Abstract
Background
Plant breeding allows altering the genetic structure of plants to meet human needs. The use of radiation technology for inducing mutations and -thereby- new phenotypic variants has become increasingly common as a tool for developing new crops. The aim of this study was to determine the effective gamma irradiation dose for inducing mutations in purple carrot.
Methods and results
Increasing gamma radiation doses [0, 50, 100, 200, 300, 400, 500, and 600 Gy] were applied to purple carrot seeds. The irradiated seeds were sown in pots and the emergence and survival rates of the seedlings were analyzed. Considering plant emergence (%) as a response variable, the LD50 dose was 387.5 Gy. Analysis of root length, root width (shoulder diameter) and plant height in control (0 Gy) and irradiated plants (50–600 Gy) revealed an inverse association between these morphological traits and radiation dose. SRAP and ISSR markers were used to identify DNA polymorphisms in irradiated and control plants. The range of amplicons per primer set revealed by ISSR and SRAP markers was 4–10 and 2–13, respectively. In the ISSR analysis of the irradiated carrots (for the 8 doses used), we obtained range values for the average Nei’s gene diversity, Shannon’s information index, and polymorphism information content (PIC) of 0.13–0.25, 0.20–0.35, and 1.39–1.67, respectively, whereas in the SRAP analysis, the range values for these parameters were 0.15–0.25, 0.23–0.37, and 0.43–0.58, respectively. Cluster analysis revealed three main groups; (a) non-irradiated (control) plants, (b) plants from the 600 Gy dose, and (c) a third group with two subgroups: one with individuals from the lowest irradiation doses (50–200 Gy) and a second group with individuals from the highest irradiation doses (300–500 Gy).
Conclusions
This is the first report on determining effective mutagen doses and genetic characterization of induced mutagenesis via gamma irradiation in purple carrot. ISSR and SRAP markers were successful in detecting variations among different levels of mutagen doses.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cultivated carrots (Daucus carota subsp. sativus) (2n = 2x = 18) are the most important members of the Apiaceae family in terms of their economic and nutritional content. Cultivated carrots are sub-classified into Eastern and Western carrots, based on their root color and other morphological and physiological features [1]. Most Eastern carrots have branched anthocyanin-pigmented purple roots, although some have yellow roots. Eastern carrots also have more pubescent and slightly dissected leaves, and a tendency for early flowering. On the other hand, Western carrots typically have unbranched carotenoid-pigmented orange, yellow, red or white roots, with less pubescent and highly dissected leaves, and a lower tendency for early flowering [2].
The Western orange carrots, which are nowadays grown worldwide, represent a rich source of carotenoids, vitamins and dietary fiber, and may contain high concentrations of the provitamin A carotenoid β-carotene [3]. In addition, due to increasing health awareness, purple carrots (Daucus carota ssp. sativus var. atrorubens Alef.) are becoming popular because of their abundant phytonutrients and the antioxidant and anti-inflammatory health-promoting effects associated with anthocyanin consumption [4].
Purple carrots accumulate mainly cyanidin derivatives, and they are particularly rich in acylated anthocyanins, with acylated forms accounting for 49.6 to 99% of the total anthocyanin content are used as food dyes because of their chemical stability (reviewed by Cavagnaro and Iorizzo, [4]). Because acylated anthocyanins are chemically more stable than their non-acylated counterparts [5], the formers are suitable natural food dyes. Thus, in addition to its health-enhancing attributes, purple carrots also represent a valuable source of anthocyanin-based dyes for the food industry.
