) Hypoxia induces the breast cancer stem cell phenotype by HIF-dependent and ALKBH5-mediated m(6)A-demethylation of NANOG mRNA. Proc Natl Acad Sci USA 113(14):E2047-2056. https://doi.org/10.1073/pnas.1602883113
Najafzadeh B, Asadzadeh Z, Motafakker Azad R, Mokhtarzadeh A, Baghbanzadeh A, Alemohammad H, Abdoli Shadbad M, Vasefifar P, Najafi S, Baradaran B (2021) The oncogenic potential of NANOG: an important cancer induction mediator. J Cell Physiol 236(4):2443–2458. https://doi.org/10.1002/jcp.30063
Zhang C, Zhi WI, Lu H, Samanta D, Chen I, Gabrielson E, Semenza GL (2016) Hypoxia-inducible factors regulate pluripotency factor expression by ZNF217- and ALKBH5-mediated modulation of RNA methylation in breast cancer cells. Oncotarget 7(40):64527–64542. https://doi.org/10.18632/oncotarget.11743
Niu Y, Lin Z, Wan A, Chen H, Liang H, Sun L, Wang Y, Li X, Xiong XF, Wei B, Wu X, Wan G (2019) RNA N6-methyladenosine demethylase FTO promotes breast tumor progression through inhibiting BNIP3. Mol Cancer 18(1):46. https://doi.org/10.1186/s12943-019-1004-4
Cai X, Wang X, Cao C, Gao Y, Zhang S, Yang Z, Liu Y, Zhang X, Zhang W, Ye L (2018) HBXIP-elevated methyltransferase METTL3 promotes the progression of breast cancer via inhibiting tumor suppressor let-7g. Cancer Lett 415:11–19. https://doi.org/10.1016/j.canlet.2017.11.018
Wang H, Xu B, Shi J (2020) N6-methyladenosine METTL3 promotes the breast cancer progression via targeting Bcl-2. Gene 722:144076. https://doi.org/10.1016/j.gene.2019.144076
Chen W, Zheng R, Baade PD, Zhang S, Zeng H, Bray F, Jemal A, Yu XQ (2015) He J (2016) Cancer statistics in China. CA Cancer J Clin 66(2):115–132. https://doi.org/10.3322/caac.21338
Yang D, Liu Y, Bai C, Wang X, Powell CA (2020) Epidemiology of lung cancer and lung cancer screening programs in China and the United States. Cancer Lett 468:82–87. https://doi.org/10.1016/j.canlet.2019.10.009
Wang H, Deng Q, Lv Z, Ling Y, Hou X, Chen Z, Dinglin X, Ma S, Li D, Wu Y, Peng Y, Huang H, Chen L (2019) N6-methyladenosine induced miR-143-3p promotes the brain metastasis of lung cancer via regulation of VASH1. Mol Cancer 18(1):181. https://doi.org/10.1186/s12943-019-1108-x
Wanna-Udom S, Terashima M, Lyu H, Ishimura A, Takino T, Sakari M, Tsukahara T, Suzuki T (2020) The m6A methyltransferase METTL3 contributes to Transforming Growth Factor-beta-induced epithelial-mesenchymal transition of lung cancer cells through the regulation of JUNB. Biochem Biophys Res Commun 524(1):150–155. https://doi.org/10.1016/j.bbrc.2020.01.042
Du M, Zhang Y, Mao Y, Mou J, Zhao J, Xue Q, Wang D, Huang J, Gao S, Gao Y (2017) MiR-33a suppresses proliferation of NSCLC cells via targeting METTL3 mRNA. Biochem Biophys Res Commun 482(4):582–589. https://doi.org/10.1016/j.bbrc.2016.11.077
Liu J, Ren D, Du Z, Wang H, Zhang H, Jin Y (2018) m(6)A demethylase FTO facilitates tumor progression in lung squamous cell carcinoma by regulating MZF1 expression. Biochem Biophys Res Commun 502(4):456–464. https://doi.org/10.1016/j.bbrc.2018.05.175