The term ‘mutation breeding’ was first used by Freisleben and Lein in [6] to describe the application of conscious induction and development of mutant lines for product development in plants. Plant breeders are still using classical breeding methods for developing new varieties with superior traits. However, the selection and breeding from existing population reduces the number of gene variants. To avoid this negative outcome, breeders may opt to use alternative breeding strategies. Mutation breeding has become an alternative method for breeders because it provides the chance to obtain new desirable traits that were not previously found in nature or they were lost throughout evolution [7]. Mutation breeding techniques are extensively applied in agriculture in order to produce genetic diversity [8]. While new genetic combinations from already existing parental genes are produced by conventional breeding, mutation breeding procedures cause new gene combinations with high mutation frequency [8]. Artificially induced mutations can be generated by means of applying chemical, physical, or biological mutagens to plant organs [7, 9]. The most widely used mutagen in mutation breeding is gamma rays, mainly due its relatively straight forward utilization, as compared to other methods. Gamma ray-induced mutations (GRIM) have been used in many species, including coriander [10], Withania somnifera [11], tomato [12], Anthurium [13], mungbean [14, 15], eggplant [16], rice [17], potato [18], and carrot [19].
GRIM produce genetic variations, and this is generally associated with an increase in DNA polymorphisms [20]. DNA markers have been used for many applications in genetic research, including the assessment of genetic stability, characterization of genetic diversity, and linkage mapping, among others. In addition, DNA markers have been used for screening and identification of mutant plants in Jatropha curcas [20], soybean [21], maize [22], potato [18, 23], Curcuma alismatifolia Gagnep. [24], faba bean [25], Acorus calamus L. [26], wheat [27], Sophpra davii [28], and Helichrysum bracteatum [29].
To date, only two studies have applied GRIM in carrot plants [30, 31] but neither of these studies examined purple-rooted carrots, nor were the irradiated plants characterized by molecular markers. The objectives of this study were to investigate the effect of different doses of gamma radiation on seed germination, seedlings survival, and morphological traits of purple carrot plants, and to investigate DNA polymorphisms in the irradiated and control (non-treated) plants using ‘inter simple sequence repeat’ (ISSR) and sequence-related amplified polymorphism (SRAP) markers.
Materials and methods
Irradiation, and analysis of plant survival and morphological traits
Seed lots of the purple carrot line (local seeds from Hatay) were irradiated with 0 (control plants), 50, 100, 200, 300, 400, 500, or 600 Gy gamma radiation, using a 60Co gamma cell at the Atomic Energy Institution, Ankara, Turkey. In order to determine survival rates, control and treated seeds were sown in 11.5-L pots containing peat and perlite in a 1:1 ratio. For each irradiation treatment, 6 replicates of 10 seeds per replicate were used. The seedlings emergence was monitored daily for 8 days after the emergence of the first seedling. Data on seedling survival rate, expressed as percentage of the total seeds sown, and irradiation doses, were used for linear regression analysis to determine the median lethal dose (LD50). Also, Pearson correlation analysis was performed between radiation dose and survival rate of the carrot seedlings.
Seeds from each irradiation treatment were sown in the experimental field of the Alata Horticultural Research Institute (Mersin, Turkey), using three replicates (20 plants per replicate) of plants per replicate according to the completely randomized design. The resulting carrot plants were grown –using conventional agricultural practices for the crop- for five months, after which the following morphological traits were analyzed: root length, root shoulder diameter and plant height (from soil level to the tip of the longest leaf). The data were analyzed using one-way ANOVA with SAS software package. Comparison of means was carried out using Duncan’s multiple range test.
Molecular analysis
DNA isolation
Fresh young leaves of control and irradiated plants (12 individuals from each dose group) were collected and frozen at − 80 °C until lyophilized and used for DNA isolation. Total genomic DNA was isolated from lyophilized and ground leaf tissue of individual plants following the CTAB protocol of Doyle and Doyle [32] with minor modifications as incorporated by Boiteux et al. [33]. Purity and concentration of the DNA was determined with a NanoDrop spectrophotometer (DeNovix DS-11 FX, USA), and aliquots of the DNA samples were diluted to a final concentration of 25 ng µL−1 for further use in polymerase chain reaction (PCR) assays and stored at − 20 °C.
SRAP analysis
For PCR amplification, nine primer combinations derived from five forward and eight reverse primers were used (Table 1). The reaction mixture and protocols were as described by Ferriol et al. [34] and Yildiz et al. [35]. Briefly, PCR reactions were performed in 25 μl containing 25 ng of genomic DNA, 1.5 mM MgCl2, 0.5 μM of primer, 0.2 mM dNTPs, 10 × Taq buffer, and 1 unit of Taq polymerase (Fermentas, Thermo Fisher Scientific Inc, US). The amplification protocol consisted of an initial denaturing step of 5 min at 94 °C, followed by five cycles of three steps of 1 min of denaturing at 94 °C, 1 min of annealing at 35 °C and 2 min of elongation at 72 °C. In the following 30 cycles the annealing temperature was increased to 50 °C, with a final extension step of 5 min at 72 °C. The amplicons were size-separated by 3% (w v−1) agarose gel electrophoresis in 0.5 × TAE buffer for 5 h and visualized with ethidium bromide. A 100 bp ladder (Fermentas) was used as molecular weight marker.
ISSR analysis
Five ISSR primers were used (Table 1). The amplification reactions were performed as previously reported by Yildiz et al. [36]. The reaction mixture (20 μl) contained 20 ng of genomic DNA, 1.5 mM MgCl2, 0.2 mM of primer, 0.2 mM dNTP, 10 × Taq buffer, 1 unit of Taq DNA polymerase (Fermentas, Thermo Fisher Scientific Inc, US). The ISSR PCR reactions were performed as follows: a first denaturing cycle of 3 min at 94 °C, followed by 45 cycles of 30 s of denaturing at 94 °C, 45 s of annealing temperature at 55 °C, and 2 min of elongation at 72 °C, and a final extension cycle of 5 min at 72 °C. PCR products were separated by 3% (w v−1) agarose gel electrophoresis in 0.5 × TAE buffer, and visualized with ethidium bromide. A 100 bp ladder (Fermentas, Thermo Fisher Scientific Inc, US) was used as molecular weight marker.
Data scoring and statistical analysis
For SRAP and ISSR marker data, only intense and legible polymorphic bands were scored as present (1) or absent (0) in all of the control and irradiated plants. For both marker systems, the total number of amplified products, the number of polymorphic bands, polymorphism percentage, polymorphism information content (PIC), and genetic diversity parameters were calculated independently for each gamma ray dose group. The PIC values for dominant markers were calculated according to De Riek et al. [37]. A dendrogram based on Jaccard similarity coefficient was constructed using UPGMA (Unweighted Pair Group Method with Arithmetic Mean). Pop Gene v.1.32 software [38] was used to estimate the polymorphic percentage, observed number of alleles (Na), effective number of alleles (Ne), Nei’s [39] gene diversity (h), Shannon’s information index (I).
Results
Significant effects (p < 0.05) of radiation on plant emergence, plant survival and morphological traits were found. Treated seeds were evaluated for survivability from different doses of gamma irradiations. Emergence of the seedlings began within 8 to 9 days of sowing in the control and all the irradiation treatments. Based on the seedlings emergence rate (Table 2), an LD50 of 387.8 Gy was obtained (Fig. 1). As shown in the regression equation presented in the graph, for each unit of increase in gamma radiation intensity (1 Gy) the plant survival rate decreased 0.113 times. This reduction was found to be statistically significant (p < 0.05). A statistically significant Pearson correlation coefficient (r) value of –0.922 (–92.2%) between radiation dose and survival rate was found.
Generally, the seedling survival rates decreased with increasing gamma irradiation doses (Table 2). The non-irradiated (control) treatment had the highest seedling survival rate (98.3%), followed by 100 Gy (85%), 50 Gy (81.6%), and 200 Gy (81.6%). Seedling survival decreased to 33.3% at the 600 Gy dose, to 38.3% at the 500 and 300 Gy, and 43.3% at 400 Gy (Table 2). The survival rates of seedlings obtained from seeds irradiated with 300–600 Gy were significantly lower than the survival rates in the control and the lowest irradiation (50–200 Gy) treatments (Table 2). No statistical differences were found between the control group and the lowest irradiation treatments.