Cayir A, Barrow TM, Guo L, Byun HM (2019) Exposure to environmental toxicants reduces global N6-methyladenosine RNA methylation and alters expression of RNA methylation modulator genes. Environ Res 175:228–234. https://doi.org/10.1016/j.envres.2019.05.011
Li J, Han Y, Zhang H, Qian Z, Jia W, Gao Y, Zheng H, Li B (2019) The m6A demethylase FTO promotes the growth of lung cancer cells by regulating the m6A level of USP7 mRNA. Biochem Biophys Res Commun 512(3):479–485. https://doi.org/10.1016/j.bbrc.2019.03.093
Li X, Tang J, Huang W, Wang F, Li P, Qin C, Qin Z, Zou Q, Wei J, Hua L, Yang H, Wang Z (2017) The M6A methyltransferase METTL3: acting as a tumor suppressor in renal cell carcinoma. Oncotarget 8(56):96103–96116. https://doi.org/10.18632/oncotarget.21726
Green NH, Galvan DL, Badal SS, Chang BH, LeBleu VS, Long J, Jonasch E, Danesh FR (2019) MTHFD2 links RNA methylation to metabolic reprogramming in renal cell carcinoma. Oncogene 38(34):6211–6225. https://doi.org/10.1038/s41388-019-0869-4
Yang F, Jin H, Que B, Chao Y, Zhang H, Ying X, Zhou Z, Yuan Z, Su J, Wu B, Zhang W, Qi D, Chen D, Min W, Lin S, Ji W (2019) Dynamic m(6)A mRNA methylation reveals the role of METTL3-m(6)A-CDCP1 signaling axis in chemical carcinogenesis. Oncogene 38(24):4755–4772. https://doi.org/10.1038/s41388-019-0755-0
Xie H, Li J, Ying Y, Yan H, Jin K, Ma X, He L, Xu X, Liu B, Wang X, Zheng X, Xie L (2020) METTL3/YTHDF2 m(6) A axis promotes tumorigenesis by degrading SETD7 and KLF4 mRNAs in bladder cancer. J Cell Mol Med 24(7):4092–4104. https://doi.org/10.1111/jcmm.15063
Gao Q, Zheng J, Ni Z, Sun P, Yang C, Cheng M, Wu M, Zhang X, Yuan L, Zhang Y, Li Y (2020) The m(6)A methylation-regulated AFF4 promotes self-renewal of bladder cancer stem cells. Stem Cells Int 2020:8849218. https://doi.org/10.1155/2020/8849218
Li J, Meng S, Xu M, Wang S, He L, Xu X, Wang X, Xie L (2018) Downregulation of N(6)-methyladenosine binding YTHDF2 protein mediated by miR-493–3p suppresses prostate cancer by elevating N(6)-methyladenosine levels. Oncotarget 9(3):3752–3764. https://doi.org/10.18632/oncotarget.23365
Ma XX, Cao ZG, Zhao SL (2020) m6A methyltransferase METTL3 promotes the progression of prostate cancer via m6A-modified LEF1. Eur Rev Med Pharmacol Sci 24(7):3565–3571. https://doi.org/10.26355/eurrev_202004_20817
Zhou S, Bai ZL, Xia D, Zhao ZJ, Zhao R, Wang YY, Zhe H (2018) FTO regulates the chemo-radiotherapy resistance of cervical squamous cell carcinoma (CSCC) by targeting beta-catenin through mRNA demethylation. Mol Carcinog 57(5):590–597. https://doi.org/10.1002/mc.22782
Liu T, Wei Q, Jin J, Luo Q, Liu Y, Yang Y, Cheng C, Li L, Pi J, Si Y, Xiao H, Li L, Rao S, Wang F, Yu J, Yu J, Zou D, Yi P (2020) The m6A reader YTHDF1 promotes ovarian cancer progression via augmenting EIF3C translation. Nucleic Acids Res 48(7):3816–3831. https://doi.org/10.1093/nar/gkaa048
Liu J, Eckert MA, Harada BT, Liu SM, Lu Z, Yu K, Tienda SM, Chryplewicz A, Zhu AC, Yang Y, Huang JT, Chen SM, Xu ZG, Leng XH, Yu XC, Cao J, Zhang Z, Liu J, Lengyel E, He C (2018) m(6)A mRNA methylation regulates AKT activity to promote the proliferation and tumorigenicity of endometrial cancer. Nat Cell Biol 20(9):1074–1083. https://doi.org/10.1038/s41556-018-0174-4