Root length (r = − 0.35), root shoulder diameter (r = − 0.28), and plant height (r = − 0.40) in control and irradiated plants were all negatively affected by gamma radiation, especially at the highest doses applied (Table 2). Root length varied from 15.9 cm (in the 500 Gy dose group) to 22.4 cm (100 Gy), whereas shoulder diameter ranged from 4.5 (400 Gy) to 5.1 cm (50 Gy), and plant height from 45.8 (600 Gy) to 62.3 cm (100 Gy).
SRAP and ISSR-based marker analysis were performed to evaluate variation at the DNA level in irradiated and control plants (Figure S1, S2). A total of 100 bands were obtained using four ISSR primers and nine SRAP primer combinations, of which 84 (84%) were polymorphic (data not shown).
The four ISSR primers yielded a total of 35 bands, of which 28 were polymorphic between at least two of the 96 individuals irradiated with eight different doses of gamma rays. In general, mean percent polymorphism decreased with increasing irradiation doses (r = −0.88) and varied from 72.5% (at 100 Gy) to 52.1% (at 600 Gy) (Table 3). PIC values followed the same general trend (r = − 0.85), and ranged from 0.51, in the non-irradiated individuals, to 0.38, in the highest irradiation dose (600 Gy). These data indicate that less DNA polymorphism was found at these four ISSR loci at higher irradiation doses.
Analysis of individual markers revealed that primer ISSR3 yielded the highest number of bands (10 bands) in individuals irradiated with 50 and 100 Gy, while the lowest number of bands (4 bands) was found at 600 Gy using primer Sola11. Among the treatments, the average polymorphic bands ranged from 3.5 (600 Gy dose group) to 6 (100 Gy dose group). The percentage of polymorphism was ranged from 0 to 100%. ISSR4 (at 0 Gy, 50 Gy, 100 Gy and 300 Gy doses) showed 100% polymorphism while Sola 11 (at 600 Gy dose) produced only monomorphic bands. Percentage of polymorphic loci for 0, 100, 200, 300, 400, 500 and 600 Gy individual plants were 69.6%, 69.9%, 72.5%, 69.2%, 62.9%, 63.2%, 63.7%, and 52.1%, respectively (Table 3).
SRAP analysis with nine primer combinations yielded a total of 65 bands of which 56 were polymorphic across 96 carrot individuals exposed to eight irradiation doses (data not shown). SRAP analysis also indicated that irradation modified the level of DNA polymorphism. Similar to ISSR analysis, increasing irradiation doses reduced mean percent polymorphism (r = − 0.81), mean PIC values (r = − 0.84) and mean na values (r = − 0.72) (Table 4).
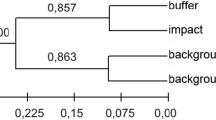
The genetic relationships among the non-irradiated and irradiated purple carrot individuals were depicted by cluster analysis using Jaccard’s genetic similarity coefficients based on marker data from the ISSR and SRAP analyses combined (Fig. 2). The dendrogram revealed three distinct clusters; (a) the non-irradiated (control) group, (b) a group with the highest irradiation dose (600 Gy), and (c) a third group with two subgroups; one with individuals from the lowest irradiation doses (50–200 Gy) and a second group with individuals from the highest irradiation doses (300–500 Gy). The dendrogram showed individuals from the highest irradiation dose (600 Gy) closer to the non-irradiated plants, and these two groups separated from the rest of the irradiated treatments.
Discussion
In mutation breeding, determining the optimal dose of a mutagen is an important prerequisite for achieving high mutation rates with fewer plant deaths, a desired outcome when trying to maximize the probabilities of rapidly attaining new phenotypic variants [39]. In the present study, survival rates decreased with increasing gamma ray doses. Reason of this decrease in the survival rate was reported by Dhakshanamoorthy et al. [20] as possibility influencing meristematic tissues of seeds by mutagen. Also, it can be considered that high level of mutagen doses cause damage in the cell, including chromosomal alteration. As comparison with our results, the LD50 for GRIM in other species was found to be 345–423 Gy [41] and 288.4–354.8 Gy [42] in rice, 985–1363 Gy in desert shrub [40], 20 Gy in palm [43], 140 Gy in pepper [44], and 149–620 Gy in cowpea [45]. The results of the study will be useful for establishing the appropriate gamma radiation dose for mutation breeding in carrot.
Low dose of mutations can stimulate [20] or reduce [10] the germination of irradiated seeds comparing to unirradiated seeds. But generally increasing mutation doses has a reducing effect. In the present study, the germination rates of irradiated with lowest doses (50, 100, 200 Gy) and control group were not different significantly. However high irradiation doses caused negative effect of germination rates. These results are in accordance with those obtained by Al-Safadi and Simon [30], who reported a strong negative correlation (r = -0.98) between dose and survival of carrot plants after seed irradiation.
Reduction of the plant height may be due to the fact that ionizing radiation generally affects cell division, thereby inhibiting or delaying the plant’s growth [46]. However, in the present study, the low-level doses of gamma irradiation (50–100 Gy) produced plants with statistically larger values for all three of these morphometric traits, suggesting that such low level of irradiation promoted -rather than inhibited- plant growth. In agreement with these results, Al-Safadi and Simon [30] reported that carrot seeds irradiated with low doses of gamma rays produced plants with 20% and 35% higher plant size and root weight than control plants, respectively. Also, another study by Bovi et al. [31] reported that carrot seeds of the orange-rooted cultivar ‘Brasilia’ exposed to 2.5 and 10.0 Gy of gamma rays produced root weight 6.0% and 1.6% more than the non-irradiated controls, respectively.
Conventional methods to determine mutant lines are influenced by external factors and less reproducible. However, molecular techniques are more safety and reproducible to confirm mutant lines [28]. Hence in the study, in addition to the morphometric analyses, ISSR and SRAP markers were used to detect polymorphism between different levels of irradiation doses due to mutation treatment. ISSR markers were favored to determine mutants before for many species; Glycine max L. [21], Curcuma alismatifolia [24], Vicia faba [25], Triticum aestivum L. [27], Sophora davidii [28], Helichrysum bracteatum L. [29] and Lilium longiflorum [47]. Polymorphism values obtained from ISSR analyses were varied between different levels of irradiation doses. These results suggest that irradiation modified the level of DNA polymorphism and ISSR markers provide relatively high discriminative power among individuals exposed to different levels of irradiation, in agreement with results from previous studies in other species [24, 28, 29, 47].
In both ISSR and SRAP analyzes, the lowest total number of bands per primer was obtained at 600 Gy dose. This suggests that the treatments of high gamma irradiation would result in DNA damage in carrot plant cells. The loss of normal bands (disappearance of bands) maybe related to events such as DNA damage (e.g., single, and double-strand breaks, modified bases), DNA–protein cross links, point mutation, and/or complex chromosomal rearrangements induced by gamma radiation [19]. In this study, the highest mean number of bands was obtained at 100 Gy and 400 Gy doses. New PCR amplification products may be generated in some oligonucleotide priming sites as a result of mutations (new annealing events), large deletions (approximation of the pre-existing annealing site) and/or homologous recombination [48]. The disappearance of bands or addition of extra bands may result in polymorphic DNA patterns in the irradiated plants [20].
The dendrogram demonstrating the relationships among the non-irradiated and irradiated purple carrot individuals grouped depending on dose level of mutation separated groups as non-irradiated, 600 Gy dose and other doses. The genetic similarity coefficient of individuals in different level of doses ranged from 0.5361 to 0.6385 calculating from both ISSR and SRAP marker data. These values of genetic similarity coefficient suggest that genetic discrimination among different dose levels in our study is stronger than those reported by Wang [28] 0.6885–0.7884 and El-Khateeb [29] 0.77–0.97. According to these results, it is confirmed that gamma ray irradiation is an effective tool to create variations in purple carrot and ISSR and SRAP markers are utilizable to detect these variations.