Luo G, Xu W, Zhao Y, Jin S, Wang S, Liu Q, Chen X, Wang J, Dong F, Hu DN, Reinach PS, Yan D (2020) RNA m(6) A methylation regulates uveal melanoma cell proliferation, migration, and invasion by targeting c-Met. J Cell Physiol 235(10):7107–7119. https://doi.org/10.1002/jcp.29608
Zhou R, Gao Y, Lv D, Wang C, Wang D, Li Q (2019) METTL3 mediated m(6)A modification plays an oncogenic role in cutaneous squamous cell carcinoma by regulating DeltaNp63. Biochem Biophys Res Commun 515(2):310–317. https://doi.org/10.1016/j.bbrc.2019.05.155
Hua W, Zhao Y, Jin X, Yu D, He J, Xie D, Duan P (2018) METTL3 promotes ovarian carcinoma growth and invasion through the regulation of AXL translation and epithelial to mesenchymal transition. Gynecol Oncol 151(2):356–365. https://doi.org/10.1016/j.ygyno.2018.09.015
Li T, Hu PS, Zuo Z, Lin JF, Li X, Wu QN, Chen ZH, Zeng ZL, Wang F, Zheng J, Chen D, Li B, Kang TB, Xie D, Lin D, Ju HQ, Xu RH (2019) METTL3 facilitates tumor progression via an m(6)A-IGF2BP2-dependent mechanism in colorectal carcinoma. Mol Cancer 18(1):112. https://doi.org/10.1186/s12943-019-1038-7
Peng W, Li J, Chen R, Gu Q, Yang P, Qian W, Ji D, Wang Q, Zhang Z, Tang J, Sun Y (2019) Upregulated METTL3 promotes metastasis of colorectal Cancer via miR-1246/SPRED2/MAPK signaling pathway. J Exp Clin Cancer Res 38(1):393. https://doi.org/10.1186/s13046-019-1408-4
Deng R, Cheng Y, Ye S, Zhang J, Huang R, Li P, Liu H, Deng Q, Wu X, Lan P, Deng Y (2019) m(6)A methyltransferase METTL3 suppresses colorectal cancer proliferation and migration through p38/ERK pathways. Onco Targets Ther 12:4391–4402. https://doi.org/10.2147/OTT.S201052
Han Z, Niu T, Chang J, Lei X, Zhao M, Wang Q, Cheng W, Wang J, Feng Y, Chai J (2010) Crystal structure of the FTO protein reveals basis for its substrate specificity. Nature 464(7292):1205–1209. https://doi.org/10.1038/nature08921
Chen B, Ye F, Yu L, Jia G, Huang X, Zhang X, Peng S, Chen K, Wang M, Gong S, Zhang R, Yin J, Li H, Yang Y, Liu H, Zhang J, Zhang H, Zhang A, Jiang H, Luo C, Yang CG (2012) Development of cell-active N6-methyladenosine RNA demethylase FTO inhibitor. J Am Chem Soc 134(43):17963–17971. https://doi.org/10.1021/ja3064149
Huang Y, Yan J, Li Q, Li J, Gong S, Zhou H, Gan J, Jiang H, Jia GF, Luo C, Yang CG (2015) Meclofenamic acid selectively inhibits FTO demethylation of m6A over ALKBH5. Nucleic Acids Res 43(1):373–384. https://doi.org/10.1093/nar/gku1276
Su R, Dong L, Li C, Nachtergaele S, Wunderlich M, Qing Y, Deng X, Wang Y, Weng X, Hu C, Yu M, Skibbe J, Dai Q, Zou D, Wu T, Yu K, Weng H, Huang H, Ferchen K, Qin X, Zhang B, Qi J, Sasaki AT, Plas DR, Bradner JE, Wei M, Marcucci G, Jiang X, Mulloy JC, Jin J, He C, Chen J (2018) R-2HG Exhibits Anti-tumor Activity by Targeting FTO/m(6)A/MYC/CEBPA Signaling. Cell 172(1–2):90–105. https://doi.org/10.1016/j.cell.2017.11.031
Peng S, Xiao W, Ju D, Sun B, Hou N, Liu Q, Wang Y, Zhao H, Gao C, Zhang S, Cao R, Li P, Huang H, Ma Y, Wang Y, Lai W, Ma Z, Zhang W, Huang S, Wang H, Zhang Z, Zhao L, Cai T, Zhao YL, Wang F, Nie Y, Zhi G, Yang YG, Zhang EE, Huang N (2019) Identification of entacapone as a chemical inhibitor of FTO mediating metabolic regulation through FOXO1. Sci Transl Med. https://doi.org/10.1126/scitranslmed.aau7116