Conclusions
In mutation breeding studies, determining the optimal dose of a mutagen is important to develop lines with the desired agronomic traits. For this reason, we were applied gamma radiation doses range of 50- 600 Gy to purple carrot seeds. Considering plant emergence (%) as a response variable, the LD50 dose was found 387.5 Gy. This study revealed that exposure of purple carrot seeds at 100 Gy is best for root length, root width and plant height among the doses studied. This is the first report on the use of SRAP and ISSR markers characterization of induced mutagenesis through gamma irradiation in purple carrots. Our results indicate that ISSR and SRAP markers were successful in the detecting DNA polymorphism among the non-irradiated and irradiated purple carrot individuals.
References
Vavilov NI (1951) The origin, variation, immunity and breeding of cultivated plants. Chronica Botonica 13:1–366
Rubatzky VE, Quiros CF, Simon PW (1999) Carrots and related vegetable umbelliferae, crop production science in horticulture. CABI Publishing, New York, pp 294
Simon PW, Pollak L, Clevidence B, Holden J, Haytowitz D (2009) Plant breeding for human nutrition. Plant Breed Rev 31:325–392
Cavagnaro PF, Iorizzo M (2019) Carrot anthocyanin diversity, genetics, and genomics. Compendium of Plant Genomes In: Simon P, Iorizzo M, Grzebelus D, Baranski R (eds) The Carrot Genome, Springer Nature Cham, pp 261–279
Mazza G, Cacace JE, Kay CD (2004) Methods of analysis for anthocyanins in plants and biological fluids. J AOAC Int 87:129–145
Freisleben RA, Lein A (1944) Möglichkeiten und praktische Durchführung der Mutationszüchtung. Kühn-Arhiv 60:211–222
Ulukapi K, Nasircilar AG (2015) Developments of gamma ray application on mutation breeding studies in recent years. In: International Conference on Advances in Agricultural, Biological & Environmental Sciences 22–23, London, pp 31–34
Piri I, Babayan M, Tavassoli A, Javaheri M (2011) The use of gamma irradiation in agriculture. Afr J Microbiol Res 5(32):5806–5811
Beyaz R, Yildiz M (2017) The use of gamma irradiation in plant mutation breeding. In: Jurić S (ed) Plant Engineering Intechopen, pp 33–46
Salve KM, More AD (2014) Effect of gamma radiation on seed germination, seedling height and seedling injury in Coriandrum sativum Linn. Int J of Life Sciences 2(3):223–225
Bhosale RS, More AD (2014) Effect of gamma radiation on seed germination, seedling height and seedling injury in Withania somnifera (L) Dunal. Int J of Life Sciences 2(3):226–228
Sikder S, Biswas P, Hazra P, Akhtar S, Chattopadhyay A, Badigannavar AM, Souza SFD (2013) Induction of mutation in tomato (Solanum lycopersicum L.) by gamma irradiation and EMS. Indian J. Genet. 73(4):392–399
Puchooa D (2005) In vitro mutation breeding of Anthurium by gamma radiation. Int J Agri Biol 7(1):11–20
Sangsiri C, Worawit S, Peerasak S (2005) Gamma radiation induced mutation in mungbean. ScienceAsia 31:251–255
Kumar S (2014) Gamma radiation induced mutations in mungbean (Vigna radiata (L.) Wilze). Scholarly J Agric Sci 4(4):240–243
Aruna J, Parakash M, Kumar BS (2010) Studies on effect of physical and chemical mutagens on seedling charecters in Brijal (Solanum melongena L.). Int J Curr Res 3:38–41
Saleem MY, Mukhtar Z, Cheema AA et al (2005) Induced mutation and in vitro techniques as a method to induce salt tolerance in Basmati rice (Oryza sauva L.). Int J Environ Sci Tech 2(2):141–145
Yaycılı O, Alikamanoğlu S (2012) Induction of Salt-Tolerant Potato (Solanum tuberosum L.) mutants with gamma irradiation and characterization of genetic variations via RAPD-PCR analysis. Turk J Biol 36:405–412
Buyukdinc DT, Kantoglu KY, Karatas A, Ipek A, Ellialtıoglu SS (2019) Determination of effective mutagen dose for carrot (Daucus carota ssp sativus var atrorubens alef and D carota) callus culture. Int J Sci Technol Res 5(3):15–23
Dhakshanamoorthy D, Selvaraj R, Chidambaram ALA (2011) Induced mutagenesis in Jatropha curcas L using gamma rays and detection of DNA polymorphism through RAPD marker. C R Biol 334(1):24–30
Mudibu J, Nkongolo KKC, Mehes-Smith M, Kalonji-Mbuyi A (2011) Genetic analysis of a soybean genetic pool using ISSR marker: effect of gamma radiation on genetic variability. Int J Plant Breed Genet 5:235–245
Rustikawati R, Suprijono E, Romeida A, Herison C, Sutjahjo SH (2012) Identification of M4 gamma irradiated maize mutant based on RAPD markers. AGRIVITA 34(2):161–165
Afrasiab H, Iqbal J (2012) Genetic analysis of somaclonal variants and induced mutants of Potato (Solanum tuberosum L.) CV. Diamant using RAPD markers. Pak J Bot 44:215–220
Taheri S, Abdullah TL, Abdullah NAP, Ahmad Z (2013) Use of inter simple sequence repeat assay for detection of DNA polymorphism induced by gamma rays in Curcuma alismatifolia. HortScience 48:1346–1351
Mejri S, Mabrouk Y, Voisinetal M (2014) Variation in quantitative characters of faba bean after seed irradiation and associated molecular changes. Afr J Biotechnol 11(34):8383–8390
Lee JH, Han TH (2014) Selection of mutants obtained by gamma ray irradiation and analysis of genetic variation using RAPD markers in Acorus calamus L. Hort Environ Biotechnol 55(3):207–212
Yaycılı O, Sen A, Alikmanoğlu S (2005) Induced of salt tolerance wheat (Triticum aestivum L) mutants with gamma radiation and determining molecular analysis by ISSR. Procedia Environ Sci 29:196
Wang P, Zhang Y, Zhao L, Mo B (2017) Luo T (2017) Effect of gamma rays on Sophora davidii and detection of DNA polymorphism through ISSR markers. Biomed Res Int 34:1–6
El-Khateeb MA, Eid RA, Mahfouze HA, Ashor HA, Mabrouk RMS (2017) Induction of mutation with gamma radiation in Helichrysum bracteatum L., and identification of mutants by molecular markers. Middle East J. Agric. Res. 6(2):282–293
Al-Safadi B, Simon PW (1996) Gamma irradiation-induced variation in carrots (Daucus carota L.). J Am soc Hortic Sci 121(4):599–603
Bovi JE, Valter A, Neto JT (2003) Use of low doses of cobalt-60 gamma radiation on carrot seeds and their effect on plant growth and yield. Acta Hortic 607:41–43
Doyle JJ, Doyle JL (1990) Isolation of plant DNA from fresh tissue. Focus 12:1315
Boiteux LS, Fonseca MEN, Simon PW (1999) Effects of plant tissue and DNA purification methods on randomly amplified polymorphic DNA based genetic fingerprinting analysis in carrot. HortScience 124:32–38
Ferriol M, Pico B, Nuez F (2003) Genetic diversity of germplasm collection of Cucurbita pepo using SRAP and AFLP markers. Theor Appl Genet 107(2):271–282
Yıldız M, Ekbiç E, Düzyaman E, Serçe S, Abak K (2016) Genetic and phenotypic variation of Turkish Okra (Abelmoschus esculentus L. Moench) accessions and their possible relationship with American, Indian and African germplasms. J Plant Biochem Biotechnol 25(3):234–244
Yıldız M, Ekbiç E, Keles D, Şensoy S, Abak K (2011) Use of ISSR, SRAP and RAPD markers to assess genetic diversity in Turkish melons. Sci Hortic 130:349–353
De Riek J, Calsyn E, Everaert I, Van Bockstaele E, De Loose M (2001) AFLP based alternatives for the assessment of distinctness, uniformity and stability of sugar beet varieties. Theor Appl Genet 103:1254–1265