Su R, Dong L, Li Y, Gao M, Han L, Wunderlich M, Deng X, Li H, Huang Y, Gao L, Li C, Zhao Z, Robinson S, Tan B, Qing Y, Qin X, Prince E, Xie J, Qin H, Li W, Shen C, Sun J, Kulkarni P, Weng H, Huang H, Chen Z, Zhang B, Wu X, Olsen MJ, Muschen M, Marcucci G, Salgia R, Li L, Fathi AT, Li Z, Mulloy JC, Wei M, Horne D, Chen J (2020) Targeting FTO suppresses cancer stem cell maintenance and immune evasion. Cancer Cell 38(1):79–96. https://doi.org/10.1016/j.ccell.2020.04.017
Huang Y, Su R, Sheng Y, Dong L, Dong Z, Xu H, Ni T, Zhang ZS, Zhang T, Li C, Han L, Zhu Z, Lian F, Wei J, Deng Q, Wang Y, Wunderlich M, Gao Z, Pan G, Zhong D, Zhou H, Zhang N, Gan J, Jiang H, Mulloy JC, Qian Z, Chen J, Yang CG (2019) Small-molecule targeting of oncogenic FTO demethylase in acute myeloid leukemia. Cancer Cell 35(4):677–691. https://doi.org/10.1016/j.ccell.2019.03.006
Yang S, Wei J, Cui YH, Park G, Shah P, Deng Y, Aplin AE, Lu Z, Hwang S, He C, He YY (2019) m(6)A mRNA demethylase FTO regulates melanoma tumorigenicity and response to anti-PD-1 blockade. Nat Commun 10(1):2782. https://doi.org/10.1038/s41467-019-10669-0
Wang L, Hui H, Agrawal K, Kang Y, Li N, Tang R, Yuan J, Rana TM (2020) m(6) A RNA methyltransferases METTL3/14 regulate immune responses to anti-PD-1 therapy. EMBO J 39(20):e104514. https://doi.org/10.15252/embj.2020104514
Li N, Kang Y, Wang L, Huff S, Tang R, Hui H, Agrawal K, Gonzalez GM, Wang Y, Patel SP, Rana TM (2020) ALKBH5 regulates anti-PD-1 therapy response by modulating lactate and suppressive immune cell accumulation in tumor microenvironment. Proc Natl Acad Sci USA 117(33):20159–20170. https://doi.org/10.1073/pnas.1918986117
Bedi RK, Huang D, Eberle SA, Wiedmer L, Sledz P, Caflisch A (2020) Small-Molecule Inhibitors of METTL3, the Major Human Epitranscriptomic Writer. ChemMedChem 15(9):744–748. https://doi.org/10.1002/cmdc.202000011
Yang G, Sun Z, Zhang N (2020) Reshaping the role of m6A modification in cancer transcriptome: a review. Cancer Cell Int 20:353. https://doi.org/10.1186/s12935-020-01445-y
Acknowledgements
This work was supported by Student Innovation Training Program of Nanjing University (202110284146Z), National Natural Science Foundation of China (31901182, 31870492), and the Natural Science Foundation of Jiangsu Province of China (BK20190316).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing financial interests.
Consent for publication
This article has been read and approved in the present form for submission by all authors.
Ethical approval
No any human participants or animal studies were included in this study.
Informed consent
Informed Consent was not applicable to the study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Chen, P., Hu, J., Han, X. et al. Advances in the functional roles of N6-methyladenosine modification in cancer progression: mechanisms and clinical implications. Mol Biol Rep 49, 4929–4941 (2022). https://doi.org/10.1007/s11033-022-07126-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-022-07126-5