Yeh FC, Yang R, Boyle TJ, Ye Z (2000) Pop Gene 32, Microsoft windows-based freeware for population genetic analysis. Version 1.32. Edmonton Canada University of Alberta
Nei M (1973) Analysis of gene diversity in subdivided populations. Proc Natl Acad Sci 70:3321–3323
Li QH, Wang SX, Zhao YM, Xu J, Gao TT, Ren WJ (2012) Irradiation dose and effect on germination and growth of desert shrub Nitraria tangutorum Bobr. with two gamma irradiation modes. Pak J Bot 44(2):661–666
Harding SS, Johnson SD, Taylor DR, Dixon CA, Turay MY (2012) Effect of gamma rays on seed germination, seedling height, survival percentage and tiller production in some rice varieties cultivated in Sierra Leone. Am J Exp Agric 2:247–255
Ramchander S, Ushakumari R, Pillai MA (2015) Lethal dose fixation and sensitivity of rice varieties to gamma radiation. Indian J Agric Res 49:24–31
Naval MM, Zuriaga E, Badenes ML (2013) AFLP analysis of mutations induced by gamma irradiation in ‘Rojo Brillante’ persimmon. ISHS Acta Horticulturae 996. In: V International Symposium on Persimmon, 20 Oct, Wuhan, pp 117–121
Jo YD, Kim SH, Hwang JE, Kim YS, Kang HS, Ki SW, Kwon SJ, Ryu J, Kim JB, Kang SY (2016) Construction of mutation populations by gamma-ray and carbon beam irradiation in chili pepper (Capsicum annuum L.). Hortic Environ Biotechnol 57(6):606–614
Olasupo FO, Olori CO, Forster BP, Bado S (2016) Mutagenic effects of gamma radiation on eight accessions of cowpea (Vigna unguiculata [L.] Walp). Am J Plant Sci 7:339–351
Alvarez-Holguin A, Morales-Nieto CR, Avendano-Arrazate CH, Corrales-Lerma R, Villarreal-Guerrero F, Santellano-Estrada E, Gomez-Simuta Y (2019) Mean lethal dose (LD50) and growth reduction (GR50) due to gamma radiation in Wilman lovegrass (Eragrostis superba). Rev Mex Cienc Pecu 10(1):227–238
Xi M, Sun L, Qiu S, Liu J, Xu J, Shi J (2012) In vitro mutagenesis and identification of mutants via ISSR in lily (Lilium longiflorum). Plant Cell Rep 31(6):1043–1051
Atienzar FA, Conradi M, Evenden AJ, Jha AN, Depledge MH (1999) Qualitative assessment of genotoxicity using random amplified polymorphic DNA: comparison of genomic template stability with key fitness parameter in Daphnia magna exposed to benzo (a) pyrene. Environ Toxicol Chem 18:2275–2282
Funding
Authors are grateful to Van Yuzuncu Yil University- Scientific Research Projects Coordinating Office for financial support (Project number: FYL-2017–6148).
Author information
Authors and Affiliations
Contributions
MY conceptualized and established the methodology, supervised the research study. MY and GY conceived the project. MY, GY, and MK conceived and designed the experiments. MY, GY, and MK performed material preparation, germination analysis and molecular analysis. MY and ND performed field experiments. MY, MK, and PC wrote the paper and organized the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
All authors declare that they have no conflict of interest.
Consent to Participate and Publish
All authors read the manuscript and showed their willingness to publish this study.
Data availability
All data needed to conduct this study is provided within the manuscript.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Yarar, G., Kocak, M., Denli, N. et al. Determination of the effective radiation dose for mutation breeding in purple carrot (Daucus carota L.) and possible variations formed. Mol Biol Rep 49, 5219–5228 (2022). https://doi.org/10.1007/s11033-021-06618-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-021-06618-0